hexsha
stringlengths 40
40
| size
int64 6
14.9M
| ext
stringclasses 1
value | lang
stringclasses 1
value | max_stars_repo_path
stringlengths 6
260
| max_stars_repo_name
stringlengths 6
119
| max_stars_repo_head_hexsha
stringlengths 40
41
| max_stars_repo_licenses
list | max_stars_count
int64 1
191k
⌀ | max_stars_repo_stars_event_min_datetime
stringlengths 24
24
⌀ | max_stars_repo_stars_event_max_datetime
stringlengths 24
24
⌀ | max_issues_repo_path
stringlengths 6
260
| max_issues_repo_name
stringlengths 6
119
| max_issues_repo_head_hexsha
stringlengths 40
41
| max_issues_repo_licenses
list | max_issues_count
int64 1
67k
⌀ | max_issues_repo_issues_event_min_datetime
stringlengths 24
24
⌀ | max_issues_repo_issues_event_max_datetime
stringlengths 24
24
⌀ | max_forks_repo_path
stringlengths 6
260
| max_forks_repo_name
stringlengths 6
119
| max_forks_repo_head_hexsha
stringlengths 40
41
| max_forks_repo_licenses
list | max_forks_count
int64 1
105k
⌀ | max_forks_repo_forks_event_min_datetime
stringlengths 24
24
⌀ | max_forks_repo_forks_event_max_datetime
stringlengths 24
24
⌀ | avg_line_length
float64 2
1.04M
| max_line_length
int64 2
11.2M
| alphanum_fraction
float64 0
1
| cells
list | cell_types
list | cell_type_groups
list |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
4ab456b4d29ba15d1a3bb0c82bab92ed4ea8340f
| 43,958 |
ipynb
|
Jupyter Notebook
|
2019-earth/2019-earth-model2.ipynb
|
hu-minghao/my_program
|
5cde81b9a9cbe0d433fbcfc016896195de23425b
|
[
"MIT"
] | null | null | null |
2019-earth/2019-earth-model2.ipynb
|
hu-minghao/my_program
|
5cde81b9a9cbe0d433fbcfc016896195de23425b
|
[
"MIT"
] | null | null | null |
2019-earth/2019-earth-model2.ipynb
|
hu-minghao/my_program
|
5cde81b9a9cbe0d433fbcfc016896195de23425b
|
[
"MIT"
] | 2 |
2020-07-07T13:14:07.000Z
|
2021-06-29T01:23:06.000Z
| 144.12459 | 17,992 | 0.873083 |
[
[
[
"def get_Earth_temp(c20,T,cloud_re):\n\n totall_cloudAndparticle_reflectfactor=cloud_re\n ocean_t=T\n totall_h20=0.0025\n #co2单位ppm\n totall_co2=c20\n totall_co2=totall_co2*ma.pow(10,-6)\n totall_radiation=1.7*10**17\n #云及颗粒物对大气的吸收及反射影响比例系数均为0.2,短波反射,长波吸收\n \n Longwave_absorptionCoeff_h20=0.8\n def get_co2_coff(n):\n return ma.log(1.2*n+0.005*ma.pow(n,2)+1.6*ma.pow(10,-6)*ma.pow(n,3)) \n Longwave_absorptionCoeff_c02=(get_co2_coff(c20)-get_co2_coff(291.4*(ma.pow(10,-6))))*3.3\n Shortwave_absorptionCoeff=0.15\n Longwave_absorptionCoeff_air=0\n land_noIce=103\n land_Ice=46\n ocean_noIce=335\n ocean_Ice=26\n h20_climate_changefactor=0.08\n c20_climate_changefactor=0.1\n Earth_R=6378.137*(10**3)\n Earth_S=ma.pi*Earth_R**2\n Earth_C=Earth_S/0.67\n M_Atmosphere=6000*10**15\n #进入到地面的辐射\n\n Earth_surface_radiation=totall_radiation*(1-Shortwave_absorptionCoeff-totall_cloudAndparticle_reflectfactor)\n #陆地及海洋受到辐射总和\n land_radiation=0.3*Earth_surface_radiation\n ocean_radiation=0.7*Earth_surface_radiation\n #陆地及海洋反射辐射,白天\n land_noIce_factor=land_noIce/(land_noIce+land_Ice)\n ocean_noIce_factor=ocean_noIce/(ocean_noIce+ocean_Ice)\n radiation_landReflect=land_radiation*(land_noIce_factor*0.18+(1-land_noIce_factor)*0.85)\n radiation_oceanReflect=ocean_radiation*(ocean_noIce_factor*0.07+(1-ocean_noIce_factor)*0.7)\n radiation_Reflect_daytime=radiation_landReflect+radiation_oceanReflect\n #白天辐射散热及大气吸收,短波吸收\n radiation_out_AtmosphereAbsorb_daytime=(totall_radiation+radiation_Reflect_daytime)*Shortwave_absorptionCoeff\n # radiation_out_totall_daytime=radiation_totall_daytime*(1-0.15)\n\n #夜晚:陆地及海洋辐射经过大气被大气吸收的辐射,考虑云层吸收\n land_absorb_radiation=land_radiation-radiation_landReflect\n ocean_absorb_radiation=ocean_radiation-radiation_oceanReflect\n AtmosphereAbsorbFactor_night=(totall_h20*Longwave_absorptionCoeff_h20*h20_climate_changefactor+totall_co2*Longwave_absorptionCoeff_c02*c20_climate_changefactor+(1-totall_h20-totall_co2)*Longwave_absorptionCoeff_air)\n #陆地吸收系数由大气成分确定,海洋则将吸收热量辐射传热给大气\n\n def get_ocean_Ra(t,x): \n #参数为海洋平均温度t,海洋吸收总辐射x\n #获得海洋发射辐射量\n factor=0.13*ma.log(1+1.5*pow(t-13.1/13.1,2))+0.32\n q_cloud=x*factor\n return q_cloud\n\n radiation_out_AtmosphereAbsorb_night=get_ocean_Ra(ocean_t,ocean_absorb_radiation)+land_absorb_radiation\n # ocean_absorb_keep=radiation_out_AtmosphereAbsorb_night*0.7*0.5\n Atmosphere_absorb_keep=(radiation_out_AtmosphereAbsorb_daytime+radiation_out_AtmosphereAbsorb_night)*AtmosphereAbsorbFactor_night\n #夜晚,地球散热\n radiation_landdirct_out_space=(land_absorb_radiation+ocean_absorb_radiation)*(1-AtmosphereAbsorbFactor_night)\n radiation_out_space=radiation_out_AtmosphereAbsorb_night*0.5\n #总散热\n # radiation_out_totall=radiation_landdirct_out_space+radiation_out_space+radiation_out_totall_daytime+radiation_in_AtmosphereReflect\n temp=Atmosphere_absorb_keep/Earth_C\n def get_cloud_reflect(t,x):\n #参数为海洋平均温度t,海洋吸收总辐射x\n #获得云总反射系数\n w=(2501.0-2382.5)/50\n qi=(2501-t*w)\n m0=44.8*ma.pow(10,12)\n u=2*0.2*0.51*x/(qi*m0)\n return u\n a=get_cloud_reflect(ocean_t,ocean_absorb_radiation)\n # print('co2的气候系数为:%f'%(ma.pow(10,-6)*Longwave_absorptionCoeff_c02*c20_climate_changefactor))\n # print('大气吸收辐射系数:%f'%AtmosphereAbsorbFactor_night)\n return temp,a,totall_cloudAndparticle_reflectfactor\n\n",
"_____no_output_____"
],
[
"year_team=[]\nyear_T=[]\ns=0\nco2=data['co2']\nT=data['sst']\nfor n,m in zip(co2,T):\n\n temps,cloud_fa,col=get_Earth_temp(n,m,0.15)\n year_team.append(temps)\n year_T.append(cloud_fa)\n \nplt.figure(figsize=(8,8))\nplt.plot(year_team)\n ",
"_____no_output_____"
],
[
"import numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport math as ma",
"_____no_output_____"
],
[
"data=pd.read_excel(r'E:\\研究生数学建模\\prediction_data.xlsx')",
"_____no_output_____"
],
[
"data.head()",
"_____no_output_____"
],
[
"prediction=pd.DataFrame({'Team':year_team})\nprediction=pd.concat([prediction,data['year']],axis=1)",
"_____no_output_____"
],
[
"prediction=prediction.set_index('year')\nprediction.to_excel('E:\\研究生数学建模\\prediction_result.xlsx')",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab468cfe58660e9d69aefd6e90ae2d29b90f9f1
| 32,256 |
ipynb
|
Jupyter Notebook
|
sm-protein-classification.ipynb
|
cxomni/amazon-sagemaker-protein-classification
|
0fff3c51b591c9356cd7905c8d091ae9f878ece2
|
[
"MIT-0"
] | 30 |
2021-01-05T18:35:49.000Z
|
2022-03-09T07:51:27.000Z
|
sm-protein-classification.ipynb
|
mani-aiml/amazon-sagemaker-protein-classification
|
479d9bbbe11bb6280479f1bc2cb9ef48c5e32150
|
[
"MIT-0"
] | 1 |
2021-01-05T21:14:04.000Z
|
2021-01-05T23:14:11.000Z
|
sm-protein-classification.ipynb
|
mani-aiml/amazon-sagemaker-protein-classification
|
479d9bbbe11bb6280479f1bc2cb9ef48c5e32150
|
[
"MIT-0"
] | 10 |
2021-01-05T18:35:55.000Z
|
2022-01-28T12:20:09.000Z
| 38.815884 | 1,051 | 0.633154 |
[
[
[
"# Fine-tuning and deploying ProtBert Model for Protein Classification using Amazon SageMaker\n",
"_____no_output_____"
],
[
"## Contents\n1. [Motivation](#Motivation)\n2. [What is ProtBert?](#What-is-ProtBert?)\n3. [Notebook Overview](#Notebook-Overview)\n - [Setup](#Setup)\n4. [Dataset](#Dataset)\n - [Download Data](#Download-Data)\n5. [Data Exploration](#Data-Exploration)\n - [Upload Data to S3](#Upload-Data-to-S3)\n6. [Training script](#Training-script)\n7. [Train on Amazon SageMaker](#Train-on-Amazon-SageMaker)\n8. [Deploy the Model on Amazon SageMaker](#Deploy-the-model-on-Amazon-SageMaker)\n - [Create a model object](#Create-a-model-object)\n - [Deploy the model on an endpoint](#Deploy-the-model-on-an-endpoint)\n9. [Predicting SubCellular Localization of Protein Sequences](#Predicting-SubCellular-Localization-of-Protein-Sequences)\n10. [References](#References)\n---",
"_____no_output_____"
],
[
"## Motivation\n<img src=\"https://upload.wikimedia.org/wikipedia/commons/6/60/Myoglobin.png\"\n alt=\"Protein Sequence\"\n style=\"float: left;\"\n height = 100\n width = 250/>\n**Proteins** are the key fundamental macromolecules governing in biological bodies. The study of protein localization is important to comprehend the function of protein and has great importance for drug design and other applications. It also plays an important role in characterizing the cellular function of hypothetical and newly discovered proteins [1]. There are several research endeavours that aim to localize whole proteomes by using high-throughput approaches [2–4]. These large datasets provide important information about protein function, and more generally global cellular processes. However, they currently do not achieve 100% coverage of proteomes, and the methodology used can in some cases cause mislocalization of subsets of proteins [5,6]. Therefore, complementary methods are necessary to address these problems. In this notebook, we will leverage Natural Language Processing (NLP) techniques for protein sequence classification. The idea is to interpret protein sequences as sentences and their constituent – amino acids –\nas single words [7]. More specifically we will fine tune Pytorch ProtBert model from Hugging Face library.",
"_____no_output_____"
],
[
"## What is ProtBert?\n\nProtBert is a pretrained model on protein sequences using a masked language modeling (MLM) objective. It is based on Bert model which is pretrained on a large corpus of protein sequences in a self-supervised fashion. This means it was pretrained on the raw protein sequences only, with no humans labelling them in any way (which is why it can use lots of publicly available data) with an automatic process to generate inputs and labels from those protein sequences [8]. For more information about ProtBert, see [`ProtTrans: Towards Cracking the Language of Life’s Code Through Self-Supervised Deep Learning and High Performance Computing`](https://www.biorxiv.org/content/10.1101/2020.07.12.199554v2.full).\n\n---\n",
"_____no_output_____"
],
[
"## Notebook Overview\n\nThis example notebook focuses on fine-tuning the Pytorch ProtBert model and deploying it using Amazon SageMaker, which is the most comprehensive and fully managed machine learning service. With SageMaker, data scientists and developers can quickly and easily build and train machine learning models, and then directly deploy them into a production-ready hosted environment. \nDuring the training, we will leverage SageMaker distributed data parallel (SDP) feature which extends SageMaker’s training capabilities on deep learning models with near-linear scaling efficiency, achieving fast time-to-train with minimal code changes.\n\n_**Note**_: Please select the Kernel as ` Python 3 (Pytorch 1.6 Python 3.6 CPU Optimized)`.\n\n---",
"_____no_output_____"
],
[
"### Setup\n\nTo start, we import some Python libraries and initialize a SageMaker session, S3 bucket and prefix, and IAM role.\n",
"_____no_output_____"
]
],
[
[
"!pip install --upgrade pip -q\n!pip install -U boto3 sagemaker -q\n!pip install seaborn -q",
"_____no_output_____"
]
],
[
[
"Next let us import the common libraries needed for the operations done later.",
"_____no_output_____"
]
],
[
[
"import re\nimport json\nimport pandas as pd\nfrom tqdm import tqdm\nimport seaborn as sns\nimport matplotlib.pyplot as plt\nimport numpy as np\nimport time\nimport os\nimport numpy as np\nimport pandas as pd\nimport sagemaker\nimport torch\nimport seaborn as sns\nimport matplotlib.pyplot as plt\n\nfrom torch import nn, optim\nfrom torch.utils.data import Dataset, DataLoader",
"_____no_output_____"
]
],
[
[
"Next, let's verify the version, create a SageMaker session and get the execution role which is the IAM role arn used to give training and hosting access to your data.",
"_____no_output_____"
]
],
[
[
"import sagemaker\nprint(sagemaker.__version__)\nsagemaker_session = sagemaker.Session()\nrole = sagemaker.get_execution_role()",
"_____no_output_____"
]
],
[
[
"Now we will specify the S3 bucket and prefix where you will store your training data and model artifacts. This should be within the same region as the Notebook Instance, training, and hosting.",
"_____no_output_____"
]
],
[
[
"bucket = sagemaker_session.default_bucket()\nprefix = \"sagemaker/DEMO-pytorch-bert\"\n",
"_____no_output_____"
]
],
[
[
"As the last step of setting up the enviroment lets set a value to a random seed so that we can reproduce the same results later.",
"_____no_output_____"
]
],
[
[
"RANDOM_SEED = 43\nnp.random.seed(RANDOM_SEED)\ntorch.manual_seed(RANDOM_SEED)",
"_____no_output_____"
]
],
[
[
"---",
"_____no_output_____"
],
[
"## Dataset",
"_____no_output_____"
],
[
"We are going to use a opensource public dataset of protein sequences available [here](http://www.cbs.dtu.dk/services/DeepLoc-1.0/data.php). The dataset is a `fasta file` composed by header and protein sequence. The header is composed by the accession number from Uniprot, the annotated subcellular localization and possibly a description field indicating if the protein was part of the test set. The subcellular localization includes an additional label, where S indicates soluble, M membrane and U unknown[9].\nSample of the data is as follows :",
"_____no_output_____"
],
[
"```\n>Q9SMX3 Mitochondrion-M test\nMVKGPGLYTEIGKKARDLLYRDYQGDQKFSVTTYSSTGVAITTTGTNKGSLFLGDVATQVKNNNFTADVKVST\nDSSLLTTLTFDEPAPGLKVIVQAKLPDHKSGKAEVQYFHDYAGISTSVGFTATPIVNFSGVVGTNGLSLGTDV\nAYNTESGNFKHFNAGFNFTKDDLTASLILNDKGEKLNASYYQIVSPSTVVGAEISHNFTTKENAITVGTQHAL>\nDPLTTVKARVNNAGVANALIQHEWRPKSFFTVSGEVDSKAIDKSAKVGIALALKP\"\n```",
"_____no_output_____"
],
[
"A sequence in FASTA format begins with a single-line description, followed by lines of sequence data. The definition line (defline) is distinguished from the sequence data by a greater-than (>) symbol at the beginning. The word following the \">\" symbol is the identifier of the sequence, and the rest of the line is the description.",
"_____no_output_____"
],
[
"### Download Data",
"_____no_output_____"
]
],
[
[
"!wget http://www.cbs.dtu.dk/services/DeepLoc-1.0/deeploc_data.fasta -P ./data -q",
"_____no_output_____"
]
],
[
[
"Since the data is in fasta format, we can leverage `Bio.SeqIO.FastaIO` library to read the dataset. Let us install the Bio package.",
"_____no_output_____"
]
],
[
[
"!pip install Bio -q\nimport Bio",
"_____no_output_____"
]
],
[
[
"Using the Bio package we will read the data directly by filtering out the columns that are of interest. We will also add a space seperater between each character in the sequence field which will be useful during model training.",
"_____no_output_____"
]
],
[
[
"def read_fasta(file_path, columns) :\n from Bio.SeqIO.FastaIO import SimpleFastaParser\n with open('./data/deeploc_data.fasta') as fasta_file: # Will close handle cleanly\n records = []\n for title, sequence in SimpleFastaParser(fasta_file):\n record = []\n title_splits = title.split(None)\n record.append(title_splits[0]) # First word is ID\n sequence = \" \".join(sequence)\n record.append(sequence)\n record.append(len(sequence))\n location_splits = title_splits[1].split(\"-\")\n record.append(location_splits[0]) # Second word is Location\n record.append(location_splits[1]) # Second word is Membrane\n\n if(len(title_splits) > 2):\n record.append(0)\n else:\n record.append(1)\n \n records.append(record)\n return pd.DataFrame(records, columns = columns)\n \ndata = read_fasta(\"./tmp/deeploc_data.fasta\", columns=[\"id\", \"sequence\", \"sequence_length\", \"location\", \"membrane\", \"is_train\"])\ndata.head()",
"_____no_output_____"
]
],
[
[
"### Data Exploration\nDataset consists of 14K sequences and 6 columns in total. We will only use the following columns during training:\n\n* _**id**_ : Unique identifier given each sequence in the dataset.\n* _**sequence**_ : Protein sequence. Each character is seperated by a \"space\". Will be useful for BERT tokernizer.\n* _**sequence_length**_ : Character length of each protein sequence. \n* _**location**_ : Classification given each sequence.\n* _**is_train**_ : Indicates whether the record be used for training or test. Will be used to seperate the dataset for traning and validation.\n",
"_____no_output_____"
],
[
"First, let's verify if there are any missing values in the dataset.",
"_____no_output_____"
]
],
[
[
"data.info()",
"_____no_output_____"
],
[
"data.isnull().values.any()",
"_____no_output_____"
]
],
[
[
"As you can see, there are **no** missing values in this dataset. ",
"_____no_output_____"
],
[
"Second, we will see the number of available classes (subcellular localization), which will be used for protein classification. ",
"_____no_output_____"
]
],
[
[
"unique_classes = data.location.unique()\nprint(\"Number of classes: \", len(unique_classes))\nunique_classes",
"_____no_output_____"
]
],
[
[
"We can see that there are 10 unique classes in the dataset.",
"_____no_output_____"
],
[
"Third, lets check the sequence length.",
"_____no_output_____"
]
],
[
[
"%matplotlib inline\n%config InlineBackend.figure_format='retina'\nsns.set(style='whitegrid', palette='muted', font_scale=1.2)\nax = sns.distplot(data['sequence_length'].values)\nax.set_xlim(0, 3000)\nplt.title(f'sequence length distribution')\nplt.grid(True)",
"_____no_output_____"
]
],
[
[
"This is an important observation as PROTBERT model receives a fixed length of sentence as input. Usually the maximum length of a sentence depends on the data we are working on. For sentences that are shorter than this maximum length, we will have to add paddings (empty tokens) to the sentences to make up the length.\n\nAs you can see from the above plot that most of the sequences lie under the length of around 1500, therefore, its a good idea to select the `max_length = 1536` but that will increase the training time for this sample notebook, therefore, we will use `max_length = 512`. You can experiment it with the bigger length and it does improves the accuracy as most of the subcellular localization information of protiens is stored at the end of the sequence. ",
"_____no_output_____"
],
[
"Next let's factorize the protein classes. ",
"_____no_output_____"
]
],
[
[
"categories = data.location.astype('category').cat\ndata['location'] = categories.codes\nclass_names = categories.categories\nnum_classes = len(class_names)\nprint(class_names)",
"_____no_output_____"
]
],
[
[
"Next, let's devide the dataset into training and test. We can leverage the `is_train` column to do the split. ",
"_____no_output_____"
]
],
[
[
"df_train = data[data.is_train == 1]\ndf_train = df_train.drop([\"is_train\"], axis = 1)\ndf_train.shape[0]",
"_____no_output_____"
],
[
"df_test = data[data.is_train == 0]\ndf_test = df_test.drop([\"is_train\"], axis = 1)\ndf_test.shape[0]",
"_____no_output_____"
]
],
[
[
"We got **11231** records as training set and **2773** records as the test set which is about 75:25 data split between the train and test. Also, the composition between multiple classes remains uniform between both datasets. ",
"_____no_output_____"
],
[
"### Upload Data to S3\nIn order to accomodate model training on SageMaker we need to upload the data to s3 location. We are going to use the `sagemaker.Session.upload_data` function to upload our datasets to an S3 location. The return value inputs identifies the location -- we will use later when we start the training job.",
"_____no_output_____"
]
],
[
[
"train_dataset_path = './data/deeploc_per_protein_train.csv'\ntest_dataset_path = './data/deeploc_per_protein_test.csv'\ndf_train.to_csv(train_dataset_path)\ndf_test.to_csv(test_dataset_path)\ninputs_train = sagemaker_session.upload_data(train_dataset_path, bucket=bucket, key_prefix=prefix)\ninputs_test = sagemaker_session.upload_data(test_dataset_path, bucket=bucket, key_prefix=prefix)",
"_____no_output_____"
],
[
"print(\"S3 location for training data: \", inputs_train )\nprint(\"S3 location for testing data: \", inputs_test )",
"_____no_output_____"
]
],
[
[
"## Training script\nWe use the [PyTorch-Transformers library](https://pytorch.org/hub/huggingface_pytorch-transformers), which contains PyTorch implementations and pre-trained model weights for many NLP models, including BERT. As mentioned above, we will use `ProtBert model` which is pre-trained on protein sequences. \n\nOur training script should save model artifacts learned during training to a file path called `model_dir`, as stipulated by the SageMaker PyTorch image. Upon completion of training, model artifacts saved in `model_dir` will be uploaded to S3 by SageMaker and will be used for deployment.\n\nWe save this script in a file named `train.py`, and put the file in a directory named `code/`. The full training script can be viewed under `code/`.\n\nIt also has the code required for distributed data parallel (DDP) training using SMDataParallel. It is very similar to a PyTorch training script you might run outside of SageMaker, but modified to run with SMDataParallel, which is a new capability in Amazon SageMaker to train deep learning models faster and cheaper. SMDataParallel's PyTorch client provides an alternative to PyTorch's native DDP. For details about how to use SMDataParallel's DDP in your native PyTorch script, see the [Getting Started with SMDataParallel tutorials](https://docs.aws.amazon.com/sagemaker/latest/dg/distributed-training.html#distributed-training-get-started).",
"_____no_output_____"
]
],
[
[
"!pygmentize code/train.py",
"_____no_output_____"
]
],
[
[
"### Train on Amazon SageMaker\nWe use Amazon SageMaker to train and deploy a model using our custom PyTorch code. The Amazon SageMaker Python SDK makes it easier to run a PyTorch script in Amazon SageMaker using its PyTorch estimator. After that, we can use the SageMaker Python SDK to deploy the trained model and run predictions. For more information on how to use this SDK with PyTorch, see [the SageMaker Python SDK documentation](https://sagemaker.readthedocs.io/en/stable/using_pytorch.html).\n\nTo start, we use the `PyTorch` estimator class to train our model. When creating our estimator, we make sure to specify a few things:\n\n* `entry_point`: the name of our PyTorch script. It contains our training script, which loads data from the input channels, configures training with hyperparameters, trains a model, and saves a model. It also contains code to load and run the model during inference.\n* `source_dir`: the location of our training scripts and requirements.txt file. \"requirements.txt\" lists packages you want to use with your script.\n* `framework_version`: the PyTorch version we want to use.\n\nThe PyTorch estimator supports both single-machine & multi-machine, distributed PyTorch training using SMDataParallel. _Our training script supports distributed training for only GPU instances_. \n\n#### Instance types\n\nSMDataParallel supports model training on SageMaker with the following instance types only:\n\n- ml.p3.16xlarge\n- ml.p3dn.24xlarge [Recommended]\n- ml.p4d.24xlarge [Recommended]\n\n#### Instance count\n\nTo get the best performance and the most out of SMDataParallel, you should use at least 2 instances, but you can also use 1 for testing this example.\n\n#### Distribution strategy\n\nNote that to use DDP mode, you update the the distribution strategy, and set it to use smdistributed dataparallel.\n\nAfter creating the estimator, we then call fit(), which launches a training job. We use the Amazon S3 URIs where we uploaded the training data earlier.",
"_____no_output_____"
]
],
[
[
"# Training job will take around 20-25 mins to execute. \n\nfrom sagemaker.pytorch import PyTorch\n\n\nTRAINING_JOB_NAME=\"protbert-training-pytorch-{}\".format(time.strftime(\"%m-%d-%Y-%H-%M-%S\")) \nprint('Training job name: ', TRAINING_JOB_NAME)\n\nestimator = PyTorch(\n entry_point=\"train.py\",\n source_dir=\"code\",\n role=role,\n framework_version=\"1.6.0\",\n py_version=\"py36\",\n instance_count=1, # this script support distributed training for only GPU instances.\n instance_type=\"ml.p3.16xlarge\",\n distribution={'smdistributed':{\n 'dataparallel':{\n 'enabled': True\n }\n }\n },\n debugger_hook_config=False,\n hyperparameters={\n \"epochs\": 3,\n \"num_labels\": num_classes,\n \"batch-size\": 4,\n \"test-batch-size\": 4,\n \"log-interval\": 100,\n \"frozen_layers\": 15,\n },\n metric_definitions=[\n {'Name': 'train:loss', 'Regex': 'Training Loss: ([0-9\\\\.]+)'},\n {'Name': 'test:accuracy', 'Regex': 'Validation Accuracy: ([0-9\\\\.]+)'},\n {'Name': 'test:loss', 'Regex': 'Validation loss: ([0-9\\\\.]+)'},\n ]\n)\nestimator.fit({\"training\": inputs_train, \"testing\": inputs_test}, job_name=TRAINING_JOB_NAME)",
"_____no_output_____"
]
],
[
[
"With `max_length=512` and running the model for only 3 epochs we get the validation accuracy of around 65%, which is pretty decent. You can optimize it further by trying bigger sequence length, increasing the number of epochs and tuning other hyperparamters. For details you can refer to the research paper: \n[`ProtTrans: Towards Cracking the Language of Life’s Code Through Self-Supervised Deep Learning and High Performance Computing`](https://arxiv.org/pdf/2007.06225.pdf).",
"_____no_output_____"
],
[
"Before, we deploy the model to an endpoint, let's first store the model to S3.",
"_____no_output_____"
]
],
[
[
"model_data = estimator.model_data\nprint(\"Storing {} as model_data\".format(model_data))\n%store model_data",
"_____no_output_____"
],
[
"%store -r model_data\n\n# If no model was found, set it manually here.\n# model_data = 's3://sagemaker-{region}-XXX/protbert-training-pytorch-XX-XX-XXXX-XX-XX-XX/output/model.tar.gz'\n\nprint(\"Using this model: {}\".format(model_data))",
"_____no_output_____"
]
],
[
[
"## Deploy the model on Amazon SageMaker",
"_____no_output_____"
],
[
"After training our model, we host it on an Amazon SageMaker Endpoint. To make the endpoint load the model and serve predictions, we implement a few methods in inference.py.\n\n- `model_fn()`: function defined to load the saved model and return a model object that can be used for model serving. The SageMaker PyTorch model server loads our model by invoking model_fn.\n- `input_fn()`: deserializes and prepares the prediction input. In this example, our request body is first serialized to JSON and then sent to model serving endpoint. Therefore, in input_fn(), we first deserialize the JSON-formatted request body and return the input as a torch.tensor, as required for BERT.\n- `predict_fn()`: performs the prediction and returns the result.\nTo deploy our endpoint, we call deploy() on our PyTorch estimator object, passing in our desired number of instances and instance type:",
"_____no_output_____"
],
[
"### Create a model object\nYou define the model object by using SageMaker SDK's PyTorchModel and pass in the model from the estimator and the entry_point. The function loads the model and sets it to use a GPU, if available.",
"_____no_output_____"
]
],
[
[
"import sagemaker\n\nfrom sagemaker.pytorch import PyTorchModel\nENDPOINT_NAME = \"protbert-inference-pytorch-1-{}\".format(time.strftime(\"%m-%d-%Y-%H-%M-%S\"))\nprint(\"Endpoint name: \", ENDPOINT_NAME)\nmodel = PyTorchModel(model_data=model_data, source_dir='code',\n entry_point='inference.py', role=role, framework_version='1.6.0', py_version='py3')",
"_____no_output_____"
]
],
[
[
"### Deploy the model on an endpoint\nYou create a predictor by using the model.deploy function. You can optionally change both the instance count and instance type.",
"_____no_output_____"
]
],
[
[
"%%time\npredictor = model.deploy(initial_instance_count=1, instance_type='ml.m5.2xlarge', endpoint_name=ENDPOINT_NAME)",
"_____no_output_____"
]
],
[
[
"## Predicting SubCellular Localization of Protein Sequences",
"_____no_output_____"
]
],
[
[
"import boto3\n\nruntime= boto3.client('runtime.sagemaker')\nclient = boto3.client('sagemaker')\n\nendpoint_desc = client.describe_endpoint(EndpointName=ENDPOINT_NAME)\nprint(endpoint_desc)\nprint('---'*30)",
"_____no_output_____"
]
],
[
[
"We then configure the predictor to use application/json for the content type when sending requests to our endpoint:",
"_____no_output_____"
]
],
[
[
"predictor.serializer = sagemaker.serializers.JSONSerializer()\npredictor.deserializer = sagemaker.deserializers.JSONDeserializer()",
"_____no_output_____"
]
],
[
[
"Finally, we use the returned predictor object to call the endpoint:",
"_____no_output_____"
]
],
[
[
"protein_sequence = 'M G K K D A S T T R T P V D Q Y R K Q I G R Q D Y K K N K P V L K A T R L K A E A K K A A I G I K E V I L V T I A I L V L L F A F Y A F F F L N L T K T D I Y E D S N N'\nprediction = predictor.predict(protein_sequence)\nprint(prediction)\nprint(f'Protein Sequence: {protein_sequence}')\nprint(\"Sequence Localization Ground Truth is: {} - prediction is: {}\".format('Endoplasmic.reticulum', class_names[prediction[0]]))",
"_____no_output_____"
],
[
"protein_sequence = 'M S M T I L P L E L I D K C I G S N L W V I M K S E R E F A G T L V G F D D Y V N I V L K D V T E Y D T V T G V T E K H S E M L L N G N G M C M L I P G G K P E'\nprediction = predictor.predict(protein_sequence)\nprint(prediction)\nprint(f'Protein Sequence: {protein_sequence}')\nprint(\"Sequence Localization Ground Truth is: {} - prediction is: {}\".format('Nucleus', class_names[prediction[0]]))",
"_____no_output_____"
],
[
"seq = 'M G G P T R R H Q E E G S A E C L G G P S T R A A P G P G L R D F H F T T A G P S K A D R L G D A A Q I H R E R M R P V Q C G D G S G E R V F L Q S P G S I G T L Y I R L D L N S Q R S T C C C L L N A G T K G M C'\nprediction = predictor.predict(seq)\nprint(prediction)\nprint(f'Protein Sequence: {seq}')\nprint(\"Sequence Localization Ground Truth is: {} - prediction is: {}\".format('Cytoplasm',class_names[prediction[0]]))",
"_____no_output_____"
]
],
[
[
"# Cleanup\n\nLastly, please remember to delete the Amazon SageMaker endpoint to avoid charges:",
"_____no_output_____"
]
],
[
[
"predictor.delete_endpoint()",
"_____no_output_____"
]
],
[
[
"## References\n- [1] Refining Protein Subcellular Localization (https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1289393/)\n- [2] Kumar A, Agarwal S, Heyman JA, Matson S, Heidtman M, et al. Subcellular localization of the yeast proteome. Genes Dev. 2002;16:707–719. [PMC free article] [PubMed] [Google Scholar]\n- [3] Huh WK, Falvo JV, Gerke LC, Carroll AS, Howson RW, et al. Global analysis of protein localization in budding yeast. Nature. 2003;425:686–691. [PubMed] [Google Scholar]\n- [4] Wiemann S, Arlt D, Huber W, Wellenreuther R, Schleeger S, et al. From ORFeome to biology: A functional genomics pipeline. Genome Res. 2004;14:2136–2144. [PMC free article] [PubMed] [Google Scholar]\n- [5] Davis TN. Protein localization in proteomics. Curr Opin Chem Biol. 2004;8:49–53. [PubMed] [Google Scholar]\n- [6] Scott MS, Thomas DY, Hallett MT. Predicting subcellular localization via protein motif co-occurrence. Genome Res. 2004;14:1957–1966. [PMC free article] [PubMed] [Google Scholar]\n- [7] ProtTrans: Towards Cracking the Language of Life's Code Through Self-Supervised Deep Learning and High Performance Computing (https://www.biorxiv.org/content/10.1101/2020.07.12.199554v2.full.pdf)\n- [8] ProtBert Hugging Face (https://huggingface.co/Rostlab/prot_bert)\n- [9] DeepLoc-1.0: Eukaryotic protein subcellular localization predictor (http://www.cbs.dtu.dk/services/DeepLoc-1.0/data.php)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
4ab46b8f79344daeb35f6496d163685c966fdb96
| 57,508 |
ipynb
|
Jupyter Notebook
|
docs/notebooks/autodiff_cookbook.ipynb
|
lucyfan7266/jax
|
36fea96185a3c222e1909be2704875625b422f27
|
[
"ECL-2.0",
"Apache-2.0"
] | 1 |
2021-03-06T16:44:45.000Z
|
2021-03-06T16:44:45.000Z
|
docs/notebooks/autodiff_cookbook.ipynb
|
lucyfan7266/jax
|
36fea96185a3c222e1909be2704875625b422f27
|
[
"ECL-2.0",
"Apache-2.0"
] | null | null | null |
docs/notebooks/autodiff_cookbook.ipynb
|
lucyfan7266/jax
|
36fea96185a3c222e1909be2704875625b422f27
|
[
"ECL-2.0",
"Apache-2.0"
] | null | null | null | 34.230952 | 800 | 0.57792 |
[
[
[
"# The Autodiff Cookbook\n\n[](https://colab.sandbox.google.com/github/google/jax/blob/master/docs/notebooks/autodiff_cookbook.ipynb)\n\n*alexbw@, mattjj@* \n\nJAX has a pretty general automatic differentiation system. In this notebook, we'll go through a whole bunch of neat autodiff ideas that you can cherry pick for your own work, starting with the basics.",
"_____no_output_____"
]
],
[
[
"import jax.numpy as jnp\nfrom jax import grad, jit, vmap\nfrom jax import random\n\nkey = random.PRNGKey(0)",
"_____no_output_____"
]
],
[
[
"## Gradients",
"_____no_output_____"
],
[
"### Starting with `grad`\n\nYou can differentiate a function with `grad`:",
"_____no_output_____"
]
],
[
[
"grad_tanh = grad(jnp.tanh)\nprint(grad_tanh(2.0))",
"0.070650935\n"
]
],
[
[
"`grad` takes a function and returns a function. If you have a Python function `f` that evaluates the mathematical function $f$, then `grad(f)` is a Python function that evaluates the mathematical function $\\nabla f$. That means `grad(f)(x)` represents the value $\\nabla f(x)$.\n\nSince `grad` operates on functions, you can apply it to its own output to differentiate as many times as you like:",
"_____no_output_____"
]
],
[
[
"print(grad(grad(jnp.tanh))(2.0))\nprint(grad(grad(grad(jnp.tanh)))(2.0))",
"-0.13621889\n0.2526544\n"
]
],
[
[
"Let's look at computing gradients with `grad` in a linear logistic regression model. First, the setup:",
"_____no_output_____"
]
],
[
[
"def sigmoid(x):\n return 0.5 * (jnp.tanh(x / 2) + 1)\n\n# Outputs probability of a label being true.\ndef predict(W, b, inputs):\n return sigmoid(jnp.dot(inputs, W) + b)\n\n# Build a toy dataset.\ninputs = jnp.array([[0.52, 1.12, 0.77],\n [0.88, -1.08, 0.15],\n [0.52, 0.06, -1.30],\n [0.74, -2.49, 1.39]])\ntargets = jnp.array([True, True, False, True])\n\n# Training loss is the negative log-likelihood of the training examples.\ndef loss(W, b):\n preds = predict(W, b, inputs)\n label_probs = preds * targets + (1 - preds) * (1 - targets)\n return -jnp.sum(jnp.log(label_probs))\n\n# Initialize random model coefficients\nkey, W_key, b_key = random.split(key, 3)\nW = random.normal(W_key, (3,))\nb = random.normal(b_key, ())",
"_____no_output_____"
]
],
[
[
"Use the `grad` function with its `argnums` argument to differentiate a function with respect to positional arguments.",
"_____no_output_____"
]
],
[
[
"# Differentiate `loss` with respect to the first positional argument:\nW_grad = grad(loss, argnums=0)(W, b)\nprint('W_grad', W_grad)\n\n# Since argnums=0 is the default, this does the same thing:\nW_grad = grad(loss)(W, b)\nprint('W_grad', W_grad)\n\n# But we can choose different values too, and drop the keyword:\nb_grad = grad(loss, 1)(W, b)\nprint('b_grad', b_grad)\n\n# Including tuple values\nW_grad, b_grad = grad(loss, (0, 1))(W, b)\nprint('W_grad', W_grad)\nprint('b_grad', b_grad)",
"W_grad [-0.16965576 -0.8774645 -1.4901344 ]\nW_grad [-0.16965576 -0.8774645 -1.4901344 ]\nb_grad -0.29227236\nW_grad [-0.16965576 -0.8774645 -1.4901344 ]\nb_grad -0.29227236\n"
]
],
[
[
"This `grad` API has a direct correspondence to the excellent notation in Spivak's classic *Calculus on Manifolds* (1965), also used in Sussman and Wisdom's [*Structure and Interpretation of Classical Mechanics*](http://mitpress.mit.edu/sites/default/files/titles/content/sicm_edition_2/book.html) (2015) and their [*Functional Differential Geometry*](https://mitpress.mit.edu/books/functional-differential-geometry) (2013). Both books are open-access. See in particular the \"Prologue\" section of *Functional Differential Geometry* for a defense of this notation.\n\nEssentially, when using the `argnums` argument, if `f` is a Python function for evaluating the mathematical function $f$, then the Python expression `grad(f, i)` evaluates to a Python function for evaluating $\\partial_i f$.",
"_____no_output_____"
],
[
"### Differentiating with respect to nested lists, tuples, and dicts",
"_____no_output_____"
],
[
"Differentiating with respect to standard Python containers just works, so use tuples, lists, and dicts (and arbitrary nesting) however you like.",
"_____no_output_____"
]
],
[
[
"def loss2(params_dict):\n preds = predict(params_dict['W'], params_dict['b'], inputs)\n label_probs = preds * targets + (1 - preds) * (1 - targets)\n return -jnp.sum(jnp.log(label_probs))\n\nprint(grad(loss2)({'W': W, 'b': b}))",
"{'W': DeviceArray([-0.16965576, -0.8774645 , -1.4901344 ], dtype=float32), 'b': DeviceArray(-0.29227236, dtype=float32)}\n"
]
],
[
[
"You can [register your own container types](https://github.com/google/jax/issues/446#issuecomment-467105048) to work with not just `grad` but all the JAX transformations (`jit`, `vmap`, etc.).",
"_____no_output_____"
],
[
"### Evaluate a function and its gradient using `value_and_grad`",
"_____no_output_____"
],
[
"Another convenient function is `value_and_grad` for efficiently computing both a function's value as well as its gradient's value:",
"_____no_output_____"
]
],
[
[
"from jax import value_and_grad\nloss_value, Wb_grad = value_and_grad(loss, (0, 1))(W, b)\nprint('loss value', loss_value)\nprint('loss value', loss(W, b))",
"loss value 3.051939\nloss value 3.051939\n"
]
],
[
[
"### Checking against numerical differences\n\nA great thing about derivatives is that they're straightforward to check with finite differences:",
"_____no_output_____"
]
],
[
[
"# Set a step size for finite differences calculations\neps = 1e-4\n\n# Check b_grad with scalar finite differences\nb_grad_numerical = (loss(W, b + eps / 2.) - loss(W, b - eps / 2.)) / eps\nprint('b_grad_numerical', b_grad_numerical)\nprint('b_grad_autodiff', grad(loss, 1)(W, b))\n\n# Check W_grad with finite differences in a random direction\nkey, subkey = random.split(key)\nvec = random.normal(subkey, W.shape)\nunitvec = vec / jnp.sqrt(jnp.vdot(vec, vec))\nW_grad_numerical = (loss(W + eps / 2. * unitvec, b) - loss(W - eps / 2. * unitvec, b)) / eps\nprint('W_dirderiv_numerical', W_grad_numerical)\nprint('W_dirderiv_autodiff', jnp.vdot(grad(loss)(W, b), unitvec))",
"b_grad_numerical -0.29563904\nb_grad_autodiff -0.29227236\nW_dirderiv_numerical -0.19788742\nW_dirderiv_autodiff -0.19909099\n"
]
],
[
[
"JAX provides a simple convenience function that does essentially the same thing, but checks up to any order of differentiation that you like:",
"_____no_output_____"
]
],
[
[
"from jax.test_util import check_grads\ncheck_grads(loss, (W, b), order=2) # check up to 2nd order derivatives",
"_____no_output_____"
]
],
[
[
"### Hessian-vector products with `grad`-of-`grad`\n\nOne thing we can do with higher-order `grad` is build a Hessian-vector product function. (Later on we'll write an even more efficient implementation that mixes both forward- and reverse-mode, but this one will use pure reverse-mode.)\n\nA Hessian-vector product function can be useful in a [truncated Newton Conjugate-Gradient algorithm](https://en.wikipedia.org/wiki/Truncated_Newton_method) for minimizing smooth convex functions, or for studying the curvature of neural network training objectives (e.g. [1](https://arxiv.org/abs/1406.2572), [2](https://arxiv.org/abs/1811.07062), [3](https://arxiv.org/abs/1706.04454), [4](https://arxiv.org/abs/1802.03451)).\n\nFor a scalar-valued function $f : \\mathbb{R}^n \\to \\mathbb{R}$ with continuous second derivatives (so that the Hessian matrix is symmetric), the Hessian at a point $x \\in \\mathbb{R}^n$ is written as $\\partial^2 f(x)$. A Hessian-vector product function is then able to evaluate\n\n$\\qquad v \\mapsto \\partial^2 f(x) \\cdot v$\n\nfor any $v \\in \\mathbb{R}^n$.\n\nThe trick is not to instantiate the full Hessian matrix: if $n$ is large, perhaps in the millions or billions in the context of neural networks, then that might be impossible to store.\n\nLuckily, `grad` already gives us a way to write an efficient Hessian-vector product function. We just have to use the identity\n\n$\\qquad \\partial^2 f (x) v = \\partial [x \\mapsto \\partial f(x) \\cdot v] = \\partial g(x)$,\n\nwhere $g(x) = \\partial f(x) \\cdot v$ is a new scalar-valued function that dots the gradient of $f$ at $x$ with the vector $v$. Notice that we're only ever differentiating scalar-valued functions of vector-valued arguments, which is exactly where we know `grad` is efficient.\n\nIn JAX code, we can just write this:",
"_____no_output_____"
]
],
[
[
"def hvp(f, x, v):\n return grad(lambda x: jnp.vdot(grad(f)(x), v))(x)",
"_____no_output_____"
]
],
[
[
"This example shows that you can freely use lexical closure, and JAX will never get perturbed or confused.\n\nWe'll check this implementation a few cells down, once we see how to compute dense Hessian matrices. We'll also write an even better version that uses both forward-mode and reverse-mode.",
"_____no_output_____"
],
[
"### Jacobians and Hessians using `jacfwd` and `jacrev`",
"_____no_output_____"
],
[
"You can compute full Jacobian matrices using the `jacfwd` and `jacrev` functions:",
"_____no_output_____"
]
],
[
[
"from jax import jacfwd, jacrev\n\n# Isolate the function from the weight matrix to the predictions\nf = lambda W: predict(W, b, inputs)\n\nJ = jacfwd(f)(W)\nprint(\"jacfwd result, with shape\", J.shape)\nprint(J)\n\nJ = jacrev(f)(W)\nprint(\"jacrev result, with shape\", J.shape)\nprint(J)",
"jacfwd result, with shape (4, 3)\n[[ 0.05981756 0.12883782 0.088576 ]\n [ 0.04015914 -0.04928622 0.00684531]\n [ 0.12188289 0.01406341 -0.3047072 ]\n [ 0.00140428 -0.00472522 0.00263777]]\njacrev result, with shape (4, 3)\n[[ 0.05981756 0.12883782 0.088576 ]\n [ 0.04015914 -0.04928622 0.00684531]\n [ 0.12188289 0.01406341 -0.3047072 ]\n [ 0.00140428 -0.00472522 0.00263777]]\n"
]
],
[
[
"These two functions compute the same values (up to machine numerics), but differ in their implementation: `jacfwd` uses forward-mode automatic differentiation, which is more efficient for \"tall\" Jacobian matrices, while `jacrev` uses reverse-mode, which is more efficient for \"wide\" Jacobian matrices. For matrices that are near-square, `jacfwd` probably has an edge over `jacrev`.",
"_____no_output_____"
],
[
"You can also use `jacfwd` and `jacrev` with container types:",
"_____no_output_____"
]
],
[
[
"def predict_dict(params, inputs):\n return predict(params['W'], params['b'], inputs)\n\nJ_dict = jacrev(predict_dict)({'W': W, 'b': b}, inputs)\nfor k, v in J_dict.items():\n print(\"Jacobian from {} to logits is\".format(k))\n print(v)",
"Jacobian from W to logits is\n[[ 0.05981756 0.12883782 0.088576 ]\n [ 0.04015914 -0.04928622 0.00684531]\n [ 0.12188289 0.01406341 -0.3047072 ]\n [ 0.00140428 -0.00472522 0.00263777]]\nJacobian from b to logits is\n[0.11503378 0.04563539 0.23439017 0.00189768]\n"
]
],
[
[
"For more details on forward- and reverse-mode, as well as how to implement `jacfwd` and `jacrev` as efficiently as possible, read on!",
"_____no_output_____"
],
[
"Using a composition of two of these functions gives us a way to compute dense Hessian matrices:",
"_____no_output_____"
]
],
[
[
"def hessian(f):\n return jacfwd(jacrev(f))\n\nH = hessian(f)(W)\nprint(\"hessian, with shape\", H.shape)\nprint(H)",
"hessian, with shape (4, 3, 3)\n[[[ 0.02285465 0.0492254 0.03384246]\n [ 0.0492254 0.10602394 0.07289147]\n [ 0.03384246 0.07289146 0.05011288]]\n\n [[-0.03195214 0.03921399 -0.00544639]\n [ 0.03921399 -0.04812626 0.0066842 ]\n [-0.00544639 0.0066842 -0.00092836]]\n\n [[-0.01583708 -0.00182736 0.03959271]\n [-0.00182736 -0.00021085 0.00456839]\n [ 0.03959271 0.00456839 -0.09898177]]\n\n [[-0.00103522 0.00348336 -0.00194453]\n [ 0.00348336 -0.01172105 0.00654308]\n [-0.00194453 0.00654308 -0.00365256]]]\n"
]
],
[
[
"This shape makes sense: if we start with a function $f : \\mathbb{R}^n \\to \\mathbb{R}^m$, then at a point $x \\in \\mathbb{R}^n$ we expect to get the shapes\n\n* $f(x) \\in \\mathbb{R}^m$, the value of $f$ at $x$,\n* $\\partial f(x) \\in \\mathbb{R}^{m \\times n}$, the Jacobian matrix at $x$,\n* $\\partial^2 f(x) \\in \\mathbb{R}^{m \\times n \\times n}$, the Hessian at $x$,\n\nand so on.\n\nTo implement `hessian`, we could have used `jacfwd(jacrev(f))` or `jacrev(jacfwd(f))` or any other composition of the two. But forward-over-reverse is typically the most efficient. That's because in the inner Jacobian computation we're often differentiating a function wide Jacobian (maybe like a loss function $f : \\mathbb{R}^n \\to \\mathbb{R}$), while in the outer Jacobian computation we're differentiating a function with a square Jacobian (since $\\nabla f : \\mathbb{R}^n \\to \\mathbb{R}^n$), which is where forward-mode wins out.",
"_____no_output_____"
],
[
"## How it's made: two foundational autodiff functions",
"_____no_output_____"
],
[
"### Jacobian-Vector products (JVPs, aka forward-mode autodiff)\n\nJAX includes efficient and general implementations of both forward- and reverse-mode automatic differentiation. The familiar `grad` function is built on reverse-mode, but to explain the difference in the two modes, and when each can be useful, we need a bit of math background.\n\n#### JVPs in math\n\nMathematically, given a function $f : \\mathbb{R}^n \\to \\mathbb{R}^m$, the Jacobian of $f$ evaluated at an input point $x \\in \\mathbb{R}^n$, denoted $\\partial f(x)$, is often thought of as a matrix in $\\mathbb{R}^m \\times \\mathbb{R}^n$:\n\n$\\qquad \\partial f(x) \\in \\mathbb{R}^{m \\times n}$.\n\nBut we can also think of $\\partial f(x)$ as a linear map, which maps the tangent space of the domain of $f$ at the point $x$ (which is just another copy of $\\mathbb{R}^n$) to the tangent space of the codomain of $f$ at the point $f(x)$ (a copy of $\\mathbb{R}^m$):\n\n$\\qquad \\partial f(x) : \\mathbb{R}^n \\to \\mathbb{R}^m$.\n\nThis map is called the [pushforward map](https://en.wikipedia.org/wiki/Pushforward_(differential)) of $f$ at $x$. The Jacobian matrix is just the matrix for this linear map in a standard basis.\n\nIf we don't commit to one specific input point $x$, then we can think of the function $\\partial f$ as first taking an input point and returning the Jacobian linear map at that input point:\n\n$\\qquad \\partial f : \\mathbb{R}^n \\to \\mathbb{R}^n \\to \\mathbb{R}^m$.\n\nIn particular, we can uncurry things so that given input point $x \\in \\mathbb{R}^n$ and a tangent vector $v \\in \\mathbb{R}^n$, we get back an output tangent vector in $\\mathbb{R}^m$. We call that mapping, from $(x, v)$ pairs to output tangent vectors, the *Jacobian-vector product*, and write it as\n\n$\\qquad (x, v) \\mapsto \\partial f(x) v$\n\n#### JVPs in JAX code\n\nBack in Python code, JAX's `jvp` function models this transformation. Given a Python function that evaluates $f$, JAX's `jvp` is a way to get a Python function for evaluating $(x, v) \\mapsto (f(x), \\partial f(x) v)$.",
"_____no_output_____"
]
],
[
[
"from jax import jvp\n\n# Isolate the function from the weight matrix to the predictions\nf = lambda W: predict(W, b, inputs)\n\nkey, subkey = random.split(key)\nv = random.normal(subkey, W.shape)\n\n# Push forward the vector `v` along `f` evaluated at `W`\ny, u = jvp(f, (W,), (v,))",
"_____no_output_____"
]
],
[
[
"In terms of Haskell-like type signatures, we could write\n\n```haskell\njvp :: (a -> b) -> a -> T a -> (b, T b)\n```\n\nwhere we use `T a` to denote the type of the tangent space for `a`. In words, `jvp` takes as arguments a function of type `a -> b`, a value of type `a`, and a tangent vector value of type `T a`. It gives back a pair consisting of a value of type `b` and an output tangent vector of type `T b`.",
"_____no_output_____"
],
[
"The `jvp`-transformed function is evaluated much like the original function, but paired up with each primal value of type `a` it pushes along tangent values of type `T a`. For each primitive numerical operation that the original function would have applied, the `jvp`-transformed function executes a \"JVP rule\" for that primitive that both evaluates the primitive on the primals and applies the primitive's JVP at those primal values.\n\nThat evaluation strategy has some immediate implications about computational complexity: since we evaluate JVPs as we go, we don't need to store anything for later, and so the memory cost is independent of the depth of the computation. In addition, the FLOP cost of the `jvp`-transformed function is about 3x the cost of just evaluating the function (one unit of work for evaluating the original function, for example `sin(x)`; one unit for linearizing, like `cos(x)`; and one unit for applying the linearized function to a vector, like `cos_x * v`). Put another way, for a fixed primal point $x$, we can evaluate $v \\mapsto \\partial f(x) \\cdot v$ for about the same marginal cost as evaluating $f$.\n\nThat memory complexity sounds pretty compelling! So why don't we see forward-mode very often in machine learning?\n\nTo answer that, first think about how you could use a JVP to build a full Jacobian matrix. If we apply a JVP to a one-hot tangent vector, it reveals one column of the Jacobian matrix, corresponding to the nonzero entry we fed in. So we can build a full Jacobian one column at a time, and to get each column costs about the same as one function evaluation. That will be efficient for functions with \"tall\" Jacobians, but inefficient for \"wide\" Jacobians.\n\nIf you're doing gradient-based optimization in machine learning, you probably want to minimize a loss function from parameters in $\\mathbb{R}^n$ to a scalar loss value in $\\mathbb{R}$. That means the Jacobian of this function is a very wide matrix: $\\partial f(x) \\in \\mathbb{R}^{1 \\times n}$, which we often identify with the Gradient vector $\\nabla f(x) \\in \\mathbb{R}^n$. Building that matrix one column at a time, with each call taking a similar number of FLOPs to evaluating the original function, sure seems inefficient! In particular, for training neural networks, where $f$ is a training loss function and $n$ can be in the millions or billions, this approach just won't scale.\n\nTo do better for functions like this, we just need to use reverse-mode.",
"_____no_output_____"
],
[
"### Vector-Jacobian products (VJPs, aka reverse-mode autodiff)\n\nWhere forward-mode gives us back a function for evaluating Jacobian-vector products, which we can then use to build Jacobian matrices one column at a time, reverse-mode is a way to get back a function for evaluating vector-Jacobian products (equivalently Jacobian-transpose-vector products), which we can use to build Jacobian matrices one row at a time.\n\n#### VJPs in math\n\nLet's again consider a function $f : \\mathbb{R}^n \\to \\mathbb{R}^m$.\nStarting from our notation for JVPs, the notation for VJPs is pretty simple:\n\n$\\qquad (x, v) \\mapsto v \\partial f(x)$,\n\nwhere $v$ is an element of the cotangent space of $f$ at $x$ (isomorphic to another copy of $\\mathbb{R}^m$). When being rigorous, we should think of $v$ as a linear map $v : \\mathbb{R}^m \\to \\mathbb{R}$, and when we write $v \\partial f(x)$ we mean function composition $v \\circ \\partial f(x)$, where the types work out because $\\partial f(x) : \\mathbb{R}^n \\to \\mathbb{R}^m$. But in the common case we can identify $v$ with a vector in $\\mathbb{R}^m$ and use the two almost interchageably, just like we might sometimes flip between \"column vectors\" and \"row vectors\" without much comment.\n\nWith that identification, we can alternatively think of the linear part of a VJP as the transpose (or adjoint conjugate) of the linear part of a JVP:\n\n$\\qquad (x, v) \\mapsto \\partial f(x)^\\mathsf{T} v$.\n\nFor a given point $x$, we can write the signature as\n\n$\\qquad \\partial f(x)^\\mathsf{T} : \\mathbb{R}^m \\to \\mathbb{R}^n$.\n\nThe corresponding map on cotangent spaces is often called the [pullback](https://en.wikipedia.org/wiki/Pullback_(differential_geometry))\nof $f$ at $x$. The key for our purposes is that it goes from something that looks like the output of $f$ to something that looks like the input of $f$, just like we might expect from a transposed linear function.\n\n#### VJPs in JAX code\n\nSwitching from math back to Python, the JAX function `vjp` can take a Python function for evaluating $f$ and give us back a Python function for evaluating the VJP $(x, v) \\mapsto (f(x), v^\\mathsf{T} \\partial f(x))$.",
"_____no_output_____"
]
],
[
[
"from jax import vjp\n\n# Isolate the function from the weight matrix to the predictions\nf = lambda W: predict(W, b, inputs)\n\ny, vjp_fun = vjp(f, W)\n\nkey, subkey = random.split(key)\nu = random.normal(subkey, y.shape)\n\n# Pull back the covector `u` along `f` evaluated at `W`\nv = vjp_fun(u)",
"_____no_output_____"
]
],
[
[
"In terms of Haskell-like type signatures, we could write\n\n```haskell\nvjp :: (a -> b) -> a -> (b, CT b -> CT a)\n```\n\nwhere we use `CT a` to denote the type for the cotangent space for `a`. In words, `vjp` takes as arguments a function of type `a -> b` and a point of type `a`, and gives back a pair consisting of a value of type `b` and a linear map of type `CT b -> CT a`.\n\nThis is great because it lets us build Jacobian matrices one row at a time, and the FLOP cost for evaluating $(x, v) \\mapsto (f(x), v^\\mathsf{T} \\partial f(x))$ is only about three times the cost of evaluating $f$. In particular, if we want the gradient of a function $f : \\mathbb{R}^n \\to \\mathbb{R}$, we can do it in just one call. That's how `grad` is efficient for gradient-based optimization, even for objectives like neural network training loss functions on millions or billions of parameters.\n\nThere's a cost, though: though the FLOPs are friendly, memory scales with the depth of the computation. Also, the implementation is traditionally more complex than that of forward-mode, though JAX has some tricks up its sleeve (that's a story for a future notebook!).\n\nFor more on how reverse-mode works, see [this tutorial video from the Deep Learning Summer School in 2017](http://videolectures.net/deeplearning2017_johnson_automatic_differentiation/).",
"_____no_output_____"
],
[
"### Vector-valued gradients with VJPs\n\nIf you're interested in taking vector-valued gradients (like `tf.gradients`):",
"_____no_output_____"
]
],
[
[
"from jax import vjp\n\ndef vgrad(f, x):\n y, vjp_fn = vjp(f, x)\n return vjp_fn(jnp.ones(y.shape))[0]\n\nprint(vgrad(lambda x: 3*x**2, jnp.ones((2, 2))))",
"[[6. 6.]\n [6. 6.]]\n"
]
],
[
[
"### Hessian-vector products using both forward- and reverse-mode",
"_____no_output_____"
],
[
"In a previous section, we implemented a Hessian-vector product function just using reverse-mode (assuming continuous second derivatives):",
"_____no_output_____"
]
],
[
[
"def hvp(f, x, v):\n return grad(lambda x: jnp.vdot(grad(f)(x), v))(x)",
"_____no_output_____"
]
],
[
[
"That's efficient, but we can do even better and save some memory by using forward-mode together with reverse-mode.\n\nMathematically, given a function $f : \\mathbb{R}^n \\to \\mathbb{R}$ to differentiate, a point $x \\in \\mathbb{R}^n$ at which to linearize the function, and a vector $v \\in \\mathbb{R}^n$, the Hessian-vector product function we want is\n\n$(x, v) \\mapsto \\partial^2 f(x) v$\n\nConsider the helper function $g : \\mathbb{R}^n \\to \\mathbb{R}^n$ defined to be the derivative (or gradient) of $f$, namely $g(x) = \\partial f(x)$. All we need is its JVP, since that will give us\n\n$(x, v) \\mapsto \\partial g(x) v = \\partial^2 f(x) v$.\n\nWe can translate that almost directly into code:",
"_____no_output_____"
]
],
[
[
"from jax import jvp, grad\n\n# forward-over-reverse\ndef hvp(f, primals, tangents):\n return jvp(grad(f), primals, tangents)[1]",
"_____no_output_____"
]
],
[
[
"Even better, since we didn't have to call `jnp.dot` directly, this `hvp` function works with arrays of any shape and with arbitrary container types (like vectors stored as nested lists/dicts/tuples), and doesn't even have a dependence on `jax.numpy`.\n\nHere's an example of how to use it:",
"_____no_output_____"
]
],
[
[
"def f(X):\n return jnp.sum(jnp.tanh(X)**2)\n\nkey, subkey1, subkey2 = random.split(key, 3)\nX = random.normal(subkey1, (30, 40))\nV = random.normal(subkey2, (30, 40))\n\nans1 = hvp(f, (X,), (V,))\nans2 = jnp.tensordot(hessian(f)(X), V, 2)\n\nprint(jnp.allclose(ans1, ans2, 1e-4, 1e-4))",
"True\n"
]
],
[
[
"Another way you might consider writing this is using reverse-over-forward:",
"_____no_output_____"
]
],
[
[
"# reverse-over-forward\ndef hvp_revfwd(f, primals, tangents):\n g = lambda primals: jvp(f, primals, tangents)[1]\n return grad(g)(primals)",
"_____no_output_____"
]
],
[
[
"That's not quite as good, though, because forward-mode has less overhead than reverse-mode, and since the outer differentiation operator here has to differentiate a larger computation than the inner one, keeping forward-mode on the outside works best:",
"_____no_output_____"
]
],
[
[
"# reverse-over-reverse, only works for single arguments\ndef hvp_revrev(f, primals, tangents):\n x, = primals\n v, = tangents\n return grad(lambda x: jnp.vdot(grad(f)(x), v))(x)\n\n\nprint(\"Forward over reverse\")\n%timeit -n10 -r3 hvp(f, (X,), (V,))\nprint(\"Reverse over forward\")\n%timeit -n10 -r3 hvp_revfwd(f, (X,), (V,))\nprint(\"Reverse over reverse\")\n%timeit -n10 -r3 hvp_revrev(f, (X,), (V,))\n\nprint(\"Naive full Hessian materialization\")\n%timeit -n10 -r3 jnp.tensordot(hessian(f)(X), V, 2)",
"Forward over reverse\n10 loops, best of 3: 6.75 ms per loop\nReverse over forward\n10 loops, best of 3: 8.4 ms per loop\nReverse over reverse\n10 loops, best of 3: 9.98 ms per loop\nNaive full Hessian materialization\n10 loops, best of 3: 15.9 ms per loop\n"
]
],
[
[
"## Composing VJPs, JVPs, and `vmap`",
"_____no_output_____"
],
[
"### Jacobian-Matrix and Matrix-Jacobian products\n\nNow that we have `jvp` and `vjp` transformations that give us functions to push-forward or pull-back single vectors at a time, we can use JAX's `vmap` [transformation](https://github.com/google/jax#auto-vectorization-with-vmap) to push and pull entire bases at once. In particular, we can use that to write fast matrix-Jacobian and Jacobian-matrix products.",
"_____no_output_____"
]
],
[
[
"# Isolate the function from the weight matrix to the predictions\nf = lambda W: predict(W, b, inputs)\n\n# Pull back the covectors `m_i` along `f`, evaluated at `W`, for all `i`.\n# First, use a list comprehension to loop over rows in the matrix M.\ndef loop_mjp(f, x, M):\n y, vjp_fun = vjp(f, x)\n return jnp.vstack([vjp_fun(mi) for mi in M])\n\n# Now, use vmap to build a computation that does a single fast matrix-matrix\n# multiply, rather than an outer loop over vector-matrix multiplies.\ndef vmap_mjp(f, x, M):\n y, vjp_fun = vjp(f, x)\n outs, = vmap(vjp_fun)(M)\n return outs\n\nkey = random.PRNGKey(0)\nnum_covecs = 128\nU = random.normal(key, (num_covecs,) + y.shape)\n\nloop_vs = loop_mjp(f, W, M=U)\nprint('Non-vmapped Matrix-Jacobian product')\n%timeit -n10 -r3 loop_mjp(f, W, M=U)\n\nprint('\\nVmapped Matrix-Jacobian product')\nvmap_vs = vmap_mjp(f, W, M=U)\n%timeit -n10 -r3 vmap_mjp(f, W, M=U)\n\nassert jnp.allclose(loop_vs, vmap_vs), 'Vmap and non-vmapped Matrix-Jacobian Products should be identical'",
"Non-vmapped Matrix-Jacobian product\n10 loops, best of 3: 291 ms per loop\n\nVmapped Matrix-Jacobian product\n10 loops, best of 3: 6.85 ms per loop\n"
],
[
"def loop_jmp(f, W, M):\n # jvp immediately returns the primal and tangent values as a tuple,\n # so we'll compute and select the tangents in a list comprehension\n return jnp.vstack([jvp(f, (W,), (mi,))[1] for mi in M])\n\ndef vmap_jmp(f, W, M):\n _jvp = lambda s: jvp(f, (W,), (s,))[1]\n return vmap(_jvp)(M)\n\nnum_vecs = 128\nS = random.normal(key, (num_vecs,) + W.shape)\n\nloop_vs = loop_jmp(f, W, M=S)\nprint('Non-vmapped Jacobian-Matrix product')\n%timeit -n10 -r3 loop_jmp(f, W, M=S)\nvmap_vs = vmap_jmp(f, W, M=S)\nprint('\\nVmapped Jacobian-Matrix product')\n%timeit -n10 -r3 vmap_jmp(f, W, M=S)\n\nassert jnp.allclose(loop_vs, vmap_vs), 'Vmap and non-vmapped Jacobian-Matrix products should be identical'",
"Non-vmapped Jacobian-Matrix product\n10 loops, best of 3: 693 ms per loop\n\nVmapped Jacobian-Matrix product\n10 loops, best of 3: 5.97 ms per loop\n"
]
],
[
[
"### The implementation of `jacfwd` and `jacrev`",
"_____no_output_____"
],
[
"Now that we've seen fast Jacobian-matrix and matrix-Jacobian products, it's not hard to guess how to write `jacfwd` and `jacrev`. We just use the same technique to push-forward or pull-back an entire standard basis (isomorphic to an identity matrix) at once.",
"_____no_output_____"
]
],
[
[
"from jax import jacrev as builtin_jacrev\n\ndef our_jacrev(f):\n def jacfun(x):\n y, vjp_fun = vjp(f, x)\n # Use vmap to do a matrix-Jacobian product.\n # Here, the matrix is the Euclidean basis, so we get all\n # entries in the Jacobian at once. \n J, = vmap(vjp_fun, in_axes=0)(jnp.eye(len(y)))\n return J\n return jacfun\n\nassert jnp.allclose(builtin_jacrev(f)(W), our_jacrev(f)(W)), 'Incorrect reverse-mode Jacobian results!'",
"_____no_output_____"
],
[
"from jax import jacfwd as builtin_jacfwd\n\ndef our_jacfwd(f):\n def jacfun(x):\n _jvp = lambda s: jvp(f, (x,), (s,))[1]\n Jt =vmap(_jvp, in_axes=1)(jnp.eye(len(x)))\n return jnp.transpose(Jt)\n return jacfun\n\nassert jnp.allclose(builtin_jacfwd(f)(W), our_jacfwd(f)(W)), 'Incorrect forward-mode Jacobian results!'",
"_____no_output_____"
]
],
[
[
"Interestingly, [Autograd](https://github.com/hips/autograd) couldn't do this. Our [implementation](https://github.com/HIPS/autograd/blob/96a03f44da43cd7044c61ac945c483955deba957/autograd/differential_operators.py#L60) of reverse-mode `jacobian` in Autograd had to pull back one vector at a time with an outer-loop `map`. Pushing one vector at a time through the computation is much less efficient than batching it all together with `vmap`.",
"_____no_output_____"
],
[
"Another thing that Autograd couldn't do is `jit`. Interestingly, no matter how much Python dynamism you use in your function to be differentiated, we could always use `jit` on the linear part of the computation. For example:",
"_____no_output_____"
]
],
[
[
"def f(x):\n try:\n if x < 3:\n return 2 * x ** 3\n else:\n raise ValueError\n except ValueError:\n return jnp.pi * x\n\ny, f_vjp = vjp(f, 4.)\nprint(jit(f_vjp)(1.))",
"(DeviceArray(3.1415927, dtype=float32),)\n"
]
],
[
[
"## Complex numbers and differentiation",
"_____no_output_____"
],
[
"JAX is great at complex numbers and differentiation. To support both [holomorphic and non-holomorphic differentiation](https://en.wikipedia.org/wiki/Holomorphic_function), it helps to think in terms of JVPs and VJPs.\n\nConsider a complex-to-complex function $f: \\mathbb{C} \\to \\mathbb{C}$ and identify it with a corresponding function $g: \\mathbb{R}^2 \\to \\mathbb{R}^2$,",
"_____no_output_____"
]
],
[
[
"def f(z):\n x, y = jnp.real(z), jnp.imag(z)\n return u(x, y) + v(x, y) * 1j\n\ndef g(x, y):\n return (u(x, y), v(x, y))",
"_____no_output_____"
]
],
[
[
"That is, we've decomposed $f(z) = u(x, y) + v(x, y) i$ where $z = x + y i$, and identified $\\mathbb{C}$ with $\\mathbb{R}^2$ to get $g$.",
"_____no_output_____"
],
[
"Since $g$ only involves real inputs and outputs, we already know how to write a Jacobian-vector product for it, say given a tangent vector $(c, d) \\in \\mathbb{R}^2$, namely\n\n$\\begin{bmatrix} \\partial_0 u(x, y) & \\partial_1 u(x, y) \\\\ \\partial_0 v(x, y) & \\partial_1 v(x, y) \\end{bmatrix}\n\\begin{bmatrix} c \\\\ d \\end{bmatrix}$.\n\nTo get a JVP for the original function $f$ applied to a tangent vector $c + di \\in \\mathbb{C}$, we just use the same definition and identify the result as another complex number, \n\n$\\partial f(x + y i)(c + d i) =\n\\begin{matrix} \\begin{bmatrix} 1 & i \\end{bmatrix} \\\\ ~ \\end{matrix}\n\\begin{bmatrix} \\partial_0 u(x, y) & \\partial_1 u(x, y) \\\\ \\partial_0 v(x, y) & \\partial_1 v(x, y) \\end{bmatrix}\n\\begin{bmatrix} c \\\\ d \\end{bmatrix}$.\n\nThat's our definition of the JVP of a $\\mathbb{C} \\to \\mathbb{C}$ function! Notice it doesn't matter whether or not $f$ is holomorphic: the JVP is unambiguous.",
"_____no_output_____"
],
[
"Here's a check:",
"_____no_output_____"
]
],
[
[
"def check(seed):\n key = random.PRNGKey(seed)\n\n # random coeffs for u and v\n key, subkey = random.split(key)\n a, b, c, d = random.uniform(subkey, (4,))\n\n def fun(z):\n x, y = jnp.real(z), jnp.imag(z)\n return u(x, y) + v(x, y) * 1j\n\n def u(x, y):\n return a * x + b * y\n\n def v(x, y):\n return c * x + d * y\n\n # primal point\n key, subkey = random.split(key)\n x, y = random.uniform(subkey, (2,))\n z = x + y * 1j\n\n # tangent vector\n key, subkey = random.split(key)\n c, d = random.uniform(subkey, (2,))\n z_dot = c + d * 1j\n\n # check jvp\n _, ans = jvp(fun, (z,), (z_dot,))\n expected = (grad(u, 0)(x, y) * c +\n grad(u, 1)(x, y) * d +\n grad(v, 0)(x, y) * c * 1j+\n grad(v, 1)(x, y) * d * 1j)\n print(jnp.allclose(ans, expected))",
"_____no_output_____"
],
[
"check(0)\ncheck(1)\ncheck(2)",
"True\nTrue\nTrue\n"
]
],
[
[
"What about VJPs? We do something pretty similar: for a cotangent vector $c + di \\in \\mathbb{C}$ we define the VJP of $f$ as\n\n$(c + di)^* \\; \\partial f(x + y i) =\n\\begin{matrix} \\begin{bmatrix} c & -d \\end{bmatrix} \\\\ ~ \\end{matrix}\n\\begin{bmatrix} \\partial_0 u(x, y) & \\partial_1 u(x, y) \\\\ \\partial_0 v(x, y) & \\partial_1 v(x, y) \\end{bmatrix}\n\\begin{bmatrix} 1 \\\\ -i \\end{bmatrix}$.\n\nWhat's with the negatives? They're just to take care of complex conjugation, and the fact that we're working with covectors.",
"_____no_output_____"
],
[
"Here's a check of the VJP rules:",
"_____no_output_____"
]
],
[
[
"def check(seed):\n key = random.PRNGKey(seed)\n\n # random coeffs for u and v\n key, subkey = random.split(key)\n a, b, c, d = random.uniform(subkey, (4,))\n\n def fun(z):\n x, y = jnp.real(z), jnp.imag(z)\n return u(x, y) + v(x, y) * 1j\n\n def u(x, y):\n return a * x + b * y\n\n def v(x, y):\n return c * x + d * y\n\n # primal point\n key, subkey = random.split(key)\n x, y = random.uniform(subkey, (2,))\n z = x + y * 1j\n\n # cotangent vector\n key, subkey = random.split(key)\n c, d = random.uniform(subkey, (2,))\n z_bar = jnp.array(c + d * 1j) # for dtype control\n\n # check vjp\n _, fun_vjp = vjp(fun, z)\n ans, = fun_vjp(z_bar)\n expected = (grad(u, 0)(x, y) * c +\n grad(v, 0)(x, y) * (-d) +\n grad(u, 1)(x, y) * c * (-1j) +\n grad(v, 1)(x, y) * (-d) * (-1j))\n assert jnp.allclose(ans, expected, atol=1e-5, rtol=1e-5)",
"_____no_output_____"
],
[
"check(0)\ncheck(1)\ncheck(2)",
"_____no_output_____"
]
],
[
[
"What about convenience wrappers like `grad`, `jacfwd`, and `jacrev`?\n\nFor $\\mathbb{R} \\to \\mathbb{R}$ functions, recall we defined `grad(f)(x)` as being `vjp(f, x)[1](1.0)`, which works because applying a VJP to a `1.0` value reveals the gradient (i.e. Jacobian, or derivative). We can do the same thing for $\\mathbb{C} \\to \\mathbb{R}$ functions: we can still use `1.0` as the cotangent vector, and we just get out a complex number result summarizing the full Jacobian:",
"_____no_output_____"
]
],
[
[
"def f(z):\n x, y = jnp.real(z), jnp.imag(z)\n return x**2 + y**2\n\nz = 3. + 4j\ngrad(f)(z)",
"_____no_output_____"
]
],
[
[
"For geneneral $\\mathbb{C} \\to \\mathbb{C}$ functions, the Jacobian has 4 real-valued degrees of freedom (as in the 2x2 Jacobian matrices above), so we can't hope to represent all of them with in a complex number. But we can for holomorphic functions! A holomorphic function is precisely a $\\mathbb{C} \\to \\mathbb{C}$ function with the special property that its derivative can be represented as a single complex number. (The [Cauchy-Riemann equations](https://en.wikipedia.org/wiki/Cauchy%E2%80%93Riemann_equations) ensure that the above 2x2 Jacobians have the special form of a scale-and-rotate matrix in the complex plane, i.e. the action of a single complex number under multiplication.) And we can reveal that one complex number using a single call to `vjp` with a covector of `1.0`.\n\nBecause this only works for holomorphic functions, to use this trick we need to promise JAX that our function is holomorphic; otherwise, JAX will raise an error when `grad` is used for a complex-output function:",
"_____no_output_____"
]
],
[
[
"def f(z):\n return jnp.sin(z)\n\nz = 3. + 4j\ngrad(f, holomorphic=True)(z)",
"_____no_output_____"
]
],
[
[
"All the `holomorphic=True` promise does is disable the error when the output is complex-valued. We can still write `holomorphic=True` when the function isn't holomorphic, but the answer we get out won't represent the full Jacobian. Instead, it'll be the Jacobian of the function where we just discard the imaginary part of the output:",
"_____no_output_____"
]
],
[
[
"def f(z):\n return jnp.conjugate(z)\n\nz = 3. + 4j\ngrad(f, holomorphic=True)(z) # f is not actually holomorphic!",
"_____no_output_____"
]
],
[
[
"There are some useful upshots for how `grad` works here:\n\n1. We can use `grad` on holomorphic $\\mathbb{C} \\to \\mathbb{C}$ functions.\n2. We can use `grad` to optimize $f : \\mathbb{C} \\to \\mathbb{R}$ functions, like real-valued loss functions of complex parameters `x`, by taking steps in the dierction of the conjugate of `grad(f)(x)`.\n3. If we have an $\\mathbb{R} \\to \\mathbb{R}$ function that just happens to use some complex-valued operations internally (some of which must be non-holomorphic, e.g. FFTs used in covolutions) then `grad` still works and we get the same result that an implementation using only real values would have given.\n\nIn any case, JVPs and VJPs are always unambiguous. And if we wanted to compute the full Jacobian matrix of a non-holomorphic $\\mathbb{C} \\to \\mathbb{C}$ function, we can do it with JVPs or VJPs!",
"_____no_output_____"
],
[
"You should expect complex numbers to work everywhere in JAX. Here's differentiating through a Cholesky decomposition of a complex matrix:",
"_____no_output_____"
]
],
[
[
"A = jnp.array([[5., 2.+3j, 5j],\n [2.-3j, 7., 1.+7j],\n [-5j, 1.-7j, 12.]])\n\ndef f(X):\n L = jnp.linalg.cholesky(X)\n return jnp.sum((L - jnp.sin(L))**2)\n\ngrad(f, holomorphic=True)(A)",
"_____no_output_____"
]
],
[
[
"## More advanced autodiff\n\nIn this notebook, we worked through some easy, and then progressively more complicated, applications of automatic differentiation in JAX. We hope you now feel that taking derivatives in JAX is easy and powerful. \n\nThere's a whole world of other autodiff tricks and functionality out there. Topics we didn't cover, but hope to in a \"Advanced Autodiff Cookbook\" include:\n\n - Gauss-Newton Vector Products, linearizing once\n - Custom VJPs and JVPs\n - Efficient derivatives at fixed-points\n - Estimating the trace of a Hessian using random Hessian-vector products.\n - Forward-mode autodiff using only reverse-mode autodiff.\n - Taking derivatives with respect to custom data types.\n - Checkpointing (binomial checkpointing for efficient reverse-mode, not model snapshotting).\n - Optimizing VJPs with Jacobian pre-accumulation.",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
4ab475be0c8db0cb75e1eb1bf0c5f58a680d1604
| 2,664 |
ipynb
|
Jupyter Notebook
|
01-reproducible_neuroimaging/01_02_fMRIDA_homework.ipynb
|
aterelak/fmri-analysis-course
|
9ed0f7e764974bf79f843f63ece36db11f46d662
|
[
"MIT"
] | 1 |
2020-05-25T11:01:58.000Z
|
2020-05-25T11:01:58.000Z
|
01-reproducible_neuroimaging/01_02_fMRIDA_homework.ipynb
|
aterelak/fmri-analysis-course
|
9ed0f7e764974bf79f843f63ece36db11f46d662
|
[
"MIT"
] | null | null | null |
01-reproducible_neuroimaging/01_02_fMRIDA_homework.ipynb
|
aterelak/fmri-analysis-course
|
9ed0f7e764974bf79f843f63ece36db11f46d662
|
[
"MIT"
] | null | null | null | 21.142857 | 138 | 0.53979 |
[
[
[
"# Advanced fMRI data analysis \n## Homework for courses: \n- #1 Introduction\n- #2 Reproducible neuroimaging",
"_____no_output_____"
],
[
"In this assignment, your task is to answer a few questions related to neuroimaging using Markdown. \n\nYou can choose to answer in Polish or English (it will not affect your grade).\n\nWrite answers in your **own words**, as you would like to explain a friend (do not copy from Wikipedia!). \n\nUse Markdown formatting options as much as possible!\n\n\n**DEADLINE:** 20-03-2020\n\n\n--------------------\n",
"_____no_output_____"
],
[
"**1. Explain in your *own words* how fMRI works (2-3 sentences).**",
"_____no_output_____"
],
[
"Your response",
"_____no_output_____"
],
[
"**2. Explain in your *own words* the term *BOLD signal* (2-3 sentences).**",
"_____no_output_____"
],
[
"Your response",
"_____no_output_____"
],
[
"**3. Make a table (two columns) with advantages and disadvantages of fMRI.**",
"_____no_output_____"
],
[
"Your response",
"_____no_output_____"
],
[
"**4. Explain in your *own words* what *reproducible research* is.**\nHighly recomended video: [From interactive exploration to reproducible data science](https://www.youtube.com/watch?v=xOKPKiAhey4).\n",
"_____no_output_____"
],
[
"Your response",
"_____no_output_____"
],
[
"**5. Make a *list* with good neuroimaging research practices that improve reproducibility of results.**",
"_____no_output_____"
],
[
"Your response",
"_____no_output_____"
]
]
] |
[
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
4ab48600f85447a8cfac890393e4a776367209c4
| 916,580 |
ipynb
|
Jupyter Notebook
|
A2/codes/Codes/Task 4/Task_4.ipynb
|
BrandonChuaa/URBAN-COMPUTING-AI6128
|
ca2106b4bdf4385512431606d867cbbd7aa6c3e9
|
[
"MIT"
] | null | null | null |
A2/codes/Codes/Task 4/Task_4.ipynb
|
BrandonChuaa/URBAN-COMPUTING-AI6128
|
ca2106b4bdf4385512431606d867cbbd7aa6c3e9
|
[
"MIT"
] | null | null | null |
A2/codes/Codes/Task 4/Task_4.ipynb
|
BrandonChuaa/URBAN-COMPUTING-AI6128
|
ca2106b4bdf4385512431606d867cbbd7aa6c3e9
|
[
"MIT"
] | null | null | null | 916,580 | 916,580 | 0.962198 |
[
[
[
"# For Google Colaboratory\nimport sys, os\nif 'google.colab' in sys.modules:\n # mount google drive\n from google.colab import drive\n drive.mount('/content/gdrive')\n\n file_name = 'Task_4.ipynb'\n import subprocess\n path_to_file = subprocess.check_output('find . -type f -name ' + str(file_name), shell=True).decode(\"utf-8\")\n path_to_file = path_to_file.replace(file_name,\"\").replace('\\n',\"\")\n\n print(path_to_file)\n\n os.chdir(path_to_file)\n !pwd",
"Mounted at /content/gdrive\n./gdrive/MyDrive/URABNA2/\n/content/gdrive/MyDrive/URABNA2\n"
],
[
"pip install osmnx==0.16.2",
"_____no_output_____"
],
[
"!pip uninstall matplotlib\n!pip install matplotlib==3.1.3",
"_____no_output_____"
],
[
"import numpy as np\nimport pandas as pd\nimport osmnx as ox\nfrom shapely.geometry import Polygon\nfrom shapely.geometry import shape\nimport json\nimport os\nimport time\nimport matplotlib.pyplot as plt\nimport geopandas as gpd\nimport math\nfrom collections import defaultdict\n%matplotlib inline\n\npd.set_option('display.max_columns', None)",
"_____no_output_____"
],
[
"def getMinMax(xMax, yMax, xMin, yMin, bFirst, xLong, yLat):\n # return the min, max of x, y coord\n \n if bFirst:\n xMin = xMax = xLong\n yMin = yMax = yLat\n bFirst = False\n else:\n if xLong > xMin:\n xMin = xLong\n elif xLong < xMax:\n xMax = xLong\n\n if yLat < yMin:\n yMin = yLat\n elif yLat > yMax:\n yMax = yLat\n \n return xMax, yMax, xMin, yMin, bFirst",
"_____no_output_____"
]
],
[
[
"# Task 4",
"_____no_output_____"
],
[
"Task 4: - Plot trajectory route onto road network",
"_____no_output_____"
]
],
[
[
"# import results trajectory dataset\ndfMapped = pd.read_csv('./data/matchedMaps.csv')\ndfMapped = pd.DataFrame(dfMapped)",
"_____no_output_____"
],
[
"mapList = []\ncount = 0\nxMax = yMax = 0\nxMin = yMin = 0\nbFirst = True\n\nfor item in dfMapped['mgeom']:\n item = item[11: -1] # remove 1st 2 & last 2 brackets\n map = []\n \n if len(item) <= 2: # []\n print('Skip empty row:', count, ', len:', len(item), ', element:', item)\n else:\n for elem in item.split(','):\n xLong, yLat = elem.split()\n map.append((float(xLong), float(yLat))) # tuple of x, y coords\n \n xMax, yMax, xMin, yMin, bFirst = getMinMax(xMax, yMax, xMin, yMin, bFirst, xLong, yLat)\n \n \n mapList.append(map)\n\n count += 1\n \nprint('xMin:', xMin, ', yMin:', yMin, ', xMax:', xMax, ', yMax:', yMax)",
"Skip empty row: 2 , len: 0 , element: \nSkip empty row: 3 , len: 0 , element: \nSkip empty row: 9 , len: 0 , element: \nSkip empty row: 15 , len: 0 , element: \nSkip empty row: 31 , len: 0 , element: \nSkip empty row: 33 , len: 0 , element: \nSkip empty row: 39 , len: 0 , element: \nSkip empty row: 57 , len: 0 , element: \nSkip empty row: 59 , len: 0 , element: \nSkip empty row: 64 , len: 0 , element: \nSkip empty row: 86 , len: 0 , element: \nSkip empty row: 101 , len: 0 , element: \nSkip empty row: 127 , len: 0 , element: \nSkip empty row: 132 , len: 0 , element: \nSkip empty row: 138 , len: 0 , element: \nSkip empty row: 184 , len: 0 , element: \nSkip empty row: 212 , len: 0 , element: \nSkip empty row: 216 , len: 0 , element: \nSkip empty row: 224 , len: 0 , element: \nSkip empty row: 230 , len: 0 , element: \nSkip empty row: 265 , len: 0 , element: \nSkip empty row: 294 , len: 0 , element: \nSkip empty row: 302 , len: 0 , element: \nSkip empty row: 303 , len: 0 , element: \nSkip empty row: 309 , len: 0 , element: \nSkip empty row: 325 , len: 0 , element: \nSkip empty row: 332 , len: 0 , element: \nSkip empty row: 345 , len: 0 , element: \nSkip empty row: 361 , len: 0 , element: \nSkip empty row: 376 , len: 0 , element: \nSkip empty row: 388 , len: 0 , element: \nSkip empty row: 411 , len: 0 , element: \nSkip empty row: 417 , len: 0 , element: \nSkip empty row: 453 , len: 0 , element: \nSkip empty row: 456 , len: 0 , element: \nSkip empty row: 460 , len: 0 , element: \nSkip empty row: 471 , len: 0 , element: \nSkip empty row: 482 , len: 0 , element: \nSkip empty row: 497 , len: 0 , element: \nSkip empty row: 519 , len: 0 , element: \nSkip empty row: 520 , len: 0 , element: \nSkip empty row: 536 , len: 0 , element: \nSkip empty row: 557 , len: 0 , element: \nSkip empty row: 574 , len: 0 , element: \nSkip empty row: 593 , len: 0 , element: \nSkip empty row: 607 , len: 0 , element: \nSkip empty row: 611 , len: 0 , element: \nSkip empty row: 618 , len: 0 , element: \nSkip empty row: 623 , len: 0 , element: \nSkip empty row: 633 , len: 0 , element: \nSkip empty row: 657 , len: 0 , element: \nSkip empty row: 675 , len: 0 , element: \nSkip empty row: 697 , len: 0 , element: \nSkip empty row: 698 , len: 0 , element: \nSkip empty row: 709 , len: 0 , element: \nSkip empty row: 721 , len: 0 , element: \nSkip empty row: 725 , len: 0 , element: \nSkip empty row: 738 , len: 0 , element: \nSkip empty row: 762 , len: 0 , element: \nSkip empty row: 783 , len: 0 , element: \nSkip empty row: 805 , len: 0 , element: \nSkip empty row: 819 , len: 0 , element: \nSkip empty row: 832 , len: 0 , element: \nSkip empty row: 848 , len: 0 , element: \nSkip empty row: 859 , len: 0 , element: \nSkip empty row: 932 , len: 0 , element: \nSkip empty row: 933 , len: 0 , element: \nSkip empty row: 950 , len: 0 , element: \nSkip empty row: 962 , len: 0 , element: \nSkip empty row: 968 , len: 0 , element: \nSkip empty row: 978 , len: 0 , element: \nSkip empty row: 979 , len: 0 , element: \nSkip empty row: 995 , len: 0 , element: \nSkip empty row: 996 , len: 0 , element: \nxMin: -8.7092708 , yMin: 41.028446 , xMax: -8.4867809 , yMax: 41.26009\n"
],
[
"# extract 10 trajectory from the list of 1000\nlimit = 10\nmapListLen = len(dfMapped)\nmapCoords = []\nxMax = yMax = 0\nxMin = yMin = 0\nbFirst = True\n#print('len:', trajListLen)\n\nfor i in range(limit):\n tmpMap = mapList[i]\n x = []\n y = []\n \n if len(tmpMap) <= 2: # []\n print('Skip empty row:', i, ', len:', len(tmpMap), ', traj:', tmpMap)\n else:\n for tmpCord in tmpMap:\n tmpX = tmpCord[0]\n tmpY = tmpCord[1]\n #print('x:', tmpX, ', y:', tmpY)\n x.append(tmpX)\n y.append(tmpY)\n \n xMax, yMax, xMin, yMin, bFirst = getMinMax(xMax, yMax, xMin, yMin, bFirst, tmpX, tmpY)\n \n mapCoords.append((x, y))\n #print('x:', x)\n #print('y:', y)\n \nprint('xMin:', xMin, ', yMin:', yMin, ', xMax:', xMax, ', yMax:', yMax)",
"xMin: -8.5621277 , yMin: 41.139909 , xMax: -8.6891266 , yMax: 41.180528\n"
],
[
"# sub graph using bounding box method\nstartTime = time.time()\ndelta = 0.01\nbounds = (xMax - delta / 2, xMin + delta / 2, yMin - delta * 0.7, yMax + delta * 0.7)\nx1, x2, y1, y2 = bounds\nboundaryPolygon = Polygon([(x1, y1), (x2, y1), (x2, y2), (x1, y2), (x1, y1)])\nG3 = ox.graph_from_polygon(boundaryPolygon, network_type = 'drive')\n\n# plot all pts\nstartTime = time.time()\n#marker = ['o', '.', ',', 'x', '+', 'v', '^', '<', '>', 's']\nmarker = ['o', 'd', 'p', 'x', '<', 'v', '^', '+', '>', 's']\nfig, ax = ox.plot_graph(G3, figsize = (20, 20), bgcolor = '#FFFFFF', \\\n node_color = '#ACFF97', node_size = 1, \\\n edge_color = '#C4D0FF', show = False, close = False)\n\nfor i in range(len(mapCoords)):\n ax.plot(mapCoords[i][0], mapCoords[i][1], marker[i], ms = 8, label = 'Route_{}'.format(i + 1))\n\nplt.legend(fontsize = 'xx-large')\nplt.tight_layout()\n# plt.savefig('./5mostoften.png', dpi = 320)\nprint(\" Duration %s seconds\" % (time.time() - startTime))",
"/usr/local/lib/python3.7/dist-packages/osmnx/utils_geo.py:281: ShapelyDeprecationWarning: Iteration over multi-part geometries is deprecated and will be removed in Shapely 2.0. Use the `geoms` property to access the constituent parts of a multi-part geometry.\n for polygon in geometry:\n/usr/local/lib/python3.7/dist-packages/osmnx/utils_geo.py:375: ShapelyDeprecationWarning: Iteration over multi-part geometries is deprecated and will be removed in Shapely 2.0. Use the `geoms` property to access the constituent parts of a multi-part geometry.\n for poly in multipoly:\n/usr/local/lib/python3.7/dist-packages/osmnx/utils_geo.py:375: ShapelyDeprecationWarning: Iteration over multi-part geometries is deprecated and will be removed in Shapely 2.0. Use the `geoms` property to access the constituent parts of a multi-part geometry.\n for poly in multipoly:\n"
]
]
] |
[
"code",
"markdown",
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
4ab48ab5eda2c6aa02667ab6ba1dbd9b85d73784
| 16,726 |
ipynb
|
Jupyter Notebook
|
intro-to-pytorch/Part 1 - Tensors in PyTorch (Solution).ipynb
|
Kshitij09/deep-learning-v2-pytorch
|
b214e63b7b560122bc5fd5b26bff6946b5078ba6
|
[
"MIT"
] | null | null | null |
intro-to-pytorch/Part 1 - Tensors in PyTorch (Solution).ipynb
|
Kshitij09/deep-learning-v2-pytorch
|
b214e63b7b560122bc5fd5b26bff6946b5078ba6
|
[
"MIT"
] | null | null | null |
intro-to-pytorch/Part 1 - Tensors in PyTorch (Solution).ipynb
|
Kshitij09/deep-learning-v2-pytorch
|
b214e63b7b560122bc5fd5b26bff6946b5078ba6
|
[
"MIT"
] | null | null | null | 39.17096 | 674 | 0.600622 |
[
[
[
"# Introduction to Deep Learning with PyTorch\n\nIn this notebook, you'll get introduced to [PyTorch](http://pytorch.org/), a framework for building and training neural networks. PyTorch in a lot of ways behaves like the arrays you love from Numpy. These Numpy arrays, after all, are just tensors. PyTorch takes these tensors and makes it simple to move them to GPUs for the faster processing needed when training neural networks. It also provides a module that automatically calculates gradients (for backpropagation!) and another module specifically for building neural networks. All together, PyTorch ends up being more coherent with Python and the Numpy/Scipy stack compared to TensorFlow and other frameworks.\n\n",
"_____no_output_____"
],
[
"## Neural Networks\n\nDeep Learning is based on artificial neural networks which have been around in some form since the late 1950s. The networks are built from individual parts approximating neurons, typically called units or simply \"neurons.\" Each unit has some number of weighted inputs. These weighted inputs are summed together (a linear combination) then passed through an activation function to get the unit's output.\n\n<img src=\"assets/simple_neuron.png\" width=400px>\n\nMathematically this looks like: \n\n$$\n\\begin{align}\ny &= f(w_1 x_1 + w_2 x_2 + b) \\\\\ny &= f\\left(\\sum_i w_i x_i +b \\right)\n\\end{align}\n$$\n\nWith vectors this is the dot/inner product of two vectors:\n\n$$\nh = \\begin{bmatrix}\nx_1 \\, x_2 \\cdots x_n\n\\end{bmatrix}\n\\cdot \n\\begin{bmatrix}\n w_1 \\\\\n w_2 \\\\\n \\vdots \\\\\n w_n\n\\end{bmatrix}\n$$",
"_____no_output_____"
],
[
"## Tensors\n\nIt turns out neural network computations are just a bunch of linear algebra operations on *tensors*, a generalization of matrices. A vector is a 1-dimensional tensor, a matrix is a 2-dimensional tensor, an array with three indices is a 3-dimensional tensor (RGB color images for example). The fundamental data structure for neural networks are tensors and PyTorch (as well as pretty much every other deep learning framework) is built around tensors.\n\n<img src=\"assets/tensor_examples.svg\" width=600px>\n\nWith the basics covered, it's time to explore how we can use PyTorch to build a simple neural network.",
"_____no_output_____"
]
],
[
[
"# First, import PyTorch\nimport torch",
"_____no_output_____"
],
[
"def activation(x):\n \"\"\" Sigmoid activation function \n \n Arguments\n ---------\n x: torch.Tensor\n \"\"\"\n return 1/(1+torch.exp(-x))",
"_____no_output_____"
],
[
"### Generate some data\ntorch.manual_seed(7) # Set the random seed so things are predictable\n\n# Features are 5 random normal variables\nfeatures = torch.randn((1, 5))\n# True weights for our data, random normal variables again\nweights = torch.randn_like(features)\n# and a true bias term\nbias = torch.randn((1, 1))",
"_____no_output_____"
]
],
[
[
"Above I generated data we can use to get the output of our simple network. This is all just random for now, going forward we'll start using normal data. Going through each relevant line:\n\n`features = torch.randn((1, 5))` creates a tensor with shape `(1, 5)`, one row and five columns, that contains values randomly distributed according to the normal distribution with a mean of zero and standard deviation of one. \n\n`weights = torch.randn_like(features)` creates another tensor with the same shape as `features`, again containing values from a normal distribution.\n\nFinally, `bias = torch.randn((1, 1))` creates a single value from a normal distribution.\n\nPyTorch tensors can be added, multiplied, subtracted, etc, just like Numpy arrays. In general, you'll use PyTorch tensors pretty much the same way you'd use Numpy arrays. They come with some nice benefits though such as GPU acceleration which we'll get to later. For now, use the generated data to calculate the output of this simple single layer network. \n> **Exercise**: Calculate the output of the network with input features `features`, weights `weights`, and bias `bias`. Similar to Numpy, PyTorch has a [`torch.sum()`](https://pytorch.org/docs/stable/torch.html#torch.sum) function, as well as a `.sum()` method on tensors, for taking sums. Use the function `activation` defined above as the activation function.",
"_____no_output_____"
]
],
[
[
"### Solution\n\n# Now, make our labels from our data and true weights\n\ny = activation(torch.sum(features * weights) + bias)\ny = activation((features * weights).sum() + bias)",
"_____no_output_____"
]
],
[
[
"You can do the multiplication and sum in the same operation using a matrix multiplication. In general, you'll want to use matrix multiplications since they are more efficient and accelerated using modern libraries and high-performance computing on GPUs.\n\nHere, we want to do a matrix multiplication of the features and the weights. For this we can use [`torch.mm()`](https://pytorch.org/docs/stable/torch.html#torch.mm) or [`torch.matmul()`](https://pytorch.org/docs/stable/torch.html#torch.matmul) which is somewhat more complicated and supports broadcasting. If we try to do it with `features` and `weights` as they are, we'll get an error\n\n```python\n>> torch.mm(features, weights)\n\n---------------------------------------------------------------------------\nRuntimeError Traceback (most recent call last)\n<ipython-input-13-15d592eb5279> in <module>()\n----> 1 torch.mm(features, weights)\n\nRuntimeError: size mismatch, m1: [1 x 5], m2: [1 x 5] at /Users/soumith/minicondabuild3/conda-bld/pytorch_1524590658547/work/aten/src/TH/generic/THTensorMath.c:2033\n```\n\nAs you're building neural networks in any framework, you'll see this often. Really often. What's happening here is our tensors aren't the correct shapes to perform a matrix multiplication. Remember that for matrix multiplications, the number of columns in the first tensor must equal to the number of rows in the second column. Both `features` and `weights` have the same shape, `(1, 5)`. This means we need to change the shape of `weights` to get the matrix multiplication to work.\n\n**Note:** To see the shape of a tensor called `tensor`, use `tensor.shape`. If you're building neural networks, you'll be using this method often.\n\nThere are a few options here: [`weights.reshape()`](https://pytorch.org/docs/stable/tensors.html#torch.Tensor.reshape), [`weights.resize_()`](https://pytorch.org/docs/stable/tensors.html#torch.Tensor.resize_), and [`weights.view()`](https://pytorch.org/docs/stable/tensors.html#torch.Tensor.view).\n\n* `weights.reshape(a, b)` will return a new tensor with the same data as `weights` with size `(a, b)` sometimes, and sometimes a clone, as in it copies the data to another part of memory.\n* `weights.resize_(a, b)` returns the same tensor with a different shape. However, if the new shape results in fewer elements than the original tensor, some elements will be removed from the tensor (but not from memory). If the new shape results in more elements than the original tensor, new elements will be uninitialized in memory. Here I should note that the underscore at the end of the method denotes that this method is performed **in-place**. Here is a great forum thread to [read more about in-place operations](https://discuss.pytorch.org/t/what-is-in-place-operation/16244) in PyTorch.\n* `weights.view(a, b)` will return a new tensor with the same data as `weights` with size `(a, b)`.\n\nI usually use `.view()`, but any of the three methods will work for this. So, now we can reshape `weights` to have five rows and one column with something like `weights.view(5, 1)`.\n\n> **Exercise**: Calculate the output of our little network using matrix multiplication.",
"_____no_output_____"
]
],
[
[
"## Solution\n\ny = activation(torch.mm(features, weights.view(5,1)) + bias)",
"_____no_output_____"
]
],
[
[
"### Stack them up!\n\nThat's how you can calculate the output for a single neuron. The real power of this algorithm happens when you start stacking these individual units into layers and stacks of layers, into a network of neurons. The output of one layer of neurons becomes the input for the next layer. With multiple input units and output units, we now need to express the weights as a matrix.\n\n<img src='assets/multilayer_diagram_weights.png' width=450px>\n\nThe first layer shown on the bottom here are the inputs, understandably called the **input layer**. The middle layer is called the **hidden layer**, and the final layer (on the right) is the **output layer**. We can express this network mathematically with matrices again and use matrix multiplication to get linear combinations for each unit in one operation. For example, the hidden layer ($h_1$ and $h_2$ here) can be calculated \n\n$$\n\\vec{h} = [h_1 \\, h_2] = \n\\begin{bmatrix}\nx_1 \\, x_2 \\cdots \\, x_n\n\\end{bmatrix}\n\\cdot \n\\begin{bmatrix}\n w_{11} & w_{12} \\\\\n w_{21} &w_{22} \\\\\n \\vdots &\\vdots \\\\\n w_{n1} &w_{n2}\n\\end{bmatrix}\n$$\n\nThe output for this small network is found by treating the hidden layer as inputs for the output unit. The network output is expressed simply\n\n$$\ny = f_2 \\! \\left(\\, f_1 \\! \\left(\\vec{x} \\, \\mathbf{W_1}\\right) \\mathbf{W_2} \\right)\n$$",
"_____no_output_____"
]
],
[
[
"### Generate some data\ntorch.manual_seed(7) # Set the random seed so things are predictable\n\n# Features are 3 random normal variables\nfeatures = torch.randn((1, 3))\n\n# Define the size of each layer in our network\nn_input = features.shape[1] # Number of input units, must match number of input features\nn_hidden = 2 # Number of hidden units \nn_output = 1 # Number of output units\n\n# Weights for inputs to hidden layer\nW1 = torch.randn(n_input, n_hidden)\n# Weights for hidden layer to output layer\nW2 = torch.randn(n_hidden, n_output)\n\n# and bias terms for hidden and output layers\nB1 = torch.randn((1, n_hidden))\nB2 = torch.randn((1, n_output))",
"_____no_output_____"
]
],
[
[
"> **Exercise:** Calculate the output for this multi-layer network using the weights `W1` & `W2`, and the biases, `B1` & `B2`. ",
"_____no_output_____"
]
],
[
[
"### Solution\n\nh = activation(torch.mm(features, W1) + B1)\noutput = activation(torch.mm(h, W2) + B2)\nprint(output)",
"tensor([[ 0.3171]])\n"
]
],
[
[
"If you did this correctly, you should see the output `tensor([[ 0.3171]])`.\n\nThe number of hidden units a parameter of the network, often called a **hyperparameter** to differentiate it from the weights and biases parameters. As you'll see later when we discuss training a neural network, the more hidden units a network has, and the more layers, the better able it is to learn from data and make accurate predictions.",
"_____no_output_____"
],
[
"## Numpy to Torch and back\n\nSpecial bonus section! PyTorch has a great feature for converting between Numpy arrays and Torch tensors. To create a tensor from a Numpy array, use `torch.from_numpy()`. To convert a tensor to a Numpy array, use the `.numpy()` method.",
"_____no_output_____"
]
],
[
[
"import numpy as np\na = np.random.rand(4,3)\na",
"_____no_output_____"
],
[
"b = torch.from_numpy(a)\nb",
"_____no_output_____"
],
[
"b.numpy()",
"_____no_output_____"
]
],
[
[
"The memory is shared between the Numpy array and Torch tensor, so if you change the values in-place of one object, the other will change as well.",
"_____no_output_____"
]
],
[
[
"# Multiply PyTorch Tensor by 2, in place\nb.mul_(2)",
"_____no_output_____"
],
[
"# Numpy array matches new values from Tensor\na",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4ab4999aea231d893f87574db07026c098bf652d
| 9,478 |
ipynb
|
Jupyter Notebook
|
exercise-grouping-and-sorting.ipynb
|
Tan7im/Data-Science-Cohort
|
98e7ee5309c07482b125b0140aee672aa3a6694c
|
[
"MIT"
] | null | null | null |
exercise-grouping-and-sorting.ipynb
|
Tan7im/Data-Science-Cohort
|
98e7ee5309c07482b125b0140aee672aa3a6694c
|
[
"MIT"
] | null | null | null |
exercise-grouping-and-sorting.ipynb
|
Tan7im/Data-Science-Cohort
|
98e7ee5309c07482b125b0140aee672aa3a6694c
|
[
"MIT"
] | null | null | null | 9,478 | 9,478 | 0.732433 |
[
[
[
"**This notebook is an exercise in the [Pandas](https://www.kaggle.com/learn/pandas) course. You can reference the tutorial at [this link](https://www.kaggle.com/residentmario/grouping-and-sorting).**\n\n---\n",
"_____no_output_____"
],
[
"# Introduction\n\nIn these exercises we'll apply groupwise analysis to our dataset.\n\nRun the code cell below to load the data before running the exercises.",
"_____no_output_____"
]
],
[
[
"import pandas as pd\n\nreviews = pd.read_csv(\"../input/wine-reviews/winemag-data-130k-v2.csv\", index_col=0)\n#pd.set_option(\"display.max_rows\", 5)\n\nfrom learntools.core import binder; binder.bind(globals())\nfrom learntools.pandas.grouping_and_sorting import *\nprint(\"Setup complete.\")\nreviews.head()",
"_____no_output_____"
]
],
[
[
"# Exercises",
"_____no_output_____"
],
[
"## 1.\nWho are the most common wine reviewers in the dataset? Create a `Series` whose index is the `taster_twitter_handle` category from the dataset, and whose values count how many reviews each person wrote.",
"_____no_output_____"
]
],
[
[
"# Your code here\nreviews_written = reviews.groupby('taster_twitter_handle')['taster_twitter_handle'].size()\n\n# Check your answer\nq1.check()",
"_____no_output_____"
],
[
"q1.hint()\n#q1.solution()",
"_____no_output_____"
]
],
[
[
"## 2.\nWhat is the best wine I can buy for a given amount of money? Create a `Series` whose index is wine prices and whose values is the maximum number of points a wine costing that much was given in a review. Sort the values by price, ascending (so that `4.0` dollars is at the top and `3300.0` dollars is at the bottom).",
"_____no_output_____"
]
],
[
[
"#max_points = reviews.groupby(['price','points']).max()\nbest_rating_per_price = reviews.groupby('price')['points'].max().sort_index()\n\n# Check your answer\nq2.check()",
"_____no_output_____"
],
[
"q2.hint()\nq2.solution()",
"_____no_output_____"
]
],
[
[
"## 3.\nWhat are the minimum and maximum prices for each `variety` of wine? Create a `DataFrame` whose index is the `variety` category from the dataset and whose values are the `min` and `max` values thereof.",
"_____no_output_____"
]
],
[
[
"price_extremes = reviews.groupby('variety').price.agg([min,max])\n\n\n# Check your answer\nq3.check()",
"_____no_output_____"
],
[
"#q3.hint()\n#q3.solution()",
"_____no_output_____"
]
],
[
[
"## 4.\nWhat are the most expensive wine varieties? Create a variable `sorted_varieties` containing a copy of the dataframe from the previous question where varieties are sorted in descending order based on minimum price, then on maximum price (to break ties).",
"_____no_output_____"
]
],
[
[
"sorted_varieties = price_extremes.sort_values(by=['min','max'], ascending = False)\n\n# Check your answer\nq4.check()",
"_____no_output_____"
],
[
"q4.hint()\nq4.solution()",
"_____no_output_____"
]
],
[
[
"## 5.\nCreate a `Series` whose index is reviewers and whose values is the average review score given out by that reviewer. Hint: you will need the `taster_name` and `points` columns.",
"_____no_output_____"
]
],
[
[
"reviewer_mean_ratings = reviews.groupby('taster_name')['points'].mean()\n\n# Check your answer\nq5.check()",
"_____no_output_____"
],
[
"#q5.hint()\n#q5.solution()",
"_____no_output_____"
]
],
[
[
"Are there significant differences in the average scores assigned by the various reviewers? Run the cell below to use the `describe()` method to see a summary of the range of values.",
"_____no_output_____"
]
],
[
[
"reviewer_mean_ratings.describe()",
"_____no_output_____"
]
],
[
[
"## 6.\nWhat combination of countries and varieties are most common? Create a `Series` whose index is a `MultiIndex`of `{country, variety}` pairs. For example, a pinot noir produced in the US should map to `{\"US\", \"Pinot Noir\"}`. Sort the values in the `Series` in descending order based on wine count.",
"_____no_output_____"
]
],
[
[
"country_variety_counts = reviews.groupby(['country', 'variety']).size().sort_values(ascending = False)\n\n# Check your answer\nq6.check()",
"_____no_output_____"
],
[
"q6.hint()\nq6.solution()",
"_____no_output_____"
]
],
[
[
"# Keep going\n\nMove on to the [**data types and missing data**](https://www.kaggle.com/residentmario/data-types-and-missing-values).",
"_____no_output_____"
],
[
"---\n\n\n\n\n*Have questions or comments? Visit the [course discussion forum](https://www.kaggle.com/learn/pandas/discussion) to chat with other learners.*",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
]
] |
4ab49db53576bdaa305ad7fa4a8550f8c305df21
| 27,246 |
ipynb
|
Jupyter Notebook
|
notebooks/CLTK data types.ipynb
|
yelircaasi/cltk
|
1583aa24682543a1f33434a21918f039ca27d60c
|
[
"MIT"
] | null | null | null |
notebooks/CLTK data types.ipynb
|
yelircaasi/cltk
|
1583aa24682543a1f33434a21918f039ca27d60c
|
[
"MIT"
] | null | null | null |
notebooks/CLTK data types.ipynb
|
yelircaasi/cltk
|
1583aa24682543a1f33434a21918f039ca27d60c
|
[
"MIT"
] | null | null | null | 29.140107 | 409 | 0.505616 |
[
[
[
"1. [`Language`](#language)\n1. [`Doc`](#doc)\n1. [`Process`](#process)\n1. [`Pipeline`](#pipeline)\n1. [`MorphosyntacticFeature`](#morpho)\n1. [`MorphosyntacticFeatureBundle`](#morpho-bundle)\n1. [`Form`](#form)\n1. [`DecisionTree`](#dt)",
"_____no_output_____"
],
[
"# `Language` <a name=\"language\"></a>\n\n`Language` are used to identify each language and keep track of their attributes (lat-long, 3-letter ISO code, etc.). Users do not normally need to create these themselves, though this type could be extended to distinguish dialects.",
"_____no_output_____"
]
],
[
[
"from cltk.core.data_types import Language",
"_____no_output_____"
],
[
"Language(\n name=\"Classical Mongolian\",\n glottolog_id=\"\",\n latitude=0.0,\n longitude=0.0,\n dates=[],\n family_id=\"\",\n parent_id=\"\",\n level=\"\",\n iso_639_3_code=\"cmg\",\n type=\"h\",\n)",
"_____no_output_____"
],
[
"from cltk.languages.glottolog import LANGUAGES",
"_____no_output_____"
],
[
"print(LANGUAGES[\"gmh\"])",
"Language(name='Middle High German', glottolog_id='midd1343', latitude=0.0, longitude=0.0, dates=[], family_id='indo1319', parent_id='midd1349', level='language', iso_639_3_code='gmh', type='h')\n"
]
],
[
[
"The following records all pre-modern languages as identified by the [Glottolog project](http://glottolog.org).",
"_____no_output_____"
]
],
[
[
"from pprint import pprint\npprint([(l.name, l.iso_639_3_code) for _, l in LANGUAGES.items()])",
"[('Aequian', 'xae'),\n ('Aghwan', 'xag'),\n ('Akkadian', 'akk'),\n ('Alanic', 'xln'),\n ('Ancient Greek', 'grc'),\n ('Ancient Hebrew', 'hbo'),\n ('Ancient Ligurian', 'xlg'),\n ('Ancient Macedonian', 'xmk'),\n ('Ancient North Arabian', 'xna'),\n ('Ancient Zapotec', 'xzp'),\n ('Andalusian Arabic', 'xaa'),\n ('Anglo-Norman', 'xno'),\n ('Aquitanian', 'xaq'),\n ('Ardhamāgadhī Prākrit', 'pka'),\n ('Armazic', 'xrm'),\n ('Avestan', 'ave'),\n ('Bactrian', 'xbc'),\n ('Bengali', 'ben'),\n ('Bolgarian', 'xbo'),\n ('Burma Pyu', 'pyx'),\n ('Camunic', 'xcc'),\n ('Carian', 'xcr'),\n ('Celtiberian', 'xce'),\n ('Church Slavic', 'chu'),\n ('Cisalpine Gaulish', 'xcg'),\n ('Classical Armenian', 'xcl'),\n ('Classical Mandaic', 'myz'),\n ('Classical Mongolian', 'cmg'),\n ('Classical Nahuatl', 'nci'),\n ('Classical Newari', 'nwc'),\n ('Classical Quechua', 'qwc'),\n ('Classical Syriac', 'syc'),\n ('Classical Tibetan', 'xct'),\n ('Coptic', 'cop'),\n ('Cumbric', 'xcb'),\n ('Cuneiform Luwian', 'xlu'),\n ('Curonian', 'xcu'),\n ('Dacian', 'xdc'),\n ('Early Irish', 'sga'),\n ('Early Tripuri', 'xtr'),\n ('Eastern Panjabi', 'pan'),\n ('Eblaite', 'xeb'),\n ('Edomite', 'xdm'),\n ('Egyptian (Ancient)', 'egy'),\n ('Elamite', 'elx'),\n ('Elymian', 'xly'),\n ('Epi-Olmec', 'xep'),\n ('Epigraphic Mayan', 'emy'),\n ('Eteocretan', 'ecr'),\n ('Eteocypriot', 'ecy'),\n ('Etruscan', 'ett'),\n ('Faliscan', 'xfa'),\n ('Galatian', 'xga'),\n ('Galindan', 'xgl'),\n ('Geez', 'gez'),\n ('Gothic', 'got'),\n ('Gujarati', 'guj'),\n ('Gāndhārī', 'pgd'),\n ('Hadrami', 'xhd'),\n ('Harami', 'xha'),\n ('Harappan', 'xiv'),\n ('Hattic', 'xht'),\n ('Hernican', 'xhr'),\n ('Hiberno-Scottish Gaelic', 'ghc'),\n ('Hieroglyphic Luwian', 'hlu'),\n ('Hindi', 'hin'),\n ('Hittite', 'hit'),\n ('Hunnic', 'xhc'),\n ('Hurrian', 'xhu'),\n ('Iberian', 'xib'),\n ('Illyrian', 'xil'),\n ('Jutish', 'jut'),\n ('Kajkavian', 'kjv'),\n ('Kannada', 'kan'),\n ('Kara (Korea)', 'zra'),\n ('Karakhanid', 'xqa'),\n ('Kaskean', 'zsk'),\n ('Kawi', 'kaw'),\n ('Khazar', 'zkz'),\n ('Khorezmian', 'zkh'),\n ('Khotanese', 'kho'),\n ('Khwarezmian', 'xco'),\n ('Kitan', 'zkt'),\n ('Koguryo', 'zkg'),\n ('Langobardic', 'lng'),\n ('Latin', 'lat'),\n ('Lemnian', 'xle'),\n ('Lepontic', 'xlp'),\n ('Liburnian', 'xli'),\n ('Linear A', 'lab'),\n ('Literary Chinese', 'lzh'),\n ('Lusitanian', 'xls'),\n ('Lycian A', 'xlc'),\n ('Lydian', 'xld'),\n ('Maek', 'hmk'),\n ('Maharastri Prakrit', 'pmh'),\n ('Malayalam', 'mal'),\n ('Manichaean Middle Persian', 'xmn'),\n ('Marrucinian', 'umc'),\n ('Marsian', 'ims'),\n ('Median', 'xme'),\n ('Meroitic', 'xmr'),\n ('Messapic', 'cms'),\n ('Middle Armenian', 'axm'),\n ('Middle Breton', 'xbm'),\n ('Middle Chinese', 'ltc'),\n ('Middle Cornish', 'cnx'),\n ('Middle Dutch', 'dum'),\n ('Middle English', 'enm'),\n ('Middle French', 'frm'),\n ('Middle High German', 'gmh'),\n ('Middle Hittite', 'htx'),\n ('Middle Irish (10-12th century)', 'mga'),\n ('Middle Korean (10th-16th cent.)', 'okm'),\n ('Middle Low German', 'gml'),\n ('Middle Mongol', 'xng'),\n ('Middle Newar', 'nwx'),\n ('Middle Welsh', 'wlm'),\n ('Milyan', 'imy'),\n ('Minaean', 'inm'),\n ('Minoan', 'omn'),\n ('Moabite', 'obm'),\n ('Mozarabic', 'mxi'),\n ('Mycenaean Greek', 'gmy'),\n ('Mysian', 'yms'),\n ('Nadruvian', 'ndf'),\n ('Neo-Hittite', 'nei'),\n ('Noric', 'nrc'),\n ('North Picene', 'nrp'),\n ('Numidian', 'nxm'),\n ('Odia', 'ory'),\n ('Official Aramaic (700-300 BCE)', 'arc'),\n ('Old Aramaic (up to 700 BCE)', 'oar'),\n ('Old Avar', 'oav'),\n ('Old Breton', 'obt'),\n ('Old Burmese', 'obr'),\n ('Old Chinese', 'och'),\n ('Old Cornish', 'oco'),\n ('Old Dutch-Old Frankish', 'odt'),\n ('Old English (ca. 450-1100)', 'ang'),\n ('Old Frankish', 'frk'),\n ('Old French (842-ca. 1400)', 'fro'),\n ('Old Frisian', 'ofs'),\n ('Old Georgian', 'oge'),\n ('Old High German (ca. 750-1050)', 'goh'),\n ('Old Hittite', 'oht'),\n ('Old Hungarian', 'ohu'),\n ('Old Japanese', 'ojp'),\n ('Old Korean (3rd-9th cent.)', 'oko'),\n ('Old Lithuanian', 'olt'),\n ('Old Manipuri', 'omp'),\n ('Old Marathi', 'omr'),\n ('Old Mon', 'omx'),\n ('Old Norse', 'non'),\n ('Old Nubian', 'onw'),\n ('Old Ossetic', 'oos'),\n ('Old Persian (ca. 600-400 B.C.)', 'peo'),\n ('Old Provençal', 'pro'),\n ('Old Russian', 'orv'),\n ('Old Saxon', 'osx'),\n ('Old Spanish', 'osp'),\n ('Old Tamil', 'oty'),\n ('Old Tibetan', 'otb'),\n ('Old Turkic', 'oui'),\n ('Old Turkish', 'otk'),\n ('Old-Middle Welsh', 'owl'),\n ('Oscan', 'osc'),\n ('Ottoman Turkish (1500-1928)', 'ota'),\n ('Paekche', 'pkc'),\n ('Paelignian', 'pgn'),\n ('Pahlavi', 'pal'),\n ('Palaic', 'plq'),\n ('Palestinian Jewish Aramaic', 'jpa'),\n ('Pali', 'pli'),\n ('Parthian', 'xpr'),\n ('Pecheneg', 'xpc'),\n ('Phoenician', 'phn'),\n ('Phrygian', 'xpg'),\n ('Pictish', 'xpi'),\n ('Pisidian', 'xps'),\n ('Primitive Irish', 'pgl'),\n ('Punic', 'xpu'),\n ('Puyo', 'xpy'),\n ('Puyo-Paekche', 'xpp'),\n ('Qatabanian', 'xqt'),\n ('Raetic', 'xrr'),\n ('Sabaic', 'xsa'),\n ('Sabine', 'sbv'),\n ('Sanskrit', 'san'),\n ('Sauraseni Prakrit', 'psu'),\n ('Scythian', 'xsc'),\n ('Sicana', 'sxc'),\n ('Sicula', 'scx'),\n ('Siculo Arabic', 'sqr'),\n ('Sidetic', 'xsd'),\n ('Skalvian', 'svx'),\n ('Sogdian', 'sog'),\n ('Sorothaptic', 'sxo'),\n ('South Picene', 'spx'),\n ('Standard Arabic', 'arb'),\n ('Sumerian', 'sux'),\n ('Tangut', 'txg'),\n ('Tartessian', 'txr'),\n ('Telugu', 'tel'),\n ('Thracian', 'txh'),\n ('Tokharian A', 'xto'),\n ('Tokharian B', 'txb'),\n ('Transalpine Gaulish', 'xtg'),\n ('Tumshuqese', 'xtq'),\n ('Ugaritic', 'uga'),\n ('Umbrian', 'xum'),\n ('Urartian', 'xur'),\n ('Urdu', 'urd'),\n ('Vandalic', 'xvn'),\n ('Venetic', 'xve'),\n ('Vestinian', 'xvs'),\n ('Volscian', 'xvo'),\n ('Western Farsi', 'pes'),\n ('Zhangzhung', 'xzh')]\n"
]
],
[
[
"# `Word` <a name=\"word\"></a>\n\n`Word` is one of the most important objects. It contains all token-level information generated by a `Process`.",
"_____no_output_____"
]
],
[
[
"from cltk.core.data_types import Word",
"_____no_output_____"
],
[
"Word(index_char_start=0, index_char_stop=6, index_token=0, string=\"Gallia\", pos=\"nom\")",
"_____no_output_____"
]
],
[
[
"# `Doc` <a name=\"doc\"></a>\n\n`Doc` is the other of the two most important types. It too is build up each time a `Process` runs.\n\nToken-level information is stored at `Doc.words`, while larger units of information (e.g., an original input string) are kept elsewhere here.",
"_____no_output_____"
]
],
[
[
"from cltk.core.data_types import Doc",
"_____no_output_____"
],
[
"Doc(raw=\"Gallia est omnis divisa in partes tres\")",
"_____no_output_____"
]
],
[
[
"It contains a number of helper methods, too, which read (usually) from `.word` and return convenient data structures.",
"_____no_output_____"
]
],
[
[
"from cltk import NLP\ncltk_nlp = NLP(language=\"lat\")\ncltk_doc = cltk_nlp.analyze(text=\"Gallia est omnis divisa in partes tres\")",
"𐤀 CLTK version '1.0.9'.\nPipeline for language 'Latin' (ISO: 'lat'): `LatinNormalizeProcess`, `LatinStanzaProcess`, `LatinEmbeddingsProcess`, `StopsProcess`, `LatinNERProcess`, `LatinLexiconProcess`.\n"
],
[
"cltk_doc.tokens # List[str]",
"_____no_output_____"
]
],
[
[
"# `Process` <a name=\"process\"></a>\n\nA `Process` is a Python `class` that wraps a particular algorithm type for a particular language (e.g., Sanskrit tokenization). It is designed to invoked by the `Pipeline`, though a user may call it directly, too.\n\nSee notebook [Make custom Process and add to Pipeline](https://github.com/cltk/cltk/blob/dev/notebooks/Make%20custom%20Process%20and%20add%20to%20Pipeline.ipynb) for a demonstration.",
"_____no_output_____"
]
],
[
[
"from dataclasses import dataclass, field\nfrom typing import List, Type\nfrom boltons.cacheutils import cachedproperty\nfrom cltk.core.data_types import Process",
"_____no_output_____"
],
[
"@dataclass\nclass StopsProcess(Process):\n \"\"\"\n\n >>> from cltk.core.data_types import Doc, Word\n >>> from cltk.stops.processes import StopsProcess\n >>> from cltk.languages.example_texts import get_example_text\n >>> lang = \"lat\"\n >>> words = [Word(string=token) for token in split_punct_ws(get_example_text(lang))]\n >>> stops_process = StopsProcess(language=lang)\n >>> output_doc = stops_process.run(Doc(raw=get_example_text(lang), words=words))\n >>> output_doc.words[1].string\n 'est'\n >>> output_doc.words[1].stop\n True\n \"\"\"\n\n @cachedproperty\n def algorithm(self):\n return Stops(iso_code=self.language).get_stopwords()\n\n def run(self, input_doc: Doc) -> Doc:\n \"\"\"Note this marks a word a stop if there is a match on\n either the inflected form (``Word.string``) or the\n lemma (``Word.lemma``).\n \"\"\"\n output_doc = deepcopy(input_doc)\n stops_list = self.algorithm\n\n for index, word_obj in enumerate(output_doc.words):\n if (word_obj.string in stops_list) or (word_obj.lemma in stops_list):\n word_obj.stop = True\n else:\n word_obj.stop = False\n output_doc.words[index] = word_obj\n\n return output_doc",
"_____no_output_____"
]
],
[
[
"# `Pipeline` <a name=\"pipeline\"></a>\n\nA `Pipeline` is a list containing the algorithms, in order, as they are to be invoked by `NLP()`.\n\nSee notebook [Make custom Process and add to Pipeline](https://github.com/cltk/cltk/blob/dev/notebooks/Make%20custom%20Process%20and%20add%20to%20Pipeline.ipynb) for a demonstration.",
"_____no_output_____"
]
],
[
[
"from cltk.core.data_types import Pipeline\nfrom cltk.languages.utils import get_lang",
"_____no_output_____"
],
[
"@dataclass\nclass AkkadianPipeline(Pipeline):\n \"\"\"Default ``Pipeline`` for Akkadian.\n\n >>> from cltk.languages.pipelines import AkkadianPipeline\n >>> a_pipeline = AkkadianPipeline()\n >>> a_pipeline.description\n 'Pipeline for the Akkadian language.'\n >>> a_pipeline.language\n Language(name='Akkadian', glottolog_id='akka1240', latitude=33.1, longitude=44.1, dates=[], family_id='afro1255', parent_id='east2678', level='language', iso_639_3_code='akk', type='a')\n >>> a_pipeline.language.name\n 'Akkadian'\n >>> a_pipeline.processes[0]\n <class 'cltk.tokenizers.processes.AkkadianTokenizationProcess'>\n \"\"\"\n\n description: str = \"Pipeline for the Akkadian language.\"\n language: Language = get_lang(\"akk\")\n processes: List[Type[Process]] = field(\n default_factory=lambda: [AkkadianTokenizationProcess, StopsProcess]\n )",
"_____no_output_____"
]
],
[
[
"# `MorphosyntacticFeature` <a name=\"morpho\"></a>\n\nThis model inherits from the Python builtin `IntEnum`. A `MorphosyntacticFeature` (at [cltk/morphology/universal_dependencies_features.py]()) has been made for each [morphsyntactic tag defined by v2 of the Universal Dependencies project](https://universaldependencies.org/u/feat/all.html).",
"_____no_output_____"
]
],
[
[
"from enum import auto\nfrom cltk.morphology.universal_dependencies_features import Case, Gender, MorphosyntacticFeature, Number",
"_____no_output_____"
],
[
"# 'hercule' ('by Hercules, assuredly, indeed')\n# http://www.perseus.tufts.edu/hopper/text?doc=Perseus:text:1999.04.0060:entry=hercule\ngender = Gender.masculine\nprint(\"Gender:\", gender)\ncase = Case.vocative\nprint(\"Case:\", case)\nnumber = Number.singular\nprint(\"Number\", number)",
"Gender: masculine\nCase: vocative\nNumber singular\n"
],
[
"type(case)",
"_____no_output_____"
],
[
"print(\"`MorphosyntacticFeature.name`:\", case.name)\nprint(\"`MorphosyntacticFeature.value`\", case.value)",
"`MorphosyntacticFeature.name`: vocative\n`MorphosyntacticFeature.value` 17\n"
],
[
"# Example declaration\nclass Case(MorphosyntacticFeature):\n \"\"\"The case of a noun phrase.\n see https://universaldependencies.org/u/feat/Case.html\n \"\"\"\n\n # structural cases\n nominative = auto()\n accusative = auto()\n ergative = auto()\n absolutive = auto()\n\n # oblique cases\n abessive = auto()\n befefactive = auto()\n causative = auto()\n comparative = auto()\n considerative = auto()\n comitative = auto()\n dative = auto()\n distributive = auto()\n equative = auto()\n genitive = auto()\n instrumental = auto()\n partitive = auto()\n vocative = auto()\n\n # spatiotemporal cases\n ablative = auto()\n additive = auto()\n adessive = auto()\n allative = auto()\n delative = auto()\n elative = auto()\n essive = auto()\n illative = auto()\n inessive = auto()\n lative = auto()\n locative = auto()\n perlative = auto()\n sublative = auto()\n superessive = auto()\n terminative = auto()\n temporal = auto()\n translative = auto()",
"_____no_output_____"
],
[
"# Users can learn a bit about these features, too\n#help(case)",
"_____no_output_____"
]
],
[
[
"Note there is a distinction between POS and a word's features. POS tags are more general categories for general classes of words, like noun, verb, etc.",
"_____no_output_____"
]
],
[
[
"from cltk.morphology.universal_dependencies_features import POS",
"_____no_output_____"
],
[
"# so the Latin word 'hercule' would be\npos = POS.interjection\nprint(pos)",
"interjection\n"
]
],
[
[
"# `MorphosyntacticFeatureBundle` <a name=\"morpho-bundle\"></a>\n\n`MorphosyntacticFeature`s are brought together to describe a word with `MorphosyntacticFeatureBundle`.",
"_____no_output_____"
]
],
[
[
"from cltk.morphology.morphosyntax import MorphosyntacticFeatureBundle",
"_____no_output_____"
],
[
"bundle = MorphosyntacticFeatureBundle(case, gender, number)",
"_____no_output_____"
],
[
"print(type(bundle))",
"<class 'cltk.morphology.morphosyntax.MorphosyntacticFeatureBundle'>\n"
],
[
"print(bundle)",
"{Case: [vocative], Gender: [masculine], Number: [singular]}\n"
],
[
"# How to access features within a bundle\nprint(\"Gender\", bundle[\"Gender\"])\nprint(\"Case\", bundle[\"Case\"])\nprint(\"Number\", bundle[\"Number\"])",
"Gender [masculine]\nCase [vocative]\nNumber [singular]\n"
]
],
[
[
"`POS` and `MorphosyntacticFeatureBundle` are intended to be placed within a `Word`instance.",
"_____no_output_____"
]
],
[
[
"Word(string=\"hercule\", pos=pos, features=bundle)",
"_____no_output_____"
]
],
[
[
"# `Form` <a name=\"form\"></a>\n\nSince the `Doc.words` is a flat list of `Word` objects, it can be difficult to model hierearchical data, such as dependency trees. To help such modeling, the `Form` and `DependencyTree` types were created.\n\n`Form` inherits from the builtin `xml` package's `Element` and `DependencyTree` from `ElementTree`.\n\nSee [Modeling syntax with Form and DependencyTree](https://github.com/cltk/cltk/blob/dev/notebooks/CLTK%20Demonstration.ipynb) for a full example.",
"_____no_output_____"
]
],
[
[
"from cltk.dependency.tree import Form",
"_____no_output_____"
]
],
[
[
"# `DecisionTree` <a name=\"dt\"></a>\n\nSee [Modeling syntax with Form and DependencyTree](https://github.com/cltk/cltk/blob/dev/notebooks/CLTK%20Demonstration.ipynb) for a full example.",
"_____no_output_____"
]
],
[
[
"from cltk.dependency.tree import DependencyTree",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ab4a5ff98cb83c7d3499f46e4721f86362a2044
| 47,826 |
ipynb
|
Jupyter Notebook
|
pipeline/complete_analysis/eQTL_analysis.ipynb
|
floutt/xqtl-pipeline
|
5ee370b6ff783166f0715d0b8c7c66497fc98a93
|
[
"MIT"
] | null | null | null |
pipeline/complete_analysis/eQTL_analysis.ipynb
|
floutt/xqtl-pipeline
|
5ee370b6ff783166f0715d0b8c7c66497fc98a93
|
[
"MIT"
] | null | null | null |
pipeline/complete_analysis/eQTL_analysis.ipynb
|
floutt/xqtl-pipeline
|
5ee370b6ff783166f0715d0b8c7c66497fc98a93
|
[
"MIT"
] | null | null | null | 40.257576 | 438 | 0.562142 |
[
[
[
"# Bulk RNA-seq eQTL analysis\n\nThis notebook provide a master control on the XQTL workflow so it can works on multiple data collection as proposed.\nInput:\n A recipe file,each row is a data collection and with the following column:\n \n Theme\n name of dataset, must be different, each uni_study analysis will be performed in a folder named after each, meta analysis will be performed in a folder named as {study1}_{study2}\n \n The column name must contain the # and be the first column\n \n genotype_list\n {Path to file}\n \n molecular_pheno\n {Path to file}\n region_list (list of regions to be analzed)\n {Path to file}\n \n covariate_file\n {Path to file}\n \n factor_analysis_opt\n \"APEX\" vs \"PEER\" for factor analysis\n \n LD options:\n \"In-sample\" LD vs {path to reference panel}\n \n QTL_tool_option\n \"APEX\" vs \"TensorQTL\" for QTL association\n \n QTL_analysis_option\n {Int for cis window} vs \"trans\"\n \n Populations\n The populations from which of the samples was drawn\n \n Conditions:\n The nature of molecular phenotype\n \n ### note: Only data collection from the same Populations and conditions will me merged to perform Fix effect meta analysis\n \nOutput:\n ...",
"_____no_output_____"
],
[
"## Generation of MWE\nThis is the code to generate the mwe recipe and LD_recipe on csg cluster\n\n",
"_____no_output_____"
]
],
[
[
"Recipe_temp = pd.DataFrame( {\"Theme\" : [\"AC\",\"DLPFC\",\"PCC\"] ,\n \"genotype_list\" : [\"/home/hs3163/GIT/ADSPFG-xQTL/MWE/mwe_genotype_list\",\n \"/home/hs3163/GIT/ADSPFG-xQTL/MWE/mwe_genotype_list\",\n \"/home/hs3163/GIT/ADSPFG-xQTL/MWE/mwe_genotype_list\"],\n \"molecular_pheno\" : [\"/home/hs3163/Project/Rosmap/data/gene_exp/AC/geneTpmResidualsAgeGenderAdj_rename.txt\",\n \"/home/hs3163/Project/Rosmap/data/gene_exp/DLPFC/geneTpmResidualsAgeGenderAdj_rename.txt\",\n \"/home/hs3163/Project/Rosmap/data/gene_exp/PCC/geneTpmResidualsAgeGenderAdj_rename.txt\"],\n \"region_list\" : [\"~/GIT/ADSPFG-xQTL/MWE/mwe_region\",\n \"~/GIT/ADSPFG-xQTL/MWE/mwe_region\",\n \"~/GIT/ADSPFG-xQTL/MWE/mwe_region\"] ,\n \"covariate_file\" : [\"/home/hs3163/GIT/ADSPFG-xQTL/MWE/MWE.cov\",\"/home/hs3163/GIT/ADSPFG-xQTL/MWE/MWE.cov\",\"/home/hs3163/GIT/ADSPFG-xQTL/MWE/MWE.cov\"],\n \"factor_analysis_opt\" : [\"APEX\",\"APEX\",\"APEX\"],\n \"LD_Recipe\": [\"~/GIT/ADSPFG-xQTL/MWE/LD_Recipe\",\"~/GIT/ADSPFG-xQTL/MWE/LD_Recipe\",\"~/GIT/ADSPFG-xQTL/MWE/LD_Recipe\"],\n \"QTL_tool_option\" : [\"APEX\",\"APEX\",\"APEX\"],\n \"QTL_analysis_option\" : [\"cis\",\"cis\",\"cis\"],\n \"cis_windows\" : [500000,500000,5000000],\n \"Metal\" : [\"T\",\"T\",\"F\"]}).to_csv(\"/home/hs3163/GIT/ADSPFG-xQTL/MWE/mwe_recipe_example\",\"\\t\")\n\n ### note: Only data collection from the same Populations and conditions will me merged to perform Fix effect meta analysis\n ",
"_____no_output_____"
],
[
"pd.DataFrame( {\"Theme\" : [\"AC\",\"DLPFC\",\"PCC\"] ,\n \"genotype_list\" : [\" /mnt/mfs/statgen/ROSMAP_xqtl/Rosmap_wgs_genotype_list.txt\",\n \" /mnt/mfs/statgen/ROSMAP_xqtl/Rosmap_wgs_genotype_list.txt\",\n \" /mnt/mfs/statgen/ROSMAP_xqtl/Rosmap_wgs_genotype_list.txt\"],\n \"molecular_pheno\" : [\"/home/hs3163/Project/Rosmap/data/gene_exp/AC/geneTpmResidualsAgeGenderAdj_rename.txt\",\n \"/home/hs3163/Project/Rosmap/data/gene_exp/DLPFC/geneTpmResidualsAgeGenderAdj_rename.txt\",\n \"/home/hs3163/Project/Rosmap/data/gene_exp/PCC/geneTpmResidualsAgeGenderAdj_rename.txt\"],\n \"region_list\" : [\"/home/hs3163/Project/Rosmap/data/gene_exp/AC/geneTpmResidualsAgeGenderAdj_rename_region_list.txt\",\n \"/home/hs3163/Project/Rosmap/data/gene_exp/AC/geneTpmResidualsAgeGenderAdj_rename_region_list.txt\",\n \"/home/hs3163/Project/Rosmap/data/gene_exp/AC/geneTpmResidualsAgeGenderAdj_rename_region_list.txt\"] ,\n \"covariate_file\" : [\"None\",\"None\",\"None\"],\n \"factor_analysis_opt\" : [\"BiCV\",\"BiCV\",\"BiCV\"],\n \"LD_Recipe\": [\"~/GIT/ADSPFG-xQTL/MWE/LD_Recipe\",\"~/GIT/ADSPFG-xQTL/MWE/LD_Recipe\",\"~/GIT/ADSPFG-xQTL/MWE/LD_Recipe\"],\n \"QTL_tool_option\" : [\"APEX\",\"APEX\",\"APEX\"],\n \"QTL_analysis_option\" : [\"cis\",\"cis\",\"cis\"],\n \"cis_windows\" : [500000,500000,500000],\n \"Metal\" : [\"T\",\"T\",\"F\"]}).to_csv(\"/home/hs3163/GIT/xqtl-pipeline/ROSMAP_recipe_example\",\"\\t\", index = 0)",
"_____no_output_____"
],
[
"home/hs3163/GIT/xqtl-pipeline/ROSMAP_recipe_example",
"_____no_output_____"
],
[
"pd.DataFrame({\"ld_file_prefix\" : [\"/mnt/mfs/statgen/neuro-twas/mv_wg/cache_arch/cache/geneTpmResidualsAgeGenderAdj_rename.\",\"/mnt/mfs/statgen/neuro-twas/mv_wg/cache_arch/cache/geneTpmResidualsAgeGenderAdj_rename.\"],\n \"ld_file_surfix\" : [\".merged.ld.rds\",\".merged.ld.rds\"]}).to_csv(\"~/GIT/ADSPFG-xQTL/MWE/LD_Recipe\",sep = \"\\t\")\n\n\n\n",
"_____no_output_____"
],
[
"nohup sos run /home/hs3163/GIT/xqtl-pipeline/pipeline/complete_analysis/eQTL_analysis.ipynb QTL \\\n --recipe /home/hs3163/GIT/ADSPFG-xQTL/MWE/mwe_recipe_example \\\n --wd ./ \\\n --exe_dir \"/home/hs3163/GIT/xqtl-pipeline/pipeline/\" &",
"_____no_output_____"
],
[
"nohup sos dryrun /home/hs3163/GIT/xqtl-pipeline/pipeline/complete_analysis/eQTL_analysis.ipynb mash_to_vcf \\\n--recipe /home/hs3163/GIT/xqtl-pipeline/ROSMAP_recipe_example --wd ./ --exe_dir \"~/GIT/xqtl-pipeline/pipeline/\" -s build &\n\nnohup sos dryrun /home/hs3163/GIT/xqtl-pipeline/pipeline/complete_analysis/eQTL_analysis.ipynb phenotype_reformatting_by_gene \\\n--recipe /home/hs3163/GIT/xqtl-pipeline/ROSMAP_recipe_example --wd ./ --exe_dir \"~/GIT/xqtl-pipeline/pipeline/\" -s build &\n\nnohup sos dryrun /home/hs3163/GIT/xqtl-pipeline/pipeline/complete_analysis/eQTL_analysis.ipynb genotype_reformatting_per_gene \\\n--recipe /home/hs3163/GIT/xqtl-pipeline/ROSMAP_recipe_example --wd ./ --exe_dir \"~/GIT/xqtl-pipeline/pipeline/\" -s build &\n\nnohup sos dryrun /home/hs3163/GIT/xqtl-pipeline/pipeline/complete_analysis/eQTL_analysis.ipynb mixture_prior \\\n--recipe /home/hs3163/GIT/xqtl-pipeline/ROSMAP_recipe_example --wd ./ --exe_dir \"~/GIT/xqtl-pipeline/pipeline/\" -s build &\n\n\nnohup sos run ~/GIT/bioworkflows/GWAS/PCA.ipynb flashpca \\\n --genoFile /mnt/mfs/statgen/xqtl_workflow_testing/ROSMAP/data_preprocessing/genotype/qc/PCC.mergrd.filtered.prune.unrelated.bed \\\n --name PCC \\\n --container_lmm /mnt/mfs/statgen/containers/xqtl_pipeline_sif/flashpcaR.sif \\\n --cwd /mnt/mfs/statgen/xqtl_workflow_testing/demo/test_pca/ \\\n -J 200 -q csg -c /home/hs3163/GIT/ADSPFG-xQTL/code/csg.yml &\n\nnohup sos run ~/GIT/bioworkflows/GWAS/PCA.ipynb project_samples:1 \\\n --genoFile /mnt/mfs/statgen/xqtl_workflow_testing/ROSMAP/data_preprocessing/genotype/qc/PCC.mergrd.filtered.prune.related.bed \\\n --pca_model /mnt/mfs/statgen/xqtl_workflow_testing/demo/test_pca/PCC.mergrd.filtered.prune.unrelated.PCC.pca.rds \\\n --name PCC \\\n --container_lmm /mnt/mfs/statgen/containers/xqtl_pipeline_sif/flashpcaR.sif \\\n --cwd /mnt/mfs/statgen/xqtl_workflow_testing/demo/test_pca/ \\\n -J 200 -q csg -c /home/hs3163/GIT/ADSPFG-xQTL/code/csg.yml &",
"_____no_output_____"
]
],
[
[
"## Example for running the workflow\nThis will run the workflow from via several submission and save the output to nohup.out",
"_____no_output_____"
],
[
"## Other example workflow:\nThese command run each of the substep to test them individually",
"_____no_output_____"
]
],
[
[
"\n",
"_____no_output_____"
],
[
"[global]\n## The aforementioned input recipe\nparameter: recipe = path\n## Overall wd, the file structure of analysis is wd/[steps]/[sub_dir for each steps]\nparameter: wd = path(\".\")\n## Diretory to the excutable\nparameter: exe_dir = path(\"~/GIT/ADSPFG-xQTL/workflow\")\n\nparameter: container = '/mnt/mfs/statgen/containers/twas_latest.sif'\nparameter: container_base_bioinfo = '/mnt/mfs/statgen/containers/xqtl_pipeline_sif/base-bioinfo.sif'\nparameter: container_apex = '/mnt/mfs/statgen/containers/xqtl_pipeline_sif/apex.sif'\nparameter: container_PEER = '/mnt/mfs/statgen/containers/xqtl_pipeline_sif/PEER.sif'\nparameter: container_TensorQTL = '/mnt/mfs/statgen/containers/xqtl_pipeline_sif/TensorQTL.sif'\nparameter: container_mvsusie = '/mnt/mfs/statgen/containers/twas_latest.sif'\nparameter: container_METAL = '/mnt/mfs/statgen/containers/xqtl_pipeline_sif/METAL.sif'\nparameter: container_flashpca = '/mnt/mfs/statgen/containers/xqtl_pipeline_sif/flashpcaR.sif'\nparameter: yml = \"/home/hs3163/GIT/ADSPFG-xQTL/code/csg.yml\"\nimport pandas as pd\ninput_inv = pd.read_csv(recipe, sep = \"\\t\")\nMetal_theme = input_inv.query(\"Metal == 'T'\")[\"Theme\"].to_list()\nMetal_theme_str = \"-\".join(Metal_theme)\nNon_Metal_theme = input_inv.query(\"Metal != 'T'\")[\"Theme\"].to_list()\nNon_Metal_theme.append(Metal_theme_str)\nTheme_Prefix = \"_\".join(Non_Metal_theme)\nparameter: LD_Recipe = path(input_inv[\"LD_Recipe\"][0])\ninput_inv = input_inv.to_dict(\"records\")\nimport os",
"_____no_output_____"
]
],
[
[
"## Molecular Phenotype Calling",
"_____no_output_____"
],
[
"## Data Preprocessing\n### Molecular Phenotype Processing",
"_____no_output_____"
]
],
[
[
"#[Normalization]\n#import os\n#input: for_each = \"input_inv\"\n#skip_if( os.path.exists(_input_inv[\"molecular_pheno\"]))\n#output: f'{wd:a}/data_preprocessing/normalization/{name}.mol_phe.bed.gz'\n#bash: expand = \"$[ ]\", stderr = f'{_output}.stderr', stdout = f'{_output}.stdout'\n# sos run $[exe_dir]/data_preprocessing/phenotype/GWAS_QC.ipynb output \\\n# --counts_gct $[_input_inv[\"genecount_table\"]] \\\n# --tpm_gct $[_input_inv[\"geneTpm_table\"]] \\\n# --sample_participant_lookup $[_input_inv[\"sample_index\"]] \\\n# --vcf_chr_list $[_input_inv[\"vcf_chr_list\"]] \\\n# --container $[container_gtex] \\\n# --name $[_input_inv[\"Theme\"]] \\\n# --wd $[wd:a]/data_preprocessing/normalization/ \\\n# --container $[container_base_bioinfo] \\\n# -J 200 -q csg -c $[yml] &",
"_____no_output_____"
],
[
"[annotation]\n## Must be ran with internet connection\nimport os\ninput: for_each = \"input_inv\"\noutput: f'{wd:a}/data_preprocessing/annotation/{_input_inv[\"Theme\"]}.{path(_input_inv[\"molecular_pheno\"]):bn}.annotated.bed.gz'\nbash: expand = \"$[ ]\", stderr = f'{_output}.stderr', stdout = f'{_output}.stdout'\n sos run $[exe_dir]/data_preprocessing/phenotype/annotation.ipynb annotation \\\n --molecular_pheno_whole $[_input_inv[\"molecular_pheno\"]] \\\n --wd $[wd:a]/data_preprocessing/annotation \\\n --name $[_input_inv[\"Theme\"]] --container $[container_base_bioinfo] -s build &",
"_____no_output_____"
],
[
"[phenotype_reformatting]\ninput: output_from(\"residual_phenotype\"),group_with = \"input_inv\"\noutput: per_chrom_pheno_list = f'{wd:a}/data_preprocessing/phenotype_reformat/{_input_inv[\"Theme\"]}.processed_phenotype.per_chrom.recipe',\n pheno_mod = f'{wd:a}/data_preprocessing/phenotype_reformat/{_input_inv[\"Theme\"]}.for_pca.mol_phe.exp'\nbash: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n sos run $[exe_dir]/data_preprocessing/phenotype/phenotype_formatting.ipynb reformat \\\n --molecular_pheno_whole $[_input] \\\n --region_list $[_input_inv[\"region_list\"]] \\\n --wd $[wd:a]/data_preprocessing/phenotype_reformat/ \\\n --name $[_input_inv[\"Theme\"]] --container $[container_base_bioinfo] \\\n -J 200 -q csg -c $[yml] ",
"_____no_output_____"
]
],
[
[
"#### The reformatiing by gene is particularly lenghthy, so to avoid exceesive waiting time, it is set to be a seperate substep",
"_____no_output_____"
]
],
[
[
"[phenotype_reformatting_by_gene]\ninput: output_from(\"residual_phenotype\"),group_with = \"input_inv\"\noutput: per_gene_pheno_list = f'{wd:a}/data_preprocessing/phenotype_reformat/{_input_inv[\"Theme\"]}.processed_phenotype.per_gene.recipe' \nbash: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n sos run $[exe_dir]/data_preprocessing/phenotype/phenotype_formatting.ipynb partition_by_gene \\\n --molecular_pheno_whole $[_input] \\\n --region_list $[_input_inv[\"region_list\"]] \\\n --wd $[wd:a]/data_preprocessing/phenotype_reformat/ \\\n --name $[_input_inv[\"Theme\"]] --container $[container_base_bioinfo] \\\n -J 200 -q csg -c $[yml] ",
"_____no_output_____"
]
],
[
[
"### Genotype Processing",
"_____no_output_____"
]
],
[
[
"[genotype_QC]\ninput: for_each = \"input_inv\"\noutput: merged_plink = f'{wd:a}/data_preprocessing/genotype/qc/{_input_inv[\"Theme\"]}.mergrd.filtered.prune.bed',\n unrelated = f'{wd:a}/data_preprocessing/genotype/qc/{_input_inv[\"Theme\"]}.mergrd.filtered.prune.unrelated.bed',\n related = f'{wd:a}/data_preprocessing/genotype/qc/{_input_inv[\"Theme\"]}.mergrd.filtered.prune.related.bed'\n#task: trunk_workers = 1, trunk_size = 1, walltime = '24h', mem = '40G', tags = f'{step_name}_{_output[0]:bn}'\nbash: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n sos run $[exe_dir]/data_preprocessing/genotype/GWAS_QC.ipynb qc \\\n --genotype_list $[_input_inv[\"genotype_list\"]] \\\n --name $[_input_inv[\"Theme\"]] \\\n --container_lmm $[container_base_bioinfo] \\\n --cwd $[wd:a]/data_preprocessing/genotype/qc/ \\\n -J 200 -q csg -c $[yml] ",
"_____no_output_____"
],
[
"[genotype_reformatting]\nimport pandas as pd\ninput: output_from(\"genotype_QC\")[\"merged_plink\"], group_with = \"input_inv\"\nname = _input_inv[\"Theme\"]\noutput: vcf_list = f'{wd}/data_preprocessing/genotype/{name}_per_chrom_vcf/{name}.vcf_chrom_list.txt',\n per_chrom_plink_list = f'{wd}/data_preprocessing/genotype/{name}_per_chrom_plink/{name}.plink_chrom_list.txt'\n#task: trunk_workers = 1, trunk_size = 1, walltime = '24h', mem = '40G', tags = f'{step_name}_{_output[0]:bn}'\nbash: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n sos run $[exe_dir]/data_preprocessing/genotype/genotype_formatting.ipynb plink2vcf \\\n --genoFile $[_input] \\\n --name $[_input_inv[\"Theme\"]] \\\n --container $[container_base_bioinfo] \\\n --region_list $[_input_inv[\"region_list\"]] \\\n --wd $[wd:a]/data_preprocessing/genotype/ \\\n -J 200 -q csg -c /home/hs3163/GIT/ADSPFG-xQTL/code/csg.yml \n \n sos run $[exe_dir]/data_preprocessing/genotype/genotype_formatting.ipynb plink_by_chrom \\\n --genoFile $[_input] \\\n --name $[_input_inv[\"Theme\"]] \\\n --region_list $[_input_inv[\"region_list\"]] \\\n --container $[container_base_bioinfo] \\\n --wd $[wd:a]/data_preprocessing/genotype/ \\\n -J 200 -q csg -c $[yml] ",
"_____no_output_____"
]
],
[
[
"#### The reformatiing by gene is particularly lenghthy, so to avoid exceesive waiting time, it is set to be a seperate substep",
"_____no_output_____"
]
],
[
[
"[genotype_reformatting_per_gene]\nimport pandas as pd\ninput: output_from(\"genotype_QC\")[\"merged_plink\"], group_with = \"input_inv\"\nname = _input_inv[\"Theme\"]\noutput: per_gene_plink = f'{wd}/data_preprocessing/genotype/{name}_per_gene_plink/{name}.plink_gene_list.txt'\nbash: expand = \"$[ ]\", stderr = f'{_output}.stderr', stdout = f'{_output}.stdout'\n sos run $[exe_dir]/data_preprocessing/genotype/genotype_formatting.ipynb plink_by_gene \\\n --genoFile $[_input] \\\n --name $[_input_inv[\"Theme\"]] \\\n --region_list $[_input_inv[\"region_list\"]] \\\n --container $[container_base_bioinfo] \\\n --region_list $[_input_inv[\"region_list\"]] \\\n --wd $[wd:a]/data_preprocessing/genotype/ \\\n -J 2000 -q csg -c $[yml] ",
"_____no_output_____"
],
[
"[LD]\nimport pandas as pd\ninput: output_from(\"genotype_reformatting\")[\"per_gene_plink\"],group_with = \"input_inv\"\noutput: f'{wd}/data_preprocessing/genotype/LD/{_input_inv[\"Theme\"]}._LD_recipe'\n#task: trunk_workers = 1, trunk_size = 1, walltime = '24h', mem = '40G', tags = f'{step_name}_{_output[0]:bn}'\nbash: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n sos run $[exe_dir]/data_preprocessing/genotype/LD.ipynb LD \\\n --genotype_list $[_input] \\\n --name $[_input_inv[\"Theme\"]] \\\n --container $[container_base_bioinfo] \\\n --wd $[wd:a]/data_preprocessing/genotype/LD/ \\\n -J 200 -q csg -c $[yml] ",
"_____no_output_____"
],
[
"[LD_Recipe]\ninput: output_from(\"LD\"), group_by = \"all\"\noutput: f'{wd:a}/data_preprocessing/genotype/LD/sumstat_list'\npython: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n import pandas as pd\n input_list = [$[_input:r,]]\n ld_recipe = pd.read_csv(input_list[0],sep = \"\\t\")\n for x in range(1,len(input_list)):\n ld_recipe = ld_recipe.append(pd.read_csv(input_list[x],sep = \"\\t\"))\n ld_recipe.to_csv(\"$[_output]\", index = 0 , sep = \"\\t\")",
"_____no_output_____"
],
[
"[GRM]\nimport pandas as pd\ninput: output_from(\"genotype_reformatting\")[\"per_chrom_plink_list\"],group_with = \"input_inv\"\noutput: f'{wd}/data_preprocessing/genotype/grm/{_input_inv[\"Theme\"]}.grm_list.txt'\n#task: trunk_workers = 1, trunk_size = 1, walltime = '24h', mem = '40G', tags = f'{step_name}_{_output[0]:bn}'\nbash: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n sos run $[exe_dir]/data_preprocessing/genotype/GRM.ipynb GRM \\\n --genotype_list $[_input] \\\n --name $[_input_inv[\"Theme\"]] \\\n --container $[container_base_bioinfo] \\\n --wd $[wd:a]/data_preprocessing/genotype/grm/ \\\n -J 200 -q csg -c $[yml] ",
"_____no_output_____"
]
],
[
[
"## Factor analysis",
"_____no_output_____"
]
],
[
[
"[factor]\ninput: output_from(\"genotype_reformatting\")[\"vcf_list\"],output_from(\"annotation\"),group_with = \"input_inv\"\noutput: f'{wd}/data_preprocessing/covariate/{_input_inv[\"Theme\"]}.{_input_inv[\"factor_analysis_opt\"]}.cov'\n#task: trunk_workers = 1, trunk_size = 1, walltime = '24h', mem = '40G', tags = f'{step_name}_{_output[0]:bn}'\nbash: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n sos run $[exe_dir]/data_preprocessing/covariate/$[_input_inv[\"factor_analysis_opt\"]]_factor.ipynb $[_input_inv[\"factor_analysis_opt\"]] \\\n --molecular_pheno $[_input[1]] \\\n --genotype_list $[_input[0]] \\\n --name $[_input_inv[\"Theme\"]] \\\n --wd $[wd:a]/data_preprocessing/covariate/ \\\n -J 200 -q csg -c $[yml] $[f'--covariate {_input_inv[\"covariate_file\"]}' if os.path.exists(_input_inv[\"covariate_file\"]) else f''] \\\n --container $[container_apex if _input_inv[\"factor_analysis_opt\"] == \"BiCV\" else container_PEER] ",
"_____no_output_____"
],
[
"[residual_phenotype]\ninput: output_from(\"factor\"), output_from(\"annotation\"),group_with = \"input_inv\"\noutput: f'{wd}/data_preprocessing/phenotype/{_input_inv[\"Theme\"]}.mol_phe.resid.bed.gz'\n#task: trunk_workers = 1, trunk_size = 1, walltime = '24h', mem = '40G', tags = f'{step_name}_{_output[0]:bn}'\nbash: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n sos run $[exe_dir]/data_preprocessing/covariate/remove_covariates.ipynb Residual_Y \\\n --molecular_pheno_whole $[_input[1]] \\\n --factor $[_input[0]] \\\n --wd $[wd]/data_preprocessing/phenotype \\\n --name $[_input_inv[\"Theme\"]] --container $[container_base_bioinfo] \\\n -J 200 -q csg -c $[yml] ",
"_____no_output_____"
],
[
"[pca]\nimport pandas as pd\ninput: output_from(\"genotype_QC\")[\"related\"],output_from(\"genotype_QC\")[\"unrelated\"],group_with = \"input_inv\"\noutput: f'{wd}/data_preprocessing/covariate/pca/{_input[0]:bn}.{_input_inv[\"Theme\"]}.pca.projected.rds'\n#task: trunk_workers = 1, trunk_size = 1, walltime = '24h', mem = '40G', tags = f'{step_name}_{_output[0]:bn}'\nbash: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n sos run $[exe_dir]/data_preprocessing/covariate/PCA.ipynb flashpca \\\n --genoFile $[_input[1]] \\\n --name $[_input_inv[\"Theme\"]] \\\n --container_lmm $[container_flashpca] \\\n --cwd $[wd:a]/data_preprocessing/covariate/pca/ \\\n -J 200 -q csg -c $[yml] \n \n sos run $[exe_dir]/data_preprocessing/covariate/PCA.ipynb project_samples:1 \\\n --genoFile $[_input[0]] \\\n --pca_model $[wd:a]/data_preprocessing/covariate/pca/$[_input[1]:bn].$[_input_inv[\"Theme\"]].pca.rds \\\n --name $[_input_inv[\"Theme\"]] \\\n --container_lmm $[container_flashpca] \\\n --cwd $[wd:a]/data_preprocessing/covariate/pca/ \\\n -J 200 -q csg -c $[yml] ",
"_____no_output_____"
],
[
"[pca_factor_merge]\nimport pandas as pd\ninput: output_from(\"pca\"),output_from(\"factor\"),group_with = \"input_inv\"\noutput: f'{wd}/data_preprocessing/covariate/{_input[1]:bn}.pca.cov'\n#task: trunk_workers = 1, trunk_size = 1, walltime = '24h', mem = '40G', tags = f'{step_name}_{_output[0]:bn}'\nbash: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n sos run $[exe_dir]/data_preprocessing/covariate/merge_covariate.ipynb pca_factor_merge \\\n --factor_and_covariate $[_input[1]] \\\n --PC $[_input[0]] \\\n --container $[container_base_bioinfo] \\\n --wd $[wd:a]/data_preprocessing/covariate/ \\\n -J 200 -q csg -c $[yml] ",
"_____no_output_____"
]
],
[
[
"## QTL associations",
"_____no_output_____"
]
],
[
[
"[QTL_1]\ninput: output_from(\"pca_factor_merge\"),output_from(\"GRM\"),output_from(\"phenotype_reformatting\")[\"per_chrom_pheno_list\"],output_from(\"genotype_reformatting\")[\"vcf_list\"], output_from(\"genotype_reformatting\")[\"per_chrom_plink_list\"] ,group_with = \"input_inv\"\noutput: f'{wd:a}/association_scan/{_input_inv[\"QTL_tool_option\"]}/{_input_inv[\"QTL_analysis_option\"]}/{_input_inv[\"Theme\"]}.{_input_inv[\"QTL_tool_option\"]}_QTL_recipe.tsv'\n#task: trunk_workers = 1, trunk_size = 1, walltime = '24h', mem = '40G', tags = f'{step_name}_{_output[0]:bn}'\nbash: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n sos run $[exe_dir]/association_scan/$[_input_inv[\"QTL_tool_option\"]]/$[_input_inv[\"QTL_tool_option\"]].ipynb $[_input_inv[\"QTL_tool_option\"]]_$[_input_inv[\"QTL_analysis_option\"]] \\\n --molecular_pheno_list $[_input[2]] \\\n --covariate $[_input[0]]\\\n --genotype_file_list $[_input[3]] \\\n --container $[container_apex if _input_inv[\"QTL_tool_option\"] == \"APEX\" else container_TensorQTL] \\\n --window $[_input_inv[\"cis_windows\"]] \\\n --name $[_input_inv[\"Theme\"]] \\\n --wd $[wd:a]/association_scan/$[_input_inv[\"QTL_tool_option\"]]/$[_input_inv[\"QTL_analysis_option\"]]/ \\\n -J 200 -q csg -c $[yml] $[f'--grm_list {_input[1]}' if _input_inv[\"QTL_tool_option\"] == \"APEX\" else f''] ",
"_____no_output_____"
]
],
[
[
"#### Example:\n\nsos run /home/hs3163/GIT/ADSPFG-xQTL/workflow/QTL_association/QTL_association.ipynb APEX_cis_Recipe \\\n --recipe data_preprocessing/PCC.data_proc_output_recipe.tsv \\\n --container /mnt/mfs/statgen/containers/apex.sif \\\n --window 500000 \\\n --name PCC \\\n --wd /mnt/mfs/statgen/xqtl_workflow_testing/testing_no_cov/QTL_association/ \\\n -J 200 -q csg -c /home/hs3163/GIT/ADSPFG-xQTL/code/csg.yml",
"_____no_output_____"
]
],
[
[
"[QTL_2]\ninput: group_by = \"all\" \noutput: f'{_input[0]:d}/sumstat_list'\npython: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n import pandas as pd\n input_list = [$[_input:r,]]\n input_inv = $[input_inv]\n sumstat_list = pd.read_csv(input_list[0],sep = \"\\t\")\n sumstat_list = sumstat_list.sort_values('#chr')\n for x in range(1,len(input_list)):\n sumstat_list = sumstat_list.merge(pd.read_csv(input_list[x],sep = \"\\t\"), on = \"#chr\")\n\n sumstat_list.columns = [\"#chr\"] + pd.DataFrame(input_inv)[\"Theme\"].values.tolist()\n sumstat_list.to_csv(\"$[_output]\", index = 0 , sep = \"\\t\")",
"_____no_output_____"
]
],
[
[
"## Meta Analysis\nInput: \n1. A recipe generated from the combination of previouse steps\n\nOutput: \n1. Recipe for Prior, Vhat, rds input, resid corr\n3. vcf",
"_____no_output_____"
]
],
[
[
"[METAL]\ninput: output_from(\"QTL_2\")\nMETAL_sumstat_list = f'{_input}.METAL.tsv'\nsumstat_list = pd.read_csv(_input,sep = \"\\t\")[[\"#chr\"] + Metal_theme].to_csv(METAL_sumstat_list,sep = \"\\t\", index = 0)\noutput: f'{wd}/multivariate/METAL/{Metal_theme_str}.METAL_list.txt'\n##task: trunk_workers = 1, trunk_size = 1, walltime = '24h', mem = '40G', tags = f'{step_name}_{_output[0]:bn}'\nbash: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n sos run $[exe_dir]/multivariate/METAL/METAL.ipynb METAL \\\n --sumstat_list_path $[METAL_sumstat_list] \\\n --wd $[wd:a]/multivariate/METAL/ --container $[container_METAL] \\\n -J 200 -q csg -c $[yml] ",
"_____no_output_____"
]
],
[
[
"## MASH",
"_____no_output_____"
]
],
[
[
"[sumstat_merger_1]\nparameter: sumstat_list = f'{wd}/multivariate/METAL/{Metal_theme_str}.METAL_list.txt'\ninput: output_from(\"QTL_2\")\noutput: yml_list = f'{wd}/multivariate/MASH/Prep/yml_list.txt',\n qced_sumstat_list = f'{wd}/multivariate/MASH/Prep/qc_sumstat_list.txt'\n##task: trunk_workers = 1, trunk_size = 1, walltime = '24h', mem = '40G', tags = f'{step_name}_{_output[0]:bn}'\nbash: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n sos run $[exe_dir]/misc/yml_generator.ipynb yml_list \\\n --sumstat_list_path $[_input] \\\n --wd $[wd:a]/multivariate/MASH/Prep/ --container $[container_base_bioinfo]\n\n sos run $[exe_dir]/misc/summary_stats_merger.ipynb \\\n --yml_list $[_output[0]] \\\n --cwd $[wd:a]/multivariate/MASH/Prep/ --container $[container_base_bioinfo] --keep_ambiguous True \\\n -J 200 -q csg -c $[yml] \n[sumstat_merger_2]\ninput: named_output(\"qced_sumstat_list\")\nname = \"_\".join(pd.DataFrame(input_inv)[\"Theme\"].values.tolist())\noutput: f'{wd}/multivariate/MASH/Prep/merge/RDS/{name}.analysis_unit'\n##task: trunk_workers = 1, trunk_size = 1, walltime = '24h', mem = '40G', tags = f'{step_name}_{_output[0]:bn}'\nbash: expand = \"$[ ]\", stderr = f'{_output}.stderr', stdout = f'{_output}.stdout'\n sos run $[exe_dir]/multivariate/MASH/sumstat_processing.ipynb processing \\\n --sumstat_list_path $[_input] \\\n --region_list $[input_inv[0][\"region_list\"]] \\\n --wd $[wd:a]/multivariate/MASH/Prep/ --container $[container_base_bioinfo] \\\n -J 2000 -q csg -c $[yml] ",
"_____no_output_____"
],
[
"[extract_effect]\ninput: output_from(\"sumstat_merger\")\nname = \"_\".join(pd.DataFrame(input_inv)[\"Theme\"].values.tolist())\noutput: f'{wd}/multivariate/MASH/Prep/{name}.rds'\nbash: expand = \"$[ ]\", stderr = f'{_output}.stderr', stdout = f'{_output}.stdout'\n sos run $[exe_dir]/multivariate/MASH/Signal_Extraction.ipynb extract_effects \\\n --cwd $[wd:a]/multivariate/MASH/Prep/ \\\n --container $[container_base_bioinfo] \\\n --name $[name] \\\n --analysis_units $[_input] \\\n -J 2000 -q csg -c $[yml] ",
"_____no_output_____"
],
[
"[mash_model]\ninput: output_from(\"extract_effect\")\nname = \"_\".join(pd.DataFrame(input_inv)[\"Theme\"].values.tolist())\noutput: MASH_model = f\"{wd}/multivariate/MASH/{name}.EZ.V_simple.mash_model.rds\",\n resid_corr = f\"{wd}/multivariate/MASH/{name}.EZ.V_simple.rds\"\nbash: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n sos run $[exe_dir]/multivariate/MASH/mashr.ipynb mash \\\n --cwd $[wd:a]/multivariate/MASH/ \\\n --container $[container_mvsusie] \\\n --output_prefix $[name] \\\n --data $[_input] \\\n -J 200 -q csg -c $[yml] \n\n[mash_posterior]\ninput: output_from(\"mash_model\")[\"MASH_model\"], output_from(\"sumstat_merger\")\nname = \"_\".join(pd.DataFrame(input_inv)[\"Theme\"].values.tolist())\nparameter: analysis_unit = _input[1]\noutput: f'{wd}/multivariate/MASH/mash_output_list'\nbash: expand = \"$[ ]\", stderr = f'{_output}.stderr', stdout = f'{_output}.stdout'\n sos run $[exe_dir]/multivariate/MASH/mashr.ipynb posterior \\\n --cwd $[wd:a]/multivariate/MASH/ \\\n --container $[container_mvsusie] \\\n --output_prefix $[name] \\\n --analysis_units $[analysis_unit] \\\n -J 2000 -q csg -c $[yml] \n\n[mash_to_vcf]\ninput: output_from(\"mash_posterior\")\nname = \"_\".join(pd.DataFrame(input_inv)[\"Theme\"].values.tolist())\noutput: f'{wd}/multivariate/MASH/mash_vcf/vcf_output_list.txt'\nbash: expand = \"$[ ]\", stderr = f'{_output}.stderr', stdout = f'{_output}.stdout'\n sos run $[exe_dir]/misc/rds_to_vcf.ipynb rds_to_vcf \\\n --wd $[wd:a]/multivariate/MASH/ \\\n --name $[name] \\\n --analysis_units $[_input] \\\n -J 2000 -q csg -c $[yml] ",
"_____no_output_____"
]
],
[
[
"## Fine mapping",
"_____no_output_____"
]
],
[
[
"[mixture_prior]\ninput: output_from(\"mash_model\")[\"MASH_model\"], output_from(\"extract_effect\")\nname = \"_\".join(pd.DataFrame(input_inv)[\"Theme\"].values.tolist())\noutput: f'{wd}/fine_mapping/mixture_prior/{name}.ed_bovy.V_simple.rds'\nbash: expand = \"$[ ]\", stderr = f'{_output}.stderr', stdout = f'{_output}.stdout'\n sos run $[exe_dir]/multivariate/MASH/mixture_prior.ipynb ed_bovy \\\n --cwd $[wd:a]/fine_mapping/mixture_prior/ \\\n --container $[container_mvsusie] \\\n --name $[name] \\\n --data $[_input[1]] \\\n --mixture_components_dir $[_input[0]:d] \\\n -J 200 -q csg -c $[yml] ",
"_____no_output_____"
],
[
"nohup sos run /home/hs3163/GIT/xqtl-pipeline/pipeline/multivariate/MASH/mixture_prior.ipynb ed_bovy --model_data fine_mapping/mixture_prior/AC_DLPFC_PCC.ed_bovy.V_simple.rds --cwd ./ --container /mnt/mfs/statgen/containers/twas_latest.sif --name AC_DLPFC_PCC --data multivariate/MASH/Prep/AC_DLPFC_PCC.rds --mixture_components_dir multivariate/MASH -J 200 -q csg -c /home/hs3163/GIT/ADSPFG-xQTL/code/csg.yml &",
"_____no_output_____"
],
[
"[mvsusie_rss]\ninput: output_from(\"mixture_prior\"), output_from(\"sumstat_merger\"), output_from(\"mash_model\")[\"resid_corr\"]\nname = \"_\".join(pd.DataFrame(input_inv)[\"Theme\"].values.tolist())\nparameter: analysis_unit = _input[1]\noutput: f'{wd:a}/fine_mapping/mvsusie_rss/{name}.mvsusie_rss.output_list.txt'\nbash: expand = \"$[ ]\", stderr = f'{_output}.stderr', stdout = f'{_output}.stdout'\n sos run $[exe_dir]/fine_mapping/SuSiE/SuSiE_RSS.ipynb MvSuSiE_summary_stats_analysis \\\n --merged_analysis_unit $[analysis_unit] \\\n --resid_cor $[_input[2]] \\\n --prior $[_input[0]] \\\n --LD_Recipe /home/hs3163/GIT/ADSPFG-xQTL/MWE/LD_Recipe \\\n --container $[container_mvsusie] \\\n --wd $[wd:a]/fine_mapping/mvsusie_rss/ \\\n --Theme_prefix $[name] -J 200 -q csg -c $[yml] ",
"_____no_output_____"
],
[
"nohup sos run /home/hs3163/GIT/xqtl-pipeline/pipeline/fine_mapping/SuSiE/SuSiE_RSS.ipynb MvSuSiE_summary_stats_analysis \\\n --merged_analysis_unit /mnt/mfs/statgen/xqtl_workflow_testing/ROSMAP/showcase_gene \\\n --resid_cor multivariate/MASH/AC_DLPFC_PCC.EZ.V_simple.rds \\\n --prior /mnt/mfs/statgen/xqtl_workflow_testing/ROSMAP/fine_mapping/mixture_prior/AC_DLPFC_PCC.ed_bovy.V_simple.rds \\\n --LD_Recipe /home/hs3163/GIT/ADSPFG-xQTL/MWE/LD_Recipe \\\n --container /mnt/mfs/statgen/containers/twas_latest.sif \\\n --wd /mnt/mfs/statgen/xqtl_workflow_testing/ROSMAP/fine_mapping/mvsusie_rss/ \\\n --Theme_prefix AC_DLPFC_PCC -J 200 -q csg -c /home/hs3163/GIT/ADSPFG-xQTL/code/csg.yml -s build &",
"_____no_output_____"
],
[
"[unisusie_rss]\ninput: output_from(\"mixture_prior\"), output_from(\"sumstat_merger\"), output_from(\"mash_model\")[\"resid_corr\"]\nname = \"_\".join(pd.DataFrame(input_inv)[\"Theme\"].values.tolist())\nparameter: analysis_unit = _input[1]\noutput: f'{wd:a}/fine_mapping/unisusie_rss/{name}.unisusie_rss.output_list.txt'\nbash: expand = \"$[ ]\", stderr = f'{_output}.stderr', stdout = f'{_output}.stdout'\n sos run $[exe_dir]/fine_mapping/SuSiE/SuSiE_RSS.ipynb UniSuSiE_summary_stats_analysis \\\n --merged_analysis_unit $[analysis_unit] \\\n --LD_Recipe /home/hs3163/GIT/ADSPFG-xQTL/MWE/LD_Recipe \\\n --container $[container_mvsusie] \\\n --wd $[wd:a]/fine_mapping/unisusie_rss/ \\\n --Theme_prefix $[name]",
"_____no_output_____"
],
[
"sos run /home/hs3163/GIT/xqtl-pipeline/pipeline/multivariate/MASH/mashr.ipynb mash \\\n --cwd ./ \\\n --container /mnt/mfs/statgen/containers/xqtl_pipeline_sif/mvsusie.sif \\\n --output_prefix AC_DLPFC_PCC \\\n --data /mnt/mfs/statgen/xqtl_workflow_testing/ROSMAP/multivariate/MASH/Prep/AC_DLPFC_PCC.rds \\\n -J 200 -q csg -c /home/hs3163/GIT/ADSPFG-xQTL/code/csg.yml &",
"_____no_output_____"
],
[
"[unisusie]\ninput: output_from(\"phenotype_reformatting_per_gene\"),output_from(\"genotype_reformatting_per_gene\"), group_with = \"input_inv\"\noutput: f'{wd:a}/fine_mapping/unisusie/{name}/{name}.unisusie.output_list.txt'\n#task: trunk_workers = 1, trunk_size = 1, walltime = '24h', mem = '40G', tags = f'{step_name}_{_output[0]:bn}'\nbash: expand = \"$[ ]\", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'\n sos run $[exe_dir]/fine_mapping/SuSiE/SuSiE.ipynb UniSuSiE_summary_stats_analysis uni_susie \\\n --phenotype_list $[_input[0]] \\\n --genotype_list $[_input[1]] \\\n --container $[container_mvsusie] \\\n --region_list $[_input_inv[\"region_list\"]] \\\n --name $[_input_inv[\"Theme\"]] \\\n --wd $[wd:a]/fine_mapping/unisusie/$[name]/ \\\n -J 200 -q csg -c $[yml] &",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab4ae533e92b85803962a8e68a6665a4319e133
| 3,499 |
ipynb
|
Jupyter Notebook
|
notebooks/book2/33/ab_test_demo.ipynb
|
patel-zeel/pyprobml
|
027ef3c13a2a63d958e05fdedb68fd7b8f0e0261
|
[
"MIT"
] | null | null | null |
notebooks/book2/33/ab_test_demo.ipynb
|
patel-zeel/pyprobml
|
027ef3c13a2a63d958e05fdedb68fd7b8f0e0261
|
[
"MIT"
] | 1 |
2022-03-27T04:59:50.000Z
|
2022-03-27T04:59:50.000Z
|
notebooks/book2/33/ab_test_demo.ipynb
|
patel-zeel/pyprobml
|
027ef3c13a2a63d958e05fdedb68fd7b8f0e0261
|
[
"MIT"
] | 2 |
2022-03-26T11:52:36.000Z
|
2022-03-27T05:17:48.000Z
| 30.426087 | 78 | 0.48471 |
[
[
[
"empty"
]
]
] |
[
"empty"
] |
[
[
"empty"
]
] |
4ab4b06991735f6de84d74e48d7caa0b314c07c2
| 8,368 |
ipynb
|
Jupyter Notebook
|
IceApp_pets.ipynb
|
airctic/icevision-gradio
|
d7bca667b8bbcd11d63581aa1b76dee91c15a869
|
[
"Apache-2.0"
] | 3 |
2020-09-23T19:31:43.000Z
|
2021-11-20T04:12:22.000Z
|
IceApp_pets.ipynb
|
airctic/icevision-gradio
|
d7bca667b8bbcd11d63581aa1b76dee91c15a869
|
[
"Apache-2.0"
] | 7 |
2020-09-23T19:40:38.000Z
|
2021-07-13T17:40:44.000Z
|
IceApp_pets.ipynb
|
airctic/icevision-gradio
|
d7bca667b8bbcd11d63581aa1b76dee91c15a869
|
[
"Apache-2.0"
] | 2 |
2020-09-23T19:29:41.000Z
|
2021-09-17T19:15:55.000Z
| 28.756014 | 234 | 0.472036 |
[
[
[
"<a href=\"https://colab.research.google.com/github/airctic/icevision-gradio/blob/master/IceApp_pets.ipynb\" target=\"_parent\"><img src=\"https://colab.research.google.com/assets/colab-badge.svg\" alt=\"Open In Colab\"/></a>",
"_____no_output_____"
],
[
"# IceVision Deployment App Example: PETS Dataset\nThis example uses Faster RCNN trained weights using the [PETS dataset](https://airctic.github.io/icedata/pets/)\n\n[IceVision](https://github.com/airctic/IceVision) features:\n\n✔ Data curation/cleaning with auto-fix \n\n✔ Exploratory data analysis dashboard \n\n✔ Pluggable transforms for better model generalization \n\n✔ Access to hundreds of neural net models (Torchvision, MMDetection, EfficientDet, Timm)\n\n✔ Access to multiple training loop libraries (Pytorch-Lightning, Fastai)\n\n✔ Multi-task training to efficiently combine object \ndetection, segmentation, and classification models ",
"_____no_output_____"
],
[
"## Installing packages",
"_____no_output_____"
]
],
[
[
"!wget https://raw.githubusercontent.com/airctic/icevision/master/install_icevision_inference.sh\n!bash install_icevision_inference.sh colab",
"_____no_output_____"
],
[
"!echo \"- Installing gradio\"\n!pip install gradio -U -q",
"_____no_output_____"
],
[
"# Restart kernel\nimport IPython\nIPython.Application.instance().kernel.do_shutdown(True)",
"_____no_output_____"
]
],
[
[
"## Imports",
"_____no_output_____"
]
],
[
[
"from icevision.all import *\nimport icedata\nimport PIL, requests\nimport torch\nfrom torchvision import transforms\nimport gradio as gr",
"_____no_output_____"
]
],
[
[
"## Loading trained model",
"_____no_output_____"
]
],
[
[
"_CLASSES = sorted(\n {\n \"Abyssinian\",\n \"great_pyrenees\",\n \"Bombay\",\n \"Persian\",\n \"samoyed\",\n \"Maine_Coon\",\n \"havanese\",\n \"beagle\",\n \"yorkshire_terrier\",\n \"pomeranian\",\n \"scottish_terrier\",\n \"saint_bernard\",\n \"Siamese\",\n \"chihuahua\",\n \"Birman\",\n \"american_pit_bull_terrier\",\n \"miniature_pinscher\",\n \"japanese_chin\",\n \"British_Shorthair\",\n \"Bengal\",\n \"Russian_Blue\",\n \"newfoundland\",\n \"wheaten_terrier\",\n \"Ragdoll\",\n \"leonberger\",\n \"english_cocker_spaniel\",\n \"english_setter\",\n \"staffordshire_bull_terrier\",\n \"german_shorthaired\",\n \"Egyptian_Mau\",\n \"boxer\",\n \"shiba_inu\",\n \"keeshond\",\n \"pug\",\n \"american_bulldog\",\n \"basset_hound\",\n \"Sphynx\",\n }\n)",
"_____no_output_____"
],
[
"class_map = ClassMap(_CLASSES)\nclass_map",
"_____no_output_____"
],
[
"# Loading model from IceZoo (IceVision Hub)\nmodel = icedata.pets.trained_models.faster_rcnn_resnet50_fpn()",
"_____no_output_____"
],
[
"# Transforms\nimage_size = 384\nvalid_tfms = tfms.A.Adapter([*tfms.A.resize_and_pad(image_size), tfms.A.Normalize()])",
"_____no_output_____"
]
],
[
[
"## Defining the `show_preds` method: called by `gr.Interface(fn=show_preds, ...)`",
"_____no_output_____"
]
],
[
[
"# Setting the model type: used in end2end_detect() method here below\nmodel_type = models.torchvision.faster_rcnn",
"_____no_output_____"
],
[
"def show_preds(input_image, display_label, display_bbox, detection_threshold):\n\n if detection_threshold==0: detection_threshold=0.5\n\n img = PIL.Image.fromarray(input_image, 'RGB')\n\n pred_dict = model_type.end2end_detect(img, valid_tfms, model, class_map=class_map, detection_threshold=detection_threshold,\n display_label=display_label, display_bbox=display_bbox, return_img=True, \n font_size=40, label_color=\"#FF59D6\")\n\n return pred_dict['img']",
"_____no_output_____"
]
],
[
[
"## Gradio User Interface",
"_____no_output_____"
]
],
[
[
"display_chkbox_label = gr.inputs.Checkbox(label=\"Label\", default=True)\ndisplay_chkbox_box = gr.inputs.Checkbox(label=\"Box\", default=True)\n\ndetection_threshold_slider = gr.inputs.Slider(minimum=0, maximum=1, step=0.1, default=0.5, label=\"Detection Threshold\")\n\noutputs = gr.outputs.Image(type=\"pil\")\n\ngr_interface = gr.Interface(fn=show_preds, inputs=[\"image\", display_chkbox_label, display_chkbox_box, detection_threshold_slider], outputs=outputs, title='IceApp - PETS')\ngr_interface.launch(inline=False, share=True, debug=True)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ab4c805708a228b3ddb5cfff10f54cb7b4f9c5c
| 46,534 |
ipynb
|
Jupyter Notebook
|
notebooks/PicnicInSF.ipynb
|
mrpozzi/mrpozzi.github.io
|
840831d8e739b654db21212cf4e62975235896ef
|
[
"MIT"
] | null | null | null |
notebooks/PicnicInSF.ipynb
|
mrpozzi/mrpozzi.github.io
|
840831d8e739b654db21212cf4e62975235896ef
|
[
"MIT"
] | null | null | null |
notebooks/PicnicInSF.ipynb
|
mrpozzi/mrpozzi.github.io
|
840831d8e739b654db21212cf4e62975235896ef
|
[
"MIT"
] | null | null | null | 131.824363 | 35,212 | 0.846521 |
[
[
[
"--- \nlayout: \npost title: \"Picnic in San Francisco\" \ndate: 2018-04-07 8:30:00 \ncategories: applications, data mining \nfeatured_image: /images/cover.jpg \n---",
"_____no_output_____"
]
],
[
[
"My beloved [SF Tsunami Master Team](http://sftsunami.org/) had planned a great picnic for this week end. For the second year in a row the plan had to be canceled due to inclement weather. \n\nI admit I sneered at the idea of having the picnic the same month as last year, considering that it got canceled once. However, forming opinions based on a sample size of two with a sprinkle of gut feeling is not the way a Scientist does things, so I thought it would be interesting and constructive to pull some data to validate or disprove my prejudice and to provide a valid alternative.",
"_____no_output_____"
],
[
"My hypothesis is quite simple: in April chances of rain are way higher than in May, while temperatures are pretty constant, so the latter would be a better option to plan outdoor activities.",
"_____no_output_____"
]
],
[
[
"%matplotlib inline\n%load_ext autoreload\n%autoreload 2\nimport pandas as pd\n\nimport urllib.request\nfrom bs4 import BeautifulSoup\n\nimport matplotlib\nfrom matplotlib import pyplot as plt\nplt.style.use('ggplot')\n\nmatplotlib.rcParams['figure.figsize'] = (20.0, 10.0)",
"The autoreload extension is already loaded. To reload it, use:\n %reload_ext autoreload\n"
]
],
[
[
"With a quick Google search I ran into [this](ggweather.com) site which reports monthly and daily information about temperature and precipitations. \n\nThe format is quite easy to scrape. The data I'm interested in are the monthly average temperatures and the number of rainy days per month.",
"_____no_output_____"
]
],
[
[
"class PicNicPlanner(object):\n \n RAIN_URL = 'http://ggweather.com/sf/daily.html'\n TEMP_URL = 'http://ggweather.com/sf/monthly%20mean%20temps.html'\n \n def __init__(self):\n self.rain_table = None\n self.temperature_table = None\n \n def _read_soup(self, url, split='\\t'):\n flob = urllib.request.urlopen(url)\n s = flob.read()\n flob.close()\n soup = BeautifulSoup(s, \"lxml\")\n return [s for s in soup.findAll('table')[1].get_text().split(split) if len(s)>0]\n \n def _clean_rain(self, row):\n return pd.Series(row.strip().split('\\n')[1:]).astype(float) \n \n def get_rains(self):\n if self.rain_table is None:\n raw_rows = self._read_soup(self.RAIN_URL, '\\xa0')\n cleaned_rows = pd.concat(\n [self._clean_rain(row) for row in raw_rows if 'Days' in row and 'Rain' not in row], \n axis=1)\n cleaned_rows.index = ['Jul', 'Aug', 'Sep', 'Oct', 'Nov', 'Dec', \n 'Jan', 'Feb', 'Mar', 'Apr', 'May', 'Jun']\n self.rain_table = cleaned_rows.transpose()\n self.rain_table.index = list(range(2008,2018))\n return self.rain_table\n \n def _clean_temperatures(self, row):\n if len(row) > 1 and not (\n 'Copyright' in row or \n 'Reproduction' in row or \n 'San Francisco' in row):\n return pd.Series(row.strip().split('\\n'))\n \n def get_temperatures(self):\n if self.temperature_table is None:\n raw_rows = self._read_soup(self.TEMP_URL)\n cleaned_rows = pd.concat([self._clean_temperatures(row) \n for row in raw_rows[2:]],axis=1)\n cleaned_rows.columns = cleaned_rows.iloc[0]\n cleaned_rows = cleaned_rows.drop(0).dropna(axis=0)\n cleaned_rows.index = ['Jan', 'Feb', 'Mar', 'Apr', 'May', 'Jun', \n 'Jul', 'Aug', 'Sep', 'Oct', 'Nov', 'Dec', 'Annual']\n self.temperature_table = cleaned_rows.transpose()\n self.temperature_table = self.temperature_table.astype(float)\n return self.temperature_table\n \n \ndef num_days(month):\n if month in ['Nov', 'Apr', 'Jun', 'Sep']:\n return 30\n if month == 'Feb':\n return 28\n return 31\n ",
"_____no_output_____"
],
[
"planner = PicNicPlanner()\ntemp = planner.get_temperatures()\nrains = planner.get_rains()",
"_____no_output_____"
]
],
[
[
"The data seems to \n\n",
"_____no_output_____"
],
[
"So, it turns out that the data confirms that having a picnic in SF in April is probably not the best idea if you want to frolic in the Sun, while your chances of having a successful event in May are almost **three times higher!** In the figure belowwe can see how April has a 23% chances of rain! Basically one day out of 4. As late as November we can have better conditions than in April, and yet I doubt people would consider reasonable to organize a picnic in November.",
"_____no_output_____"
]
],
[
[
"fig, axes = plt.subplots(nrows=2, ncols=1)\nmonths = ['Jan', 'Feb', 'Mar', 'Apr', 'May', 'Jun', 'Jul', 'Aug', 'Sep', 'Oct', 'Nov', 'Dec']\n#axes[0].set_title('Average Rainy Days by Month')\naxes[0].set_title('Changes of Rain by Month')\naxes[1].set_title('Monthly Average Temperatures')\naxes[0].axhline(10, color='r', linestyle='--')\n\n(100*rains.mean() / rains.columns.map(num_days))[months].plot(kind='bar', ax=axes[0], sharex=True, color='#000099')\naxes[0].set_ylabel('Days')\ntemp.mean()[months].plot(kind='bar', ax=axes[1], color='#7f0000')\naxes[1].set_ylabel('Hours')\naxes[1].set_xlabel('Month')\naxes[1].set_ylim([50,65]) # never too hot, never too warm",
"_____no_output_____"
],
[
"pd.DataFrame({'Daily Chances of Rain': 100*rains.mean() / rains.columns.map(num_days)}).loc[months]",
"_____no_output_____"
]
],
[
[
"The takehome message should be:\n- April is too soon to plan a picnic\n- May is quite dry\n- SF's weather is good enough to allow you to hang out outside as late as November!",
"_____no_output_____"
],
[
"[source](https://github.com/mrpozzi/mrpozzi.github.io/blob/master/notebooks/PicnicInSF.ipynb)",
"_____no_output_____"
]
]
] |
[
"raw",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"raw"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
]
] |
4ab4d1bda1a863f0c9084bc85be910938c976492
| 37,156 |
ipynb
|
Jupyter Notebook
|
notebooks/01-probability.ipynb
|
nablabits/ThinkBayes2
|
84d08578da607f62ac6cd77937d2e411230a28c4
|
[
"MIT"
] | null | null | null |
notebooks/01-probability.ipynb
|
nablabits/ThinkBayes2
|
84d08578da607f62ac6cd77937d2e411230a28c4
|
[
"MIT"
] | null | null | null |
notebooks/01-probability.ipynb
|
nablabits/ThinkBayes2
|
84d08578da607f62ac6cd77937d2e411230a28c4
|
[
"MIT"
] | null | null | null | 28.603541 | 421 | 0.586554 |
[
[
[
"# Probability",
"_____no_output_____"
],
[
"Think Bayes, Second Edition\n\nCopyright 2020 Allen B. Downey\n\nLicense: [Attribution-NonCommercial-ShareAlike 4.0 International (CC BY-NC-SA 4.0)](https://creativecommons.org/licenses/by-nc-sa/4.0/)",
"_____no_output_____"
],
[
"The foundation of Bayesian statistics is Bayes's Theorem, and the foundation of Bayes's Theorem is conditional probability. \n\nIn this chapter, we'll start with conditional probability, derive Bayes's Theorem, and demonstrate it using a real dataset. In the next chapter, we'll use Bayes's Theorem to solve problems related to conditional probability. In the chapters that follow, we'll make the transition from Bayes's Theorem to Bayesian statistics, and I'll explain the difference.",
"_____no_output_____"
],
[
"## Linda the Banker\n\nTo introduce conditional probability, I'll use an example from a [famous experiment by Tversky and Kahneman](https://en.wikipedia.org/wiki/Conjunction_fallacy), who posed the following question:\n\n> Linda is 31 years old, single, outspoken, and very bright. She majored in philosophy. As a student, she was deeply concerned with issues of discrimination and social justice, and also participated in anti-nuclear demonstrations. Which is more probable?\n> 1. Linda is a bank teller.\n> 2. Linda is a bank teller and is active in the feminist movement.\n\nMany people choose the second answer, presumably because it seems more consistent with the description. It seems uncharacteristic if Linda is *just* a bank teller; it seems more consistent if she is also a feminist.",
"_____no_output_____"
],
[
"But the second answer cannot be \"more probable\", as the question asks. Suppose we find 1000 people who fit Linda's description and 10 of them work as bank tellers. How many of them are also feminists? At most, all 10 of them are; in that case, the two options are *equally* probable. If fewer than 10 are, the second option is *less* probable. But there is no way the second option can be *more* probable.\n\nIf you were inclined to choose the second option, you are in good company. The biologist [Stephen J. Gould wrote](https://sci-hub.tw/https://doi.org/10.1080/09332480.1989.10554932) :\n\n> I am particularly fond of this example because I know that the [second] statement is least probable, yet a little [homunculus](https://en.wikipedia.org/wiki/Homunculus_argument) in my head continues to jump up and down, shouting at me, \"but she can't just be a bank teller; read the description.\"\n\nIf the little person in your head is still unhappy, maybe this chapter will help.",
"_____no_output_____"
],
[
"## Probability\n\nAt this point I should provide a definition of \"probability\", but that [turns out to be surprisingly difficult](https://en.wikipedia.org/wiki/Probability_interpretations). To avoid getting stuck before we start, we will use a simple definition for now and refine it later: A **probability** is a fraction of a finite set.\n\nFor example, if we survey 1000 people, and 20 of them are bank tellers, the fraction that work as bank tellers is 0.02 or 2\\%. If we choose a person from this population at random, the probability that they are a bank teller is 2\\%.\nBy \"at random\" I mean that every person in the dataset has the same chance of being chosen.\n\nWith this definition and an appropriate dataset, we can compute probabilities by counting.\nTo demonstrate, I'll use data from the [General Social Survey](http://gss.norc.org/) (GSS). ",
"_____no_output_____"
],
[
"The following cell downloads the data.",
"_____no_output_____"
]
],
[
[
"# Load the data file\n\nimport os\n\nif not os.path.exists('gss_bayes.csv'):\n !wget https://github.com/AllenDowney/BiteSizeBayes/raw/master/gss_bayes.csv",
"_____no_output_____"
]
],
[
[
"I'll use Pandas to read the data and store it in a `DataFrame`.",
"_____no_output_____"
]
],
[
[
"import pandas as pd\n\ngss = pd.read_csv('gss_bayes.csv', index_col=0)\ngss.head()",
"_____no_output_____"
]
],
[
[
"The `DataFrame` has one row for each person surveyed and one column for each variable I selected.\n\nThe columns are\n\n* `caseid`: Respondent id (which is the index of the table).\n\n* `year`: Year when the respondent was surveyed.\n\n* `age`: Respondent's age when surveyed.\n\n* `sex`: Male or female.\n\n* `polviews`: Political views on a range from liberal to conservative.\n\n* `partyid`: Political party affiliation, Democrat, Independent, or Republican.\n\n* `indus10`: [Code](https://www.census.gov/cgi-bin/sssd/naics/naicsrch?chart=2007) for the industry the respondent works in.\n\nLet's look at these variables in more detail, starting with `indus10`.",
"_____no_output_____"
],
[
"## Fraction of Bankers\n\nThe code for \"Banking and related activities\" is 6870, so we can select bankers like this:",
"_____no_output_____"
]
],
[
[
"banker = (gss['indus10'] == 6870)\nbanker.head()",
"_____no_output_____"
]
],
[
[
"The result is a Pandas `Series` that contains the Boolean values `True` and `False`.\n\nIf we use the `sum` function on this `Series`, it treats `True` as 1 and `False` as 0, so the total is the number of bankers.",
"_____no_output_____"
]
],
[
[
"banker.sum()",
"_____no_output_____"
]
],
[
[
"In this dataset, there are 728 bankers.\n\nTo compute the *fraction* of bankers, we can use the `mean` function, which computes the fraction of `True` values in the `Series`:",
"_____no_output_____"
]
],
[
[
"banker.mean()",
"_____no_output_____"
]
],
[
[
"About 1.5% of the respondents work in banking, so if we choose a random person from the dataset, the probability they are a banker is about 1.5%.",
"_____no_output_____"
],
[
"## The Probability Function\n\nI'll put the code from the previous section in a function that takes a Boolean series and returns a probability:",
"_____no_output_____"
]
],
[
[
"def prob(A):\n \"\"\"Computes the probability of a proposition, A.\"\"\" \n return A.mean()",
"_____no_output_____"
]
],
[
[
"So we can compute the fraction of bankers like this:",
"_____no_output_____"
]
],
[
[
"prob(banker)",
"_____no_output_____"
]
],
[
[
"Now let's look at another variable in this dataset.\nThe values of the column `sex` are encoded like this:\n\n```\n1 Male\n2 Female\n```\n\nSo we can make a Boolean series that is `True` for female respondents and `False` otherwise.",
"_____no_output_____"
]
],
[
[
"female = (gss['sex'] == 2)",
"_____no_output_____"
]
],
[
[
"And use it to compute the fraction of respondents who are women.",
"_____no_output_____"
]
],
[
[
"prob(female)",
"_____no_output_____"
]
],
[
[
"The fraction of women in this dataset is higher than in the adult U.S. population because [the GSS does not include people living in institutions](https://gss.norc.org/faq) like prisons and military housing, and those populations are more likely to be male.",
"_____no_output_____"
],
[
"## Political Views and Parties\n\nThe other variables we'll consider are `polviews`, which describes the political views of the respondents, and `partyid`, which describes their affiliation with a political party.\n\nThe values of `polviews` are on a seven-point scale:\n\n```\n1\tExtremely liberal\n2\tLiberal\n3\tSlightly liberal\n4\tModerate\n5\tSlightly conservative\n6\tConservative\n7\tExtremely conservative\n```\n\nI'll define `liberal` to be `True` for anyone whose response is \"Extremely liberal\", \"Liberal\", or \"Slightly liberal\".",
"_____no_output_____"
]
],
[
[
"liberal = (gss['polviews'] <= 3)",
"_____no_output_____"
]
],
[
[
"Here's the fraction of respondents who are liberal by this definition.",
"_____no_output_____"
]
],
[
[
"prob(liberal)",
"_____no_output_____"
]
],
[
[
"If we choose a random person in this dataset, the probability they are liberal is about 27%.",
"_____no_output_____"
],
[
"The values of `partyid` are encoded like this:\n\n```\n0\tStrong democrat\n1\tNot strong democrat\n2\tIndependent, near democrat\n3\tIndependent\n4\tIndependent, near republican\n5\tNot strong republican\n6\tStrong republican\n7\tOther party\n```\n\nI'll define `democrat` to include respondents who chose \"Strong democrat\" or \"Not strong democrat\":",
"_____no_output_____"
]
],
[
[
"democrat = (gss['partyid'] <= 1)",
"_____no_output_____"
]
],
[
[
"And here's the fraction of respondents who are Democrats, by this definition.",
"_____no_output_____"
]
],
[
[
"prob(democrat)",
"_____no_output_____"
]
],
[
[
"## Conjunction\n\nNow that we have a definition of probability and a function that computes it, let's move on to conjunction.\n\n\"Conjunction\" is another name for the logical `and` operation. If you have two [propositions](https://en.wikipedia.org/wiki/Proposition), `A` and `B`, the conjunction `A and B` is `True` if both `A` and `B` are `True`, and `False` otherwise.\n\nIf we have two Boolean series, we can use the `&` operator to compute their conjunction.\nFor example, we have already computed the probability that a respondent is a banker.",
"_____no_output_____"
]
],
[
[
"prob(banker)",
"_____no_output_____"
]
],
[
[
"And the probability that they are a Democrat:",
"_____no_output_____"
]
],
[
[
"prob(democrat)",
"_____no_output_____"
]
],
[
[
"Now we can compute the probability that a respondent is a banker *and* a Democrat:",
"_____no_output_____"
]
],
[
[
"prob(banker & democrat)",
"_____no_output_____"
]
],
[
[
"As we should expect, `prob(banker & democrat)` is less than `prob(banker)`, because not all bankers are Democrats.\n\nWe expect conjunction to be commutative; that is, `A & B` should be the same as `B & A`. To check, we can also compute `prob(democrat & banker)`:",
"_____no_output_____"
]
],
[
[
"prob(democrat & banker)",
"_____no_output_____"
]
],
[
[
"As expected, they are the same.",
"_____no_output_____"
],
[
"## Conditional Probability\n\nConditional probability is a probability that depends on a condition, but that might not be the most helpful definition. Here are some examples:\n\n* What is the probability that a respondent is a Democrat, given that they are liberal?\n\n* What is the probability that a respondent is female, given that they are a banker?\n\n* What is the probability that a respondent is liberal, given that they are female?\n\nLet's start with the first one, which we can interpret like this: \"Of all the respondents who are liberal, what fraction are Democrats?\"\n\nWe can compute this probability in two steps:\n\n1. Select all respondents who are liberal.\n\n2. Compute the fraction of the selected respondents who are Democrats.\n\nTo select liberal respondents, we can use the bracket operator, `[]`, like this:",
"_____no_output_____"
]
],
[
[
"selected = democrat[liberal]",
"_____no_output_____"
]
],
[
[
"`selected` contains the values of `democrat` for liberal respondents, so `prob(selected)` is the fraction of liberals who are Democrats:",
"_____no_output_____"
]
],
[
[
"prob(selected)",
"_____no_output_____"
]
],
[
[
"A little more than half of liberals are Democrats. If that result is lower than you expected, keep in mind:\n\n1. We used a somewhat strict definition of \"Democrat\", excluding Independents who \"lean\" democratic.\n\n2. The dataset includes respondents as far back as 1974; in the early part of this interval, there was less alignment between political views and party affiliation, compared to the present.\n\nLet's try the second example, \"What is the probability that a respondent is female, given that they are a banker?\"\nWe can interpret that to mean, \"Of all respondents who are bankers, what fraction are female?\"\n\nAgain, we'll use the bracket operator to select only the bankers and `prob` to compute the fraction that are female.",
"_____no_output_____"
]
],
[
[
"selected = female[banker]\nprob(selected)",
"_____no_output_____"
]
],
[
[
"About 77% of the bankers in this dataset are female.\n\nLet's wrap this computation in a function.\nI'll define `conditional` to take two Boolean series, `proposition` and `given`, and compute the conditional probability of `proposition` conditioned on `given`:",
"_____no_output_____"
]
],
[
[
"def conditional(proposition, given):\n \"\"\"Probability of A conditioned on given.\"\"\"\n return prob(proposition[given])",
"_____no_output_____"
]
],
[
[
"We can use `conditional` to compute the probability that a respondent is liberal given that they are female.",
"_____no_output_____"
]
],
[
[
"conditional(liberal, given=female)",
"_____no_output_____"
]
],
[
[
"About 28% of female respondents are liberal.\n\nI included the keyword, `given`, along with the parameter, `female`, to make this expression more readable.",
"_____no_output_____"
],
[
"## Conditional Probability Is Not Commutative\n\nWe have seen that conjunction is commutative; that is, `prob(A & B)` is always equal to `prob(B & A)`.\n\nBut conditional probability is *not* commutative; that is, `conditional(A, B)` is not the same as `conditional(B, A)`.\n\nThat should be clear if we look at an example. Previously, we computed the probability a respondent is female, given that they are banker.",
"_____no_output_____"
]
],
[
[
"conditional(female, given=banker)",
"_____no_output_____"
]
],
[
[
"The result shows that the majority of bankers are female. That is not the same as the probability that a respondent is a banker, given that they are female:",
"_____no_output_____"
]
],
[
[
"conditional(banker, given=female)",
"_____no_output_____"
]
],
[
[
"Only about 2% of female respondents are bankers.\n\nI hope this example makes it clear that conditional probability is not commutative, and maybe it was already clear to you. Nevertheless, it is a common error to confuse `conditional(A, B)` and `conditional(B, A)`. We'll see some examples later.",
"_____no_output_____"
],
[
"## Condition and Conjunction\n\nWe can combine conditional probability and conjunction. For example, here's the probability a respondent is female, given that they are a liberal Democrat.",
"_____no_output_____"
]
],
[
[
"conditional(female, given=liberal & democrat)",
"_____no_output_____"
]
],
[
[
"About 57% of liberal Democrats are female.\n\nAnd here's the probability they are a liberal female, given that they are a banker:",
"_____no_output_____"
]
],
[
[
"conditional(liberal & female, given=banker)",
"_____no_output_____"
]
],
[
[
"About 17% of bankers are liberal women.",
"_____no_output_____"
],
[
"## Laws of Probability\n\nIn the next few sections, we'll derive three relationships between conjunction and conditional probability:\n\n* Theorem 1: Using a conjunction to compute a conditional probability.\n\n* Theorem 2: Using a conditional probability to compute a conjunction.\n\n* Theorem 3: Using `conditional(A, B)` to compute `conditional(B, A)`.\n\nTheorem 3 is also known as Bayes's Theorem.\n\nI'll write these theorems using mathematical notation for probability:\n\n* $P(A)$ is the probability of proposition $A$.\n\n* $P(A~\\mathrm{and}~B)$ is the probability of the conjunction of $A$ and $B$, that is, the probability that both are true.\n\n* $P(A | B)$ is the conditional probability of $A$ given that $B$ is true. The vertical line between $A$ and $B$ is pronounced \"given\". \n\nWith that, we are ready for Theorem 1.",
"_____no_output_____"
],
[
"### Theorem 1\n\nWhat fraction of bankers are female? We have already seen one way to compute the answer:\n\n1. Use the bracket operator to select the bankers, then\n\n2. Use `mean` to compute the fraction of bankers who are female.\n\nWe can write these steps like this:",
"_____no_output_____"
]
],
[
[
"female[banker].mean()",
"_____no_output_____"
]
],
[
[
"Or we can use the `conditional` function, which does the same thing:",
"_____no_output_____"
]
],
[
[
"conditional(female, given=banker)",
"_____no_output_____"
]
],
[
[
"But there is another way to compute this conditional probability, by computing the ratio of two probabilities:\n\n1. The fraction of respondents who are female bankers, and\n\n2. The fraction of respondents who are bankers.\n\nIn other words: of all the bankers, what fraction are female bankers?\nHere's how we compute this ratio.",
"_____no_output_____"
]
],
[
[
"prob(female & banker) / prob(banker)",
"_____no_output_____"
]
],
[
[
"The result is the same. This example demonstrates a general rule that relates conditional probability and conjunction. Here's what it looks like in math notation:\n\n$$P(A|B) = \\frac{P(A~\\mathrm{and}~B)}{P(B)}$$\n\nAnd that's Theorem 1.",
"_____no_output_____"
],
[
"### Theorem 2\n\nIf we start with Theorem 1 and multiply both sides by $P(B)$, we get Theorem 2.\n\n$$P(A~\\mathrm{and}~B) = P(B) ~ P(A|B)$$\n\nThis formula suggests a second way to compute a conjunction: instead of using the `&` operator, we can compute the product of two probabilities.\n\nLet's see if it works for `liberal` and `democrat`. Here's the result using `&`:",
"_____no_output_____"
]
],
[
[
"prob(liberal & democrat)",
"_____no_output_____"
]
],
[
[
"And here's the result using Theorem 2:",
"_____no_output_____"
]
],
[
[
"prob(democrat) * conditional(liberal, democrat)",
"_____no_output_____"
]
],
[
[
"They are the same.",
"_____no_output_____"
],
[
"### Theorem 3\n\nWe have established that conjunction is commutative. In math notation, that means:\n\n$$P(A~\\mathrm{and}~B) = P(B~\\mathrm{and}~A)$$\n\nIf we apply Theorem 2 to both sides, we have\n\n$$P(B) P(A|B) = P(A) P(B|A)$$\n\nHere's one way to interpret that: if you want to check $A$ and $B$, you can do it in either order:\n\n1. You can check $B$ first, then $A$ conditioned on $B$, or\n\n2. You can check $A$ first, then $B$ conditioned on $A$.\n\nIf we divide through by $P(B)$, we get Theorem 3:\n\n$$P(A|B) = \\frac{P(A) P(B|A)}{P(B)}$$\n\nAnd that, my friends, is Bayes's Theorem.\n\nTo see how it works, let's compute the fraction of bankers who are liberal, first using `conditional`:",
"_____no_output_____"
]
],
[
[
"conditional(liberal, given=banker)",
"_____no_output_____"
]
],
[
[
"Now using Bayes's Theorem:",
"_____no_output_____"
]
],
[
[
"prob(liberal) * conditional(banker, liberal) / prob(banker)",
"_____no_output_____"
]
],
[
[
"They are the same.",
"_____no_output_____"
],
[
"### The Law of Total Probability\n\nIn addition to these three theorems, there's one more thing we'll need to do Bayesian statistics: the law of total probability.\nHere's one form of the law, expressed in mathematical notation:\n\n$$P(A) = P(B_1 \\mathrm{and} A) + P(B_2 \\mathrm{and} A)$$\n\nIn words, the total probability of $A$ is the sum of two possibilities: either $B_1$ and $A$ are true or $B_2$ and $A$ are true.\nBut this law applies only if $B_1$ and $B_2$ are:\n\n* Mutually exclusive, which means that only one of them can be true, and\n\n* Collectively exhaustive, which means that one of them must be true.\n\nAs an example, let's use this law to compute the probability that a respondent is a banker.\nWe can compute it directly like this:",
"_____no_output_____"
]
],
[
[
"prob(banker)",
"_____no_output_____"
]
],
[
[
"So let's confirm that we get the same thing if we compute male and female bankers separately.\n\nIn this dataset all respondents are designated male or female. Recently, the GSS Board of Overseers announced that they will add more inclusive gender questions to the survey (you can read more about this issue, and their decision, at https://gender.stanford.edu/news-publications/gender-news/more-inclusive-gender-questions-added-general-social-survey).\n\n\n\nWe already have a Boolean `Series` that is `True` for female respondents.\nHere's the complementary `Series` for male respondents.",
"_____no_output_____"
]
],
[
[
"male = (gss['sex'] == 1)",
"_____no_output_____"
]
],
[
[
"Now we can compute the total probability of `banker` like this. ",
"_____no_output_____"
]
],
[
[
"prob(male & banker) + prob(female & banker)",
"_____no_output_____"
]
],
[
[
"Because `male` and `female` are mutually exclusive and collectively exhaustive (MECE), we get the same result we got by computing the probability of `banker` directly.\n\nApplying Theorem 2, we can also write the law of total probability like this:\n\n$$P(A) = P(B_1) P(A|B_1) + P(B_2) P(A|B_2)$$\n\nAnd we can test it with the same example:",
"_____no_output_____"
]
],
[
[
"(prob(male) * conditional(banker, given=male) +\nprob(female) * conditional(banker, given=female))",
"_____no_output_____"
]
],
[
[
"When there are more than two conditions, it is more concise to write the law of total probability as a summation:\n\n$$P(A) = \\sum_i P(B_i) P(A|B_i)$$\n\nAgain, this holds as long as the conditions, $B_i$ are mutually exclusive and collectively exhaustive.\nAs an example, let's consider `polviews`, which has seven different values.",
"_____no_output_____"
]
],
[
[
"B = gss['polviews']\nB.value_counts().sort_index()",
"_____no_output_____"
]
],
[
[
"On this scale, `4.0` represents \"Moderate\".\nSo we can compute the probability of a moderate banker like this:",
"_____no_output_____"
]
],
[
[
"i = 4\nprob(B==i) * conditional(banker, B==i)",
"_____no_output_____"
]
],
[
[
"And we can use `sum` and a [generator expression](https://www.johndcook.com/blog/2020/01/15/generator-expression/) to compute the summation.",
"_____no_output_____"
]
],
[
[
"sum(prob(B==i) * conditional(banker, B==i)\n for i in range(1, 8))",
"_____no_output_____"
]
],
[
[
"The result is the same.\n\nIn this example, using the law of total probability is a lot more work than computing the probability directly, but it will turn out to be useful, I promise.",
"_____no_output_____"
],
[
"## Summary\n\nHere's what we have so far:\n\n**Theorem 1** gives us a way to compute a conditional probability using a conjunction:\n\n$$P(A|B) = \\frac{P(A~\\mathrm{and}~B)}{P(B)}$$\n\n**Theorem 2** gives us a way to compute a conjunction using a conditional probability:\n\n$$P(A~\\mathrm{and}~B) = P(B) P(A|B)$$\n\n**Theorem 3**, also known as Bayes's Theorem, gives us a way to get from $P(A|B)$ to $P(B|A)$, or the other way around:\n\n$$P(A|B) = \\frac{P(A) P(B|A)}{P(B)}$$\n\n**The Law of Total Probability** provides a way to compute probabilities by adding up the pieces:\n\n$$P(A) = \\sum_i P(B_i) P(A|B_i)$$\n\nAt this point you might ask, \"So what?\" If we have all of the data, we can compute any probability we want, any conjunction, or any conditional probability, just by counting. We don't have to use these formulas.\n\nAnd you are right, *if* we have all of the data. But often we don't, and in that case, these formulas can be pretty useful -- especially Bayes's Theorem.\nIn the next chapter, we'll see how.",
"_____no_output_____"
],
[
"## Exercises",
"_____no_output_____"
],
[
"**Exercise:** Let's use the tools in this chapter to solve a variation of the Linda problem.\n\n> Linda is 31 years old, single, outspoken, and very bright. She majored in philosophy. As a student, she was deeply concerned with issues of discrimination and social justice, and also participated in anti-nuclear demonstrations. Which is more probable?\n> 1. Linda is a banker.\n> 2. Linda is a banker and considers herself a liberal Democrat.\n\nTo answer this question, compute \n\n* The probability that Linda is a female banker,\n\n* The probability that Linda is a liberal female banker, and\n\n* The probability that Linda is a liberal female banker and a Democrat.",
"_____no_output_____"
]
],
[
[
"# Solution goes here",
"_____no_output_____"
],
[
"# Solution goes here",
"_____no_output_____"
],
[
"# Solution goes here",
"_____no_output_____"
]
],
[
[
"**Exercise:** Use `conditional` to compute the following probabilities:\n\n* What is the probability that a respondent is liberal, given that they are a Democrat?\n\n* What is the probability that a respondent is a Democrat, given that they are liberal?\n\nThink carefully about the order of the arguments you pass to `conditional`.",
"_____no_output_____"
]
],
[
[
"# Solution goes here",
"_____no_output_____"
],
[
"# Solution goes here",
"_____no_output_____"
]
],
[
[
"**Exercise:** There's a [famous quote](https://quoteinvestigator.com/2014/02/24/heart-head/) about young people, old people, liberals, and conservatives that goes something like:\n\n> If you are not a liberal at 25, you have no heart. If you are not a conservative at 35, you have no brain.\n\nWhether you agree with this proposition or not, it suggests some probabilities we can compute as an exercise.\nRather than use the specific ages 25 and 35, let's define `young` and `old` as under 30 or over 65:",
"_____no_output_____"
]
],
[
[
"young = (gss['age'] < 30)\nprob(young)",
"_____no_output_____"
],
[
"old = (gss['age'] >= 65)\nprob(old)",
"_____no_output_____"
]
],
[
[
"For these thresholds, I chose round numbers near the 20th and 80th percentiles. Depending on your age, you may or may not agree with these definitions of \"young\" and \"old\".\n\nI'll define `conservative` as someone whose political views are \"Conservative\", \"Slightly Conservative\", or \"Extremely Conservative\".",
"_____no_output_____"
]
],
[
[
"conservative = (gss['polviews'] >= 5)\nprob(conservative)",
"_____no_output_____"
]
],
[
[
"Use `prob` and `conditional` to compute the following probabilities.\n\n* What is the probability that a randomly chosen respondent is a young liberal?\n\n* What is the probability that a young person is liberal?\n\n* What fraction of respondents are old conservatives?\n\n* What fraction of conservatives are old?\n\nFor each statement, think about whether it is expressing a conjunction, a conditional probability, or both.\n\nFor the conditional probabilities, be careful about the order of the arguments.\nIf your answer to the last question is greater than 30%, you have it backwards!",
"_____no_output_____"
]
],
[
[
"# Solution goes here",
"_____no_output_____"
],
[
"# Solution goes here",
"_____no_output_____"
],
[
"# Solution goes here",
"_____no_output_____"
],
[
"# Solution goes here",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
4ab4e8a14b80437b339def480d25ec584858d0fb
| 5,005 |
ipynb
|
Jupyter Notebook
|
Copy_of_LS_DS_144_Real_world_Experiment_Design.ipynb
|
mowgli28/DS-Unit-1-Sprint-4-Statistical-Tests-and-Experiments
|
36b3b9abcbcf03b4ca02338e9cc6b2088eeb1070
|
[
"MIT"
] | null | null | null |
Copy_of_LS_DS_144_Real_world_Experiment_Design.ipynb
|
mowgli28/DS-Unit-1-Sprint-4-Statistical-Tests-and-Experiments
|
36b3b9abcbcf03b4ca02338e9cc6b2088eeb1070
|
[
"MIT"
] | null | null | null |
Copy_of_LS_DS_144_Real_world_Experiment_Design.ipynb
|
mowgli28/DS-Unit-1-Sprint-4-Statistical-Tests-and-Experiments
|
36b3b9abcbcf03b4ca02338e9cc6b2088eeb1070
|
[
"MIT"
] | null | null | null | 38.79845 | 436 | 0.590809 |
[
[
[
"<a href=\"https://colab.research.google.com/github/mowgli28/DS-Unit-1-Sprint-4-Statistical-Tests-and-Experiments/blob/master/Copy_of_LS_DS_144_Real_world_Experiment_Design.ipynb\" target=\"_parent\"><img src=\"https://colab.research.google.com/assets/colab-badge.svg\" alt=\"Open In Colab\"/></a>",
"_____no_output_____"
],
[
"# Lambda School Data Science Module 144\n## Real-world Experiment Design\n\n\n\n[Induction experiment, Wikipedia](https://commons.wikimedia.org/wiki/File:Induction_experiment.png)",
"_____no_output_____"
],
[
"## Prepare - Learn about JavaScript and Google Analytics",
"_____no_output_____"
],
[
"Python is great - but with web applications, it's impossible to avoid JavaScript. The lingua franca of the web, JavaScript runs in all browsers, and thus all front-end code must either be JS or transpiled to it. As a data scientist you don't have to learn JavaScript - but you do have to be aware of it, and being able to figure out snippets of it is an invaluable skill to connect your skills with real-world applications.\n\nSo, we leave the warm comfort of Python, and venture to a bigger world - check out the [LambdaSchool/AB-Demo repo](https://github.com/LambdaSchool/AB-Demo) and [live experiment](https://lambdaschool.github.io/AB-Demo/) before class.\n\nAdditionally, sign up for [Google Analytics](https://www.google.com/analytics) - if you're not sure on the steps or what \"property\" to give it, you can put a placeholder or wait until the live lecture. Google also has [Analytics documentation](https://support.google.com/analytics/) that is worth a look.\n\nNote - if you use any of the various tracker blocking techniques, it's quite likely you won't show up in Google Analytics. You'll have to disable them to be able to fully test your experiment.",
"_____no_output_____"
],
[
"## Live Lecture - Using Google Analytics with a live A/B test",
"_____no_output_____"
],
[
"Again we won't do much Python here, but we'll put a few notes and results in the notebook as we go.",
"_____no_output_____"
],
[
"## Assignment - Set up your own A/B test!",
"_____no_output_____"
],
[
"For a baseline, a straight fork of the Lambda School repo is OK. Getting that working with your own Analytics profile is already a task. But if you get through that, stretch goals:\n\n1. Explore Google Analytics - it's big and changes frequently, but powerful (can track conversions and events, flows, etc.)\n2. Customize the experiment to be more interesting/different (try colors!)\n3. Check out the various tools for setting up A/B experiments (e.g. [Optimizely](https://www.optimizely.com/) and [alternatives](https://alternativeto.net/software/optimizely/))\n4. Try to get enough traffic to actually have more real data (don't spam people, but do share with friends)\n5. If you do get more traffic, don't just apply a t-test - dig into the results and use both math and writing to describe your findings",
"_____no_output_____"
]
],
[
[
"https://mowgli28.github.io/AB-Demo/",
"_____no_output_____"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
]
] |
4ab4e96722d0168669d4494afc914827e6ee532d
| 101,730 |
ipynb
|
Jupyter Notebook
|
Artificial Neural Network/detect.ipynb
|
kkzzo1o1/kkzzo1o1
|
ad9b2503c1385226db733f5b4d2e1aca5af218a4
|
[
"MIT"
] | null | null | null |
Artificial Neural Network/detect.ipynb
|
kkzzo1o1/kkzzo1o1
|
ad9b2503c1385226db733f5b4d2e1aca5af218a4
|
[
"MIT"
] | null | null | null |
Artificial Neural Network/detect.ipynb
|
kkzzo1o1/kkzzo1o1
|
ad9b2503c1385226db733f5b4d2e1aca5af218a4
|
[
"MIT"
] | null | null | null | 45.577957 | 216 | 0.406154 |
[
[
[
"# 目標:利用openpose和神經網路辨識T-POSE和DAB姿勢。",
"_____no_output_____"
],
[
"# 組員:\n# 應數一 108701011 游能澤\n# 應數一 108701034 柯里橫\n# 應數一 108701018 池欣霓\n# 應數一 108701019 許辰宇",
"_____no_output_____"
],
[
"# 分工:\n# 構想與建設環境:游能澤、柯里橫\n# 程式建構:柯里橫",
"_____no_output_____"
],
[
"# 由於一直無法使用pyopenpose函式,無法透過\n# openpose取得樣本,只好找別人做好的。",
"_____no_output_____"
],
[
"# 到https://github.com/burningion/dab-and-tpose-controlled-lights/tree/master/data\n# 載下作者生成好的數據,開個data資料夾放進去\n# dabs.npy tposes.npy other.npy\n# more-dabs.npy more-tposes.npy more-other.npy\n# test-dabs.npy",
"_____no_output_____"
]
],
[
[
"import numpy as np",
"_____no_output_____"
]
],
[
[
"# 這幾個檔案是以numpy儲存的二進制文件\n# 裡頭有我們需要的特徵",
"_____no_output_____"
]
],
[
[
"dabDataset = np.load('data/dabs.npy')\ntposeDataset = np.load('data/tposes.npy')\notherDataset = np.load('data/other.npy')",
"_____no_output_____"
],
[
"dabDataset[0]",
"_____no_output_____"
],
[
"dabDataset.shape",
"_____no_output_____"
],
[
"dabDataset[0].shape",
"_____no_output_____"
]
],
[
[
"# Adding our Labels\n# Our labels come from the [BODY_25 Pose Output format](https://github.com/CMU-Perceptual-Computing-Lab/openpose/blob/master/doc/output.md#pose-output-format-body_25) available at the repo.",
"_____no_output_____"
]
],
[
[
"labels = [\"Nose\", \"Neck\", \"RShoulder\", \"RElbow\", \"RWrist\", \"LShoulder\", \"LElbow\",\n \"LWrist\", \"MidHip\", \"RHip\", \"RKnee\", \"RAnkle\", \"LHip\", \"LKnee\", \"LAnkle\",\n \"REye\", \"LEye\", \"REar\", \"LEar\", \"LBigToe\", \"LSmallToe\", \"LHeel\", \"RBigToe\",\n \"RSmallToe\", \"RHeel\", \"Background\"]",
"_____no_output_____"
]
],
[
[
"## 看數據有三個維度分別是X、Y、Confidence,不需要用到Confidence。",
"_____no_output_____"
]
],
[
[
"properLabels = []\nfor label in labels:\n properLabels.append(label + 'X')\n properLabels.append(label + 'Y')\n properLabels.append(label + 'Confidence')",
"_____no_output_____"
],
[
"import csv\n\nwith open('data/dabs.csv', 'w+') as dabcsv:\n dabwriter = csv.writer(dabcsv, delimiter=',')\n dabwriter.writerow(properLabels)\n for cell in dabDataset:\n dabwriter.writerow(cell.flatten())\n \nwith open('data/tposes.csv', 'w+') as tposecsv:\n tposewriter = csv.writer(tposecsv, delimiter=',')\n tposewriter.writerow(properLabels)\n for cell in tposeDataset:\n tposewriter.writerow(cell.flatten())\n \nwith open('data/other.csv', 'w+') as othercsv:\n otherwriter = csv.writer(othercsv, delimiter=',')\n otherwriter.writerow(properLabels)\n for cell in otherDataset:\n otherwriter.writerow(cell.flatten())",
"_____no_output_____"
]
],
[
[
"# 用CSV檔看訓練資料略少,但還是試著訓練看看。",
"_____no_output_____"
],
[
"# Creating a Labeled Dataset for Training and Testing\n# We'll use 0 for other poses, 1 for dabs, and 2 for tposes.",
"_____no_output_____"
]
],
[
[
"labels = np.zeros(len(otherDataset))\nlabels = np.append(labels, np.full((len(dabDataset)), 1))\nlabels = np.append(labels, np.full((len(tposeDataset)), 2))\nprint(labels)\nprint(\"%i total examples for training.\" % len(labels))",
"[0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.\n 0. 0. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 2. 2. 2. 2. 2. 2. 2. 2.\n 2. 2. 2. 2. 2. 2. 2. 2.]\n56 total examples for training.\n"
],
[
"dataset = np.append(otherDataset, dabDataset, axis=0)\ndataset = np.append(dataset, tposeDataset, axis=0)\nprint(dataset)",
"[[[488.3213 147.51425 0.83340967]\n [494.22372 284.5734 0.8012297 ]\n [386.4863 270.83716 0.66853976]\n ...\n [ 0. 0. 0. ]\n [ 0. 0. 0. ]\n [ 0. 0. 0. ]]\n\n [[515.7737 112.18818 0.83487195]\n [478.48004 274.7029 0.8005627 ]\n [368.77948 257.2105 0.6782713 ]\n ...\n [ 0. 0. 0. ]\n [ 0. 0. 0. ]\n [ 0. 0. 0. ]]\n\n [[547.1316 112.15151 0.79948723]\n [464.79065 268.88403 0.73338044]\n [360.98135 243.43745 0.62600124]\n ...\n [ 0. 0. 0. ]\n [ 0. 0. 0. ]\n [ 0. 0. 0. ]]\n\n ...\n\n [[509.97504 257.06958 0.892523 ]\n [460.9663 351.17117 0.7867987 ]\n [372.75305 333.54434 0.6111988 ]\n ...\n [ 0. 0. 0. ]\n [ 0. 0. 0. ]\n [ 0. 0. 0. ]]\n\n [[554.90063 286.50854 0.88104486]\n [496.1236 374.6841 0.7804795 ]\n [415.7811 353.13678 0.74092144]\n ...\n [ 0. 0. 0. ]\n [ 0. 0. 0. ]\n [ 0. 0. 0. ]]\n\n [[570.5084 268.88422 0.855286 ]\n [509.95016 370.85114 0.7977039 ]\n [431.50925 353.08978 0.7552353 ]\n ...\n [ 0. 0. 0. ]\n [ 0. 0. 0. ]\n [ 0. 0. 0. ]]]\n"
],
[
"dataset.shape",
"_____no_output_____"
]
],
[
[
"## 讓數值變成0~1之間",
"_____no_output_____"
]
],
[
[
"dataset[:,:,1] / 1280",
"_____no_output_____"
]
],
[
[
"# Shuffle labels and features",
"_____no_output_____"
]
],
[
[
"from sklearn.utils import shuffle\nX, y = shuffle(dataset, labels)\nfrom keras.utils.np_utils import to_categorical\ny = to_categorical(y, 3)\nprint(y.shape[1])",
"Using TensorFlow backend.\n"
]
],
[
[
"# 架設神經網路",
"_____no_output_____"
]
],
[
[
"from keras.models import Sequential\nfrom keras.layers import Dense, Dropout, Activation, Flatten\nfrom keras.optimizers import SGD",
"_____no_output_____"
],
[
"X = X.reshape(len(X), 75)\ndataset[0]",
"_____no_output_____"
],
[
"model = Sequential()\nmodel.add(Dense(128, activation = 'relu', input_shape = (75,)))\nmodel.add(Dropout(0.5))\nmodel.add(Dense(128, activation = 'relu'))\nmodel.add(Dropout(0.5))\nmodel.add(Dense(y.shape[1], activation = 'softmax'))\nmodel.compile(optimizer = SGD(lr = 0.005),\n loss = 'categorical_crossentropy',\n metrics = ['accuracy'])\nmodel.fit(X, y, epochs = 200, batch_size = 64)",
"Epoch 1/200\n56/56 [==============================] - 0s 3ms/step - loss: 349.1770 - accuracy: 0.3571\nEpoch 2/200\n56/56 [==============================] - 0s 53us/step - loss: 10615.8975 - accuracy: 0.2500\nEpoch 3/200\n56/56 [==============================] - 0s 35us/step - loss: 40285.3789 - accuracy: 0.4643\nEpoch 4/200\n56/56 [==============================] - 0s 32us/step - loss: 39818.2070 - accuracy: 0.2857\nEpoch 5/200\n56/56 [==============================] - 0s 66us/step - loss: 6458.7578 - accuracy: 0.4643\nEpoch 6/200\n56/56 [==============================] - 0s 85us/step - loss: 1.0930 - accuracy: 0.3393\nEpoch 7/200\n56/56 [==============================] - 0s 67us/step - loss: 1.0814 - accuracy: 0.4643\nEpoch 8/200\n56/56 [==============================] - 0s 40us/step - loss: 1.1014 - accuracy: 0.4643\nEpoch 9/200\n56/56 [==============================] - 0s 39us/step - loss: 1.0853 - accuracy: 0.4643\nEpoch 10/200\n56/56 [==============================] - 0s 37us/step - loss: 1.0844 - accuracy: 0.4643\nEpoch 11/200\n56/56 [==============================] - 0s 36us/step - loss: 1.0920 - accuracy: 0.4643\nEpoch 12/200\n56/56 [==============================] - 0s 34us/step - loss: 1.1036 - accuracy: 0.4643\nEpoch 13/200\n56/56 [==============================] - 0s 71us/step - loss: 1.0882 - accuracy: 0.4643\nEpoch 14/200\n56/56 [==============================] - 0s 71us/step - loss: 1.0975 - accuracy: 0.4643\nEpoch 15/200\n56/56 [==============================] - 0s 69us/step - loss: 1.0845 - accuracy: 0.4643\nEpoch 16/200\n56/56 [==============================] - 0s 37us/step - loss: 1.0877 - accuracy: 0.4643\nEpoch 17/200\n56/56 [==============================] - 0s 32us/step - loss: 1.0809 - accuracy: 0.4643\nEpoch 18/200\n56/56 [==============================] - 0s 36us/step - loss: 1.0869 - accuracy: 0.4643\nEpoch 19/200\n56/56 [==============================] - 0s 35us/step - loss: 1.0983 - accuracy: 0.4643\nEpoch 20/200\n56/56 [==============================] - 0s 35us/step - loss: 1.0788 - accuracy: 0.4643\nEpoch 21/200\n56/56 [==============================] - 0s 62us/step - loss: 1.0950 - accuracy: 0.4643\nEpoch 22/200\n56/56 [==============================] - 0s 66us/step - loss: 1.0877 - accuracy: 0.4643\nEpoch 23/200\n56/56 [==============================] - 0s 42us/step - loss: 1.0841 - accuracy: 0.4643\nEpoch 24/200\n56/56 [==============================] - 0s 41us/step - loss: 1.0747 - accuracy: 0.4643\nEpoch 25/200\n56/56 [==============================] - 0s 48us/step - loss: 1.0943 - accuracy: 0.4643\nEpoch 26/200\n56/56 [==============================] - 0s 81us/step - loss: 1.0724 - accuracy: 0.4643\nEpoch 27/200\n56/56 [==============================] - 0s 64us/step - loss: 1.0853 - accuracy: 0.4643\nEpoch 28/200\n56/56 [==============================] - 0s 38us/step - loss: 1.0973 - accuracy: 0.4643\nEpoch 29/200\n56/56 [==============================] - 0s 60us/step - loss: 1.0826 - accuracy: 0.4643\nEpoch 30/200\n56/56 [==============================] - 0s 38us/step - loss: 1.0759 - accuracy: 0.4643\nEpoch 31/200\n56/56 [==============================] - 0s 40us/step - loss: 1.0886 - accuracy: 0.4643\nEpoch 32/200\n56/56 [==============================] - 0s 39us/step - loss: 1.0937 - accuracy: 0.4643\nEpoch 33/200\n56/56 [==============================] - 0s 59us/step - loss: 1.0934 - accuracy: 0.4643\nEpoch 34/200\n56/56 [==============================] - 0s 41us/step - loss: 1.0710 - accuracy: 0.4643\nEpoch 35/200\n56/56 [==============================] - 0s 33us/step - loss: 1.0782 - accuracy: 0.4643\nEpoch 36/200\n56/56 [==============================] - 0s 37us/step - loss: 1.0830 - accuracy: 0.4643\nEpoch 37/200\n56/56 [==============================] - 0s 64us/step - loss: 1.0779 - accuracy: 0.4643\nEpoch 38/200\n56/56 [==============================] - 0s 61us/step - loss: 1.0944 - accuracy: 0.4643\nEpoch 39/200\n56/56 [==============================] - 0s 35us/step - loss: 1.0917 - accuracy: 0.4643\nEpoch 40/200\n56/56 [==============================] - 0s 63us/step - loss: 1.0916 - accuracy: 0.4643\nEpoch 41/200\n56/56 [==============================] - 0s 36us/step - loss: 1.0827 - accuracy: 0.4643\nEpoch 42/200\n56/56 [==============================] - 0s 47us/step - loss: 1.0747 - accuracy: 0.4643\nEpoch 43/200\n56/56 [==============================] - 0s 84us/step - loss: 1.0671 - accuracy: 0.4643\nEpoch 44/200\n56/56 [==============================] - 0s 53us/step - loss: 1.0827 - accuracy: 0.4643\nEpoch 45/200\n56/56 [==============================] - 0s 43us/step - loss: 1.0939 - accuracy: 0.4643\nEpoch 46/200\n56/56 [==============================] - 0s 114us/step - loss: 1.0915 - accuracy: 0.4643\nEpoch 47/200\n56/56 [==============================] - 0s 55us/step - loss: 1.0804 - accuracy: 0.4643\nEpoch 48/200\n56/56 [==============================] - 0s 95us/step - loss: 1.0774 - accuracy: 0.4643\nEpoch 49/200\n56/56 [==============================] - 0s 38us/step - loss: 1.1006 - accuracy: 0.4643\nEpoch 50/200\n56/56 [==============================] - 0s 34us/step - loss: 1.0789 - accuracy: 0.4643\nEpoch 51/200\n56/56 [==============================] - 0s 54us/step - loss: 1.0823 - accuracy: 0.4643\nEpoch 52/200\n56/56 [==============================] - 0s 67us/step - loss: 1.0895 - accuracy: 0.4643\nEpoch 53/200\n56/56 [==============================] - 0s 38us/step - loss: 1.0844 - accuracy: 0.4643\nEpoch 54/200\n56/56 [==============================] - 0s 32us/step - loss: 1.0918 - accuracy: 0.4643\nEpoch 55/200\n56/56 [==============================] - 0s 44us/step - loss: 1.0826 - accuracy: 0.4643\nEpoch 56/200\n56/56 [==============================] - 0s 35us/step - loss: 1.0893 - accuracy: 0.4643\nEpoch 57/200\n56/56 [==============================] - 0s 53us/step - loss: 1.0884 - accuracy: 0.4643\nEpoch 58/200\n56/56 [==============================] - 0s 33us/step - loss: 1.0941 - accuracy: 0.4643\nEpoch 59/200\n56/56 [==============================] - 0s 50us/step - loss: 1.0852 - accuracy: 0.4643\nEpoch 60/200\n56/56 [==============================] - 0s 42us/step - loss: 1.0921 - accuracy: 0.4643\nEpoch 61/200\n56/56 [==============================] - 0s 51us/step - loss: 1.0724 - accuracy: 0.4643\nEpoch 62/200\n56/56 [==============================] - 0s 42us/step - loss: 1.0787 - accuracy: 0.4643\nEpoch 63/200\n56/56 [==============================] - 0s 38us/step - loss: 1.0809 - accuracy: 0.4643\nEpoch 64/200\n56/56 [==============================] - 0s 43us/step - loss: 1.0701 - accuracy: 0.4643\nEpoch 65/200\n56/56 [==============================] - 0s 59us/step - loss: 1.0770 - accuracy: 0.4643\nEpoch 66/200\n56/56 [==============================] - 0s 38us/step - loss: 1.0892 - accuracy: 0.4643\nEpoch 67/200\n56/56 [==============================] - 0s 54us/step - loss: 1.0868 - accuracy: 0.4643\nEpoch 68/200\n56/56 [==============================] - 0s 46us/step - loss: 1.0700 - accuracy: 0.4643\nEpoch 69/200\n56/56 [==============================] - 0s 44us/step - loss: 1.0705 - accuracy: 0.4643\nEpoch 70/200\n56/56 [==============================] - 0s 44us/step - loss: 1.0801 - accuracy: 0.4643\nEpoch 71/200\n56/56 [==============================] - 0s 48us/step - loss: 1.0841 - accuracy: 0.4643\nEpoch 72/200\n56/56 [==============================] - 0s 41us/step - loss: 1.0663 - accuracy: 0.4643\nEpoch 73/200\n56/56 [==============================] - 0s 41us/step - loss: 1.0660 - accuracy: 0.4643\nEpoch 74/200\n56/56 [==============================] - 0s 35us/step - loss: 1.0947 - accuracy: 0.4643\nEpoch 75/200\n56/56 [==============================] - 0s 59us/step - loss: 1.0669 - accuracy: 0.4643\nEpoch 76/200\n56/56 [==============================] - 0s 36us/step - loss: 1.0807 - accuracy: 0.4643\nEpoch 77/200\n56/56 [==============================] - 0s 40us/step - loss: 1.0718 - accuracy: 0.4643\nEpoch 78/200\n56/56 [==============================] - 0s 43us/step - loss: 1.0821 - accuracy: 0.4643\nEpoch 79/200\n56/56 [==============================] - 0s 72us/step - loss: 1.0778 - accuracy: 0.4643\nEpoch 80/200\n56/56 [==============================] - 0s 39us/step - loss: 1.0854 - accuracy: 0.4643\nEpoch 81/200\n56/56 [==============================] - 0s 41us/step - loss: 1.0859 - accuracy: 0.4643\nEpoch 82/200\n"
]
],
[
[
"# Cleaning up data further¶",
"_____no_output_____"
],
[
"## accuracy too low",
"_____no_output_____"
]
],
[
[
"X, y = shuffle(dataset, labels)\ny = to_categorical(y, 3)\nprint(X.shape)\nX[:,:,0] = X[:,:,0] / 720 \nX[:,:,1] = X[:,:,1] / 1280\nX = X[:,:,:2]\nprint(X.shape)\nX = X.reshape(56, 50) \nprint(X.shape)",
"(56, 25, 3)\n(56, 25, 2)\n(56, 50)\n"
],
[
"model = Sequential()\nmodel.add(Dense(128, activation='relu', input_shape=(50,)))\nmodel.add(Dropout(0.5))\nmodel.add(Dense(128, activation='relu'))\nmodel.add(Dropout(0.5))\nmodel.add(Dense(y.shape[1], activation='softmax'))\nmodel.compile(optimizer='Adam',\n loss='categorical_crossentropy',\n metrics=['accuracy'])\nmodel.fit(X, y, epochs=200,batch_size=64)",
"Epoch 1/200\n56/56 [==============================] - 0s 3ms/step - loss: 1.0406 - accuracy: 0.4821\nEpoch 2/200\n56/56 [==============================] - 0s 36us/step - loss: 1.1201 - accuracy: 0.4107\nEpoch 3/200\n56/56 [==============================] - 0s 59us/step - loss: 1.0904 - accuracy: 0.3750\nEpoch 4/200\n56/56 [==============================] - 0s 52us/step - loss: 1.1830 - accuracy: 0.3036\nEpoch 5/200\n56/56 [==============================] - 0s 39us/step - loss: 1.1125 - accuracy: 0.3214\nEpoch 6/200\n56/56 [==============================] - 0s 36us/step - loss: 1.0818 - accuracy: 0.5000\nEpoch 7/200\n56/56 [==============================] - 0s 72us/step - loss: 1.1210 - accuracy: 0.4643\nEpoch 8/200\n56/56 [==============================] - 0s 59us/step - loss: 1.1126 - accuracy: 0.4107\nEpoch 9/200\n56/56 [==============================] - 0s 49us/step - loss: 1.1226 - accuracy: 0.4464\nEpoch 10/200\n56/56 [==============================] - 0s 61us/step - loss: 1.0506 - accuracy: 0.5000\nEpoch 11/200\n56/56 [==============================] - 0s 54us/step - loss: 1.0489 - accuracy: 0.5179\nEpoch 12/200\n56/56 [==============================] - 0s 45us/step - loss: 1.0744 - accuracy: 0.4821\nEpoch 13/200\n56/56 [==============================] - 0s 43us/step - loss: 1.1476 - accuracy: 0.3750\nEpoch 14/200\n56/56 [==============================] - 0s 44us/step - loss: 1.0980 - accuracy: 0.3571\nEpoch 15/200\n56/56 [==============================] - 0s 39us/step - loss: 1.0293 - accuracy: 0.5179\nEpoch 16/200\n56/56 [==============================] - 0s 35us/step - loss: 1.0451 - accuracy: 0.4464\nEpoch 17/200\n56/56 [==============================] - 0s 47us/step - loss: 1.0533 - accuracy: 0.5000\nEpoch 18/200\n56/56 [==============================] - 0s 56us/step - loss: 1.0400 - accuracy: 0.4464\nEpoch 19/200\n56/56 [==============================] - 0s 49us/step - loss: 1.0453 - accuracy: 0.5000\nEpoch 20/200\n56/56 [==============================] - 0s 44us/step - loss: 1.0191 - accuracy: 0.5179\nEpoch 21/200\n56/56 [==============================] - 0s 57us/step - loss: 0.9748 - accuracy: 0.5714\nEpoch 22/200\n56/56 [==============================] - 0s 48us/step - loss: 1.0180 - accuracy: 0.5536\nEpoch 23/200\n56/56 [==============================] - 0s 37us/step - loss: 1.0769 - accuracy: 0.4107\nEpoch 24/200\n56/56 [==============================] - 0s 38us/step - loss: 1.0039 - accuracy: 0.4643\nEpoch 25/200\n56/56 [==============================] - 0s 54us/step - loss: 1.0341 - accuracy: 0.4286\nEpoch 26/200\n56/56 [==============================] - 0s 38us/step - loss: 1.0186 - accuracy: 0.4464\nEpoch 27/200\n56/56 [==============================] - 0s 62us/step - loss: 0.9989 - accuracy: 0.5000\nEpoch 28/200\n56/56 [==============================] - 0s 46us/step - loss: 0.9547 - accuracy: 0.6250\nEpoch 29/200\n56/56 [==============================] - 0s 48us/step - loss: 0.9977 - accuracy: 0.4643\nEpoch 30/200\n56/56 [==============================] - 0s 57us/step - loss: 0.9668 - accuracy: 0.4821\nEpoch 31/200\n56/56 [==============================] - 0s 38us/step - loss: 0.9866 - accuracy: 0.5179\nEpoch 32/200\n56/56 [==============================] - 0s 39us/step - loss: 0.9320 - accuracy: 0.5714\nEpoch 33/200\n56/56 [==============================] - 0s 36us/step - loss: 0.9227 - accuracy: 0.5000\nEpoch 34/200\n56/56 [==============================] - 0s 38us/step - loss: 0.9132 - accuracy: 0.5893\nEpoch 35/200\n56/56 [==============================] - 0s 53us/step - loss: 0.9124 - accuracy: 0.6250\nEpoch 36/200\n56/56 [==============================] - 0s 66us/step - loss: 0.9734 - accuracy: 0.5714\nEpoch 37/200\n56/56 [==============================] - 0s 66us/step - loss: 0.9819 - accuracy: 0.5179\nEpoch 38/200\n56/56 [==============================] - 0s 49us/step - loss: 0.9442 - accuracy: 0.5536\nEpoch 39/200\n56/56 [==============================] - 0s 74us/step - loss: 0.9597 - accuracy: 0.5357\nEpoch 40/200\n56/56 [==============================] - 0s 41us/step - loss: 0.9376 - accuracy: 0.5536\nEpoch 41/200\n56/56 [==============================] - 0s 42us/step - loss: 0.9535 - accuracy: 0.5893\nEpoch 42/200\n56/56 [==============================] - 0s 37us/step - loss: 0.8973 - accuracy: 0.6964\nEpoch 43/200\n56/56 [==============================] - 0s 78us/step - loss: 0.9176 - accuracy: 0.6429\nEpoch 44/200\n56/56 [==============================] - 0s 38us/step - loss: 0.9357 - accuracy: 0.5357\nEpoch 45/200\n56/56 [==============================] - 0s 70us/step - loss: 0.9040 - accuracy: 0.5714\nEpoch 46/200\n56/56 [==============================] - 0s 85us/step - loss: 0.8752 - accuracy: 0.6429\nEpoch 47/200\n56/56 [==============================] - 0s 54us/step - loss: 0.8456 - accuracy: 0.6071\nEpoch 48/200\n56/56 [==============================] - 0s 64us/step - loss: 0.8921 - accuracy: 0.6429\nEpoch 49/200\n56/56 [==============================] - 0s 93us/step - loss: 0.8815 - accuracy: 0.6786\nEpoch 50/200\n56/56 [==============================] - 0s 57us/step - loss: 0.8611 - accuracy: 0.6250\nEpoch 51/200\n56/56 [==============================] - 0s 52us/step - loss: 0.8316 - accuracy: 0.6786\nEpoch 52/200\n56/56 [==============================] - 0s 59us/step - loss: 0.8723 - accuracy: 0.6607\nEpoch 53/200\n56/56 [==============================] - 0s 46us/step - loss: 0.8413 - accuracy: 0.6429\nEpoch 54/200\n56/56 [==============================] - 0s 44us/step - loss: 0.8631 - accuracy: 0.6786\nEpoch 55/200\n56/56 [==============================] - 0s 46us/step - loss: 0.8321 - accuracy: 0.6607\nEpoch 56/200\n56/56 [==============================] - 0s 42us/step - loss: 0.8484 - accuracy: 0.7143\nEpoch 57/200\n56/56 [==============================] - 0s 65us/step - loss: 0.8510 - accuracy: 0.6786\nEpoch 58/200\n56/56 [==============================] - 0s 40us/step - loss: 0.8072 - accuracy: 0.6786\nEpoch 59/200\n56/56 [==============================] - 0s 42us/step - loss: 0.7828 - accuracy: 0.7857\nEpoch 60/200\n56/56 [==============================] - 0s 37us/step - loss: 0.7619 - accuracy: 0.8036\nEpoch 61/200\n56/56 [==============================] - 0s 75us/step - loss: 0.8104 - accuracy: 0.7143\nEpoch 62/200\n56/56 [==============================] - 0s 66us/step - loss: 0.7332 - accuracy: 0.7143\nEpoch 63/200\n56/56 [==============================] - 0s 52us/step - loss: 0.7541 - accuracy: 0.6786\nEpoch 64/200\n56/56 [==============================] - 0s 36us/step - loss: 0.7779 - accuracy: 0.6786\nEpoch 65/200\n56/56 [==============================] - 0s 54us/step - loss: 0.7512 - accuracy: 0.6964\nEpoch 66/200\n56/56 [==============================] - 0s 35us/step - loss: 0.7310 - accuracy: 0.7679\nEpoch 67/200\n56/56 [==============================] - 0s 54us/step - loss: 0.7744 - accuracy: 0.6250\nEpoch 68/200\n56/56 [==============================] - 0s 53us/step - loss: 0.7224 - accuracy: 0.7679\nEpoch 69/200\n56/56 [==============================] - 0s 56us/step - loss: 0.7512 - accuracy: 0.7321\nEpoch 70/200\n56/56 [==============================] - 0s 53us/step - loss: 0.7768 - accuracy: 0.6964\nEpoch 71/200\n56/56 [==============================] - 0s 62us/step - loss: 0.7715 - accuracy: 0.7321\nEpoch 72/200\n56/56 [==============================] - 0s 45us/step - loss: 0.7261 - accuracy: 0.6964\nEpoch 73/200\n56/56 [==============================] - 0s 53us/step - loss: 0.7031 - accuracy: 0.7857\nEpoch 74/200\n56/56 [==============================] - 0s 47us/step - loss: 0.7318 - accuracy: 0.7500\nEpoch 75/200\n56/56 [==============================] - 0s 47us/step - loss: 0.7131 - accuracy: 0.6607\nEpoch 76/200\n56/56 [==============================] - 0s 38us/step - loss: 0.6899 - accuracy: 0.7679\nEpoch 77/200\n56/56 [==============================] - 0s 81us/step - loss: 0.6441 - accuracy: 0.8036\nEpoch 78/200\n56/56 [==============================] - 0s 45us/step - loss: 0.7058 - accuracy: 0.7321\nEpoch 79/200\n56/56 [==============================] - 0s 40us/step - loss: 0.6638 - accuracy: 0.7679\nEpoch 80/200\n56/56 [==============================] - 0s 49us/step - loss: 0.6369 - accuracy: 0.7679\nEpoch 81/200\n56/56 [==============================] - 0s 38us/step - loss: 0.6866 - accuracy: 0.7679\nEpoch 82/200\n56/56 [==============================] - 0s 42us/step - loss: 0.6356 - accuracy: 0.7857\n"
]
],
[
[
"# Adding More Data and Beginning Data Augmentation",
"_____no_output_____"
]
],
[
[
"dabDataset = np.load('data/more-dabs.npy')\ntposeDataset = np.load('data/more-tposes.npy')\notherDataset = np.load('data/more-other.npy')\nlabels1 = np.zeros(len(otherDataset))\nlabels1 = np.append(labels1, np.full((len(dabDataset)), 1))\nlabels1 = np.append(labels1, np.full((len(tposeDataset)), 2))\nprint(labels1)\nprint(\"%i total new samples\" % len(labels1))",
"[0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.\n 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.\n 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0. 0.\n 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.\n 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1. 1.\n 1. 1. 1. 1. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2.\n 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2. 2.\n 2. 2. 2.]\n171 total new samples\n"
],
[
"dataset1 = np.append(otherDataset, dabDataset, axis=0)\ndataset1 = np.append(dataset1, tposeDataset, axis=0)\nX1, y1 = shuffle(dataset1, labels1)\ny1 = to_categorical(y1, 3)\nprint(X1.shape)\nX1[:,:,0] = X1[:,:,0] / 720\nX1[:,:,1] = X1[:,:,1] / 1280\nX1 = X1[:,:,:2]\nprint(X1.shape)\nX1 = X1.reshape(len(X1), 50) \nprint(X1.shape)",
"(171, 25, 3)\n(171, 25, 2)\n(171, 50)\n"
],
[
"model = Sequential()\nmodel.add(Dense(128, activation = 'relu', input_shape = (50,)))\nmodel.add(Dropout(0.5))\nmodel.add(Dense(128, activation = 'relu'))\nmodel.add(Dropout(0.5))\nmodel.add(Dense(y.shape[1], activation = 'softmax'))\nmodel.compile(optimizer = 'Adam',\n loss = 'categorical_crossentropy',\n metrics = ['accuracy'])\nhistory = model.fit(X1, y1, epochs = 200, batch_size = 64)",
"Epoch 1/200\n171/171 [==============================] - 0s 817us/step - loss: 1.1506 - accuracy: 0.3977\nEpoch 2/200\n171/171 [==============================] - 0s 41us/step - loss: 1.1650 - accuracy: 0.3801\nEpoch 3/200\n171/171 [==============================] - 0s 40us/step - loss: 1.1043 - accuracy: 0.3918\nEpoch 4/200\n171/171 [==============================] - 0s 44us/step - loss: 1.1017 - accuracy: 0.3977\nEpoch 5/200\n171/171 [==============================] - 0s 41us/step - loss: 1.0634 - accuracy: 0.3801\nEpoch 6/200\n171/171 [==============================] - 0s 41us/step - loss: 0.9900 - accuracy: 0.5380\nEpoch 7/200\n171/171 [==============================] - 0s 45us/step - loss: 0.9947 - accuracy: 0.4912\nEpoch 8/200\n171/171 [==============================] - 0s 60us/step - loss: 1.0302 - accuracy: 0.4503\nEpoch 9/200\n171/171 [==============================] - 0s 50us/step - loss: 0.9533 - accuracy: 0.5906\nEpoch 10/200\n171/171 [==============================] - 0s 49us/step - loss: 0.9815 - accuracy: 0.5088\nEpoch 11/200\n171/171 [==============================] - 0s 54us/step - loss: 0.9358 - accuracy: 0.5731\nEpoch 12/200\n171/171 [==============================] - 0s 59us/step - loss: 0.9097 - accuracy: 0.5789\nEpoch 13/200\n171/171 [==============================] - 0s 43us/step - loss: 0.9493 - accuracy: 0.5146\nEpoch 14/200\n171/171 [==============================] - 0s 52us/step - loss: 0.8765 - accuracy: 0.6140\nEpoch 15/200\n171/171 [==============================] - 0s 66us/step - loss: 0.8655 - accuracy: 0.6140\nEpoch 16/200\n171/171 [==============================] - 0s 55us/step - loss: 0.8718 - accuracy: 0.6082\nEpoch 17/200\n171/171 [==============================] - 0s 60us/step - loss: 0.8780 - accuracy: 0.5965\nEpoch 18/200\n171/171 [==============================] - 0s 64us/step - loss: 0.8196 - accuracy: 0.6550\nEpoch 19/200\n171/171 [==============================] - 0s 68us/step - loss: 0.7922 - accuracy: 0.6316\nEpoch 20/200\n171/171 [==============================] - 0s 44us/step - loss: 0.7818 - accuracy: 0.6725\nEpoch 21/200\n171/171 [==============================] - 0s 50us/step - loss: 0.7933 - accuracy: 0.6433\nEpoch 22/200\n171/171 [==============================] - 0s 61us/step - loss: 0.7709 - accuracy: 0.6959\nEpoch 23/200\n171/171 [==============================] - 0s 42us/step - loss: 0.7785 - accuracy: 0.6784\nEpoch 24/200\n171/171 [==============================] - 0s 38us/step - loss: 0.7445 - accuracy: 0.7135\nEpoch 25/200\n171/171 [==============================] - 0s 61us/step - loss: 0.7130 - accuracy: 0.7485\nEpoch 26/200\n171/171 [==============================] - 0s 45us/step - loss: 0.7177 - accuracy: 0.7135\nEpoch 27/200\n171/171 [==============================] - 0s 49us/step - loss: 0.7596 - accuracy: 0.6784\nEpoch 28/200\n171/171 [==============================] - 0s 59us/step - loss: 0.7091 - accuracy: 0.7135\nEpoch 29/200\n171/171 [==============================] - 0s 43us/step - loss: 0.6479 - accuracy: 0.7719\nEpoch 30/200\n171/171 [==============================] - 0s 48us/step - loss: 0.6127 - accuracy: 0.8070\nEpoch 31/200\n171/171 [==============================] - 0s 47us/step - loss: 0.6317 - accuracy: 0.7368\nEpoch 32/200\n171/171 [==============================] - 0s 42us/step - loss: 0.6049 - accuracy: 0.7953\nEpoch 33/200\n171/171 [==============================] - 0s 54us/step - loss: 0.6023 - accuracy: 0.7895\nEpoch 34/200\n171/171 [==============================] - 0s 53us/step - loss: 0.5617 - accuracy: 0.7895\nEpoch 35/200\n171/171 [==============================] - 0s 90us/step - loss: 0.6283 - accuracy: 0.7310\nEpoch 36/200\n171/171 [==============================] - 0s 55us/step - loss: 0.5837 - accuracy: 0.7836\nEpoch 37/200\n171/171 [==============================] - 0s 71us/step - loss: 0.5429 - accuracy: 0.8012\nEpoch 38/200\n171/171 [==============================] - 0s 79us/step - loss: 0.5143 - accuracy: 0.8246\nEpoch 39/200\n171/171 [==============================] - 0s 67us/step - loss: 0.5448 - accuracy: 0.8363\nEpoch 40/200\n171/171 [==============================] - 0s 84us/step - loss: 0.4946 - accuracy: 0.8246\nEpoch 41/200\n171/171 [==============================] - 0s 55us/step - loss: 0.4395 - accuracy: 0.8772\nEpoch 42/200\n171/171 [==============================] - 0s 65us/step - loss: 0.4571 - accuracy: 0.8363\nEpoch 43/200\n171/171 [==============================] - 0s 74us/step - loss: 0.4033 - accuracy: 0.8596\nEpoch 44/200\n171/171 [==============================] - 0s 70us/step - loss: 0.4970 - accuracy: 0.8012\nEpoch 45/200\n171/171 [==============================] - 0s 66us/step - loss: 0.3911 - accuracy: 0.8538\nEpoch 46/200\n171/171 [==============================] - 0s 55us/step - loss: 0.4045 - accuracy: 0.8596\nEpoch 47/200\n171/171 [==============================] - 0s 60us/step - loss: 0.3606 - accuracy: 0.9064\nEpoch 48/200\n171/171 [==============================] - 0s 63us/step - loss: 0.4457 - accuracy: 0.8012\nEpoch 49/200\n171/171 [==============================] - 0s 71us/step - loss: 0.3713 - accuracy: 0.8830\nEpoch 50/200\n171/171 [==============================] - 0s 66us/step - loss: 0.4062 - accuracy: 0.8538\nEpoch 51/200\n171/171 [==============================] - 0s 50us/step - loss: 0.3907 - accuracy: 0.8713\nEpoch 52/200\n171/171 [==============================] - 0s 60us/step - loss: 0.3823 - accuracy: 0.8480\nEpoch 53/200\n171/171 [==============================] - 0s 51us/step - loss: 0.3507 - accuracy: 0.9006\nEpoch 54/200\n171/171 [==============================] - 0s 51us/step - loss: 0.3545 - accuracy: 0.8830\nEpoch 55/200\n171/171 [==============================] - 0s 52us/step - loss: 0.3434 - accuracy: 0.9064\nEpoch 56/200\n171/171 [==============================] - 0s 45us/step - loss: 0.3197 - accuracy: 0.9123\nEpoch 57/200\n171/171 [==============================] - 0s 53us/step - loss: 0.2924 - accuracy: 0.8830\nEpoch 58/200\n171/171 [==============================] - 0s 57us/step - loss: 0.3072 - accuracy: 0.9064\nEpoch 59/200\n171/171 [==============================] - 0s 40us/step - loss: 0.2644 - accuracy: 0.9298\nEpoch 60/200\n171/171 [==============================] - 0s 52us/step - loss: 0.2598 - accuracy: 0.9064\nEpoch 61/200\n171/171 [==============================] - 0s 60us/step - loss: 0.2782 - accuracy: 0.9064\nEpoch 62/200\n171/171 [==============================] - 0s 50us/step - loss: 0.2678 - accuracy: 0.9240\nEpoch 63/200\n171/171 [==============================] - 0s 54us/step - loss: 0.2839 - accuracy: 0.9064\nEpoch 64/200\n171/171 [==============================] - 0s 63us/step - loss: 0.3064 - accuracy: 0.8889\nEpoch 65/200\n171/171 [==============================] - 0s 82us/step - loss: 0.2564 - accuracy: 0.9240\nEpoch 66/200\n171/171 [==============================] - 0s 73us/step - loss: 0.2666 - accuracy: 0.8889\nEpoch 67/200\n171/171 [==============================] - 0s 58us/step - loss: 0.2616 - accuracy: 0.9181\nEpoch 68/200\n171/171 [==============================] - 0s 53us/step - loss: 0.2441 - accuracy: 0.9357\nEpoch 69/200\n171/171 [==============================] - 0s 59us/step - loss: 0.2297 - accuracy: 0.9357\nEpoch 70/200\n171/171 [==============================] - 0s 54us/step - loss: 0.2818 - accuracy: 0.9006\nEpoch 71/200\n171/171 [==============================] - 0s 54us/step - loss: 0.2296 - accuracy: 0.9181\nEpoch 72/200\n171/171 [==============================] - 0s 57us/step - loss: 0.1971 - accuracy: 0.9298\nEpoch 73/200\n171/171 [==============================] - 0s 53us/step - loss: 0.1906 - accuracy: 0.9649\nEpoch 74/200\n171/171 [==============================] - 0s 50us/step - loss: 0.2046 - accuracy: 0.9591\nEpoch 75/200\n171/171 [==============================] - 0s 54us/step - loss: 0.1654 - accuracy: 0.9649\nEpoch 76/200\n171/171 [==============================] - 0s 46us/step - loss: 0.2305 - accuracy: 0.9240\nEpoch 77/200\n171/171 [==============================] - 0s 51us/step - loss: 0.1695 - accuracy: 0.9591\nEpoch 78/200\n171/171 [==============================] - 0s 53us/step - loss: 0.1804 - accuracy: 0.9532\nEpoch 79/200\n171/171 [==============================] - 0s 49us/step - loss: 0.2005 - accuracy: 0.9474\nEpoch 80/200\n"
],
[
"model.predict_classes(X)",
"_____no_output_____"
],
[
"X",
"_____no_output_____"
],
[
"y",
"_____no_output_____"
],
[
"model.test_on_batch(X, y)",
"_____no_output_____"
],
[
"model.save('data/dab-tpose-other.h5')",
"_____no_output_____"
],
[
"import keras\nmodello = keras.models.load_model('data/dab-tpose-other.h5')",
"_____no_output_____"
],
[
"dabDataset = np.load('data/test-dabs.npy')\ndabDataset[:,:,0] = dabDataset[:,:,0] / 720\ndabDataset[:,:,1] = dabDataset[:,:,1] / 1280\ndabDataset = dabDataset[:,:,:2]\ndabDataset = dabDataset.reshape(len(dabDataset), 50)\nmodello.predict_classes(dabDataset)",
"_____no_output_____"
]
],
[
[
"# 預測結果全是1,從前面我們設定的數字,1代表dabs,預測成功。",
"_____no_output_____"
],
[
"# 關於專案分工的部分預計資料前處理和使用模型交給另外兩個人,但費勁了洪荒之力還是做不出來,最後放棄做那些部分,剩下上訓練模型的部分。",
"_____no_output_____"
],
[
"# 作者在他自己的blog上發這篇(https://www.makeartwithpython.com/blog/dab-and-tpose-controlled-lights/,裡頭提到了「All of the code, models, and training data are freely available on Github.」,可能在是否能公開發表上有模擬地帶,下次會注意授權問題,並且先寄信詢問。\n# 模型是做好了,但卻無法使用,由於導入pyopenpose的不成功,無法在python裡呼叫openpose來取得測試資料,沒辦法實際應用。",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
]
] |
4ab4f35369b6f6c33d7978d915b4fcb684d72bef
| 16,056 |
ipynb
|
Jupyter Notebook
|
docs/Examples/High Molecular Weight Petroleum Pseudocomponents.ipynb
|
andr1976/thermo
|
42d10b3702373aacc88167d4046ea9af92abd570
|
[
"MIT"
] | null | null | null |
docs/Examples/High Molecular Weight Petroleum Pseudocomponents.ipynb
|
andr1976/thermo
|
42d10b3702373aacc88167d4046ea9af92abd570
|
[
"MIT"
] | null | null | null |
docs/Examples/High Molecular Weight Petroleum Pseudocomponents.ipynb
|
andr1976/thermo
|
42d10b3702373aacc88167d4046ea9af92abd570
|
[
"MIT"
] | null | null | null | 37.426573 | 342 | 0.53575 |
[
[
[
"# High Molecular Weight Petroleum Pseudocomponents",
"_____no_output_____"
],
[
"Thermo is a general phase equilibrium engine, and if the user provides enough properties for the components, there is no issue adding your own components. In this basic example below, a made-up extended gas analysis is used to specify a gas consisting of the standard real components and three heavier fractions, C10+, C12+ and C15+.\n\nA bare minimum of basic properties are estimated using the Kesler-Lee method (1976), and the estimated fraction molecular weights are turned into atomic compositions. The heat capacities of each pseudocomponent is found with\nthe similarity variable concept of Lastovka and Shaw (2013) based on atomic composition.\n\nThis example ends with calculating a flash at 270 Kelvin and 1 bar.",
"_____no_output_____"
]
],
[
[
"from math import log, exp\nimport numpy as np\nfrom scipy.constants import psi\nfrom thermo import *\nfrom chemicals import *\n\ndef Tc_Kesler_Lee_SG_Tb(SG, Tb):\n r'''Estimates critical temperature of a hydrocarbon compound or petroleum\n fraction using only its specific gravity and boiling point, from\n [1]_ as presented in [2]_.\n\n .. math::\n T_c = 341.7 + 811.1SG + [0.4244 + 0.1174SG]T_b\n + \\frac{[0.4669 - 3.26238SG]10^5}{T_b}\n\n Parameters\n ----------\n SG : float\n Specific gravity of the fluid at 60 degrees Farenheight [-]\n Tb : float\n Boiling point the fluid [K]\n\n Returns\n -------\n Tc : float\n Estimated critical temperature [K]\n\n Notes\n -----\n Model shows predictions for Tc, Pc, MW, and omega.\n Original units in degrees Rankine.\n\n Examples\n --------\n Example 2.2 from [2]_, but with K instead of R.\n\n >>> Tc_Kesler_Lee_SG_Tb(0.7365, 365.555)\n 545.0124354151242\n\n References\n ----------\n .. [1] Kesler, M. G., and B. I. Lee. \"Improve Prediction of Enthalpy of\n Fractions.\" Hydrocarbon Processing (March 1976): 153-158.\n .. [2] Ahmed, Tarek H. Equations of State and PVT Analysis: Applications\n for Improved Reservoir Modeling. Gulf Pub., 2007.\n '''\n Tb = 9/5.*Tb # K to R\n Tc = 341.7 + 811.1*SG + (0.4244 + 0.1174*SG)*Tb + ((0.4669 - 3.26238*SG)*1E5)/Tb\n Tc = 5/9.*Tc # R to K\n return Tc\n\ndef Pc_Kesler_Lee_SG_Tb(SG, Tb):\n r'''Estimates critical pressure of a hydrocarbon compound or petroleum\n fraction using only its specific gravity and boiling point, from\n [1]_ as presented in [2]_.\n\n .. math::\n \\ln(P_c) = 8.3634 - \\frac{0.0566}{SG} - \\left[0.24244 + \\frac{2.2898}\n {SG} + \\frac{0.11857}{SG^2}\\right]10^{-3}T_b\n + \\left[1.4685 + \\frac{3.648}{SG} + \\frac{0.47227}{SG^2}\\right]\n 10^{-7}T_b^2-\\left[0.42019 + \\frac{1.6977}{SG^2}\\right]10^{-10}T_b^3\n\n Parameters\n ----------\n SG : float\n Specific gravity of the fluid at 60 degrees Farenheight [-]\n Tb : float\n Boiling point the fluid [K]\n\n Returns\n -------\n Pc : float\n Estimated critical pressure [Pa]\n\n Notes\n -----\n Model shows predictions for Tc, Pc, MW, and omega.\n Original units in degrees Rankine and psi.\n\n Examples\n --------\n Example 2.2 from [2]_, but with K instead of R and Pa instead of psi.\n\n >>> Pc_Kesler_Lee_SG_Tb(0.7365, 365.555)\n 3238323.346840464\n\n References\n ----------\n .. [1] Kesler, M. G., and B. I. Lee. \"Improve Prediction of Enthalpy of\n Fractions.\" Hydrocarbon Processing (March 1976): 153-158.\n .. [2] Ahmed, Tarek H. Equations of State and PVT Analysis: Applications\n for Improved Reservoir Modeling. Gulf Pub., 2007.\n '''\n Tb = 9/5.*Tb # K to R\n Pc = exp(8.3634 - 0.0566/SG - (0.24244 + 2.2898/SG + 0.11857/SG**2)*1E-3*Tb\n + (1.4685 + 3.648/SG + 0.47227/SG**2)*1E-7*Tb**2\n -(0.42019 + 1.6977/SG**2)*1E-10*Tb**3)\n Pc = Pc*psi\n return Pc\n\ndef MW_Kesler_Lee_SG_Tb(SG, Tb):\n r'''Estimates molecular weight of a hydrocarbon compound or petroleum\n fraction using only its specific gravity and boiling point, from\n [1]_ as presented in [2]_.\n\n .. math::\n MW = -12272.6 + 9486.4SG + [4.6523 - 3.3287SG]T_b + [1-0.77084SG\n - 0.02058SG^2]\\left[1.3437 - \\frac{720.79}{T_b}\\right]\\frac{10^7}{T_b}\n + [1-0.80882SG + 0.02226SG^2][1.8828 - \\frac{181.98}{T_b}]\n \\frac{10^{12}}{T_b^3}\n\n Parameters\n ----------\n SG : float\n Specific gravity of the fluid at 60 degrees Farenheight [-]\n Tb : float\n Boiling point the fluid [K]\n\n Returns\n -------\n MW : float\n Estimated molecular weight [g/mol]\n\n Notes\n -----\n Model shows predictions for Tc, Pc, MW, and omega.\n Original units in degrees Rankine.\n\n Examples\n --------\n Example 2.2 from [2]_, but with K instead of R and Pa instead of psi.\n\n >>> MW_Kesler_Lee_SG_Tb(0.7365, 365.555)\n 98.70887589833501\n\n References\n ----------\n .. [1] Kesler, M. G., and B. I. Lee. \"Improve Prediction of Enthalpy of\n Fractions.\" Hydrocarbon Processing (March 1976): 153-158.\n .. [2] Ahmed, Tarek H. Equations of State and PVT Analysis: Applications\n for Improved Reservoir Modeling. Gulf Pub., 2007.\n '''\n Tb = 9/5.*Tb # K to R\n MW = (-12272.6 + 9486.4*SG + (4.6523 - 3.3287*SG)*Tb + (1.-0.77084*SG - 0.02058*SG**2)*\n (1.3437 - 720.79/Tb)*1E7/Tb + (1.-0.80882*SG + 0.02226*SG**2)*\n (1.8828 - 181.98/Tb)*1E12/Tb**3)\n return MW\n\ndef omega_Kesler_Lee_SG_Tb_Tc_Pc(SG, Tb, Tc=None, Pc=None):\n r'''Estimates accentric factor of a hydrocarbon compound or petroleum\n fraction using only its specific gravity and boiling point, from\n [1]_ as presented in [2]_. If Tc and Pc are provided, the Kesler-Lee\n routines for estimating them are not used.\n\n For Tbr > 0.8:\n .. math::\n \\omega = -7.904 + 0.1352K - 0.007465K^2 + 8.359T_{br}\n + ([1.408-0.01063K]/T_{br})\n\n Otherwise:\n .. math::\n \\omega = \\frac{-\\ln\\frac{P_c}{14.7} - 5.92714 + \\frac{6.09648}{T_{br}}\n + 1.28862\\ln T_{br} - 0.169347T_{br}^6}{15.2518 - \\frac{15.6875}{T_{br}}\n - 13.4721\\ln T_{br} + 0.43577T_{br}^6}\n\n K = \\frac{T_b^{1/3}}{SG}\n\n T_{br} = \\frac{T_b}{T_c}\n\n Parameters\n ----------\n SG : float\n Specific gravity of the fluid at 60 degrees Farenheight [-]\n Tb : float\n Boiling point the fluid [K]\n Tc : float, optional\n Estimated critical temperature [K]\n Pc : float, optional\n Estimated critical pressure [Pa]\n\n Returns\n -------\n omega : float\n Acentric factor [-]\n\n Notes\n -----\n Model shows predictions for Tc, Pc, MW, and omega.\n Original units in degrees Rankine and psi.\n\n Examples\n --------\n Example 2.2 from [2]_, but with K instead of R and Pa instead of psi.\n\n >>> omega_Kesler_Lee_SG_Tb_Tc_Pc(0.7365, 365.555, 545.012, 3238323.)\n 0.306392118159797\n\n References\n ----------\n .. [1] Kesler, M. G., and B. I. Lee. \"Improve Prediction of Enthalpy of\n Fractions.\" Hydrocarbon Processing (March 1976): 153-158.\n .. [2] Ahmed, Tarek H. Equations of State and PVT Analysis: Applications\n for Improved Reservoir Modeling. Gulf Pub., 2007.\n '''\n if Tc is None:\n Tc = Tc_Kesler_Lee_SG_Tb(SG, Tb)\n if Pc is None:\n Pc = Pc_Kesler_Lee_SG_Tb(SG, Tb)\n Tb = 9/5.*Tb # K to R\n Tc = 9/5.*Tc # K to R\n K = Tb**(1/3.)/SG\n Tbr = Tb/Tc\n if Tbr > 0.8:\n omega = -7.904 + 0.1352*K - 0.007465*K**2 + 8.359*Tbr + ((1.408-0.01063*K)/Tbr)\n else:\n omega = ((-log(Pc/101325.) - 5.92714 + 6.09648/Tbr + 1.28862*log(Tbr)\n - 0.169347*Tbr**6) / (15.2518 - 15.6875/Tbr - 13.4721*log(Tbr) +0.43577*Tbr**6))\n return omega\n",
"_____no_output_____"
],
[
"# Basic composition and names. All pure component properties are obtained from Chemicals and Thermo.\npure_constants = ChemicalConstantsPackage.constants_from_IDs(\n ['water', 'hydrogen', 'helium', 'nitrogen', 'carbon dioxide', 'hydrogen sulfide', 'methane',\n 'ethane', 'propane', 'isobutane', 'n-butane', 'isopentane', 'n-pentane', 'hexane', \n 'heptane', 'octane', 'nonane'])\npure_fractions = [.02, .00005, .00018, .009, .02, .002, .82, .08, .031,\n .009, .0035, .0033, .0003, .0007, .0004, .00005, .00002]",
"_____no_output_____"
],
[
"pseudo_names = ['C10-C11', 'C12-C14', 'C15+']\npseudo_carbon_numbers = [10.35, 12.5, 16.9]\npseudo_SGs = [.73, .76, .775] # Specific gravity values are based of the alkane series\npseudo_Tbs = [447, 526, 589] \n\n# Using the estimation methods defined earlier, we obtain some critical properties\npseudo_Tcs = [Tc_Kesler_Lee_SG_Tb(SG, Tb) for SG, Tb in zip(pseudo_SGs, pseudo_Tbs)]\npseudo_Pcs = [Pc_Kesler_Lee_SG_Tb(SG, Tb) for SG, Tb in zip(pseudo_SGs, pseudo_Tbs)]\npseudo_MWs = [MW_Kesler_Lee_SG_Tb(SG, Tb) for SG, Tb in zip(pseudo_SGs, pseudo_Tbs)]\npseudo_omegas = [omega_Kesler_Lee_SG_Tb_Tc_Pc(SG, Tb) for SG, Tb in zip(pseudo_SGs, pseudo_Tbs)]\n# Estimate the hydroen counts\nhydrogen_counts = [(MW - C*periodic_table.C.MW)/periodic_table.H.MW \n for C, MW in zip(pseudo_carbon_numbers, pseudo_MWs)]\n# Get the atomic compositions\npseudo_atoms = [{'C': C, 'H': H} for C, H in zip(pseudo_carbon_numbers, hydrogen_counts)]\n# Calculate the similarity variable of each species\nsimilarity_variables = [similarity_variable(atoms=atoms) for atoms in pseudo_atoms]\n\npseudo_fractions = [.0003, .00015, .00005]",
"_____no_output_____"
],
[
"pseudos = ChemicalConstantsPackage(names=pseudo_names, MWs=pseudo_MWs, Tbs=pseudo_Tbs,\n atomss=pseudo_atoms,\n Tcs=pseudo_Tcs, Pcs=pseudo_Pcs, omegas=pseudo_omegas,\n similarity_variables=similarity_variables)\n# Add the pure components and the pseudocomponents to create a new package of constant values\n# which will be used by the phase and flash objects\nconstants = pure_constants + pseudos\n# Obtain the temperature and pressure dependent objects\nproperties = PropertyCorrelationsPackage(constants=constants)\n# This is the feed composition\nzs = normalize(pure_fractions + pseudo_fractions)\nT = 270 # K\nP = 1e5 # bar",
"_____no_output_____"
],
[
"kijs = np.zeros((constants.N, constants.N)).tolist() # kijs left as zero in this example\neos_kwargs = dict(Tcs=constants.Tcs, Pcs=constants.Pcs, omegas=constants.omegas, kijs=kijs)\n\n# The API SRK equation of state is used, but other cubic equations of state can be uesd instead\ngas = CEOSGas(APISRKMIX, eos_kwargs, HeatCapacityGases=properties.HeatCapacityGases, T=T, P=P, zs=zs)\nliq = CEOSLiquid(APISRKMIX, eos_kwargs, HeatCapacityGases=properties.HeatCapacityGases, T=T, P=P, zs=zs)\nliq2 = CEOSLiquid(APISRKMIX, eos_kwargs, HeatCapacityGases=properties.HeatCapacityGases, T=T, P=P, zs=zs)\nphase_list = [gas, liq, liq]\n\n# Set up the three phase flash engine\nflashN = FlashVLN(constants, properties, liquids=[liq, liq2], gas=gas)",
"_____no_output_____"
],
[
"# Do the flash, and get some properties\nres = flashN.flash(T=T, P=P, zs=zs)\nres.phase_count, res.gas_beta, res.liquids_betas",
"_____no_output_____"
],
[
"res.H(), res.Cp_mass(), res.MW(), res.gas.mu(), res.gas.k()",
"_____no_output_____"
],
[
"res.heaviest_liquid.rho_mass(), res.lightest_liquid.rho_mass()",
"_____no_output_____"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab4f65d201d90d61867008279a6f7f16331a194
| 33,640 |
ipynb
|
Jupyter Notebook
|
module3-nosql-and-document-oriented-databases/mongo_db_example.ipynb
|
ianforrest11/DS-Unit-3-Sprint-2-SQL-and-Databases
|
f578b049dd49e1243fe42969500926644818a7bf
|
[
"MIT"
] | null | null | null |
module3-nosql-and-document-oriented-databases/mongo_db_example.ipynb
|
ianforrest11/DS-Unit-3-Sprint-2-SQL-and-Databases
|
f578b049dd49e1243fe42969500926644818a7bf
|
[
"MIT"
] | null | null | null |
module3-nosql-and-document-oriented-databases/mongo_db_example.ipynb
|
ianforrest11/DS-Unit-3-Sprint-2-SQL-and-Databases
|
f578b049dd49e1243fe42969500926644818a7bf
|
[
"MIT"
] | null | null | null | 31.409897 | 1,550 | 0.519084 |
[
[
[
"import pymongo",
"_____no_output_____"
],
[
"# using 3.4 connection string for clarity/workingness\nclient = pymongo.MongoClient(\"mongodb://admin:[email protected]:27017,cluster0-shard-00-01-ooj9o.mongodb.net:27017,cluster0-shard-00-02-ooj9o.mongodb.net:27017/test?ssl=true&replicaSet=Cluster0-shard-0&authSource=admin&retryWrites=true&w=majority\")\ndb = client.test",
"_____no_output_____"
],
[
"db",
"_____no_output_____"
],
[
"client.nodes",
"_____no_output_____"
],
[
"dir(db.test)",
"_____no_output_____"
],
[
"help(db.test.insert_many)",
"Help on method insert_many in module pymongo.collection:\n\ninsert_many(documents, ordered=True, bypass_document_validation=False, session=None) method of pymongo.collection.Collection instance\n Insert an iterable of documents.\n \n >>> db.test.count_documents({})\n 0\n >>> result = db.test.insert_many([{'x': i} for i in range(2)])\n >>> result.inserted_ids\n [ObjectId('54f113fffba522406c9cc20e'), ObjectId('54f113fffba522406c9cc20f')]\n >>> db.test.count_documents({})\n 2\n \n :Parameters:\n - `documents`: A iterable of documents to insert.\n - `ordered` (optional): If ``True`` (the default) documents will be\n inserted on the server serially, in the order provided. If an error\n occurs all remaining inserts are aborted. If ``False``, documents\n will be inserted on the server in arbitrary order, possibly in\n parallel, and all document inserts will be attempted.\n - `bypass_document_validation`: (optional) If ``True``, allows the\n write to opt-out of document level validation. Default is\n ``False``.\n - `session` (optional): a\n :class:`~pymongo.client_session.ClientSession`.\n \n :Returns:\n An instance of :class:`~pymongo.results.InsertManyResult`.\n \n .. seealso:: :ref:`writes-and-ids`\n \n .. note:: `bypass_document_validation` requires server version\n **>= 3.2**\n \n .. versionchanged:: 3.6\n Added ``session`` parameter.\n \n .. versionchanged:: 3.2\n Added bypass_document_validation support\n \n .. versionadded:: 3.0\n\n"
],
[
"db.test.count_documents({'x':1})",
"_____no_output_____"
],
[
"db.test.insert_one({'x':1})",
"_____no_output_____"
],
[
"db.test.count_documents({'x':1})",
"_____no_output_____"
],
[
"db.test.insert_one({'x':1})",
"_____no_output_____"
],
[
"db.test.count_documents({'x':1})",
"_____no_output_____"
],
[
"db.test.find_one({'x':1})",
"_____no_output_____"
],
[
"curs = db.test.find({'x':1})\n# gives us SQL cursor",
"_____no_output_____"
],
[
"dir(curs)",
"_____no_output_____"
],
[
"list(curs)",
"_____no_output_____"
],
[
"ian_doc = {\n 'favorite_animal': ['Red Panda', 'Firefox']\n}\n\ndodd_doc = {\n 'favorite_animal': 'Dog',\n 'favorite_color': 'Ocean Blue'\n}\n\nfor_doc = {\n 'favorite_animal': 'Whale'\n}\n\njul_doc = {\n 'favorite_animal': 'Zebra'\n}",
"_____no_output_____"
],
[
"type(jul_doc)",
"_____no_output_____"
],
[
"db.test.insert_many([ian_doc, dodd_doc, for_doc, jul_doc])",
"_____no_output_____"
],
[
"# similar to SQL fetchall command/ SELECT *\nlist(db.test.find())",
"_____no_output_____"
],
[
"more_docs = []\nfor i in range(10):\n doc = {'even': i % 2 ==0}\n doc['value'] = i\n more_docs.append(doc)\n\nmore_docs",
"_____no_output_____"
],
[
"db.test.insert_many(more_docs)",
"_____no_output_____"
],
[
"list(db.test.find({'even':True}))",
"_____no_output_____"
],
[
"list(db.test.find({'even':False}))",
"_____no_output_____"
],
[
"# CRUD applications = create, read, update, delete\nlist(db.test.find({'favorite_animal': 'Dog'}))",
"_____no_output_____"
],
[
"db.test.update_one({'value': 4}, \n {'$inc': {'value': 5}})",
"_____no_output_____"
],
[
"list(db.test.find())",
"_____no_output_____"
],
[
"db.test.update_many({'even': True}, \n {'$inc': {'value': 10}})\n\nlist(db.test.find({'even': True}))",
"_____no_output_____"
],
[
"db.test.delete_many({'even': False})\nlist(db.test.find())",
"_____no_output_____"
],
[
"rpg_character = (1, 'King Bob', 10, 3, 0, 0, 0)",
"_____no_output_____"
],
[
"type(rpg_character)",
"_____no_output_____"
],
[
"db.test.insert_one(rpg_character)",
"_____no_output_____"
],
[
"db.test.insert_one({'rpg_character': rpg_character})",
"_____no_output_____"
],
[
"db.test.find_one({'rpg_character': rpg_character})",
"_____no_output_____"
],
[
"db.test.insert_one({\n 'sql_id': rpg_character[0],\n 'name': rpg_character[1],\n 'hp': rpg_character[2],\n 'level': rpg_character[3]\n})",
"_____no_output_____"
],
[
"list(db.test.find())",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab4f71334b73cb11e851cb14070428abfaae1ad
| 9,389 |
ipynb
|
Jupyter Notebook
|
tutorials/1-skimage_intro.ipynb
|
mbeyeler/2018-neurohack-cv
|
5d69f7b55dc148e10bfeaff843eca085a188cd4c
|
[
"MIT"
] | 3 |
2018-08-05T06:10:00.000Z
|
2021-10-06T21:41:17.000Z
|
tutorials/1-skimage_intro.ipynb
|
mbeyeler/2018-neurohack-skimage
|
5d69f7b55dc148e10bfeaff843eca085a188cd4c
|
[
"MIT"
] | null | null | null |
tutorials/1-skimage_intro.ipynb
|
mbeyeler/2018-neurohack-skimage
|
5d69f7b55dc148e10bfeaff843eca085a188cd4c
|
[
"MIT"
] | null | null | null | 24.970745 | 300 | 0.544999 |
[
[
[
"To participate, you'll need to git clone (or download the .zip from GitHub):\n\nhttps://github.com/mbeyeler/2018-neurohack-skimage\n\nYou can do that in git using:\n\n git clone https://github.com/mbeyeler/2018-neurohack-skimage\n\nIf you have already cloned the material, please issue `git pull` now and reload the notebook to ensure that you have the latest updates.",
"_____no_output_____"
],
[
"# Tutorial 1: Image Manipulation\n\nThis tutorial was adapted from https://github.com/scikit-image/skimage-tutorials/blob/master/lectures/00_images_are_arrays.ipynb.",
"_____no_output_____"
]
],
[
[
"%matplotlib inline",
"_____no_output_____"
]
],
[
[
"## Images are NumPy arrays",
"_____no_output_____"
],
[
"Images are represented in ``scikit-image`` using standard ``numpy`` arrays. This allows maximum inter-operability with other libraries in the scientific Python ecosystem, such as ``matplotlib`` and ``scipy``.\n\nLet's see how to build a grayscale image as a 2D array:",
"_____no_output_____"
]
],
[
[
"import numpy as np\nfrom matplotlib import pyplot as plt\n\nrandom_image = np.random.random([500, 500])\n\nplt.imshow(random_image, cmap='gray')\nplt.colorbar();",
"_____no_output_____"
]
],
[
[
"The same holds for \"real-world\" images:",
"_____no_output_____"
]
],
[
[
"from skimage import data\n\ncoins = data.coins()\n\nprint('Type:', type(coins))\nprint('dtype:', coins.dtype)\nprint('shape:', coins.shape)\n\nplt.imshow(coins, cmap='gray');",
"_____no_output_____"
]
],
[
[
"A color image is a 3D array, where the last dimension has size 3 and represents the red, green, and blue channels:",
"_____no_output_____"
]
],
[
[
"cat = data.chelsea()\nprint(\"Shape:\", cat.shape)\nprint(\"Values min/max:\", cat.min(), cat.max())\n\nplt.imshow(cat);",
"_____no_output_____"
]
],
[
[
"These are *just NumPy arrays*. E.g., we can make a red square by using standard array slicing and manipulation:",
"_____no_output_____"
]
],
[
[
"cat[10:110, 10:110, :] = [255, 0, 0] # [red, green, blue]\nplt.imshow(cat);",
"_____no_output_____"
]
],
[
[
"Images can also include transparent regions by adding a 4th dimension, called an *alpha layer*.",
"_____no_output_____"
],
[
"### Other shapes, and their meanings\n\n|Image type|Coordinates|\n|:---|:---|\n|2D grayscale|(row, column)|\n|2D multichannel|(row, column, channel)|\n|3D grayscale (or volumetric) |(plane, row, column)|\n|3D multichannel|(plane, row, column, channel)|\n\n### Data types and image values\n\nIn literature, one finds different conventions for representing image values:\n\n```\n 0 - 255 where 0 is black, 255 is white\n 0 - 1 where 0 is black, 1 is white\n```\n\n``scikit-image`` supports both conventions--the choice is determined by the\ndata-type of the array.\n\nE.g., here, I generate two valid images:",
"_____no_output_____"
],
[
"## Displaying images using matplotlib",
"_____no_output_____"
]
],
[
[
"from skimage import data\n\nimg0 = data.chelsea()\nimg1 = data.rocket()",
"_____no_output_____"
],
[
"import matplotlib.pyplot as plt\n\nf, (ax0, ax1) = plt.subplots(ncols=2, figsize=(20, 10))\n\nax0.imshow(img0)\nax0.set_title('Cat', fontsize=18)\nax0.axis('off')\n\nax1.imshow(img1)\nax1.set_title('Rocket', fontsize=18)\nax1.set_xlabel(r'Launching position $\\alpha=320$')\n\nax1.vlines([202, 450], 0, img1.shape[0], colors='white', linewidth=3, label='Side tower position')\n\nax1.legend();",
"_____no_output_____"
]
],
[
[
"## Drawing",
"_____no_output_____"
]
],
[
[
"from skimage import draw\n\n# Draw a circle with radius 50 at (200, 150):\nr, c = draw.circle(200, 150, 50)\n# Change only the green channel:\nimg1[r, c, 1] = 255\n\nplt.imshow(img1)",
"_____no_output_____"
]
],
[
[
"For more on plotting, see the [Matplotlib documentation](https://matplotlib.org/gallery/index.html#images-contours-and-fields) and [pyplot API](https://matplotlib.org/api/pyplot_summary.html).",
"_____no_output_____"
],
[
"## Image I/O\n\nMostly, we won't be using input images from the scikit-image example data sets. Those images are typically stored in JPEG or PNG format. Since scikit-image operates on NumPy arrays, *any* image reader library that provides arrays will do. Options include imageio, matplotlib, pillow, etc.\n\nscikit-image conveniently wraps many of these in the `io` submodule, and will use whichever of the libraries mentioned above are installed:",
"_____no_output_____"
]
],
[
[
"from skimage import io\n\nimage = io.imread('../img/skimage-logo.png')\n\nprint(type(image))\nprint(image.dtype)\nprint(image.shape)\nprint(image.min(), image.max())\n\nplt.imshow(image);",
"_____no_output_____"
]
],
[
[
"We also have the ability to load multiple images, or multi-layer TIFF images:",
"_____no_output_____"
]
],
[
[
"ic = io.ImageCollection('../img/*.jpg')\n\nprint('Type:', type(ic))\n\nic.files",
"_____no_output_____"
]
],
[
[
"# Exercise: Visualizing RGB channels\n\nDisplay the different color channels of the image along (each as a gray-scale image). Start with the following template:",
"_____no_output_____"
]
],
[
[
"# --- read in the image ---\n\nimage = io.imread('../img/skimage-logo.png')\n\n# --- assign each color channel to a different variable ---\n\nr = ...\ng = ...\nb = ...\n\n# --- display the image and r, g, b channels ---\n\nf, axes = plt.subplots(1, 4, figsize=(16, 5))\n\nfor ax in axes:\n ax.axis('off')\n\n(ax_r, ax_g, ax_b, ax_color) = axes\n \nax_r.imshow(r, cmap='gray')\nax_r.set_title('red channel')\n\nax_g.imshow(g, cmap='gray')\nax_g.set_title('green channel')\n\nax_b.imshow(b, cmap='gray')\nax_b.set_title('blue channel')\n\n# --- Here, we stack the R, G, and B layers again\n# to form a color image ---\nax_color.imshow(np.stack([r, g, b], axis=2))\nax_color.set_title('all channels');",
"_____no_output_____"
]
],
[
[
"Now, take a look at the following R, G, and B channels. How would their combination look? (Write some code to confirm your intuition.)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
4ab504acb6b0e99a67dff38892273fe4eedc9971
| 14,259 |
ipynb
|
Jupyter Notebook
|
rojenkin.ipynb
|
rojenkin/astr-119_hw-5
|
52ead50ab2a390b88bf9b65ebd3a9e1d6461274c
|
[
"MIT"
] | null | null | null |
rojenkin.ipynb
|
rojenkin/astr-119_hw-5
|
52ead50ab2a390b88bf9b65ebd3a9e1d6461274c
|
[
"MIT"
] | 1 |
2018-11-20T17:45:55.000Z
|
2018-12-09T03:19:47.000Z
|
rojenkin.ipynb
|
rojenkin/astr-119_hw-5
|
52ead50ab2a390b88bf9b65ebd3a9e1d6461274c
|
[
"MIT"
] | null | null | null | 27.741245 | 119 | 0.436847 |
[
[
[
"Grade 5/7\n\nNo implementation of Cash-Karp coefficients\n\nImplementation of adaptive step is not set by Cash Karp algorithm",
"_____no_output_____"
],
[
"## Create a notebook to perform Runge-Kutta integration for multiple coupled variables.",
"_____no_output_____"
]
],
[
[
"%matplotlib inline\nimport matplotlib.pyplot as plt\nimport numpy as np",
"_____no_output_____"
]
],
[
[
"This cell and the one following it are not a requirement, it is only for looks",
"_____no_output_____"
]
],
[
[
"#use colors.subclass(or command; e.g bold).colorname to print\n#examples: print(colors.bold, colors.fg.blue, \"this will be bold and blue\")\n#everything after this will have that format until the following command\n#is given: print(colors.reset, \"now, this text will be normal\")\n\nclass colors:\n reset='\\033[0m' #reset all colors with colors.reset\n bold='\\033[01m'\n underline='\\033[04m'\n strikethrough='\\033[09m'\n reverse='\\033[07m'\n class fg: #foreground subclass\n black='\\033[30m'\n red='\\033[31m'\n green='\\033[32m'\n orange='\\033[33m'\n blue='\\033[34m'\n purple='\\033[35m'\n cyan='\\033[36m'\n lightgrey='\\033[37m'\n darkgrey='\\033[90m'\n lightred='\\033[91m'\n lightgreen='\\033[92m'\n yellow='\\033[93m'\n lightblue='\\033[94m'\n pink='\\033[95m'\n lightcyan='\\033[96m'\n class bg: #background subclass\n black='\\033[40m'\n red='\\033[41m'\n green='\\033[42m'\n orange='\\033[43m'\n blue='\\033[44m'\n purple='\\033[45m'\n cyan='\\033[46m'\n lightgrey='\\033[47m'",
"_____no_output_____"
]
],
[
[
"The above code was provided by https://www.geeksforgeeks.org/print-colors-python-terminal/",
"_____no_output_____"
],
[
"### Define our coupled derivatives to integrate",
"_____no_output_____"
]
],
[
[
"def dydx(x,y):\n \n #set the derivatives\n \n #so we can write \n #dydx = z\n #dzdx = -y\n \n #we will set y = y[0]\n #we will set z = y[1]\n \n #declare an array\n y_derivs = np.zeros(2)\n \n #set dydx = z\n y_derivs[0] = y[1]\n \n #set dzdx = -y\n y_derivs[1] = -1*y[0]\n \n #here we have to return an array with dydx\n return y_derivs",
"_____no_output_____"
]
],
[
[
"### Define the 4th order RK method",
"_____no_output_____"
]
],
[
[
"def rk4_mv_core(dydx,xi,yi,nv,h):\n \n #declare k? arrays; (? is a wildcard, used for k1,k2,...,kn)\n k1=np.zeros(nv)\n k2=np.zeros(nv)\n k3=np.zeros(nv)\n k4=np.zeros(nv)\n \n #define x at 1/2 step\n x_ipoh = xi + 0.5*h\n \n #define x at 1 step\n x_ipo = xi + h\n \n #declare a temp y array\n y_temp = np.zeros(nv)\n \n #get k1 values\n y_derivs = dydx(xi,yi)\n k1[:] = h*y_derivs[:]\n \n #get k2 values\n y_temp[:] = yi[:] + 0.5*k1[:]\n y_derivs = dydx(x_ipoh,y_temp)\n k2[:] = h*y_derivs[:]\n \n #get k3 values\n y_temp[:] = yi[:] + 0.5*k2[:]\n y_derivs = dydx(x_ipoh,y_temp)\n k3[:] = h*y_derivs[:]\n \n #get k4 values\n y_temp[:] = yi[:] + k3[:]\n y_derivs = dydx(x_ipo,y_temp)\n k4[:] = h*y_derivs[:]\n \n \n #advance y by a step h\n yipo = yi + (k1 + 2*k2 + 2*k3 + k4)/6.\n \n return yipo",
"_____no_output_____"
]
],
[
[
"### Define an adaptive step size driver for RK4",
"_____no_output_____"
]
],
[
[
"def rk4_mv_ad(dydx,x_i,y_i,nv,h,tol):\n \n #define safety scale\n SAFETY = 0.9\n H_NEW_FAC = 2.0\n \n #set a max number of iterations\n imax = 10000\n \n #set an iteration variable\n i = 0\n \n #create an error\n Delta = np.full(nv,2*tol)\n \n #remember the step\n h_step = h\n \n #adjust step\n while(Delta.max()/tol > 1.0):\n \n #estimate our error by taking one step of size h\n #vs. two steps of size h/2\n y_2 = rk4_mv_core(dydx,x_i,y_i,nv,h_step)\n y_1 = rk4_mv_core(dydx,x_i,y_i,nv,0.5*h_step)\n y_11 = rk4_mv_core(dydx,x_i+0.5*h_step,y_1,nv,0.5*h_step)\n \n #compute an error\n Delta = np.fabs(y_2 - y_11)\n \n #if the error is too large, take a smaller step\n if(Delta.max()/tol > 1.0):\n \n #our error is too large, decrease the step\n h_step *= SAFETY * (Delta.max()/tol)**(-0.25)\n \n #check iteration\n if(i>=imax):\n print(\"Too many iterations in rk4_mv_ad()\")\n raise StopIteration(\"Ending after i = \",i)\n \n #iterate\n i+=1\n \n \n #next time, try to take a bigger step\n h_new = np.fmin(h_step * (Delta.max()/tol)**(-0.9), h_step*H_NEW_FAC)\n \n #return the answer, a new step, and the step we actually took\n return y_2, h_new, h_step",
"_____no_output_____"
]
],
[
[
"## Define a wrapper for RK4",
"_____no_output_____"
]
],
[
[
"def rk4_mv(dydx,a,b,y_a,tol):\n \n #dfdx is derivative w.r.t. x\n #a is lower bound\n #b is upper bound\n #y_a are boundary conditions\n #tol is tolerance for integrating y\n \n #define our starting step\n xi = a\n yi = y_a.copy()\n \n #an initial step size == make very small\n h = 1.0e-4 * (b-a)\n \n #set max number of iterations\n imax = 10000\n \n #set an iteration variable\n i = 0\n \n #set the number of coupled ODEs to the size of y_a\n nv = len(y_a)\n \n \n #set the initial conditions\n x = np.full(1,a)\n y = np.full((1,nv),y_a)\n \n #set a flag\n flag = 1\n \n #loop until we reach the right side\n while(flag):\n \n #calculate y_i+1\n yi_new, h_new, h_step = rk4_mv_ad(dydx,xi,yi,nv,h,tol)\n \n #update the step\n h = h_new\n \n #prevent an overshoot\n if(xi+h_step>b):\n \n #take a smaller step\n h = b-xi\n \n #recalculate y_i+1\n yi_new, h_new, h_step = rk4_mv_ad(dydx,xi,yi,nv,h,tol)\n \n #break\n flag = 0\n \n \n #update values\n xi += h_step\n yi[:] = yi_new[:]\n \n #add the step to the arrays\n x = np.append(x,xi)\n y_new = np.zeros((len(x),nv))\n y_new[0:len(x)-1,:] = y\n y_new[-1,:] = yi[:]\n del y\n y = y_new\n \n #prevent too many iterations\n if(i>=imax):\n \n print(colors.bold, colors.fg.red, colors.bg.black, \"Maximum iterations reached.\", colors.reset)\n raise StopIteration(\"Iteration number = \",i)\n \n \n #iterate\n i += 1\n \n #output some information\n s = \"i =%3d\\tx = %9.8f\\th = %9.8f\\tb=%9.8f\" % (i,xi,h_step,b)\n print(s)\n \n #break if new xi is == b\n if(xi==b):\n flag = 0\n \n #return the answer\n print(colors.bold, colors.fg.purple, \"Number of iterations:\", i, colors.reset)\n return x,y",
"_____no_output_____"
]
],
[
[
"### Perform the integration",
"_____no_output_____"
]
],
[
[
"a = 0.0\nb = 2.0 * np.pi\n\ny_0 = np.zeros(2)\ny_0[0] = 0.0\ny_0[1] = 1.0\nnv = 2\n\ntolerance = 1.0e-6\n\n#perform the integration\nx,y = rk4_mv(dydx,a,b,y_0,tolerance)",
"_____no_output_____"
]
],
[
[
"## Plot the result",
"_____no_output_____"
]
],
[
[
"plt.plot(x,y[:,0],'o',color='y',label='y(x)')\nplt.plot(x,y[:,1],'o',color='silver',label='dydx(x)')\nxx = np.linspace(0,2.0*np.pi,1000)\nplt.plot(xx,np.sin(xx),color='k',label='sin(x)')\nplt.plot(xx,np.cos(xx),color='purple',label='cos(x)')\nplt.xlabel('x',color='chocolate',fontsize='large')\nplt.ylabel('y, dy/dx',color='chocolate',fontsize='large')\nplt.legend(frameon=False,fontsize='medium')",
"_____no_output_____"
]
],
[
[
"## Plot the error",
"_____no_output_____"
],
[
"Notice the errors will exceed our \"tolerance\".",
"_____no_output_____"
]
],
[
[
"sin_X = np.sin(x)\ncos_X = np.cos(x)\n\ny_error = (y[:,0]-sin_X)\ndydx_error = (y[:,1]-cos_X)\n\nplt.plot(x, y_error, label=\"y(x) Error\", color='fuchsia')\nplt.plot(x, dydx_error, label=\"dydx(x) Error\", color='aqua')\nplt.legend(frameon=False, fontsize='medium')",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
]
] |
4ab5073369dc5a8f46def4cbb8f1f8b7c7b2698c
| 33,134 |
ipynb
|
Jupyter Notebook
|
advanced_functionality/multi_model_xgboost_home_value/xgboost_multi_model_endpoint_home_value.ipynb
|
P15241328/amazon-sagemaker-examples
|
00cba545be0822474f070321a62d22865187e09b
|
[
"Apache-2.0"
] | 5 |
2019-01-19T23:53:35.000Z
|
2022-01-29T14:04:31.000Z
|
advanced_functionality/multi_model_xgboost_home_value/xgboost_multi_model_endpoint_home_value.ipynb
|
P15241328/amazon-sagemaker-examples
|
00cba545be0822474f070321a62d22865187e09b
|
[
"Apache-2.0"
] | 2 |
2021-08-25T16:15:24.000Z
|
2022-02-10T02:49:50.000Z
|
advanced_functionality/multi_model_xgboost_home_value/xgboost_multi_model_endpoint_home_value.ipynb
|
P15241328/amazon-sagemaker-examples
|
00cba545be0822474f070321a62d22865187e09b
|
[
"Apache-2.0"
] | 7 |
2020-03-04T22:23:51.000Z
|
2021-07-13T14:05:46.000Z
| 38.798595 | 692 | 0.613207 |
[
[
[
"# Amazon SageMaker Multi-Model Endpoints using XGBoost\nWith [Amazon SageMaker multi-model endpoints](https://docs.aws.amazon.com/sagemaker/latest/dg/multi-model-endpoints.html), customers can create an endpoint that seamlessly hosts up to thousands of models. These endpoints are well suited to use cases where any one of a large number of models, which can be served from a common inference container to save inference costs, needs to be invokable on-demand and where it is acceptable for infrequently invoked models to incur some additional latency. For applications which require consistently low inference latency, an endpoint deploying a single model is still the best choice.\n\nAt a high level, Amazon SageMaker manages the loading and unloading of models for a multi-model endpoint, as they are needed. When an invocation request is made for a particular model, Amazon SageMaker routes the request to an instance assigned to that model, downloads the model artifacts from S3 onto that instance, and initiates loading of the model into the memory of the container. As soon as the loading is complete, Amazon SageMaker performs the requested invocation and returns the result. If the model is already loaded in memory on the selected instance, the downloading and loading steps are skipped and the invocation is performed immediately.\n\nTo demonstrate how multi-model endpoints are created and used, this notebook provides an example using a set of XGBoost models that each predict housing prices for a single location. This domain is used as a simple example to easily experiment with multi-model endpoints.\n\nThe Amazon SageMaker multi-model endpoint capability is designed to work across with Mxnet, PyTorch and Scikit-Learn machine learning frameworks (TensorFlow coming soon), SageMaker XGBoost, KNN, and Linear Learner algorithms.\n\nIn addition, Amazon SageMaker multi-model endpoints are also designed to work with cases where you bring your own container that integrates with the multi-model server library. An example of this can be found [here](https://github.com/awslabs/amazon-sagemaker-examples/tree/master/advanced_functionality/multi_model_bring_your_own) and documentation [here.](https://docs.aws.amazon.com/sagemaker/latest/dg/build-multi-model-build-container.html)",
"_____no_output_____"
],
[
"### Contents\n\n1. [Generate synthetic data for housing models](#Generate-synthetic-data-for-housing-models)\n1. [Train multiple house value prediction models](#Train-multiple-house-value-prediction-models)\n1. [Create the Amazon SageMaker MultiDataModel entity](#Create-the-Amazon-SageMaker-MultiDataModel-entity)\n 1. [Create the Multi-Model Endpoint](#Create-the-multi-model-endpoint)\n 1. [Deploy the Multi-Model Endpoint](#deploy-the-multi-model-endpoint)\n1. [Get Predictions from the endpoint](#Get-predictions-from-the-endpoint)\n1. [Additional Information](#Additional-information)\n1. [Clean up](#Clean-up)",
"_____no_output_____"
],
[
"# Generate synthetic data\n\nThe code below contains helper functions to generate synthetic data in the form of a `1x7` numpy array representing the features of a house.\n\nThe first entry in the array is the randomly generated price of a house. The remaining entries are the features (i.e. number of bedroom, square feet, number of bathrooms, etc.).\n\nThese functions will be used to generate synthetic data for training, validation, and testing. It will also allow us to submit synthetic payloads for inference to test our multi-model endpoint.",
"_____no_output_____"
]
],
[
[
"import numpy as np\nimport pandas as pd\nimport time",
"_____no_output_____"
],
[
"NUM_HOUSES_PER_LOCATION = 1000\nLOCATIONS = ['NewYork_NY', 'LosAngeles_CA', 'Chicago_IL', 'Houston_TX', 'Dallas_TX',\n 'Phoenix_AZ', 'Philadelphia_PA', 'SanAntonio_TX', 'SanDiego_CA', 'SanFrancisco_CA']\nPARALLEL_TRAINING_JOBS = 4 # len(LOCATIONS) if your account limits can handle it\nMAX_YEAR = 2019",
"_____no_output_____"
],
[
"def gen_price(house):\n _base_price = int(house['SQUARE_FEET'] * 150)\n _price = int(_base_price + (10000 * house['NUM_BEDROOMS']) + \\\n (15000 * house['NUM_BATHROOMS']) + \\\n (15000 * house['LOT_ACRES']) + \\\n (15000 * house['GARAGE_SPACES']) - \\\n (5000 * (MAX_YEAR - house['YEAR_BUILT'])))\n return _price",
"_____no_output_____"
],
[
"def gen_random_house():\n _house = {'SQUARE_FEET': int(np.random.normal(3000, 750)),\n 'NUM_BEDROOMS': np.random.randint(2, 7),\n 'NUM_BATHROOMS': np.random.randint(2, 7) / 2,\n 'LOT_ACRES': round(np.random.normal(1.0, 0.25), 2),\n 'GARAGE_SPACES': np.random.randint(0, 4),\n 'YEAR_BUILT': min(MAX_YEAR, int(np.random.normal(1995, 10)))}\n _price = gen_price(_house)\n return [_price, _house['YEAR_BUILT'], _house['SQUARE_FEET'], \n _house['NUM_BEDROOMS'], _house['NUM_BATHROOMS'], \n _house['LOT_ACRES'], _house['GARAGE_SPACES']]",
"_____no_output_____"
],
[
"def gen_houses(num_houses):\n _house_list = []\n for i in range(num_houses):\n _house_list.append(gen_random_house())\n _df = pd.DataFrame(_house_list, \n columns=['PRICE', 'YEAR_BUILT', 'SQUARE_FEET', 'NUM_BEDROOMS',\n 'NUM_BATHROOMS','LOT_ACRES', 'GARAGE_SPACES'])\n return _df",
"_____no_output_____"
]
],
[
[
"# Train multiple house value prediction models\n\nIn the follow section, we are setting up the code to train a house price prediction model for each of 4 different cities.\n\nAs such, we will launch multiple training jobs asynchronously, using the XGBoost algorithm.\n\nIn this notebook, we will be using the AWS Managed XGBoost Image for both training and inference - this image provides native support for launching multi-model endpoints.",
"_____no_output_____"
]
],
[
[
"import sagemaker\nfrom sagemaker import get_execution_role\nfrom sagemaker.amazon.amazon_estimator import image_uris\nimport boto3\n\nfrom sklearn.model_selection import train_test_split\n\ns3 = boto3.resource('s3')\n\nsagemaker_session = sagemaker.Session()\nrole = get_execution_role()\n\nBUCKET = sagemaker_session.default_bucket()\n\n# This is references the AWS managed XGBoost container\nXGBOOST_IMAGE = image_uris.retrieve(region=boto3.Session().region_name, framework='xgboost', version='1.0-1')\n\nDATA_PREFIX = 'XGBOOST_BOSTON_HOUSING'\nMULTI_MODEL_ARTIFACTS = 'multi_model_artifacts'\n\nTRAIN_INSTANCE_TYPE = 'ml.m4.xlarge'\nENDPOINT_INSTANCE_TYPE = 'ml.m4.xlarge'\n\nENDPOINT_NAME = 'mme-xgboost-housing'\n\nMODEL_NAME = ENDPOINT_NAME",
"_____no_output_____"
]
],
[
[
"### Split a given dataset into train, validation, and test\n\nThe code below will generate 3 sets of data. 1 set to train, 1 set for validation and 1 for testing.",
"_____no_output_____"
]
],
[
[
"SEED = 7\nSPLIT_RATIOS = [0.6, 0.3, 0.1]\n\ndef split_data(df):\n # split data into train and test sets\n seed = SEED\n val_size = SPLIT_RATIOS[1]\n test_size = SPLIT_RATIOS[2]\n \n num_samples = df.shape[0]\n X1 = df.values[:num_samples, 1:] # keep only the features, skip the target, all rows\n Y1 = df.values[:num_samples, :1] # keep only the target, all rows\n\n # Use split ratios to divide up into train/val/test\n X_train, X_val, y_train, y_val = \\\n train_test_split(X1, Y1, test_size=(test_size + val_size), random_state=seed)\n # Of the remaining non-training samples, give proper ratio to validation and to test\n X_test, X_test, y_test, y_test = \\\n train_test_split(X_val, y_val, test_size=(test_size / (test_size + val_size)), \n random_state=seed)\n # reassemble the datasets with target in first column and features after that\n _train = np.concatenate([y_train, X_train], axis=1)\n _val = np.concatenate([y_val, X_val], axis=1)\n _test = np.concatenate([y_test, X_test], axis=1)\n\n return _train, _val, _test",
"_____no_output_____"
]
],
[
[
"### Launch a single training job for a given housing location\nThere is nothing specific to multi-model endpoints in terms of the models it will host. They are trained in the same way as all other SageMaker models. Here we are using the XGBoost estimator and not waiting for the job to complete.",
"_____no_output_____"
]
],
[
[
"def launch_training_job(location):\n # clear out old versions of the data\n s3_bucket = s3.Bucket(BUCKET)\n full_input_prefix = f'{DATA_PREFIX}/model_prep/{location}'\n s3_bucket.objects.filter(Prefix=full_input_prefix + '/').delete()\n\n # upload the entire set of data for all three channels\n local_folder = f'data/{location}'\n inputs = sagemaker_session.upload_data(path=local_folder, key_prefix=full_input_prefix)\n print(f'Training data uploaded: {inputs}')\n \n _job = 'xgb-{}'.format(location.replace('_', '-'))\n full_output_prefix = f'{DATA_PREFIX}/model_artifacts/{location}'\n s3_output_path = f's3://{BUCKET}/{full_output_prefix}'\n\n \n xgb = sagemaker.estimator.Estimator(XGBOOST_IMAGE, role, \n instance_count=1, instance_type=TRAIN_INSTANCE_TYPE,\n output_path=s3_output_path, base_job_name=_job,\n sagemaker_session=sagemaker_session)\n \n xgb.set_hyperparameters(max_depth=5, eta=0.2, gamma=4, min_child_weight=6, subsample=0.8, silent=0, \n early_stopping_rounds=5, objective='reg:linear', num_round=25) \n \n DISTRIBUTION_MODE = 'FullyReplicated'\n \n train_input = sagemaker.inputs.TrainingInput(s3_data=inputs+'/train', \n distribution=DISTRIBUTION_MODE, content_type='csv')\n \n val_input = sagemaker.inputs.TrainingInput(s3_data=inputs+'/val', \n distribution=DISTRIBUTION_MODE, content_type='csv')\n \n remote_inputs = {'train': train_input, 'validation': val_input}\n\n xgb.fit(remote_inputs, wait=False)\n \n # Return the estimator object\n return xgb",
"_____no_output_____"
]
],
[
[
"### Kick off a model training job for each housing location",
"_____no_output_____"
]
],
[
[
"def save_data_locally(location, train, val, test):\n os.makedirs(f'data/{location}/train')\n np.savetxt( f'data/{location}/train/{location}_train.csv', train, delimiter=',', fmt='%.2f')\n \n os.makedirs(f'data/{location}/val')\n np.savetxt(f'data/{location}/val/{location}_val.csv', val, delimiter=',', fmt='%.2f')\n \n os.makedirs(f'data/{location}/test')\n np.savetxt(f'data/{location}/test/{location}_test.csv', test, delimiter=',', fmt='%.2f')",
"_____no_output_____"
],
[
"import shutil\nimport os\n\nestimators = []\n\nshutil.rmtree('data', ignore_errors=True)\n\nfor loc in LOCATIONS[:PARALLEL_TRAINING_JOBS]:\n _houses = gen_houses(NUM_HOUSES_PER_LOCATION)\n _train, _val, _test = split_data(_houses)\n save_data_locally(loc, _train, _val, _test)\n estimator = launch_training_job(loc)\n estimators.append(estimator)\n\nprint()\nprint(f'{len(estimators)} training jobs launched: {[x.latest_training_job.job_name for x in estimators]}')",
"Training data uploaded: s3://sagemaker-eu-west-1-872120163790/XGBOOST_BOSTON_HOUSING/model_prep/NewYork_NY\nTraining data uploaded: s3://sagemaker-eu-west-1-872120163790/XGBOOST_BOSTON_HOUSING/model_prep/LosAngeles_CA\nTraining data uploaded: s3://sagemaker-eu-west-1-872120163790/XGBOOST_BOSTON_HOUSING/model_prep/Chicago_IL\nTraining data uploaded: s3://sagemaker-eu-west-1-872120163790/XGBOOST_BOSTON_HOUSING/model_prep/Houston_TX\n\n4 training jobs launched: ['xgb-NewYork-NY-2020-11-23-10-50-10-689', 'xgb-LosAngeles-CA-2020-11-23-10-50-11-151', 'xgb-Chicago-IL-2020-11-23-10-50-14-171', 'xgb-Houston-TX-2020-11-23-10-50-15-164']\n"
]
],
[
[
"### Wait for all model training to finish",
"_____no_output_____"
]
],
[
[
"def wait_for_training_job_to_complete(estimator):\n job = estimator.latest_training_job.job_name\n print(f'Waiting for job: {job}')\n status = estimator.latest_training_job.describe()['TrainingJobStatus']\n while status == 'InProgress':\n time.sleep(45)\n status = estimator.latest_training_job.describe()['TrainingJobStatus']\n if status == 'InProgress':\n print(f'{job} job status: {status}')\n print(f'DONE. Status for {job} is {status}\\n')\n ",
"_____no_output_____"
],
[
"for est in estimators:\n wait_for_training_job_to_complete(est)",
"Waiting for job: xgb-NewYork-NY-2020-11-23-10-50-10-689\nxgb-NewYork-NY-2020-11-23-10-50-10-689 job status: InProgress\nxgb-NewYork-NY-2020-11-23-10-50-10-689 job status: InProgress\nDONE. Status for xgb-NewYork-NY-2020-11-23-10-50-10-689 is Completed\n\nWaiting for job: xgb-LosAngeles-CA-2020-11-23-10-50-11-151\nDONE. Status for xgb-LosAngeles-CA-2020-11-23-10-50-11-151 is Completed\n\nWaiting for job: xgb-Chicago-IL-2020-11-23-10-50-14-171\nDONE. Status for xgb-Chicago-IL-2020-11-23-10-50-14-171 is Completed\n\nWaiting for job: xgb-Houston-TX-2020-11-23-10-50-15-164\nDONE. Status for xgb-Houston-TX-2020-11-23-10-50-15-164 is Completed\n\n"
]
],
[
[
"# Create the multi-model endpoint with the SageMaker SDK",
"_____no_output_____"
],
[
"### Create a SageMaker Model from one of the Estimators",
"_____no_output_____"
]
],
[
[
"estimator = estimators[0]\nmodel = estimator.create_model(role=role, image_uri=XGBOOST_IMAGE)",
"_____no_output_____"
]
],
[
[
"### Create the Amazon SageMaker MultiDataModel entity\n\nWe create the multi-model endpoint using the [```MultiDataModel```](https://sagemaker.readthedocs.io/en/stable/api/inference/multi_data_model.html) class.\n\nYou can create a MultiDataModel by directly passing in a `sagemaker.model.Model` object - in which case, the Endpoint will inherit information about the image to use, as well as any environmental variables, network isolation, etc., once the MultiDataModel is deployed.\n\nIn addition, a MultiDataModel can also be created without explictly passing a `sagemaker.model.Model` object. Please refer to the documentation for additional details.",
"_____no_output_____"
]
],
[
[
"from sagemaker.multidatamodel import MultiDataModel",
"_____no_output_____"
],
[
"# This is where our MME will read models from on S3.\nmodel_data_prefix = f's3://{BUCKET}/{DATA_PREFIX}/{MULTI_MODEL_ARTIFACTS}/'",
"_____no_output_____"
],
[
"mme = MultiDataModel(name=MODEL_NAME,\n model_data_prefix=model_data_prefix,\n model=model,# passing our model - passes container image needed for the endpoint\n sagemaker_session=sagemaker_session)",
"_____no_output_____"
]
],
[
[
"# Deploy the Multi Model Endpoint\n\nYou need to consider the appropriate instance type and number of instances for the projected prediction workload across all the models you plan to host behind your multi-model endpoint. The number and size of the individual models will also drive memory requirements.",
"_____no_output_____"
]
],
[
[
"predictor = mme.deploy(initial_instance_count=1,\n instance_type=ENDPOINT_INSTANCE_TYPE,\n endpoint_name=ENDPOINT_NAME)",
"---------------!"
]
],
[
[
"### Our endpoint has launched! Let's look at what models are available to the endpoint!\n\nBy 'available', what we mean is, what model artfiacts are currently stored under the S3 prefix we defined when setting up the `MultiDataModel` above i.e. `model_data_prefix`.\n\nCurrently, since we have no artifacts (i.e. `tar.gz` files) stored under our defined S3 prefix, our endpoint, will have no models 'available' to serve inference requests.\n\nWe will demonstrate how to make models 'available' to our endpoint below.",
"_____no_output_____"
]
],
[
[
"# No models visible!\nlist(mme.list_models())",
"_____no_output_____"
]
],
[
[
"### Lets deploy model artifacts to be found by the endpoint\n\nWe are now using the `.add_model()` method of the `MultiDataModel` to copy over our model artifacts from where they were initially stored, during training, to where our endpoint will source model artifacts for inference requests.\n\n`model_data_source` refers to the location of our model artifact (i.e. where it was deposited on S3 after training completed)\n\n`model_data_path` is the **relative** path to the S3 prefix we specified above (i.e. `model_data_prefix`) where our endpoint will source models for inference requests.\n\nSince this is a **relative** path, we can simply pass the name of what we wish to call the model artifact at inference time (i.e. `Chicago_IL.tar.gz`)\n\n### Dynamically deploying additional models\n\nIt is also important to note, that we can always use the `.add_model()` method, as shown below, to dynamically deploy more models to the endpoint, to serve up inference requests as needed.",
"_____no_output_____"
]
],
[
[
"for est in estimators:\n artifact_path = est.latest_training_job.describe()['ModelArtifacts']['S3ModelArtifacts']\n model_name = artifact_path.split('/')[-4]+'.tar.gz'\n # This is copying over the model artifact to the S3 location for the MME.\n mme.add_model(model_data_source=artifact_path, model_data_path=model_name)",
"_____no_output_____"
]
],
[
[
"## We have added the 4 model artifacts from our training jobs!\n\nWe can see that the S3 prefix we specified when setting up `MultiDataModel` now has 4 model artifacts. As such, the endpoint can now serve up inference requests for these models.",
"_____no_output_____"
]
],
[
[
"list(mme.list_models())",
"_____no_output_____"
]
],
[
[
"# Get predictions from the endpoint\n\nRecall that ```mme.deploy()``` returns a [RealTimePredictor](https://github.com/aws/sagemaker-python-sdk/blob/master/src/sagemaker/predictor.py#L35) that we saved in a variable called ```predictor```.\n\nWe will use ```predictor``` to submit requests to the endpoint.\n\nXGBoost supports ```text/csv``` for the content type and accept type. For more information on XGBoost Input/Output Interface, please see [here.](https://docs.aws.amazon.com/sagemaker/latest/dg/xgboost.html#InputOutput-XGBoost)\n\nSince the default RealTimePredictor does not have a serializer or deserializer set for requests, we will also set these.\n\nThis will allow us to submit a python list for inference, and get back a float response.",
"_____no_output_____"
]
],
[
[
"from sagemaker.serializers import CSVSerializer\nfrom sagemaker.deserializers import JSONDeserializer\n",
"_____no_output_____"
],
[
"predictor.serializer = CSVSerializer()\npredictor.deserializer = JSONDeserializer()\n#predictor.content_type =predictor.content_type , removed as mentioned https://github.com/aws/sagemaker-python-sdk/blob/e8d16f8bc4c570f763f1129afc46ba3e0b98cdad/src/sagemaker/predictor.py#L82\n#predictor.accept = \"text/csv\" # removed also : https://github.com/aws/sagemaker-python-sdk/blob/e8d16f8bc4c570f763f1129afc46ba3e0b98cdad/src/sagemaker/predictor.py#L83",
"_____no_output_____"
]
],
[
[
"### Invoking models on a multi-model endpoint\nNotice the higher latencies on the first invocation of any given model. This is due to the time it takes SageMaker to download the model to the Endpoint instance and then load the model into the inference container. Subsequent invocations of the same model take advantage of the model already being loaded into the inference container.",
"_____no_output_____"
]
],
[
[
"start_time = time.time()\n\npredicted_value = predictor.predict(data=gen_random_house()[1:], target_model='Chicago_IL.tar.gz')\n\nduration = time.time() - start_time\nprint('${:,.2f}, took {:,d} ms\\n'.format(predicted_value[0], int(duration * 1000)))",
"$548,638.31, took 16 ms\n\n"
],
[
"start_time = time.time()\n\n#Invoke endpoint\npredicted_value = predictor.predict(data=gen_random_house()[1:], target_model='Chicago_IL.tar.gz')\n\nduration = time.time() - start_time\nprint('${:,.2f}, took {:,d} ms\\n'.format(predicted_value[0], int(duration * 1000)))",
"$517,095.44, took 19 ms\n\n"
],
[
"start_time = time.time()\n\n#Invoke endpoint\npredicted_value = predictor.predict(data=gen_random_house()[1:], target_model='Houston_TX.tar.gz')\n\nduration = time.time() - start_time\nprint('${:,.2f}, took {:,d} ms\\n'.format(predicted_value[0], int(duration * 1000)))",
"$313,395.22, took 1,013 ms\n\n"
],
[
"start_time = time.time()\n\n#Invoke endpoint\npredicted_value = predictor.predict(data=gen_random_house()[1:], target_model='Houston_TX.tar.gz')\n\nduration = time.time() - start_time\nprint('${:,.2f}, took {:,d} ms\\n'.format(predicted_value[0], int(duration * 1000)))",
"$543,326.38, took 16 ms\n\n"
]
],
[
[
"### Updating a model\nTo update a model, you would follow the same approach as above and add it as a new model. For example, if you have retrained the `NewYork_NY.tar.gz` model and wanted to start invoking it, you would upload the updated model artifacts behind the S3 prefix with a new name such as `NewYork_NY_v2.tar.gz`, and then change the `target_model` field to invoke `NewYork_NY_v2.tar.gz` instead of `NewYork_NY.tar.gz`. You do not want to overwrite the model artifacts in Amazon S3, because the old version of the model might still be loaded in the containers or on the storage volume of the instances on the endpoint. Invocations to the new model could then invoke the old version of the model.\n\nAlternatively, you could stop the endpoint and re-deploy a fresh set of models.",
"_____no_output_____"
],
[
"## Using Boto APIs to invoke the endpoint\n\nWhile developing interactively within a Jupyter notebook, since `.deploy()` returns a `RealTimePredictor` it is a more seamless experience to start invoking your endpoint using the SageMaker SDK. You have more fine grained control over the serialization and deserialization protocols to shape your request and response payloads to/from the endpoint.\n\nThis is great for iterative experimentation within a notebook. Furthermore, should you have an application that has access to the SageMaker SDK, you can always import `RealTimePredictor` and attach it to an existing endpoint - this allows you to stick to using the high level SDK if preferable.\n\nAdditional documentation on `RealTimePredictor` can be found [here.](https://sagemaker.readthedocs.io/en/stable/api/inference/predictors.html?highlight=RealTimePredictor#sagemaker.predictor.RealTimePredictor)\n\nThe lower level Boto3 SDK may be preferable if you are attempting to invoke the endpoint as a part of a broader architecture.\n\nImagine an API gateway frontend that uses a Lambda Proxy in order to transform request payloads before hitting a SageMaker Endpoint - in this example, Lambda does not have access to the SageMaker Python SDK, and as such, Boto3 can still allow you to interact with your endpoint and serve inference requests.\n\nBoto3 allows for quick injection of ML intelligence via SageMaker Endpoints into existing applications with minimal/no refactoring to existing code.\n\nBoto3 will submit your requests as a binary payload, while still allowing you to supply your desired `Content-Type` and `Accept` headers with serialization being handled by the inference container in the SageMaker Endpoint.\n\nAdditional documentation on `.invoke_endpoint()` can be found [here.](https://boto3.amazonaws.com/v1/documentation/api/latest/reference/services/sagemaker-runtime.html)",
"_____no_output_____"
]
],
[
[
"import boto3\nimport json\n\nruntime_sm_client = boto3.client(service_name='sagemaker-runtime')\n\ndef predict_one_house_value(features, model_name):\n print(f'Using model {model_name} to predict price of this house: {features}')\n \n # Notice how we alter the list into a string as the payload\n body = ','.join(map(str, features)) + '\\n'\n \n start_time = time.time()\n\n response = runtime_sm_client.invoke_endpoint(\n EndpointName=ENDPOINT_NAME,\n ContentType='text/csv',\n TargetModel=model_name,\n Body=body)\n \n predicted_value = json.loads(response['Body'].read())[0]\n\n duration = time.time() - start_time\n \n print('${:,.2f}, took {:,d} ms\\n'.format(predicted_value, int(duration * 1000)))",
"_____no_output_____"
],
[
"predict_one_house_value(gen_random_house()[1:], 'Chicago_IL.tar.gz')",
"Using model Chicago_IL.tar.gz to predict price of this house: [1990, 2269, 3, 1.0, 0.95, 3]\n$309,167.28, took 57 ms\n\n"
]
],
[
[
"## Clean up\nHere, to be sure we are not billed for endpoints we are no longer using, we clean up.",
"_____no_output_____"
]
],
[
[
"predictor.delete_endpoint()",
"_____no_output_____"
],
[
"predictor.delete_model()",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4ab508fc88769f8ca7853db9aff19ef51130e435
| 103,877 |
ipynb
|
Jupyter Notebook
|
kaggle/get-97-using-simple-yet-one-parameter-algorithm.ipynb
|
fossabot/denmune-clustering-algorithm
|
aa815150bd3dc0dfddca70d7f75c207f92b5d3b0
|
[
"BSD-3-Clause"
] | 9 |
2021-12-22T21:21:20.000Z
|
2022-02-09T18:01:53.000Z
|
kaggle/get-97-using-simple-yet-one-parameter-algorithm.ipynb
|
fossabot/denmune-clustering-algorithm
|
aa815150bd3dc0dfddca70d7f75c207f92b5d3b0
|
[
"BSD-3-Clause"
] | 3 |
2022-01-01T00:45:12.000Z
|
2022-02-05T01:21:34.000Z
|
kaggle/get-97-using-simple-yet-one-parameter-algorithm.ipynb
|
fossabot/denmune-clustering-algorithm
|
aa815150bd3dc0dfddca70d7f75c207f92b5d3b0
|
[
"BSD-3-Clause"
] | 2 |
2022-02-03T23:03:35.000Z
|
2022-02-05T07:56:39.000Z
| 103,877 | 103,877 | 0.939611 |
[
[
[
"<a href=\"https://www.kaggle.com/egyfirst/get-97-using-simple-yet-one-parameter-algorithm?scriptVersionId=86411700\" target=\"_blank\"><img align=\"left\" alt=\"Kaggle\" title=\"Open in Kaggle\" src=\"https://kaggle.com/static/images/open-in-kaggle.svg\"></a>",
"_____no_output_____"
]
],
[
[
"import pandas as pd\nimport numpy as np\nimport time\nimport os.path\n\nimport warnings\nwarnings.filterwarnings('ignore')",
"_____no_output_____"
],
[
"# install DenMune clustering algorithm using pip command from the offecial Python repository, PyPi\n# from https://pypi.org/project/denmune/\n!pip install denmune\n\n# now import it\nfrom denmune import DenMune",
"Collecting denmune\r\n Downloading denmune-0.0.9.0-py3-none-any.whl (13 kB)\r\nRequirement already satisfied: pandas>=1.0.3 in /opt/conda/lib/python3.7/site-packages (from denmune) (1.3.5)\r\nRequirement already satisfied: scikit-learn>=0.22.1 in /opt/conda/lib/python3.7/site-packages (from denmune) (0.23.2)\r\nCollecting anytree>=2.8.0\r\n Downloading anytree-2.8.0-py2.py3-none-any.whl (41 kB)\r\n |████████████████████████████████| 41 kB 92 kB/s \r\n\u001b[?25hCollecting ngt>=1.11.6\r\n Downloading ngt-1.12.2-cp37-cp37m-manylinux1_x86_64.whl (2.2 MB)\r\n |████████████████████████████████| 2.2 MB 1.6 MB/s \r\n\u001b[?25hCollecting treelib>=1.6.1\r\n Downloading treelib-1.6.1.tar.gz (24 kB)\r\n Preparing metadata (setup.py) ... \u001b[?25l-\b \bdone\r\n\u001b[?25hRequirement already satisfied: matplotlib>=3.2.1 in /opt/conda/lib/python3.7/site-packages (from denmune) (3.5.1)\r\nRequirement already satisfied: numpy>=1.18.5 in /opt/conda/lib/python3.7/site-packages (from denmune) (1.20.3)\r\nRequirement already satisfied: seaborn>=0.10.1 in /opt/conda/lib/python3.7/site-packages (from denmune) (0.11.2)\r\nRequirement already satisfied: six>=1.9.0 in /opt/conda/lib/python3.7/site-packages (from anytree>=2.8.0->denmune) (1.16.0)\r\nRequirement already satisfied: pillow>=6.2.0 in /opt/conda/lib/python3.7/site-packages (from matplotlib>=3.2.1->denmune) (8.2.0)\r\nRequirement already satisfied: pyparsing>=2.2.1 in /opt/conda/lib/python3.7/site-packages (from matplotlib>=3.2.1->denmune) (3.0.6)\r\nRequirement already satisfied: kiwisolver>=1.0.1 in /opt/conda/lib/python3.7/site-packages (from matplotlib>=3.2.1->denmune) (1.3.2)\r\nRequirement already satisfied: cycler>=0.10 in /opt/conda/lib/python3.7/site-packages (from matplotlib>=3.2.1->denmune) (0.11.0)\r\nRequirement already satisfied: fonttools>=4.22.0 in /opt/conda/lib/python3.7/site-packages (from matplotlib>=3.2.1->denmune) (4.28.4)\r\nRequirement already satisfied: python-dateutil>=2.7 in /opt/conda/lib/python3.7/site-packages (from matplotlib>=3.2.1->denmune) (2.8.0)\r\nRequirement already satisfied: packaging>=20.0 in /opt/conda/lib/python3.7/site-packages (from matplotlib>=3.2.1->denmune) (21.3)\r\nRequirement already satisfied: pybind11 in /opt/conda/lib/python3.7/site-packages (from ngt>=1.11.6->denmune) (2.9.0)\r\nRequirement already satisfied: pytz>=2017.3 in /opt/conda/lib/python3.7/site-packages (from pandas>=1.0.3->denmune) (2021.3)\r\nRequirement already satisfied: scipy>=0.19.1 in /opt/conda/lib/python3.7/site-packages (from scikit-learn>=0.22.1->denmune) (1.7.3)\r\nRequirement already satisfied: joblib>=0.11 in /opt/conda/lib/python3.7/site-packages (from scikit-learn>=0.22.1->denmune) (1.1.0)\r\nRequirement already satisfied: threadpoolctl>=2.0.0 in /opt/conda/lib/python3.7/site-packages (from scikit-learn>=0.22.1->denmune) (3.0.0)\r\nRequirement already satisfied: future in /opt/conda/lib/python3.7/site-packages (from treelib>=1.6.1->denmune) (0.18.2)\r\nBuilding wheels for collected packages: treelib\r\n Building wheel for treelib (setup.py) ... \u001b[?25l-\b \b\\\b \bdone\r\n\u001b[?25h Created wheel for treelib: filename=treelib-1.6.1-py3-none-any.whl size=18386 sha256=fef0b01779b1cc65ee2fc6b0e82bb7ec95bd727eced8b26130a3551e44f4400b\r\n Stored in directory: /root/.cache/pip/wheels/89/be/94/2c6d949ce599d1443426d83ba4dc93cd35c0f4638260930a53\r\nSuccessfully built treelib\r\nInstalling collected packages: treelib, ngt, anytree, denmune\r\nSuccessfully installed anytree-2.8.0 denmune-0.0.9.0 ngt-1.12.2 treelib-1.6.1\r\n\u001b[33mWARNING: Running pip as the 'root' user can result in broken permissions and conflicting behaviour with the system package manager. It is recommended to use a virtual environment instead: https://pip.pypa.io/warnings/venv\u001b[0m\r\n"
],
[
"data_path = '../input/digit-recognizer/' \nfile_2d = '../input/mnist-2d/mnist-2d.csv'\n\nX_train = pd.read_csv( data_path + 'train.csv', sep=',')\nX_test = pd.read_csv( data_path +'test.csv', sep=',')\ny_train = X_train['label']\nX_train = X_train.drop(['label'], axis=1)\n\ndm = DenMune(train_data=X_train,\n train_truth=y_train,\n test_data=X_test, \n k_nearest=93,\n file_2d=file_2d,\n rgn_tsne=False)\n\nlabels, validity = dm.fit_predict(show_noise=True, show_analyzer=True)\n",
"Plotting dataset Groundtruth\n"
],
[
"# prepare our output to be submitted to the dataset kaggle competition\nImageID = np.arange(len(X_test))+1\nOut = pd.DataFrame([ImageID,labels['test']]).T\nOut.to_csv('submission.csv', header = ['ImageId', 'Label' ], index = None)\n",
"_____no_output_____"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
4ab50d698dbdb77bf538473ed3fcaf0c59bde491
| 80,113 |
ipynb
|
Jupyter Notebook
|
Continuous_Control.ipynb
|
payal80/P2-DRLND
|
d7ad6569d36d94f26464752ed54cb753e11e1f06
|
[
"MIT"
] | null | null | null |
Continuous_Control.ipynb
|
payal80/P2-DRLND
|
d7ad6569d36d94f26464752ed54cb753e11e1f06
|
[
"MIT"
] | null | null | null |
Continuous_Control.ipynb
|
payal80/P2-DRLND
|
d7ad6569d36d94f26464752ed54cb753e11e1f06
|
[
"MIT"
] | null | null | null | 42.41027 | 21,172 | 0.614245 |
[
[
[
"# Continuous Control\n\n---\n\nIn this notebook, you will learn how to use the Unity ML-Agents environment for the second project of the [Deep Reinforcement Learning Nanodegree](https://www.udacity.com/course/deep-reinforcement-learning-nanodegree--nd893) program.\n\n### 1. Start the Environment\n\nWe begin by importing the necessary packages. If the code cell below returns an error, please revisit the project instructions to double-check that you have installed [Unity ML-Agents](https://github.com/Unity-Technologies/ml-agents/blob/master/docs/Installation.md) and [NumPy](http://www.numpy.org/).",
"_____no_output_____"
]
],
[
[
"from unityagents import UnityEnvironment\nimport numpy as np",
"_____no_output_____"
]
],
[
[
"Next, we will start the environment! **_Before running the code cell below_**, change the `file_name` parameter to match the location of the Unity environment that you downloaded.\n\n- **Mac**: `\"path/to/Reacher.app\"`\n- **Windows** (x86): `\"path/to/Reacher_Windows_x86/Reacher.exe\"`\n- **Windows** (x86_64): `\"path/to/Reacher_Windows_x86_64/Reacher.exe\"`\n- **Linux** (x86): `\"path/to/Reacher_Linux/Reacher.x86\"`\n- **Linux** (x86_64): `\"path/to/Reacher_Linux/Reacher.x86_64\"`\n- **Linux** (x86, headless): `\"path/to/Reacher_Linux_NoVis/Reacher.x86\"`\n- **Linux** (x86_64, headless): `\"path/to/Reacher_Linux_NoVis/Reacher.x86_64\"`\n\nFor instance, if you are using a Mac, then you downloaded `Reacher.app`. If this file is in the same folder as the notebook, then the line below should appear as follows:\n```\nenv = UnityEnvironment(file_name=\"Reacher.app\")\n```",
"_____no_output_____"
]
],
[
[
"env = UnityEnvironment(file_name='/home/payal/Documents/deep-reinforcement-learning/p2-continuous-control//Reacher_Linux_NoVis/Reacher.x86_64')",
"INFO:unityagents:\n'Academy' started successfully!\nUnity Academy name: Academy\n Number of Brains: 1\n Number of External Brains : 1\n Lesson number : 0\n Reset Parameters :\n\t\tgoal_size -> 5.0\n\t\tgoal_speed -> 1.0\nUnity brain name: ReacherBrain\n Number of Visual Observations (per agent): 0\n Vector Observation space type: continuous\n Vector Observation space size (per agent): 33\n Number of stacked Vector Observation: 1\n Vector Action space type: continuous\n Vector Action space size (per agent): 4\n Vector Action descriptions: , , , \n"
]
],
[
[
"Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.",
"_____no_output_____"
]
],
[
[
"# get the default brain\nbrain_name = env.brain_names[0]\nbrain = env.brains[brain_name]",
"_____no_output_____"
]
],
[
[
"### 2. Examine the State and Action Spaces\n\nIn this environment, a double-jointed arm can move to target locations. A reward of `+0.1` is provided for each step that the agent's hand is in the goal location. Thus, the goal of your agent is to maintain its position at the target location for as many time steps as possible.\n\nThe observation space consists of `33` variables corresponding to position, rotation, velocity, and angular velocities of the arm. Each action is a vector with four numbers, corresponding to torque applicable to two joints. Every entry in the action vector must be a number between `-1` and `1`.\n\nThe simulation contains `20` identical agents, each with its own copy of the environment.\n\nRun the code cell below to print some information about the environment.",
"_____no_output_____"
]
],
[
[
"# reset the environment\nenv_info = env.reset(train_mode=True)[brain_name]\n\n# number of agents\nnum_agents = len(env_info.agents)\nprint('Number of agents:', num_agents)\n\n# size of each action\naction_size = brain.vector_action_space_size\nprint('Size of each action:', action_size)\n\n# examine the state space \nstates = env_info.vector_observations\nstate_size = states.shape[1]\nprint('There are {} agents. Each observes a state with length: {}'.format(states.shape[0], state_size))\nprint('The state for the first agent looks like:', states[0])",
"Number of agents: 20\nSize of each action: 4\nThere are 20 agents. Each observes a state with length: 33\nThe state for the first agent looks like: [ 0.00000000e+00 -4.00000000e+00 0.00000000e+00 1.00000000e+00\n -0.00000000e+00 -0.00000000e+00 -4.37113883e-08 0.00000000e+00\n 0.00000000e+00 0.00000000e+00 0.00000000e+00 0.00000000e+00\n 0.00000000e+00 0.00000000e+00 -1.00000000e+01 0.00000000e+00\n 1.00000000e+00 -0.00000000e+00 -0.00000000e+00 -4.37113883e-08\n 0.00000000e+00 0.00000000e+00 0.00000000e+00 0.00000000e+00\n 0.00000000e+00 0.00000000e+00 5.75471878e+00 -1.00000000e+00\n 5.55726624e+00 0.00000000e+00 1.00000000e+00 0.00000000e+00\n -1.68164849e-01]\n"
]
],
[
[
"### 3. Take Random Actions in the Environment\n\nIn the next code cell, you will learn how to use the Python API to control the agent and receive feedback from the environment.\n\nOnce this cell is executed, you will watch the agent's performance, if it selects an action at random with each time step. A window should pop up that allows you to observe the agent, as it moves through the environment. \n\nOf course, as part of the project, you'll have to change the code so that the agent is able to use its experience to gradually choose better actions when interacting with the environment!",
"_____no_output_____"
]
],
[
[
"env_info = env.reset(train_mode=False)[brain_name] # reset the environment \nstates = env_info.vector_observations # get the current state (for each agent)\nscores = np.zeros(num_agents) # initialize the score (for each agent)\nwhile True:\n actions = np.random.randn(num_agents, action_size) # select an action (for each agent)\n actions = np.clip(actions, -1, 1) # all actions between -1 and 1\n env_info = env.step(actions)[brain_name] # send all actions to tne environment\n next_states = env_info.vector_observations # get next state (for each agent)\n rewards = env_info.rewards # get reward (for each agent)\n dones = env_info.local_done # see if episode finished\n scores += env_info.rewards # update the score (for each agent)\n states = next_states # roll over states to next time step\n if np.any(dones): # exit loop if episode finished\n break\nprint('Total score (averaged over agents) this episode: {}'.format(np.mean(scores)))",
"Total score (averaged over agents) this episode: 0.14999999664723873\n"
]
],
[
[
"When finished, you can close the environment.",
"_____no_output_____"
]
],
[
[
"import gym\nimport random\nimport torch\nimport numpy as np\nfrom collections import deque\nimport matplotlib.pyplot as plt\n%matplotlib inline\n\n",
"_____no_output_____"
],
[
"import numpy as np\n\nimport torch\nimport torch.nn as nn\nimport torch.nn.functional as F\n\ndef hidden_init(layer):\n fan_in = layer.weight.data.size()[0]\n lim = 1. / np.sqrt(fan_in)\n return (-lim, lim)\n\n\nclass Actor(nn.Module):\n \"\"\"Actor (Policy) Model.\"\"\"\n\n def __init__(self, state_size, action_size, seed, fc1_units=256, fc2_units=128):\n \"\"\"Initialize parameters and build model.\n Params\n ======\n state_size (int): Dimension of each state\n action_size (int): Dimension of each action\n seed (int): Random seed\n fc1_units (int): Number of nodes in first hidden layer\n fc2_units (int): Number of nodes in second hidden layer\n \"\"\"\n super(Actor, self).__init__()\n self.seed = torch.manual_seed(seed)\n self.fc1 = nn.Linear(state_size, fc1_units)\n self.fc2 = nn.Linear(fc1_units, fc2_units)\n self.fc3 = nn.Linear(fc2_units, action_size)\n self.reset_parameters()\n\n def reset_parameters(self):\n self.fc1.weight.data.uniform_(*hidden_init(self.fc1))\n self.fc2.weight.data.uniform_(*hidden_init(self.fc2))\n self.fc3.weight.data.uniform_(-3e-3, 3e-3)\n\n def forward(self, state):\n \"\"\"Build an actor (policy) network that maps states -> actions.\"\"\"\n x = F.relu(self.fc1(state))\n x = F.relu(self.fc2(x))\n return F.tanh(self.fc3(x))\n \n\nclass Critic(nn.Module):\n \"\"\"Critic (Value) Model.\"\"\"\n\n def __init__(self, state_size, action_size, seed, fcs1_units=128, fc2_units=64):\n \"\"\"Initialize parameters and build model.\n Params\n ======\n state_size (int): Dimension of each state\n action_size (int): Dimension of each action\n seed (int): Random seed\n fcs1_units (int): Number of nodes in the first hidden layer\n fc2_units (int): Number of nodes in the second hidden layer\n \"\"\"\n super(Critic, self).__init__()\n self.seed = torch.manual_seed(seed)\n self.fcs1 = nn.Linear(state_size, fcs1_units)\n self.fc2 = nn.Linear(fcs1_units+action_size, fc2_units)\n self.fc3 = nn.Linear(fc2_units, 1)\n self.reset_parameters()\n\n def reset_parameters(self):\n self.fcs1.weight.data.uniform_(*hidden_init(self.fcs1))\n self.fc2.weight.data.uniform_(*hidden_init(self.fc2))\n self.fc3.weight.data.uniform_(-3e-3, 3e-3)\n\n def forward(self, state, action):\n \"\"\"Build a critic (value) network that maps (state, action) pairs -> Q-values.\"\"\"\n state = nn.BatchNorm1d(state)\n xs = F.relu(self.fcs1(state))\n x = torch.cat((xs, action), dim=1)\n x = F.relu(self.fc2(x))\n return self.fc3(x)",
"_____no_output_____"
],
[
"import numpy as np\nimport random\nimport time \nimport copy\nfrom collections import namedtuple, deque\n\n\nimport torch\nimport torch.nn.functional as F\nimport torch.optim as optim\n\nBUFFER_SIZE = int(1e6) \nBATCH_SIZE = 1024 \nGAMMA = 0.99 \nTAU = 1e-3 \nLR_ACTOR = 1e-3 \nLR_CRITIC = 1e-3 \nWEIGHT_DECAY = 0.0000 \n\ndevice = torch.device(\"cuda:0\" if torch.cuda.is_available() else \"cpu\")\n\nclass Agent():\n \"\"\"Interacts with and learns from the environment.\"\"\"\n \n def __init__(self, state_size, action_size, random_seed):\n \"\"\"Initialize an Agent object.\n \n Params\n ======\n state_size (int): dimension of each state\n action_size (int): dimension of each action\n random_seed (int): random seed\n \"\"\"\n self.state_size = state_size\n self.action_size = action_size\n self.seed = random.seed(random_seed)\n\n self.actor_local = Actor(state_size, action_size, random_seed).to(device)\n self.actor_target = Actor(state_size, action_size, random_seed).to(device)\n self.actor_optimizer = optim.Adam(self.actor_local.parameters(), lr=LR_ACTOR)\n\n self.critic_local = Critic(state_size, action_size, random_seed).to(device)\n self.critic_target = Critic(state_size, action_size, random_seed).to(device)\n self.critic_optimizer = optim.Adam(self.critic_local.parameters(), lr=LR_CRITIC, weight_decay=WEIGHT_DECAY)\n\n self.noise = OUNoise(action_size, random_seed)\n\n self.memory = ReplayBuffer(action_size, BUFFER_SIZE, BATCH_SIZE, random_seed)\n \n def step(self, state, action, reward, next_state, done, time_step):\n\n for s, a, r, ns, d in zip(state, action, reward, next_state, done):\n self.memory.add(s, a, r, ns, d) \n \n if len(self.memory) > BATCH_SIZE and time_step % 20 == 0:\n for _ in range(10):\n experiences = self.memory.sample()\n self.learn(experiences, GAMMA)\n\n def act(self, state, add_noise=True):\n \"\"\"Returns actions for given state as per current policy.\"\"\"\n\n state = torch.from_numpy(state).float().to(device)\n\n self.actor_local.eval()\n with torch.no_grad():\n action = self.actor_local(state).cpu().data.numpy()\n self.actor_local.train()\n\n if add_noise:\n action += self.noise.sample()\n \n return action\n\n def reset(self):\n self.noise.reset()\n\n def learn(self, experiences, gamma):\n \"\"\"Update policy and value parameters using given batch of experience tuples.\n Q_targets = r + γ * critic_target(next_state, actor_target(next_state))\n where:\n actor_target(state) -> action\n critic_target(state, action) -> Q-value\n Params\n ======\n experiences (Tuple[torch.Tensor]): tuple of (s, a, r, s', done) tuples \n gamma (float): discount factor\n \"\"\"\n states, actions, rewards, next_states, dones = experiences\n\n actions_next = self.actor_target(next_states)\n Q_targets_next = self.critic_target(next_states, actions_next)\n\n Q_targets = rewards + (gamma * Q_targets_next * (1 - dones))\n Q_expected = self.critic_local(states, actions)\n critic_loss = F.mse_loss(Q_expected, Q_targets)\n\n self.critic_optimizer.zero_grad()\n critic_loss.backward()\n torch.nn.utils.clip_grad_norm_(self.critic_local.parameters(), 1)\n self.critic_optimizer.step()\n\n actions_pred = self.actor_local(states)\n actor_loss = -self.critic_local(states, actions_pred).mean()\n self.actor_optimizer.zero_grad()\n actor_loss.backward()\n self.actor_optimizer.step()\n\n self.soft_update(self.critic_local, self.critic_target, TAU)\n self.soft_update(self.actor_local, self.actor_target, TAU) \n\n def soft_update(self, local_model, target_model, tau):\n \"\"\"Soft update model parameters.\n θ_target = τ*θ_local + (1 - τ)*θ_target\n Params\n ======\n local_model: PyTorch model (weights will be copied from)\n target_model: PyTorch model (weights will be copied to)\n tau (float): interpolation parameter \n \"\"\"\n for target_param, local_param in zip(target_model.parameters(), local_model.parameters()):\n target_param.data.copy_(tau*local_param.data + (1.0-tau)*target_param.data)\n\nclass OUNoise:\n \"\"\"Ornstein-Uhlenbeck process.\"\"\"\n\n def __init__(self, size, seed, mu=0., theta=0.15, sigma=0.2):\n \"\"\"Initialize parameters and noise process.\"\"\"\n self.mu = mu * np.ones(size)\n self.theta = theta\n self.sigma = sigma\n self.seed = random.seed(seed)\n self.reset()\n\n def reset(self):\n \"\"\"Reset the internal state (= noise) to mean (mu).\"\"\"\n self.state = copy.copy(self.mu)\n\n def sample(self):\n \"\"\"Update internal state and return it as a noise sample.\"\"\"\n x = self.state\n\n dx = self.theta * (self.mu - x) + self.sigma * np.array([random.random() for i in range(len(x))])\n self.state = x + dx\n\n return self.state\n\nclass ReplayBuffer:\n \"\"\"Fixed-size buffer to store experience tuples.\"\"\"\n\n def __init__(self, action_size, buffer_size, batch_size, seed):\n \"\"\"Initialize a ReplayBuffer object.\n Params\n ======\n buffer_size (int): maximum size of buffer\n batch_size (int): size of each training batch\n \"\"\"\n self.action_size = action_size\n self.memory = deque(maxlen=buffer_size)\n self.batch_size = batch_size\n self.experience = namedtuple(\"Experience\", field_names=[\"state\", \"action\", \"reward\", \"next_state\", \"done\"])\n self.seed = random.seed(seed)\n \n def add(self, state, action, reward, next_state, done):\n \"\"\"Add a new experience to memory.\"\"\"\n e = self.experience(state, action, reward, next_state, done)\n self.memory.append(e)\n \n def sample(self):\n \"\"\"Randomly sample a batch of experiences from memory.\"\"\"\n experiences = random.sample(self.memory, k=self.batch_size)\n\n states = torch.from_numpy(np.vstack([e.state for e in experiences if e is not None])).float().to(device)\n actions = torch.from_numpy(np.vstack([e.action for e in experiences if e is not None])).float().to(device)\n rewards = torch.from_numpy(np.vstack([e.reward for e in experiences if e is not None])).float().to(device)\n next_states = torch.from_numpy(np.vstack([e.next_state for e in experiences if e is not None])).float().to(device)\n dones = torch.from_numpy(np.vstack([e.done for e in experiences if e is not None]).astype(np.uint8)).float().to(device)\n\n return (states, actions, rewards, next_states, dones)\n\n def __len__(self):\n \"\"\"Return the current size of internal memory.\"\"\"\n return len(self.memory)",
"_____no_output_____"
],
[
"agent = Agent(state_size=state_size, action_size=action_size, random_seed=0)\n\n\ndef ddpg(n_episodes=2000, max_steps=1000):\n scores_hundred_queue = deque(maxlen=100)\n scores = []\n for i_episode in range(1, n_episodes+1):\n then = time.time()\n average_score = 0\n env_info = env.reset(train_mode=True)[brain_name]\n states = env_info.vector_observations \n scores_agents = np.zeros(num_agents) \n agent.reset()\n score = 0\n for step in range(max_steps):\n actions = agent.act(states)\n env_info = env.step(actions)[brain_name] \n next_states = env_info.vector_observations \n rewards = env_info.rewards \n dones = env_info.local_done \n agent.step(states, actions, rewards, next_states, dones, step)\n states = next_states\n scores_agents += rewards\n if np.any(dones):\n break\n score = np.mean(scores_agents)\n scores_hundred_queue.append(score)\n average_score = np.mean(scores_hundred_queue)\n scores.append(score) \n \n print(\"Episode: \", i_episode)\n print(\"Min Score: {:.2f} Max Score: {:.2f}\".format(scores_agents.min(), scores_agents.max()))\n print(\"Score: {:.2f}\".format(score))\n print(\"AvgScore: {:.2f}\".format(average_score))\n now = time.time()\n print(\"Time: {:.2f} secs\\n\".format(now - then))\n if average_score > 30:\n torch.save(agent.actor_local.state_dict(), 'checkpoint_actor.pth')\n torch.save(agent.critic_local.state_dict(), 'checkpoint_critic.pth') \n break\n return scores\n\nscores = ddpg()\n",
"Episode: 1\nMin Score: 0.02 Max Score: 1.41\nScore: 0.56\nAvgScore: 0.56\nTime: 33.38 secs\n\nEpisode: 2\nMin Score: 0.10 Max Score: 1.22\nScore: 0.69\nAvgScore: 0.63\nTime: 36.57 secs\n\nEpisode: 3\nMin Score: 0.05 Max Score: 1.58\nScore: 0.52\nAvgScore: 0.59\nTime: 36.19 secs\n\nEpisode: 4\nMin Score: 0.07 Max Score: 1.39\nScore: 0.68\nAvgScore: 0.61\nTime: 35.77 secs\n\nEpisode: 5\nMin Score: 0.29 Max Score: 1.36\nScore: 0.83\nAvgScore: 0.66\nTime: 34.88 secs\n\nEpisode: 6\nMin Score: 0.24 Max Score: 1.52\nScore: 0.82\nAvgScore: 0.69\nTime: 35.70 secs\n\nEpisode: 7\nMin Score: 0.22 Max Score: 1.62\nScore: 0.82\nAvgScore: 0.71\nTime: 38.68 secs\n\nEpisode: 8\nMin Score: 0.33 Max Score: 1.66\nScore: 1.02\nAvgScore: 0.74\nTime: 39.40 secs\n\nEpisode: 9\nMin Score: 0.15 Max Score: 2.43\nScore: 1.07\nAvgScore: 0.78\nTime: 34.10 secs\n\nEpisode: 10\nMin Score: 0.54 Max Score: 2.18\nScore: 1.33\nAvgScore: 0.84\nTime: 37.18 secs\n\nEpisode: 11\nMin Score: 0.59 Max Score: 2.39\nScore: 1.16\nAvgScore: 0.87\nTime: 35.75 secs\n\nEpisode: 12\nMin Score: 0.20 Max Score: 1.96\nScore: 1.18\nAvgScore: 0.89\nTime: 37.73 secs\n\nEpisode: 13\nMin Score: 0.66 Max Score: 3.14\nScore: 1.51\nAvgScore: 0.94\nTime: 36.25 secs\n\nEpisode: 14\nMin Score: 0.85 Max Score: 3.86\nScore: 1.83\nAvgScore: 1.00\nTime: 37.03 secs\n\nEpisode: 15\nMin Score: 0.66 Max Score: 3.93\nScore: 1.86\nAvgScore: 1.06\nTime: 37.83 secs\n\nEpisode: 16\nMin Score: 1.35 Max Score: 3.60\nScore: 2.37\nAvgScore: 1.14\nTime: 37.04 secs\n\nEpisode: 17\nMin Score: 1.07 Max Score: 4.06\nScore: 2.31\nAvgScore: 1.21\nTime: 37.81 secs\n\nEpisode: 18\nMin Score: 0.66 Max Score: 3.93\nScore: 2.06\nAvgScore: 1.26\nTime: 38.25 secs\n\nEpisode: 19\nMin Score: 1.41 Max Score: 4.46\nScore: 2.63\nAvgScore: 1.33\nTime: 39.65 secs\n\nEpisode: 20\nMin Score: 1.25 Max Score: 6.17\nScore: 2.93\nAvgScore: 1.41\nTime: 40.81 secs\n\nEpisode: 21\nMin Score: 0.44 Max Score: 3.66\nScore: 2.35\nAvgScore: 1.45\nTime: 41.49 secs\n\nEpisode: 22\nMin Score: 1.67 Max Score: 7.66\nScore: 3.18\nAvgScore: 1.53\nTime: 41.72 secs\n\nEpisode: 23\nMin Score: 1.06 Max Score: 5.53\nScore: 2.83\nAvgScore: 1.59\nTime: 44.00 secs\n\nEpisode: 24\nMin Score: 1.65 Max Score: 5.56\nScore: 3.31\nAvgScore: 1.66\nTime: 42.30 secs\n\nEpisode: 25\nMin Score: 0.56 Max Score: 5.42\nScore: 3.28\nAvgScore: 1.73\nTime: 46.03 secs\n\nEpisode: 26\nMin Score: 1.71 Max Score: 8.55\nScore: 3.60\nAvgScore: 1.80\nTime: 48.43 secs\n\nEpisode: 27\nMin Score: 2.73 Max Score: 7.45\nScore: 4.23\nAvgScore: 1.89\nTime: 44.83 secs\n\nEpisode: 28\nMin Score: 1.83 Max Score: 6.39\nScore: 3.63\nAvgScore: 1.95\nTime: 44.71 secs\n\nEpisode: 29\nMin Score: 1.57 Max Score: 6.80\nScore: 4.10\nAvgScore: 2.02\nTime: 44.36 secs\n\nEpisode: 30\nMin Score: 2.05 Max Score: 9.52\nScore: 4.06\nAvgScore: 2.09\nTime: 42.52 secs\n\nEpisode: 31\nMin Score: 1.33 Max Score: 13.18\nScore: 5.02\nAvgScore: 2.19\nTime: 43.97 secs\n\nEpisode: 32\nMin Score: 2.95 Max Score: 8.92\nScore: 5.69\nAvgScore: 2.30\nTime: 43.45 secs\n\nEpisode: 33\nMin Score: 2.82 Max Score: 10.30\nScore: 5.14\nAvgScore: 2.38\nTime: 42.20 secs\n\nEpisode: 34\nMin Score: 2.64 Max Score: 7.55\nScore: 4.57\nAvgScore: 2.45\nTime: 47.80 secs\n\nEpisode: 35\nMin Score: 2.53 Max Score: 7.68\nScore: 5.09\nAvgScore: 2.52\nTime: 49.52 secs\n\nEpisode: 36\nMin Score: 2.91 Max Score: 9.05\nScore: 5.26\nAvgScore: 2.60\nTime: 43.95 secs\n\nEpisode: 37\nMin Score: 3.79 Max Score: 9.82\nScore: 5.75\nAvgScore: 2.68\nTime: 44.78 secs\n\nEpisode: 38\nMin Score: 2.75 Max Score: 10.85\nScore: 6.20\nAvgScore: 2.78\nTime: 44.27 secs\n\nEpisode: 39\nMin Score: 3.46 Max Score: 9.37\nScore: 6.35\nAvgScore: 2.87\nTime: 42.11 secs\n\nEpisode: 40\nMin Score: 4.05 Max Score: 13.63\nScore: 7.46\nAvgScore: 2.98\nTime: 43.93 secs\n\nEpisode: 41\nMin Score: 1.36 Max Score: 10.37\nScore: 6.18\nAvgScore: 3.06\nTime: 45.60 secs\n\nEpisode: 42\nMin Score: 4.95 Max Score: 12.64\nScore: 7.58\nAvgScore: 3.17\nTime: 44.02 secs\n\nEpisode: 43\nMin Score: 4.55 Max Score: 10.10\nScore: 7.19\nAvgScore: 3.26\nTime: 44.93 secs\n\nEpisode: 44\nMin Score: 5.55 Max Score: 19.29\nScore: 9.09\nAvgScore: 3.39\nTime: 44.61 secs\n\nEpisode: 45\nMin Score: 4.65 Max Score: 11.35\nScore: 7.86\nAvgScore: 3.49\nTime: 45.62 secs\n\nEpisode: 46\nMin Score: 2.68 Max Score: 11.51\nScore: 8.82\nAvgScore: 3.61\nTime: 43.24 secs\n\nEpisode: 47\nMin Score: 5.90 Max Score: 14.62\nScore: 8.32\nAvgScore: 3.71\nTime: 41.92 secs\n\nEpisode: 48\nMin Score: 1.55 Max Score: 9.99\nScore: 8.21\nAvgScore: 3.80\nTime: 42.70 secs\n\nEpisode: 49\nMin Score: 6.40 Max Score: 16.33\nScore: 9.82\nAvgScore: 3.93\nTime: 43.08 secs\n\nEpisode: 50\nMin Score: 6.77 Max Score: 13.63\nScore: 9.59\nAvgScore: 4.04\nTime: 41.23 secs\n\nEpisode: 51\nMin Score: 4.73 Max Score: 19.88\nScore: 10.15\nAvgScore: 4.16\nTime: 43.42 secs\n\nEpisode: 52\nMin Score: 3.62 Max Score: 13.47\nScore: 9.10\nAvgScore: 4.25\nTime: 42.42 secs\n\nEpisode: 53\nMin Score: 4.61 Max Score: 20.85\nScore: 10.65\nAvgScore: 4.37\nTime: 42.13 secs\n\nEpisode: 54\nMin Score: 7.46 Max Score: 13.28\nScore: 10.11\nAvgScore: 4.48\nTime: 43.94 secs\n\nEpisode: 55\nMin Score: 3.07 Max Score: 13.70\nScore: 10.54\nAvgScore: 4.59\nTime: 42.78 secs\n\nEpisode: 56\nMin Score: 6.88 Max Score: 14.51\nScore: 10.52\nAvgScore: 4.70\nTime: 41.41 secs\n\nEpisode: 57\nMin Score: 7.35 Max Score: 13.78\nScore: 10.59\nAvgScore: 4.80\nTime: 42.80 secs\n\nEpisode: 58\nMin Score: 6.27 Max Score: 14.03\nScore: 10.35\nAvgScore: 4.90\nTime: 42.75 secs\n\nEpisode: 59\nMin Score: 7.94 Max Score: 17.17\nScore: 11.84\nAvgScore: 5.01\nTime: 41.84 secs\n\nEpisode: 60\nMin Score: 6.98 Max Score: 19.02\nScore: 12.26\nAvgScore: 5.13\nTime: 43.00 secs\n\nEpisode: 61\nMin Score: 3.98 Max Score: 17.59\nScore: 11.21\nAvgScore: 5.23\nTime: 43.97 secs\n\nEpisode: 62\nMin Score: 6.04 Max Score: 14.77\nScore: 11.54\nAvgScore: 5.34\nTime: 51.14 secs\n\nEpisode: 63\nMin Score: 9.25 Max Score: 14.70\nScore: 12.12\nAvgScore: 5.44\nTime: 44.36 secs\n\nEpisode: 64\nMin Score: 9.35 Max Score: 21.04\nScore: 12.93\nAvgScore: 5.56\nTime: 44.95 secs\n\nEpisode: 65\nMin Score: 8.49 Max Score: 18.45\nScore: 13.96\nAvgScore: 5.69\nTime: 44.44 secs\n\nEpisode: 66\nMin Score: 9.15 Max Score: 18.09\nScore: 13.33\nAvgScore: 5.81\nTime: 44.63 secs\n\nEpisode: 67\nMin Score: 8.86 Max Score: 17.82\nScore: 13.60\nAvgScore: 5.92\nTime: 48.76 secs\n\nEpisode: 68\nMin Score: 5.60 Max Score: 23.12\nScore: 14.78\nAvgScore: 6.05\nTime: 46.60 secs\n\nEpisode: 69\nMin Score: 11.87 Max Score: 21.97\nScore: 17.11\nAvgScore: 6.21\nTime: 46.17 secs\n\nEpisode: 70\nMin Score: 8.19 Max Score: 28.37\nScore: 15.83\nAvgScore: 6.35\nTime: 43.00 secs\n\nEpisode: 71\nMin Score: 7.53 Max Score: 26.74\nScore: 15.98\nAvgScore: 6.49\nTime: 45.41 secs\n\nEpisode: 72\nMin Score: 7.95 Max Score: 24.17\nScore: 15.34\nAvgScore: 6.61\nTime: 45.33 secs\n\nEpisode: 73\nMin Score: 12.15 Max Score: 19.59\nScore: 15.57\nAvgScore: 6.73\nTime: 46.11 secs\n\nEpisode: 74\nMin Score: 11.77 Max Score: 18.07\nScore: 15.26\nAvgScore: 6.85\nTime: 46.70 secs\n\nEpisode: 75\nMin Score: 12.21 Max Score: 20.48\nScore: 15.67\nAvgScore: 6.96\nTime: 46.30 secs\n\nEpisode: 76\nMin Score: 6.47 Max Score: 25.88\nScore: 16.18\nAvgScore: 7.09\nTime: 44.33 secs\n\nEpisode: 77\nMin Score: 11.56 Max Score: 21.30\nScore: 16.44\nAvgScore: 7.21\nTime: 45.15 secs\n\nEpisode: 78\nMin Score: 10.20 Max Score: 27.41\nScore: 18.56\nAvgScore: 7.35\nTime: 45.23 secs\n\nEpisode: 79\nMin Score: 11.79 Max Score: 21.97\nScore: 16.45\nAvgScore: 7.47\nTime: 44.99 secs\n\nEpisode: 80\nMin Score: 10.74 Max Score: 19.13\nScore: 15.53\nAvgScore: 7.57\nTime: 45.24 secs\n\nEpisode: 81\nMin Score: 10.08 Max Score: 21.44\nScore: 15.15\nAvgScore: 7.66\nTime: 46.19 secs\n\nEpisode: 82\nMin Score: 7.02 Max Score: 21.13\nScore: 16.50\nAvgScore: 7.77\nTime: 46.31 secs\n\nEpisode: 83\nMin Score: 7.98 Max Score: 24.39\nScore: 17.36\nAvgScore: 7.89\nTime: 48.62 secs\n\nEpisode: 84\nMin Score: 7.66 Max Score: 25.46\nScore: 18.06\nAvgScore: 8.01\nTime: 46.74 secs\n\nEpisode: 85\nMin Score: 11.60 Max Score: 25.53\nScore: 18.07\nAvgScore: 8.12\nTime: 48.34 secs\n\nEpisode: 86\nMin Score: 8.98 Max Score: 23.18\nScore: 16.51\nAvgScore: 8.22\nTime: 46.15 secs\n\nEpisode: 87\nMin Score: 12.86 Max Score: 22.65\nScore: 16.51\nAvgScore: 8.32\nTime: 45.49 secs\n\nEpisode: 88\nMin Score: 11.47 Max Score: 21.70\nScore: 16.36\nAvgScore: 8.41\nTime: 51.48 secs\n\nEpisode: 89\nMin Score: 13.27 Max Score: 23.07\nScore: 17.94\nAvgScore: 8.52\nTime: 45.30 secs\n\nEpisode: 90\nMin Score: 14.36 Max Score: 23.60\nScore: 19.16\nAvgScore: 8.63\nTime: 45.80 secs\n\nEpisode: 91\nMin Score: 4.89 Max Score: 24.50\nScore: 17.44\nAvgScore: 8.73\nTime: 42.50 secs\n\nEpisode: 92\nMin Score: 7.79 Max Score: 40.67\nScore: 19.93\nAvgScore: 8.85\nTime: 43.75 secs\n\nEpisode: 93\nMin Score: 8.42 Max Score: 20.21\nScore: 16.26\nAvgScore: 8.93\nTime: 44.87 secs\n\nEpisode: 94\nMin Score: 11.49 Max Score: 23.84\nScore: 19.03\nAvgScore: 9.04\nTime: 44.41 secs\n\nEpisode: 95\nMin Score: 16.09 Max Score: 39.04\nScore: 23.13\nAvgScore: 9.19\nTime: 45.77 secs\n\nEpisode: 96\nMin Score: 8.11 Max Score: 30.16\nScore: 19.08\nAvgScore: 9.29\nTime: 45.12 secs\n\nEpisode: 97\nMin Score: 8.51 Max Score: 27.75\nScore: 18.70\nAvgScore: 9.39\nTime: 44.09 secs\n\nEpisode: 98\nMin Score: 13.44 Max Score: 30.03\nScore: 21.59\nAvgScore: 9.51\nTime: 44.79 secs\n\nEpisode: 99\nMin Score: 11.51 Max Score: 28.98\nScore: 20.82\nAvgScore: 9.63\nTime: 46.71 secs\n\nEpisode: 100\nMin Score: 15.95 Max Score: 26.85\nScore: 21.42\nAvgScore: 9.75\nTime: 44.03 secs\n\nEpisode: 101\nMin Score: 10.72 Max Score: 39.40\nScore: 20.82\nAvgScore: 9.95\nTime: 44.44 secs\n\nEpisode: 102\nMin Score: 14.55 Max Score: 40.75\nScore: 24.58\nAvgScore: 10.19\nTime: 42.95 secs\n\nEpisode: 103\nMin Score: 19.64 Max Score: 29.02\nScore: 24.37\nAvgScore: 10.43\nTime: 44.29 secs\n\nEpisode: 104\nMin Score: 17.70 Max Score: 32.08\nScore: 25.06\nAvgScore: 10.67\nTime: 43.74 secs\n\nEpisode: 105\nMin Score: 21.62 Max Score: 31.77\nScore: 27.94\nAvgScore: 10.94\nTime: 42.94 secs\n\nEpisode: 106\nMin Score: 24.41 Max Score: 32.08\nScore: 27.59\nAvgScore: 11.21\nTime: 43.43 secs\n\nEpisode: 107\nMin Score: 17.03 Max Score: 36.30\nScore: 28.81\nAvgScore: 11.49\nTime: 46.85 secs\n\nEpisode: 108\nMin Score: 24.28 Max Score: 33.83\nScore: 27.93\nAvgScore: 11.76\nTime: 47.73 secs\n\nEpisode: 109\nMin Score: 20.89 Max Score: 34.72\nScore: 28.20\nAvgScore: 12.03\nTime: 45.06 secs\n\nEpisode: 110\nMin Score: 20.78 Max Score: 30.05\nScore: 25.36\nAvgScore: 12.27\nTime: 44.72 secs\n\nEpisode: 111\nMin Score: 22.54 Max Score: 33.54\nScore: 27.60\nAvgScore: 12.53\nTime: 45.49 secs\n\nEpisode: 112\nMin Score: 22.46 Max Score: 36.56\nScore: 28.96\nAvgScore: 12.81\nTime: 48.92 secs\n\nEpisode: 113\nMin Score: 18.91 Max Score: 31.91\nScore: 26.48\nAvgScore: 13.06\nTime: 51.54 secs\n\nEpisode: 114\nMin Score: 22.00 Max Score: 35.06\nScore: 29.00\nAvgScore: 13.33\nTime: 48.24 secs\n\nEpisode: 115\nMin Score: 23.05 Max Score: 34.93\nScore: 27.09\nAvgScore: 13.58\nTime: 43.40 secs\n\nEpisode: 116\nMin Score: 17.15 Max Score: 29.93\nScore: 25.30\nAvgScore: 13.81\nTime: 46.86 secs\n\nEpisode: 117\nMin Score: 23.56 Max Score: 33.46\nScore: 29.35\nAvgScore: 14.08\nTime: 46.21 secs\n\nEpisode: 118\nMin Score: 17.41 Max Score: 32.85\nScore: 27.90\nAvgScore: 14.34\nTime: 45.15 secs\n\nEpisode: 119\nMin Score: 23.36 Max Score: 31.01\nScore: 28.19\nAvgScore: 14.60\nTime: 46.50 secs\n\nEpisode: 120\nMin Score: 22.31 Max Score: 37.82\nScore: 28.21\nAvgScore: 14.85\nTime: 45.96 secs\n\nEpisode: 121\nMin Score: 20.40 Max Score: 36.65\nScore: 29.81\nAvgScore: 15.13\nTime: 44.66 secs\n\nEpisode: 122\nMin Score: 20.82 Max Score: 36.29\nScore: 27.34\nAvgScore: 15.37\nTime: 45.76 secs\n\nEpisode: 123\nMin Score: 21.87 Max Score: 35.60\nScore: 28.30\nAvgScore: 15.62\nTime: 45.02 secs\n\nEpisode: 124\nMin Score: 23.46 Max Score: 38.94\nScore: 30.32\nAvgScore: 15.89\nTime: 44.62 secs\n\nEpisode: 125\nMin Score: 25.24 Max Score: 35.79\nScore: 30.73\nAvgScore: 16.17\nTime: 44.66 secs\n\nEpisode: 126\nMin Score: 23.73 Max Score: 38.79\nScore: 32.11\nAvgScore: 16.45\nTime: 45.12 secs\n\nEpisode: 127\nMin Score: 20.49 Max Score: 34.81\nScore: 28.94\nAvgScore: 16.70\nTime: 44.04 secs\n\nEpisode: 128\nMin Score: 26.87 Max Score: 38.84\nScore: 34.29\nAvgScore: 17.01\nTime: 45.33 secs\n\nEpisode: 129\nMin Score: 23.89 Max Score: 37.16\nScore: 31.13\nAvgScore: 17.28\nTime: 46.90 secs\n\nEpisode: 130\nMin Score: 23.26 Max Score: 37.35\nScore: 31.91\nAvgScore: 17.55\nTime: 47.13 secs\n\nEpisode: 131\nMin Score: 25.15 Max Score: 40.62\nScore: 32.98\nAvgScore: 17.83\nTime: 45.09 secs\n\nEpisode: 132\nMin Score: 22.69 Max Score: 37.38\nScore: 30.20\nAvgScore: 18.08\nTime: 45.59 secs\n\nEpisode: 133\nMin Score: 19.82 Max Score: 38.78\nScore: 31.10\nAvgScore: 18.34\nTime: 42.67 secs\n\nEpisode: 134\nMin Score: 23.29 Max Score: 38.14\nScore: 32.47\nAvgScore: 18.62\nTime: 44.69 secs\n\nEpisode: 135\nMin Score: 22.67 Max Score: 39.96\nScore: 31.32\nAvgScore: 18.88\nTime: 44.20 secs\n\nEpisode: 136\nMin Score: 25.66 Max Score: 38.93\nScore: 33.33\nAvgScore: 19.16\nTime: 45.03 secs\n\nEpisode: 137\nMin Score: 25.98 Max Score: 39.76\nScore: 34.05\nAvgScore: 19.44\nTime: 44.97 secs\n\nEpisode: 138\nMin Score: 24.46 Max Score: 41.26\nScore: 34.06\nAvgScore: 19.72\nTime: 45.35 secs\n\nEpisode: 139\nMin Score: 23.01 Max Score: 36.76\nScore: 31.46\nAvgScore: 19.97\nTime: 44.90 secs\n\nEpisode: 140\nMin Score: 25.30 Max Score: 39.56\nScore: 34.41\nAvgScore: 20.24\nTime: 47.21 secs\n\nEpisode: 141\nMin Score: 22.78 Max Score: 37.19\nScore: 31.13\nAvgScore: 20.49\nTime: 44.30 secs\n\nEpisode: 142\nMin Score: 22.41 Max Score: 38.66\nScore: 34.79\nAvgScore: 20.76\nTime: 44.77 secs\n\nEpisode: 143\nMin Score: 28.23 Max Score: 39.65\nScore: 34.31\nAvgScore: 21.04\nTime: 44.80 secs\n\nEpisode: 144\nMin Score: 21.50 Max Score: 39.93\nScore: 34.71\nAvgScore: 21.29\nTime: 43.75 secs\n\nEpisode: 145\nMin Score: 26.47 Max Score: 41.46\nScore: 35.02\nAvgScore: 21.56\nTime: 45.26 secs\n\nEpisode: 146\nMin Score: 21.88 Max Score: 39.43\nScore: 32.29\nAvgScore: 21.80\nTime: 49.01 secs\n\nEpisode: 147\nMin Score: 24.39 Max Score: 38.51\nScore: 32.40\nAvgScore: 22.04\nTime: 44.23 secs\n\nEpisode: 148\nMin Score: 26.44 Max Score: 39.43\nScore: 33.68\nAvgScore: 22.29\nTime: 42.22 secs\n\nEpisode: 149\nMin Score: 22.86 Max Score: 40.97\nScore: 35.33\nAvgScore: 22.55\nTime: 45.08 secs\n\nEpisode: 150\nMin Score: 16.55 Max Score: 37.42\nScore: 31.85\nAvgScore: 22.77\nTime: 42.24 secs\n\nEpisode: 151\nMin Score: 20.93 Max Score: 38.32\nScore: 30.61\nAvgScore: 22.98\nTime: 47.12 secs\n\nEpisode: 152\nMin Score: 24.01 Max Score: 35.60\nScore: 31.15\nAvgScore: 23.20\nTime: 49.59 secs\n\nEpisode: 153\nMin Score: 21.82 Max Score: 35.26\nScore: 30.90\nAvgScore: 23.40\nTime: 53.89 secs\n\nEpisode: 154\nMin Score: 26.82 Max Score: 36.25\nScore: 31.18\nAvgScore: 23.61\nTime: 49.76 secs\n\nEpisode: 155\nMin Score: 23.64 Max Score: 35.83\nScore: 30.67\nAvgScore: 23.81\nTime: 47.32 secs\n\nEpisode: 156\nMin Score: 25.37 Max Score: 37.46\nScore: 31.82\nAvgScore: 24.02\nTime: 44.01 secs\n\nEpisode: 157\nMin Score: 27.69 Max Score: 36.76\nScore: 32.28\nAvgScore: 24.24\nTime: 43.58 secs\n\nEpisode: 158\nMin Score: 24.11 Max Score: 36.16\nScore: 29.22\nAvgScore: 24.43\nTime: 43.24 secs\n\nEpisode: 159\nMin Score: 26.93 Max Score: 36.23\nScore: 31.66\nAvgScore: 24.63\nTime: 43.16 secs\n\nEpisode: 160\nMin Score: 24.91 Max Score: 36.53\nScore: 30.55\nAvgScore: 24.81\nTime: 42.30 secs\n\nEpisode: 161\nMin Score: 14.46 Max Score: 39.32\nScore: 29.20\nAvgScore: 24.99\nTime: 43.29 secs\n\nEpisode: 162\nMin Score: 19.74 Max Score: 32.42\nScore: 27.41\nAvgScore: 25.15\nTime: 43.74 secs\n\nEpisode: 163\nMin Score: 18.77 Max Score: 36.44\nScore: 30.09\nAvgScore: 25.33\nTime: 42.18 secs\n\nEpisode: 164\nMin Score: 24.89 Max Score: 41.92\nScore: 33.02\nAvgScore: 25.53\nTime: 43.72 secs\n\nEpisode: 165\nMin Score: 21.55 Max Score: 34.05\nScore: 28.45\nAvgScore: 25.67\nTime: 42.62 secs\n\nEpisode: 166\nMin Score: 18.91 Max Score: 35.07\nScore: 27.17\nAvgScore: 25.81\nTime: 43.03 secs\n\nEpisode: 167\nMin Score: 14.43 Max Score: 37.96\nScore: 29.03\nAvgScore: 25.97\nTime: 42.77 secs\n\nEpisode: 168\nMin Score: 21.19 Max Score: 39.64\nScore: 30.60\nAvgScore: 26.12\nTime: 42.51 secs\n\nEpisode: 169\nMin Score: 20.83 Max Score: 35.98\nScore: 28.21\nAvgScore: 26.24\nTime: 43.06 secs\n\nEpisode: 170\nMin Score: 15.88 Max Score: 34.35\nScore: 25.86\nAvgScore: 26.34\nTime: 43.23 secs\n\nEpisode: 171\nMin Score: 18.25 Max Score: 35.91\nScore: 27.28\nAvgScore: 26.45\nTime: 42.75 secs\n\nEpisode: 172\nMin Score: 18.20 Max Score: 32.89\nScore: 25.72\nAvgScore: 26.55\nTime: 46.56 secs\n\nEpisode: 173\nMin Score: 14.71 Max Score: 33.10\nScore: 24.62\nAvgScore: 26.64\nTime: 43.31 secs\n\nEpisode: 174\nMin Score: 18.61 Max Score: 34.74\nScore: 25.93\nAvgScore: 26.75\nTime: 42.35 secs\n\nEpisode: 175\nMin Score: 15.75 Max Score: 30.48\nScore: 23.78\nAvgScore: 26.83\nTime: 43.54 secs\n\nEpisode: 176\nMin Score: 18.55 Max Score: 30.17\nScore: 23.66\nAvgScore: 26.91\nTime: 42.92 secs\n\nEpisode: 177\nMin Score: 18.26 Max Score: 29.49\nScore: 22.27\nAvgScore: 26.96\nTime: 42.75 secs\n\nEpisode: 178\nMin Score: 18.33 Max Score: 33.96\nScore: 26.28\nAvgScore: 27.04\nTime: 43.01 secs\n\nEpisode: 179\nMin Score: 19.06 Max Score: 30.06\nScore: 24.07\nAvgScore: 27.12\nTime: 43.59 secs\n\nEpisode: 180\nMin Score: 19.98 Max Score: 41.14\nScore: 26.38\nAvgScore: 27.23\nTime: 43.34 secs\n\nEpisode: 181\nMin Score: 20.70 Max Score: 36.28\nScore: 27.41\nAvgScore: 27.35\nTime: 43.12 secs\n\nEpisode: 182\nMin Score: 20.54 Max Score: 32.59\nScore: 27.20\nAvgScore: 27.46\nTime: 43.54 secs\n\nEpisode: 183\nMin Score: 21.15 Max Score: 30.36\nScore: 26.05\nAvgScore: 27.54\nTime: 42.94 secs\n\nEpisode: 184\nMin Score: 22.76 Max Score: 30.76\nScore: 26.18\nAvgScore: 27.62\nTime: 43.79 secs\n\nEpisode: 185\nMin Score: 22.09 Max Score: 32.13\nScore: 27.72\nAvgScore: 27.72\nTime: 42.88 secs\n\nEpisode: 186\nMin Score: 20.37 Max Score: 32.78\nScore: 26.96\nAvgScore: 27.83\nTime: 43.21 secs\n\nEpisode: 187\nMin Score: 21.41 Max Score: 32.16\nScore: 26.72\nAvgScore: 27.93\nTime: 43.33 secs\n\nEpisode: 188\nMin Score: 22.31 Max Score: 35.21\nScore: 28.31\nAvgScore: 28.05\nTime: 43.22 secs\n\nEpisode: 189\nMin Score: 25.40 Max Score: 31.71\nScore: 28.88\nAvgScore: 28.16\nTime: 42.48 secs\n\nEpisode: 190\nMin Score: 24.11 Max Score: 33.09\nScore: 29.45\nAvgScore: 28.26\nTime: 43.21 secs\n\nEpisode: 191\nMin Score: 23.57 Max Score: 36.37\nScore: 30.59\nAvgScore: 28.39\nTime: 42.95 secs\n\nEpisode: 192\nMin Score: 22.07 Max Score: 36.50\nScore: 29.97\nAvgScore: 28.49\nTime: 42.81 secs\n\nEpisode: 193\nMin Score: 25.59 Max Score: 36.33\nScore: 31.08\nAvgScore: 28.64\nTime: 43.45 secs\n\nEpisode: 194\nMin Score: 22.22 Max Score: 35.90\nScore: 29.43\nAvgScore: 28.74\nTime: 42.14 secs\n\nEpisode: 195\nMin Score: 25.75 Max Score: 38.80\nScore: 31.39\nAvgScore: 28.83\nTime: 43.15 secs\n\nEpisode: 196\nMin Score: 26.38 Max Score: 36.38\nScore: 30.32\nAvgScore: 28.94\nTime: 44.07 secs\n\nEpisode: 197\nMin Score: 25.61 Max Score: 32.55\nScore: 30.04\nAvgScore: 29.05\nTime: 42.14 secs\n\nEpisode: 198\nMin Score: 22.89 Max Score: 36.42\nScore: 29.06\nAvgScore: 29.13\nTime: 43.21 secs\n\nEpisode: 199\nMin Score: 24.12 Max Score: 34.57\nScore: 30.23\nAvgScore: 29.22\nTime: 43.56 secs\n\nEpisode: 200\nMin Score: 23.11 Max Score: 38.95\nScore: 30.53\nAvgScore: 29.31\nTime: 42.95 secs\n\nEpisode: 201\nMin Score: 26.28 Max Score: 41.12\nScore: 32.51\nAvgScore: 29.43\nTime: 44.54 secs\n\nEpisode: 202\nMin Score: 24.88 Max Score: 35.30\nScore: 29.53\nAvgScore: 29.48\nTime: 42.80 secs\n\nEpisode: 203\nMin Score: 25.08 Max Score: 36.82\nScore: 31.27\nAvgScore: 29.55\nTime: 42.72 secs\n\nEpisode: 204\nMin Score: 26.56 Max Score: 35.46\nScore: 31.45\nAvgScore: 29.61\nTime: 43.32 secs\n\nEpisode: 205\nMin Score: 25.72 Max Score: 36.55\nScore: 31.64\nAvgScore: 29.65\nTime: 42.52 secs\n\nEpisode: 206\nMin Score: 25.21 Max Score: 38.17\nScore: 32.09\nAvgScore: 29.69\nTime: 41.70 secs\n\nEpisode: 207\nMin Score: 26.96 Max Score: 36.96\nScore: 33.31\nAvgScore: 29.74\nTime: 44.44 secs\n\nEpisode: 208\nMin Score: 27.47 Max Score: 39.45\nScore: 33.16\nAvgScore: 29.79\nTime: 42.77 secs\n\nEpisode: 209\nMin Score: 27.84 Max Score: 38.48\nScore: 33.05\nAvgScore: 29.84\nTime: 42.47 secs\n\nEpisode: 210\nMin Score: 31.55 Max Score: 38.08\nScore: 34.12\nAvgScore: 29.93\nTime: 43.25 secs\n\nEpisode: 211\nMin Score: 26.34 Max Score: 38.20\nScore: 32.78\nAvgScore: 29.98\nTime: 42.96 secs\n\nEpisode: 212\nMin Score: 24.03 Max Score: 41.58\nScore: 31.50\nAvgScore: 30.00\nTime: 43.38 secs\n\n"
],
[
"fig = plt.figure()\nax = fig.add_subplot(111)\nplt.plot(np.arange(1, len(scores)+1), scores)\nplt.ylabel('Score')\nplt.xlabel('Episode #')\n\nplt.show()",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
]
] |
4ab513dc1194998062bd1bebaa6c67e4638d7ae6
| 9,488 |
ipynb
|
Jupyter Notebook
|
Contig_size.ipynb
|
AnimalGenomicsETH/bovine-assembly
|
10dcdcb7166b1e7bb30d77549d7acf3693b965dd
|
[
"MIT"
] | 2 |
2021-05-26T08:48:14.000Z
|
2021-09-16T08:47:30.000Z
|
Contig_size.ipynb
|
AnimalGenomicsETH/bovine-assembly
|
10dcdcb7166b1e7bb30d77549d7acf3693b965dd
|
[
"MIT"
] | 1 |
2022-03-17T03:04:44.000Z
|
2022-03-17T03:04:44.000Z
|
Contig_size.ipynb
|
AnimalGenomicsETH/bovine-assembly
|
10dcdcb7166b1e7bb30d77549d7acf3693b965dd
|
[
"MIT"
] | null | null | null | 27.661808 | 527 | 0.447407 |
[
[
[
"%matplotlib widget\nimport pandas as pd\nimport seaborn as sns\nfrom matplotlib import pyplot as plt\n",
"_____no_output_____"
],
[
"from IPython.core.display import HTML\nstyle = \"\"\"\n<style>\ndiv.output_area {\n overflow-x: scroll;\n}\ndiv.output_area img {\n max-width: unset;\n}\n</style>\n\"\"\"\nHTML(style) ",
"_____no_output_____"
],
[
"df = pd.read_csv('breed.lengths.csv')\nreads = {'hifiasm':'HiFi','canu':'HiFi','peregrine':'HiFi','shasta':'ONT','flye':'ONT','raven':'ONT'}\ndf['read'] = [reads[asm] for asm in df['assembler']]\nrefs = {\"1\":158534110,\"2\":136231102,\"3\":121005158,\"4\":120000601,\"5\":120089316,\"6\":117806340,\"7\":110682743,\"8\":113319770,\"9\":105454467,\"10\":103308737,\"11\":106982474,\"12\":87216183,\"13\":83472345,\"14\":82403003,\"15\":85007780,\"16\":81013979,\"17\":73167244,\"18\":65820629,\"19\":63449741,\"20\":71974595,\"21\":69862954,\"22\":60773035,\"23\":52498615,\"24\":62317253,\"25\":42350435,\"26\":51992305,\"27\":45612108,\"28\":45940150,\"29\":51098607,\"MT\":16340,\"X\":139009144,\"Y\":43300181}\npalette = {'hifiasm':'r','canu':'orange','peregrine':'gold','shasta':'b','flye':'c','raven':'purple'}\ndf['delta'] = [l-refs[x] for x,l in zip(df['chromosome'],df['length'])]",
"_____no_output_____"
],
[
"plt.close('all')\n\n#g = sns.FacetGrid(data=df, col='chromosome', col_wrap=6,sharey=False,col_order=refs.keys())\n#for i,m in zip(['HiFi','ONT'],('D','o')):\n# df_x = df[df['read']==i]\n# g.map_dataframe(sns.stripplot,x='assembler',y='length',hue='assembler',marker=m,data=df_x,palette=palette)\n\n#g.add_legend()\ng= sns.catplot(data=df,kind='strip',x='breed',y='delta',col='chromosome',palette=palette,col_wrap=6,hue='assembler',col_order=refs.keys(),legend=True,sharey=False)\nfor c,ax in zip(refs.keys(),g.axes):\n ax.axhline(y=0,c='k',ls='--')#refs[str(c)])\n",
"_____no_output_____"
],
[
"f,ax = plt.subplots()\ndfx = df[(df['assembler']=='hifiasm')|(df['assembler']=='shasta')]\nfor i,m in zip(['HiFi','ONT'],('D','o')):\n df_x = dfx[dfx['read']==i]\n f2=sns.stripplot(data=df_x,x='chromosome',y='delta',hue='breed',marker=m,order=refs.keys(),ax=ax,dodge=True)\n#ax.scatter(refs.keys(),refs.values(),marker='D',ec='k',fc='none')\nax.axhline(y=0)",
"_____no_output_____"
],
[
"df.groupby(['read'])['delta'].mean()",
"_____no_output_____"
],
[
"dfx.groupby(['read'])['delta'].mean()",
"_____no_output_____"
],
[
"df_m = df[df['chromosome']=='MT']\ndf_m.groupby('breed').count()",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab51a36147ad67c23e0626d56b73a19f5e1785b
| 556 |
ipynb
|
Jupyter Notebook
|
Python Types.ipynb
|
JohnCrickett/PythonExample
|
42059d1d58fef0d05de19e3861d15aa057b5bc63
|
[
"MIT"
] | 2 |
2017-05-01T20:06:49.000Z
|
2017-06-10T13:18:23.000Z
|
Python Types.ipynb
|
JohnCrickett/PythonExamples
|
42059d1d58fef0d05de19e3861d15aa057b5bc63
|
[
"MIT"
] | null | null | null |
Python Types.ipynb
|
JohnCrickett/PythonExamples
|
42059d1d58fef0d05de19e3861d15aa057b5bc63
|
[
"MIT"
] | null | null | null | 16.352941 | 34 | 0.521583 |
[
[
[
"# Python Types\nExamples",
"_____no_output_____"
]
]
] |
[
"markdown"
] |
[
[
"markdown"
]
] |
4ab53050d4bc8480df8464774fbbacbcd3ff0621
| 486,670 |
ipynb
|
Jupyter Notebook
|
ces/carbon.ipynb
|
csaladenes/sustainable-energy-transitions
|
84b78141e41cf8564c04e5efa0f388380dfaabf8
|
[
"MIT"
] | 11 |
2017-06-26T16:49:32.000Z
|
2021-03-01T22:45:22.000Z
|
ces/carbon.ipynb
|
csaladenes/set
|
84b78141e41cf8564c04e5efa0f388380dfaabf8
|
[
"MIT"
] | 1 |
2017-04-21T01:55:40.000Z
|
2017-04-21T01:57:06.000Z
|
ces/carbon.ipynb
|
csaladenes/set
|
84b78141e41cf8564c04e5efa0f388380dfaabf8
|
[
"MIT"
] | 7 |
2017-04-21T06:43:58.000Z
|
2022-01-16T03:08:15.000Z
| 1,175.531401 | 198,156 | 0.940428 |
[
[
[
"If not explicitly mentioned otherwise we assume:\n- RCP2.6 scenario or the lowest ppm concentration reported (stabilized around 400-420)\n- Linear phase-out of fossil fuels from model start time (2000-2015) by 2100\n- BAU scenario would lead to RCP6 or higher\n- as it is widely accepcted that in order to obtain RCP2.6, emissions must at least cease or turn into removals in the geological near-term (throughout this century), therefore whenever the carbon price is given in terms of percentage reduction from current levels, a linear 100% reduction is assumed from model start time (2000-2015) by 2100\n- if ranges are reported, the mean is taken\n- if the model reports price in dollar per ton of carbon, it is converted to dollar per ton of carbon dioxide",
"_____no_output_____"
]
],
[
[
"import pandas as pd, numpy as np, matplotlib.pyplot as plt, matplotlib as mpl\n%matplotlib inline\nmpl.style.use('classic')",
"_____no_output_____"
],
[
"d=[]\n#d.append(pd.read_csv('carbon/alberth_hope2006.csv',header=None))\n#d.append(pd.read_csv('carbon/alberth_hope2006_2.csv',header=None))\nd.append(pd.read_csv('carbon/bauer2012.csv',header=None))\nd.append(pd.read_csv('carbon/bauer2012_2a.csv',header=None))\nd.append(pd.read_csv('carbon/bauer2012_2b.csv',header=None))\nd.append(pd.read_csv('carbon/bauer2012_2c.csv',header=None))\nd.append(pd.read_csv('carbon/bosetti2014a.csv',header=None))\nd.append(pd.read_csv('carbon/bosetti2014b.csv',header=None))\nd.append(pd.read_csv('carbon/bosetti2014c.csv',header=None))\nd.append(pd.read_csv('carbon/cai2015.csv',header=None))\nd.append(pd.read_csv('carbon/chen2005.csv',header=None))\nd.append(pd.read_csv('carbon/edmonds_GCAM1994.csv',header=None))\nd.append(pd.read_csv('carbon/kriegler2015_2.csv',header=None))\n#d.append(pd.read_csv('carbon/luderer_REMIND2015.csv',header=None))\nd.append(pd.read_csv('carbon/manne_richels_MERGE2005.csv',header=None))\nd.append(pd.read_csv('carbon/paltsev2005.csv',header=None))\nd.append(pd.read_csv('carbon/russ_POLES2012.csv',header=None))\nd.append(pd.read_csv('carbon/wilkerson2015.csv',header=None))",
"_____no_output_____"
],
[
"from scipy.interpolate import interp1d\nkd=[]\nfd=[]",
"_____no_output_____"
],
[
"for z in range(len(d)):\n kd.append({})\n for i in range(len(d[z][0])):\n if ~np.isnan(d[z][0][i]):\n kd[z][np.round(d[z][0][i],0)]=d[z][1][i]\n fd.append(interp1d(sorted(kd[z].keys()),[kd[z][j] for j in sorted(kd[z].keys())]))",
"_____no_output_____"
],
[
"for z in range(len(d)):\n #plt.scatter(d[z][0],d[z][1])\n years=range(int(min(d[z][0]))+1,int(max(d[z][0]))+1)\n plt.plot(years,fd[z](years))",
"_____no_output_____"
],
[
"labels=['Bauer, Hilaire et al.\\n2012 | REMIND-R',\\\n'Luderer, Bosetti et al.\\n2011 | IMACLIM-R',\\\n'Luderer, Bosetti et al.\\n2011 | REMIND-R',\\\n'Luderer, Bosetti et al.\\n2011 | WITCH',\\\n'Bosetti, Marangoni et al.\\n2015 | GCAM',\\\n'Bosetti, Marangoni et al.\\n2015 | MARKAL US',\\\n'Bosetti, Marangoni et al.\\n2015 | WITCH',\\\n'Cai, Newth et al.\\n2015 | GTEM-C',\\\n'Chen, 2005\\nMARKAL-MACRO',\\\n'Edmonds, Wise, MacCracken\\n1994 | GCAM',\\\n'Kriegler, Petermann, et al.\\n2015 | multiple',\\\n'Manne, Richels\\n2005 | MERGE',\\\n'Paltsev, Reilly et al.\\n2005 | MIT EPPA',\\\n'Russ, Ciscar et al.\\n2009 | POLES',\\\n'Wilkerson, Leibowicz et al.\\n2015 | multiple'\\\n]\nco2=[1,1,1,1,0,0,0,1,0,0,1,0,0,0,1]\nz=14\nplt.scatter(d[z][0],d[z][1])\nyears=range(int(min(d[z][0]))+1,int(max(d[z][0]))+1)\nplt.plot(years,fd[z](years))",
"_____no_output_____"
],
[
"def plotter(ax,x,y,c,l,z=2,zz=2,step=2,w=-50,w2=30):\n yrs=range(x[0]-40,x[len(x)-1]+10)\n maxi=[0,0]\n maxv=-100\n \n #try a few initial values for maximum rsquared\n i=0\n for k in range(1,5):\n p0 = [1., 1., x[len(x)*k/5]]\n fit2 = optimize.leastsq(errfunc,p0,args=(x,y),full_output=True)\n ss_err=(fit2[2]['fvec']**2).sum()\n ss_tot=((y-y.mean())**2).sum()\n rsquared=1-(ss_err/ss_tot)\n if rsquared>maxv:\n maxi=[i,k]\n maxv=rsquared\n\n i=maxi[0]\n k=maxi[1]\n \n p0 = [1., 1., x[len(x)*k/5], -1+i*0.5]\n fit2 = optimize.leastsq(errfunc,p0,args=(x,y),full_output=True)\n ss_err=(fit2[2]['fvec']**2).sum()\n ss_tot=((y-y.mean())**2).sum()\n rsquared=1-(ss_err/ss_tot)\n\n \n ax.scatter(x[::step],y[::step],lw*3,color=c)\n #ax.plot(yrs,logist(fit2[0],yrs),color=\"#006d2c\",lw=lw)\n ax.plot(yrs,logist(fit2[0],yrs),color=\"#444444\",lw=lw)\n #ax.plot(yrs,logist(fit2[0],yrs),color=c,lw=1)\n \n yk=logist([fit2[0][0],fit2[0][1],fit2[0][2],fit2[0][3]],range(3000))\n mint=0\n maxt=3000\n perc=0.1\n for i in range(3000):\n if yk[i]<perc: mint=i\n if yk[i]<1-perc: maxt=i\n \n if z>-1:\n coord=len(x)*z/5\n ax.annotate('$R^2 = '+str(np.round(rsquared,2))+'$\\n'+\\\n '$\\\\alpha = '+str(np.round(fit2[0][0],2))+'$\\n'+\\\n '$\\\\beta = '+str(np.round(fit2[0][1],2))+'$\\n'+\\\n '$\\\\Delta t = '+str(int(maxt-mint))+'$', xy=(yrs[coord], logist(fit2[0],yrs)[coord]),\\\n xycoords='data', \n xytext=(w, w2), textcoords='offset points', color=\"#444444\",\n arrowprops=dict(arrowstyle=\"->\",color='#444444')) \n \n coord=len(x)*zz/5\n ax.annotate(l, xy=(yrs[coord], logist(fit2[0],yrs)[coord]),\\\n xycoords='data', \n xytext=(w, w2), textcoords='offset points',\n arrowprops=dict(arrowstyle=\"->\")) ",
"_____no_output_____"
],
[
"fig, ax = plt.subplots(1,1,subplot_kw=dict(axisbg='#EEEEEE',axisbelow=True),figsize=(10,5))\nlw=2\n\ncolors=[\"#756bb1\",\"#d95f0e\",\"#444444\"]\n\nax.grid(color='white', linestyle='solid')\nax.set_xlabel('Years')\nax.set_ylabel('Carbon tax $[\\$/tonCO_2]$')\nax.set_xlim([2000,2100])\nax.set_ylim([0,5000])\n#ax.set_yscale('log')\nax.set_title('Carbon price estimations from various IAM models',size=13,y=1.04)\n\nloc=[2088,2083,2084,2080,2031,2047,2043,2088,2015,2072,2050,2075,2095,2020,2062]\nlz=[(-70, 20),(-70, 20),(-20, 10),(-40, 20),(-100, 40),(-110, 20),(-130, 20),(-15, 15),\\\n (-70, 20),(-105, 20),(-80, 20),(-60, 12),(-120, -5),(-70, 50),(-30, 7)]\n\nfor z in range(len(d))[:15]:\n #ax.scatter(d[z][0],d[z][1])\n years=range(int(min(d[z][0]))+1,int(max(d[z][0]))+1)\n if (co2[z]==1):k=1\n else: k=44.0/12.0\n ax.plot(years,fd[z](years)*k,lw=lw,color=colors[z%3])\n ax.annotate(labels[z]+str(z), xy=(loc[z],fd[z]([loc[z]])*k),\\\n xycoords='data', \n xytext=lz[z], textcoords='offset points',fontsize=9, color=colors[z%3],\n arrowprops=dict(arrowstyle=\"->\",color=colors[z%3]))\n \n#plt.savefig('ces9.png',bbox_inches = 'tight', pad_inches = 0.1, dpi=150)\nplt.show()",
"_____no_output_____"
],
[
"fig, ax = plt.subplots(1,1,subplot_kw=dict(axisbg='#EEEEEE',axisbelow=True),figsize=(10,5))\nlw=2\n\ncolors=[\"#756bb1\",\"#d95f0e\",\"#444444\"]\n\nax.grid(color='white', linestyle='solid')\nax.set_xlabel('Years')\nax.set_ylabel('$MAC$ $[\\$/tonCO_2]$')\nax.set_xlim([2000,2100])\nax.set_ylim([0,5000])\n#ax.set_yscale('log')\nax.set_title(u'Marginal abatement cost $(MAC)$ estimations from various IAM models',size=13,y=1.04)\n\nloc=[2088,2070,2084,2070,2031,2047,2043,2088,2015,2072,2065,2075,2095,2019,2062]\nlz=[(-60, 20),(-75, 20),(-20, 10),(-70, 20),(-100, 40),(-110, 20),(-130, 20),(-15, 15),\\\n (-70, 20),(-90, 20),(-70, 20),(-70, 12),(-120, -5),(-60, 50),(-30, 7)]\n\nfor z in range(len(d))[:15]:\n #ax.scatter(d[z][0],d[z][1])\n if z not in {0,9,14}:\n years=range(int(min(d[z][0]))+1,int(max(d[z][0]))+1)\n if (co2[z]==1):k=1\n else: k=44.0/12.0\n if z in {3,6,7,12}:\n lw=3\n c=colors[2]\n elif z in {0,1,2,5}: \n lw=1\n c=colors[1]\n else:\n lw=1\n c=colors[0]\n ax.plot(years,fd[z](years)*k,lw=lw,color=c)\n ax.annotate(labels[z], xy=(loc[z],fd[z]([loc[z]])*k),\\\n xycoords='data', \n xytext=lz[z], textcoords='offset points',fontsize=9, color=c,\n arrowprops=dict(arrowstyle=\"->\",color=c))\n \nplt.savefig('ces9b.png',bbox_inches = 'tight', pad_inches = 0.1, dpi=150)\nplt.show()",
"_____no_output_____"
],
[
"for z in range(len(d))[:15]:\n print labels[z]",
"Bauer, Hilaire et al.\n2012 | REMIND-R\nLuderer, Bosetti et al.\n2011 | IMACLIM-R\nLuderer, Bosetti et al.\n2011 | REMIND-R\nLuderer, Bosetti et al.\n2011 | WITCH\nBosetti, Marangoni et al.\n2015 | GCAM\nBosetti, Marangoni et al.\n2015 | MARKAL US\nBosetti, Marangoni et al.\n2015 | WITCH\nCai, Newth et al.\n2015 | GTEM-C\nChen, 2005\nMARKAL-MACRO\nEdmonds, Wise, MacCracken\n1994 | GCAM\nKriegler, Petermann, et al.\n2015 | multiple\nManne, Richels\n2005 | MERGE\nPaltsev, Reilly et al.\n2005 | MIT EPPA\nRuss, Ciscar et al.\n2009 | POLES\nWilkerson, Leibowicz et al.\n2015 | multiple\n"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab534514a166229521a52179ae902e9da7a82d1
| 755,196 |
ipynb
|
Jupyter Notebook
|
examples/select_data.ipynb
|
ocefpaf/xroms
|
763d6e678e28fe074e0aaab26fecd2b74e51a8b0
|
[
"MIT"
] | 4 |
2020-01-21T21:24:17.000Z
|
2020-10-02T03:09:32.000Z
|
examples/select_data.ipynb
|
ocefpaf/xroms
|
763d6e678e28fe074e0aaab26fecd2b74e51a8b0
|
[
"MIT"
] | 1 |
2020-04-08T00:11:39.000Z
|
2020-04-25T08:03:45.000Z
|
examples/select_data.ipynb
|
ocefpaf/xroms
|
763d6e678e28fe074e0aaab26fecd2b74e51a8b0
|
[
"MIT"
] | 1 |
2020-04-06T06:42:36.000Z
|
2020-04-06T06:42:36.000Z
| 130.815174 | 192,764 | 0.773953 |
[
[
[
"import xarray as xr\nimport xroms\nimport pandas as pd\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport cmocean.cm as cmo\nimport cartopy",
"_____no_output_____"
]
],
[
[
"# How to select data\n\nThe [load_data](load_data.ipynb) notebook demonstrates how to load in data, but now how to select out parts of it?",
"_____no_output_____"
],
[
"### Load in data\n\nMore information at in [load_data notebook](load_data.ipynb)",
"_____no_output_____"
]
],
[
[
"loc = 'http://barataria.tamu.edu:8080/thredds/dodsC/forecast_latest/txla2_his_f_latest.nc'\nchunks = {'ocean_time':1}\nds = xr.open_dataset(loc, chunks=chunks)",
"_____no_output_____"
],
[
"# set up grid\nds, grid = xroms.roms_dataset(ds)",
"_____no_output_____"
]
],
[
[
"## Select",
"_____no_output_____"
],
[
"### Slices by index or keyword",
"_____no_output_____"
],
[
"#### Surface layer slice\n\nThe surface in ROMS is given by the last index in the vertical dimension. The easiest way to access this is by indexing into `s_rho`. While normally it is better to access coordinates through keywords to be human-readable, it's not easy to tell what value of `s_rho` gives the surface. In this instance, it's easier to just go by index.",
"_____no_output_____"
]
],
[
[
"ds.salt.isel(s_rho=-1)",
"_____no_output_____"
]
],
[
[
"#### x/y index slice\n\nFor a curvilinear ROMS grid, selecting by the dimensions `xi_rho` or `eta_rho` (or for whichever is the relevant grid) is not very meaningful because they are given by index. Thus the following is possible to get a slice along the index, but it cannot be used to find a slice based on the lon/lat values.",
"_____no_output_____"
]
],
[
[
"ds.temp.sel(xi_rho=20)",
"_____no_output_____"
]
],
[
[
"#### Single time\n\nFind the forecast model output available that is closest to now. Note that the `method` keyword argument is not necessary if the desired date/time is exactly a model output time.",
"_____no_output_____"
]
],
[
[
"now = pd.Timestamp.today()",
"_____no_output_____"
],
[
"ds.salt.isel(s_rho=-1).sel(ocean_time=now, method='nearest')",
"_____no_output_____"
]
],
[
[
"#### Range of time",
"_____no_output_____"
]
],
[
[
"ds.salt.sel(ocean_time=slice(now,now+pd.Timedelta('2 days')))",
"_____no_output_____"
]
],
[
[
"### Calculate slice",
"_____no_output_____"
],
[
"#### Cross-section along a longitude value\n\nBecause the example grid is curvilinear, a slice along a grid dimension is not the same as a slice along a longitude or latitude (or projected $x$/$y$) value. This needs to be calculated and we can use the `xisoslice` function to do this. The calculation is done lazily. We calculate only part of the slice, on the continental shelf. Renaming the subsetted dataset (below, as `dss`) is convenient because this variable can be used in place of `ds` for all related function calls to be consistent and only have to subset one time.",
"_____no_output_____"
]
],
[
[
"# want salinity along this constant value\nlon0 = -91.5\n\n# This is the array we want projected onto the longitude value. \n# Note that we are requesting multiple times at once.\ndss = ds.isel(ocean_time=slice(0,10), eta_rho=slice(50,-1))\n\n# Projecting 3rd input onto constant value lon0 in iso_array ds.lon_rho\nsl = xroms.xisoslice(dss.lon_rho, lon0, dss.salt, 'xi_rho')\n\nsl",
"_____no_output_____"
],
[
"fig, axes = plt.subplots(1, 2, figsize=(15,6))\n\nsl.isel(ocean_time=0).plot(ax=axes[0])\nsl.isel(ocean_time=-1).plot(ax=axes[1])",
"_____no_output_____"
]
],
[
[
"Better plot: use coordinates and one colorbar to compare.",
"_____no_output_____"
]
],
[
[
"# calculate z values (s_rho)\nslz = xroms.xisoslice(dss.lon_rho, lon0, dss.z_rho, 'xi_rho')\n\n# calculate latitude values (eta_rho)\nsllat = xroms.xisoslice(dss.lon_rho, lon0, dss.lat_rho, 'xi_rho')\n\n# assign these as coords to be used in plot\nsl = sl.assign_coords(z=slz, lat=sllat)\n\n# points that should be masked\nslmask = xroms.xisoslice(dss.lon_rho, lon0, dss.mask_rho, 'xi_rho')\n\n# drop masked values\nsl = sl.where(slmask==1, drop=True)\n\n# find min and max of the slice itself (without values that should be masked)\nvmin = sl.min().values\nvmax = sl.max().values\n",
"_____no_output_____"
],
[
"fig, axes = plt.subplots(1, 2, figsize=(15,6), sharey=True)\n\nsl.isel(ocean_time=0).plot(x='lat', y='z', ax=axes[0], vmin=vmin, vmax=vmax, add_colorbar=False)\nmappable = sl.isel(ocean_time=-1).plot(x='lat', y='z', ax=axes[1], vmin=vmin, vmax=vmax, add_colorbar=False)\n\nfig.colorbar(ax=axes, mappable=mappable, orientation='horizontal').set_label('salt')",
"_____no_output_____"
]
],
[
[
"Verify performance of isoslice by comparing slice at surface with planview surface plot.",
"_____no_output_____"
]
],
[
[
"vmin = dss.salt.min().values\nvmax = dss.salt.max().values\n\n\nfig, ax = plt.subplots(1, 1, figsize=(15,15))\nds.salt.isel(ocean_time=0, s_rho=-1).plot(ax=ax, x='lon_rho', y='lat_rho')\nax.scatter(lon0*np.ones_like(sl.lat[::10]), sl.lat[::10], c=sl.isel(ocean_time=0, s_rho=-1)[::10], \n s=100, vmin=vmin, vmax=vmax, zorder=10, edgecolor='k')",
"_____no_output_____"
]
],
[
[
"#### Variable at constant z value",
"_____no_output_____"
]
],
[
[
"# want temperature along this constant depth value\nz0 = -10\n\n# This is the array we want projected \ndss = ds.isel(ocean_time=0)\n\n# Projecting 3rd input onto constant value z0 in iso_array (1st input)\nsl = xroms.xisoslice(dss.z_rho, z0, dss.temp, 's_rho')\n\nsl",
"_____no_output_____"
],
[
"sl.plot(cmap=cmo.thermal, x='lon_rho', y='lat_rho')",
"/Users/kthyng/opt/miniconda3/envs/env/lib/python3.7/site-packages/dask/core.py:121: RuntimeWarning: invalid value encountered in true_divide\n return func(*(_execute_task(a, cache) for a in args))\n"
]
],
[
[
"#### Variable at constant z depth, in time",
"_____no_output_____"
]
],
[
[
"# want temperature along this constant depth value\nz0 = -10\n\n# Projecting 3rd input onto constant value z0 in iso_array (1st input)\nsl = xroms.xisoslice(ds.z_rho, z0, ds.temp, 's_rho')\n\nsl",
"_____no_output_____"
]
],
[
[
"#### zeta at constant z depth, in time\n\n... to verify that xisoslice does act in time across zeta.",
"_____no_output_____"
]
],
[
[
"# want temperature along this constant depth value\nz0 = -10\n\n# Projecting 3rd input onto constant value z0 in iso_array (1st input)\nzeta_s_rho = ds.zeta.expand_dims({'s_rho': ds.s_rho}).transpose('ocean_time','s_rho',...)\nsl = xroms.xisoslice(ds.z_rho, z0, zeta_s_rho, 's_rho')",
"_____no_output_____"
],
[
"sl.sel(eta_rho=30,xi_rho=20).plot()",
"/Users/kthyng/opt/miniconda3/envs/env/lib/python3.7/site-packages/dask/core.py:121: RuntimeWarning: invalid value encountered in true_divide\n return func(*(_execute_task(a, cache) for a in args))\n"
]
],
[
[
"#### Depth of isohaline surface\n\nCalculate the depth of a specific isohaline.\n\nNote that in this case there are a few wonky values, so we should filter them out or control the vmin/vmax values on the plot.",
"_____no_output_____"
]
],
[
[
"# want the depth of this constant salinity value\nS0 = 33\n\n# This is the array we want projected \ndss = ds.isel(ocean_time=0)\n\n# Projecting 3rd input onto constant value z0 in iso_array (1st input)\nsl = xroms.xisoslice(dss.salt, S0, dss.z_rho, 's_rho')\n\nsl.plot(cmap=cmo.deep, x='lon_rho', y='lat_rho', vmin=-20, vmax=0, figsize=(10, 10))",
"/Users/kthyng/opt/miniconda3/envs/env/lib/python3.7/site-packages/dask/core.py:121: RuntimeWarning: invalid value encountered in true_divide\n return func(*(_execute_task(a, cache) for a in args))\n"
]
],
[
[
"### Select region\n\nSelect a boxed region by min/max lon and lat values.",
"_____no_output_____"
]
],
[
[
"# want model output only within the box defined by these lat/lon values\nlon = np.array([-97, -96])\nlat = np.array([28, 29])",
"_____no_output_____"
],
[
"# this condition defines the region of interest\nbox = ((lon[0] < ds.lon_rho) & (ds.lon_rho < lon[1]) & (lat[0] < ds.lat_rho) & (ds.lat_rho < lat[1])).compute()",
"_____no_output_____"
]
],
[
[
"Plot the model output in the box at the surface",
"_____no_output_____"
]
],
[
[
"dss = ds.where(box).salt.isel(s_rho=-1, ocean_time=0)\ndss.plot(x='lon_rho', y='lat_rho')",
"_____no_output_____"
]
],
[
[
"Can calculate a metric within the box:",
"_____no_output_____"
]
],
[
[
"dss.mean().values",
"_____no_output_____"
]
],
[
[
"### Find nearest model output in two dimensions\n\nThis matters for a curvilinear grid.\n\nCan't use `sel` because it will only search in one coordinate for the nearest value and the coordinates are indices which are not necessarily geographic distance. Instead need to use a search for distance and use that for the `where` condition from the previous example.\n\nFind the model output at the grid node nearest the point (lon0, lat0). You can create the projection to use for the distance calculation in `sel2d` and input it into the function, or you can let it choose a default for you.",
"_____no_output_____"
]
],
[
[
"lon0, lat0 = -96, 27\ndl = 0.05\nproj = cartopy.crs.LambertConformal(central_longitude=-98, central_latitude=30)",
"_____no_output_____"
],
[
"dssub = xroms.sel2d(ds, lon0, lat0, proj)",
"_____no_output_____"
]
],
[
[
"Or, if you instead want the indices of the nearest grid node returned, you can call `argsel2d`:",
"_____no_output_____"
]
],
[
[
"ix, iy = xroms.argsel2d(ds, lon0, lat0, proj)",
"_____no_output_____"
]
],
[
[
"Check this function, just to be sure:",
"_____no_output_____"
]
],
[
[
"box = (ds.lon_rho>lon0-dl) & (ds.lon_rho<lon0+dl) & (ds.lat_rho>lat0-dl) & (ds.lat_rho<lat0+dl)\ndss = ds.where(box).salt.isel(ocean_time=0, s_rho=-1)\n\nvmin = dss.min().values\nvmax = dss.max().values\n\ndss.plot(x='lon_rho', y='lat_rho')\nplt.scatter(lon0, lat0, c=dssub.salt.isel(s_rho=-1, ocean_time=0), s=200, edgecolor='k', vmin=vmin, vmax=vmax)\nplt.xlim(lon0-dl,lon0+dl)\nplt.ylim(lat0-dl, lat0+dl)",
"_____no_output_____"
]
],
[
[
"Note that the `sel2d` function returned a time series since that was input, and it worked fine. Getting the numbers take time.",
"_____no_output_____"
]
],
[
[
"dssub.salt.isel(s_rho=-1, ocean_time=slice(0,5)).plot()",
"/Users/kthyng/opt/miniconda3/envs/env/lib/python3.7/site-packages/pandas/plotting/_matplotlib/converter.py:256: MatplotlibDeprecationWarning: \nThe epoch2num function was deprecated in Matplotlib 3.3 and will be removed two minor releases later.\n base = dates.epoch2num(dt.asi8 / 1.0e9)\n"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ab54db17e6ec7c3ec15342074533d94f7fb80f4
| 193,101 |
ipynb
|
Jupyter Notebook
|
Inverse Design/InverseDesign.ipynb
|
mranneberg/viiflow-examples
|
3d1ddb52360199bd674954a04e6e8b69bbb76212
|
[
"MIT"
] | 3 |
2019-05-04T13:38:58.000Z
|
2021-01-02T17:00:29.000Z
|
Inverse Design/InverseDesign.ipynb
|
mranneberg/viiflow-examples
|
3d1ddb52360199bd674954a04e6e8b69bbb76212
|
[
"MIT"
] | null | null | null |
Inverse Design/InverseDesign.ipynb
|
mranneberg/viiflow-examples
|
3d1ddb52360199bd674954a04e6e8b69bbb76212
|
[
"MIT"
] | null | null | null | 55.345658 | 310 | 0.514736 |
[
[
[
"## Viscous Inverse Design\nThis notebook demonstrates the use of gradients from viiflow for fully viscous inverse design.\nIt defines a target pressure distribution from one airfoil and, coming from another airfoil, tries to find the shape necessary to arrive at this target pressure.\nIt uses virtual displacements, which do not necessitate the recalculation of the panel operator.\nInstead, it uses the same model used for the effect of boundary layer thickness onto the flow for modification of the airfoil shape.\n\nThe heart of this notebook is a Gauss-Newton iteration which solves for these virtual displacements.\nInstead of trying to solve the pressure distribution exactly, the iteration sovles a least-squares problem that joins the pressure difference with regularizing terms.\nFully viscous inverse design is not a straightforward problem. There are several ways an optimizer may *cheat*, for example\n\n* The velocity is defined by the inviscid solution of the airfoil shape plus boundary layer thickness. An optimizer can therefore choose to reduce the thickness of the airfoil if for some reason a thick boundary layer leads to the target velocity distribution.\n* Kinks in the desired velocity are, in the case below, due to laminar-turbulent transition. However, an optimizer can choose to model this kink by an actual kink in the airfoil.\n\nTo alleviate this, the pressure error is appended by a regularizing term that penalizes non-smooth displacements - simply by adding $ \\frac{\\mathrm{d}^2}{\\mathrm{d}^2 s} \\delta_{virtual}(s) $ at every point along the airfoil surface coordinate $s$ to the Least-Squares problem.\n\nThe parameters chosen to increrase/decrease the penalties were chosen ad-hoc by trial and error.\nIn addition, the nodes very close to the stagnation point are not modified.\n\nIn addition, the residual $r$ of the viiflow solver itself is added to the Least-Squares problem and scaled such that at convergence its error is sufficiently low.\nEvery iteration then performs for dispalcements $y$ and the viiflow variables $x$\n$$ \ny^{k+1} = y^k - \\lambda {\\Delta y}^k\\\\\nx^{k+1} = x^k - \\lambda {\\Delta x}^k\\\\\n{\\Delta y}^k, {\\Delta x}^k = \\min_{\\Delta y,\\Delta x} \\| F(y^k,x^k) - \\frac{\\partial F}{\\partial y}(y^k,x^k) \\Delta y - \\frac{\\partial F}{\\partial x}(y^k,x^k) \\Delta x\\|^2\\\\\n\\|F(y,x)\\|^2 = \\gamma_{cp}^2\\|ue(y)-ue_{target}\\|^2 + \\gamma_y^2\\| \\frac{\\mathrm{d}^2}{\\mathrm{d}^2 s} y \\|^2 + \\gamma_r^2 \\|r(y,x)\\|^2\n$$\nThis may seem like a large problem, but the effort for solving the overdetermined least-squares problem grows largely with the degrees of freedom, not the amount of equations.\n\nBelow, this procedure is used to morph the S805 airfoil into the S825 airfoil. Even with the regularizing terms, little dips that enforce the laminar-turbulent transition can still be seen when zooming in.\n\nWhile this solves for an airfoil shape of a specified pressure distribution, it is probably not a very smart idea to use this for actual design. A better idea is to use first an inviscid inverse design method, e.g. conformal mapping [1, 2], and remove the discrepancies using a fully viscid iteration.\nThe benefit of this Gauss-Newton approach is how straightforward additional constraints can be included, e.g. only fit the suction side from .1c onwards or fit multiple target distributions at multiple angles of attack.",
"_____no_output_____"
]
],
[
[
"import viiflow as vf\nimport viiflowtools.vf_tools as vft\nimport viiflowtools.vf_plots as vfp\nimport numpy as np\nimport matplotlib\nimport matplotlib.pyplot as plt",
"_____no_output_____"
],
[
"# Analysis Settings\nRE = 1e6\nncrit =5\nMach = 0.0\nalpha = 4.0\n\nN = 300\n\n# Read Airfoils\nBASE = vft.repanel(vft.read_selig(\"S805.dat\"),N,KAPFAC=2)\nTARGET = vft.repanel(vft.read_selig(\"S825.dat\"),N,KAPFAC=2)\n# Solve target for our target cp (or more precisely edge velocity)\ns = vf.setup(Re=RE,Ma=Mach,Ncrit=ncrit,Alpha=alpha)\n\n# Internal iterations\ns.Itermax = 100\n\n# Set-up and initialize based on inviscid panel solution\n[p,bl,x] = vf.init([TARGET],s)\nres = None\ngrad = None\n\n\n# Solve aerodynamic problem of target airfoil\nvf.iter(x,bl,p,s,None,None)\nXT0 = p.foils[0].X[0,:].copy()\nUT = p.gamma_viscid[0:p.foils[0].N].copy()\n\n# Set-up and initialize based on inviscid panel solution\n[p,bl,x0] = vf.init([BASE],s)\nres = None\ngrad = None\n\n# Solve aerodynamic problem of current airfoil and save for later plotting\n[x0,_,res,grad,_] = vf.iter(x0,bl,p,s,None,None)\nXC0 = p.foils[0].X[0,:].copy()\nUC = p.gamma_viscid[0:p.foils[0].N].copy()\n\n# To interpolate from one grid to the next, suction and pressure side must have unique grid points\n# That is why below a grid is created where the pressure side is appended with *-1 at the nose\nXT = XT0.copy()\nXC = XC0.copy()\nXT[np.argmin(XT0)+1::] = 2*XT0[np.argmin(XT0)]-XT0[np.argmin(XT0)+1::]\nXC[np.argmin(XC0)+1::] = 2*XC0[np.argmin(XC0)]-XC0[np.argmin(XC0)+1::]\n\n# Interpolate target pressure onto current airfoil grid\nUT = np.interp(-XC.flatten(),-XT.flatten(),np.asarray(UT[:,0]).flatten())",
"Iteration 10, |res| 0.000092, lam 0.984502\nIteration 7, |res| 0.000093, lam 1.000000\n"
],
[
"# Weighting factors for Gauss-Newton\nfacx = 500 # Penalty for smooth dioscplacement\nfac_err = 5 #Weighting of cp error w.r.t. above penalties\nfac_res = 1e4\ns.Gradients = True\n\nNAERO = x.shape[0]\nNVD = len(XC)\n\n# Set-up and initialize based on inviscid panel solution\n[p,bl,x0] = vf.init([BASE],s)\nres = None\ngrad = None\n\n# Solve aerodynamic problem to convergence\n[x,_,_,_,_] = vf.iter(x0,bl,p,s,None,None)\nfprev = np.inf\n# Find ST and do not change near there\nII = np.logical_and(np.fabs(XT-XT[bl[0].sti])>0.001,p.foils[0].X[0,:].ravel()>np.amin(p.foils[0].X[0,:].ravel()))\nII[0]=False\nII[NVD-1]=False\niter = 0\nlam = 1.0\ny = np.zeros(NVD)\nwhile True:\n iter+=1\n\n # Solve Aerodynamic problem\n s.Itermax = 0\n s.Silent = True\n [_,_,res,grad,gradients] = vf.iter(x,bl,p,s,None,None,[y])\n\n # Residual \n RESy = fac_err*(p.gamma_viscid[0:p.foils[0].N].A1-UT)\n dRESydy = fac_err*gradients.partial.gam_vd[0:NVD,:]\n dRESydx = fac_err*gradients.partial.gam_x[0:NVD,:]\n \n # Penalty for thick boundary layer\n #REGdelta = bl[0].bl_fl.nodes.delta*facx\n #dREGdeltady = gradients.total.delta_vd[0:NVD,:]*facx\n \n # Penalty for smooth displacement\n difforder = 2\n REGdelta = np.diff(y,difforder)*facx\n dREGdeltady = np.diff(np.eye(NVD),difforder,0)*facx\n dREGdeltadx = np.zeros((len(REGdelta),len(x)))\n \n # Gauss-Newton step from all terms\n F = np.r_[RESy,REGdelta,res*fac_res]\n fcurr = np.sqrt(F.T@F)\n \n y0 = y\n fprev = fcurr\n # Find ST and do not change near there\n II = np.logical_and(np.fabs(XT-XT[bl[0].sti])>0.001,p.foils[0].X[0,:].ravel()>np.amin(p.foils[0].X[0,:].ravel()))\n II[0]=False\n II[NVD-1]=False\n \n dFdy = np.r_[dRESydy,dREGdeltady,gradients.partial.res_vd*fac_res]\n dFdx = np.r_[dRESydx,dREGdeltadx,grad*fac_res]\n dF = np.c_[dFdy[:,II],dFdx]\n dX = -np.linalg.lstsq(dF,F,rcond=None)[0]\n dy = dX[0:np.sum(II)]\n dx = dX[np.sum(II)::]\n lam = 1\n \n # Print\n resaero = np.sqrt(np.matmul(res,res.T))\n\n \n\n\n # Ad-hoc Damping\n for k in range(len(dy)):\n lam = np.fmin(lam,0.005/abs(dy[k])) # Do not move virtual displacement more than 1mm\n for k in range(len(x)):\n lam = np.fmin(lam,.2/(abs(dx[k]/x[k])))\n \n print(\"iter %u res p:%f resaero: %f dvd:%f lam:%f\"%(iter, np.sqrt(np.matmul(F,F.T)), \\\n resaero,np.sqrt(np.matmul(dy,dy.T)),lam))\n \n if np.sqrt(np.matmul(dy,dy.T))<1e-4 and resaero<1e-4:\n print(\"Converged\")\n break\n \n if iter>100:\n print(\"Not Converged (iteration)\")\n break\n \n j =0\n for k in np.argwhere(II):\n y[k] += lam*dy[j]\n j+=1\n x += lam*dx\n",
"Iteration 6, |res| 0.000097, lam 1.000000\niter 1 res p:11.894336 resaero: 0.000097 dvd:0.266439 lam:0.140405\niter 2 res p:42.820851 resaero: 0.004158 dvd:0.234428 lam:0.123880\niter 3 res p:85.835201 resaero: 0.008537 dvd:0.206997 lam:0.119415\niter 4 res p:93.909179 resaero: 0.009357 dvd:0.162562 lam:0.064107\niter 5 res p:96.672841 resaero: 0.009638 dvd:0.137987 lam:0.228048\niter 6 res p:99.598568 resaero: 0.009939 dvd:0.126887 lam:0.236911\niter 7 res p:120.501384 resaero: 0.012039 dvd:0.126665 lam:0.237082\niter 8 res p:168.659996 resaero: 0.016861 dvd:0.102393 lam:0.316430\niter 9 res p:131.675839 resaero: 0.013164 dvd:0.075484 lam:0.215973\niter 10 res p:113.098834 resaero: 0.011307 dvd:0.062747 lam:0.508614\niter 11 res p:72.376591 resaero: 0.007235 dvd:0.035579 lam:0.377960\niter 12 res p:64.119543 resaero: 0.006410 dvd:0.025200 lam:0.547697\niter 13 res p:84.933741 resaero: 0.008492 dvd:0.014152 lam:0.502831\niter 14 res p:55.367172 resaero: 0.005535 dvd:0.008584 lam:0.493695\niter 15 res p:41.002317 resaero: 0.004098 dvd:0.005812 lam:0.650318\niter 16 res p:35.989745 resaero: 0.003597 dvd:0.003268 lam:0.771179\niter 17 res p:24.019512 resaero: 0.002399 dvd:0.002626 lam:1.000000\niter 18 res p:11.607647 resaero: 0.001156 dvd:0.000428 lam:1.000000\niter 19 res p:7.275695 resaero: 0.000720 dvd:0.001337 lam:1.000000\niter 20 res p:2.221979 resaero: 0.000198 dvd:0.000219 lam:1.000000\niter 21 res p:1.019085 resaero: 0.000019 dvd:0.000062 lam:1.000000\nConverged\n"
],
[
"%matplotlib inline\n%config InlineBackend.figure_format = 'svg'\nmatplotlib.rcParams['figure.figsize'] = [11, 5.5]\nfig,ax = plt.subplots(1,1)\nax.plot(p.foils[0].X[0,:],np.power(UC,2)-1,'-k')\nax.plot(p.foils[0].X[0,:],np.power(p.gamma_viscid[0:p.foils[0].N].A1,2)-1,'-',color=(0.6,0.6,0.6))\nax.plot(p.foils[0].X[0,:],np.power(UT,2)-1,'2k')\nax.legend(['Initial Pressure','Found Pressure','Target Pressure'])\nxlim = ax.get_xlim()\n\nfig,ax = plt.subplots(1,1)\nlines = None\nax.plot(TARGET[0,:],TARGET[1,:],'2k')\nlines = vfp.plot_geometry(ax,p,bl,lines)\nax.legend(['Target Airfoil','Initial Geometry','Found Geometry'])\nax.set_xlim(xlim)",
"_____no_output_____"
]
],
[
[
"[1] Selig, Michael S., and Mark D. Maughmer. *Generalized multipoint inverse airfoil design.* AIAA journal 30.11 (1992): 2618-2625.\n\n[2] Drela, Mark. *XFOIL: An analysis and design system for low Reynolds number airfoils.* Low Reynolds number aerodynamics. Springer, Berlin, Heidelberg, 1989. 1-12.",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
]
] |
4ab5512896ff1ec59ac6478e146a817a64c75097
| 5,317 |
ipynb
|
Jupyter Notebook
|
bit_manipulation/insert_m_into_n/insert_m_into_n_solution.ipynb
|
benkeesey/interactive-coding-challenges
|
4994452a729f4bcfab5c8a4225f2b5e004b79075
|
[
"Apache-2.0"
] | 27,173 |
2015-07-06T12:36:05.000Z
|
2022-03-31T23:56:41.000Z
|
bit_manipulation/insert_m_into_n/insert_m_into_n_solution.ipynb
|
benkeesey/interactive-coding-challenges
|
4994452a729f4bcfab5c8a4225f2b5e004b79075
|
[
"Apache-2.0"
] | 143 |
2015-07-07T05:13:11.000Z
|
2021-12-07T17:05:54.000Z
|
bit_manipulation/insert_m_into_n/insert_m_into_n_solution.ipynb
|
benkeesey/interactive-coding-challenges
|
4994452a729f4bcfab5c8a4225f2b5e004b79075
|
[
"Apache-2.0"
] | 4,657 |
2015-07-06T13:28:02.000Z
|
2022-03-31T10:11:28.000Z
| 24.615741 | 185 | 0.471695 |
[
[
[
"This notebook was prepared by [Donne Martin](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).",
"_____no_output_____"
],
[
"# Solution Notebook",
"_____no_output_____"
],
[
"## Problem: Given two 16 bit numbers, n and m, and two indices i, j, insert m into n such that m starts at bit j and ends at bit i.\n\n* [Constraints](#Constraints)\n* [Test Cases](#Test-Cases)\n* [Algorithm](#Algorithm)\n* [Code](#Code)\n* [Unit Test](#Unit-Test)",
"_____no_output_____"
],
[
"## Constraints\n\n* Can we assume j > i?\n * Yes\n* Can we assume i through j have enough space for m?\n * Yes\n* Can we assume the inputs are valid?\n * No\n* Can we assume this fits memory?\n * Yes",
"_____no_output_____"
],
[
"## Test Cases\n\n* None as an input -> Exception\n* Negative index for i or j -> Exception\n* General case\n\n<pre>\ni = 2, j = 6\n j i\nn = 0000 0100 0011 1101\nm = 0000 0000 0001 0011\nresult = 0000 0100 0100 1101\n</pre>",
"_____no_output_____"
],
[
"## Algorithm\n\n<pre>\n j i\nn = 0000 0100 0011 1101\nm = 0000 0000 0001 0011\n\nlmask = 1111 1111 1111 1111 -1\nlmask = 1111 1111 1000 0000 -1 << (j + 1)\n\nrmask = 0000 0000 0000 0001 1\nrmask = 0000 0000 0000 0100 1 << i\nrmask = 0000 0000 0000 0011 (1 << i) -1\n\nmask = 1111 1111 1000 0011 lmask | rmask\n\nn = 0000 0100 0011 1101\nmask = 1111 1111 1000 0011 n & mask \n--------------------------------------------------\nn2 = 0000 0100 0000 0001\n\nn2 = 0000 0100 0000 0001\nmask2 = 0000 0000 0100 1100 m << i\n--------------------------------------------------\nresult = 0000 0100 0100 1101 n2 | mask2\n</pre>\n\nComplexity:\n* Time: O(b), where b is the number of bits\n* Space: O(b), where b is the number of bits",
"_____no_output_____"
],
[
"## Code",
"_____no_output_____"
]
],
[
[
"class Bits(object):\n\n def insert_m_into_n(self, m, n, i, j):\n if None in (m, n, i, j):\n raise TypeError('Argument cannot be None')\n if i < 0 or j < 0:\n raise ValueError('Index cannot be negative')\n left_mask = -1 << (j + 1)\n right_mask = (1 << i) - 1\n n_mask = left_mask | right_mask\n # Clear bits from j to i, inclusive\n n_cleared = n & n_mask\n # Shift m into place before inserting it into n\n m_mask = m << i\n return n_cleared | m_mask",
"_____no_output_____"
]
],
[
[
"## Unit Test",
"_____no_output_____"
]
],
[
[
"%%writefile test_insert_m_into_n.py\nimport unittest\n\n\nclass TestBit(unittest.TestCase):\n\n def test_insert_m_into_n(self):\n n = int('0000010000111101', base=2)\n m = int('0000000000010011', base=2)\n expected = int('0000010001001101', base=2)\n bits = Bits()\n self.assertEqual(bits.insert_m_into_n(m, n, i=2, j=6), expected)\n print('Success: test_insert_m_into_n')\n\n\ndef main():\n test = TestBit()\n test.test_insert_m_into_n()\n\n\nif __name__ == '__main__':\n main()",
"Overwriting test_insert_m_into_n.py\n"
],
[
"%run -i test_insert_m_into_n.py",
"Success: test_insert_m_into_n\n"
]
]
] |
[
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4ab5611f0cc925619b3ad28dbb0185461b7864b9
| 16,406 |
ipynb
|
Jupyter Notebook
|
py4e_class3_2.ipynb
|
pbj0812/TIL
|
ab765ceecd348ac56aa1315bca96c2a2c227ea1d
|
[
"MIT"
] | null | null | null |
py4e_class3_2.ipynb
|
pbj0812/TIL
|
ab765ceecd348ac56aa1315bca96c2a2c227ea1d
|
[
"MIT"
] | 3 |
2018-05-05T12:15:23.000Z
|
2018-05-05T12:25:42.000Z
|
py4e_class3_2.ipynb
|
pbj0812/TIL
|
ab765ceecd348ac56aa1315bca96c2a2c227ea1d
|
[
"MIT"
] | null | null | null | 30.494424 | 307 | 0.568268 |
[
[
[
"import socket",
"_____no_output_____"
],
[
"mysock = socket.socket(socket.AF_INET, socket.SOCK_STREAM)\nmysock.connect( ('data.pr4e.org', 80) )",
"_____no_output_____"
],
[
"import socket",
"_____no_output_____"
],
[
"mysock = socket.socket(socket.AF_INET, socket.SOCK_STREAM)\nmysock.connect(('data.pr4e.org', 80))\ncmd = 'GET http://data.pr4e.org/romeo.txt HTTP/1.0\\r\\n\\r\\n'.encode()\nmysock.send(cmd)",
"_____no_output_____"
],
[
"while True:\n data = mysock.recv(512)\n if(len(data) < 1):\n break\n print(data.decode(), end = '')\nmysock.close()",
"HTTP/1.1 200 OK\r\nDate: Sat, 26 May 2018 09:48:08 GMT\r\nServer: Apache/2.4.18 (Ubuntu)\r\nLast-Modified: Sat, 13 May 2017 11:22:22 GMT\r\nETag: \"a7-54f6609245537\"\r\nAccept-Ranges: bytes\r\nContent-Length: 167\r\nCache-Control: max-age=0, no-cache, no-store, must-revalidate\r\nPragma: no-cache\r\nExpires: Wed, 11 Jan 1984 05:00:00 GMT\r\nConnection: close\r\nContent-Type: text/plain\r\n\r\nBut soft what light through yonder window breaks\nIt is the east and Juliet is the sun\nArise fair sun and kill the envious moon\nWho is already sick and pale with grief\n"
],
[
"# ASCII\nprint(ord('H'))\nprint(ord('e'))\nprint(ord('\\n'))",
"72\n101\n10\n"
],
[
"x='이광춘'\nprint(type(x))\nx=u'이광춘'\nprint(type(x))",
"<class 'str'>\n<class 'str'>\n"
],
[
"'GET http://data.pr4e.org/romeo.txt HTTP/1.0\\n\\n'.encode()",
"_____no_output_____"
],
[
"import socket\n\nmysock = socket.socket(socket.AF_INET, socket.SOCK_STREAM)\nmysock.connect(('data.pr4e.org', 80))\ncmd = 'GET http://data.pr4e.org/romeo.txt HTTP/1.0\\n\\n'.encode()\nmysock.send(cmd)",
"_____no_output_____"
],
[
"while True:\n data = mysock.recv(512)\n if (len(data) < 1):\n break\n print(data.decode())\nmysock.close()",
"HTTP/1.1 400 Bad Request\r\nDate: Sat, 26 May 2018 16:10:34 GMT\r\nServer: Apache/2.4.18 (Ubuntu)\r\nContent-Length: 308\r\nConnection: close\r\nContent-Type: text/html; charset=iso-8859-1\r\n\r\n<!DOCTYPE HTML PUBLIC \"-//IETF//DTD HTML 2.0//EN\">\n<html><head>\n<title>400 Bad Request</title>\n</head><body>\n<h1>Bad Request</h1>\n<p>Your browser sent a request that this server could not understand.<br />\n</p>\n<hr>\n<address>Apache/2.4.18 (Ubuntu) Server at do1.dr-chuck.com Port 80</address>\n</body></html>\n\n"
],
[
"import urllib.request, urllib.parse, urllib.error\n\nfhand = urllib.request.urlopen('http://data.pr4e.org/romeo.txt')\nfor line in fhand:\n print(line.decode().strip())",
"But soft what light through yonder window breaks\nIt is the east and Juliet is the sun\nArise fair sun and kill the envious moon\nWho is already sick and pale with grief\n"
],
[
"import urllib.request, urllib.parse, urllib.error\n\nfhand = urllib.request.urlopen('http://data.pr4e.org/romeo.txt')\n\ncounts = dict()\nfor line in fhand:\n words = line.decode().split()\n for word in words:\n counts[word] = counts.get(word, 0) + 1\nprint(counts)",
"{'window': 1, 'It': 1, 'sun': 2, 'Arise': 1, 'already': 1, 'sick': 1, 'what': 1, 'kill': 1, 'fair': 1, 'east': 1, 'breaks': 1, 'light': 1, 'yonder': 1, 'pale': 1, 'But': 1, 'is': 3, 'through': 1, 'envious': 1, 'with': 1, 'the': 3, 'Juliet': 1, 'Who': 1, 'grief': 1, 'moon': 1, 'and': 3, 'soft': 1}\n"
],
[
"import urllib.request, urllib.parse, urllib.error\n\nfhand = urllib.request.urlopen('http://www.dr-chuck.com/page1.htm')\nfor line in fhand:\n print(line.decode().strip())",
"<h1>The First Page</h1>\n<p>\nIf you like, you can switch to the\n<a href=\"http://www.dr-chuck.com/page2.htm\">\nSecond Page</a>.\n</p>\n"
],
[
"import urllib.request, urllib.parse, urllib.error\nfrom bs4 import BeautifulSoup\n\nurl = input('Enter - ')\nhtml = urllib.request.urlopen(url).read()\nsoup = BeautifulSoup(html, 'html.parser')\n\n# Retrieve all of the anchor tags\ntags = soup('a')\nfor tag in tags:\n print(tag.get('href', None))",
"Enter - http://www.facebook.com\n#\n#\nhttps://www.facebook.com/\nhttps://www.facebook.com/recover/initiate?lwv=110\n#\n/legal/terms/update\n/about/privacy/update\n/policies/cookies/\nhttps://www.facebook.com/about/privacy/update?ref=old_policy\n/about/basics\n/help/1561485474074139\n/help/1561485474074139\nhttps://research.facebook.com/\n/about/privacy/update#what-kinds-of-information-do-we-collect\n/about/ads/#568137493302217\n/about/ads\n/help/379220725465972\n/policies/cookies/\n/help/203805466323736\n/help/203805466323736\n/notes/facebook-safety/details-on-social-reporting/196124227075034\n/help/211813265517027\n/help/203805466323736\n/settings?tab=applications\n/help/111814505650678\n/ad_guidelines.php\n/help/769828729705201/\n/me/allactivity\n/help/405183566203254/\n/help/125338004213029\nhttps://l.facebook.com/l.php?u=https%3A%2F%2Ffeedback-form.truste.com%2Fwatchdog%2Frequest&h=ATM2VX0-hHE_H1xaSbTWJr6AlwQgW4urDwExEJlxLBjtXStKKdejuLqx1Anl4FOMb8Zr_dx5lOt9BJPRu7Jj0oCXo-AGrs1Axi7VC43E8lSJgmtjauO71xny38j0FnOb\n/about/basics\n/help/contact/173545232710000\n/help/contact/173545232710000\nhttps://www.facebook.com/legal/terms/update?ref=old_policy\nhttps://www.facebook.com/principles.php\n/help/1561485474074139\n/help/1561485474074139\n/about/privacy/\n/about/privacy/\n/settings/?tab=privacy\n/settings/?tab=applications\n/settings/?tab=privacy\n/settings/?tab=applications\n/about/privacy/\nhttps://developers.facebook.com/docs/\n/help/399224883474207\nhttps://www.facebook.com/payments_terms\nhttps://developers.facebook.com/policy\n/legal/self_service_ads_terms\n/ad_guidelines.php\nhttps://www.facebook.com/page_guidelines.php\nhttps://www.facebook.com/terms/provisions/german/index.php\nhttps://l.facebook.com/l.php?u=https%3A%2F%2Fwww.facebookbrand.com%2Ftrademarks%2F&h=ATMv8vY7eBklD23NZrHW13vmF9mVFaH1qrP77AP5Onq2Xggp1QzHqNDxwytlD9gmu9li2_5oId8-jyy4VoAmaBNB-NxGSy7lwnOBTqZ5jUZFnteCybQ63qW5HW4j-_sR\n/about/privacy/\nhttps://www.facebook.com/payments_terms\n/platform\nhttps://developers.facebook.com/policy/\n/ad_guidelines.php\n/legal/self_service_ads_terms\n/page_guidelines.php#promotionsguidelines\nhttps://l.facebook.com/l.php?u=https%3A%2F%2Fwww.facebookbrand.com%2F&h=ATPA_R7ngujU9bREZ_iy_mCRAFhbQ8h6oia3hg8naQUfMUS8faZ5HAXJRFgN7OlxaikOLcd6YmbMHWHmxJe2y6V5TSpGpwCAf5EJJ8idy2wLGS1oyWs56QRVDN7z9SsN\n/help/399224883474207\nhttps://www.facebook.com/page_guidelines.php\n/communitystandards\n/about/location\n/help/325807937506242\n/help/256333951065527\n#\n/pages/create/?ref_type=registration_form\nhttps://www.facebook.com/\nhttps://vi-vn.facebook.com/\nhttps://id-id.facebook.com/\nhttps://th-th.facebook.com/\nhttps://es-la.facebook.com/\nhttps://zh-cn.facebook.com/\nhttps://ja-jp.facebook.com/\nhttps://pt-br.facebook.com/\nhttps://fr-fr.facebook.com/\nhttps://de-de.facebook.com/\n#\n/r.php\n/login/\nhttps://messenger.com/\n/lite/\n/mobile/?ref=pf\n/find-friends?ref=pf\n/directory/people/\n/directory/pages/\n/places/\n/games/\n/directory/places/\n/directory/marketplace/\n/directory/pages_videos/\n/directory/groups/\n/recipes/\n/sport/\n/look/directory/\nhttp://l.facebook.com/l.php?u=http%3A%2F%2Fmomentsapp.com%2F&h=ATOQef4dXu0oWAutiZQBTRg_8JhbtU34dgK0r04hvLVeSSMC0uEzbqWYb3aZDcES2bnoaGjvwWQ9RT462IljVga26PB9CWyQ2oIXfucqfEJSP0ITqod_dlIWt56If3EI\nhttps://l.facebook.com/l.php?u=https%3A%2F%2Finstagram.com%2F&h=ATOv5qb5ZnzxKh0hV9eB0vlA7L5lSpa1ku0aQKUGNu63vIz6F4HHWuLyd4dKm5BPxHNGZ3OnbP0lRxid37slP1yYKNtupwJ8zK0zxgxcQf4whJsbcSDirWGh-tfEwPMy\n/local/lists/245019872666104/\n/facebook\n/ad_campaign/landing.php?placement=pflo&campaign_id=402047449186&extra_1=auto\n/pages/create/?ref_type=sitefooter\nhttps://developers.facebook.com/?ref=pf\n/careers/?ref=pf\n/privacy/explanation\n/policies/cookies/\nhttps://www.facebook.com/help/568137493302217\n/policies?ref=pf\n/help/?ref=pf\n/settings\n/allactivity?privacy_source=activity_log_top_menu\n"
],
[
"# To run this, you can install BeautifulSoup\n# https://pypi.python.org/pypi/beautifulsoup4\n\n# Or download the file\n# http://www.py4e.com/code3/bs4.zip\n# and unzip it in the same directory as this file\n\nimport urllib.request, urllib.parse, urllib.error\nfrom bs4 import BeautifulSoup\nimport ssl\n\n# Ignore SSL certificate errors\nctx = ssl.create_default_context()\nctx.check_hostname = False\nctx.verify_mode = ssl.CERT_NONE\n\nurl = input('Enter - ')\nhtml = urllib.request.urlopen(url, context=ctx).read()\nsoup = BeautifulSoup(html, 'html.parser')\n\n# Retrieve all of the anchor tags\ntags = soup('a')\nfor tag in tags:\n print(tag.get('href', None))",
"Enter - http://www.dr-chuck.com/\nhttps://www.dr-chuck.com/csev-blog/\nhttps://www.si.umich.edu/\nhttps://www.ratemyprofessors.com/ShowRatings.jsp?tid=1159280\nhttps://www.dr-chuck.com/csev-blog/\nhttps://www.twitter.com/drchuck/\nhttps://www.dr-chuck.com/dr-chuck/resume/speaking.htm\nhttps://www.slideshare.net/csev\n/dr-chuck/resume/index.htm\nhttps://amzn.to/1K5Q81K\nhttp://afs.dr-chuck.com/papers/\nhttps://itunes.apple.com/us/podcast/computing-conversations/id731495760\nhttps://www.youtube.com/playlist?list=PLHJB2bhmgB7dFuY7HmrXLj5BmHGKTD-3R\nhttps://developers.imsglobal.org/\nhttps://www.youtube.com/user/csev\nhttps://vimeo.com/drchuck/videos\nhttps://backpack.openbadges.org/share/4f76699ddb399d162a00b89a452074b3/\nhttps://www.linkedin.com/in/charlesseverance/\nhttps://www.researchgate.net/profile/Charles_Severance/\nhttps://www.tsugicloud.org/\n/office\nhttps://www.coursera.org/course/pythonlearn\nhttps://www.coursera.org/course/insidetheinternet\nhttps://open.umich.edu/education/si/si502/winter2009/\nhttp://www.pythonlearn.com\nhttp://www.php-intro.com/\nhttp://www.appenginelearn.com/\nhttp://www.pythonlearn.com/\n/sakai-book\nhttp://www.amazon.com/gp/product/1624311393/ref=as_li_ss_tl?ie=UTF8&camp=1789&creative=390957&creativeASIN=1624311393&linkCode=as2&tag=drchu02-20\nhttp://www.amazon.com/gp/product/059680069X/ref=as_li_ss_tl?ie=UTF8&camp=1789&creative=390957&creativeASIN=059680069X&linkCode=as2&tag=drchu02-20\nhttp://www.amazon.com/Performance-Computing-Architectures-Optimization-Benchmarks/dp/156592312X/\nhttp://oreilly.com/catalog/9781565923126/\nhttp://cnx.org/content/col11136/latest/\nhttp://www.youtube.com/playlist?list=PLHJB2bhmgB7dFuY7HmrXLj5BmHGKTD-3R\nhttps://www.vimeo.com/17207620\nhttps://www.youtube.com/watch?v=BVKpW02hsrU\nhttps://www.youtube.com/watch?v=sa2WsgCvn7c\nhttps://www.vimeo.com/17213019\nhttps://www.youtube.com/watch?v=FJ078sO35M0\nhttp://afs.dr-chuck.com/citoolkit\nhttps://www.sakaiproject.org/\nhttps://www.tsugi.org/\nhttps://developers.imsglobal.org/\n/obi-sample\nhttps://twitter.com/drchuck\n"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab56cb3738bfb3f9b43641e6452ff2be2e95e3d
| 22,034 |
ipynb
|
Jupyter Notebook
|
analysis/eda_twconv_word_count.ipynb
|
johnnytorres/recsys_twconv_s2s
|
864524915988ad38e664bb7843800ec714407702
|
[
"MIT"
] | null | null | null |
analysis/eda_twconv_word_count.ipynb
|
johnnytorres/recsys_twconv_s2s
|
864524915988ad38e664bb7843800ec714407702
|
[
"MIT"
] | 2 |
2020-09-26T00:44:27.000Z
|
2022-02-10T01:13:05.000Z
|
analysis/eda_twconv_word_count.ipynb
|
johnnytorres/recsys_twconv_s2s
|
864524915988ad38e664bb7843800ec714407702
|
[
"MIT"
] | null | null | null | 173.496063 | 9,944 | 0.915767 |
[
[
[
"%matplotlib inline\nimport matplotlib.pyplot as plt\nimport pandas as pd\nimport numpy as np\nimport os",
"_____no_output_____"
],
[
"def get_word_counts(dataset):\n data_dir=f'~/data/twconv/{dataset}'\n data_dir=os.path.expanduser(data_dir)\n fpath = os.path.join(data_dir, 'alldata', 'train.csv')\n print(fpath)\n train = pd.read_csv(fpath)\n train['source_wcount'] = train.source.apply(lambda x: len(x.split(' ')))\n train['target_wcount'] = train.target.apply(lambda x: len(x.split(' ')))\n plt.figure(figsize=(20,8))\n plt.subplot(231)\n train['source_wcount'].plot.hist(bins=50)\n plt.yscale('log')\n plt.subplot(232)\n train['target_wcount'].plot.hist(bins=50)\n plt.yscale('log')",
"_____no_output_____"
],
[
"get_word_counts('twconv_2011_trec')",
"/Users/johnnytorres/data/twconv/twconv_2011_trec/alldata/train.csv\n"
],
[
"get_word_counts('twconv_2016_usersec')",
"/Users/johnnytorres/data/twconv/twconv_2016_usersec/alldata/train.csv\n"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code"
]
] |
4ab58aff5c84e6fa57e406306a79045f655e66b1
| 523,130 |
ipynb
|
Jupyter Notebook
|
3_ml_start_knn_examples/approximate_nearest_neighbors.ipynb
|
codingmoh/alw
|
91884cf58970e86aa5618668bfde485cfb050273
|
[
"MIT"
] | null | null | null |
3_ml_start_knn_examples/approximate_nearest_neighbors.ipynb
|
codingmoh/alw
|
91884cf58970e86aa5618668bfde485cfb050273
|
[
"MIT"
] | null | null | null |
3_ml_start_knn_examples/approximate_nearest_neighbors.ipynb
|
codingmoh/alw
|
91884cf58970e86aa5618668bfde485cfb050273
|
[
"MIT"
] | null | null | null | 1,260.554217 | 496,600 | 0.949991 |
[
[
[
"! pip install annoy nmslib",
"Requirement already satisfied: nmslib in c:\\users\\moh\\anaconda3\\lib\\site-packages (2.1.1)\nRequirement already satisfied: psutil in c:\\users\\moh\\anaconda3\\lib\\site-packages (from nmslib) (5.7.0)\nRequirement already satisfied: numpy>=1.10.0; python_version >= \"3.5\" in c:\\users\\moh\\anaconda3\\lib\\site-packages (from nmslib) (1.18.5)\nRequirement already satisfied: pybind11<2.6.2 in c:\\users\\moh\\anaconda3\\lib\\site-packages (from nmslib) (2.6.1)\n"
],
[
"%matplotlib inline",
"_____no_output_____"
]
],
[
[
"\n# Approximate nearest neighbors in TSNE\n\nThis example presents how to chain KNeighborsTransformer and TSNE in a\npipeline. It also shows how to wrap the packages `annoy` and `nmslib` to\nreplace KNeighborsTransformer and perform approximate nearest neighbors.\nThese packages can be installed with `pip install annoy nmslib`.\n\nNote: Currently `TSNE(metric='precomputed')` does not modify the precomputed\ndistances, and thus assumes that precomputed euclidean distances are squared.\nIn future versions, a parameter in TSNE will control the optional squaring of\nprecomputed distances (see #12401).\n\nNote: In KNeighborsTransformer we use the definition which includes each\ntraining point as its own neighbor in the count of `n_neighbors`, and for\ncompatibility reasons, one extra neighbor is computed when\n`mode == 'distance'`. Please note that we do the same in the proposed wrappers.\n\nSample output::\n\n Benchmarking on MNIST_2000:\n ---------------------------\n AnnoyTransformer: 0.583 sec\n NMSlibTransformer: 0.321 sec\n KNeighborsTransformer: 1.225 sec\n TSNE with AnnoyTransformer: 4.903 sec\n TSNE with NMSlibTransformer: 5.009 sec\n TSNE with KNeighborsTransformer: 6.210 sec\n TSNE with internal NearestNeighbors: 6.365 sec\n\n Benchmarking on MNIST_10000:\n ----------------------------\n AnnoyTransformer: 4.457 sec\n NMSlibTransformer: 2.080 sec\n KNeighborsTransformer: 30.680 sec\n TSNE with AnnoyTransformer: 30.225 sec\n TSNE with NMSlibTransformer: 43.295 sec\n TSNE with KNeighborsTransformer: 64.845 sec\n TSNE with internal NearestNeighbors: 64.984 sec\n",
"_____no_output_____"
]
],
[
[
"# Author: Tom Dupre la Tour\n#\n# License: BSD 3 clause\nimport time\nimport sys\n\ntry:\n import annoy\nexcept ImportError:\n print(\"The package 'annoy' is required to run this example.\")\n sys.exit()\n\ntry:\n import nmslib\nexcept ImportError:\n print(\"The package 'nmslib' is required to run this example.\")\n sys.exit()\n\nimport numpy as np\nimport matplotlib.pyplot as plt\nfrom matplotlib.ticker import NullFormatter\nfrom scipy.sparse import csr_matrix\n\nfrom sklearn.base import BaseEstimator, TransformerMixin\nfrom sklearn.neighbors import KNeighborsTransformer\nfrom sklearn.utils._testing import assert_array_almost_equal\nfrom sklearn.datasets import fetch_openml\nfrom sklearn.pipeline import make_pipeline\nfrom sklearn.manifold import TSNE\nfrom sklearn.utils import shuffle\n\nprint(__doc__)\n\n\nclass NMSlibTransformer(TransformerMixin, BaseEstimator):\n \"\"\"Wrapper for using nmslib as sklearn's KNeighborsTransformer\"\"\"\n\n def __init__(self, n_neighbors=5, metric='euclidean', method='sw-graph',\n n_jobs=1):\n self.n_neighbors = n_neighbors\n self.method = method\n self.metric = metric\n self.n_jobs = n_jobs\n\n def fit(self, X):\n self.n_samples_fit_ = X.shape[0]\n\n # see more metric in the manual\n # https://github.com/nmslib/nmslib/tree/master/manual\n space = {\n 'sqeuclidean': 'l2',\n 'euclidean': 'l2',\n 'cosine': 'cosinesimil',\n 'l1': 'l1',\n 'l2': 'l2',\n }[self.metric]\n\n self.nmslib_ = nmslib.init(method=self.method, space=space)\n self.nmslib_.addDataPointBatch(X)\n self.nmslib_.createIndex()\n return self\n\n def transform(self, X):\n n_samples_transform = X.shape[0]\n\n # For compatibility reasons, as each sample is considered as its own\n # neighbor, one extra neighbor will be computed.\n n_neighbors = self.n_neighbors + 1\n\n results = self.nmslib_.knnQueryBatch(X, k=n_neighbors,\n num_threads=self.n_jobs)\n indices, distances = zip(*results)\n indices, distances = np.vstack(indices), np.vstack(distances)\n\n if self.metric == 'sqeuclidean':\n distances **= 2\n\n indptr = np.arange(0, n_samples_transform * n_neighbors + 1,\n n_neighbors)\n kneighbors_graph = csr_matrix((distances.ravel(), indices.ravel(),\n indptr), shape=(n_samples_transform,\n self.n_samples_fit_))\n\n return kneighbors_graph\n\n\nclass AnnoyTransformer(TransformerMixin, BaseEstimator):\n \"\"\"Wrapper for using annoy.AnnoyIndex as sklearn's KNeighborsTransformer\"\"\"\n\n def __init__(self, n_neighbors=5, metric='euclidean', n_trees=10,\n search_k=-1):\n self.n_neighbors = n_neighbors\n self.n_trees = n_trees\n self.search_k = search_k\n self.metric = metric\n\n def fit(self, X):\n self.n_samples_fit_ = X.shape[0]\n metric = self.metric if self.metric != 'sqeuclidean' else 'euclidean'\n self.annoy_ = annoy.AnnoyIndex(X.shape[1], metric=metric)\n for i, x in enumerate(X):\n self.annoy_.add_item(i, x.tolist())\n self.annoy_.build(self.n_trees)\n return self\n\n def transform(self, X):\n return self._transform(X)\n\n def fit_transform(self, X, y=None):\n return self.fit(X)._transform(X=None)\n\n def _transform(self, X):\n \"\"\"As `transform`, but handles X is None for faster `fit_transform`.\"\"\"\n\n n_samples_transform = self.n_samples_fit_ if X is None else X.shape[0]\n\n # For compatibility reasons, as each sample is considered as its own\n # neighbor, one extra neighbor will be computed.\n n_neighbors = self.n_neighbors + 1\n\n indices = np.empty((n_samples_transform, n_neighbors),\n dtype=int)\n distances = np.empty((n_samples_transform, n_neighbors))\n\n if X is None:\n for i in range(self.annoy_.get_n_items()):\n ind, dist = self.annoy_.get_nns_by_item(\n i, n_neighbors, self.search_k, include_distances=True)\n\n indices[i], distances[i] = ind, dist\n else:\n for i, x in enumerate(X):\n indices[i], distances[i] = self.annoy_.get_nns_by_vector(\n x.tolist(), n_neighbors, self.search_k,\n include_distances=True)\n\n if self.metric == 'sqeuclidean':\n distances **= 2\n\n indptr = np.arange(0, n_samples_transform * n_neighbors + 1,\n n_neighbors)\n kneighbors_graph = csr_matrix((distances.ravel(), indices.ravel(),\n indptr), shape=(n_samples_transform,\n self.n_samples_fit_))\n\n return kneighbors_graph\n\n\ndef test_transformers():\n \"\"\"Test that AnnoyTransformer and KNeighborsTransformer give same results\n \"\"\"\n X = np.random.RandomState(42).randn(10, 2)\n\n knn = KNeighborsTransformer()\n Xt0 = knn.fit_transform(X)\n\n ann = AnnoyTransformer()\n Xt1 = ann.fit_transform(X)\n\n nms = NMSlibTransformer()\n Xt2 = nms.fit_transform(X)\n\n assert_array_almost_equal(Xt0.toarray(), Xt1.toarray(), decimal=5)\n assert_array_almost_equal(Xt0.toarray(), Xt2.toarray(), decimal=5)\n\n\ndef load_mnist(n_samples):\n \"\"\"Load MNIST, shuffle the data, and return only n_samples.\"\"\"\n mnist = fetch_openml(\"mnist_784\")\n X, y = shuffle(mnist.data, mnist.target, random_state=2)\n return X[:n_samples] / 255, y[:n_samples]\n\n\ndef run_benchmark():\n datasets = [\n ('MNIST_2000', load_mnist(n_samples=2000)),\n ('MNIST_10000', load_mnist(n_samples=10000)),\n ]\n\n n_iter = 500\n perplexity = 30\n # TSNE requires a certain number of neighbors which depends on the\n # perplexity parameter.\n # Add one since we include each sample as its own neighbor.\n n_neighbors = int(3. * perplexity + 1) + 1\n\n transformers = [\n ('AnnoyTransformer', AnnoyTransformer(n_neighbors=n_neighbors,\n metric='sqeuclidean')),\n ('NMSlibTransformer', NMSlibTransformer(n_neighbors=n_neighbors,\n metric='sqeuclidean')),\n ('KNeighborsTransformer', KNeighborsTransformer(\n n_neighbors=n_neighbors, mode='distance', metric='sqeuclidean')),\n ('TSNE with AnnoyTransformer', make_pipeline(\n AnnoyTransformer(n_neighbors=n_neighbors, metric='sqeuclidean'),\n TSNE(metric='precomputed', perplexity=perplexity,\n method=\"barnes_hut\", random_state=42, n_iter=n_iter), )),\n ('TSNE with NMSlibTransformer', make_pipeline(\n NMSlibTransformer(n_neighbors=n_neighbors, metric='sqeuclidean'),\n TSNE(metric='precomputed', perplexity=perplexity,\n method=\"barnes_hut\", random_state=42, n_iter=n_iter), )),\n ('TSNE with KNeighborsTransformer', make_pipeline(\n KNeighborsTransformer(n_neighbors=n_neighbors, mode='distance',\n metric='sqeuclidean'),\n TSNE(metric='precomputed', perplexity=perplexity,\n method=\"barnes_hut\", random_state=42, n_iter=n_iter), )),\n ('TSNE with internal NearestNeighbors',\n TSNE(metric='sqeuclidean', perplexity=perplexity, method=\"barnes_hut\",\n random_state=42, n_iter=n_iter)),\n ]\n\n # init the plot\n nrows = len(datasets)\n ncols = np.sum([1 for name, model in transformers if 'TSNE' in name])\n fig, axes = plt.subplots(nrows=nrows, ncols=ncols, squeeze=False,\n figsize=(5 * ncols, 4 * nrows))\n axes = axes.ravel()\n i_ax = 0\n\n for dataset_name, (X, y) in datasets:\n\n msg = 'Benchmarking on %s:' % dataset_name\n print('\\n%s\\n%s' % (msg, '-' * len(msg)))\n\n for transformer_name, transformer in transformers:\n start = time.time()\n Xt = transformer.fit_transform(X)\n duration = time.time() - start\n\n # print the duration report\n longest = np.max([len(name) for name, model in transformers])\n whitespaces = ' ' * (longest - len(transformer_name))\n print('%s: %s%.3f sec' % (transformer_name, whitespaces, duration))\n\n # plot TSNE embedding which should be very similar across methods\n if 'TSNE' in transformer_name:\n axes[i_ax].set_title(transformer_name + '\\non ' + dataset_name)\n axes[i_ax].scatter(Xt[:, 0], Xt[:, 1], c=y.astype(np.int32),\n alpha=0.2, cmap=plt.cm.viridis)\n axes[i_ax].xaxis.set_major_formatter(NullFormatter())\n axes[i_ax].yaxis.set_major_formatter(NullFormatter())\n axes[i_ax].axis('tight')\n i_ax += 1\n\n fig.tight_layout()\n plt.show()\n\n\nif __name__ == '__main__':\n test_transformers()\n run_benchmark()",
"Automatically created module for IPython interactive environment\n\nBenchmarking on MNIST_2000:\n---------------------------\nAnnoyTransformer: 2.460 sec\nNMSlibTransformer: 0.173 sec\nKNeighborsTransformer: 2.206 sec\nTSNE with AnnoyTransformer: 7.510 sec\nTSNE with NMSlibTransformer: 5.929 sec\nTSNE with KNeighborsTransformer: 8.114 sec\nTSNE with internal NearestNeighbors: 9.213 sec\n\nBenchmarking on MNIST_10000:\n----------------------------\nAnnoyTransformer: 12.359 sec\nNMSlibTransformer: 1.494 sec\nKNeighborsTransformer: 52.123 sec\nTSNE with AnnoyTransformer: 51.923 sec\nTSNE with NMSlibTransformer: 49.194 sec\n"
]
]
] |
[
"code",
"markdown",
"code"
] |
[
[
"code",
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ab59fbb12168713cd4927f611d5b90d4c328b26
| 24,159 |
ipynb
|
Jupyter Notebook
|
site/ja/beta/tutorials/keras/feature_columns.ipynb
|
crypdra/docs
|
41ab06fd14b3a3dff933bb80b19ce46c7c5781cf
|
[
"Apache-2.0"
] | 2 |
2019-10-25T18:51:16.000Z
|
2019-10-25T18:51:18.000Z
|
site/ja/beta/tutorials/keras/feature_columns.ipynb
|
crypdra/docs
|
41ab06fd14b3a3dff933bb80b19ce46c7c5781cf
|
[
"Apache-2.0"
] | null | null | null |
site/ja/beta/tutorials/keras/feature_columns.ipynb
|
crypdra/docs
|
41ab06fd14b3a3dff933bb80b19ce46c7c5781cf
|
[
"Apache-2.0"
] | null | null | null | 32.428188 | 544 | 0.55321 |
[
[
[
"##### Copyright 2019 The TensorFlow Authors.",
"_____no_output_____"
]
],
[
[
"#@title Licensed under the Apache License, Version 2.0 (the \"License\");\n# you may not use this file except in compliance with the License.\n# You may obtain a copy of the License at\n#\n# https://www.apache.org/licenses/LICENSE-2.0\n#\n# Unless required by applicable law or agreed to in writing, software\n# distributed under the License is distributed on an \"AS IS\" BASIS,\n# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.\n# See the License for the specific language governing permissions and\n# limitations under the License.",
"_____no_output_____"
]
],
[
[
"# 構造化されたデータの分類",
"_____no_output_____"
],
[
"<table class=\"tfo-notebook-buttons\" align=\"left\">\n <td>\n <a target=\"_blank\" href=\"https://www.tensorflow.org/beta/tutorials/keras/feature_columns\">\n <img src=\"https://www.tensorflow.org/images/tf_logo_32px.png\" />\n View on TensorFlow.org</a>\n </td>\n <td>\n <a target=\"_blank\" href=\"https://colab.research.google.com/github/tensorflow/docs/blob/master/site/ja/beta/tutorials/keras/feature_columns.ipynb\">\n <img src=\"https://www.tensorflow.org/images/colab_logo_32px.png\" />\n Run in Google Colab</a>\n </td>\n <td>\n <a target=\"_blank\" href=\"https://github.com/tensorflow/docs/blob/master/site/ja/beta/tutorials/keras/feature_columns.ipynb\">\n <img src=\"https://www.tensorflow.org/images/GitHub-Mark-32px.png\" />\n View source on GitHub</a>\n </td>\n</table>",
"_____no_output_____"
],
[
"Note: これらのドキュメントは私たちTensorFlowコミュニティが翻訳したものです。コミュニティによる 翻訳は**ベストエフォート**であるため、この翻訳が正確であることや[英語の公式ドキュメント](https://www.tensorflow.org/?hl=en)の 最新の状態を反映したものであることを保証することはできません。 この翻訳の品質を向上させるためのご意見をお持ちの方は、GitHubリポジトリ[tensorflow/docs](https://github.com/tensorflow/docs)にプルリクエストをお送りください。 コミュニティによる翻訳やレビューに参加していただける方は、 [[email protected] メーリングリスト](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ja)にご連絡ください。",
"_____no_output_____"
],
[
"このチュートリアルでは、(例えばCSVファイルに保存された表形式データのような)構造化されたデータをどうやって分類するかを示します。ここでは、モデルの定義に[Keras](https://www.tensorflow.org/guide/keras)を、[feature columns](https://www.tensorflow.org/guide/feature_columns)をCSVファイルの列をモデルを訓練するための特徴量にマッピングするための橋渡し役として使用します。このチュートリアルには、下記のことを行うコードすべてが含まれています。\n\n* [Pandas](https://pandas.pydata.org/)を使用したCSVファイルの読み込み\n* [tf.data](https://www.tensorflow.org/guide/datasets)を使用して行データをシャッフルし、バッチ化するための入力パイプライン構築\n* feature columnsを使ったCSVの列のモデル訓練用の特徴量へのマッピング\n* Kerasを使ったモデルの構築と、訓練及び評価\n\n## データセット\n\nここでは、Cleveland Clinic Foundation for Heart Diseaseが提供している小さな[データセット](https://archive.ics.uci.edu/ml/datasets/heart+Disease)を使用します。このCSVファイルには数百行が含まれています。行が患者を、列がその属性を表します。この情報を使用して、患者が心臓疾患を持っているかを予測します。このデータセットの場合には二値分類タスクとなります。\n\n下記はこのデータセットの[說明](https://archive.ics.uci.edu/ml/machine-learning-databases/heart-disease/heart-disease.names)です。数値列とカテゴリー列があることに注目してください。\n\n>列| 說明| 特徴量の型 | データ型\n>------------|--------------------|----------------------|-----------------\n>Age | 年齢 | 数値型 | 整数\n>Sex | (1 = 男性; 0 = 女性) | カテゴリー型 | 整数\n>CP | 胸痛のタイプ (0, 1, 2, 3, 4) | カテゴリー型 | 整数\n>Trestbpd | 安静時血圧 (単位:mm Hg 入院時) | 数値型 | 整数\n>Chol | 血清コレステロール 単位:mg/dl | 数値型 | 整数\n>FBS | (空腹時血糖 > 120 mg/dl) (1 = 真; 0 = 偽) | カテゴリー型 | 整数\n>RestECG | 安静時心電図の診断結果 (0, 1, 2) | カテゴリー型 | 整数\n>Thalach | 最大心拍数 | 数値型 | 整数\n>Exang | 運動誘発狭心症 (1 = はい; 0 = いいえ) | カテゴリー型 | 整数\n>Oldpeak | 安静時と比較した運動時のST低下 | 数値型 | 整数\n>Slope | ピーク運動STセグメントの勾配 | 数値型 | 浮動小数点数\n>CA | 蛍光透視法によって着色された主要血管の数(0−3) | 数値型 | 整数\n>Thal | 3 = 正常; 6 = 固定欠陥; 7 = 可逆的欠陥 | カテゴリー型 | 文字列\n>Target | 心臓疾患の診断 (1 = 真; 0 = 偽) | 分類 | 整数",
"_____no_output_____"
],
[
"## TensorFlow他ライブラリのインポート",
"_____no_output_____"
]
],
[
[
"!pip install sklearn",
"_____no_output_____"
],
[
"from __future__ import absolute_import, division, print_function, unicode_literals\n\nimport numpy as np\nimport pandas as pd\n\ntry:\n # Colab only\n %tensorflow_version 2.x\nexcept Exception:\n pass\nimport tensorflow as tf\n\nfrom tensorflow import feature_column\nfrom tensorflow.keras import layers\nfrom sklearn.model_selection import train_test_split",
"_____no_output_____"
]
],
[
[
"## Pandasを使ったデータフレーム作成\n\n[Pandas](https://pandas.pydata.org/)は、構造化データの読み込みや操作のための便利なユーティリティを持つPythonのライブラリです。ここでは、Pandasを使ってURLからデータをダウンロードし、データフレームに読み込みます。",
"_____no_output_____"
]
],
[
[
"URL = 'https://storage.googleapis.com/applied-dl/heart.csv'\ndataframe = pd.read_csv(URL)\ndataframe.head()",
"_____no_output_____"
]
],
[
[
"## データフレームを、訓練用、検証用、テスト用に分割\n\nダウンロードしたデータセットは1つのCSVファイルです。これを、訓練用、検証用、テスト用のデータセットに分割します。",
"_____no_output_____"
]
],
[
[
"train, test = train_test_split(dataframe, test_size=0.2)\ntrain, val = train_test_split(train, test_size=0.2)\nprint(len(train), 'train examples')\nprint(len(val), 'validation examples')\nprint(len(test), 'test examples')",
"_____no_output_____"
]
],
[
[
"## tf.dataを使った入力パイプラインの構築\n\n次に、[tf.data](https://www.tensorflow.org/guide/datasets)を使ってデータフレームをラップします。こうすることで、feature columns をPandasデータフレームの列をモデル訓練用の特徴量へのマッピングするための橋渡し役として使うことができます。(メモリに収まらないぐらいの)非常に大きなCSVファイルを扱う場合には、tf.dataを使ってディスクから直接CSVファイルを読み込むことになります。この方法は、このチュートリアルでは扱いません。",
"_____no_output_____"
]
],
[
[
"# Pandasデータフレームからtf.dataデータセットを作るためのユーティリティメソッド\ndef df_to_dataset(dataframe, shuffle=True, batch_size=32):\n dataframe = dataframe.copy()\n labels = dataframe.pop('target')\n ds = tf.data.Dataset.from_tensor_slices((dict(dataframe), labels))\n if shuffle:\n ds = ds.shuffle(buffer_size=len(dataframe))\n ds = ds.batch(batch_size)\n return ds",
"_____no_output_____"
],
[
"batch_size = 5 # デモ用として小さなバッチサイズを使用\ntrain_ds = df_to_dataset(train, batch_size=batch_size)\nval_ds = df_to_dataset(val, shuffle=False, batch_size=batch_size)\ntest_ds = df_to_dataset(test, shuffle=False, batch_size=batch_size)",
"_____no_output_____"
]
],
[
[
"## 入力パイプラインを理解する\n\n入力パイプラインを構築したので、それが返すデータのフォーマットを見るために呼び出してみましょう。出力を読みやすくするためにバッチサイズを小さくしてあります。",
"_____no_output_____"
]
],
[
[
"for feature_batch, label_batch in train_ds.take(1):\n print('Every feature:', list(feature_batch.keys()))\n print('A batch of ages:', feature_batch['age'])\n print('A batch of targets:', label_batch )",
"_____no_output_____"
]
],
[
[
"データセットが(データフレームにある)列名からなるディクショナリを返すことがわかります。列名から、データフレームの行に含まれる列の値が得られます。",
"_____no_output_____"
],
[
"## feature columnsの様々な型の例\n\nTensorFlowにはたくさんの型のfeature columnがあります。このセクションでは、いくつかの型のfeature columnsを作り、データフレームの列をどのように変換しているかを示します。",
"_____no_output_____"
]
],
[
[
"# いくつかの型のfeature columnsを例示するためこのバッチを使用する\nexample_batch = next(iter(train_ds))[0]",
"_____no_output_____"
],
[
"# feature columnsを作りデータのバッチを変換する\n# ユーティリティメソッド\ndef demo(feature_column):\n feature_layer = layers.DenseFeatures(feature_column)\n print(feature_layer(example_batch).numpy())",
"_____no_output_____"
]
],
[
[
"### 数値コラム\n\nfeature columnsの出力はモデルへの入力になります(上記で定義したdemo関数を使うと、データフレームの列がどのように変換されるかをつぶさに見ることができます)。[数値コラム](https://www.tensorflow.org/api_docs/python/tf/feature_column/numeric_column)は、最も単純な型のコラムです。数値コラムは実数特徴量を表現するのに使われます。このコラムを使う場合、モデルにはデータフレームの列の値がそのまま渡されます。",
"_____no_output_____"
]
],
[
[
"age = feature_column.numeric_column(\"age\")\ndemo(age)",
"_____no_output_____"
]
],
[
[
"心臓疾患データセットでは、データフレームのほとんどの列が数値型です。",
"_____no_output_____"
],
[
"### バケット化コラム\n\n数値をそのままモデルに入力するのではなく、値の範囲に基づいた異なるカテゴリーに分割したいことがあります。例えば、人の年齢を表す生データを考えてみましょう。[バケット化コラム](https://www.tensorflow.org/api_docs/python/tf/feature_column/bucketized_column)を使うと年齢を数値コラムとして表現するのではなく、年齢をいくつかのバケットに分割できます。下記のワンホット値が、各行がどの年齢範囲にあるかを表していることに注目してください。",
"_____no_output_____"
]
],
[
[
"age_buckets = feature_column.bucketized_column(age, boundaries=[18, 25, 30, 35, 40, 45, 50, 55, 60, 65])\ndemo(age_buckets)",
"_____no_output_____"
]
],
[
[
"### カテゴリー型コラム\n\nこのデータセットでは、Thalは('fixed'、'normal'、'reversible'のような)文字列として表現されています。文字列を直接モデルに入力することはできません。まず、文字列を数値にマッピングする必要があります。categorical vocabulary コラムを使うと、(上記で示した年齢バケットのように)文字列をワンホットベクトルとして表現することができます。カテゴリーを表す語彙(vocabulary)は[categorical_column_with_vocabulary_list](https://www.tensorflow.org/api_docs/python/tf/feature_column/categorical_column_with_vocabulary_list)を使ってリストで渡すか、[categorical_column_with_vocabulary_file](https://www.tensorflow.org/api_docs/python/tf/feature_column/categorical_column_with_vocabulary_file)を使ってファイルから読み込むことができます。",
"_____no_output_____"
]
],
[
[
"thal = feature_column.categorical_column_with_vocabulary_list(\n 'thal', ['fixed', 'normal', 'reversible'])\n\nthal_one_hot = feature_column.indicator_column(thal)\ndemo(thal_one_hot)",
"_____no_output_____"
]
],
[
[
"より複雑なデータセットでは、たくさんの列がカテゴリー型(例えば文字列)であることでしょう。feature columns はカテゴリー型データを扱う際に最も役に立ちます。このデータセットでは、カテゴリー型コラムは1つだけですが、他のデータセットを扱う際に使用できるいくつかの重要な型のfeature columnsを紹介するために、この列を使用することにします。",
"_____no_output_____"
],
[
"### 埋め込み型コラム\n\n数種類の候補となる文字列ではなく、カテゴリー毎に数千(あるいはそれ以上)の値があるとしましょう。カテゴリーの数が多くなってくると、様々な理由から、ワンホットエンコーディングを使ってニューラルネットワークを訓練することが難しくなります。埋込み型コラムを使うと、こうした制約を克服することが可能です。[埋込み型コラム](https://www.tensorflow.org/api_docs/python/tf/feature_column/embedding_column)は、データを多次元のワンホットベクトルとして表すのではなく、セルの値が0か1かだけではなく、どんな数値でもとれるような密な低次元ベクトルとして表現します。埋め込みのサイズ(下記の例では8)は、チューニングが必要なパラメータです。\n\nキーポイント:カテゴリー型コラムがたくさんの選択肢を持つ場合、埋め込み型コラムを使用することが最善の方法です。ここでは例を一つ示しますので、今後様々なデータセットを扱う際には、この例を参考にしてください。",
"_____no_output_____"
]
],
[
[
"# この埋込み型コラムの入力は、先程作成したカテゴリ型コラムであることに注意\nthal_embedding = feature_column.embedding_column(thal, dimension=8)\ndemo(thal_embedding)",
"_____no_output_____"
]
],
[
[
"### ハッシュ化特徴コラム\n\n値の種類が多いカテゴリー型コラムを表現するもう一つの方法が、[categorical_column_with_hash_bucket](https://www.tensorflow.org/api_docs/python/tf/feature_column/categorical_column_with_hash_bucket)を使う方法です。このfeature columnは文字列をエンコードするために入力のハッシュ値を計算し、`hash_bucket_size`個のバケットの中から1つを選択します。このコラムを使用する場合には、語彙を用意する必要はありません。また、スペースの節約のために、実際のカテゴリー数に比べて極めて少ないバケット数を選択することも可能です。\n\nキーポイント:この手法の重要な欠点の一つは、異なる文字列が同じバケットにマッピングされるというハッシュ値の衝突が起きることです。実務上は、データセットによっては、この問題を無視できることがあります。",
"_____no_output_____"
]
],
[
[
"thal_hashed = feature_column.categorical_column_with_hash_bucket(\n 'thal', hash_bucket_size=1000)\ndemo(feature_column.indicator_column(thal_hashed))",
"_____no_output_____"
]
],
[
[
"### クロスフィーチャーコラム\n\n複数の特徴量をまとめて1つの特徴量にする、[フィーチャークロス](https://developers.google.com/machine-learning/glossary/#feature_cross)として知られている手法は、モデルが特徴量の組み合わせの一つ一つに別々の重みを学習することを可能にします。ここでは年齢とThalをクロスさせて新しい特徴量を作ってみます。交差列(`crossed_column`)が、起こりうるすべての組み合わせ全体のテーブル(これは非常に大きくなる可能性があります)を作るものではないことに注意してください。クロスフィーチャーコラムは、代わりにバックエンドとしてハッシュ化コラムを使用しているため、テーブルの大きさを選択することができます。",
"_____no_output_____"
]
],
[
[
"crossed_feature = feature_column.crossed_column([age_buckets, thal], hash_bucket_size=1000)\ndemo(feature_column.indicator_column(crossed_feature))",
"_____no_output_____"
]
],
[
[
"## 使用するコラムを選択する\n\nこれまで、いくつかのfeature columnの使い方を見てきました。いよいよモデルの訓練にそれらを使用することにします。このチュートリアルの目的は、feature columnsを使うのに必要な完全なコード(いわば力学)を示すことです。以下ではモデルを訓練するための列を適当に選びました。\n\nキーポイント:正確なモデルを構築するのが目的である場合には、できるだけ大きなデータセットを使用して、どの特徴量を含めるのがもっとも意味があるのかや、それらをどう表現したらよいかを、慎重に検討してください。",
"_____no_output_____"
]
],
[
[
"feature_columns = []\n\n# 数値コラム\nfor header in ['age', 'trestbps', 'chol', 'thalach', 'oldpeak', 'slope', 'ca']:\n feature_columns.append(feature_column.numeric_column(header))\n\n# バケット化コラム\nage_buckets = feature_column.bucketized_column(age, boundaries=[18, 25, 30, 35, 40, 45, 50, 55, 60, 65])\nfeature_columns.append(age_buckets)\n\n# インジケーター(カテゴリー型)コラム\nthal = feature_column.categorical_column_with_vocabulary_list(\n 'thal', ['fixed', 'normal', 'reversible'])\nthal_one_hot = feature_column.indicator_column(thal)\nfeature_columns.append(thal_one_hot)\n\n# 埋め込み型コラム\nthal_embedding = feature_column.embedding_column(thal, dimension=8)\nfeature_columns.append(thal_embedding)\n\n# クロスフィーチャーコラム\ncrossed_feature = feature_column.crossed_column([age_buckets, thal], hash_bucket_size=1000)\ncrossed_feature = feature_column.indicator_column(crossed_feature)\nfeature_columns.append(crossed_feature)",
"_____no_output_____"
]
],
[
[
"### 特徴量層の構築\n\nfeature columnsを定義し終わったので、次に[DenseFeatures](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/layers/DenseFeatures)層を使ってKerasモデルへの入力とします。",
"_____no_output_____"
]
],
[
[
"feature_layer = tf.keras.layers.DenseFeatures(feature_columns)",
"_____no_output_____"
]
],
[
[
"これまでは、feature columnsの働きを見るため、小さなバッチサイズを使ってきました。ここではもう少し大きなバッチサイズの新しい入力パイプラインを作ります。",
"_____no_output_____"
]
],
[
[
"batch_size = 32\ntrain_ds = df_to_dataset(train, batch_size=batch_size)\nval_ds = df_to_dataset(val, shuffle=False, batch_size=batch_size)\ntest_ds = df_to_dataset(test, shuffle=False, batch_size=batch_size)",
"_____no_output_____"
]
],
[
[
"## モデルの構築、コンパイルと訓練",
"_____no_output_____"
]
],
[
[
"model = tf.keras.Sequential([\n feature_layer,\n layers.Dense(128, activation='relu'),\n layers.Dense(128, activation='relu'),\n layers.Dense(1, activation='sigmoid')\n])\n\nmodel.compile(optimizer='adam',\n loss='binary_crossentropy',\n metrics=['accuracy'])\n\nmodel.fit(train_ds, \n validation_data=val_ds, \n epochs=5)",
"_____no_output_____"
],
[
"loss, accuracy = model.evaluate(test_ds)\nprint(\"Accuracy\", accuracy)",
"_____no_output_____"
]
],
[
[
"キーポイント:一般的に、ディープラーニングが最良の結果となるのは、もっと大きくて、もっと複雑なデータセットです。この例のように小さなデータセットを使用する際には、強固なベースラインとして、決定木やランダムフォレストを使うことをおすすめします。このチュートリアルの目的は、訓練により正確なモデルを得ることではなく、構造化データの使い方をデモすることです。今後ご自分のデータセットに取り組まれる際の出発点として、これらのコードをお使いください。",
"_____no_output_____"
],
[
"## 次のステップ\n\n構造化データの分類について更に多くのことを学ぶためには、自分自身で試してみることです。別のデータセットを見つけ、上記と同様のコードを使って、それを分類するモデルを訓練してみてください。正解率を上げるためには、モデルにどの特徴量を含めたらよいかや、その特徴量をどのように表現すべきかをじっくり考えてください。",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
]
] |
4ab5a38cddd2ef8758d3948928b860fb711ebba2
| 5,786 |
ipynb
|
Jupyter Notebook
|
ml/04-sklearn/LinearRegression.ipynb
|
sdyz5210/python
|
78f9999f94d92d9ca7fde6f18acec7d3abd422ef
|
[
"BSD-3-Clause"
] | null | null | null |
ml/04-sklearn/LinearRegression.ipynb
|
sdyz5210/python
|
78f9999f94d92d9ca7fde6f18acec7d3abd422ef
|
[
"BSD-3-Clause"
] | null | null | null |
ml/04-sklearn/LinearRegression.ipynb
|
sdyz5210/python
|
78f9999f94d92d9ca7fde6f18acec7d3abd422ef
|
[
"BSD-3-Clause"
] | null | null | null | 24.832618 | 89 | 0.400622 |
[
[
[
"# 多元线性回归",
"_____no_output_____"
]
],
[
[
"from sklearn.linear_model import LinearRegression as LR\nfrom sklearn.model_selection import cross_val_score\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.datasets import fetch_california_housing as fch\nimport pandas as pd",
"_____no_output_____"
],
[
"housevalue = fch()",
"_____no_output_____"
],
[
"X = pd.DataFrame(housevalue.data)\ny = housevalue.target\nX.head()",
"_____no_output_____"
],
[
"Xtrain, Xtest, Ytrain, Ytest = train_test_split(X,y,test_size=0.3,random_state=420)",
"_____no_output_____"
],
[
"for i in [Xtrain, Xtest]: \n i.index = range(i.shape[0])\nXtrain.shape",
"_____no_output_____"
],
[
"#建模\nreg = LR().fit(Xtrain, Ytrain)\nyhat = reg.predict(Xtest)\nyhat",
"_____no_output_____"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab5a72587cf6a787ed74428822c40e8ec69a37d
| 558,263 |
ipynb
|
Jupyter Notebook
|
TP1-DataCleaning-Properati/Desafio 1 - Grupo 4.ipynb
|
stephanieizquierdo/DigitalHouse-DataScience
|
3b2d27c0d78b2d2289884e46e5774aa8f5f38752
|
[
"MIT"
] | null | null | null |
TP1-DataCleaning-Properati/Desafio 1 - Grupo 4.ipynb
|
stephanieizquierdo/DigitalHouse-DataScience
|
3b2d27c0d78b2d2289884e46e5774aa8f5f38752
|
[
"MIT"
] | null | null | null |
TP1-DataCleaning-Properati/Desafio 1 - Grupo 4.ipynb
|
stephanieizquierdo/DigitalHouse-DataScience
|
3b2d27c0d78b2d2289884e46e5774aa8f5f38752
|
[
"MIT"
] | null | null | null | 55.8263 | 52,744 | 0.62784 |
[
[
[
"## Importación de librerías",
"_____no_output_____"
]
],
[
[
"import numpy as np\nimport pandas as pd\nimport re\nimport matplotlib.pyplot as plt\nimport seaborn as sns",
"_____no_output_____"
]
],
[
[
"## Cargo el dataset",
"_____no_output_____"
]
],
[
[
"data = pd.read_csv('../Data/properatti.csv',index_col=0)\n#data.head()",
"_____no_output_____"
]
],
[
[
"## Valores Faltantes del dataset",
"_____no_output_____"
]
],
[
[
"data.isnull().sum()",
"_____no_output_____"
],
[
"data.loc[data['description'].isna(),['price','currency','price_aprox_usd','price_usd_per_m2','rooms','floor','expenses']]",
"_____no_output_____"
],
[
"nodesc_index = data.loc[data['description'].isna()].index\ndata.drop(index=nodesc_index,inplace=True)\ndata.isna().sum()",
"_____no_output_____"
]
],
[
[
"## Tipos de datos",
"_____no_output_____"
]
],
[
[
"data.dtypes",
"_____no_output_____"
]
],
[
[
"# Columna Currency\n\n¿La moneda de publicación es pesos argentinos y dólares? ¿O hay otra?",
"_____no_output_____"
]
],
[
[
"data['currency'].value_counts()",
"_____no_output_____"
]
],
[
[
"Las publicaciones en Sol Peruano y Peso Uruguayo se deberán pasar a dólares.",
"_____no_output_____"
],
[
"### Tipo de Cambio (conversión de pesos a dolares y viceversa)",
"_____no_output_____"
]
],
[
[
"data['currency']==\"USD\"\ndata_USD= data.loc[data['currency']==\"USD\", [\"price\", \"price_aprox_local_currency\",\"price_aprox_usd\"]]\ndata_USD.head(5)",
"_____no_output_____"
],
[
"y=data_USD[\"price_aprox_local_currency\"]/data_USD['price_aprox_usd']\ny=y.round(decimals=4)\ny.unique()",
"_____no_output_____"
]
],
[
[
"1 dolar = 17.6445 pesos argentinos. (A este precio, yo compro!)",
"_____no_output_____"
]
],
[
[
"#cambio_dolar_a_peso = 17.6445\ndef dolar_a_peso(price_usd):\n #return round(price_usd*17.6445,2)\n return price_usd*17.6445\ndef peso_a_dolar(price_ars):\n #return round(price_ars/17.6445,2)\n return price_ars/17.6445",
"_____no_output_____"
]
],
[
[
"Las filas donde falta el precio en pesos, tambien falta el precio en dólares:",
"_____no_output_____"
]
],
[
[
"data.loc[(data['price_aprox_local_currency'].isna())&(data['price_aprox_usd'].isna())].shape",
"_____no_output_____"
],
[
"# data.loc[(data['price'].notnull())&(data['currency'].notnull())&(data['currency']=='ARS')&(data['price']!=data['price_aprox_local_currency']),['price','currency','price_aprox_local_currency','price_aprox_usd']].head(10)",
"_____no_output_____"
]
],
[
[
"# Precio por metro cuadrado",
"_____no_output_____"
],
[
"En cuanto al metro cuadrado, solo en 28295 filas falta el valor de ambos. Podemos calcular el valor del precio por metro cuadrado de una columna a partir de la otra.",
"_____no_output_____"
]
],
[
[
"data.loc[(data['price_usd_per_m2'].isna())&(data['price_per_m2'].isna())].shape",
"_____no_output_____"
],
[
"data.loc[(data['currency']=='USD'),['currency','price_aprox_local_currency','price','surface_covered_in_m2','price_per_m2',\\\n 'price_aprox_usd','surface_total_in_m2','price_usd_per_m2']].head(10)",
"_____no_output_____"
],
[
"data.loc[(data['currency']=='ARS'),['currency','price_aprox_local_currency','price','surface_covered_in_m2','price_per_m2',\\\n 'price_aprox_usd','surface_total_in_m2','price_usd_per_m2']].head(10)",
"_____no_output_____"
]
],
[
[
"$price\\_usd\\_per\\_m2 = \\frac{price\\_aprox\\_usd}{surface\\_total\\_in\\_m2}$\n\n$price\\_per\\_m2 = \\frac{price}{surface\\_covered\\_in\\_m2}$\n\nEl problema se da que a veces la superficie total es menor que la superficie cubierta. Por lo tanto, se decidió dropear estas 1106 filas.",
"_____no_output_____"
]
],
[
[
"data.loc[data['surface_covered_in_m2']>data['surface_total_in_m2']]#.shape",
"_____no_output_____"
],
[
"sup_invalida = data.loc[data['surface_covered_in_m2']>data['surface_total_in_m2']].index\ndata.drop(index=sup_invalida,inplace=True)",
"_____no_output_____"
],
[
"data.loc[(data['price_usd_per_m2'].isna())&(data['price_aprox_usd'].notnull())&(data['surface_total_in_m2'].notnull())\\\n &(data['surface_total_in_m2']!=0),\n ['price_usd_per_m2','price_aprox_usd','surface_total_in_m2']]",
"_____no_output_____"
],
[
"data.loc[(data['price_per_m2'].isna())&(data['price'].notnull())&(data['surface_covered_in_m2'].notnull())\\\n ,['price_per_m2','price','surface_covered_in_m2']]",
"_____no_output_____"
],
[
"data.loc[data['surface_total_in_m2']==0,'surface_total_in_m2']=np.nan\ndata.loc[data['surface_covered_in_m2']==0,'surface_covered_in_m2']=np.nan\n",
"_____no_output_____"
]
],
[
[
"# Valores faltantes de Currency",
"_____no_output_____"
]
],
[
[
"data.loc[(data['currency'].isna())&(data['price_aprox_local_currency'].isna())&(data['price_aprox_usd'].isna()),['price','price_aprox_local_currency','price_aprox_usd']]",
"_____no_output_____"
],
[
"data.loc[(data['currency'].isna())&(data['price_aprox_local_currency'].notnull())&(data['price_aprox_usd'].notnull()),['price','price_aprox_local_currency','price_aprox_usd']]",
"_____no_output_____"
]
],
[
[
"De lo visto, no puedo calcular valores faltantes de currency a partir de price, price_aprox_usd y price_aprox_local_currency",
"_____no_output_____"
],
[
"## Cálculo de superficie, precio y precio por m2",
"_____no_output_____"
],
[
"La idea es ver en que filas tengo 2 valores y obtener el tercero.\nPrecio por m2 = Precio / Superficie.\n\nPara esto, es necesario verificar primero que los valores sean válidos, es decir, valores no nulos.\n\n**Verificación valores de superficie**",
"_____no_output_____"
]
],
[
[
"min_sup_valida = 10\ndata.loc[data['surface_total_in_m2']<min_sup_valida,'surface_total_in_m2']=np.nan",
"_____no_output_____"
]
],
[
[
"**Verificación valores de precio por m2**\n\nDespués de un análisis de la información, no se encuentra un patrón para establecer un mínimo o un máximo para verificar la validez de un dato. Mientras no hayan valores nulos (cero), se dejará todo como esta",
"_____no_output_____"
]
],
[
[
"#min_pricem2_valido = \ndata.loc[data['price_usd_per_m2']<=5,'price_usd_per_m2'].value_counts()",
"_____no_output_____"
]
],
[
[
"**Verificación valores de precio en dolares**\n\nSólo se encontró un valor nulo (cero).",
"_____no_output_____"
]
],
[
[
"print(data.loc[data['price_aprox_usd']<=4000,['price_aprox_usd','description']].values)\ndata.loc[data['price_aprox_usd']<=4000,'price_aprox_usd']=np.nan",
"[[0.0 'Casa en Venta de 2 dorm. en Armenia']]\n"
]
],
[
[
"Si hay valor de precio y hay valor superficie total, se puede calcular el valor del precio por metro cuadrado.\nPero este caso no ocurre.",
"_____no_output_____"
]
],
[
[
"data.loc[(data['price'].notnull())&(data['surface_total_in_m2'].notnull())&(data['price_usd_per_m2'].isna())].shape",
"_____no_output_____"
]
],
[
[
"Si hay valor de precio y hay valor de precio por metro cuadrado, se puede calcular el valor de la superficie. Pero este caso no ocurre.",
"_____no_output_____"
]
],
[
[
"data.loc[(data['price'].notnull())&(data['surface_total_in_m2'].isna())&(data['price_usd_per_m2'].notnull())].shape",
"_____no_output_____"
]
],
[
[
"Si hay valor de superficie total y hay valor de precio por metro cuadrado, se puede calcular el valor del precio.\nPero este caso no ocurre.",
"_____no_output_____"
]
],
[
[
"data.loc[(data['price'].isna())&(data['surface_total_in_m2'].notnull())&(data['price_usd_per_m2'].notnull())].shape",
"_____no_output_____"
]
],
[
[
"# Valores faltantes de 'place_name'\n\nSe aprecia que todos los valores faltantes de 'place_name' corresponden a Tigre. Podríamos generar una función para tomar este valore de la columna 'place_with_parent_name', pero lo vamos a hardcodear.",
"_____no_output_____"
]
],
[
[
"data.loc[data['place_name'].isna(),['place_name','place_with_parent_names','country_name','state_name']]",
"_____no_output_____"
],
[
"def get_city(place):\n a=place.split(sep='|')\n new_a=[item for item in a if len(item)>0]\n return(new_a[-1])\n\ndef get_place_names(text):\n place_names = text.split(sep='|')\n place_names = [item for item in place_names if len(item)>0]\n place_names = place_names[0:-1]\n complete_names = '|'.join(place_names)\n return '|'+complete_names+'|'",
"_____no_output_____"
],
[
"data['place_name'].fillna('Tigre',inplace=True)\ndata.groupby('place_name')['place_name'].unique()",
"_____no_output_____"
]
],
[
[
"Vemos un lugar que se llama 'coordenadas 34.255511'. Vamos a verificar qué lugar es en realidad.\n",
"_____no_output_____"
]
],
[
[
"data.loc[data['place_name']=='coordenadas 34.255511','place_with_parent_names'].values",
"_____no_output_____"
]
],
[
[
"Es Tigre. También hay que arreglarlo.",
"_____no_output_____"
]
],
[
[
"data.loc[data['place_name']=='coordenadas 34.255511','place_with_parent_names']=get_place_names(data.loc[data['place_name']=='coordenadas 34.255511','place_with_parent_names'].values[0])\ndata.loc[data['place_name']=='coordenadas 34.255511','place_name']='Tigre'",
"_____no_output_____"
],
[
"data.loc[data['place_name']=='coordenadas 34.255511']",
"_____no_output_____"
]
],
[
[
"# Valores Faltantes de 'floor'",
"_____no_output_____"
],
[
"### Funciones para Limpieza y Búsqueda de Datos",
"_____no_output_____"
]
],
[
[
"#!pip install unidecode\nimport unidecode\n\ndef quitar_caracteres(entrada):\n return str.lower(unidecode.unidecode(entrada))",
"_____no_output_____"
],
[
"# def pb_search(text):\n# pattern_fl = r'(?:planta baja|pb|p.b.)'\n# regex_fl = re.compile(pattern_fl, flags = re.IGNORECASE)\n# floors_lst = regex_fl.findall(text)",
"_____no_output_____"
],
[
"#Revisar y ver que pasa con varias ocurrencias\ndef text_floor_search(text):\n floor_map = {\n 'primer': 1.0,'segundo': 2.0,'tercer': 3.0,'cuarto': 4.0,'quinto': 5.0,\n 'sexto': 6.0,'septimo': 7.0,'octavo': 8.0,'noveno': 9.0,'decimo': 10.0\n }\n \n pattern_fl = r'(\\w{2,7})\\s?(?:piso)'\n regex_fl = re.compile(pattern_fl, flags = re.IGNORECASE)\n floors_lst = regex_fl.findall(text)\n \n if not floors_lst:\n return np.nan\n if len(floors_lst)!=1:\n return np.nan\n if quitar_caracteres(floors_lst[0]) not in floor_map.keys():\n return np.nan\n floor = floor_map[quitar_caracteres(floors_lst[0])]\n return floor",
"_____no_output_____"
],
[
"#Revisar y ver que pasa con varias ocurrencias\ndef text_search_floor(text):\n floor_map = {\n 'un': 1.0,'uno': 1.0,'dos': 2.0,'tres': 3.0,'cuatro': 4.0,'cinco': 5.0,\n 'seis': 6.0,'siete': 7.0,'ocho': 8.0,'nueve': 9.0,'diez': 10.0,'once': 11.0,\n 'doce': 12.0,'trece': 13.0,'catorce': 14.0,'quince': 15.0\n }\n \n pattern_fl = r'(?:piso)\\s?(\\w{2,7})'\n regex_fl = re.compile(pattern_fl, flags = re.IGNORECASE)\n floors_lst = regex_fl.findall(text)\n \n if not floors_lst:\n return np.nan\n if len(floors_lst)==0:\n return np.nan\n if len(floors_lst)==1:\n num = floors_lst[0]\n if quitar_caracteres(num) not in floor_map.keys():\n return np.nan\n floor = floor_map[quitar_caracteres(num)]\n return floor\n else:\n floors_l = [item for item in floors_lst]\n floors_val = [int(item in floor_map) for item in floors_l]\n for floor,val in zip(floors_l,floors_val):\n if val==1:\n return floor_map[floor]\n return np.nan",
"_____no_output_____"
],
[
"#Revisar y ver que pasa con varias ocurrencias\ndef num_floor_search(text):\n pattern_fl = r'(\\d{1,2})\\.?(°|\\w{2})\\s?(?:piso)'\n regex_fl = re.compile(pattern_fl, flags = re.IGNORECASE)\n floors_lst = regex_fl.findall(text)\n \n abb_lst = ['er','do','to','mo','vo','no','deg','ero','']\n\n if not floors_lst:\n return np.nan\n if len(floors_lst)!=1:\n return np.nan\n sufijo=quitar_caracteres(floors_lst[0][1])\n if sufijo not in abb_lst:\n return np.nan\n num=float(floors_lst[0][0])\n# if num<15 and num>9:\n# print(\"Piso: {}\".format(num))\n# print(text)\n if num>15:\n return np.nan\n else:\n return num\n \n #return(floors_lst)",
"_____no_output_____"
],
[
"#Revisar y ver que pasa con varias ocurrencias\ndef num_search_floor(text):\n pattern_fl = r'\\s?(?:piso)(\\d{1,2})'\n regex_fl = re.compile(pattern_fl, flags = re.IGNORECASE)\n floors_lst = regex_fl.findall(text)\n \n if not floors_lst:\n return np.nan\n if len(floors_lst)!=1:\n return np.nan\n num = float(floors_lst[0])\n if num>40:\n return np.nan\n else:\n return num\n ",
"_____no_output_____"
],
[
"def clean_temp_col(df,tempCol):\n \"\"\"Cleans a tempCol column from a df dataframe. Writes nan on every row.\n \n Parameters\n ----------\n df : pandas.DataFrame\n The dataframe you want to clean a temporary column.\n tempCol: str\n The name passed as a string of the column to clean.\n \"\"\"\n df.loc[:,tempCol]=np.nan",
"_____no_output_____"
],
[
"def new_found(df,tempCol,cleanCol):\n \"\"\"Shows a message about the new data found.\n \n Parameters\n ----------\n df : pandas.DataFrame\n The dataframe to take data from.\n tempCol: str\n The name passed as a string of the temporary column.\n cleanCol: str\n the name passed as a string of the clean column.\n \"\"\"\n print(\"Nuevos: {}.\".format(df[tempCol].notnull().sum()))\n print(\"Total: {}.\".format(df[cleanCol].notnull().sum()))",
"_____no_output_____"
],
[
"import scipy.stats as stats\n\n#valor_mas_chico=0\n\ndef calculate_iqr_values(array):\n \"\"\"Calculates inferior and superior limits of the iqr plus a margin.\n Returns\n ---------\n lim_inf: float\n q1-1.5*iqr\n lim_sup: float\n q3+1.5*iqr\n \"\"\"\n q1,q3=np.percentile(array,[25 ,75])\n arr_iqr = stats.iqr(array)\n lim_inf = q1-1.5*arr_iqr\n# if lim_inf<valor_mas_chico:\n# lim_inf=valor_mas_chico\n lim_sup = q3+1.5*arr_iqr\n return lim_inf,lim_sup\n\ndef check_outliers(x,min_lim,max_lim):\n if x<min_lim:\n return np.nan\n elif x>max_lim:\n return np.nan\n else:\n return x",
"_____no_output_____"
]
],
[
[
"## Tipos de propiedad\n\nEl número de piso tiene mayor sentido e importancia para las propiedades de tipo 'apartment' que para house, PH y store, las cuales suelen ser en planta baja o primer piso.",
"_____no_output_____"
]
],
[
[
"data['property_type'].value_counts()",
"_____no_output_____"
]
],
[
[
"## Análisis de 'floor' para Casas",
"_____no_output_____"
],
[
"El valor de piso para una casa suele ser 0 (PB). Hay casas que se encuentran en un primer piso al subir por una escalera. Por lo cual, se decidió poner en cero el piso de todas las casas que digan estar en un segundo piso o superior.",
"_____no_output_____"
],
[
"**¿En qué piso dice que están las casas?**\n\nSi nos fijamos hay valores que carecen de sentido y se le atribuye a un error en el cargado de los datos.",
"_____no_output_____"
]
],
[
[
"#data.loc[data['property_type']=='house']['floor'].value_counts()\nnp.set_printoptions(suppress=True) \nnp.sort(data.loc[data['property_type']=='house']['floor'].unique())\ndata.loc[data['property_type']=='house']['floor'].unique()",
"_____no_output_____"
]
],
[
[
"Con un chequeo 'automático' de outliers, reduzco los valores a los siguientes:",
"_____no_output_____"
]
],
[
[
"iqr_values=calculate_iqr_values(data.loc[(data['floor'].notnull())&(data['property_type']=='PH'),'floor'])\ndata.loc[(data['property_type']=='house')&(data['floor'].notnull()),'floor'].apply(check_outliers,args=iqr_values).unique()",
"_____no_output_____"
]
],
[
[
"Si bien algunos valores no parece tener sentido, como una casa en el piso 6 o 7, luego de revisar las descripciones se vio que algunas están mal catalogadas como casas (y son departamentos), o habla de duplex en en el piso 7. Por lo tanto, se decide dejar estos como valores válidos, y el resto asignarles NaN.",
"_____no_output_____"
],
[
"## Corrección de los datos de Casas",
"_____no_output_____"
]
],
[
[
"#data.loc[(data['property_type']=='house') & (data['floor']==1)]['description'].values\niqr_values_house=calculate_iqr_values(data.loc[(data['floor'].notnull())&(data['property_type']=='PH'),'floor'])\n\ndata.loc[(data['property_type']=='house')&(data['floor']>iqr_values[1]),'floor']=np.nan\n# print(data.loc[data['property_type']=='house','floor'].value_counts(dropna=False))\n\nvariantes = ['1er piso por escalera','1° piso por escalera','1 piso por escalera',\n 'primer piso por escalera','planta alta por escalera']\n\nfor text in variantes:\n house_PA=data.loc[(data['property_type']=='house')&(data['floor'].isna())]['description']\\\n .str.contains(text,case=False,regex=False).astype(float).apply(lambda x: x if x else np.nan)\n #data.loc[(data['property_type']=='house')&(data['floor'].isna()),'floor'].fillna(house_PA,inplace=True)\n data.loc[(data['property_type']=='house')&(data['floor'].isna()),'floor']=data.loc[(data['property_type']=='house')&(data['floor'].isna()),'floor'].add(house_PA,fill_value=0)\n #print(data.loc[data['property_type']=='house','floor'].value_counts(dropna=False))\n",
"_____no_output_____"
]
],
[
[
"¿Y ahora?",
"_____no_output_____"
]
],
[
[
"#data.loc[data['property_type']=='house']['floor'].value_counts()\nnp.set_printoptions(suppress=True) \nnp.sort(data.loc[data['property_type']=='house']['floor'].unique())\ndata.loc[data['property_type']=='house']['floor'].value_counts(dropna=False, sort=True)",
"_____no_output_____"
]
],
[
[
"Los valores faltantes de casas (39456 datos) los completo con 0 (las casas suelen estar en PB).",
"_____no_output_____"
]
],
[
[
"data.loc[(data['property_type']=='house')&(data['floor'].isnull()),'floor']=0\ndata.loc[data['property_type']=='house']['floor'].value_counts(dropna=False, sort=True)",
"_____no_output_____"
]
],
[
[
"## Análisis de 'floor' para PH\n\nEl valor de piso de un PH suele ser PB o 1° piso. Pero hay casos donde se ve que está en un segundo o un tercer piso. Para mayor número de piso, encontramos muchos valores incorrectos. Por eso, un número de piso 4 o mayor, será tomado como incorrecto y se pondrá a cero, para realizar una corrección similar a la hecha con las casas",
"_____no_output_____"
]
],
[
[
"np.sort(data.loc[data['property_type']=='PH']['floor'].unique())\n#data.loc[data['property_type']=='PH']['floor'].value_counts(dropna=False, sort=True)",
"_____no_output_____"
],
[
"iqr_values_ph=calculate_iqr_values(data.loc[(data['floor'].notnull())&(data['property_type']=='PH'),'floor'])\ndata.loc[(data['property_type']=='PH')&(data['floor'].notnull()),'floor'].apply(check_outliers,args=iqr_values).unique()",
"_____no_output_____"
]
],
[
[
"Por outliers elimino una buena cantidad de valores extraños para el número de piso de un PH.",
"_____no_output_____"
]
],
[
[
"#data.loc[(data['property_type']=='PH')&(data['floor']<=iqr_values[1]),['floor','description']].values",
"_____no_output_____"
]
],
[
[
"## Corrección de los datos de PH",
"_____no_output_____"
]
],
[
[
"data.loc[(data['property_type']=='PH')]['floor'].value_counts(dropna=False,sort=True)",
"_____no_output_____"
]
],
[
[
"Los datos de piso>7 los tomamos como inválidos y los ponemos en NaN.",
"_____no_output_____"
]
],
[
[
"data.loc[(data['property_type']=='PH')&(data['floor']>iqr_values[1]),'floor']=np.nan\ndata.loc[(data['property_type']=='PH')]['floor'].value_counts(dropna=False,sort=True)",
"_____no_output_____"
]
],
[
[
"Al resto de valores (los 5246 NaN) los paso por una búsqueda de piso (entre 0 y 3).",
"_____no_output_____"
]
],
[
[
"patterns_pb = ['planta baja','PB','P.B.']\nacc_1 = ['primer piso','1er piso','1° piso']\nacc_2 = ['segundo piso','2do piso','2° piso']\nacc_3 = ['tercer piso','3er piso','3° piso']\n\naccepted = [patterns_pb,acc_1,acc_2,acc_3]\npiso = 0.0\nfor acc in accepted:\n for text in acc:\n ph_piso = data.loc[(data['property_type']=='PH')&(data['floor'].isna()),'description']\\\n .str.contains(text,case=False,regex=False)\n data.loc[(data['property_type']=='PH')&(ph_piso),'floor']=piso\n piso+=1\ndata.loc[(data['property_type']=='PH')]['floor'].value_counts(dropna=False,sort=True)",
"_____no_output_____"
]
],
[
[
"Al resto de NaN los pongo en cero.",
"_____no_output_____"
]
],
[
[
"data.loc[(data['property_type']=='PH')&(data['floor'].isna()),'floor']=0\n#data.loc[(data['property_type']=='PH')&(data['floor'].isna()),'floor']\ndata.loc[(data['property_type']=='PH')]['floor'].value_counts(dropna=False,sort=True)",
"_____no_output_____"
]
],
[
[
"## Análisis de 'floor' para store",
"_____no_output_____"
]
],
[
[
"#np.sort(data.loc[data['property_type']=='store']['floor'].unique())\ndata.loc[data['property_type']=='store']['floor'].value_counts(dropna=False, sort=True)",
"_____no_output_____"
]
],
[
[
"Luego de una inspección de los datos y algunas descripciones, se decide que hay valores válidos hasta piso 10 (oficina). Luego, tomamos como valores inválidos (mal cargados), y seguirán el mismo camino que el análisis de la descripción que las categorías de 'house' y 'PH'.",
"_____no_output_____"
]
],
[
[
"iqr_values_store=calculate_iqr_values(data.loc[(data['floor'].notnull())&(data['property_type']=='store'),'floor'])\ndata.loc[(data['property_type']=='store')&(data['floor'].notnull()),'floor'].apply(check_outliers,args=iqr_values).unique()",
"_____no_output_____"
]
],
[
[
"## Corrección de los datos de store",
"_____no_output_____"
],
[
"Luego de revisar algunas descripciones, se encontró que los valores de piso válidos encontrados son hasta el piso 10. Hay oficinas clasificadas como store. Haciendo un filtro por outliers, los datos válidos son hasta el piso 2. Tomamos como despreciable los datos perdidos.",
"_____no_output_____"
]
],
[
[
"data.loc[(data['property_type']=='store')]['floor'].value_counts(dropna=False,sort=True)",
"_____no_output_____"
],
[
"data.loc[(data['property_type']=='store') & (data['floor']>iqr_values_store[1]),'floor']=0\ndata.loc[(data['property_type']=='store')]['floor'].value_counts(dropna=False,sort=True)",
"_____no_output_____"
],
[
"data['clean_fl']=data['floor']\ndata['temp_fl']=np.nan\n\nfunc_lst = [text_floor_search,text_search_floor,num_floor_search,num_search_floor]\nfor func in func_lst:\n# print(func.__name__)\n# print(data.loc[(data['property_type']=='store')&(data['floor'].isna())]['description'].apply(func).value_counts()\n# )\n clean_temp_col(data,'temp_fl')\n data.loc[data['property_type']=='store','temp_fl']=data.loc[(data['property_type']=='store')&(data['floor'].isna())]['description'].apply(func)\n data.loc[data['property_type']=='store','temp_fl'] = data.loc[data['property_type']=='store','temp_fl'].apply(check_outliers,args=iqr_values_store)\n data.loc[data['property_type']=='store','clean_fl']=data.loc[data['property_type']=='store','clean_fl'].add(data.loc[data['property_type']=='store','temp_fl'],fill_value=0)\n\ndata.loc[(data['property_type']=='store'),'floor'] = data[data['property_type']=='store']['floor'].add(data[data['property_type']=='store']['clean_fl'],fill_value=0)\n#data.loc[(data['property_type']=='store')&(data['clean_fl']).notnull()] \ndata.loc[(data['property_type']=='store')]['floor'].value_counts(dropna=False,sort=True)",
"_____no_output_____"
]
],
[
[
"El resto de los NaN los pongo a cero.",
"_____no_output_____"
]
],
[
[
"data.loc[(data['property_type']=='store')&(data['floor'].isna()),'floor']=0\ndata.loc[(data['property_type']=='store')]['floor'].value_counts(dropna=False,sort=True)",
"_____no_output_____"
],
[
"data['floor'].isna().sum()",
"_____no_output_____"
]
],
[
[
"## Análisis de 'floor' para apartment\n\nDespués de una análisis exploratorio y lectura de algunas descripciones se encontró que el piso válido más alto es el piso 40 (departamento en puerto madero). Los valores mayores se pondrán a NaN y se buscará el valor correcto en title y description.",
"_____no_output_____"
]
],
[
[
"np.sort(data.loc[data['property_type']=='apartment']['floor'].unique())\n#data.loc[(data['property_type']=='apartment')&(data['floor']<=40)&(data['floor']>=10),['floor','description']].sample(10).values",
"_____no_output_____"
]
],
[
[
"Filtrando por outliers, me quedan estos valores:",
"_____no_output_____"
]
],
[
[
"iqr_values_apt=calculate_iqr_values(data.loc[(data['floor'].notnull())&(data['property_type']=='apartment'),'floor'])\ndata.loc[(data['property_type']=='apartment')&(data['floor'].notnull()),'floor'].apply(check_outliers,args=iqr_values).unique()",
"_____no_output_____"
]
],
[
[
"Pero se ha hecho un análisis donde se encontró departamentos hasta en piso 40. Por lo tanto, se tomará como valores válidos para estos datos hasta piso 40. Lo mismo se tomará para los demás valores buscados en la descripción de la propiedad.",
"_____no_output_____"
]
],
[
[
"data.loc[(data['property_type']=='apartment')&(data['floor']>40),'floor']=np.nan\ndata.loc[(data['floor']<=15)|(data['floor'].isna()),'floor'].value_counts(dropna=False)",
"_____no_output_____"
]
],
[
[
"## Columnas temporales",
"_____no_output_____"
]
],
[
[
"data['clean_fl']=data['floor']\ndata['temp_fl']=np.nan",
"_____no_output_____"
]
],
[
[
"## Búsqueda de piso para apartment \n\nItero con las columnas 'title' y 'description'",
"_____no_output_____"
]
],
[
[
"#print(apartment_df.columns)\n\ndata['description'] = data['description'].fillna(\"-\")\nfl_func_lst = [text_floor_search,num_floor_search,text_search_floor,num_search_floor]\ncol_lst = ['title','description']\nfor col in col_lst:\n print(\"------------------------\")\n print(\"Columna: {}\".format(col))\n print(\"------------------------\")\n for func in fl_func_lst:\n clean_temp_col(data,'temp_fl')\n data.loc[data['clean_fl'].isna(),'temp_fl'] = data[data['clean_fl'].isna()][col].apply(func)\n data.loc[data['property_type']=='apartment','temp_fl'] = data.loc[data['property_type']=='apartment','temp_fl'].apply(check_outliers,args=(0,40))\n data.loc[data['clean_fl'].isna(),'clean_fl'] = data['clean_fl'].add(data['temp_fl'],fill_value=0)\n print(func.__name__)\n new_found(data,'temp_fl','clean_fl')\n",
"------------------------\nColumna: title\n------------------------\ntext_floor_search\nNuevos: 85.\nTotal: 55830.\nnum_floor_search\nNuevos: 180.\nTotal: 56010.\ntext_search_floor\nNuevos: 13.\nTotal: 56023.\nnum_search_floor\nNuevos: 3.\nTotal: 56026.\n------------------------\nColumna: description\n------------------------\ntext_floor_search\nNuevos: 2281.\nTotal: 58307.\nnum_floor_search\nNuevos: 2223.\nTotal: 60530.\ntext_search_floor\nNuevos: 157.\nTotal: 60687.\nnum_search_floor\nNuevos: 33.\nTotal: 60720.\n"
],
[
"data.loc[data['floor'].isna(),'floor']=data.loc[data['floor'].isna(),'clean_fl']\ndata['floor'].isna().sum()",
"_____no_output_____"
]
],
[
[
"### Imputar valores faltantes por mediana por place. ",
"_____no_output_____"
]
],
[
[
"apartment_df = data.groupby('property_type').get_group('apartment')\nfloor_place_dict = apartment_df[['place_name','floor']].groupby('place_name').aggregate(np.median).to_dict()\nfloor_place_dict = floor_place_dict['floor']\nfloor_place_dict\n\nfor key in floor_place_dict.keys():\n data.loc[(data['place_name']==key)&(data['floor'].isna()),'floor']=floor_place_dict[key]\n#data.loc[(data['floor']<=15)|(data['floor'].isna()),'floor'].value_counts(dropna=False)",
"_____no_output_____"
]
],
[
[
"## Elimino columnas temporales",
"_____no_output_____"
]
],
[
[
"data.drop(['clean_fl','temp_fl'],axis=1,inplace=True)\ndata.isna().sum()",
"_____no_output_____"
]
],
[
[
"# Valores Faltantes de 'rooms'",
"_____no_output_____"
],
[
"### Funciones para la búsqueda de ambientes con RegEx\n\nDefino algunas funciones con algunos patrones de RegEx para buscar el número de ambientes por dígitos o por texto.",
"_____no_output_____"
]
],
[
[
"def mono_amb_search(text):\n '''Busca si aparece algun monoambiente en el texto.\n '''\n #pattern_1amb = r'mono\\s?ambiente'\n #este funciona mejor, captura más.\n pattern_1amb = r'mono\\s?h?amb\\.?\\w+' \n regex_1amb = re.compile(pattern_1amb, flags = re.IGNORECASE)\n mono_amb = regex_1amb.findall(text)\n if(mono_amb):\n return 1\n else:\n return np.nan",
"_____no_output_____"
],
[
"def more_amb_search(text): \n '''Busca la cantidad de ambientes con dígitos.\n '''\n \n accepted_lst = ['ambientes','ambientes','amb.','amvientes','anvientes','hambientes',\n 'hamvientes','hanvientes','ambintes','ambtes','ambiemtes','ambietes',\n 'ambieintes','ambeintes','ambentes','ambs','ambietnes','ambienes',\n 'ambientessan','ambc']\n \n accepted = ';'.join(accepted_lst)\n \n pattern_amb = r'(\\d+)\\s*(h?(?:amb|anv|amv)\\.?\\w*)'\n regex_amb = re.compile(pattern_amb, flags = re.IGNORECASE)\n ambientes = regex_amb.findall(text)\n \n if not ambientes:\n return np.nan\n if len(ambientes)>1:\n #si es cero, no encontré nada. si es mayor a uno, es un proyecto o un complejo.\n #lo dejo para después.\n #return np.nan\n amb_lst = [float(item[0]) for item in ambientes if item[1].lower() in accepted]\n \n if amb_lst:\n amb=max(amb_lst)\n if amb==0:\n return np.nan\n else:\n return(max(amb_lst))\n else:\n return np.nan\n \n if len(ambientes)==1:\n if ambientes[0][1].lower() in accepted:\n amb = float(ambientes[0][0])\n if amb==0:\n return np.nan\n# if amb<20:\n# return amb\n# else:\n# return np.nan \n return amb",
"_____no_output_____"
],
[
"def text_amb_search(text):\n \n '''Buscar por un número con palabras y convertirlo a float.\n '''\n \n convert_dict = {\n 'un': 1.0,'uno': 1.0,'dos': 2.0,'tres': 3.0,'cuatro': 4.0,'cinco': 5.0,\n 'seis': 6.0,'siete': 7.0,'ocho': 8.0,'nueve': 9.0,'diez': 10.0,'once': 11.0,\n 'doce': 12.0,'trece': 13.0,'catorce': 14.0,'quince': 15.0\n }\n \n accepted_lst = ['ambientes','ambientes','amb.','amvientes','anvientes','hambientes',\n 'hamvientes','hanvientes','ambintes','ambtes','ambiemtes','ambietes',\n 'ambieintes','ambeintes','ambentes','ambs','ambietnes','ambienes',\n 'ambientessan','ambc']\n \n accepted = ';'.join(accepted_lst)\n pattern_amb = r'([a-z]{2,7})\\s*(h?(?:amb|anv|amv)\\.?\\w*)'\n regex_amb = re.compile(pattern_amb, flags = re.IGNORECASE)\n ambientes = regex_amb.findall(text)\n if not ambientes:\n return np.nan\n if len(ambientes)!=1:\n amb_lst = [convert_dict[item[0].lower()] for item in ambientes if item[1].lower() in accepted\\\n and item[0].lower() in convert_dict.keys()]\n if amb_lst:\n amb = max(amb_lst)\n if amb==0:\n return np.nan\n return amb\n else:\n return np.nan\n else:\n if ambientes[0][1].lower() not in accepted:\n return np.nan\n else:\n if ambientes[0][0].lower() not in convert_dict.keys():\n return np.nan\n else:\n amb = convert_dict[ambientes[0][0].lower()]\n if amb==0:\n return np.nan\n return amb\n ",
"_____no_output_____"
],
[
"def dorm_search(text):\n pattern_dorm = r'([0-9]+)\\s*(?:dorm|habit|habiat|amplios dorm|buenos dorm)\\.?(?:itorio|cion|acion|ación)?(?:s|es)?'\n regex_dorm = re.compile(pattern_dorm, flags = re.IGNORECASE)\n dorms = regex_dorm.findall(text)\n if not dorms:\n return np.nan\n if len(dorms)>1:\n dorm_lst = [float(item) for item in dorms]\n return max(dorm_lst)\n if len(dorms)==1:\n amb = float(dorms[0])+1\n if amb<=20:\n return amb\n else:\n return np.nan",
"_____no_output_____"
],
[
"def dorm_search2(text):\n pattern_dorm = r'(?:dorm|habit|habiat)\\.?(?:itorio|cion|acion|ación)?(?:s|es)?:?\\s*([0-9]+)'\n regex_dorm = re.compile(pattern_dorm, flags = re.IGNORECASE)\n dorms = regex_dorm.findall(text)\n if not dorms:\n return np.nan\n if len(dorms)>1:\n dorm_lst = [float(item) for item in dorms]\n return max(dorm_lst)+1\n if len(dorms)==1:\n amb = float(dorms[0])+1\n if amb<=20:\n return amb\n else:\n return np.nan",
"_____no_output_____"
],
[
"def text_dorm_search(text):\n convert_dict = {\n 'un': 1.0,'uno': 1.0,'dos': 2.0,'tres': 3.0,'cuatro': 4.0,\n 'cinco': 5.0,'seis': 6.0,'siete': 7.0,'ocho': 8.0,'nueve': 9.0,\n 'diez': 10.0,'once': 11.0,'doce': 12.0,'trece': 13.0,\n 'catorce': 14.0,'quince': 15.0\n }\n \n pattern_dorm = r'([a-z]{2,7})\\s*(?:dorm|habit|amplios dorm)\\.?(?:itorio|acion|ación)?(?:s|es)?'\n regex_dorm = re.compile(pattern_dorm, flags = re.IGNORECASE)\n dorms = regex_dorm.findall(text)\n \n if not dorms:\n return np.nan \n if len(dorms)>1:\n dorms_lst = [convert_dict[item.lower()] for item in dorms if item.lower() in convert_dict.keys()]\n if dorms_lst:\n return max(dorms_lst)\n else:\n return np.nan\n if len(dorms)==1:\n if dorms[0].lower() not in convert_dict.keys():\n return np.nan\n else:\n amb = convert_dict[dorms[0].lower()] + 1.0\n if amb<=20:\n return amb\n else:\n return np.nan",
"_____no_output_____"
],
[
"def clean_temp_col(df,tempCol):\n \"\"\"Cleans a tempCol column from a df dataframe. Writes nan on every row.\n \n Parameters\n ----------\n df : pandas.DataFrame\n The dataframe you want to clean a temporary column.\n tempCol: str\n The name passed as a string of the column to clean.\n \"\"\"\n df.loc[:,tempCol]=np.nan",
"_____no_output_____"
],
[
"def new_found(df,tempCol,cleanCol):\n \"\"\"Shows a message about the new data found.\n \n Parameters\n ----------\n df : pandas.DataFrame\n The dataframe to take data from.\n tempCol: str\n The name passed as a string of the temporary column.\n cleanCol: str\n the name passed as a string of the clean column.\n \"\"\"\n print(\"Nuevos: {}.\".format(df[tempCol].notnull().sum()))\n print(\"Total: {}.\".format(df[cleanCol].notnull().sum()))",
"_____no_output_____"
],
[
"import scipy.stats as stats\n\n#valor_mas_chico=0\n\ndef calculate_iqr_values(array):\n \"\"\"Calculates inferior and superior limits of the iqr plus a margin.\n Returns\n ---------\n lim_inf: float\n q1-1.5*iqr\n lim_sup: float\n q3+1.5*iqr\n \"\"\"\n q1,q3=np.percentile(array,[25 ,75])\n arr_iqr = stats.iqr(array)\n lim_inf = q1-1.5*arr_iqr\n# if lim_inf<valor_mas_chico:\n# lim_inf=valor_mas_chico\n lim_sup = q3+1.5*arr_iqr\n return lim_inf,lim_sup\n\ndef check_outliers(x,min_lim,max_lim):\n if x<min_lim:\n return np.nan\n elif x>max_lim:\n return np.nan\n else:\n return x",
"_____no_output_____"
],
[
"data['property_type'].value_counts(dropna=False)",
"_____no_output_____"
],
[
"plt.figure(figsize=(8,8))\nplt.grid(True) \nax = sns.boxplot(x=\"property_type\", y=\"rooms\", data=data, palette=\"Set1\")\nplt.title('Distribución de ambientes por tipo de propiedad')\nplt.xlabel('Tipo de Propiedad')\nplt.ylabel('N° de Ambientes')",
"_____no_output_____"
]
],
[
[
"## Verificar outliers para house",
"_____no_output_____"
]
],
[
[
"data.loc[data['property_type']=='house','rooms'].value_counts(dropna=False)",
"_____no_output_____"
],
[
"iqr_values_house=calculate_iqr_values(data.loc[(data['rooms'].notnull())&(data['property_type']=='house'),'rooms'])\nprint(data.loc[(data['property_type']=='house')&(data['rooms'].notnull()),'rooms'].apply(check_outliers,args=iqr_values_house).unique())\nprint(iqr_values_house)",
"[ 5. 6. 2. 3. 4. 7. nan 8. 1.]\n(0.0, 8.0)\n"
],
[
"#data.loc[(data['property_type']=='house')&(data['rooms']>iqr_values_house[1]),['rooms','description']].sample(10).values\ndata.loc[(data['property_type']=='house')&(data['rooms']>iqr_values_house[1]),'rooms']=np.nan\ndata.loc[data['property_type']=='house','rooms'].value_counts(dropna=False)",
"_____no_output_____"
]
],
[
[
"## Búsqueda de ambientes para house",
"_____no_output_____"
]
],
[
[
"data['clean_amb']=data['rooms']\ndata['temp_amb']=np.nan\n\nfunc_lst = [mono_amb_search,more_amb_search,text_amb_search,dorm_search,text_dorm_search,dorm_search2]\nfor func in func_lst:\n# print(func.__name__)\n# print(data.loc[(data['property_type']=='store')&(data['floor'].isna())]['description'].apply(func).value_counts()\n# )\n clean_temp_col(data,'temp_amb')\n data.loc[(data['property_type']=='house')&(data['rooms'].isna()),'temp_amb']=data.loc[(data['property_type']=='house')&(data['rooms'].isna())]['description'].apply(func)\n data.loc[data['property_type']=='house','temp_amb'] = data.loc[data['property_type']=='house','temp_amb'].apply(check_outliers,args=(1,iqr_values_house[1]))\n data.loc[(data['property_type']=='house')&(data['clean_amb'].isna()),'clean_amb']=data.loc[(data['property_type']=='house')&(data['clean_amb'].isna()),'clean_amb'].add(data.loc[data['property_type']=='house','temp_amb'],fill_value=0)\n\ndata.loc[(data['property_type']=='house')&(data['rooms'].isna()),'rooms'] = data[(data['property_type']=='house')&(data['rooms'].isna())]['rooms'].add(data[data['property_type']=='house']['clean_amb'],fill_value=0)\n#data.loc[(data['property_type']=='store')&(data['clean_fl']).notnull()] \ndata.loc[(data['property_type']=='house')]['rooms'].value_counts(dropna=False,sort=True)",
"_____no_output_____"
]
],
[
[
"## Verificar outliers para PH",
"_____no_output_____"
]
],
[
[
"data.loc[data['property_type']=='PH','rooms'].value_counts(dropna=False)",
"_____no_output_____"
],
[
"iqr_values_PH=calculate_iqr_values(data.loc[(data['rooms'].notnull())&(data['property_type']=='PH'),'rooms'])\nprint(data.loc[(data['property_type']=='PH')&(data['rooms'].notnull()),'rooms'].apply(check_outliers,args=iqr_values_PH).unique())\nprint(iqr_values_PH)",
"[ 4. 6. 2. 3. 1. nan 5. 7.]\n(-1.0, 7.0)\n"
]
],
[
[
"Luego de revisar los outliers de los ambientes de los PH y sus descripciones, se notó que todos salvo el de 20 son correctos.",
"_____no_output_____"
]
],
[
[
"#data.loc[(data['property_type']=='house')&(data['rooms']>iqr_values_house[1]),['rooms','description']].sample(10).values\ndata.loc[(data['property_type']=='PH')&(data['rooms']>iqr_values_PH[1]),'rooms']=np.nan\ndata.loc[data['property_type']=='PH','rooms'].value_counts(dropna=False)",
"_____no_output_____"
]
],
[
[
"## Búsqueda de ambientes para PH",
"_____no_output_____"
]
],
[
[
"data['clean_amb']=data['rooms']\ndata['temp_amb']=np.nan\n\nfunc_lst = [mono_amb_search,more_amb_search,text_amb_search,dorm_search,text_dorm_search,dorm_search2]\nfor func in func_lst:\n# print(func.__name__)\n# print(data.loc[(data['property_type']=='store')&(data['floor'].isna())]['description'].apply(func).value_counts()\n# )\n clean_temp_col(data,'temp_amb')\n data.loc[(data['property_type']=='PH')&(data['rooms'].isna()),'temp_amb']=data.loc[(data['property_type']=='PH')&(data['rooms'].isna())]['description'].apply(func)\n data.loc[data['property_type']=='PH','temp_amb'] = data.loc[data['property_type']=='PH','temp_amb'].apply(check_outliers,args=iqr_values_PH)\n data.loc[(data['property_type']=='PH')&(data['clean_amb'].isna()),'clean_amb']=data.loc[(data['property_type']=='PH')&(data['clean_amb'].isna()),'clean_amb'].add(data.loc[data['property_type']=='PH','temp_amb'],fill_value=0)\n\ndata.loc[(data['property_type']=='PH')&(data['rooms'].isna()),'rooms'] = data[(data['property_type']=='PH')&(data['rooms'].isna())]['rooms'].add(data[data['property_type']=='PH']['clean_amb'],fill_value=0)\n#data.loc[(data['property_type']=='store')&(data['clean_fl']).notnull()] \ndata.loc[(data['property_type']=='PH')]['rooms'].value_counts(dropna=False,sort=True)",
"_____no_output_____"
],
[
"#data.loc[(data['property_type']=='PH')&(data['rooms'].isna()),'description'].sample(10).values",
"_____no_output_____"
]
],
[
[
"## Verificar outliers para store",
"_____no_output_____"
]
],
[
[
"data.loc[data['property_type']=='store','rooms'].value_counts(dropna=False)",
"_____no_output_____"
],
[
"iqr_values_store=calculate_iqr_values(data.loc[(data['rooms'].notnull())&(data['property_type']=='store'),'rooms'])\nprint(data.loc[(data['property_type']=='store')&(data['rooms'].notnull()),'rooms'].apply(check_outliers,args=iqr_values_store).unique())\nprint(iqr_values_store)",
"[ 1. nan 2. 3. 6. 4. 5.]\n(-2.0, 6.0)\n"
],
[
"#data.loc[(data['property_type']=='store')&(data['rooms']>iqr_values_store[1]),['rooms','description']].values",
"_____no_output_____"
],
[
"data.loc[(data['property_type']=='store')&(data['rooms']>iqr_values_store[1]),'rooms']=np.nan\ndata.loc[data['property_type']=='store','rooms'].value_counts(dropna=False)",
"_____no_output_____"
]
],
[
[
"## Búsqueda de ambientes para store",
"_____no_output_____"
]
],
[
[
"data['clean_amb']=data['rooms']\ndata['temp_amb']=np.nan\n\nfunc_lst = [mono_amb_search,more_amb_search,text_amb_search,dorm_search,text_dorm_search,dorm_search2]\nfor func in func_lst:\n# print(func.__name__)\n# print(data.loc[(data['property_type']=='store')&(data['floor'].isna())]['description'].apply(func).value_counts()\n# )\n clean_temp_col(data,'temp_amb')\n data.loc[(data['property_type']=='store')&(data['rooms'].isna()),'temp_amb']=data.loc[(data['property_type']=='store')&(data['rooms'].isna())]['description'].apply(func)\n data.loc[data['property_type']=='store','temp_amb'] = data.loc[data['property_type']=='store','temp_amb'].apply(check_outliers,args=iqr_values_store)\n data.loc[(data['property_type']=='store')&(data['clean_amb'].isna()),'clean_amb']=data.loc[(data['property_type']=='store')&(data['clean_amb'].isna()),'clean_amb'].add(data.loc[data['property_type']=='store','temp_amb'],fill_value=0)\n\ndata.loc[(data['property_type']=='store')&(data['rooms'].isna()),'rooms'] = data[(data['property_type']=='store')&(data['rooms'].isna())]['rooms'].add(data[data['property_type']=='PH']['clean_amb'],fill_value=0)\n#data.loc[(data['property_type']=='store')&(data['clean_fl']).notnull()] \ndata.loc[(data['property_type']=='store')&(data['rooms'].isna())&(data['surface_total_in_m2']<50),'rooms']=1\ndata.loc[(data['property_type']=='store')]['rooms'].value_counts(dropna=False,sort=True)",
"_____no_output_____"
]
],
[
[
"## Verificar outliers para apartment",
"_____no_output_____"
]
],
[
[
"data.loc[data['property_type']=='apartment','rooms'].value_counts(dropna=False)",
"_____no_output_____"
],
[
"iqr_values_apartment=calculate_iqr_values(data.loc[(data['rooms'].notnull())&(data['property_type']=='apartment'),'rooms'])\nprint(data.loc[(data['property_type']=='apartment')&(data['rooms'].notnull()),'rooms'].apply(check_outliers,args=iqr_values_apartment).unique())\nprint(iqr_values_apartment)",
"[ 1. 3. 4. 2. nan]\n(0.5, 4.5)\n"
],
[
"#data.loc[(data['property_type']=='apartment')&(data['rooms']==iqr_values_apartment[1]+3),['rooms','description']].sample(10).values",
"_____no_output_____"
]
],
[
[
"Luego de revisar los outliers de los ambientes de los apartment y sus descripciones, se decidió tomar como válidos a todos los que aparecen hasta 7 ambientes. Esto es debido a que existe una buena cantidad de publicaciones donde se venden varias propiedades juntas o departamentos de varios pisos.",
"_____no_output_____"
]
],
[
[
"#data.loc[(data['property_type']=='apartment')&(data['rooms']>iqr_values_apartment[1]),['rooms','description']].sample(10).values\ndata.loc[(data['property_type']=='apartment')&(data['rooms']>7),'rooms']=np.nan\ndata.loc[data['property_type']=='apartment','rooms'].value_counts(dropna=False)",
"_____no_output_____"
]
],
[
[
"## Búsqueda de ambientes para apartment",
"_____no_output_____"
]
],
[
[
"print(iqr_values_apartment)",
"(0.5, 4.5)\n"
],
[
"data['clean_amb']=data['rooms']\ndata['temp_amb']=np.nan\n\nfunc_lst = [mono_amb_search,more_amb_search,text_amb_search,dorm_search,text_dorm_search,dorm_search2]\nfor func in func_lst:\n# print(func.__name__)\n# print(data.loc[(data['property_type']=='store')&(data['floor'].isna())]['description'].apply(func).value_counts()\n# )\n clean_temp_col(data,'temp_amb')\n data.loc[(data['property_type']=='apartment')&(data['rooms'].isna()),'temp_amb']=data.loc[(data['property_type']=='apartment')&(data['rooms'].isna())]['description'].apply(func)\n data.loc[data['property_type']=='apartment','temp_amb'] = data.loc[data['property_type']=='apartment','temp_amb'].apply(check_outliers,args=(iqr_values_apartment[0],iqr_values_apartment[1]+2.5))\n data.loc[(data['property_type']=='apartment')&(data['clean_amb'].isna()),'clean_amb']=data.loc[(data['property_type']=='apartment')&(data['clean_amb'].isna()),'clean_amb'].add(data.loc[data['property_type']=='apartment','temp_amb'],fill_value=0)\n\ndata.loc[(data['property_type']=='apartment')&(data['rooms'].isna()),'rooms'] = data[(data['property_type']=='apartment')&(data['rooms'].isna())]['rooms'].add(data[data['property_type']=='apartment']['clean_amb'],fill_value=0)\n#data.loc[(data['property_type']=='store')&(data['clean_fl']).notnull()] \ndata.loc[(data['property_type']=='apartment')]['rooms'].value_counts(dropna=False,sort=True)",
"_____no_output_____"
],
[
"#data.loc[(data['property_type']=='PH')&(data['rooms'].isna()),'description'].sample(10).values",
"_____no_output_____"
],
[
"data.drop(['clean_amb','temp_amb'],axis=1,inplace=True)\ndata.isna().sum()",
"_____no_output_____"
],
[
"plt.figure(figsize=(8,8))\nplt.grid(True) \nax = sns.boxplot(x=\"property_type\", y=\"rooms\", data=data, palette=\"Set1\")\nplt.title('Distribución de ambientes por tipo de propiedad')\nplt.xlabel('Tipo de Propiedad')\nplt.ylabel('N° de Ambientes')",
"_____no_output_____"
]
],
[
[
"# Variables Dummy: Amenities\n\nSe busca crear variables dummy de características que sumen para la regresión del desafío 2.",
"_____no_output_____"
]
],
[
[
"sin_cochera = data.loc[:,'description'].str.contains(r\"(?:sin cochera|sin estacionamiento|sin garage)\",flags=re.IGNORECASE,regex=True)\nsin_cochera = sin_cochera.apply(lambda x: 0 if x else np.nan)\n\ncochera = data.loc[sin_cochera.isna(),'description'].str.contains(r\"(?:cochera|estacionamiento|garage)\",flags=re.IGNORECASE,regex=True).astype(float)\ncochera = cochera.add(sin_cochera,fill_value=0)\n\nsin_gimnasio = data.loc[:,'description'].str.contains(r\"(?:sin gym|sin gimnasio|sin gimnasia)\",flags=re.IGNORECASE,regex=True)\nsin_gimnasio = sin_gimnasio.apply(lambda x: 0 if x else np.nan)\n\ngimnasio = data.loc[:,'description'].str.contains(r\"(?:gym|gimnasio|gimnasia)\",flags=re.IGNORECASE,regex=True).astype(float)\ngimnasio = gimnasio.add(sin_gimnasio,fill_value=0)\n\nsin_piscina = data.loc[:,'description'].str.contains(r\"(?:sin piscina|sin pisicina|sin pileta)\",flags=re.IGNORECASE,regex=True)\nsin_piscina = sin_piscina.apply(lambda x: 0 if x else np.nan)\n\npiscina = data.loc[:,'description'].str.contains(r\"(?:piscina|pisicina|pileta)\",flags=re.IGNORECASE,regex=True).astype(float)\npiscina = piscina.add(sin_piscina,fill_value=0)\n\nsin_sum = data.loc[:,'description'].str.contains(r\"(?:sin SUM|sin salón|sin salon|sin playroom)\",flags=re.IGNORECASE,regex=True)\nsin_sum = sin_sum.apply(lambda x: 0 if x else np.nan)\n\nSUM = data.loc[:,'description'].str.contains(r\"(?:SUM|salón|salon|playroom)\",flags=re.IGNORECASE,regex=True).astype(float)\nSUM = SUM.add(sin_gimnasio,fill_value=0)\n\nsin_laundry = data.loc[:,'description'].str.contains(r\"(?:sin laundy|sin laundry|sin lavadero)\",flags=re.IGNORECASE,regex=True)\nsin_laundry = sin_laundry.apply(lambda x: 0 if x else np.nan)\n\nlaundry = data.loc[:,'description'].str.contains(r\"(?:laundy|laundry|lavadero)\",flags=re.IGNORECASE,regex=True).astype(float)\nlaundry = laundry.add(sin_gimnasio,fill_value=0)\n\nsin_balcon = data.loc[:,'description'].str.contains(r\"(?:sin balcon|sin balcón)\",flags=re.IGNORECASE,regex=True)\nsin_balcon = sin_balcon.apply(lambda x: 0 if x else np.nan)\n\nbalcon = data.loc[:,'description'].str.contains(r\"(?:balcon|balcón)\",flags=re.IGNORECASE,regex=True).astype(float)\nbalcon = balcon.add(sin_gimnasio,fill_value=0)\n\nsin_patio = data.loc[:,'description'].str.contains(r\"(?:sin patio|sin jardín|sin jardin)\",flags=re.IGNORECASE,regex=True)\nsin_patio = sin_patio.apply(lambda x: 0 if x else np.nan)\n\npatio = data.loc[:,'description'].str.contains(r\"(?:patio|jardín|jardin)\",flags=re.IGNORECASE,regex=True).astype(float)\npatio = patio.add(sin_gimnasio,fill_value=0)\n\namenities = {\n 'cochera':cochera,\n 'gimnasio':gimnasio,\n 'piscina':piscina,\n 'SUM':SUM,\n 'laundry':laundry,\n 'balcon':balcon,\n 'patio':patio,\n}\n\nfor key,item in amenities.items():\n data[key]=item\ndata.columns",
"_____no_output_____"
],
[
"data.isna().sum()",
"_____no_output_____"
]
],
[
[
"# Limpieza de Superficie (Total y Cubierta)\n\nDe las inspecciónes de la columnas \"surface_total_in_m2\" y \"surface_covered_in_m2\", vemos que los valores comienzan desde el 0.0, por ende consideraremos superficies validas aquellas que sean igual o superen los diez metros cuadrados.",
"_____no_output_____"
]
],
[
[
"min_sup_valida = 10\ndata.loc[data['surface_total_in_m2']<min_sup_valida,'surface_total_in_m2']=np.nan\ndata.loc[data['surface_covered_in_m2']<min_sup_valida,'surface_total_in_m2']=np.nan",
"_____no_output_____"
]
],
[
[
"La mediana de las superficies por la cantidad de ambientes. En los casos de 1 y 2 ambientes parece lógico. ",
"_____no_output_____"
]
],
[
[
"apartment_df = data.groupby('property_type').get_group('apartment')\n\napartment_df[['rooms','surface_total_in_m2','surface_covered_in_m2']].groupby('rooms').median()\ndata.loc[(data['property_type']=='apartment'),'surface_total_in_m2'].isna().sum()",
"_____no_output_____"
],
[
"apartment_df = data.loc[data['property_type']=='apartment'].copy()\nhouse_df = data.loc[data['property_type']=='house'].copy()\nph_df = data.loc[data['property_type']=='PH'].copy()\nstore_df = data.loc[data['property_type']=='store'].copy()\n\nplt.figure(figsize=(14,10))\nplt.subplot(221)\nplt.grid(True) \n#ax = sns.boxplot(x=\"rooms\", y=\"surface_total_in_m2\", data=apartment_df, palette=\"Set1\")\nax = sns.boxplot(x=\"rooms\", y=\"surface_covered_in_m2\", data=apartment_df, palette=\"Set1\")\nplt.title('Superficie por número de ambientes (departamentos)')\nplt.xlabel('Número de Ambientes')\nplt.ylabel('Superficie [m2]')\n\nplt.subplot(222)\nplt.grid(True) \n#ax = sns.boxplot(x=\"rooms\", y=\"surface_total_in_m2\", data=house_df, palette=\"Set1\")\nax = sns.boxplot(x=\"rooms\", y=\"surface_covered_in_m2\", data=house_df, palette=\"Set1\")\nplt.title('Superficie por número de ambientes (Casa)')\nplt.xlabel('Número de Ambientes')\nplt.ylabel('Superficie [m2]')\n\nplt.subplot(223)\nplt.grid(True) \n#ax = sns.boxplot(x=\"rooms\", y=\"surface_total_in_m2\", data=ph_df, palette=\"Set1\")\nax = sns.boxplot(x=\"rooms\", y=\"surface_covered_in_m2\", data=ph_df, palette=\"Set1\")\nplt.title('Superficie por número de ambientes (PH)')\nplt.xlabel('Número de Ambientes')\nplt.ylabel('Superficie [m2]')\n\nplt.subplot(224)\nplt.grid(True) \n#ax = sns.boxplot(x=\"rooms\", y=\"surface_total_in_m2\", data=store_df, palette=\"Set1\")\nax = sns.boxplot(x=\"rooms\", y=\"surface_covered_in_m2\", data=store_df, palette=\"Set1\")\nplt.title('Superficie por número de ambientes (Store)')\nplt.xlabel('Número de Ambientes')\nplt.ylabel('Superficie [m2]')\nplt.savefig(\"../Images/Rooms-Before.png\")",
"_____no_output_____"
]
],
[
[
"## Limpiando Superficies ",
"_____no_output_____"
],
[
"### apartment",
"_____no_output_____"
]
],
[
[
"sup_tot_limits_apt = []\nsup_cub_limits_apt = []\nfor i in range(1,8,1):\n a=calculate_iqr_values(data.loc[(data['rooms']==i)&(data['property_type']=='apartment')&(data['surface_total_in_m2'].notnull()),'surface_total_in_m2'])[1]\n sup_tot_limits_apt.append((10,a))\n b=calculate_iqr_values(data.loc[(data['rooms']==i)&(data['property_type']=='apartment')&(data['surface_covered_in_m2'].notnull()),'surface_covered_in_m2'])[1]\n sup_cub_limits_apt.append((10,b))\nprint(sup_tot_limits_apt)\nprint(sup_cub_limits_apt)\n\nfor i in range(7):\n data.loc[(data['property_type']=='apartment')&(data['surface_total_in_m2']>sup_tot_limits_apt[i][1]),'surface_total_in_m2']=np.nan\n data.loc[(data['property_type']=='apartment')&(data['surface_covered_in_m2']>sup_cub_limits_apt[i][1]),'surface_covered_in_m2']=np.nan\n data.loc[(data['property_type']=='apartment')&(data['surface_total_in_m2']<sup_tot_limits_apt[i][0]),'surface_total_in_m2']=np.nan\n data.loc[(data['property_type']=='apartment')&(data['surface_covered_in_m2']<sup_cub_limits_apt[i][0]),'surface_covered_in_m2']=np.nan\napartment_df[['rooms','surface_total_in_m2','surface_covered_in_m2']].groupby('rooms').median()",
"[(10, 64.5), (10, 89.0), (10, 154.0), (10, 252.0), (10, 410.0), (10, 531.75), (10, 537.5)]\n[(10, 55.0), (10, 75.0), (10, 124.0), (10, 210.0), (10, 361.25), (10, 502.625), (10, 477.5)]\n"
]
],
[
[
"### house",
"_____no_output_____"
]
],
[
[
"sup_tot_limits_hs = []\nsup_cub_limits_hs = []\nfor i in range(1,8,1):\n a=calculate_iqr_values(data.loc[(data['rooms']==i)&(data['property_type']=='house')&(data['surface_total_in_m2'].notnull()),'surface_total_in_m2'])[1]\n sup_tot_limits_hs.append((10,a))\n b=calculate_iqr_values(data.loc[(data['rooms']==i)&(data['property_type']=='house')&(data['surface_covered_in_m2'].notnull()),'surface_covered_in_m2'])[1]\n sup_cub_limits_hs.append((10,b))\nprint(sup_tot_limits_hs)\nprint(sup_cub_limits_hs)\n\nfor i in range(7):\n data.loc[(data['property_type']=='house')&(data['surface_total_in_m2']>sup_tot_limits_hs[i][1]),'surface_total_in_m2']=np.nan\n data.loc[(data['property_type']=='house')&(data['surface_covered_in_m2']>sup_cub_limits_hs[i][1]),'surface_covered_in_m2']=np.nan\n data.loc[(data['property_type']=='house')&(data['surface_total_in_m2']<sup_tot_limits_hs[i][0]),'surface_total_in_m2']=np.nan\n data.loc[(data['property_type']=='house')&(data['surface_covered_in_m2']<sup_cub_limits_hs[i][0]),'surface_covered_in_m2']=np.nan\n#apartment_df[['rooms','surface_total_in_m2','surface_covered_in_m2']].groupby('rooms').median()",
"[(10, 945.0), (10, 900.0), (10, 827.5), (10, 880.0), (10, 1126.5), (10, 1365.0), (10, 1561.0)]\n[(10, 462.5), (10, 465.0), (10, 400.0), (10, 428.5), (10, 521.875), (10, 612.875), (10, 683.75)]\n"
]
],
[
[
"### PH",
"_____no_output_____"
]
],
[
[
"sup_tot_limits_ph = []\nsup_cub_limits_ph = []\nfor i in range(1,6,1):\n a=calculate_iqr_values(data.loc[(data['rooms']==i)&(data['property_type']=='PH')&(data['surface_total_in_m2'].notnull()),'surface_total_in_m2'])[1]\n sup_tot_limits_ph.append((10,a))\n b=calculate_iqr_values(data.loc[(data['rooms']==i)&(data['property_type']=='PH')&(data['surface_covered_in_m2'].notnull()),'surface_covered_in_m2'])[1]\n sup_cub_limits_ph.append((10,b))\nprint(sup_tot_limits_ph)\nprint(sup_cub_limits_ph)\n\nfor i in range(5):\n data.loc[(data['property_type']=='PH')&(data['surface_total_in_m2']>sup_tot_limits_ph[i][1]),'surface_total_in_m2']=np.nan\n data.loc[(data['property_type']=='PH')&(data['surface_covered_in_m2']>sup_cub_limits_ph[i][1]),'surface_covered_in_m2']=np.nan\n data.loc[(data['property_type']=='PH')&(data['surface_total_in_m2']<sup_tot_limits_ph[i][0]),'surface_total_in_m2']=np.nan\n data.loc[(data['property_type']=='PH')&(data['surface_covered_in_m2']<sup_cub_limits_ph[i][0]),'surface_covered_in_m2']=np.nan\n#apartment_df[['rooms','surface_total_in_m2','surface_covered_in_m2']].groupby('rooms').median()",
"[(10, 224.0), (10, 109.5), (10, 198.0), (10, 294.375), (10, 372.75)]\n[(10, 147.125), (10, 90.0), (10, 138.0), (10, 197.5), (10, 257.5)]\n"
]
],
[
[
"### store",
"_____no_output_____"
]
],
[
[
"sup_tot_limits_str = []\nsup_cub_limits_str = []\nfor i in range(1,5,1):\n a=calculate_iqr_values(data.loc[(data['rooms']==i)&(data['property_type']=='store')&(data['surface_total_in_m2'].notnull()),'surface_total_in_m2'])[1]\n sup_tot_limits_str.append((10,a))\n b=calculate_iqr_values(data.loc[(data['rooms']==i)&(data['property_type']=='store')&(data['surface_covered_in_m2'].notnull()),'surface_covered_in_m2'])[1]\n sup_cub_limits_str.append((10,b))\nprint(sup_tot_limits_str)\nprint(sup_cub_limits_str)\n\nfor i in range(4):\n data.loc[(data['property_type']=='store')&(data['surface_total_in_m2']>sup_tot_limits_str[i][1]),'surface_total_in_m2']=np.nan\n data.loc[(data['property_type']=='store')&(data['surface_covered_in_m2']>sup_cub_limits_str[i][1]),'surface_covered_in_m2']=np.nan\n data.loc[(data['property_type']=='store')&(data['surface_total_in_m2']<sup_tot_limits_str[i][0]),'surface_total_in_m2']=np.nan\n data.loc[(data['property_type']=='store')&(data['surface_covered_in_m2']<sup_cub_limits_str[i][0]),'surface_covered_in_m2']=np.nan\n#apartment_df[['rooms','surface_total_in_m2','surface_covered_in_m2']].groupby('rooms').median()",
"[(10, 67.5), (10, 295.0), (10, 3128.375), (10, 356.0)]\n[(10, 65.0), (10, 177.25), (10, 424.25), (10, 198.125)]\n"
]
],
[
[
"Me armo un dataframe con los datos de apartment y en cada fila la superficie segun número de ambientes.",
"_____no_output_____"
]
],
[
[
"apartment_df = data.loc[data['property_type']=='apartment'].copy()\nhouse_df = data.loc[data['property_type']=='house'].copy()\nph_df = data.loc[data['property_type']=='PH'].copy()\nstore_df = data.loc[data['property_type']=='store'].copy()\n\nplt.figure(figsize=(14,10))\nplt.subplot(221)\nplt.grid(True) \n#ax = sns.boxplot(x=\"rooms\", y=\"surface_total_in_m2\", data=apartment_df, palette=\"Set1\")\nax = sns.boxplot(x=\"rooms\", y=\"surface_covered_in_m2\", data=apartment_df, palette=\"Set1\")\nplt.title('Superficie por número de ambientes (departamentos)')\nplt.xlabel('Número de Ambientes')\nplt.ylabel('Superficie [m2]')\n\nplt.subplot(222)\nplt.grid(True) \n#ax = sns.boxplot(x=\"rooms\", y=\"surface_total_in_m2\", data=house_df, palette=\"Set1\")\nax = sns.boxplot(x=\"rooms\", y=\"surface_covered_in_m2\", data=house_df, palette=\"Set1\")\nplt.title('Superficie por número de ambientes (Casa)')\nplt.xlabel('Número de Ambientes')\nplt.ylabel('Superficie [m2]')\n\nplt.subplot(223)\nplt.grid(True) \n#ax = sns.boxplot(x=\"rooms\", y=\"surface_total_in_m2\", data=ph_df, palette=\"Set1\")\nax = sns.boxplot(x=\"rooms\", y=\"surface_covered_in_m2\", data=ph_df, palette=\"Set1\")\nplt.title('Superficie por número de ambientes (PH)')\nplt.xlabel('Número de Ambientes')\nplt.ylabel('Superficie [m2]')\n\nplt.subplot(224)\nplt.grid(True) \n#ax = sns.boxplot(x=\"rooms\", y=\"surface_total_in_m2\", data=store_df, palette=\"Set1\")\nax = sns.boxplot(x=\"rooms\", y=\"surface_covered_in_m2\", data=store_df, palette=\"Set1\")\nplt.title('Superficie por número de ambientes (Store)')\nplt.xlabel('Número de Ambientes')\nplt.ylabel('Superficie [m2]')\nplt.savefig(\"../Images/Rooms-After.png\")",
"_____no_output_____"
]
],
[
[
"## Funciones para búsqueda de superficie",
"_____no_output_____"
]
],
[
[
"def sup_total_search(text):\n pattern = r'(?:lote de|sup\\.\\s?total|superficie total|sup\\.\\s?tot\\.)\\s?:?\\s*(\\d+[,|\\.]?\\d*)\\s?(?:m2|mts2|mtrs2|metros\\s?cuadrados|m²)'\n regex = re.compile(pattern, flags = re.IGNORECASE)\n sup_tot_lst = regex.findall(text)\n \n if not sup_tot_lst:\n return np.nan\n if len(sup_tot_lst)!=1:\n return np.nan\n sup=pd.to_numeric(sup_tot_lst[0].replace(',','.'))\n if sup<10:\n return np.nan\n return sup\ndef sup_total_search2(text):\n pattern = r'(\\d+[,|\\.]?\\d*)\\s?(?:m2|mts2|mtrs2|metros\\s?cuadrados|m²)'\n regex = re.compile(pattern, flags = re.IGNORECASE)\n sup_tot_lst = regex.findall(text)\n \n if not sup_tot_lst:\n return np.nan\n if len(sup_tot_lst)>1:\n sup_lst = [pd.to_numeric(item.replace(',','.')) for item in sup_tot_lst]\n if not sup_lst:\n return np.nan\n elif len(sup_lst)>1:\n return np.nan\n elif sup_lst<10:\n return np.nan\n else:\n return sup_lst[0]\n \n if len(sup_tot_lst)==1:\n return pd.to_numeric(sup_tot_lst[0].replace(',','.'))\n\ndef sup_cub_search(text):\n pattern = r'(?:superficie\\s?cubierta:?|sup\\.?\\s?cub.?:?|sub\\.?\\s?cubierta:?|metros.\\s?cubiertos:?|cub\\.?)\\s?:?\\s?(\\d+,?\\.?\\d*)\\s?(?:m2|mt2|mtrs2|mtrs\\.?|mts\\.?||m²)'\n regex = re.compile(pattern, flags = re.IGNORECASE)\n sup_cub_lst = regex.findall(text)\n if not sup_cub_lst:\n return np.nan\n sup_cub_lst = [item.replace(',','.') for item in sup_cub_lst]\n if len(sup_cub_lst)==1:\n sup=pd.to_numeric(sup_cub_lst[0])\n if sup<10:\n return np.nan\n else:\n return sup\n else:\n return np.nan\n #return(max(pd.to_numeric(sup_cub_lst)))\n\ndef sup_cub_search2(text):\n pattern = r'(\\d+,?\\.?\\d*)\\s?(?:m2|mtrs2|mtrs\\.?|mts\\.?|m²|metros|metros\\s?cuadrados|metros\\s?cuadrados\\s?de|),?\\s?(?:construidos|cubiertos|cub\\.?)'\n regex = re.compile(pattern, flags = re.IGNORECASE)\n sup_cub_lst = regex.findall(text)\n if not sup_cub_lst:\n return np.nan\n sup_cub_lst = [item.replace(',','.') for item in sup_cub_lst]\n if len(sup_cub_lst)==1:\n sup=pd.to_numeric(sup_cub_lst[0])\n if sup<10:\n return np.nan\n else:\n return sup\n else:\n return np.nan\n #return(max(pd.to_numeric(sup_cub_lst)))\n ",
"_____no_output_____"
]
],
[
[
"## Columnas temporales\n\nColumnas temporales:\n\ntemp_amb: columna para obtener el resultado del apply de alguna función con RegEx. Puede usarse sobre 'title' o 'description'. empieza en np.nan\n\nclean_amb: columna para ir acumulando los resultados de cada función con RegEx. Puede usarse sobre 'title' o 'description'. empieza inicializada con los valores originales de 'rooms'.",
"_____no_output_____"
]
],
[
[
"#Columnas temporales para superficie cubierta\ndata['clean_sup_cub']=data['surface_covered_in_m2']\ndata['temp_sup_cub']=np.nan\n\n#Columnas temporales para superficie total\ndata['clean_sup_tot']=data['surface_total_in_m2']\ndata['temp_sup_tot']=np.nan",
"_____no_output_____"
]
],
[
[
"## Búsqueda de superficie cubierta",
"_____no_output_____"
]
],
[
[
"#data['description'] = data['description'].fillna(\"-\")\nsup_cub_func_lst = [sup_cub_search,sup_cub_search2]\ncol_lst = ['title','description']\nfor col in col_lst:\n print(\"------------------------\")\n print(\"Columna: {}\".format(col))\n print(\"------------------------\")\n for func in sup_cub_func_lst:\n clean_temp_col(data,'temp_sup_cub')\n data.loc[data['clean_sup_cub'].isna(),'temp_sup_cub'] = data[data['clean_sup_cub'].isna()][col].apply(func)\n data.loc[data['clean_sup_cub'].isna(),'clean_sup_cub'] = data['clean_sup_cub'].add(data['temp_sup_cub'],fill_value=0)\n print(func.__name__)\n new_found(data,'temp_sup_cub','clean_sup_cub')",
"------------------------\nColumna: title\n------------------------\nsup_cub_search\nNuevos: 0.\nTotal: 67098.\nsup_cub_search2\nNuevos: 78.\nTotal: 67176.\n------------------------\nColumna: description\n------------------------\nsup_cub_search\nNuevos: 994.\nTotal: 68170.\nsup_cub_search2\nNuevos: 2303.\nTotal: 70473.\n"
]
],
[
[
"## Búsqueda de superficie total",
"_____no_output_____"
]
],
[
[
"sup_tot_func_lst = [sup_total_search,sup_total_search2]\ncol_lst = ['title','description']\nfor col in col_lst:\n print(\"------------------------\")\n print(\"Columna: {}\".format(col))\n print(\"------------------------\")\n for func in sup_tot_func_lst:\n clean_temp_col(data,'temp_sup_tot')\n data.loc[data['clean_sup_tot'].isna(),'temp_sup_tot'] = data[data['clean_sup_tot'].isna()][col].apply(func)\n data.loc[data['clean_sup_tot'].isna(),'clean_sup_tot'] = data['clean_sup_tot'].add(data['temp_sup_tot'],fill_value=0)\n print(func.__name__)\n new_found(data,'temp_sup_tot','clean_sup_tot')",
"------------------------\nColumna: title\n------------------------\nsup_total_search\nNuevos: 47.\nTotal: 52208.\nsup_total_search2\nNuevos: 3054.\nTotal: 55262.\n------------------------\nColumna: description\n------------------------\nsup_total_search\nNuevos: 1605.\nTotal: 56867.\nsup_total_search2\nNuevos: 8889.\nTotal: 65756.\n"
]
],
[
[
"Filtrar los nuevos valores con los outliers de antes.",
"_____no_output_____"
]
],
[
[
"sup_tot_limits = [sup_tot_limits_apt,sup_tot_limits_hs,sup_tot_limits_ph,sup_tot_limits_str]\nsup_cub_limits = [sup_cub_limits_apt,sup_cub_limits_hs,sup_cub_limits_ph,sup_cub_limits_str]\nproperty_type = ('apartment','house','PH','store')",
"_____no_output_____"
],
[
"for tipo,limites_tot,limites_cub in zip(property_type,sup_tot_limits,sup_cub_limits):\n print(tipo,limites_tot,limites_cub)",
"apartment [(10, 64.5), (10, 89.0), (10, 154.0), (10, 252.0), (10, 410.0), (10, 531.75), (10, 537.5)] [(10, 55.0), (10, 75.0), (10, 124.0), (10, 210.0), (10, 361.25), (10, 502.625), (10, 477.5)]\nhouse [(10, 945.0), (10, 900.0), (10, 827.5), (10, 880.0), (10, 1126.5), (10, 1365.0), (10, 1561.0)] [(10, 462.5), (10, 465.0), (10, 400.0), (10, 428.5), (10, 521.875), (10, 612.875), (10, 683.75)]\nPH [(10, 224.0), (10, 109.5), (10, 198.0), (10, 294.375), (10, 372.75)] [(10, 147.125), (10, 90.0), (10, 138.0), (10, 197.5), (10, 257.5)]\nstore [(10, 67.5), (10, 295.0), (10, 3128.375), (10, 356.0)] [(10, 65.0), (10, 177.25), (10, 424.25), (10, 198.125)]\n"
],
[
"data['surface_total_in_m2'] = data['clean_sup_tot']\ndata['surface_covered_in_m2'] = data['clean_sup_cub']\n\ndata.drop(['clean_sup_tot','temp_sup_tot','clean_sup_cub','temp_sup_cub'],axis=1,inplace=True)",
"_____no_output_____"
],
[
"for tipo,limites_tot,limites_cub in zip(property_type,sup_tot_limits,sup_cub_limits):\n for i in range(len(limites_tot)): \n data.loc[(data['property_type']==str(tipo))&(data['surface_total_in_m2']>limites_tot[i][1]),'surface_total_in_m2']=np.nan\n data.loc[(data['property_type']==str(tipo))&(data['surface_covered_in_m2']>limites_cub[i][1]),'surface_covered_in_m2']=np.nan\n data.loc[(data['property_type']==str(tipo))&(data['surface_total_in_m2']<limites_tot[i][0]),'surface_total_in_m2']=np.nan\n data.loc[(data['property_type']==str(tipo))&(data['surface_covered_in_m2']<limites_cub[i][0]),'surface_covered_in_m2']=np.nan",
"_____no_output_____"
],
[
"#data.to_csv('../Data/falta_sup_price.csv')",
"_____no_output_____"
]
],
[
[
"## Imputación de superficies\n\nSe calcula el índice de construcción por zona y por tipo de propiedad.\n\nEl índice se define como:\n\n<center>$Indice\\ de\\ Construcción = \\frac{Superficie Cubierta}{Superficie Total}$</center>\n\nLuego, se calcula los valores faltantes de Superficie Cubierta multiplicando al índice por la Superficie Total.\nY, la Superficie Total como la Superficie Cubierta dividido el Indice",
"_____no_output_____"
]
],
[
[
"data['construction_index']=data['surface_covered_in_m2']/data['surface_total_in_m2']\nplace_const_index = data[['property_type','place_name','construction_index']].groupby(['place_name','property_type']).mean()\ndata.drop('construction_index',axis=1,inplace=True)\n\ndata = pd.merge(data,place_const_index,on=['place_name','property_type'],how='inner')\n\nfalta_sup_cub_index = data.loc[data['construction_index'].notnull()&(data['surface_covered_in_m2'].isna())&(data['surface_total_in_m2'].notnull()),['surface_covered_in_m2','surface_total_in_m2','construction_index']].index\ndata.loc[falta_sup_cub_index,'surface_covered_in_m2'] = data.loc[falta_sup_cub_index,'construction_index']*data.loc[falta_sup_cub_index,'surface_total_in_m2']\n\nfalta_sup_tot_index = data.loc[data['construction_index'].notnull()&(data['surface_covered_in_m2'].notnull())&(data['surface_total_in_m2'].isna()),['surface_covered_in_m2','surface_total_in_m2','construction_index']].index\ndata.loc[falta_sup_tot_index,'surface_total_in_m2'] = data.loc[falta_sup_tot_index,'surface_covered_in_m2']/data.loc[falta_sup_tot_index,'construction_index']",
"_____no_output_____"
],
[
"data.isna().sum()",
"_____no_output_____"
]
],
[
[
"# ANÁLISIS DE LAS DIFERENTES COTIZACIONES",
"_____no_output_____"
],
[
"En primer lugar evaluamos las diferentes monedas que existen en el dataset. ",
"_____no_output_____"
]
],
[
[
"data['currency'].value_counts()",
"_____no_output_____"
]
],
[
[
"de la inspección de la columna \"currency\", podemos ver que salvo 3 registros, toda la información está cotizada en \nPesos Argentinos o Dólares. La decisión será chequear si esos 3 registros se tratan de errores de imputación y corregir o \nconvertir los mismos a dólares para manejar una sola divisa de cambio.\nLa elección se basa además en la descripción del dataset recibido ya que en \"currency\" sólo contempla la moneda ARS y DOL.",
"_____no_output_____"
],
[
"#### Chequeamos Tipos de Cambio presentes en el Dataset",
"_____no_output_____"
],
[
"#### Valores con currency ARS",
"_____no_output_____"
]
],
[
[
"((data.loc[data['currency']=='ARS','price_aprox_local_currency'])/(data.loc[data['currency']=='ARS','price_aprox_usd'])).round(decimals=4).value_counts()",
"_____no_output_____"
],
[
"((data.loc[data['currency']=='ARS','price'])/(data.loc[data['currency']=='ARS','price_aprox_usd'])).round(decimals=4).value_counts()",
"_____no_output_____"
]
],
[
[
"vemos que cuando el aviso es ingresado con currency==ARS están presentes en el dataset dos tipos de cambio :\n\nuno para convertir ARS a USD = TC Comprador\nOtro para convertir USD a ARS = TC Vendedor\n\nDefinimos las variables",
"_____no_output_____"
]
],
[
[
"TC_comprador=((data.loc[data['currency']=='ARS','price_aprox_local_currency'])/(data.loc[data['currency']=='ARS','price_aprox_usd'])).round(decimals=4).value_counts()\nTC_comprador",
"_____no_output_____"
],
[
"TC_vendedor=((data.loc[data['currency']=='ARS','price'])/(data.loc[data['currency']=='ARS','price_aprox_usd'])).round(decimals=4).mean()\nTC_vendedor",
"_____no_output_____"
]
],
[
[
"#### Valores con currency USD",
"_____no_output_____"
]
],
[
[
"((data.loc[data['currency']=='USD','price'])==(data.loc[data['currency']=='USD','price_aprox_usd'])).value_counts()\n",
"_____no_output_____"
],
[
"((data.loc[data['currency']=='USD','price_aprox_local_currency'])/(data.loc[data['currency']=='USD','price_aprox_usd'])).round(decimals=4).value_counts()",
"_____no_output_____"
]
],
[
[
"Para los valores ingresados en dólares 'price' coincide con 'price_aprox_usd'\nEl tipo de cambio entre 'price_aprox_local_currency' (moneda local, ARS) y el dolar es el ya visto como \"TC_comprador\"",
"_____no_output_____"
],
[
"Hacemos ahora un filtro del registro ingresado con moneda uruguaya (UYU), calculando el tipo de cambio original (columna \"price\") y revisamos además el correspondiente a la moneda americana",
"_____no_output_____"
]
],
[
[
"data['currency']==\"UYU\"\ndata_UYU= data.loc[data['currency']==\"UYU\", [\"price\", \"price_aprox_local_currency\",\"price_aprox_usd\",\"place_with_parent_names\"]]\ndata_UYU\n\ndata_UYU_TC=data_UYU[\"price\"]/data_UYU['price_aprox_usd']\ndata_USD_TC=data_UYU[\"price_aprox_local_currency\"]/data_UYU['price_aprox_usd']\nprint(data_UYU)\nprint(data_UYU_TC)\nprint(data_USD_TC)",
" price price_aprox_local_currency price_aprox_usd \\\n116625 650000000.0 4.054773e+08 22980378.29 \n\n place_with_parent_names \n116625 |Argentina|Mendoza|San Roque| \n116625 28.285\ndtype: float64\n116625 17.6445\ndtype: float64\n"
]
],
[
[
"En este caso vemos que se trata de una propiedad vendida en territorio argentino (provincia de Mendoza), que se ingresó con 3 cotizaciones diferentes. Vamos a depurar la fila, dejando los precios establecidos en ARS ya que Price no coincide con Price_aprox_usd y utiliza TC_comprador",
"_____no_output_____"
]
],
[
[
"data.loc[data['currency']==\"UYU\",\"price\"]=data.loc[data['currency']==\"UYU\",\"price_aprox_usd\"]*TC_vendedor",
"_____no_output_____"
],
[
"data['currency']==\"UYU\"\ndata_UYU= data.loc[data['currency']==\"UYU\", [\"price\", \"price_aprox_local_currency\",\"price_aprox_usd\",\"place_with_parent_names\"]]\ndata_UYU\n\ndata_UYU_TC=data_UYU[\"price\"]/data_UYU['price_aprox_usd']\ndata_USD_TC=data_UYU[\"price_aprox_local_currency\"]/data_UYU['price_aprox_usd']\nprint(data_UYU)\nprint(data_UYU_TC)\nprint(data_USD_TC)\nprint(data_UYU.index)",
" price price_aprox_local_currency price_aprox_usd \\\n116625 4.093825e+08 4.054773e+08 22980378.29 \n\n place_with_parent_names \n116625 |Argentina|Mendoza|San Roque| \n116625 17.814437\ndtype: float64\n116625 17.6445\ndtype: float64\nInt64Index([116625], dtype='int64')\n"
]
],
[
[
"resta sólo cambiar la moneda",
"_____no_output_____"
]
],
[
[
"data.loc[data['currency']=='UYU', 'currency']=\"ARS\"",
"_____no_output_____"
]
],
[
[
"chequeamos que el cambio se haya efectuado correctamente",
"_____no_output_____"
]
],
[
[
"data.iloc[data_UYU.index,:]",
"_____no_output_____"
]
],
[
[
"Hacemos el mismo procedimiento con las propiedades cotizadas con moneda peruana (\"PEN\")",
"_____no_output_____"
]
],
[
[
"#data['currency']==\"PEN\"\ndata_PEN=data.loc[data['currency']==\"PEN\", [\"price\", \"price_aprox_local_currency\",\"price_aprox_usd\",\"country_name\",\"place_with_parent_names\"]]\n\ndata_PEN_TC=data_PEN[\"price\"]/data_PEN['price_aprox_usd']\ndata_USD_TC=data_PEN[\"price_aprox_local_currency\"]/data_PEN['price_aprox_usd']\n\nprint(data_PEN)\nprint(data_PEN_TC)\nprint(data_USD_TC)",
" price price_aprox_local_currency price_aprox_usd country_name \\\n86677 380000.0 2066864.90 117139.33 Argentina \n86678 950000.0 5167162.35 292848.33 Argentina \n\n place_with_parent_names \n86677 |Argentina|Capital Federal| \n86678 |Argentina|Capital Federal| \n86677 3.244\n86678 3.244\ndtype: float64\n86677 17.6445\n86678 17.6445\ndtype: float64\n"
]
],
[
[
"Al igual que en caso anterior, se tratn de propiedades vendidas en CABA, que cuentan con dos tipos de cambio diferentes. \nSe decide unificar a ARS",
"_____no_output_____"
]
],
[
[
"data.loc[data['currency']==\"PEN\",\"price\"]=data.loc[data['currency']==\"PEN\",\"price_aprox_usd\"]*TC_vendedor",
"_____no_output_____"
],
[
"data.iloc[data_PEN.index,:]",
"_____no_output_____"
],
[
"data.loc[data['currency']=='PEN', 'currency']=\"ARS\"",
"_____no_output_____"
]
],
[
[
"Verificamos que se hayan aplicado corréctamente los cambios",
"_____no_output_____"
]
],
[
[
"data.iloc[data_PEN.index,:][['price', \"currency\", \"price_aprox_local_currency\",\"price_aprox_usd\",\"country_name\",\"place_with_parent_names\"]]",
"_____no_output_____"
]
],
[
[
"## Filtrado y limpieza de OUTLIERS en las Columnas de Precios",
"_____no_output_____"
],
[
"## Columna \"price_per_m2\"",
"_____no_output_____"
]
],
[
[
"data.loc[(data['currency']=='ARS')&(data['price_per_m2'].notnull()),'price_per_m2'].describe()",
"_____no_output_____"
],
[
"price_m2_ARS=data.loc[(data['currency']=='ARS')&(data['price_per_m2'].notnull()),'price_per_m2']\n\nprint ('La mediana de precios por m2 en pesos es: {0:.2f}'.format(price_m2_ARS.median()),'(registros con currency ARS)')\nprint ('La media de de precios por m2 en pesos es: {0:.2f}'.format(price_m2_ARS.mean()),'(registros con currency ARS)')\nprint ('El desvio de precios por m2 en pesos es : {0:.2f}'.format(price_m2_ARS.std()),'(registros con currency ARS)')\n",
"La mediana de precios por m2 en pesos es: 30276.23 (registros con currency ARS)\nLa media de de precios por m2 en pesos es: 33507.36 (registros con currency ARS)\nEl desvio de precios por m2 en pesos es : 55825.37 (registros con currency ARS)\n"
],
[
"data.loc[(data['currency']=='USD')&(data['price_per_m2'].notnull()),'price_per_m2'].describe()",
"_____no_output_____"
],
[
"price_per_m2_USD=data.loc[(data['currency']=='USD')&(data['price_per_m2'].notnull()),'price_per_m2']\n\nprint ('La mediana de precios por m2 en dólares es: {0:.2f}'.format(price_m2_ARS.median()),'(registros con currency USD)')\nprint ('La media de de precios por m2 en dólares es: {0:.2f}'.format(price_m2_ARS.mean()),'(registros con currency USD)')\nprint ('El desvio de precios por m2 en dólares es: {0:.2f}'.format(price_m2_ARS.std()),'(registros con currency USD)')",
"La mediana de precios por m2 en dólares es: 30276.23 (registros con currency USD)\nLa media de de precios por m2 en dólares es: 33507.36 (registros con currency USD)\nEl desvio de precios por m2 en dólares es: 55825.37 (registros con currency USD)\n"
]
],
[
[
"## Estado Inicial de Outliers",
"_____no_output_____"
]
],
[
[
"plt.figure(figsize=(8,8))\nplt.grid(True) \n\n\nax = sns.boxplot(x=\"currency\", y=\"price_per_m2\", data=data, palette=\"Set1\")\nplt.title('price_per_m2 / Outliers Estado Inicial')\nplt.xlabel('Currency')\nplt.ylabel('Precio por M2');\nplt.savefig('../Images/Price_m2_before.png')",
"_____no_output_____"
]
],
[
[
"## Cálculo de outliers con currency=ARS",
"_____no_output_____"
]
],
[
[
"min_price_ARS_per_m2 = 1800\niqr_values_ARS=calculate_iqr_values(data.loc[(data['price_per_m2'].notnull())&(data['currency']=='ARS'),'price_per_m2'])\nprint(data.loc[(data['currency']=='ARS')&(data['price_per_m2'].notnull()),'price_per_m2'].apply(check_outliers,args=iqr_values_ARS).unique())\nprint(iqr_values_ARS)\ndata.loc[(data['currency']=='ARS')&((data['price_per_m2']>iqr_values_ARS[1])|(data['price_per_m2']<min_price_ARS_per_m2)),'price_per_m2']=np.nan",
"[31016.94915254 32666.66666667 32711.86440678 ... 4666.66666667\n 2444.44444444 8965.51724138]\n(-5769.608865110535, 66257.42891659836)\n"
]
],
[
[
"## Cálculo de outliers con currency=USD",
"_____no_output_____"
]
],
[
[
"min_price_USD_per_m2 = 100\niqr_values_USD=calculate_iqr_values(data.loc[(data['price_per_m2'].notnull())&(data['currency']=='USD'),'price_per_m2'])\nprint(data.loc[(data['currency']=='USD')&(data['price_per_m2'].notnull()),'price_per_m2'].apply(check_outliers,args=iqr_values_USD).unique())\nprint(iqr_values_USD)\ndata.loc[(data['currency']=='USD')&((data['price_per_m2']>iqr_values_ARS[1])|(data['price_per_m2']<min_price_USD_per_m2)),'price_per_m2']=np.nan",
"[1550. 2438.7755102 1892.85714286 ... 528.45528455 2016.28571429\n 2018.19047619]\n(-569.0284855002349, 4889.540837119539)\n"
]
],
[
[
"## Revisamos La Distribución Corregida de Valores",
"_____no_output_____"
]
],
[
[
"plt.figure(figsize=(8,8))\nplt.grid(True) \nax = sns.boxplot(x=\"currency\", y=\"price_per_m2\", data=data, palette=\"Set1\")\nplt.title('price_per_m2 Distribución ')\nplt.xlabel('Currency USD')\nplt.ylabel('Precio por M2');\nplt.savefig('../Images/Price_m2_after.png')",
"_____no_output_____"
]
],
[
[
"### Verificar Outliers para Columna \"price_usd_per_m2\"",
"_____no_output_____"
]
],
[
[
"plt.figure(figsize=(15,6))\n\nplt.grid(True) \nax=sns.boxplot(x='price_usd_per_m2', data=data, orient='h', palette=\"Set1\") \nplt.title('Distribución de Precios por m2 en USD')\nplt.xlabel('Valores')\nplt.ylabel('Precio por Metro2')\nplt.show()\nplt.savefig(\"../Images/Price_USD_m2-before.png\")\n",
"_____no_output_____"
],
[
"calculate_iqr_values(data.loc[data['price_usd_per_m2'].notnull(), \"price_usd_per_m2\"])",
"_____no_output_____"
],
[
"min_price2_USD_per_m2 = 150\niqr_values_price_USD=calculate_iqr_values(data.loc[data['price_usd_per_m2'].notnull(), \"price_usd_per_m2\"])\ndata.loc[(data['price_usd_per_m2']>iqr_values_price_USD[1])|(data['price_usd_per_m2']<min_price2_USD_per_m2),'price_usd_per_m2']=np.nan",
"_____no_output_____"
],
[
"plt.figure(figsize=(15,6))\n\nplt.grid(True) \nax=sns.boxplot(x='price_usd_per_m2', data=data, orient='h', palette=\"Set1\") \nplt.title('Distribución de Precios por m2 en USD')\nplt.xlabel('Valores')\nplt.ylabel('Precio por Metro2')\nplt.show()\n\nplt.savefig(\"../Images/Price_USD_m2-after.png\")",
"_____no_output_____"
]
],
[
[
"## Usamos regex para buscar los precios faltantes en las columnas \"description\" y \"title\"",
"_____no_output_____"
]
],
[
[
"def price_search(text):\n pattern = r'(?:USD|U\\$D|DOLL|DOL.|U\\$S|dolares|dólares|precio|precio total)\\s?(\\d+,?\\.?\\d*,?\\.?\\d*)'\n regex = re.compile(pattern, flags = re.IGNORECASE)\n price_lst = regex.findall(text)\n price_lst = [float(item.replace(',','').replace('.','')) for item in price_lst]\n if not price_lst:\n return np.nan\n if len(price_lst)>1:\n if max(price_lst)>35000:\n return max(price_lst)\n else:\n return np.nan\n if price_lst[0]>35000:\n return price_lst[0]\n else:\n return np.nan",
"_____no_output_____"
],
[
"data.loc[data['price_aprox_usd'].isna(),'description'].apply(price_search).notnull().sum()",
"_____no_output_____"
],
[
"data.loc[data['price_aprox_usd'].isna(),'title'].apply(price_search).notnull().sum()",
"_____no_output_____"
],
[
"def price_search2(text):\n pattern = r'(\\d+,?\\.?\\d*,?\\.?\\d*)\\s?(?:USD|U\\$D|DOLL|DOL.|U\\$S|dolares|dólares)+'\n regex = re.compile(pattern, flags = re.IGNORECASE)\n price_lst = regex.findall(text)\n price_lst = [float(item.replace(',','').replace('.','')) for item in price_lst]\n \n if not price_lst:\n return np.nan\n if len(price_lst)>1:\n if max(price_lst)>35000:\n return max(price_lst)\n else:\n return np.nan\n if price_lst[0]>35000:\n return price_lst[0]\n else:\n return np.nan",
"_____no_output_____"
]
],
[
[
"Aplicamos las dos funciones a las dos columnas ",
"_____no_output_____"
]
],
[
[
"data.iloc[:,10:18].isnull().sum()",
"_____no_output_____"
],
[
"data[\"price_aprox_usd\"].fillna(value=data.loc[data['price_aprox_usd'].isna(),'title'].apply(price_search), inplace=True)\ndata[\"price_aprox_usd\"].fillna(value=data.loc[data['price_aprox_usd'].isna(),'title'].apply(price_search2), inplace=True)\ndata[\"price_aprox_usd\"].fillna(value=data.loc[data['price_aprox_usd'].isna(),'description'].apply(price_search), inplace=True)\ndata[\"price_aprox_usd\"].fillna(value=data.loc[data['price_aprox_usd'].isna(),'description'].apply(price_search2), inplace=True)",
"_____no_output_____"
],
[
"data.iloc[:,10:18].isnull().sum()",
"_____no_output_____"
],
[
"#REVISAR SI ESTA FILA DESAPARECE ANTES\n\n(data.loc[(data['price_aprox_local_currency'].notnull())&(data['currency'].isna())&(data['price'].notnull()),['price','price_aprox_usd','price_aprox_local_currency','currency']])",
"_____no_output_____"
]
],
[
[
"## Completando los valores expresados en pesos",
"_____no_output_____"
]
],
[
[
"def price_search_pesos(text):\n pattern = r'(?:\\$)\\s?(\\d+,?\\.?\\d*,?\\.?\\d*)'\n regex = re.compile(pattern, flags = re.IGNORECASE)\n price_lst = regex.findall(text)\n price_lst = [float(item.replace(',','').replace('.','')) for item in price_lst]\n if not price_lst:\n return np.nan\n if len(price_lst)>1:\n if max(price_lst)>20000:\n return max(price_lst)\n else:\n return np.nan\n \n if len(price_lst)==1:\n if price_lst[0]>35000:\n return price_lst[0]\n \n else:\n return np.nan\n",
"_____no_output_____"
],
[
"data.iloc[:,10:18].isnull().sum()",
"_____no_output_____"
],
[
"data[\"price_aprox_local_currency\"].fillna(value=data.loc[data['price_aprox_local_currency'].isna(),'title'].apply(price_search_pesos), inplace=True)\ndata[\"price_aprox_local_currency\"].fillna(value=data.loc[data['price_aprox_local_currency'].isna(),'description'].apply(price_search_pesos), inplace=True)\n",
"_____no_output_____"
],
[
"data.iloc[:,10:18].isnull().sum()",
"_____no_output_____"
]
],
[
[
"# CÁLCULO DE VALORES FALTANTES CORRESPONDIENTES A VARIABLES ALEATORIAS LINEALMENTE DEPENDIENTES",
"_____no_output_____"
],
[
"En el paso anterior se vio, y se hizo una reducción de los tipos de cambio presentes en el dataset. En esa instancia se decidió\ntrabajar sólo con moneda local (pesos argentinos) y dólar como única divisa.\nA continuación, vamos a analizar las variables aleatorias presentes en el dataset, en primer lugar, el peso que se le dá a los metros cubiertos y descubiertos en el precio de la propiedad\n",
"_____no_output_____"
]
],
[
[
"data.iloc[:,10:18].isnull().sum()",
"_____no_output_____"
]
],
[
[
"#### Variables Aleatorias Linealmente Dependientes",
"_____no_output_____"
],
[
"Las siguientes columnas, están formadas por variables que son LD unas de otras. Por lo cuál, vamos a recuperar valores\nrealizando operaciones algebraicas entre ellas\n\nLos datos recuperados en 'price' se calcularán directamente en dólares",
"_____no_output_____"
]
],
[
[
"data.iloc[1:5,:][['price', 'currency', 'price_aprox_local_currency','price_aprox_usd','surface_total_in_m2', 'price_usd_per_m2', \"price_per_m2\"]]",
"_____no_output_____"
]
],
[
[
"Partimos con estos datos faltantes",
"_____no_output_____"
]
],
[
[
"data.iloc[:,10:18].isnull().sum()",
"_____no_output_____"
]
],
[
[
"#### Operaciones con Tipo de Cambio",
"_____no_output_____"
],
[
"Sabemos que price=price_aprox_usd cuando \"currency\"==DOL\ny que con ese ingreso la relación entre \"price\" y \"price_aprox_local_currency\" es el \"TC_comprador\" ya calculado",
"_____no_output_____"
]
],
[
[
"price_new=data.loc[data[\"price_aprox_usd\"].notnull(), \"price_aprox_usd\"]\n\ndata[\"price\"].fillna(value=price_new, inplace=True) ",
"_____no_output_____"
],
[
"data[\"price_aprox_usd\"].fillna(value=data.loc[data[\"price\"].notnull(), \"price\"], inplace=True) \n",
"_____no_output_____"
],
[
"TC_comprador=((data.loc[data['currency']=='ARS','price_aprox_local_currency'])/(data.loc[data['currency']=='ARS','price_aprox_usd'])).round(decimals=4)\nTC_comprador.value_counts()",
"_____no_output_____"
],
[
"price_aprox_local_currency_new =data[\"price\"]*17.6445\ndata[\"price_aprox_local_currency\"].fillna(value=price_aprox_local_currency_new, inplace=True) \n",
"_____no_output_____"
]
],
[
[
"#### Precios por M2",
"_____no_output_____"
]
],
[
[
"price_usd_per_m2_new=data['price_aprox_usd']/data['surface_total_in_m2']\nprice_per_m2_new=data['price']/data['surface_covered_in_m2']\n\ndata[\"price_usd_per_m2\"].fillna(value=price_usd_per_m2_new, inplace=True) \ndata[\"price_per_m2\"].fillna(value=price_per_m2_new, inplace=True) \n\n",
"_____no_output_____"
]
],
[
[
"#### Cálculo de superficies",
"_____no_output_____"
],
[
"Calculamos la superficie a partir de los precios en ambas monedas y el correspondiente por m2",
"_____no_output_____"
]
],
[
[
"surface_total_in_m2_new= data[\"price_aprox_usd\"]/data['price_usd_per_m2']\nsurface_covered_in_m2_new= data[\"price\"]/data['price_per_m2']\n \ndata[\"surface_total_in_m2\"].fillna(value=surface_total_in_m2_new, inplace=True)\ndata[\"surface_covered_in_m2\"].fillna(value=surface_covered_in_m2_new, inplace=True)\n\n# for tipo,limites_tot,limites_cub in zip(property_type,sup_tot_limits,sup_cub_limits):\n# for i in range(len(limites_tot)): \n# data.loc[(data['property_type']==str(tipo))&(data['surface_total_in_m2']>limites_tot[i][1]),'surface_total_in_m2']=np.nan\n# data.loc[(data['property_type']==str(tipo))&(data['surface_covered_in_m2']>limites_cub[i][1]),'surface_covered_in_m2']=np.nan\n# data.loc[(data['property_type']==str(tipo))&(data['surface_total_in_m2']<limites_tot[i][0]),'surface_total_in_m2']=np.nan\n# data.loc[(data['property_type']==str(tipo))&(data['surface_covered_in_m2']<limites_cub[i][0]),'surface_covered_in_m2']=np.nan",
"_____no_output_____"
]
],
[
[
"#### Cálculo de los precios (Columnas \"price\" y \"price_aprox_usd\" a partir de los correspondientes precios por m2 ",
"_____no_output_____"
]
],
[
[
"price_aprox_usd_new=data['price_usd_per_m2']*data[\"surface_total_in_m2\"]\ndata[\"price_aprox_usd\"].fillna(value=price_aprox_usd_new, inplace=True)",
"_____no_output_____"
],
[
"price_new=data['price_per_m2']*data[\"surface_covered_in_m2\"]\nvalues_price = {'price': price_new, 'currency': 'USD'}\ndata[\"price\"].fillna(value=values_price, inplace=True)",
"_____no_output_____"
]
],
[
[
"#### Cálculo del precio por m2 en pesos a partir del precio y superficie (operación inversa)",
"_____no_output_____"
]
],
[
[
"price_usd_per_m2_new= data[\"price_aprox_usd\"]/data['surface_total_in_m2']\nprice_per_m2_new= data[\"price\"]/data['surface_covered_in_m2']\n\ndata[\"price_usd_per_m2\"].fillna(value=price_usd_per_m2_new, inplace=True)\ndata[\"price_per_m2\"].fillna(value=price_per_m2_new, inplace=True) \n",
"_____no_output_____"
]
],
[
[
"# Columna 'expenses'\n\nLa columna de expensas tenía más de 100.000 datos faltantes. Se intentó hacer una búsqueda de esos datos, pero sin mucho éxito (no se pudieron recuperar los suficientes). Además, tomando en cuenta que para el modelo de regresión no tiene valor, se decidió dropear toda esta columna.",
"_____no_output_____"
],
[
"## Limpieza Final\n\nEliminar columnas que no servirán para el siguiente desafío (regresión).\nEliminar datos nulos.\nGuardar en un csv los datos limpios.",
"_____no_output_____"
]
],
[
[
"data.columns",
"_____no_output_____"
],
[
"data2 = data.copy()\n\ndata3 = data2.drop(['geonames_id','lat-lon','lat','lon','expenses','image_thumbnail','construction_index','title','description','properati_url'],axis=1).dropna().copy()\n\ndata3.reset_index(inplace=True)\n\ndata3.to_csv('../Data/properati_clean.csv')\n\ndata3.head()",
"_____no_output_____"
],
[
"data3.shape",
"_____no_output_____"
],
[
"data3.operation.value_counts()",
"_____no_output_____"
],
[
"data3.currency.value_counts()",
"_____no_output_____"
],
[
"data3.country_name.value_counts()",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab5b8ebb7db20144eb60124bd5de776885c31ca
| 11,651 |
ipynb
|
Jupyter Notebook
|
01_Python_Basics.ipynb
|
dhavidshajan/Introdution-To-Python
|
873da6b80798bfef3c12c07cba750c1a1794b590
|
[
"MIT"
] | 11 |
2022-03-09T10:04:52.000Z
|
2022-03-09T10:17:43.000Z
|
01_Python_Basics.ipynb
|
dhavidshajan/Introdution-To-Python
|
873da6b80798bfef3c12c07cba750c1a1794b590
|
[
"MIT"
] | null | null | null |
01_Python_Basics.ipynb
|
dhavidshajan/Introdution-To-Python
|
873da6b80798bfef3c12c07cba750c1a1794b590
|
[
"MIT"
] | null | null | null | 20.621239 | 67 | 0.371985 |
[
[
[
"10+20\n",
"_____no_output_____"
],
[
"30*40",
"_____no_output_____"
],
[
"15/10",
"_____no_output_____"
],
[
"20-30",
"_____no_output_____"
],
[
"x=10",
"_____no_output_____"
],
[
"x",
"_____no_output_____"
],
[
"x=16",
"_____no_output_____"
],
[
"x",
"_____no_output_____"
],
[
"x=10\ny=40\nx+y",
"_____no_output_____"
],
[
"x-y",
"_____no_output_____"
],
[
"x*y",
"_____no_output_____"
],
[
"x/y",
"_____no_output_____"
],
[
"x=90\ny=70\nz=100\nx+y-z",
"_____no_output_____"
],
[
"p=10000\nn=5\nr=9\n",
"_____no_output_____"
],
[
"i=p*n*r/100",
"_____no_output_____"
],
[
"i\n",
"_____no_output_____"
],
[
"p+i",
"_____no_output_____"
],
[
"x='india'",
"_____no_output_____"
],
[
"x",
"_____no_output_____"
],
[
"x=10\ny=10.5\nz='india'",
"_____no_output_____"
],
[
"type(x)",
"_____no_output_____"
],
[
"type(y)",
"_____no_output_____"
],
[
"type(z)",
"_____no_output_____"
],
[
"x=input('enter a name')\nprint(x)",
"enter a nameDhavid\nDhavid\n"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab5b9c26dcb5731c471ae610afc9a70593c9808
| 448,029 |
ipynb
|
Jupyter Notebook
|
labs/respuestas_lab_11.ipynb
|
Nico-Rivera/mat281_portfolio
|
0f6d0626e683f0aa8bbd963ef04de7f79e196549
|
[
"MIT"
] | null | null | null |
labs/respuestas_lab_11.ipynb
|
Nico-Rivera/mat281_portfolio
|
0f6d0626e683f0aa8bbd963ef04de7f79e196549
|
[
"MIT"
] | null | null | null |
labs/respuestas_lab_11.ipynb
|
Nico-Rivera/mat281_portfolio
|
0f6d0626e683f0aa8bbd963ef04de7f79e196549
|
[
"MIT"
] | null | null | null | 334.100671 | 94,052 | 0.918554 |
[
[
[
"# MAT281 - Laboratorio N°11\n\n<a id='p1'></a>\n## I.- Problema 01\n\nLista de actos delictivos registrados por el Service de police de la Ville de Montréal (SPVM).\n\n\n<img src=\"http://henriquecapriles.com/wp-content/uploads/2017/02/femina_detenida-1080x675.jpg\" width=\"480\" height=\"360\" align=\"center\"/>\n\nEl conjunto de datos en estudio `interventionscitoyendo.csv` corresponde a todos los delitos entre 2015 y agosto de 2020en Montreal. Cada delito está asociado en grandes categorías, y hay información sobre la ubicación, el momento del día, etc.\n\n> **Nota**: Para más información seguir el siguiente el [link](https://donnees.montreal.ca/ville-de-montreal/actes-criminels).",
"_____no_output_____"
]
],
[
[
"# librerias \n\nimport os\nimport pandas as pd\nimport numpy as np \nimport matplotlib.pyplot as plt \nimport seaborn as sns\nimport random\nfrom matplotlib.colors import rgb_to_hsv\nfrom statsmodels.tsa.statespace.sarimax import SARIMAX\nfrom metrics_regression import *\n\n\n# graficos incrustados\nplt.style.use('fivethirtyeight')\n%matplotlib inline\n\n# parametros esteticos de seaborn\nsns.set_palette(\"deep\", desat=.6)\nsns.set_context(rc={\"figure.figsize\": (12, 4)})",
"_____no_output_____"
],
[
"# read data\n\nvalidate_categorie = [\n 'Introduction', 'Méfait','Vol dans / sur véhicule à moteur', 'Vol de véhicule à moteur',\n]\n\ndf = pd.read_csv(os.path.join(\"data\",\"interventionscitoyendo.csv\"), sep=\",\", encoding='latin-1')\ndf.columns = df.columns.str.lower()\ndf['date_str'] = df['date'].apply(lambda x: x[:7])\ndf['date'] = pd.to_datetime(df['date'], format='%Y-%m-%d')\n\ndf = df.loc[lambda x: x['categorie'].isin(validate_categorie)]\ndf.head()",
"_____no_output_____"
]
],
[
[
"Como tenemos muchos datos por categoría a nivel de día, agruparemos a nivel de **semanas** y crearemos un nuevo conjuntos de datos llamados `df_week`.",
"_____no_output_____"
]
],
[
[
"df['weekofyear'] = df['date'].dt.week\ndf['year'] = df['date'].dt.year\n\ndf_week = df.groupby(['categorie','weekofyear','year'])['pdq'].sum().reset_index()\ndates = df_week.year*100+df_week.weekofyear\ndf_week['date'] = pd.to_datetime(dates.astype(str) + '0', format='%Y%W%w')\ndf_week.head()",
"<ipython-input-3-171270f4f4f6>:1: FutureWarning: Series.dt.weekofyear and Series.dt.week have been deprecated. Please use Series.dt.isocalendar().week instead.\n df['weekofyear'] = df['date'].dt.week\n"
]
],
[
[
"El objetivo de este laboratorio es poder realizar un análisis completo del conjunto de datos en estudio, para eso debe responder las siguientes preguntas:\n\n1. Para cada categoría grafique la serie temporal correspondiente (columna `categorie`).\n2. Elegir una categoría dentro de `validate_categorie`, luego cree el conjunto de datos llamado `df_categorie`.\n3. Realice un análisis exploratorio de la serie temporal escogida.\n4. Aplicar el modelo de pronóstico $SARIMA(p,d,q)x(P,D,Q,S)$, probando varias configuraciones de los hiperparámetros. Encuentre la mejor configuración. Concluya.\n5. Para el mejor modelo encontrado, verificar si el residuo corresponde a un ruido blanco.\n",
"_____no_output_____"
]
],
[
[
"for cat in validate_categorie:\n mask= lambda df: (df[\"categorie\"] == cat)\n plt.figure(figsize=(10, 6))\n sns.lineplot(\n x='date',\n y='pdq',\n hue='categorie',\n data=df_week[mask],\n ci = None\n )\n plt.show()\n",
"_____no_output_____"
],
[
"random.seed(666)\ncategorie = validate_categorie[random.randint(0,3)]",
"_____no_output_____"
],
[
"mask = lambda df: df[\"categorie\"] == categorie\ndf_categorie = df_week[mask]\ndf_categorie.head()",
"_____no_output_____"
],
[
"#Exploracion de datos\n\n\n# Resumen de la informacion\n\ndef resumen_por_columna(df,cols):\n pd_series = df[cols]\n \n # elementos distintos \n l_unique = pd_series.unique()\n \n # elementos vacios\n \n l_vacios = pd_series[pd_series.isna()]\n \n df_info = pd.DataFrame({\n 'columna': [cols],\n 'unicos': [len(l_unique)],\n 'vacios': [len(l_vacios)]\n })\n \n return df_info\n\n#exploracion de datos nulos\n\nframes = []\n\nfor col in df_categorie.columns:\n aux_df = resumen_por_columna(df_categorie,col)\n frames.append(aux_df)\n \ndf_info = pd.concat(frames).reset_index(drop=True)\ndf_info",
"_____no_output_____"
],
[
"#revisar la integridad de la columna year\n\ndf_categorie[\"year\"].unique()",
"_____no_output_____"
],
[
"#eliminar datos donde el año no coincide con la fecha\n\n\nmask = lambda df: df[\"year\"] == df[\"date\"].dt.year\ndf_categorie = df_categorie[mask]\ndf_categorie",
"_____no_output_____"
],
[
"#modelo de pronostico\n\ndf_y = df_categorie[[\"pdq\",\"date\"]].set_index('date').resample('M').mean()\n\ntarget_date = \"2021-01-01\"\n\nmask_ds = df_y.index < target_date\n\ny_train = df_y[mask_ds]\ny_test = df_y[~mask_ds]\n\n#plotting the data\ny_train[\"pdq\"].plot()\ny_test[\"pdq\"].plot()\nplt.show()",
"_____no_output_____"
],
[
"#definir clase para automatizar el proceso\n\nclass SarimaModels:\n def __init__(self,params):\n\n self.params = params\n \n \n @property\n def name_model(self):\n return f\"SARIMA_{self.params[0]}X{self.params[1]}\".replace(' ','')\n \n @staticmethod\n def test_train_model(y,date):\n mask_ds = y.index < date\n\n y_train = y[mask_ds]\n y_test = y[~mask_ds] \n \n return y_train, y_test\n \n def fit_model(self,y,date):\n y_train, y_test = self.test_train_model(y,date )\n model = SARIMAX(y_train,\n order=self.params[0],\n seasonal_order=self.params[1],\n enforce_stationarity=False,\n enforce_invertibility=False)\n \n model_fit = model.fit(disp=0)\n\n return model_fit\n \n def df_testig(self,y,date):\n y_train, y_test = self.test_train_model(y,date )\n model = SARIMAX(y_train,\n order=self.params[0],\n seasonal_order=self.params[1],\n enforce_stationarity=False,\n enforce_invertibility=False)\n \n model_fit = model.fit(disp=0)\n \n start_index = y_test.index.min()\n end_index = y_test.index.max()\n\n preds = model_fit.get_prediction(start=start_index,end=end_index, dynamic=False)\n df_temp = pd.DataFrame(\n {\n 'y':y_test['pdq'],\n 'yhat': preds.predicted_mean\n }\n )\n \n return df_temp\n \n def metrics(self,y,date):\n df_temp = self.df_testig(y,date)\n df_metrics = summary_metrics(df_temp)\n df_metrics['model'] = self.name_model\n \n return df_metrics",
"_____no_output_____"
],
[
"# definir parametros \n\nimport itertools\n\np = d = q = range(0, 2)\npdq = list(itertools.product(p, d, q))\nseasonal_pdq = [(x[0], x[1], x[2], 12) for x in list(itertools.product(p, d, q))]\n\nparams = list(itertools.product(pdq,seasonal_pdq))\ntarget_date = '2021-01-01'",
"_____no_output_____"
],
[
"# iterar para los distintos escenarios\n\nframes = []\nfor param in params:\n try:\n sarima_model = SarimaModels(param)\n df_metrics = sarima_model.metrics(df_y,target_date)\n frames.append(df_metrics)\n\n except:\n pass",
"_____no_output_____"
],
[
"# juntar resultados de las métricas y comparar\ndf_metrics_result = pd.concat(frames)\ndf_metrics_result.sort_values(['mae','mape'])",
"_____no_output_____"
],
[
"# ajustar mejor modelo\n\nparam = [(0,1,0),(1,0,0,12)]\nsarima_model = SarimaModels(param)\nmodel_fit = sarima_model.fit_model(df_y,target_date)\nbest_model = sarima_model.df_testig(df_y,target_date)\nbest_model.head()",
"_____no_output_____"
],
[
"# graficar mejor modelo\n\npreds = best_model['yhat']\nax = df_y['2015':].plot(label='observed')\npreds.plot(ax=ax, label='Forecast', alpha=.7, figsize=(14, 7))\nax.set_xlabel('Date')\nax.set_ylabel('pdq')\nplt.legend()\nplt.show()",
"_____no_output_____"
],
[
"# resultados del error \nmodel_fit.plot_diagnostics(figsize=(16, 8))\nplt.show()",
"_____no_output_____"
]
],
[
[
"Segun los gra",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
]
] |
4ab5c55fa9f50600599eece01c95e4480060d389
| 26,882 |
ipynb
|
Jupyter Notebook
|
6 aerial cactus/solving-aerial-cactus-challenge-using-fast-ai.ipynb
|
MLVPRASAD/KaggleProjects
|
379e062cf58d83ff57a456552bb956df68381fdd
|
[
"MIT"
] | 2 |
2020-01-25T08:31:14.000Z
|
2022-03-23T18:24:03.000Z
|
6 aerial cactus/solving-aerial-cactus-challenge-using-fast-ai.ipynb
|
MLVPRASAD/KaggleProjects
|
379e062cf58d83ff57a456552bb956df68381fdd
|
[
"MIT"
] | null | null | null |
6 aerial cactus/solving-aerial-cactus-challenge-using-fast-ai.ipynb
|
MLVPRASAD/KaggleProjects
|
379e062cf58d83ff57a456552bb956df68381fdd
|
[
"MIT"
] | null | null | null | 56.593684 | 14,636 | 0.733167 |
[
[
[
"**About this challenge**\n\nTo assess the impact of climate change on Earth's flora and fauna, it is vital to quantify how human activities such as logging, mining, and agriculture are impacting our protected natural areas. Researchers in Mexico have created the VIGIA project, which aims to build a system for autonomous surveillance of protected areas. A first step in such an effort is the ability to recognize the vegetation inside the protected areas. In this competition, you are tasked with creation of an algorithm that can identify a specific type of cactus in aerial imagery.\n\nIn this kernel we will be trying to solve this challenge using CNN through **fast.ai library**\n\n",
"_____no_output_____"
],
[
"**Loading necessary libraries**",
"_____no_output_____"
]
],
[
[
"from fastai.vision import *\nfrom fastai import *\nimport os\nimport pandas as pd\nimport numpy as np\nprint(os.listdir(\"../input/\"))",
"['test', 'train', 'train.csv', 'sample_submission.csv']\n"
],
[
"train_dir=\"../input/train/train\"\ntest_dir=\"../input/test/test\"\ntrain = pd.read_csv('../input/train.csv')\ntest = pd.read_csv(\"../input/sample_submission.csv\")\ndata_folder = Path(\"../input\")\n",
"_____no_output_____"
]
],
[
[
"**Analysing the given data**",
"_____no_output_____"
]
],
[
[
"train.head(5)\n",
"_____no_output_____"
],
[
"train.describe()",
"_____no_output_____"
]
],
[
[
"**Getting the Data. **\n[reference](https://docs.fast.ai/vision.data.html)",
"_____no_output_____"
]
],
[
[
"test_img = ImageList.from_df(test, path=data_folder/'test', folder='test')\n# Applying Data augmentation\ntrfm = get_transforms(do_flip=True, flip_vert=True, max_rotate=10.0, max_zoom=1.1, max_lighting=0.2, max_warp=0.2, p_affine=0.75, p_lighting=0.75)\ntrain_img = (ImageList.from_df(train, path=data_folder/'train', folder='train')\n .split_by_rand_pct(0.01)\n .label_from_df()\n .add_test(test_img)\n .transform(trfm, size=128)\n .databunch(path='.', bs=64, device= torch.device('cuda:0'))\n .normalize(imagenet_stats)\n )",
"_____no_output_____"
]
],
[
[
"**Training the data using appropriate model. We have used [densenet](https://pytorch.org/docs/stable/torchvision/models.html) here**",
"_____no_output_____"
]
],
[
[
"learn = cnn_learner(train_img, models.densenet161, metrics=[error_rate, accuracy])\n",
"Downloading: \"https://download.pytorch.org/models/densenet161-8d451a50.pth\" to /tmp/.torch/models/densenet161-8d451a50.pth\n115730790it [00:01, 99358993.39it/s]\n"
]
],
[
[
"**Finding the suitable learning rate**",
"_____no_output_____"
]
],
[
[
"learn.lr_find()\n\n",
"_____no_output_____"
]
],
[
[
"**Plotting the Learning Rate**",
"_____no_output_____"
]
],
[
[
"learn.recorder.plot()",
"_____no_output_____"
]
],
[
[
"**Now training the data based on suitable learning rate**",
"_____no_output_____"
]
],
[
[
"lr = 1e-02\nlearn.fit_one_cycle(3, slice(lr))\n\n",
"_____no_output_____"
],
[
"preds,_ = learn.get_preds(ds_type=DatasetType.Test)",
"_____no_output_____"
],
[
"test.has_cactus = preds.numpy()[:, 0]",
"_____no_output_____"
],
[
"test.to_csv('submission.csv', index=False)",
"_____no_output_____"
]
],
[
[
"**References**\n* https://docs.fast.ai/\n* https://www.kaggle.com/kenseitrg/simple-fastai-exercise\n* https://www.kaggle.com/shahules/getting-started-with-cnn-and-vgg16\n",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
]
] |
4ab5d9f0d7c5f9f3c5d264f8ad15bd40ff493fcc
| 21,697 |
ipynb
|
Jupyter Notebook
|
Exercises/04_Numpy_Exercises.ipynb
|
abhisheksethi02/MicroHackML
|
dd2f740e9b3d05ee7429de050453502beb3d7473
|
[
"MIT"
] | null | null | null |
Exercises/04_Numpy_Exercises.ipynb
|
abhisheksethi02/MicroHackML
|
dd2f740e9b3d05ee7429de050453502beb3d7473
|
[
"MIT"
] | null | null | null |
Exercises/04_Numpy_Exercises.ipynb
|
abhisheksethi02/MicroHackML
|
dd2f740e9b3d05ee7429de050453502beb3d7473
|
[
"MIT"
] | 2 |
2020-09-20T01:03:37.000Z
|
2020-09-21T07:10:34.000Z
| 24.16147 | 170 | 0.388902 |
[
[
[
"___\n\n<a href='http://www.pieriandata.com'> <img src='../Pierian_Data_Logo.png' /></a>\n___",
"_____no_output_____"
],
[
"# NumPy Exercises \n\nNow that we've learned about NumPy let's test your knowledge. We'll start off with a few simple tasks, and then you'll be asked some more complicated questions.",
"_____no_output_____"
],
[
"#### Import NumPy as np",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
],
[
[
"#### Create an array of 10 zeros ",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
],
[
[
"#### Create an array of 10 ones",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
],
[
[
"#### Create an array of 10 fives",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
],
[
[
"#### Create an array of the integers from 10 to 50",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
],
[
[
"#### Create an array of all the even integers from 10 to 50",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
],
[
[
"#### Create a 3x3 matrix with values ranging from 0 to 8",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
],
[
[
"#### Create a 3x3 identity matrix",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
],
[
[
"#### Use NumPy to generate a random number between 0 and 1",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
],
[
[
"#### Use NumPy to generate an array of 25 random numbers sampled from a standard normal distribution",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
],
[
[
"#### Create the following matrix:",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
],
[
[
"#### Create an array of 20 linearly spaced points between 0 and 1:",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
],
[
[
"## Numpy Indexing and Selection\n\nNow you will be given a few matrices, and be asked to replicate the resulting matrix outputs:",
"_____no_output_____"
]
],
[
[
"mat = np.arange(1,26).reshape(5,5)\nmat",
"_____no_output_____"
],
[
"# WRITE CODE HERE THAT REPRODUCES THE OUTPUT OF THE CELL BELOW\n# BE CAREFUL NOT TO RUN THE CELL BELOW, OTHERWISE YOU WON'T\n# BE ABLE TO SEE THE OUTPUT ANY MORE",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"# WRITE CODE HERE THAT REPRODUCES THE OUTPUT OF THE CELL BELOW\n# BE CAREFUL NOT TO RUN THE CELL BELOW, OTHERWISE YOU WON'T\n# BE ABLE TO SEE THE OUTPUT ANY MORE",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"# WRITE CODE HERE THAT REPRODUCES THE OUTPUT OF THE CELL BELOW\n# BE CAREFUL NOT TO RUN THE CELL BELOW, OTHERWISE YOU WON'T\n# BE ABLE TO SEE THE OUTPUT ANY MORE",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"# WRITE CODE HERE THAT REPRODUCES THE OUTPUT OF THE CELL BELOW\n# BE CAREFUL NOT TO RUN THE CELL BELOW, OTHERWISE YOU WON'T\n# BE ABLE TO SEE THE OUTPUT ANY MORE",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"# WRITE CODE HERE THAT REPRODUCES THE OUTPUT OF THE CELL BELOW\n# BE CAREFUL NOT TO RUN THE CELL BELOW, OTHERWISE YOU WON'T\n# BE ABLE TO SEE THE OUTPUT ANY MORE",
"_____no_output_____"
],
[
"",
"_____no_output_____"
]
],
[
[
"### Now do the following",
"_____no_output_____"
],
[
"#### Get the sum of all the values in mat",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
],
[
[
"#### Get the standard deviation of the values in mat",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
],
[
[
"#### Get the sum of all the columns in mat",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
],
[
[
"# Great Job!",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
4ab5e351a421a40ed8cff258feef6574e196a56a
| 139,479 |
ipynb
|
Jupyter Notebook
|
tutorials/AxTutorials/Tune_a_CNN_on_MNIST.ipynb
|
sudhu26/data-science-portfolio
|
88f7a350cbd9245e4f92ff1829e49c5d378c609d
|
[
"MIT"
] | null | null | null |
tutorials/AxTutorials/Tune_a_CNN_on_MNIST.ipynb
|
sudhu26/data-science-portfolio
|
88f7a350cbd9245e4f92ff1829e49c5d378c609d
|
[
"MIT"
] | null | null | null |
tutorials/AxTutorials/Tune_a_CNN_on_MNIST.ipynb
|
sudhu26/data-science-portfolio
|
88f7a350cbd9245e4f92ff1829e49c5d378c609d
|
[
"MIT"
] | 1 |
2021-03-26T11:47:37.000Z
|
2021-03-26T11:47:37.000Z
| 158.679181 | 51,819 | 0.769521 |
[
[
[
"https://www.ax.dev/tutorials/",
"_____no_output_____"
],
[
"__Tune a CNN on MNIST__\n\n1. [Import](#Import)\n1. [Load MNIST data](#Load-MNIST-data)\n1. [Define a function to optimize](#Define-a-function-to-optimize)\n1. [Run the optimization loop](#Run-the-optimization-loop)\n1. [Plot response surface](#Plot-response-surface)\n1. [Plot best objective as function of the iteration](#Plot-best-objective-as-function-of-the-iteration)\n1. [Train CNN with best hyperparameters and evaluate on test set](#Train-CNN-with-best-hyperparameters-and-evaluate-on-test-set)\n\n",
"_____no_output_____"
],
[
"# Import",
"_____no_output_____"
],
[
"<a id = 'Import'></a>",
"_____no_output_____"
]
],
[
[
"import torch\nimport numpy as np\n\nfrom ax.plot.contour import plot_contour\nfrom ax.plot.trace import optimization_trace_single_method\nfrom ax.service.managed_loop import optimize\nfrom ax.utils.notebook.plotting import render, init_notebook_plotting\nfrom ax.utils.tutorials.cnn_utils import load_mnist, train, evaluate\n\ninit_notebook_plotting()",
"_____no_output_____"
]
],
[
[
"# Load MNIST data\n",
"_____no_output_____"
],
[
"<a id = 'Load-MNIST-data'></a>",
"_____no_output_____"
]
],
[
[
"#\ntrain_loader, valid_loader, test_loader = load_mnist()",
"_____no_output_____"
]
],
[
[
"# Define a function to optimize\n",
"_____no_output_____"
],
[
"<a id = 'Define-a-function-to-optimize'></a>",
"_____no_output_____"
]
],
[
[
"#\ndtype = torch.float\ndevice = torch.device(\"cuda\" if torch.cuda.is_available() else \"cpu\")\n\n\ndef train_evaluate(parameterization):\n net = train(\n train_loader=train_loader,\n parameters=parameterization,\n dtype=dtype,\n device=device,\n )\n return evaluate(net=net, data_loader=valid_loader, dtype=dtype, device=device)",
"_____no_output_____"
]
],
[
[
"# Run the optimization loop\n\n",
"_____no_output_____"
],
[
"<a id = 'Run-the-optimization-loop'></a>",
"_____no_output_____"
]
],
[
[
"#\nbest_parameters, values, experiment, model = optimize(\n parameters=[\n {\"name\": \"lr\", \"type\": \"range\", \"bounds\": [1e-6, 0.4], \"log_scale\": True},\n {\"name\": \"momentum\", \"type\": \"range\", \"bounds\": [0.0, 1.0]},\n ],\n evaluation_function=train_evaluate,\n objective_name=\"accuracy\",\n)",
"[INFO 05-14 04:40:09] ax.service.utils.dispatch: Using Bayesian Optimization generation strategy. Iterations after 5 will take longer to generate due to model-fitting.\n[INFO 05-14 04:40:09] ax.service.managed_loop: Started full optimization with 20 steps.\n[INFO 05-14 04:40:09] ax.service.managed_loop: Running optimization trial 1...\n[INFO 05-14 04:40:18] ax.service.managed_loop: Running optimization trial 2...\n[INFO 05-14 04:40:27] ax.service.managed_loop: Running optimization trial 3...\n[INFO 05-14 04:40:35] ax.service.managed_loop: Running optimization trial 4...\n[INFO 05-14 04:40:44] ax.service.managed_loop: Running optimization trial 5...\n[INFO 05-14 04:40:53] ax.service.managed_loop: Running optimization trial 6...\n[INFO 05-14 04:41:02] ax.service.managed_loop: Running optimization trial 7...\n[INFO 05-14 04:41:12] ax.service.managed_loop: Running optimization trial 8...\n[INFO 05-14 04:41:22] ax.service.managed_loop: Running optimization trial 9...\n[INFO 05-14 04:41:32] ax.service.managed_loop: Running optimization trial 10...\n[INFO 05-14 04:41:43] ax.service.managed_loop: Running optimization trial 11...\n[INFO 05-14 04:41:53] ax.service.managed_loop: Running optimization trial 12...\n[INFO 05-14 04:42:03] ax.service.managed_loop: Running optimization trial 13...\n[INFO 05-14 04:42:13] ax.service.managed_loop: Running optimization trial 14...\n[INFO 05-14 04:42:23] ax.service.managed_loop: Running optimization trial 15...\n[INFO 05-14 04:42:33] ax.service.managed_loop: Running optimization trial 16...\n[INFO 05-14 04:42:42] ax.service.managed_loop: Running optimization trial 17...\n[INFO 05-14 04:42:52] ax.service.managed_loop: Running optimization trial 18...\n[INFO 05-14 04:43:03] ax.service.managed_loop: Running optimization trial 19...\n/usr/local/lib/python3.6/site-packages/gpytorch/utils/cholesky.py:42: RuntimeWarning:\n\nA not p.d., added jitter of 1e-08 to the diagonal\n\n[INFO 05-14 04:43:13] ax.service.managed_loop: Running optimization trial 20...\n"
],
[
"#\nbest_parameters",
"_____no_output_____"
],
[
"#\nmeans, covariances = values\nprint(means)\nprint(covariances)",
"{'accuracy': 0.9708328909297179}\n{'accuracy': {'accuracy': 1.0481821595415295e-08}}\n"
]
],
[
[
"# Plot response surface\n\n",
"_____no_output_____"
],
[
"<a id = 'Plot-response-surface'></a>",
"_____no_output_____"
]
],
[
[
"#\nrender(\n plot_contour(model=model, param_x=\"lr\", param_y=\"momentum\", metric_name=\"accuracy\")\n)",
"_____no_output_____"
]
],
[
[
"# Plot best objective as function of the iteration\n\n",
"_____no_output_____"
],
[
"<a id = 'Plot-best-objective-as-function-of-the-iteration'></a>",
"_____no_output_____"
]
],
[
[
"#\nbest_objectives = np.array(\n [[trial.objective_mean * 100 for trial in experiment.trials.values()]]\n)\nbest_objective_plot = optimization_trace_single_method(\n y=np.maximum.accumulate(best_objectives, axis=1),\n title=\"Model performance vs. # of iters\",\n ylabel=\"Classification accuracy %\",\n)\nrender(best_objective_plot)",
"_____no_output_____"
]
],
[
[
"# Train CNN with best hyperparameters and evaluate on test set\n\n",
"_____no_output_____"
],
[
"<a id = 'Train-CNN-with-best-hyperparameters-and-evaluate-on-test-set'></a>",
"_____no_output_____"
]
],
[
[
"#\ndata = experiment.fetch_data()\ndf = data.df\nbest_arm_name = df.arm_name[df[\"mean\"] == df[\"mean\"].max()].values[0]\nbest_arm = experiment.arms_by_name[best_arm_name]\nbest_arm",
"_____no_output_____"
],
[
"#\nnet = train(\n train_loader=train_loader,\n parameters=best_arm.parameters,\n dtype=dtype,\n device=device,\n)\ntest_accuracy = evaluate(net=net, data_loader=test_loader, dtype=dtype, device=device)",
"_____no_output_____"
],
[
"test_accuracy",
"_____no_output_____"
],
[
"#\nprint(\"Classification\")",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
4ab6067b0f1382148f068bec78e1d4bfac1d61b8
| 31,000 |
ipynb
|
Jupyter Notebook
|
sagemaker-pipelines/tabular/tuning-step/sagemaker-pipelines-tuning-step.ipynb
|
eugeneteoh/amazon-sagemaker-examples
|
15c006d367d27371a407706953e2e962fe3bbe48
|
[
"Apache-2.0"
] | 1 |
2022-01-29T15:06:57.000Z
|
2022-01-29T15:06:57.000Z
|
sagemaker-pipelines/tabular/tuning-step/sagemaker-pipelines-tuning-step.ipynb
|
eugeneteoh/amazon-sagemaker-examples
|
15c006d367d27371a407706953e2e962fe3bbe48
|
[
"Apache-2.0"
] | 2 |
2020-09-26T01:31:38.000Z
|
2020-10-07T22:23:56.000Z
|
sagemaker-pipelines/tabular/tuning-step/sagemaker-pipelines-tuning-step.ipynb
|
eugeneteoh/amazon-sagemaker-examples
|
15c006d367d27371a407706953e2e962fe3bbe48
|
[
"Apache-2.0"
] | null | null | null | 36.686391 | 439 | 0.574581 |
[
[
[
"#### SageMaker Pipelines Tuning Step\n\nThis notebook illustrates how a Hyperparameter Tuning Job can be run as a step in a SageMaker Pipeline. \n\nThe steps in this pipeline include -\n* Preprocessing the abalone dataset\n* Running a Hyperparameter Tuning job\n* Creating the 2 best models\n* Evaluating the performance of the top performing model of the HPO step\n* Registering the top model in the model registry using a conditional step based on evaluation metrics",
"_____no_output_____"
]
],
[
[
"import sys\n\n!{sys.executable} -m pip install \"sagemaker>=2.48.0\"",
"_____no_output_____"
],
[
"import os\n\nimport boto3\nimport sagemaker\n\nfrom sagemaker.estimator import Estimator\nfrom sagemaker.inputs import TrainingInput\n\nfrom sagemaker.processing import (\n ProcessingInput,\n ProcessingOutput,\n Processor,\n ScriptProcessor,\n)\n\nfrom sagemaker import Model\nfrom sagemaker.xgboost import XGBoostPredictor\nfrom sagemaker.sklearn.processing import SKLearnProcessor\nfrom sagemaker.model_metrics import (\n MetricsSource,\n ModelMetrics,\n)\nfrom sagemaker.workflow.parameters import (\n ParameterInteger,\n ParameterString,\n)\nfrom sagemaker.workflow.pipeline import Pipeline\nfrom sagemaker.workflow.properties import PropertyFile\nfrom sagemaker.workflow.steps import (\n ProcessingStep,\n CacheConfig,\n TuningStep,\n)\nfrom sagemaker.workflow.step_collections import RegisterModel, CreateModelStep\nfrom sagemaker.workflow.conditions import ConditionLessThanOrEqualTo\nfrom sagemaker.workflow.condition_step import ConditionStep\n\nfrom sagemaker.workflow.functions import Join, JsonGet\nfrom sagemaker.workflow.execution_variables import ExecutionVariables\n\nfrom sagemaker.tuner import (\n ContinuousParameter,\n HyperparameterTuner,\n WarmStartConfig,\n WarmStartTypes,\n)",
"_____no_output_____"
],
[
"# Create the SageMaker Session\n\nregion = sagemaker.Session().boto_region_name\nsm_client = boto3.client(\"sagemaker\")\nboto_session = boto3.Session(region_name=region)\nsagemaker_session = sagemaker.session.Session(boto_session=boto_session, sagemaker_client=sm_client)",
"_____no_output_____"
],
[
"# Define variables and parameters needed for the Pipeline steps\n\nrole = sagemaker.get_execution_role()\ndefault_bucket = sagemaker_session.default_bucket()\nbase_job_prefix = \"tuning-step-example\"\nmodel_package_group_name = \"tuning-job-model-packages\"\n\nprocessing_instance_count = ParameterInteger(name=\"ProcessingInstanceCount\", default_value=1)\nprocessing_instance_type = ParameterString(\n name=\"ProcessingInstanceType\", default_value=\"ml.m5.xlarge\"\n)\ntraining_instance_type = ParameterString(name=\"TrainingInstanceType\", default_value=\"ml.m5.xlarge\")\nmodel_approval_status = ParameterString(\n name=\"ModelApprovalStatus\", default_value=\"PendingManualApproval\"\n)\ninput_data = ParameterString(\n name=\"InputDataUrl\",\n default_value=f\"s3://sagemaker-servicecatalog-seedcode-{region}/dataset/abalone-dataset.csv\",\n)\nmodel_approval_status = ParameterString(\n name=\"ModelApprovalStatus\", default_value=\"PendingManualApproval\"\n)\n\n# Cache Pipeline steps to reduce execution time on subsequent executions\ncache_config = CacheConfig(enable_caching=True, expire_after=\"30d\")",
"_____no_output_____"
]
],
[
[
"#### Data Preparation\n\nAn SKLearn processor is used to prepare the dataset for the Hyperparameter Tuning job. Using the script `preprocess.py`, the dataset is featurized and split into train, test, and validation datasets. \n\nThe output of this step is used as the input to the TuningStep",
"_____no_output_____"
]
],
[
[
"%%writefile preprocess.py\n\n\"\"\"Feature engineers the abalone dataset.\"\"\"\nimport argparse\nimport logging\nimport os\nimport pathlib\nimport requests\nimport tempfile\n\nimport boto3\nimport numpy as np\nimport pandas as pd\n\nfrom sklearn.compose import ColumnTransformer\nfrom sklearn.impute import SimpleImputer\nfrom sklearn.pipeline import Pipeline\nfrom sklearn.preprocessing import StandardScaler, OneHotEncoder\n\nlogger = logging.getLogger()\nlogger.setLevel(logging.INFO)\nlogger.addHandler(logging.StreamHandler())\n\n\n# Since we get a headerless CSV file we specify the column names here.\nfeature_columns_names = [\n \"sex\",\n \"length\",\n \"diameter\",\n \"height\",\n \"whole_weight\",\n \"shucked_weight\",\n \"viscera_weight\",\n \"shell_weight\",\n]\nlabel_column = \"rings\"\n\nfeature_columns_dtype = {\n \"sex\": str,\n \"length\": np.float64,\n \"diameter\": np.float64,\n \"height\": np.float64,\n \"whole_weight\": np.float64,\n \"shucked_weight\": np.float64,\n \"viscera_weight\": np.float64,\n \"shell_weight\": np.float64,\n}\nlabel_column_dtype = {\"rings\": np.float64}\n\n\ndef merge_two_dicts(x, y):\n \"\"\"Merges two dicts, returning a new copy.\"\"\"\n z = x.copy()\n z.update(y)\n return z\n\n\nif __name__ == \"__main__\":\n logger.debug(\"Starting preprocessing.\")\n parser = argparse.ArgumentParser()\n parser.add_argument(\"--input-data\", type=str, required=True)\n args = parser.parse_args()\n\n base_dir = \"/opt/ml/processing\"\n pathlib.Path(f\"{base_dir}/data\").mkdir(parents=True, exist_ok=True)\n input_data = args.input_data\n bucket = input_data.split(\"/\")[2]\n key = \"/\".join(input_data.split(\"/\")[3:])\n\n logger.info(\"Downloading data from bucket: %s, key: %s\", bucket, key)\n fn = f\"{base_dir}/data/abalone-dataset.csv\"\n s3 = boto3.resource(\"s3\")\n s3.Bucket(bucket).download_file(key, fn)\n\n logger.debug(\"Reading downloaded data.\")\n df = pd.read_csv(\n fn,\n header=None,\n names=feature_columns_names + [label_column],\n dtype=merge_two_dicts(feature_columns_dtype, label_column_dtype),\n )\n os.unlink(fn)\n\n logger.debug(\"Defining transformers.\")\n numeric_features = list(feature_columns_names)\n numeric_features.remove(\"sex\")\n numeric_transformer = Pipeline(\n steps=[\n (\"imputer\", SimpleImputer(strategy=\"median\")),\n (\"scaler\", StandardScaler()),\n ]\n )\n\n categorical_features = [\"sex\"]\n categorical_transformer = Pipeline(\n steps=[\n (\"imputer\", SimpleImputer(strategy=\"constant\", fill_value=\"missing\")),\n (\"onehot\", OneHotEncoder(handle_unknown=\"ignore\")),\n ]\n )\n\n preprocess = ColumnTransformer(\n transformers=[\n (\"num\", numeric_transformer, numeric_features),\n (\"cat\", categorical_transformer, categorical_features),\n ]\n )\n\n logger.info(\"Applying transforms.\")\n y = df.pop(\"rings\")\n X_pre = preprocess.fit_transform(df)\n y_pre = y.to_numpy().reshape(len(y), 1)\n\n X = np.concatenate((y_pre, X_pre), axis=1)\n\n logger.info(\"Splitting %d rows of data into train, validation, test datasets.\", len(X))\n np.random.shuffle(X)\n train, validation, test = np.split(X, [int(0.7 * len(X)), int(0.85 * len(X))])\n\n logger.info(\"Writing out datasets to %s.\", base_dir)\n pd.DataFrame(train).to_csv(f\"{base_dir}/train/train.csv\", header=False, index=False)\n pd.DataFrame(validation).to_csv(\n f\"{base_dir}/validation/validation.csv\", header=False, index=False\n )\n pd.DataFrame(test).to_csv(f\"{base_dir}/test/test.csv\", header=False, index=False)",
"_____no_output_____"
],
[
"# Process the training data step using a python script.\n# Split the training data set into train, test, and validation datasets\n# When defining the ProcessingOutput destination as a dynamic value using the\n# Pipeline Execution ID, caching will not be in effect as each time the step runs,\n# the step definition changes resulting in new execution. If caching is required,\n# the ProcessingOutput definition should be status\n\nsklearn_processor = SKLearnProcessor(\n framework_version=\"0.23-1\",\n instance_type=processing_instance_type,\n instance_count=processing_instance_count,\n base_job_name=f\"{base_job_prefix}/sklearn-abalone-preprocess\",\n sagemaker_session=sagemaker_session,\n role=role,\n)\nstep_process = ProcessingStep(\n name=\"PreprocessAbaloneDataForHPO\",\n processor=sklearn_processor,\n outputs=[\n ProcessingOutput(\n output_name=\"train\",\n source=\"/opt/ml/processing/train\",\n destination=Join(\n on=\"/\",\n values=[\n \"s3:/\",\n default_bucket,\n base_job_prefix,\n ExecutionVariables.PIPELINE_EXECUTION_ID,\n \"PreprocessAbaloneDataForHPO\",\n ],\n ),\n ),\n ProcessingOutput(\n output_name=\"validation\",\n source=\"/opt/ml/processing/validation\",\n destination=Join(\n on=\"/\",\n values=[\n \"s3:/\",\n default_bucket,\n base_job_prefix,\n ExecutionVariables.PIPELINE_EXECUTION_ID,\n \"PreprocessAbaloneDataForHPO\",\n ],\n ),\n ),\n ProcessingOutput(\n output_name=\"test\",\n source=\"/opt/ml/processing/test\",\n destination=Join(\n on=\"/\",\n values=[\n \"s3:/\",\n default_bucket,\n base_job_prefix,\n ExecutionVariables.PIPELINE_EXECUTION_ID,\n \"PreprocessAbaloneDataForHPO\",\n ],\n ),\n ),\n ],\n code=\"preprocess.py\",\n job_arguments=[\"--input-data\", input_data],\n)",
"_____no_output_____"
]
],
[
[
"#### Hyperparameter Tuning\n\nAmazon SageMaker automatic model tuning, also known as hyperparameter tuning, finds the best version of a model by running many training jobs on your dataset using the algorithm and ranges of hyperparameters that you specify. It then chooses the hyperparameter values that result in a model that performs the best, as measured by a metric that you choose. \n\n[Valid metrics](https://github.com/dmlc/xgboost/blob/master/doc/parameter.rst#learning-task-parameters) for XGBoost Tuning Job\n\nYou can learn more about [Hyperparameter Tuning](https://docs.aws.amazon.com/sagemaker/latest/dg/automatic-model-tuning-how-it-works.html) in the SageMaker docs.",
"_____no_output_____"
]
],
[
[
"# Define the output path for the model artifacts from the Hyperparameter Tuning Job\nmodel_path = f\"s3://{default_bucket}/{base_job_prefix}/AbaloneTrain\"\n\nimage_uri = sagemaker.image_uris.retrieve(\n framework=\"xgboost\",\n region=region,\n version=\"1.0-1\",\n py_version=\"py3\",\n instance_type=training_instance_type,\n)\n\nxgb_train = Estimator(\n image_uri=image_uri,\n instance_type=training_instance_type,\n instance_count=1,\n output_path=model_path,\n base_job_name=f\"{base_job_prefix}/abalone-train\",\n sagemaker_session=sagemaker_session,\n role=role,\n)\n\nxgb_train.set_hyperparameters(\n eval_metric=\"rmse\",\n objective=\"reg:squarederror\", # Define the object metric for the training job\n num_round=50,\n max_depth=5,\n eta=0.2,\n gamma=4,\n min_child_weight=6,\n subsample=0.7,\n silent=0,\n)\n\nobjective_metric_name = \"validation:rmse\"\n\nhyperparameter_ranges = {\n \"alpha\": ContinuousParameter(0.01, 10, scaling_type=\"Logarithmic\"),\n \"lambda\": ContinuousParameter(0.01, 10, scaling_type=\"Logarithmic\"),\n}\n\ntuner_log = HyperparameterTuner(\n xgb_train,\n objective_metric_name,\n hyperparameter_ranges,\n max_jobs=3,\n max_parallel_jobs=3,\n strategy=\"Random\",\n objective_type=\"Minimize\",\n)\n\nstep_tuning = TuningStep(\n name=\"HPTuning\",\n tuner=tuner_log,\n inputs={\n \"train\": TrainingInput(\n s3_data=step_process.properties.ProcessingOutputConfig.Outputs[\"train\"].S3Output.S3Uri,\n content_type=\"text/csv\",\n ),\n \"validation\": TrainingInput(\n s3_data=step_process.properties.ProcessingOutputConfig.Outputs[\n \"validation\"\n ].S3Output.S3Uri,\n content_type=\"text/csv\",\n ),\n },\n cache_config=cache_config,\n)",
"_____no_output_____"
]
],
[
[
"#### Warm start for Hyperparameter Tuning Job\n\nUse warm start to start a hyperparameter tuning job using one or more previous tuning jobs as a starting point. The results of previous tuning jobs are used to inform which combinations of hyperparameters to search over in the new tuning job. Hyperparameter tuning uses either Bayesian or random search to choose combinations of hyperparameter values from ranges that you specify.\n\nFind more information on [Warm Starts](https://docs.aws.amazon.com/sagemaker/latest/dg/automatic-model-tuning-warm-start.html) in the SageMaker docs.\n\nIn a training pipeline, the parent tuning job name can be provided as a pipeline parameter if there is an already complete Hyperparameter tuning job that should be used as the basis for the warm start. \n\nThis step is left out of the pipeline steps in this notebook. It can be added into the steps while defining the pipeline and the appropriate parent tuning job should be specified. ",
"_____no_output_____"
]
],
[
[
"# This is an example to illustrate how a the name of the tuning job from the previous step can be used as the parent tuning job, in practice,\n# it is unlikely to have the parent job run before the warm start job on each run. Typically the first tuning job would run and the pipeline\n# would be altered to use tuning jobs with a warm start using the first job as the parent job.\n\nparent_tuning_job_name = (\n step_tuning.properties.HyperParameterTuningJobName\n) # Use the parent tuning job specific to the use case\n\nwarm_start_config = WarmStartConfig(\n WarmStartTypes.IDENTICAL_DATA_AND_ALGORITHM, parents={parent_tuning_job_name}\n)\n\ntuner_log_warm_start = HyperparameterTuner(\n xgb_train,\n objective_metric_name,\n hyperparameter_ranges,\n max_jobs=3,\n max_parallel_jobs=3,\n strategy=\"Random\",\n objective_type=\"Minimize\",\n warm_start_config=warm_start_config,\n)\n\nstep_tuning_warm_start = TuningStep(\n name=\"HPTuningWarmStart\",\n tuner=tuner_log_warm_start,\n inputs={\n \"train\": TrainingInput(\n s3_data=step_process.properties.ProcessingOutputConfig.Outputs[\"train\"].S3Output.S3Uri,\n content_type=\"text/csv\",\n ),\n \"validation\": TrainingInput(\n s3_data=step_process.properties.ProcessingOutputConfig.Outputs[\n \"validation\"\n ].S3Output.S3Uri,\n content_type=\"text/csv\",\n ),\n },\n cache_config=cache_config,\n)",
"_____no_output_____"
]
],
[
[
"#### Creating and Registering the best models\n\nAfter successfully completing the Hyperparameter Tuning job. You can either create SageMaker models from the model artifacts created by the training jobs from the TuningStep or register the models into the Model Registry. \n\nWhen using the model Registry, if you register multiple models from the TuningStep, they will be registered as versions within the same model package group unless unique model package groups are specified for each RegisterModelStep that is part of the pipeline. \n\nIn this example, the two best models from the TuningStep are added to the same model package group in the Model Registry as v0 and v1.\n\nYou use the `get_top_model_s3_uri` method of the TuningStep class to get the model artifact from one of the top performing model versions",
"_____no_output_____"
]
],
[
[
"# Creating 2 SageMaker Models\n\nmodel_bucket_key = f\"{default_bucket}/{base_job_prefix}/AbaloneTrain\"\nbest_model = Model(\n image_uri=image_uri,\n model_data=step_tuning.get_top_model_s3_uri(top_k=0, s3_bucket=model_bucket_key),\n sagemaker_session=sagemaker_session,\n role=role,\n predictor_cls=XGBoostPredictor,\n)\n\nstep_create_first = CreateModelStep(\n name=\"CreateTopModel\",\n model=best_model,\n inputs=sagemaker.inputs.CreateModelInput(instance_type=\"ml.m4.large\"),\n)\n\nsecond_best_model = Model(\n image_uri=image_uri,\n model_data=step_tuning.get_top_model_s3_uri(top_k=1, s3_bucket=model_bucket_key),\n sagemaker_session=sagemaker_session,\n role=role,\n predictor_cls=XGBoostPredictor,\n)\n\nstep_create_second = CreateModelStep(\n name=\"CreateSecondBestModel\",\n model=second_best_model,\n inputs=sagemaker.inputs.CreateModelInput(instance_type=\"ml.m4.large\"),\n)",
"_____no_output_____"
]
],
[
[
"#### Evaluate the top model\n\nUse a processing job to evaluate the top model from the tuning step",
"_____no_output_____"
]
],
[
[
"%%writefile evaluate.py\n\n\"\"\"Evaluation script for measuring mean squared error.\"\"\"\nimport json\nimport logging\nimport pathlib\nimport pickle\nimport tarfile\n\nimport numpy as np\nimport pandas as pd\nimport xgboost\n\nfrom sklearn.metrics import mean_squared_error\n\nlogger = logging.getLogger()\nlogger.setLevel(logging.INFO)\nlogger.addHandler(logging.StreamHandler())\n\n\nif __name__ == \"__main__\":\n logger.debug(\"Starting evaluation.\")\n model_path = \"/opt/ml/processing/model/model.tar.gz\"\n with tarfile.open(model_path) as tar:\n tar.extractall(path=\".\")\n\n logger.debug(\"Loading xgboost model.\")\n model = pickle.load(open(\"xgboost-model\", \"rb\"))\n\n logger.debug(\"Reading test data.\")\n test_path = \"/opt/ml/processing/test/test.csv\"\n df = pd.read_csv(test_path, header=None)\n\n logger.debug(\"Reading test data.\")\n y_test = df.iloc[:, 0].to_numpy()\n df.drop(df.columns[0], axis=1, inplace=True)\n X_test = xgboost.DMatrix(df.values)\n\n logger.info(\"Performing predictions against test data.\")\n predictions = model.predict(X_test)\n\n logger.debug(\"Calculating mean squared error.\")\n mse = mean_squared_error(y_test, predictions)\n std = np.std(y_test - predictions)\n report_dict = {\n \"regression_metrics\": {\n \"mse\": {\"value\": mse, \"standard_deviation\": std},\n },\n }\n\n output_dir = \"/opt/ml/processing/evaluation\"\n pathlib.Path(output_dir).mkdir(parents=True, exist_ok=True)\n\n logger.info(\"Writing out evaluation report with mse: %f\", mse)\n evaluation_path = f\"{output_dir}/evaluation.json\"\n with open(evaluation_path, \"w\") as f:\n f.write(json.dumps(report_dict))",
"_____no_output_____"
],
[
"# A ProcessingStep is used to evaluate the performance of a selected model from the HPO step. In this case, the top performing model\n# is evaluated. Based on the results of the evaluation, the model is registered into the Model Registry using a ConditionStep.\n\nscript_eval = ScriptProcessor(\n image_uri=image_uri,\n command=[\"python3\"],\n instance_type=processing_instance_type,\n instance_count=1,\n base_job_name=f\"{base_job_prefix}/script-tuning-step-eval\",\n sagemaker_session=sagemaker_session,\n role=role,\n)\n\nevaluation_report = PropertyFile(\n name=\"BestTuningModelEvaluationReport\",\n output_name=\"evaluation\",\n path=\"evaluation.json\",\n)\n\n# This can be extended to evaluate multiple models from the HPO step\nstep_eval = ProcessingStep(\n name=\"EvaluateTopModel\",\n processor=script_eval,\n inputs=[\n ProcessingInput(\n source=step_tuning.get_top_model_s3_uri(top_k=0, s3_bucket=model_bucket_key),\n destination=\"/opt/ml/processing/model\",\n ),\n ProcessingInput(\n source=step_process.properties.ProcessingOutputConfig.Outputs[\"test\"].S3Output.S3Uri,\n destination=\"/opt/ml/processing/test\",\n ),\n ],\n outputs=[\n ProcessingOutput(output_name=\"evaluation\", source=\"/opt/ml/processing/evaluation\"),\n ],\n code=\"evaluate.py\",\n property_files=[evaluation_report],\n cache_config=cache_config,\n)\n\nmodel_metrics = ModelMetrics(\n model_statistics=MetricsSource(\n s3_uri=\"{}/evaluation.json\".format(\n step_eval.arguments[\"ProcessingOutputConfig\"][\"Outputs\"][0][\"S3Output\"][\"S3Uri\"]\n ),\n content_type=\"application/json\",\n )\n)",
"_____no_output_____"
],
[
"# Register the model in the Model Registry\n# Multiple models can be registered into the Model Registry using multiple RegisterModel steps. These models can either be added to the\n# same model package group as different versions within the group or the models can be added to different model package groups.\n\nstep_register_best = RegisterModel(\n name=\"RegisterBestAbaloneModel\",\n estimator=xgb_train,\n model_data=step_tuning.get_top_model_s3_uri(top_k=0, s3_bucket=model_bucket_key),\n content_types=[\"text/csv\"],\n response_types=[\"text/csv\"],\n inference_instances=[\"ml.t2.medium\", \"ml.m5.large\"],\n transform_instances=[\"ml.m5.large\"],\n model_package_group_name=model_package_group_name,\n approval_status=model_approval_status,\n)",
"_____no_output_____"
],
[
"# condition step for evaluating model quality and branching execution\n\ncond_lte = ConditionLessThanOrEqualTo(\n left=JsonGet(\n step_name=step_eval.name,\n property_file=evaluation_report,\n json_path=\"regression_metrics.mse.value\",\n ),\n right=6.0,\n)\nstep_cond = ConditionStep(\n name=\"CheckMSEAbaloneEvaluation\",\n conditions=[cond_lte],\n if_steps=[step_register_best],\n else_steps=[],\n)",
"_____no_output_____"
],
[
"pipeline = Pipeline(\n name=\"tuning-step-pipeline\",\n parameters=[\n processing_instance_type,\n processing_instance_count,\n training_instance_type,\n input_data,\n model_approval_status,\n ],\n steps=[\n step_process,\n step_tuning,\n step_create_first,\n step_create_second,\n step_eval,\n step_cond,\n ],\n sagemaker_session=sagemaker_session,\n)",
"_____no_output_____"
]
],
[
[
"#### Execute the Pipeline",
"_____no_output_____"
]
],
[
[
"import json\n\ndefinition = json.loads(pipeline.definition())\ndefinition",
"_____no_output_____"
],
[
"pipeline.upsert(role_arn=role)",
"_____no_output_____"
],
[
"pipeline.start()",
"_____no_output_____"
]
],
[
[
"#### Cleaning up resources\n\nUsers are responsible for cleaning up resources created when running this notebook. Specify the ModelName, ModelPackageName, and ModelPackageGroupName that need to be deleted. The model names are generated by the CreateModel step of the Pipeline and the property values are available only in the Pipeline context. To delete the models created by this pipeline, navigate to the Model Registry and Console to find the models to delete.",
"_____no_output_____"
]
],
[
[
"# # Create a SageMaker client\n# sm_client = boto3.client(\"sagemaker\")\n\n# # Delete SageMaker Models\n# sm_client.delete_model(ModelName=\"...\")\n\n# # Delete Model Packages\n# sm_client.delete_model_package(ModelPackageName=\"...\")\n\n# # Delete the Model Package Group\n# sm_client.delete_model_package_group(ModelPackageGroupName=\"...\")\n\n# # Delete the Pipeline\n# sm_client.delete_pipeline(PipelineName=\"tuning-step-pipeline\")",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ab60a97a9509e158190c36b6dd8b18fb3da6026
| 541,442 |
ipynb
|
Jupyter Notebook
|
intro-to-data-analysis/python_overview/Demo.ipynb
|
bhetey/become_data_analyst
|
494193e337456dc5681fc937d933ab84c1109109
|
[
"MIT"
] | null | null | null |
intro-to-data-analysis/python_overview/Demo.ipynb
|
bhetey/become_data_analyst
|
494193e337456dc5681fc937d933ab84c1109109
|
[
"MIT"
] | null | null | null |
intro-to-data-analysis/python_overview/Demo.ipynb
|
bhetey/become_data_analyst
|
494193e337456dc5681fc937d933ab84c1109109
|
[
"MIT"
] | null | null | null | 16.833266 | 1,825 | 0.344384 |
[
[
[
"# Python Demo\n _This is a demo for python_",
"_____no_output_____"
]
],
[
[
"firstnumber = 5\nsecondnumber = 5\nadditon = firstnumber + secondnumber\nprint (additon)",
"10\n"
]
],
[
[
"## List of number or strings ",
"_____no_output_____"
]
],
[
[
"arr = [1,2,3,3,4,4,4,4,4,4]\nprint(arr)",
"[1, 2, 3, 3, 4, 4, 4, 4, 4, 4]\n"
],
[
"a = input('Write your first number : ')\nb = input('Write your second number : ')\ndef addNumber (firstnumber , secondNUmber):\n return int(firstnumber) + int(secondNUmber)\n\naddNumber(a, b)\nprint (type(addNumber))",
"Write your first number : 3\nWrite your second number : 3\n<class 'function'>\n"
],
[
"length = len('my_string')\nprint (type(length))\nprint (length)",
"<class 'int'>\n9\n"
],
[
"## Set of no \nmySet = set([1,2,3,3,3,3,3,3,3,3,3,3,3,3,3,3,3,4,5,5,7])\nprint(mySet)",
"{1, 2, 3, 4, 5, 7}\n"
],
[
"numb = 1\nprint (type(str(numb)) )",
"<class 'str'>\n"
],
[
"first = 3\nsecond = 5\nthird = 6\naddition = first + second + third\nprint ('This is ' , addition)",
"This is 14\n"
]
],
[
[
"## For loop demo",
"_____no_output_____"
]
],
[
[
"## for loops example \nsampleData = [1,'iseg', 'pingo', 'parliamento', 12435]\nfor e in sampleData:\n if e is not 'iseg':\n print (e)",
"1\npingo\nparliamento\n12435\n"
],
[
"a = 10\nb = 0\nwhile a == 10:\n b += 1\n print(b)",
"1\n2\n3\n4\n5\n6\n7\n8\n9\n10\n11\n12\n13\n14\n15\n16\n17\n18\n19\n20\n21\n22\n23\n24\n25\n26\n27\n28\n29\n30\n31\n32\n33\n34\n35\n36\n37\n38\n39\n40\n41\n42\n43\n44\n45\n46\n47\n48\n49\n50\n51\n52\n53\n54\n55\n56\n57\n58\n59\n60\n61\n62\n63\n64\n65\n66\n67\n68\n69\n70\n71\n72\n73\n74\n75\n76\n77\n78\n79\n80\n81\n82\n83\n84\n85\n86\n87\n88\n89\n90\n91\n92\n93\n94\n95\n96\n97\n98\n99\n100\n101\n102\n103\n104\n105\n106\n107\n108\n109\n110\n111\n112\n113\n114\n115\n116\n117\n118\n119\n120\n121\n122\n123\n124\n125\n126\n127\n128\n129\n130\n131\n132\n133\n134\n135\n136\n137\n138\n139\n140\n141\n142\n143\n144\n145\n146\n147\n148\n149\n150\n151\n152\n153\n154\n155\n156\n157\n158\n159\n160\n161\n162\n163\n164\n165\n166\n167\n168\n169\n170\n171\n172\n173\n174\n175\n176\n177\n178\n179\n180\n181\n182\n183\n184\n185\n186\n187\n188\n189\n190\n191\n192\n193\n194\n195\n196\n197\n198\n199\n200\n201\n202\n203\n204\n205\n206\n207\n208\n209\n210\n211\n212\n213\n214\n215\n216\n217\n218\n219\n220\n221\n222\n223\n224\n225\n226\n227\n228\n229\n230\n231\n232\n233\n234\n235\n236\n237\n238\n239\n240\n241\n242\n243\n244\n245\n246\n247\n248\n249\n250\n251\n252\n253\n254\n255\n256\n257\n258\n259\n260\n261\n262\n263\n264\n265\n266\n267\n268\n269\n270\n271\n272\n273\n274\n275\n276\n277\n278\n279\n280\n281\n282\n283\n284\n285\n286\n287\n288\n289\n290\n291\n292\n293\n294\n295\n296\n297\n298\n299\n300\n301\n302\n303\n304\n305\n306\n307\n308\n309\n310\n311\n312\n313\n314\n315\n316\n317\n318\n319\n320\n321\n322\n323\n324\n325\n326\n327\n328\n329\n330\n331\n332\n333\n334\n335\n336\n337\n338\n339\n340\n341\n342\n343\n344\n345\n346\n347\n348\n349\n350\n351\n352\n353\n354\n355\n356\n357\n358\n359\n360\n361\n362\n363\n364\n365\n366\n367\n368\n369\n370\n371\n372\n373\n374\n375\n376\n377\n378\n379\n380\n381\n382\n383\n384\n385\n386\n387\n388\n389\n390\n391\n392\n393\n394\n395\n396\n397\n398\n399\n400\n401\n402\n403\n404\n405\n406\n407\n408\n409\n410\n411\n412\n413\n414\n415\n416\n417\n418\n419\n420\n421\n422\n423\n424\n425\n426\n427\n428\n429\n430\n431\n432\n433\n434\n435\n436\n437\n438\n439\n440\n441\n442\n443\n444\n445\n446\n447\n448\n449\n450\n451\n452\n453\n454\n455\n456\n457\n458\n459\n460\n461\n462\n463\n464\n465\n466\n467\n468\n469\n470\n471\n472\n473\n474\n475\n476\n477\n478\n479\n480\n481\n482\n483\n484\n485\n486\n487\n488\n489\n490\n491\n492\n493\n494\n495\n496\n497\n498\n499\n500\n501\n502\n503\n504\n505\n506\n507\n508\n509\n510\n511\n512\n513\n514\n515\n516\n517\n518\n519\n520\n521\n522\n523\n524\n525\n526\n527\n528\n529\n530\n531\n532\n533\n534\n535\n536\n537\n538\n539\n540\n541\n542\n543\n544\n545\n546\n547\n548\n549\n550\n551\n552\n553\n554\n555\n556\n557\n558\n559\n560\n561\n562\n563\n564\n565\n566\n567\n568\n569\n570\n571\n572\n573\n574\n575\n576\n577\n578\n579\n580\n581\n582\n583\n584\n585\n586\n587\n588\n589\n590\n591\n592\n593\n594\n595\n596\n597\n598\n599\n600\n601\n602\n603\n604\n605\n606\n607\n608\n609\n610\n611\n612\n613\n614\n615\n616\n617\n618\n619\n620\n621\n622\n623\n624\n625\n626\n627\n628\n629\n630\n631\n632\n633\n634\n635\n636\n637\n638\n639\n640\n641\n642\n643\n644\n645\n646\n647\n648\n649\n650\n651\n652\n653\n654\n655\n656\n657\n658\n659\n660\n661\n662\n663\n664\n665\n666\n667\n668\n669\n670\n671\n672\n673\n674\n675\n676\n677\n678\n679\n680\n681\n682\n683\n684\n685\n686\n687\n688\n689\n690\n691\n692\n693\n694\n695\n696\n697\n698\n699\n700\n701\n702\n703\n704\n705\n706\n707\n708\n709\n710\n711\n712\n713\n714\n715\n716\n717\n718\n719\n720\n721\n722\n723\n724\n725\n726\n727\n728\n729\n730\n731\n732\n733\n734\n735\n736\n737\n738\n739\n740\n741\n742\n743\n744\n745\n746\n747\n748\n749\n750\n751\n752\n753\n754\n755\n756\n757\n758\n759\n760\n761\n762\n763\n764\n765\n766\n767\n768\n769\n770\n771\n772\n773\n774\n775\n776\n777\n778\n779\n780\n781\n782\n783\n784\n785\n786\n787\n788\n789\n790\n791\n792\n793\n794\n795\n796\n797\n798\n799\n800\n801\n802\n803\n804\n805\n806\n807\n808\n809\n810\n811\n812\n813\n814\n815\n816\n817\n818\n819\n820\n821\n822\n823\n824\n825\n826\n827\n828\n829\n830\n831\n832\n833\n834\n835\n836\n837\n838\n839\n840\n841\n842\n843\n844\n845\n846\n847\n848\n849\n850\n851\n852\n853\n854\n855\n856\n857\n858\n859\n860\n861\n862\n863\n864\n865\n866\n867\n868\n869\n870\n871\n872\n873\n874\n875\n876\n877\n878\n879\n880\n881\n882\n883\n884\n885\n886\n887\n888\n889\n890\n891\n892\n893\n894\n895\n896\n897\n898\n899\n900\n901\n902\n903\n904\n905\n906\n907\n908\n909\n910\n911\n912\n913\n914\n915\n916\n917\n918\n919\n920\n921\n922\n923\n924\n925\n926\n927\n928\n929\n930\n931\n932\n933\n934\n935\n936\n937\n938\n939\n940\n941\n942\n943\n944\n945\n946\n947\n948\n949\n950\n951\n952\n953\n954\n955\n956\n957\n958\n959\n960\n961\n962\n963\n964\n965\n966\n967\n968\n969\n970\n971\n972\n973\n974\n975\n976\n977\n978\n979\n980\n981\n982\n983\n984\n985\n986\n987\n988\n989\n990\n991\n992\n993\n994\n995\n996\n997\n998\n999\n1000\n1001\n1002\n1003\n1004\n1005\n1006\n1007\n1008\n1009\n1010\n1011\n1012\n1013\n1014\n1015\n1016\n1017\n1018\n1019\n1020\n1021\n1022\n1023\n1024\n1025\n1026\n1027\n1028\n1029\n1030\n1031\n1032\n1033\n1034\n1035\n1036\n1037\n1038\n1039\n1040\n1041\n1042\n1043\n1044\n1045\n1046\n1047\n1048\n1049\n1050\n1051\n1052\n1053\n1054\n1055\n1056\n1057\n1058\n1059\n1060\n1061\n1062\n1063\n1064\n1065\n1066\n1067\n1068\n1069\n1070\n1071\n1072\n1073\n1074\n1075\n1076\n1077\n1078\n1079\n1080\n1081\n1082\n1083\n1084\n1085\n1086\n1087\n1088\n1089\n1090\n1091\n1092\n1093\n1094\n1095\n1096\n1097\n1098\n1099\n1100\n1101\n1102\n1103\n1104\n1105\n1106\n1107\n1108\n1109\n1110\n1111\n1112\n1113\n1114\n1115\n1116\n1117\n1118\n1119\n1120\n1121\n1122\n1123\n1124\n1125\n1126\n1127\n1128\n1129\n1130\n1131\n1132\n1133\n1134\n1135\n1136\n1137\n1138\n1139\n1140\n1141\n1142\n1143\n1144\n1145\n1146\n1147\n1148\n1149\n1150\n1151\n1152\n1153\n1154\n1155\n1156\n1157\n1158\n1159\n1160\n1161\n1162\n1163\n1164\n1165\n1166\n1167\n1168\n1169\n1170\n1171\n1172\n1173\n1174\n1175\n1176\n1177\n1178\n1179\n1180\n1181\n1182\n1183\n1184\n1185\n1186\n1187\n1188\n1189\n1190\n1191\n1192\n1193\n1194\n1195\n1196\n1197\n1198\n1199\n1200\n1201\n1202\n1203\n1204\n1205\n1206\n1207\n1208\n1209\n1210\n1211\n1212\n1213\n1214\n1215\n1216\n1217\n1218\n1219\n1220\n1221\n1222\n1223\n1224\n1225\n1226\n1227\n1228\n1229\n1230\n1231\n1232\n1233\n1234\n1235\n1236\n1237\n1238\n1239\n1240\n1241\n1242\n1243\n1244\n1245\n1246\n1247\n1248\n1249\n1250\n1251\n1252\n1253\n1254\n1255\n1256\n1257\n1258\n1259\n1260\n1261\n1262\n1263\n1264\n1265\n1266\n1267\n1268\n1269\n1270\n1271\n1272\n1273\n1274\n1275\n1276\n1277\n1278\n1279\n1280\n1281\n1282\n1283\n1284\n1285\n1286\n1287\n1288\n1289\n1290\n1291\n1292\n1293\n1294\n1295\n1296\n1297\n1298\n1299\n1300\n1301\n1302\n1303\n1304\n1305\n1306\n1307\n1308\n1309\n1310\n1311\n1312\n1313\n1314\n1315\n1316\n1317\n1318\n1319\n1320\n1321\n1322\n1323\n1324\n1325\n1326\n1327\n1328\n1329\n1330\n1331\n1332\n1333\n1334\n1335\n1336\n1337\n1338\n1339\n1340\n1341\n1342\n1343\n1344\n1345\n1346\n1347\n1348\n1349\n1350\n1351\n1352\n1353\n1354\n1355\n1356\n1357\n1358\n1359\n1360\n1361\n1362\n1363\n1364\n1365\n1366\n1367\n1368\n1369\n1370\n1371\n1372\n1373\n1374\n1375\n1376\n1377\n1378\n1379\n1380\n1381\n1382\n1383\n1384\n1385\n1386\n1387\n1388\n1389\n1390\n1391\n1392\n1393\n1394\n1395\n1396\n1397\n1398\n1399\n1400\n1401\n1402\n1403\n1404\n1405\n1406\n1407\n1408\n1409\n1410\n1411\n1412\n1413\n1414\n1415\n1416\n1417\n1418\n1419\n1420\n1421\n1422\n1423\n1424\n1425\n1426\n1427\n1428\n1429\n1430\n1431\n1432\n1433\n1434\n1435\n1436\n1437\n1438\n1439\n1440\n1441\n1442\n1443\n1444\n1445\n1446\n1447\n1448\n1449\n1450\n1451\n1452\n1453\n1454\n1455\n1456\n1457\n1458\n1459\n1460\n1461\n1462\n1463\n1464\n1465\n1466\n1467\n1468\n1469\n1470\n1471\n1472\n1473\n1474\n1475\n1476\n1477\n1478\n1479\n1480\n1481\n1482\n1483\n1484\n1485\n1486\n1487\n1488\n1489\n1490\n1491\n1492\n1493\n1494\n1495\n1496\n1497\n1498\n1499\n1500\n1501\n1502\n1503\n1504\n1505\n1506\n1507\n1508\n1509\n1510\n1511\n1512\n1513\n1514\n1515\n1516\n1517\n1518\n1519\n1520\n1521\n1522\n1523\n1524\n1525\n1526\n1527\n1528\n1529\n1530\n1531\n1532\n1533\n1534\n1535\n1536\n1537\n1538\n1539\n1540\n1541\n1542\n1543\n1544\n1545\n1546\n1547\n1548\n1549\n1550\n1551\n1552\n1553\n1554\n1555\n1556\n1557\n1558\n1559\n1560\n1561\n1562\n1563\n1564\n1565\n1566\n1567\n1568\n1569\n1570\n1571\n1572\n1573\n1574\n1575\n1576\n1577\n1578\n1579\n1580\n1581\n1582\n1583\n1584\n1585\n1586\n1587\n1588\n1589\n1590\n1591\n1592\n1593\n1594\n1595\n1596\n1597\n1598\n1599\n1600\n1601\n1602\n1603\n1604\n1605\n1606\n1607\n1608\n1609\n1610\n1611\n1612\n1613\n1614\n1615\n1616\n1617\n1618\n1619\n1620\n1621\n1622\n1623\n1624\n1625\n1626\n1627\n1628\n1629\n1630\n1631\n1632\n1633\n1634\n1635\n1636\n1637\n1638\n1639\n1640\n1641\n1642\n1643\n1644\n1645\n1646\n1647\n1648\n1649\n1650\n1651\n1652\n1653\n1654\n1655\n1656\n1657\n1658\n1659\n1660\n1661\n1662\n1663\n1664\n1665\n1666\n1667\n1668\n1669\n1670\n1671\n1672\n1673\n1674\n1675\n1676\n1677\n1678\n1679\n1680\n1681\n1682\n1683\n1684\n1685\n1686\n1687\n1688\n1689\n1690\n1691\n1692\n1693\n1694\n1695\n1696\n1697\n1698\n1699\n1700\n1701\n1702\n1703\n1704\n1705\n1706\n1707\n1708\n1709\n1710\n1711\n1712\n1713\n1714\n1715\n1716\n1717\n1718\n1719\n1720\n1721\n1722\n1723\n1724\n1725\n1726\n1727\n1728\n1729\n1730\n1731\n1732\n1733\n1734\n1735\n1736\n1737\n1738\n1739\n1740\n1741\n1742\n1743\n1744\n1745\n1746\n1747\n1748\n1749\n1750\n1751\n1752\n1753\n1754\n1755\n1756\n1757\n1758\n1759\n1760\n1761\n1762\n1763\n1764\n1765\n1766\n1767\n1768\n1769\n1770\n1771\n1772\n1773\n1774\n1775\n1776\n1777\n1778\n1779\n1780\n1781\n1782\n1783\n1784\n1785\n1786\n1787\n1788\n1789\n1790\n1791\n1792\n1793\n1794\n1795\n1796\n1797\n1798\n1799\n1800\n1801\n1802\n1803\n1804\n1805\n1806\n1807\n1808\n1809\n1810\n1811\n1812\n1813\n1814\n1815\n1816\n1817\n1818\n1819\n1820\n1821\n1822\n1823\n1824\n1825\n1826\n1827\n1828\n1829\n1830\n1831\n1832\n1833\n1834\n1835\n1836\n1837\n1838\n1839\n1840\n1841\n1842\n1843\n1844\n1845\n1846\n1847\n1848\n1849\n1850\n1851\n1852\n1853\n1854\n1855\n1856\n1857\n1858\n1859\n1860\n1861\n1862\n1863\n1864\n1865\n1866\n1867\n1868\n1869\n1870\n1871\n1872\n1873\n1874\n1875\n1876\n1877\n1878\n1879\n1880\n1881\n1882\n1883\n1884\n1885\n1886\n1887\n1888\n1889\n1890\n1891\n1892\n1893\n1894\n1895\n1896\n1897\n1898\n1899\n1900\n1901\n1902\n1903\n1904\n1905\n1906\n1907\n1908\n1909\n1910\n1911\n1912\n1913\n1914\n1915\n1916\n1917\n1918\n1919\n1920\n1921\n1922\n1923\n1924\n1925\n1926\n1927\n1928\n1929\n1930\n1931\n1932\n1933\n1934\n1935\n1936\n1937\n1938\n1939\n1940\n1941\n1942\n1943\n1944\n1945\n1946\n1947\n1948\n1949\n1950\n1951\n1952\n1953\n1954\n1955\n1956\n1957\n1958\n1959\n1960\n1961\n1962\n1963\n1964\n1965\n1966\n1967\n1968\n1969\n1970\n1971\n1972\n1973\n1974\n1975\n1976\n1977\n1978\n1979\n1980\n1981\n1982\n1983\n1984\n1985\n1986\n1987\n1988\n1989\n1990\n1991\n1992\n1993\n1994\n1995\n1996\n1997\n1998\n1999\n2000\n2001\n2002\n2003\n2004\n2005\n2006\n2007\n2008\n2009\n2010\n2011\n2012\n2013\n2014\n2015\n2016\n2017\n2018\n2019\n2020\n2021\n2022\n2023\n2024\n2025\n2026\n2027\n2028\n2029\n2030\n2031\n2032\n2033\n2034\n2035\n2036\n2037\n2038\n2039\n2040\n2041\n2042\n2043\n2044\n2045\n2046\n2047\n2048\n2049\n2050\n2051\n2052\n2053\n2054\n2055\n2056\n2057\n2058\n2059\n2060\n2061\n2062\n2063\n2064\n2065\n2066\n2067\n2068\n2069\n2070\n2071\n2072\n2073\n2074\n2075\n2076\n2077\n2078\n2079\n2080\n2081\n2082\n2083\n2084\n2085\n2086\n2087\n2088\n2089\n2090\n2091\n2092\n2093\n2094\n2095\n2096\n2097\n2098\n2099\n2100\n2101\n2102\n2103\n2104\n2105\n2106\n2107\n2108\n2109\n2110\n2111\n2112\n2113\n2114\n2115\n2116\n2117\n2118\n2119\n2120\n2121\n2122\n2123\n2124\n2125\n2126\n2127\n2128\n2129\n2130\n2131\n2132\n2133\n2134\n2135\n2136\n2137\n2138\n2139\n2140\n2141\n2142\n2143\n2144\n2145\n2146\n2147\n2148\n2149\n2150\n2151\n2152\n2153\n2154\n2155\n2156\n2157\n2158\n2159\n2160\n2161\n2162\n2163\n2164\n2165\n2166\n2167\n2168\n2169\n2170\n2171\n2172\n2173\n2174\n2175\n2176\n2177\n2178\n2179\n2180\n2181\n2182\n2183\n2184\n2185\n2186\n2187\n2188\n2189\n2190\n2191\n2192\n2193\n2194\n2195\n2196\n2197\n2198\n2199\n2200\n2201\n2202\n2203\n2204\n2205\n2206\n2207\n2208\n2209\n2210\n2211\n2212\n2213\n2214\n2215\n2216\n2217\n2218\n2219\n2220\n2221\n2222\n2223\n2224\n2225\n2226\n2227\n2228\n2229\n2230\n2231\n2232\n2233\n2234\n2235\n2236\n2237\n2238\n2239\n2240\n2241\n2242\n2243\n2244\n2245\n2246\n2247\n2248\n2249\n2250\n2251\n2252\n2253\n2254\n2255\n2256\n2257\n2258\n2259\n2260\n2261\n2262\n2263\n2264\n2265\n2266\n2267\n2268\n2269\n2270\n2271\n2272\n2273\n2274\n2275\n2276\n2277\n2278\n2279\n2280\n2281\n2282\n2283\n2284\n2285\n2286\n2287\n2288\n2289\n2290\n2291\n2292\n2293\n2294\n2295\n2296\n2297\n2298\n2299\n2300\n2301\n2302\n2303\n2304\n2305\n2306\n2307\n2308\n2309\n2310\n2311\n2312\n2313\n2314\n2315\n2316\n2317\n2318\n2319\n2320\n2321\n2322\n2323\n2324\n2325\n2326\n2327\n2328\n2329\n2330\n2331\n2332\n2333\n2334\n2335\n2336\n2337\n2338\n2339\n2340\n2341\n2342\n2343\n2344\n2345\n2346\n2347\n2348\n2349\n2350\n2351\n2352\n2353\n2354\n2355\n2356\n2357\n2358\n2359\n2360\n2361\n2362\n2363\n2364\n2365\n2366\n2367\n2368\n2369\n2370\n2371\n2372\n2373\n2374\n2375\n2376\n2377\n2378\n2379\n2380\n2381\n2382\n2383\n2384\n2385\n2386\n2387\n2388\n2389\n2390\n2391\n2392\n2393\n2394\n2395\n2396\n2397\n2398\n2399\n2400\n2401\n2402\n2403\n2404\n2405\n2406\n2407\n2408\n2409\n2410\n2411\n2412\n2413\n2414\n2415\n2416\n2417\n2418\n2419\n2420\n2421\n2422\n2423\n2424\n2425\n2426\n2427\n2428\n2429\n2430\n2431\n2432\n2433\n2434\n2435\n2436\n2437\n2438\n2439\n2440\n2441\n2442\n2443\n2444\n2445\n2446\n2447\n2448\n2449\n2450\n2451\n2452\n2453\n2454\n2455\n2456\n2457\n2458\n2459\n2460\n2461\n2462\n2463\n2464\n2465\n2466\n2467\n2468\n2469\n2470\n2471\n2472\n2473\n2474\n2475\n2476\n2477\n2478\n2479\n2480\n2481\n2482\n2483\n2484\n2485\n2486\n2487\n2488\n2489\n2490\n2491\n2492\n2493\n2494\n2495\n2496\n2497\n2498\n2499\n2500\n2501\n2502\n2503\n2504\n2505\n2506\n2507\n2508\n2509\n2510\n2511\n2512\n2513\n2514\n2515\n2516\n2517\n2518\n2519\n2520\n2521\n2522\n2523\n2524\n2525\n2526\n2527\n2528\n2529\n2530\n2531\n2532\n2533\n2534\n2535\n2536\n2537\n2538\n2539\n2540\n2541\n2542\n2543\n2544\n2545\n2546\n2547\n2548\n2549\n2550\n2551\n2552\n2553\n2554\n2555\n2556\n2557\n2558\n2559\n2560\n2561\n2562\n2563\n2564\n2565\n2566\n2567\n2568\n2569\n2570\n2571\n2572\n2573\n2574\n2575\n2576\n2577\n2578\n2579\n2580\n2581\n2582\n2583\n2584\n2585\n2586\n2587\n2588\n2589\n2590\n2591\n2592\n2593\n2594\n2595\n2596\n2597\n2598\n2599\n2600\n2601\n2602\n2603\n2604\n2605\n2606\n2607\n2608\n2609\n2610\n2611\n2612\n2613\n2614\n2615\n2616\n2617\n2618\n2619\n2620\n2621\n2622\n2623\n2624\n2625\n2626\n2627\n2628\n2629\n2630\n2631\n2632\n2633\n2634\n2635\n2636\n2637\n2638\n2639\n2640\n2641\n2642\n2643\n2644\n2645\n2646\n2647\n2648\n2649\n2650\n2651\n2652\n2653\n2654\n2655\n2656\n2657\n2658\n2659\n2660\n2661\n2662\n2663\n2664\n2665\n2666\n2667\n2668\n2669\n2670\n2671\n2672\n2673\n2674\n2675\n2676\n2677\n2678\n2679\n2680\n2681\n2682\n2683\n2684\n2685\n2686\n2687\n2688\n2689\n2690\n2691\n2692\n2693\n2694\n2695\n2696\n2697\n2698\n2699\n2700\n2701\n2702\n2703\n2704\n2705\n2706\n2707\n2708\n2709\n2710\n2711\n2712\n2713\n2714\n2715\n2716\n2717\n2718\n2719\n2720\n2721\n2722\n2723\n2724\n2725\n2726\n2727\n2728\n2729\n2730\n2731\n2732\n2733\n2734\n2735\n2736\n2737\n2738\n2739\n2740\n2741\n2742\n2743\n2744\n2745\n2746\n2747\n2748\n2749\n2750\n2751\n2752\n2753\n2754\n2755\n2756\n2757\n2758\n2759\n2760\n2761\n2762\n2763\n2764\n2765\n2766\n2767\n2768\n2769\n2770\n2771\n2772\n2773\n2774\n2775\n2776\n2777\n2778\n2779\n2780\n2781\n2782\n2783\n2784\n2785\n2786\n2787\n2788\n2789\n2790\n2791\n2792\n2793\n2794\n2795\n2796\n2797\n2798\n2799\n2800\n2801\n2802\n2803\n2804\n2805\n2806\n2807\n2808\n2809\n2810\n2811\n2812\n2813\n2814\n2815\n2816\n2817\n2818\n2819\n2820\n2821\n2822\n2823\n2824\n2825\n"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4ab614a35ee8e142f87d9e5171d4075d812fcd1f
| 5,438 |
ipynb
|
Jupyter Notebook
|
formation/TP-01-Python pour Data Scientists/Titanic.ipynb
|
meteofrance/formation-machine-learning
|
aa693473ed2f43af019de4fd4abee297e6744a01
|
[
"Apache-2.0"
] | 3 |
2020-12-25T23:48:31.000Z
|
2021-03-17T16:18:28.000Z
|
formation/TP-01-Python pour Data Scientists/Titanic.ipynb
|
meteofrance/formation-machine-learning
|
aa693473ed2f43af019de4fd4abee297e6744a01
|
[
"Apache-2.0"
] | null | null | null |
formation/TP-01-Python pour Data Scientists/Titanic.ipynb
|
meteofrance/formation-machine-learning
|
aa693473ed2f43af019de4fd4abee297e6744a01
|
[
"Apache-2.0"
] | 3 |
2020-11-23T14:31:55.000Z
|
2021-10-01T16:43:22.000Z
| 19.919414 | 114 | 0.529423 |
[
[
[
"# Prédiction des survivants du Titanic",
"_____no_output_____"
]
],
[
[
"import pandas \nimport matplotlib.pyplot as plt",
"_____no_output_____"
]
],
[
[
"## Chargement des données\n",
"_____no_output_____"
]
],
[
[
"fichier_titanic = \"titanic/train.csv\"",
"_____no_output_____"
]
],
[
[
"Signification des coolonnes du jeu de données:\n- survival: \tSurvie (0 = No, 1 = Yes)\n- pclass: \tTicket class \t(1 = 1ère Cl., 2 = 2ème Cl., 3 = 3ème Cl.)\n- sex: \tSexe du passager \t\n- Age: \tAge en nombre d'années \t\n- sibsp: \tnombre de frères et soeurs avec leurs conjoints à bord\n- parch \tnombre de parents/enfants à bord \t\n- ticket: numéro de ticket \t\n- fare: \tprix du billet \t\n- cabin: \tnuméro de cabine \t\n- embarked: \tport d'embarcation (C = Cherbourg, Q = Queenstown, S = Southampton)",
"_____no_output_____"
]
],
[
[
"# A COMPLETER\n\n# charger les données",
"_____no_output_____"
]
],
[
[
"# Description des données",
"_____no_output_____"
]
],
[
[
"# A completer \n\n# quels types de données\n# données manquantes ?",
"_____no_output_____"
]
],
[
[
"## Traitement des données\n### nettoyage des données",
"_____no_output_____"
]
],
[
[
"# mettre la colonne PassengerId en index\n# éliminer les colonnes Name, Ticket\n# creer une unique varible Family = SibSp + Parch\n",
"_____no_output_____"
]
],
[
[
"### Données manquantes",
"_____no_output_____"
]
],
[
[
"# Assigner les ages manquants à la moyenne\n# Creer une nouvelle variable With_cabin égale à True si le passager possède un num de cabine, False sinon\n",
"_____no_output_____"
]
],
[
[
"### Données catégorielles",
"_____no_output_____"
]
],
[
[
"# traiter les données catégorielles pour les rendre exploitables pour la modélisation",
"_____no_output_____"
]
],
[
[
"### Variables",
"_____no_output_____"
]
],
[
[
"# créer un jeu de données X,y\n\nX, y = None, None",
"_____no_output_____"
]
],
[
[
"## Modèles de machine learning",
"_____no_output_____"
]
],
[
[
"from sklearn.ensemble import RandomForestClassifier\nfrom sklearn.neighbors import KNeighborsClassifier",
"_____no_output_____"
],
[
"# définir 2 modèles : un random forrest et un KNN",
"_____no_output_____"
]
],
[
[
"## Evalutation des modèles",
"_____no_output_____"
]
],
[
[
"from sklearn.model_selection import cross_validate",
"_____no_output_____"
]
],
[
[
"- Cross-validation du random forest",
"_____no_output_____"
]
],
[
[
"# calculer les perfomances du random forest en utilisant la cross validation (cv=10)",
"_____no_output_____"
]
],
[
[
"- Cross-validation du KNN",
"_____no_output_____"
]
],
[
[
"# calculer les perfomances du KNN en utilisant la cross validation (cv=10)",
"_____no_output_____"
],
[
"# Quel est le meilleur modèle? Tester différentes valeur des hyperparamètres",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4ab622e13650e9a7bd66035f6ce71e4a9f5c452a
| 30,536 |
ipynb
|
Jupyter Notebook
|
notebooks/nb074_ensembles.ipynb
|
hoelzl/ML-Course
|
efa7ccb7c6583753675bbcda569d3184d1ca98d2
|
[
"MIT"
] | 1 |
2022-03-02T15:59:11.000Z
|
2022-03-02T15:59:11.000Z
|
notebooks/nb074_ensembles.ipynb
|
hoelzl/ML-Course
|
efa7ccb7c6583753675bbcda569d3184d1ca98d2
|
[
"MIT"
] | null | null | null |
notebooks/nb074_ensembles.ipynb
|
hoelzl/ML-Course
|
efa7ccb7c6583753675bbcda569d3184d1ca98d2
|
[
"MIT"
] | null | null | null | 23.471176 | 126 | 0.537333 |
[
[
[
"# Ensembles",
"_____no_output_____"
]
],
[
[
"import numpy as np\nimport matplotlib.pyplot as plt\nimport seaborn as sns\nfrom sklearn.metrics import mean_absolute_error, mean_squared_error\nfrom sklearn.linear_model import LinearRegression\nfrom sklearn.tree import DecisionTreeRegressor\n\nsns.set_theme()",
"_____no_output_____"
],
[
"rng = np.random.default_rng(42)\n\nx = rng.uniform(size=(150, 1), low=0.0, high=10.0)\nx_train, x_test = x[:100], x[100:]\n\nx_plot = np.linspace(0, 10, 500).reshape(-1, 1)",
"_____no_output_____"
],
[
"def lin(x):\n return 0.85 * x - 1.5",
"_____no_output_____"
],
[
"def fun(x):\n return 2 * np.sin(x) + 0.1 * x ** 2 - 2",
"_____no_output_____"
],
[
"def randomize(fun, x, scale=0.5):\n return fun(x) + rng.normal(size=x.shape, scale=scale)",
"_____no_output_____"
],
[
"def evaluate_non_random_regressor(reg_type, f_y, *args, **kwargs):\n reg = reg_type(*args, **kwargs)\n\n y_train = f_y(x_train).reshape(-1)\n y_test = f_y(x_test).reshape(-1)\n\n reg.fit(x_train, y_train)\n y_pred = reg.predict(x_test)\n\n x_plot = np.linspace(0, 10, 500).reshape(-1, 1)\n fig, ax = plt.subplots(figsize=(20, 8))\n sns.lineplot(x=x_plot[:, 0], y=reg.predict(x_plot), ax=ax)\n sns.lineplot(x=x_plot[:, 0], y=f_y(x_plot[:, 0]), ax=ax)\n sns.scatterplot(x=x_train[:, 0], y=y_train, ax=ax)\n plt.show()\n\n mae = mean_absolute_error(y_test, y_pred)\n mse = mean_squared_error(y_test, y_pred)\n rmse = np.sqrt(mean_squared_error(y_test, y_pred))\n print(\n \"\\nNo randomness: \" f\"MAE = {mae:.2f}, MSE = {mse:.2f}, RMSE = {rmse:.2f}\"\n )\n\n return reg",
"_____no_output_____"
],
[
"def plot_graphs(f_y, reg, reg_rand, reg_chaos, y_train, y_rand_train, y_chaos_train):\n x_plot = np.linspace(0, 10, 500).reshape(-1, 1)\n fig, ax = plt.subplots(figsize=(20, 12))\n sns.lineplot(x=x_plot[:, 0], y=reg.predict(x_plot), ax=ax)\n sns.scatterplot(x=x_train[:, 0], y=y_train, ax=ax)\n\n sns.lineplot(x=x_plot[:, 0], y=reg_rand.predict(x_plot), ax=ax)\n sns.scatterplot(x=x_train[:, 0], y=y_rand_train, ax=ax)\n\n sns.lineplot(x=x_plot[:, 0], y=reg_chaos.predict(x_plot), ax=ax)\n sns.scatterplot(x=x_train[:, 0], y=y_chaos_train, ax=ax)\n\n sns.lineplot(x=x_plot[:, 0], y=f_y(x_plot[:, 0]), ax=ax)\n plt.show() ",
"_____no_output_____"
],
[
"def print_evaluation(y_test, y_pred, y_rand_test, y_rand_pred, y_chaos_test, y_chaos_pred):\n mae = mean_absolute_error(y_test, y_pred)\n mae_rand = mean_absolute_error(y_rand_test, y_rand_pred)\n mae_chaos = mean_absolute_error(y_chaos_test, y_chaos_pred)\n\n mse = mean_squared_error(y_test, y_pred)\n mse_rand = mean_squared_error(y_rand_test, y_rand_pred)\n mse_chaos = mean_squared_error(y_chaos_test, y_chaos_pred)\n\n rmse = np.sqrt(mean_squared_error(y_test, y_pred))\n rmse_rand = np.sqrt(mean_squared_error(y_rand_test, y_rand_pred))\n rmse_chaos = np.sqrt(mean_squared_error(y_chaos_test, y_chaos_pred))\n\n print(\n \"\\nNo randomness: \" f\"MAE = {mae:.2f}, MSE = {mse:.2f}, RMSE = {rmse:.2f}\"\n )\n print(\n \"Some randomness: \"\n f\"MAE = {mae_rand:.2f}, MSE = {mse_rand:.2f}, RMSE = {rmse_rand:.2f}\"\n )\n print(\n \"Lots of randomness: \"\n f\"MAE = {mae_chaos:.2f}, MSE = {mse_chaos:.2f}, RMSE = {rmse_chaos:.2f}\"\n )",
"_____no_output_____"
],
[
"def evaluate_regressor(reg_type, f_y, *args, **kwargs):\n reg = reg_type(*args, **kwargs)\n reg_rand = reg_type(*args, **kwargs)\n reg_chaos = reg_type(*args, **kwargs)\n \n y_train = f_y(x_train).reshape(-1)\n y_test = f_y(x_test).reshape(-1)\n y_pred = reg.fit(x_train, y_train).predict(x_test)\n \n y_rand_train = randomize(f_y, x_train).reshape(-1)\n y_rand_test = randomize(f_y, x_test).reshape(-1)\n y_rand_pred = reg_rand.fit(x_train, y_rand_train).predict(x_test)\n\n y_chaos_train = randomize(f_y, x_train, 1.5).reshape(-1)\n y_chaos_test = randomize(f_y, x_test, 1.5).reshape(-1)\n y_chaos_pred = reg_chaos.fit(x_train, y_chaos_train).predict(x_test)\n\n plot_graphs(f_y, reg, reg_rand, reg_chaos, y_train, y_rand_train, y_chaos_train)\n print_evaluation(y_test, y_pred, y_rand_test, y_rand_pred, y_chaos_test, y_chaos_pred)",
"_____no_output_____"
]
],
[
[
"# Ensembles, Random Forests, Gradient Boosted Trees",
"_____no_output_____"
],
[
"## Ensemble Methods\n\nIdea: combine several estimators to improve their overal performance.\n\n- Averaging methods: \n - Independent estimators, average predictions\n - Reduces variance (overfitting)\n - Bagging, random forests\n- Boosting methods:\n - Train estimators sequentially\n - Each estimator is trained to reduce the bias of its (combined) predecessors",
"_____no_output_____"
],
[
"### Bagging\n\n- Averaging method: build several estimators of the same type, average their results\n- Needs some way to introduce differences between estimators\n - Otherwise variance is not reduced\n - Train on random subsets of the training data\n- Reduce overfitting\n- Work best with strong estimators (e.g., decision trees with (moderately) large depth)",
"_____no_output_____"
],
[
"### Random Forests\n\n- Bagging classifier/regressor using decision trees\n- For each tree in the forest:\n - Subset of training data\n - Subset of features\n- Often significant reduction in variance (overfitting)\n- Sometimes increase in bias",
"_____no_output_____"
]
],
[
[
"from sklearn.ensemble import RandomForestRegressor",
"_____no_output_____"
],
[
"evaluate_non_random_regressor(RandomForestRegressor, lin, random_state=42);",
"_____no_output_____"
],
[
"evaluate_non_random_regressor(RandomForestRegressor, fun, random_state=42);",
"_____no_output_____"
],
[
"evaluate_non_random_regressor(\n RandomForestRegressor, fun, n_estimators=25, criterion=\"absolute_error\", random_state=42\n);",
"_____no_output_____"
],
[
"evaluate_regressor(RandomForestRegressor, lin, random_state=42);",
"_____no_output_____"
],
[
"evaluate_regressor(\n RandomForestRegressor, lin, n_estimators=500, max_depth=3, random_state=42\n)",
"_____no_output_____"
],
[
"evaluate_regressor(\n RandomForestRegressor, lin, n_estimators=500, min_samples_leaf=6, random_state=42\n)",
"_____no_output_____"
],
[
"evaluate_regressor(RandomForestRegressor, fun, random_state=42)",
"_____no_output_____"
],
[
"evaluate_regressor(\n RandomForestRegressor,\n fun,\n n_estimators=1000,\n min_samples_leaf=6,\n random_state=43,\n n_jobs=-1,\n)",
"_____no_output_____"
]
],
[
[
"## Gradient Boosted Trees\n\n- Boosting method for both regression and classification\n- Requires differentiable loss function",
"_____no_output_____"
]
],
[
[
"from sklearn.ensemble import GradientBoostingRegressor",
"_____no_output_____"
],
[
"evaluate_non_random_regressor(GradientBoostingRegressor, lin);",
"_____no_output_____"
],
[
"evaluate_non_random_regressor(GradientBoostingRegressor, fun);",
"_____no_output_____"
],
[
"evaluate_regressor(GradientBoostingRegressor, lin);",
"_____no_output_____"
],
[
"evaluate_regressor(GradientBoostingRegressor, lin, n_estimators=200, learning_rate=0.05, loss=\"absolute_error\");",
"_____no_output_____"
],
[
"evaluate_regressor(GradientBoostingRegressor, lin, n_estimators=500, learning_rate=0.01,\n loss=\"absolute_error\", subsample=0.1, random_state=46);",
"_____no_output_____"
],
[
"evaluate_regressor(GradientBoostingRegressor, fun, n_estimators=500, learning_rate=0.01,\n loss=\"absolute_error\", subsample=0.1, random_state=44);",
"_____no_output_____"
]
],
[
[
"### Multiple Features",
"_____no_output_____"
]
],
[
[
"from sklearn.datasets import make_regression\nfrom sklearn.model_selection import train_test_split\nnp.set_printoptions(precision=1)",
"_____no_output_____"
],
[
"x, y, coef = make_regression(n_samples=250, n_features=4, n_informative=1, coef=True, random_state=42)\nx.shape, y.shape, coef",
"_____no_output_____"
],
[
"fig, axs = plt.subplots(ncols=2, nrows=2, figsize=(20, 12))\nfor i, ax in enumerate(axs.reshape(-1)):\n sns.scatterplot(x=x[:, i], y=y, ax=ax)",
"_____no_output_____"
],
[
"x, y, coef = make_regression(n_samples=250, n_features=20, n_informative=10, coef=True, random_state=42)\nx.shape, y.shape, coef",
"_____no_output_____"
],
[
"x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.4)\nx_train.shape, x_test.shape, y_train.shape, y_test.shape",
"_____no_output_____"
],
[
"fig, axs = plt.subplots(ncols=2, nrows=2, figsize=(20, 12))\nfor i in range(2):\n sns.scatterplot(x=x[:, i], y=y, ax=axs[0, i]);\nfor i in range(2):\n sns.scatterplot(x=x[:, i + 6], y=y, ax=axs[1, i]);",
"_____no_output_____"
],
[
"lr_clf = LinearRegression()\nlr_clf.fit(x_train, y_train)\ny_lr_pred = lr_clf.predict(x_test)\n\nmean_absolute_error(y_test, y_lr_pred), mean_squared_error(y_test, y_lr_pred)",
"_____no_output_____"
],
[
"lr_clf.coef_.astype(np.int32), coef.astype(np.int32)",
"_____no_output_____"
],
[
"dt_clf = DecisionTreeRegressor()\ndt_clf.fit(x_train, y_train)\ny_dt_pred = dt_clf.predict(x_test)\n\nmean_absolute_error(y_test, y_dt_pred), mean_squared_error(y_test, y_dt_pred)",
"_____no_output_____"
],
[
"rf_clf = RandomForestRegressor()\nrf_clf.fit(x_train, y_train)\ny_rf_pred = rf_clf.predict(x_test)\n\nmean_absolute_error(y_test, y_rf_pred), mean_squared_error(y_test, y_rf_pred)",
"_____no_output_____"
],
[
"gb_clf = GradientBoostingRegressor()\ngb_clf.fit(x_train, y_train)\ny_gb_pred = gb_clf.predict(x_test)\n\nmean_absolute_error(y_test, y_gb_pred), mean_squared_error(y_test, y_gb_pred)",
"_____no_output_____"
],
[
"x, y, coef = make_regression(n_samples=250, n_features=20, n_informative=10, noise=100.0, coef=True, random_state=42)\nx.shape, y.shape, coef",
"_____no_output_____"
],
[
"x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.4)\nx_train.shape, x_test.shape, y_train.shape, y_test.shape",
"_____no_output_____"
],
[
"lr_clf = LinearRegression()\nlr_clf.fit(x_train, y_train)\ny_lr_pred = lr_clf.predict(x_test)\n\nmean_absolute_error(y_test, y_lr_pred), mean_squared_error(y_test, y_lr_pred)",
"_____no_output_____"
],
[
"dt_clf = DecisionTreeRegressor()\ndt_clf.fit(x_train, y_train)\ny_dt_pred = dt_clf.predict(x_test)\n\nmean_absolute_error(y_test, y_dt_pred), mean_squared_error(y_test, y_dt_pred)",
"_____no_output_____"
],
[
"rf_clf = RandomForestRegressor()\nrf_clf.fit(x_train, y_train)\ny_rf_pred = rf_clf.predict(x_test)\n\nmean_absolute_error(y_test, y_rf_pred), mean_squared_error(y_test, y_rf_pred)",
"_____no_output_____"
],
[
"gb_clf = GradientBoostingRegressor()\ngb_clf.fit(x_train, y_train)\ny_gb_pred = gb_clf.predict(x_test)\n\nmean_absolute_error(y_test, y_gb_pred), mean_squared_error(y_test, y_gb_pred)",
"_____no_output_____"
],
[
"x, y, coef = make_regression(n_samples=250, n_features=20, n_informative=10, noise=100.0,\n coef=True, random_state=42)\ny += (20 * x[:, 1]) ** 2\nx.shape, y.shape, coef",
"_____no_output_____"
],
[
"x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.4)\nx_train.shape, x_test.shape, y_train.shape, y_test.shape",
"_____no_output_____"
],
[
"fig, axs = plt.subplots(ncols=2, nrows=2, figsize=(20, 12))\nfor i in range(2):\n sns.scatterplot(x=x[:, i], y=y, ax=axs[0, i]);\nfor i in range(2):\n sns.scatterplot(x=x[:, i + 6], y=y, ax=axs[1, i]);",
"_____no_output_____"
],
[
"lr_clf = LinearRegression()\nlr_clf.fit(x_train, y_train)\ny_lr_pred = lr_clf.predict(x_test)\n\nmean_absolute_error(y_test, y_lr_pred), mean_squared_error(y_test, y_lr_pred)",
"_____no_output_____"
],
[
"dt_clf = DecisionTreeRegressor()\ndt_clf.fit(x_train, y_train)\ny_dt_pred = dt_clf.predict(x_test)\n\nmean_absolute_error(y_test, y_dt_pred), mean_squared_error(y_test, y_dt_pred)",
"_____no_output_____"
],
[
"rf_clf = RandomForestRegressor()\nrf_clf.fit(x_train, y_train)\ny_rf_pred = rf_clf.predict(x_test)\n\nmean_absolute_error(y_test, y_rf_pred), mean_squared_error(y_test, y_rf_pred)",
"_____no_output_____"
],
[
"gb_clf = GradientBoostingRegressor()\ngb_clf.fit(x_train, y_train)\ny_gb_pred = gb_clf.predict(x_test)\n\nmean_absolute_error(y_test, y_gb_pred), mean_squared_error(y_test, y_gb_pred)",
"_____no_output_____"
]
],
[
[
"\n ## Feature Engineering",
"_____no_output_____"
]
],
[
[
"x = rng.uniform(size=(150, 1), low=0.0, high=10.0)\nx_train, x_test = x[:100], x[100:]\nx_plot = np.linspace(0, 10, 500)\nx_train[:3]",
"_____no_output_____"
],
[
"y_lin_train = lin(x_train).reshape(-1)\ny_lin_test = lin(x_test).reshape(-1)\ny_fun_train = fun(x_train.reshape(-1))\ny_fun_test = fun(x_test).reshape(-1)",
"_____no_output_____"
],
[
"x_squares = x * x\nx_squares[:3]",
"_____no_output_____"
],
[
"x_sins = np.sin(x)\nx_sins[:3]",
"_____no_output_____"
],
[
"x_train_aug = np.concatenate([x_train, x_train * x_train, np.sin(x_train)], axis=1)\nx_train_aug[:3]",
"_____no_output_____"
],
[
"x_test_aug = np.concatenate([x_test, x_test * x_test, np.sin(x_test)], axis=1)",
"_____no_output_____"
],
[
"# from sklearn.linear_model import Ridge\n# lr_aug_lin = Ridge()\nlr_aug_lin = LinearRegression()\nlr_aug_lin.fit(x_train_aug, y_lin_train);",
"_____no_output_____"
],
[
"lr_aug_lin.coef_, lr_aug_lin.intercept_",
"_____no_output_____"
],
[
"y_aug_lin_pred = lr_aug_lin.predict(x_test_aug)",
"_____no_output_____"
],
[
"mean_absolute_error(y_lin_test, y_aug_lin_pred), mean_squared_error(\n y_lin_test, y_aug_lin_pred\n)",
"_____no_output_____"
],
[
"x_test.shape, x_plot.shape",
"_____no_output_____"
],
[
"def train_and_plot_aug(f_y, scale=0.5):\n y_plot = f_y(x_plot)\n \n f_r = lambda x: randomize(f_y, x, scale=scale)\n y_train = f_r(x_train_aug[:, 0])\n y_test = f_r(x_test)\n \n lr_aug = LinearRegression() # Try with Ridge() as well...\n lr_aug.fit(x_train_aug, y_train)\n y_pred_test = lr_aug.predict(\n np.concatenate([x_test, x_test * x_test, np.sin(x_test)], axis=1)\n )\n x_plot2 = x_plot.reshape(-1, 1)\n y_pred_plot = lr_aug.predict(\n np.concatenate([x_plot2, x_plot2 * x_plot2, np.sin(x_plot2)], axis=1)\n )\n \n fig, ax = plt.subplots(figsize=(12, 6))\n sns.scatterplot(x=x_plot2[:, 0], y=y_plot, color=\"orange\")\n sns.scatterplot(x=x_plot2[:, 0], y=y_pred_plot, color=\"red\")\n sns.scatterplot(x=x_train_aug[:, 0], y=y_train, color=\"green\")\n plt.show()\n\n mae_in = mean_absolute_error(y_test, y_pred_test)\n mse_in = mean_absolute_error(y_test, y_pred_test)\n rmse_in = np.sqrt(mse_in)\n\n y_nr = f_y(x_test)\n mae_true = mean_absolute_error(y_nr, y_pred_test)\n mse_true = mean_absolute_error(y_nr, y_pred_test)\n rmse_true = np.sqrt(mse_true)\n\n print(f\"Vs. input: MAE: {mae_in:.2f}, MSE: {mse_in:.2f}, RMSE: {rmse_in:.2f}\")\n print(f\"True: MAE: {mae_true:.2f}, MSE: {mse_true:.2f}, RMSE: {rmse_true:.2f}\")\n print(f\"Parameters: {lr_aug.coef_}, {lr_aug.intercept_}\")",
"_____no_output_____"
],
[
"train_and_plot_aug(lin)",
"_____no_output_____"
],
[
"train_and_plot_aug(fun, scale=0.0)",
"_____no_output_____"
],
[
"train_and_plot_aug(fun, scale=0.5)",
"_____no_output_____"
],
[
"train_and_plot_aug(fun, scale=1.5)",
"_____no_output_____"
],
[
"train_and_plot_aug(fun, scale=3)",
"_____no_output_____"
],
[
"def fun2(x): return 2.8 * np.sin(x) + 0.3 * x + 0.08 * x ** 2 - 2.5\n\ntrain_and_plot_aug(fun2, scale=1.5)",
"_____no_output_____"
],
[
"train_and_plot_aug(lambda x: np.select([x<=6, x>6], [-0.5, 3.5]))",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab62be053d4d5885dd415b5877ab522f42c30f6
| 243,149 |
ipynb
|
Jupyter Notebook
|
Pt_1/04_05_Visualizing_time_series/04_05_end.ipynb
|
vladcuevas/Python-Data-Science-Labs
|
eb7b2fa94402f396ed6fd5d4de62f89f930da458
|
[
"MIT"
] | null | null | null |
Pt_1/04_05_Visualizing_time_series/04_05_end.ipynb
|
vladcuevas/Python-Data-Science-Labs
|
eb7b2fa94402f396ed6fd5d4de62f89f930da458
|
[
"MIT"
] | null | null | null |
Pt_1/04_05_Visualizing_time_series/04_05_end.ipynb
|
vladcuevas/Python-Data-Science-Labs
|
eb7b2fa94402f396ed6fd5d4de62f89f930da458
|
[
"MIT"
] | null | null | null | 1,482.615854 | 143,599 | 0.750634 |
[
[
[
"# Chapter 4 - Practical Data Visualization\n## Segment 5 - Visualizing time series",
"_____no_output_____"
]
],
[
[
"import numpy as np\nfrom numpy.random import randn\nimport pandas as pd\nfrom pandas import Series, DataFrame\n\nimport matplotlib.pyplot as plt\nfrom pylab import rcParams",
"_____no_output_____"
],
[
"%matplotlib inline\nrcParams['figure.figsize'] = 5, 4",
"_____no_output_____"
]
],
[
[
"### The simplest time series plot",
"_____no_output_____"
]
],
[
[
"import pathlib\nimport os\naddress = pathlib.Path(os.getcwd()).parent\naddress = pathlib.Path(os.path.join(address, 'Data/Superstore-Sales.csv'))",
"_____no_output_____"
],
[
"df = pd.read_csv(address, index_col='Order Date', encoding='cp1252', parse_dates=True)\ndf.head()",
"_____no_output_____"
],
[
"df['Order Quantity'].plot()",
"_____no_output_____"
],
[
"df2 = df.sample(n=100, random_state=25, axis=0)\n\nplt.xlabel('Order Date')\nplt.ylabel('Order Quantity')\nplt.title('Superstore Sales')\n\ndf2['Order Quantity'].plot()",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
4ab62c49026d8aae16346a2630feefe45b0a5e92
| 57,017 |
ipynb
|
Jupyter Notebook
|
4_convolutional_neural_network/week2_Keras+-+Tutorial+-+Happy+House+v1.ipynb
|
hongta/DeepLearningAI
|
fb6744099670351da5189fda33b14cc0a5247d7b
|
[
"MIT"
] | 2 |
2017-11-06T18:45:30.000Z
|
2018-02-08T14:42:24.000Z
|
4_convolutional_neural_network/week2_Keras+-+Tutorial+-+Happy+House+v1.ipynb
|
hongta/DeepLearningAI
|
fb6744099670351da5189fda33b14cc0a5247d7b
|
[
"MIT"
] | null | null | null |
4_convolutional_neural_network/week2_Keras+-+Tutorial+-+Happy+House+v1.ipynb
|
hongta/DeepLearningAI
|
fb6744099670351da5189fda33b14cc0a5247d7b
|
[
"MIT"
] | null | null | null | 88.39845 | 27,832 | 0.759931 |
[
[
[
"# Keras tutorial - the Happy House\n\nWelcome to the first assignment of week 2. In this assignment, you will:\n1. Learn to use Keras, a high-level neural networks API (programming framework), written in Python and capable of running on top of several lower-level frameworks including TensorFlow and CNTK. \n2. See how you can in a couple of hours build a deep learning algorithm.\n\nWhy are we using Keras? Keras was developed to enable deep learning engineers to build and experiment with different models very quickly. Just as TensorFlow is a higher-level framework than Python, Keras is an even higher-level framework and provides additional abstractions. Being able to go from idea to result with the least possible delay is key to finding good models. However, Keras is more restrictive than the lower-level frameworks, so there are some very complex models that you can implement in TensorFlow but not (without more difficulty) in Keras. That being said, Keras will work fine for many common models. \n\nIn this exercise, you'll work on the \"Happy House\" problem, which we'll explain below. Let's load the required packages and solve the problem of the Happy House!",
"_____no_output_____"
]
],
[
[
"import numpy as np\nfrom keras import layers\nfrom keras.layers import Input, Dense, Activation, ZeroPadding2D, BatchNormalization, Flatten, Conv2D\nfrom keras.layers import AveragePooling2D, MaxPooling2D, Dropout, GlobalMaxPooling2D, GlobalAveragePooling2D\nfrom keras.models import Model\nfrom keras.preprocessing import image\nfrom keras.utils import layer_utils\nfrom keras.utils.data_utils import get_file\nfrom keras.applications.imagenet_utils import preprocess_input\nimport pydot\nfrom IPython.display import SVG\nfrom keras.utils.vis_utils import model_to_dot\nfrom keras.utils import plot_model\nfrom kt_utils import *\n\nimport keras.backend as K\nK.set_image_data_format('channels_last')\nimport matplotlib.pyplot as plt\nfrom matplotlib.pyplot import imshow\n\n%matplotlib inline",
"Using TensorFlow backend.\n"
]
],
[
[
"**Note**: As you can see, we've imported a lot of functions from Keras. You can use them easily just by calling them directly in the notebook. Ex: `X = Input(...)` or `X = ZeroPadding2D(...)`.",
"_____no_output_____"
],
[
"## 1 - The Happy House \n\nFor your next vacation, you decided to spend a week with five of your friends from school. It is a very convenient house with many things to do nearby. But the most important benefit is that everybody has commited to be happy when they are in the house. So anyone wanting to enter the house must prove their current state of happiness.\n\n<img src=\"images/happy-house.jpg\" style=\"width:350px;height:270px;\">\n<caption><center> <u> <font color='purple'> **Figure 1** </u><font color='purple'> : **the Happy House**</center></caption>\n\n\nAs a deep learning expert, to make sure the \"Happy\" rule is strictly applied, you are going to build an algorithm which that uses pictures from the front door camera to check if the person is happy or not. The door should open only if the person is happy. \n\nYou have gathered pictures of your friends and yourself, taken by the front-door camera. The dataset is labbeled. \n\n<img src=\"images/house-members.png\" style=\"width:550px;height:250px;\">\n\nRun the following code to normalize the dataset and learn about its shapes.",
"_____no_output_____"
]
],
[
[
"X_train_orig, Y_train_orig, X_test_orig, Y_test_orig, classes = load_dataset()\n\n# Normalize image vectors\nX_train = X_train_orig/255.\nX_test = X_test_orig/255.\n\n# Reshape\nY_train = Y_train_orig.T\nY_test = Y_test_orig.T\n\nprint (\"number of training examples = \" + str(X_train.shape[0]))\nprint (\"number of test examples = \" + str(X_test.shape[0]))\nprint (\"X_train shape: \" + str(X_train.shape))\nprint (\"Y_train shape: \" + str(Y_train.shape))\nprint (\"X_test shape: \" + str(X_test.shape))\nprint (\"Y_test shape: \" + str(Y_test.shape))",
"number of training examples = 600\nnumber of test examples = 150\nX_train shape: (600, 64, 64, 3)\nY_train shape: (600, 1)\nX_test shape: (150, 64, 64, 3)\nY_test shape: (150, 1)\n"
]
],
[
[
"**Details of the \"Happy\" dataset**:\n- Images are of shape (64,64,3)\n- Training: 600 pictures\n- Test: 150 pictures\n\nIt is now time to solve the \"Happy\" Challenge.",
"_____no_output_____"
],
[
"## 2 - Building a model in Keras\n\nKeras is very good for rapid prototyping. In just a short time you will be able to build a model that achieves outstanding results.\n\nHere is an example of a model in Keras:\n\n```python\ndef model(input_shape):\n # Define the input placeholder as a tensor with shape input_shape. Think of this as your input image!\n X_input = Input(input_shape)\n\n # Zero-Padding: pads the border of X_input with zeroes\n X = ZeroPadding2D((3, 3))(X_input)\n\n # CONV -> BN -> RELU Block applied to X\n X = Conv2D(32, (7, 7), strides = (1, 1), name = 'conv0')(X)\n X = BatchNormalization(axis = 3, name = 'bn0')(X)\n X = Activation('relu')(X)\n\n # MAXPOOL\n X = MaxPooling2D((2, 2), name='max_pool')(X)\n\n # FLATTEN X (means convert it to a vector) + FULLYCONNECTED\n X = Flatten()(X)\n X = Dense(1, activation='sigmoid', name='fc')(X)\n\n # Create model. This creates your Keras model instance, you'll use this instance to train/test the model.\n model = Model(inputs = X_input, outputs = X, name='HappyModel')\n \n return model\n```\n\nNote that Keras uses a different convention with variable names than we've previously used with numpy and TensorFlow. In particular, rather than creating and assigning a new variable on each step of forward propagation such as `X`, `Z1`, `A1`, `Z2`, `A2`, etc. for the computations for the different layers, in Keras code each line above just reassigns `X` to a new value using `X = ...`. In other words, during each step of forward propagation, we are just writing the latest value in the commputation into the same variable `X`. The only exception was `X_input`, which we kept separate and did not overwrite, since we needed it at the end to create the Keras model instance (`model = Model(inputs = X_input, ...)` above). \n\n**Exercise**: Implement a `HappyModel()`. This assignment is more open-ended than most. We suggest that you start by implementing a model using the architecture we suggest, and run through the rest of this assignment using that as your initial model. But after that, come back and take initiative to try out other model architectures. For example, you might take inspiration from the model above, but then vary the network architecture and hyperparameters however you wish. You can also use other functions such as `AveragePooling2D()`, `GlobalMaxPooling2D()`, `Dropout()`. \n\n**Note**: You have to be careful with your data's shapes. Use what you've learned in the videos to make sure your convolutional, pooling and fully-connected layers are adapted to the volumes you're applying it to.",
"_____no_output_____"
]
],
[
[
"# GRADED FUNCTION: HappyModel\n\ndef HappyModel(input_shape):\n \"\"\"\n Implementation of the HappyModel.\n \n Arguments:\n input_shape -- shape of the images of the dataset\n\n Returns:\n model -- a Model() instance in Keras\n \"\"\"\n \n ### START CODE HERE ###\n # Feel free to use the suggested outline in the text above to get started, and run through the whole\n # exercise (including the later portions of this notebook) once. The come back also try out other\n # network architectures as well. \n X_input = Input(shape=input_shape)\n X = ZeroPadding2D((3,3))(X_input)\n \n # Conv -> BN -> ReLU\n X = Conv2D(32, (7, 7), strides=(1,1), name='conv0')(X)\n X = BatchNormalization(axis=3, name='bn0')(X)\n X = Activation('relu')(X)\n \n # Max-pool\n X = MaxPooling2D((2,2), name='max_pool')(X)\n \n X = Flatten()(X)\n X = Dense(1, activation='sigmoid', name='fc')(X)\n \n model = Model(X_input, X, name='HappyModel')\n ### END CODE HERE ###\n \n return model",
"_____no_output_____"
]
],
[
[
"You have now built a function to describe your model. To train and test this model, there are four steps in Keras:\n1. Create the model by calling the function above\n2. Compile the model by calling `model.compile(optimizer = \"...\", loss = \"...\", metrics = [\"accuracy\"])`\n3. Train the model on train data by calling `model.fit(x = ..., y = ..., epochs = ..., batch_size = ...)`\n4. Test the model on test data by calling `model.evaluate(x = ..., y = ...)`\n\nIf you want to know more about `model.compile()`, `model.fit()`, `model.evaluate()` and their arguments, refer to the official [Keras documentation](https://keras.io/models/model/).\n\n**Exercise**: Implement step 1, i.e. create the model.",
"_____no_output_____"
]
],
[
[
"### START CODE HERE ### (1 line)\nhappyModel = HappyModel((64,64,3))\n### END CODE HERE ###",
"_____no_output_____"
]
],
[
[
"**Exercise**: Implement step 2, i.e. compile the model to configure the learning process. Choose the 3 arguments of `compile()` wisely. Hint: the Happy Challenge is a binary classification problem.",
"_____no_output_____"
]
],
[
[
"### START CODE HERE ### (1 line)\nhappyModel.compile(optimizer='adam',\n loss='binary_crossentropy',\n metrics=['accuracy'])\n### END CODE HERE ###",
"_____no_output_____"
]
],
[
[
"**Exercise**: Implement step 3, i.e. train the model. Choose the number of epochs and the batch size.",
"_____no_output_____"
]
],
[
[
"### START CODE HERE ### (1 line)\nhappyModel.fit(X_train,\n Y_train,\n epochs=10,\n batch_size=32)\n### END CODE HERE ###",
"Epoch 1/10\n600/600 [==============================] - 13s - loss: 2.2303 - acc: 0.6100 \nEpoch 2/10\n600/600 [==============================] - 13s - loss: 0.6234 - acc: 0.7933 \nEpoch 3/10\n600/600 [==============================] - 13s - loss: 0.3350 - acc: 0.8683 \nEpoch 4/10\n600/600 [==============================] - 12s - loss: 0.1527 - acc: 0.9433 \nEpoch 5/10\n600/600 [==============================] - 12s - loss: 0.1657 - acc: 0.9283 \nEpoch 6/10\n600/600 [==============================] - 12s - loss: 0.0916 - acc: 0.9767 \nEpoch 7/10\n600/600 [==============================] - 13s - loss: 0.0760 - acc: 0.9833 \nEpoch 8/10\n600/600 [==============================] - 12s - loss: 0.0686 - acc: 0.9833 \nEpoch 9/10\n600/600 [==============================] - 12s - loss: 0.0720 - acc: 0.9767 \nEpoch 10/10\n600/600 [==============================] - 13s - loss: 0.0744 - acc: 0.9800 \n"
]
],
[
[
"Note that if you run `fit()` again, the `model` will continue to train with the parameters it has already learnt instead of reinitializing them.\n\n**Exercise**: Implement step 4, i.e. test/evaluate the model.",
"_____no_output_____"
]
],
[
[
"### START CODE HERE ### (1 line)\npreds = happyModel.evaluate(X_test, Y_test)\n### END CODE HERE ###\nprint()\nprint (\"Loss = \" + str(preds[0]))\nprint (\"Test Accuracy = \" + str(preds[1]))",
"150/150 [==============================] - 1s \n\nLoss = 0.245946276983\nTest Accuracy = 0.939999997616\n"
]
],
[
[
"If your `happyModel()` function worked, you should have observed much better than random-guessing (50%) accuracy on the train and test sets. To pass this assignment, you have to get at least 75% accuracy. \n\nTo give you a point of comparison, our model gets around **95% test accuracy in 40 epochs** (and 99% train accuracy) with a mini batch size of 16 and \"adam\" optimizer. But our model gets decent accuracy after just 2-5 epochs, so if you're comparing different models you can also train a variety of models on just a few epochs and see how they compare. \n\nIf you have not yet achieved 75% accuracy, here're some things you can play around with to try to achieve it:\n\n- Try using blocks of CONV->BATCHNORM->RELU such as:\n```python\nX = Conv2D(32, (3, 3), strides = (1, 1), name = 'conv0')(X)\nX = BatchNormalization(axis = 3, name = 'bn0')(X)\nX = Activation('relu')(X)\n```\nuntil your height and width dimensions are quite low and your number of channels quite large (≈32 for example). You are encoding useful information in a volume with a lot of channels. You can then flatten the volume and use a fully-connected layer.\n- You can use MAXPOOL after such blocks. It will help you lower the dimension in height and width.\n- Change your optimizer. We find Adam works well. \n- If the model is struggling to run and you get memory issues, lower your batch_size (12 is usually a good compromise)\n- Run on more epochs, until you see the train accuracy plateauing. \n\nEven if you have achieved 75% accuracy, please feel free to keep playing with your model to try to get even better results. \n\n**Note**: If you perform hyperparameter tuning on your model, the test set actually becomes a dev set, and your model might end up overfitting to the test (dev) set. But just for the purpose of this assignment, we won't worry about that here.\n",
"_____no_output_____"
],
[
"## 3 - Conclusion\n\nCongratulations, you have solved the Happy House challenge! \n\nNow, you just need to link this model to the front-door camera of your house. We unfortunately won't go into the details of how to do that here. ",
"_____no_output_____"
],
[
"<font color='blue'>\n**What we would like you to remember from this assignment:**\n- Keras is a tool we recommend for rapid prototyping. It allows you to quickly try out different model architectures. Are there any applications of deep learning to your daily life that you'd like to implement using Keras? \n- Remember how to code a model in Keras and the four steps leading to the evaluation of your model on the test set. Create->Compile->Fit/Train->Evaluate/Test.",
"_____no_output_____"
],
[
"## 4 - Test with your own image (Optional)\n\nCongratulations on finishing this assignment. You can now take a picture of your face and see if you could enter the Happy House. To do that:\n 1. Click on \"File\" in the upper bar of this notebook, then click \"Open\" to go on your Coursera Hub.\n 2. Add your image to this Jupyter Notebook's directory, in the \"images\" folder\n 3. Write your image's name in the following code\n 4. Run the code and check if the algorithm is right (0 is unhappy, 1 is happy)!\n \nThe training/test sets were quite similar; for example, all the pictures were taken against the same background (since a front door camera is always mounted in the same position). This makes the problem easier, but a model trained on this data may or may not work on your own data. But feel free to give it a try! ",
"_____no_output_____"
]
],
[
[
"### START CODE HERE ###\nimg_path = 'images/my_image.jpg'\n### END CODE HERE ###\nimg = image.load_img(img_path, target_size=(64, 64))\nimshow(img)\n\nx = image.img_to_array(img)\nx = np.expand_dims(x, axis=0)\nx = preprocess_input(x)\n\nprint(happyModel.predict(x))",
"[[ 0.]]\n"
]
],
[
[
"## 5 - Other useful functions in Keras (Optional)\n\nTwo other basic features of Keras that you'll find useful are:\n- `model.summary()`: prints the details of your layers in a table with the sizes of its inputs/outputs\n- `plot_model()`: plots your graph in a nice layout. You can even save it as \".png\" using SVG() if you'd like to share it on social media ;). It is saved in \"File\" then \"Open...\" in the upper bar of the notebook.\n\nRun the following code.",
"_____no_output_____"
]
],
[
[
"happyModel.summary()",
"_________________________________________________________________\nLayer (type) Output Shape Param # \n=================================================================\ninput_1 (InputLayer) (None, 64, 64, 3) 0 \n_________________________________________________________________\nzero_padding2d_1 (ZeroPaddin (None, 70, 70, 3) 0 \n_________________________________________________________________\nconv0 (Conv2D) (None, 64, 64, 32) 4736 \n_________________________________________________________________\nbn0 (BatchNormalization) (None, 64, 64, 32) 128 \n_________________________________________________________________\nactivation_1 (Activation) (None, 64, 64, 32) 0 \n_________________________________________________________________\nmax_pool (MaxPooling2D) (None, 32, 32, 32) 0 \n_________________________________________________________________\nflatten_1 (Flatten) (None, 32768) 0 \n_________________________________________________________________\nfc (Dense) (None, 1) 32769 \n=================================================================\nTotal params: 37,633\nTrainable params: 37,569\nNon-trainable params: 64\n_________________________________________________________________\n"
],
[
"plot_model(happyModel, to_file='HappyModel.png')\nSVG(model_to_dot(happyModel).create(prog='dot', format='svg'))",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4ab63928e70ccc22d12824950ead314faf6ae0f0
| 8,018 |
ipynb
|
Jupyter Notebook
|
test_exp_tmin.ipynb
|
etneil/model_average_paper
|
96d3572badb745107135e9b1e05fcb6966f0cd64
|
[
"MIT"
] | 2 |
2021-07-27T18:15:08.000Z
|
2022-02-18T18:20:28.000Z
|
test_exp_tmin.ipynb
|
etneil/model_average_paper
|
96d3572badb745107135e9b1e05fcb6966f0cd64
|
[
"MIT"
] | null | null | null |
test_exp_tmin.ipynb
|
etneil/model_average_paper
|
96d3572badb745107135e9b1e05fcb6966f0cd64
|
[
"MIT"
] | 2 |
2021-08-11T18:12:32.000Z
|
2021-11-18T17:29:07.000Z
| 30.957529 | 143 | 0.538289 |
[
[
[
"%matplotlib inline\n\nimport lsqfit\nfrom model_avg_paper import *\nfrom model_avg_paper.test_tmin import test_vary_tmin_SE",
"_____no_output_____"
],
[
"p0_test_ME = {\n 'A0': 2.0,\n 'E0': 0.8,\n 'A1': 10.4,\n 'E1': 1.16,\n}\nNt = 32\nnoise_params = {\n 'noise_amp': 0.3,\n 'noise_samples': 500,\n 'frac_noise': True,\n 'cross_val': False,\n 'cv_frac': 0.1,\n}\nobs_name='E0'\n\ncorrelated_data = True\nrho=0.6\n\n# Set seed for consistency of outcome\n#np.random.seed(10911) # Fig 3, subfig A; Fig 4\n#np.random.seed(81890) # Fig 3, subfig B\n#np.random.seed(87414) # Fig 3, subfig C\nnp.random.seed(77700) # Fig 3, subfig D\n\n\ndef ME_model(x,p):\n return multi_exp_model(x,p,Nexc=2)",
"_____no_output_____"
],
[
"if correlated_data:\n test_data = gen_synth_data_corr(\n np.arange(0,Nt), \n p0_test_ME, \n ME_model,\n rho=rho,\n **noise_params)\nelse:\n test_data = gen_synth_data(\n np.arange(0,Nt), \n p0_test_ME, \n ME_model,\n **noise_params)",
"_____no_output_____"
],
[
"test_res = test_vary_tmin_SE(test_data, Nt=Nt, max_tmin=26, obs_name=obs_name, IC='AIC', \n cross_val=noise_params['cross_val'])\nprint(test_res['obs_avg'])",
"_____no_output_____"
],
[
"## Figure 3\n\nimport matplotlib.ticker as ticker\n\ngs = plt.GridSpec(2, 1, height_ratios=[3,1])\ngs.update(hspace=0.06)\n\nax1 = plt.subplot(gs[0])\n\nplot_gvcorr([test_res['obs_avg']], x=np.array([1.5]), color='red', markersize=7, marker='s', open_symbol=True, label='Model avg.')\nplot_gvcorr(test_res['obs'], x=test_res['tmin'], label='Individual fits')\n\nax1.plot(np.array([-1,34]), 0*np.array([0,0])+p0_test_ME[obs_name], linestyle='--', color='k', label='Model truth')\n#ax1.set_xlabel('$N_p$')\nax1.set_ylabel('$E_0$')\n\nax1.legend(loc='center left', bbox_to_anchor=(1,0.5))\nax1.set_xlim(0.7,27.3)\n\nplt.setp(ax1.get_xticklabels(), visible=False)\n\nax2 = plt.subplot(gs[1])\n\np_norm = test_res['probs'] / np.sum(test_res['probs'])\nQ_norm = test_res['Qs'] / np.sum(test_res['Qs'])\nplt.plot(test_res['tmin'], p_norm, color='orange', label='pr$(M|D)$')\nplt.plot(test_res['tmin'], Q_norm, color='blue', linestyle='-.', label='Fit p-value') # Note: fit prob != model prob! \n\ntick_spacing = 4\nax2.xaxis.set_major_locator(ticker.MultipleLocator(tick_spacing))\nplt.yticks([0,np.max(p_norm)])\nax2.yaxis.set_major_formatter(ticker.FuncFormatter(lambda x, pos: '0' if x == 0 else '{:.2f}'.format(x)))\nax2.set_xlim(0.7,27.3)\n\n\n\n\n# Put a legend to the right of the current axis\nax2.legend(loc='center left', bbox_to_anchor=(1, 0.5))\nax2.set_xlabel(r'$t_{\\rm min}$')\nax2.set_ylabel('p')\n\n# Uncomment to save figure to disk\n#plt.savefig('plots/exp_avg_4.pdf', bbox_inches = \"tight\")\n",
"_____no_output_____"
],
[
"# Scaling w/number of samples\nNsamp_array = np.array([20, 40, 80, 160, 320, 640, 2040, 4096, 4096*2, 4096*4])\nNsamp_max = Nsamp_array[-1]\n\nnoise_params['noise_samples'] = Nsamp_max\nif correlated_data:\n scale_data = gen_synth_data_corr(\n np.arange(0,Nt), \n p0_test_ME, \n ME_model,\n rho=rho,\n **noise_params) \nelse:\n scale_data = gen_synth_data(\n np.arange(0,Nt), \n p0_test_ME, \n ME_model,\n **noise_params)\n\nmodel_avg_vs_Nsamp = []\nnaive_avg_vs_Nsamp = []\nfixed_tmin_vs_Nsamp = []\nfixed_tmin_2_vs_Nsamp = []\nfw_vs_Nsamp = []\n\nfix_tmin = 14\nfix_tmin_2 = 8\n\n\nfor Nsamp in Nsamp_array:\n test_data_scale = cut_synth_data_Nsamp(scale_data, Nsamp)\n test_res_scale = test_vary_tmin_SE(test_data_scale, Nt=Nt, max_tmin=Nt-4, obs_name=obs_name, IC='AIC')\n test_res_scale_naive = test_vary_tmin_SE(test_data_scale, Nt=Nt, max_tmin=Nt-4, obs_name=obs_name, \n IC='naive') \n \n model_avg_vs_Nsamp.append(test_res_scale['obs_avg'])\n naive_avg_vs_Nsamp.append(test_res_scale_naive['obs_avg'])\n fixed_tmin_vs_Nsamp.append(test_res_scale['obs'][fix_tmin])\n fixed_tmin_2_vs_Nsamp.append(test_res_scale['obs'][fix_tmin_2])\n fw_vs_Nsamp.append(obs_avg_full_width(test_res_scale['obs'], test_res_scale['Qs'], test_res_scale['fits'], bf_i=None))\n\n\n\n\n",
"_____no_output_____"
],
[
"## Figure 4\n\nplot_gvcorr(model_avg_vs_Nsamp, x=np.log(Nsamp_array)+0.1, label='Model avg. (AIC)')\nplot_gvcorr(fixed_tmin_vs_Nsamp, x=np.log(Nsamp_array)+0.2, color='red', marker='s', markersize=6, label=r'Fixed $t_{\\rm min} = 14$')\nplot_gvcorr(fw_vs_Nsamp, x=np.log(Nsamp_array)+0.3, marker='X', markersize=8, color='orange', label='Full-width systematic')\nplot_gvcorr(naive_avg_vs_Nsamp, x=np.log(Nsamp_array)+0.4, color='silver', marker='v', markersize=8, label=r'Model avg. (naive)')\n\nplt.plot(np.arange(0,10), 0*np.arange(0,10)+p0_test_ME[obs_name], linestyle='--', color='k', label='Model truth')\nplt.xlabel(r'$\\log(N_s)$')\nplt.ylabel(r'$E_0$')\nplt.xlim(2.7,7.)\nplt.ylim(0.78,0.82)\n\n\n# Put a legend to the right of the current axis\nax = plt.subplot(111)\nax.legend(loc='center left', bbox_to_anchor=(1, 0.5))\n\n# Uncomment to save figure to disk\n#plt.savefig('plots/exp_N_scaling.pdf', bbox_inches = \"tight\")\n",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab63ae008b948abe9464ef3f5b2130b0343ef08
| 18,229 |
ipynb
|
Jupyter Notebook
|
13 - Classes.ipynb
|
prabal5ghosh/python-primer
|
0ab2faefa952113a2dceb14f3243b1ec59e886e1
|
[
"MIT"
] | 2 |
2020-04-07T13:23:54.000Z
|
2020-04-07T13:23:57.000Z
|
13 - Classes.ipynb
|
B2BDA/python-primer
|
0ab2faefa952113a2dceb14f3243b1ec59e886e1
|
[
"MIT"
] | null | null | null |
13 - Classes.ipynb
|
B2BDA/python-primer
|
0ab2faefa952113a2dceb14f3243b1ec59e886e1
|
[
"MIT"
] | null | null | null | 25.247922 | 596 | 0.553952 |
[
[
[
"### Classes",
"_____no_output_____"
],
[
"Finally we get to classes.\n\nI assume you already have some knowledge of classes and OOP in general, so I'll focus on the semantics of creating classes and some of the differences with Java classes.",
"_____no_output_____"
],
[
"First, the question of visibility. There is no such thing as private or public in Python. Everything is public, period. So we don't have to specify the visibility of functions and attributes in Python.",
"_____no_output_____"
],
[
"Class instantiations are done in two steps - the instance is created first, and then the instance is initialized. In general we hook into the initialization phase and leave the object creation alone. We do this by using a special method in the class called `__init__`. We'll see how large a role special methods play in Python.",
"_____no_output_____"
],
[
"The important thing to note is that by the time `__init__` is called in our class, the object (instance) has **already** been created.",
"_____no_output_____"
],
[
"We use the `class` keyword to create classes:",
"_____no_output_____"
]
],
[
[
"class Person:\n pass",
"_____no_output_____"
]
],
[
[
"`pass` is something we can use in Python to indicate \"do nothing\" (a so called \"no-op\" operation). Here I use it to supply a body for the class definition, but don't actually want to specify any functionality.",
"_____no_output_____"
],
[
"So now we can actually create instances of the `Person` object at this point. They will be pretty useless since we have not implemented any functionality yet.",
"_____no_output_____"
],
[
"We create instances of classes by **calling** the class - remember how we call things in Python, we use `()`:",
"_____no_output_____"
]
],
[
[
"p = Person()",
"_____no_output_____"
],
[
"id(p), type(p)",
"_____no_output_____"
]
],
[
[
"So now we may want to add some functions to the class. Whenever we define a function in a class, we have to understand what happens when we call that function from an instance, using dot notation:",
"_____no_output_____"
],
[
"For example, if we write \n\n```p.say_hello()```\n\nthen we are calling the `say_hello()` function from the instance, and Python will **bind** that function to the specific instance - i.e. it creates an association between the instance used to call the function, and the function. \n\nThe way it does this is by passing in the instance reference to the function as the first positional argument - in this case it would actually call `say_hello(p)`. And our `say_hello` function now has access to the instance it was called from, including any internal state.\n\nWhen functions are bound to an instance, they are called **methods** of the class, and, in particular **instance methods** because they are bound to instances of the class when called. (There are other types of functions that can be bound to the class, called *class methods*, but this is beyond the scope of this primer).",
"_____no_output_____"
],
[
"Let's see how this works:",
"_____no_output_____"
]
],
[
[
"class Person:\n def say_hello(instance, name):\n return f'{instance} says hello to {name}'",
"_____no_output_____"
]
],
[
[
"So here, we had to make sure the first argument of our function was created to receive the instance it is being called from. After that we are free to add our own arguments as needed.",
"_____no_output_____"
]
],
[
[
"p = Person()",
"_____no_output_____"
]
],
[
[
"Let's see what `p` looks like when we print it:",
"_____no_output_____"
]
],
[
[
"p",
"_____no_output_____"
]
],
[
[
"And now let's call the `say_hello` method from the instance `p`:",
"_____no_output_____"
]
],
[
[
"p.say_hello('Alex')",
"_____no_output_____"
]
],
[
[
"You'll notice that we did not pass `p` as the first argument to `say_hello` - Python did that for us since we wrote `p.say_hello`.",
"_____no_output_____"
],
[
"By convention, that `instance` argument I wrote above, is usually named `self`, for obvious reasons. But it is just a convention, although one you should stick to.",
"_____no_output_____"
]
],
[
[
"class Person:\n def say_hello(self, name):\n return f'{self} says hello to {name}'",
"_____no_output_____"
],
[
"p = Person()\np.say_hello('Alex')",
"_____no_output_____"
]
],
[
[
"So now let's turn our attention to instance attributes. ",
"_____no_output_____"
],
[
"Python is dynamic, so instance attributes do not have to be defined at the time the class is created. In fact we rarely do so.\n\nLet's see how we can add an attribute to an instance after it's been created:",
"_____no_output_____"
]
],
[
[
"p = Person()",
"_____no_output_____"
],
[
"p.name = 'Alex'",
"_____no_output_____"
]
],
[
[
"That's it, `p` now has an attribute called `name`:",
"_____no_output_____"
]
],
[
[
"p.name",
"_____no_output_____"
]
],
[
[
"But this is specific to this instance, not other instances:",
"_____no_output_____"
]
],
[
[
"p2 = Person()\np2.name",
"_____no_output_____"
]
],
[
[
"So instance attributes are **specific** to the instance (hence the name).\n\nOf course we can define attributes by calling methods in our class - let's see an example of this:",
"_____no_output_____"
]
],
[
[
"class Person:\n def set_name(self, name):\n self.name = name\n \n def get_name(self):\n return self.name",
"_____no_output_____"
],
[
"p = Person()",
"_____no_output_____"
]
],
[
[
"At this point `p` does **not** have a `name` attribute (it hasn't been set yet!):",
"_____no_output_____"
]
],
[
[
"p.get_name()",
"_____no_output_____"
]
],
[
[
"But we can easily set it using the `set_name` method:",
"_____no_output_____"
]
],
[
[
"p.set_name('Alex')",
"_____no_output_____"
],
[
"p.get_name()",
"_____no_output_____"
]
],
[
[
"And of course the attribute is called `name` and is easily accessible directly as well:",
"_____no_output_____"
]
],
[
[
"p.name",
"_____no_output_____"
]
],
[
[
"This is what is called a *bare* attribute - it is not hidden by getter and setter methods like we would normally do in Java (remember we do not have private variables).\n\nYou'll notice the issue we had - we would get an exception if we tried to access the attribute before it was actually created.\n\nFor this reason, best practice is to create these instance attributes (even setting them to a default value or `None`) when the class instance is being created.\n\nThe best place to do this is in the *initialization* phase of the class (remember that class instantiation has two phases - creation and initialization).",
"_____no_output_____"
],
[
"To do this we use the special method `__init__`. \n\nThis is going to be a functionm in our class, and will be bound to the instance when it is called (by that time the instance has already been created), so just like our `set_name` method, we'll need to allow for the instance to be received as the first argument:",
"_____no_output_____"
]
],
[
[
"class Person:\n def __init__(self, name):\n self.name = name",
"_____no_output_____"
]
],
[
[
"So the `__init__` method is basically doing the same thing as our `set_name` method - the difference is in how it is called.\n\nWhen we create an instance using `Person()`, Python looks for, and if available calls, the `__init__` method (that's why it's called a *special method*).\n\n\nIn our case here, the first argument will receive the just created object, but we have one additional argument, `name`. So we need to pass that in when we create the instance:",
"_____no_output_____"
]
],
[
[
"p = Person('name')",
"_____no_output_____"
]
],
[
[
"The `__init__` method was actually called - let's see this:",
"_____no_output_____"
]
],
[
[
"class Person:\n def __init__(self, name):\n print(f'__init__ called for object {self}')\n self.name = name",
"_____no_output_____"
],
[
"p = Person('Alex')",
"__init__ called for object <__main__.Person object at 0x1081fe320>\n"
]
],
[
[
"And in fact, the memory address of `p` is:",
"_____no_output_____"
]
],
[
[
"hex(id(p))",
"_____no_output_____"
]
],
[
[
"which as you can see, is excactly the same object `self` was set to when `__init__` was called.",
"_____no_output_____"
],
[
"And our instance `p` now has an attribute called `name`:",
"_____no_output_____"
]
],
[
[
"p.name",
"_____no_output_____"
]
],
[
[
"We can create another instance:",
"_____no_output_____"
]
],
[
[
"p2 = Person('Eric')",
"__init__ called for object <__main__.Person object at 0x1081fe0b8>\n"
],
[
"hex(id(p2)), p2.name",
"_____no_output_____"
]
],
[
[
"And that has not affected the `name` of `p` - since `name` is an instance attribute (it is specific to the instance):",
"_____no_output_____"
]
],
[
[
"p.name",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ab67a139c049271f7bec9fb723a7b5365649b7f
| 43,747 |
ipynb
|
Jupyter Notebook
|
assignments/seminars/Seminar_1_python_&_numpy.ipynb
|
AMSolin/dlcourse_ai
|
f719c98ab33f268b760f269229638da99af30b61
|
[
"MIT"
] | null | null | null |
assignments/seminars/Seminar_1_python_&_numpy.ipynb
|
AMSolin/dlcourse_ai
|
f719c98ab33f268b760f269229638da99af30b61
|
[
"MIT"
] | null | null | null |
assignments/seminars/Seminar_1_python_&_numpy.ipynb
|
AMSolin/dlcourse_ai
|
f719c98ab33f268b760f269229638da99af30b61
|
[
"MIT"
] | null | null | null | 21.624815 | 141 | 0.459506 |
[
[
[
"# Массивы",
"_____no_output_____"
]
],
[
[
"import numpy\nimport numpy as np",
"_____no_output_____"
],
[
"a = numpy.zeros((2, 3), dtype=numpy.float32) # ничего не вывело",
"_____no_output_____"
],
[
"a # выведет а, по оси Y левая координата (внешняя), по оси X правая (внутренняя) ",
"_____no_output_____"
],
[
"a; # не выведет а",
"_____no_output_____"
],
[
"type(a)",
"_____no_output_____"
],
[
"a.__class__",
"_____no_output_____"
],
[
"a = numpy.zeros((2, 3, 4), dtype=numpy.float32);\na # выведет новый а, по оси X правая координата, по оси Y -- внешние координаты",
"_____no_output_____"
],
[
"a += 1; \ndisplay(a); \na *=2; \ndisplay(a); # сложение со скаляром, умножение на скаляр; display -- вывод с помощью jupyter (colab)",
"_____no_output_____"
],
[
"ones = lambda *shape: numpy.ones(shape, dtype=numpy.float32)",
"_____no_output_____"
],
[
"ones(2,2,2)",
"_____no_output_____"
]
],
[
[
"# Задача",
"_____no_output_____"
]
],
[
[
"\"\"\"\nЗадача: \nвывести красиво операцию A + B = C на экран\n\"\"\"\nmy_input = [ones(2, 3), '+', ones(2, 3), '=', ones(2,3)*2]\nprint(my_input)\nprint(*my_input)\ndisplay(*my_input)",
"[array([[1., 1., 1.],\n [1., 1., 1.]], dtype=float32), '+', array([[1., 1., 1.],\n [1., 1., 1.]], dtype=float32), '=', array([[2., 2., 2.],\n [2., 2., 2.]], dtype=float32)]\n[[1. 1. 1.]\n [1. 1. 1.]] + [[1. 1. 1.]\n [1. 1. 1.]] = [[2. 2. 2.]\n [2. 2. 2.]]\n"
],
[
"# пусть наша функция имеет такой формат:\ndef pp(*lst):\n pass\npp(*my_input)",
"_____no_output_____"
],
[
"lines = [repr(a).split('\\n') for a in my_input]; lines",
"_____no_output_____"
],
[
"lines = [str(a).split('\\n') for a in my_input]; lines",
"_____no_output_____"
],
[
"max_lines = max([len(line) for line in lines])\nmax_lines",
"_____no_output_____"
],
[
"lines = [line+['']*(max_lines-len(line)) for line in lines]; lines",
"_____no_output_____"
]
],
[
[
"### транспонирование, reshape, операция получения максимума",
"_____no_output_____"
]
],
[
[
"lines_arr = numpy.array(lines); lines_arr.T",
"_____no_output_____"
],
[
"widths = [len(x) for x in lines]; display(widths)",
"_____no_output_____"
],
[
"lines_arr.shape, lines_arr.flatten().shape",
"_____no_output_____"
],
[
"widths = numpy.array([len(x) for x in lines_arr.flatten()]); display(widths)\nwidths = widths.reshape(len(lines_arr), -1); display(widths)",
"_____no_output_____"
],
[
"widths = widths.reshape(*lines_arr.shape); display(widths)",
"_____no_output_____"
],
[
"numpy.max(widths)",
"_____no_output_____"
],
[
"widths",
"_____no_output_____"
],
[
"widths.max()",
"_____no_output_____"
],
[
"widths.max(axis=1) # складывает по внутренней оси, ",
"_____no_output_____"
],
[
"line_widths = widths.max(axis=1)\nline_widths",
"_____no_output_____"
]
],
[
[
"### форматирование всё равно придётся делать поэлементно. зашли в тупик...",
"_____no_output_____"
]
],
[
[
"lines = [str(a).split('\\n') for a in my_input]; lines\nmax_lines = max([len(line) for line in lines])\nmax_lines",
"_____no_output_____"
],
[
"def fmt_item(lines, max_lines=0):\n max_width = max([len(line) for line in lines])\n empty = ' '*max_width\n lines = [line.ljust(max_width) for line in lines]\n lines += [empty] * (max_lines - len(lines))\n return lines \n \nresults = fmt_item(str(numpy.arange(1, 7).reshape(2,-1)).split('\\n'), max_lines=3)\nprint('\\n'.join(results))",
"[[1 2 3] \n [4 5 6]]\n \n"
],
[
"# а что делать дальше, когда у нас есть такие массивы? \n# воспользуемся array().T",
"_____no_output_____"
],
[
"def pp(*lst):\n lines = [str(item).split('\\n') for item in lst]\n max_lines = max([len(item) for item in lines])\n lines = [fmt_item(item, max_lines=max_lines) for item in lines]\n lines_t = numpy.array(lines).T\n print('\\n'.join([' '.join(line) for line in lines_t]))\n \npp(*my_input)\n",
"[[1. 1. 1.] + [[1. 1. 1.] = [[2. 2. 2.] \n [1. 1. 1.]] [1. 1. 1.]] [2. 2. 2.]]\n"
]
],
[
[
"### Попробуем нашу функцию:",
"_____no_output_____"
]
],
[
[
"A = numpy.arange(1, 11).reshape(5,2)\nB = numpy.linspace(10, 30, 10).reshape(5,2)\nR = A * B\npp(A, '*', B, '=', R)",
"[[ 1 2] * [[10. 12.22222222] = [[ 10. 24.44444444] \n [ 3 4] [14.44444444 16.66666667] [ 43.33333333 66.66666667] \n [ 5 6] [18.88888889 21.11111111] [ 94.44444444 126.66666667] \n [ 7 8] [23.33333333 25.55555556] [163.33333333 204.44444444] \n [ 9 10]] [27.77777778 30. ]] [250. 300. ]]\n"
],
[
"import operator\n#operator.add = lambda x, y: x + y\n\ndef op(x, name, y, func):\n r = func(x, y)\n pp(x, name, y, '=', r)\n \nop(numpy.array([[1,2],[3,4]]), '+', numpy.array([[5,6],[7,8]]), operator.add)",
"[[1 2] + [[5 6] = [[ 6 8] \n [3 4]] [7 8]] [10 12]]\n"
],
[
"# вспомним классы, оформим в виде класса\nclass Op:\n def __init__(self, name, func):\n self.name = name\n self.func = func\n \n def __call__(self, x, y):\n r = self.func(x, y)\n pp(x, self.name, y, '=', r)\n\nSSum = Op('+', lambda x, y: x + y)\nSMul = Op('*', lambda x, y: x * y)\nMMul = Op('@', lambda x, y: x @ y) # python 3.5+ \n\nSMul(A, B)",
"[[ 1 2] * [[10. 12.22222222] = [[ 10. 24.44444444] \n [ 3 4] [14.44444444 16.66666667] [ 43.33333333 66.66666667] \n [ 5 6] [18.88888889 21.11111111] [ 94.44444444 126.66666667] \n [ 7 8] [23.33333333 25.55555556] [163.33333333 204.44444444] \n [ 9 10]] [27.77777778 30. ]] [250. 300. ]]\n"
],
[
"MMul(A.T, B)",
"[[ 1 3 5 7 9] @ [[10. 12.22222222] = [[561.11111111 616.66666667] \n [ 2 4 6 8 10]] [14.44444444 16.66666667] [655.55555556 722.22222222]]\n [18.88888889 21.11111111] \n [23.33333333 25.55555556] \n [27.77777778 30. ]] \n"
],
[
"tr = numpy.array([[9,8],[7,6],[5,4]])\nte = numpy.array([[1,2],[3,4]])",
"_____no_output_____"
],
[
"num_train = tr.shape[0]\nnum_test = te.shape[0]\n\ndists = np.zeros((num_test, num_train), np.float32)\npp(dists)\nfor i_test in range(num_test):\n for i_train in range(num_train):\n dists[i_test, i_train] = np.abs(np.add.reduce(te[i_test] - tr[i_train]))\n pp(dists[i_test, i_train])\npp(dists)",
"[[0. 0. 0.] \n [0. 0. 0.]]\n14.0\n10.0\n6.0\n10.0\n6.0\n2.0\n[[14. 10. 6.] \n [10. 6. 2.]]\n"
],
[
"num_train = tr.shape[0]\nnum_test = te.shape[0]\n\ndists = np.zeros((num_test, num_train), np.float32)\npp(dists)\nfor i_test in range(num_test):\n dists[i_test] = np.sum(np.abs(te[i_test]- tr), axis=1)\npp(dists)",
"[[0. 0. 0.] \n [0. 0. 0.]]\n[[14. 10. 6.] \n [10. 6. 2.]]\n"
],
[
"new_te = te[:, np.newaxis]\ndists = np.sum(np.abs(new_te - tr), axis=2)\ndists",
"_____no_output_____"
],
[
"for i in range(dists.shape[0]):\n print(np.sort(dists[i])[:3])",
"[ 6 10 14]\n[ 2 6 10]\n"
],
[
"a = np.array([9, 4, 4, 3, 3, 9, 0, 4, 6, 0])\nb = np.array([False, False, False, False, False, False, False, False, False, False])\n\nind = np.argpartition(a, 1)#[:3]\nind #[6, 9, 4]\n#b[ind] #array([ True, False, True])\n#c = np.bincount(b[ind]) # array([1, 2]\n#np.argmax(c)",
"_____no_output_____"
],
[
"tr",
"_____no_output_____"
],
[
"a = np.array_split(tr, 3)\nfor i in range(len(a)):\n b = np.delete(a.copy(), i, 0)\n print(b, ',')\na =np.concatenate(a)\na",
"[[[7 6]]\n\n [[5 4]]] ,\n[[[9 8]]\n\n [[5 4]]] ,\n[[[9 8]]\n\n [[7 6]]] ,\n"
],
[
"# 0, 1, 2, 3, 4, 5, 6, 7\narr = np.array([1100, 1, 9, 2, 0, 17, 17, 5])\nc = np.bincount(arr)\nnp.argmax(c)",
"_____no_output_____"
],
[
"tr",
"_____no_output_____"
],
[
"tr_n = tr - np",
"_____no_output_____"
]
],
[
[
"# Простая индексация",
"_____no_output_____"
]
],
[
[
"pred = np.zeros(5, np.bool)\npred",
"_____no_output_____"
],
[
"pp(B[0, :], B[-1,-2], B[-2:-1, 0])",
"_____no_output_____"
]
],
[
[
"# Broadcasting и stack",
"_____no_output_____"
]
],
[
[
"SSum(A, B[0]) # что произойдёт? это преобразование называется broadcasting",
"_____no_output_____"
],
[
"SMul(A, B[0]) # смотрим ещё раз, на умножении",
"_____no_output_____"
],
[
"SMul(A, numpy.stack(list(B[0:1])*5))",
"_____no_output_____"
]
],
[
[
"# Бинарные и логические операции с массивами",
"_____no_output_____"
]
],
[
[
"pp(A, A>3, B, B>=10.) # сравнение float -- дело неточное",
"_____no_output_____"
],
[
"1<<23",
"_____no_output_____"
]
],
[
[
"Float 32 format: 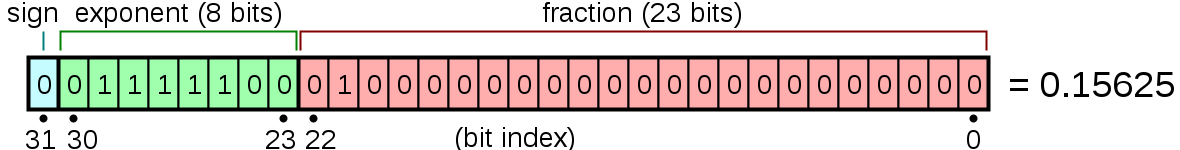 ",
"_____no_output_____"
]
],
[
[
"pp(numpy.isclose(B, 10), (10 - 1e-7 <= B) & (B <= 10 + 1e-7))",
"_____no_output_____"
],
[
"idx = numpy.isclose(B, 10) | numpy.isclose(B, 30)\nB[idx]",
"_____no_output_____"
]
],
[
[
"# Индексация массивов",
"_____no_output_____"
]
],
[
[
"pp(B)\npp(numpy.isclose(B[1:-1, 1:-1], 10)) # классическая индексация питона\npp(numpy.isclose(B[0], 10)) # берём первую строку\npp(numpy.isclose(B[:, 1], 10)) # берём второй столбец",
"_____no_output_____"
]
],
[
[
"# Cлучайные значения",
"_____no_output_____"
]
],
[
[
"pp(numpy.random.uniform(0, 1), numpy.random.randn(2, 3), numpy.random.rand(2, 3))",
"_____no_output_____"
],
[
"import random\nrandom.seed(10)\nnumpy.random.seed(10)\nnr = numpy.random.uniform(0, 1)\nnr2 = numpy.random.uniform(0, 1)\nrr = random.uniform(0, 1)\nrr2 = random.uniform(0, 1)\npp(rr, nr, rr2, nr2)\n\nrandom.seed(10)\nnumpy.random.seed(10)\nrr = random.uniform(0, 1)\nnr = numpy.random.uniform(0, 1)\nrr2 = random.uniform(0, 1)\nnr2 = numpy.random.uniform(0, 1)\npp(rr, nr, rr2, nr2)",
"_____no_output_____"
],
[
"import matplotlib\n# значение по умолчанию в colab, но не в локальных ноутбуках\nmatplotlib.rcParams['axes.grid'] = True\n\nimport matplotlib.pyplot as plt\n# нарисуем график для 11 случайных нормально распределённых чисел.\nx = numpy.linspace(0, 10, 11)\ndots = numpy.random.randn(11)\nplt.plot(x, dots);",
"_____no_output_____"
],
[
"import matplotlib\nmatplotlib.rcParams['axes.grid'] = False # убираем белую клетку (настройки colab)\n\nimport matplotlib.pyplot as plt\nplt.plot(x, dots); # рисуем линию ещё раз",
"_____no_output_____"
],
[
"# если надо просто вывести точки, без линий между ними\nplt.scatter(x, dots);",
"_____no_output_____"
],
[
"# выводим двумерный массив, используя цветовую схему 'hot'.\n# matplotlib автоматически определяет минимум и максимум\n# по нему настраивает цвета\nplt.imshow(numpy.arange(0, 120).reshape(6, 20), 'hot');",
"_____no_output_____"
]
],
[
[
"# Строки и словари",
"_____no_output_____"
]
],
[
[
"# Задача: определить, какой символ встречается чаще всего в данном тексте,\n# не считая пробелов!\na = \"Кот сидел на крыше тише мыши\"\npp(set(list(a)))",
"_____no_output_____"
],
[
"from collections import Counter\n# приближённое решение.\nCounter(list(a)).most_common()\n# убрать пробел и будет то, что надо. попробуйте сами.",
"_____no_output_____"
],
[
"# напишем вручную код аналогичный тому, что происходит внутри Counter\ncounter = {}\nfor ch in list(a.lower()):\n if ch == ' ': continue\n if ch in counter:\n counter[ch] += 1\n else:\n counter[ch] = 1\n \npp(counter)",
"_____no_output_____"
],
[
"pp(type(ch)) # в питоне даже отдельные символы -- это строчки",
"_____no_output_____"
],
[
"max(counter.values()) # нашли максимум",
"_____no_output_____"
],
[
"# алгоритм для поиска максимума\nwinner_ch = None\nwinner_count = 0\nfor ch, count in counter.items():\n if count > winner_count:\n winner_count = count\n winner_ch = ch\nprint(winner_ch, winner_count) # и вот наш символ-победитель\n# имейте в виду, победителя на самом деле три.\n# исправьте код, чтобы вывести и второй и третий.",
"_____no_output_____"
]
],
[
[
" ---",
"_____no_output_____"
],
[
"---",
"_____no_output_____"
],
[
"---",
"_____no_output_____"
]
],
[
[
"A",
"_____no_output_____"
],
[
"a=A[0]\nb=B[0]\nc=B[0].T\nprint(a.shape, b.shape, c.shape)\npp(a, ' ', b, ' ', a+b)",
"_____no_output_____"
],
[
"a=A[0:1]\nb=B[0:1]\nc=B[0:1].T\npp('shapes:', a.shape, b.shape, c.shape)\npp('arrays:', a, ' ', c, ' ', a+c)",
"_____no_output_____"
],
[
"a = A[0]\nb = A[0].reshape(-1, 1)\nc = A[0][:,None] # or [:, numpy.newaxis]\nd = A[0][None, :] # or [numpy.newaxis, :]\npp(a.shape, ' ', b.shape, ' ', c.shape, ' ', d.shape)\nprint()\npp(a, ' ', b, ' ', c, ' ', d)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
4ab68a61cbcdff488b4496c67749b149cde4619a
| 80,633 |
ipynb
|
Jupyter Notebook
|
docs/source/do_why_estimation_methods.ipynb
|
djinnome/dowhy
|
3b9132d7cc394f6237b1bb6a80857bc38610ff94
|
[
"MIT"
] | null | null | null |
docs/source/do_why_estimation_methods.ipynb
|
djinnome/dowhy
|
3b9132d7cc394f6237b1bb6a80857bc38610ff94
|
[
"MIT"
] | null | null | null |
docs/source/do_why_estimation_methods.ipynb
|
djinnome/dowhy
|
3b9132d7cc394f6237b1bb6a80857bc38610ff94
|
[
"MIT"
] | null | null | null | 123.292049 | 59,892 | 0.856969 |
[
[
[
"# DoWhy: Different estimation methods for causal inference\nThis is quick introduction to DoWhy causal inference library.\nWe will load in a sample dataset and use different methods for estimating causal effect from a (pre-specified)treatment variable to a (pre-specified) outcome variable.\n\nFirst, let us add required path for python to find DoWhy code and load required packages.",
"_____no_output_____"
]
],
[
[
"import os, sys\nsys.path.append(os.path.abspath(\"../../\"))",
"_____no_output_____"
],
[
"import numpy as np\nimport pandas as pd\nimport logging\n\nimport dowhy\nfrom dowhy.do_why import CausalModel\nimport dowhy.datasets ",
"_____no_output_____"
]
],
[
[
"Let us first load a dataset. For simplicity, we simulate a dataset with linear relationships between common causes and treatment, and common causes and outcome. \n\nBeta is the true causal effect. ",
"_____no_output_____"
]
],
[
[
"data = dowhy.datasets.linear_dataset(beta=10,\n num_common_causes=5, \n num_instruments = 2,\n num_samples=10000,\n treatment_is_binary=True)\ndf = data[\"df\"]\n",
"_____no_output_____"
]
],
[
[
"Note that we are using a pandas dataframe to load the data.",
"_____no_output_____"
],
[
"## Identifying the causal estimand",
"_____no_output_____"
],
[
"We now input a causal graph in the DOT graph format.",
"_____no_output_____"
]
],
[
[
"# With graph\nmodel=CausalModel(\n data = df,\n treatment=data[\"treatment_name\"],\n outcome=data[\"outcome_name\"],\n graph=data[\"dot_graph\"],\n instruments=data[\"instrument_names\"],\n logging_level = logging.INFO\n )",
"Model to find the causal effect of treatment v on outcome y\n"
],
[
"model.view_model()",
"_____no_output_____"
],
[
"from IPython.display import Image, display\ndisplay(Image(filename=\"causal_model.png\"))",
"_____no_output_____"
]
],
[
[
"We get a causal graph. Now identification and estimation is done. ",
"_____no_output_____"
]
],
[
[
"identified_estimand = model.identify_effect()\nprint(identified_estimand)",
"INFO:dowhy.causal_identifier:Common causes of treatment and outcome:{'X3', 'Z1', 'U', 'X2', 'X0', 'X4', 'X1', 'Z0'}\n"
]
],
[
[
"## Method 1: Regression\n\nUse linear regression.",
"_____no_output_____"
]
],
[
[
"causal_estimate_reg = model.estimate_effect(identified_estimand,\n method_name=\"backdoor.linear_regression\",\n test_significance=True)\nprint(causal_estimate_reg)\nprint(\"Causal Estimate is \" + str(causal_estimate_reg.value))",
"LinearRegressionEstimator\n"
]
],
[
[
"## Method 2: Stratification\n\nWe will be using propensity scores to stratify units in the data.",
"_____no_output_____"
]
],
[
[
"causal_estimate_strat = model.estimate_effect(identified_estimand,\n method_name=\"backdoor.propensity_score_stratification\")\nprint(causal_estimate_strat)\nprint(\"Causal Estimate is \" + str(causal_estimate_strat.value))",
"INFO:dowhy.causal_estimator:INFO: Using Propensity Score Stratification Estimator\nINFO:dowhy.causal_estimator:b: y~v+X3+Z1+X2+X0+X4+X1+Z0\n"
]
],
[
[
"## Method 3: Matching\n\nWe will be using propensity scores to match units in the data.",
"_____no_output_____"
]
],
[
[
"causal_estimate_match = model.estimate_effect(identified_estimand,\n method_name=\"backdoor.propensity_score_matching\")\nprint(causal_estimate_match)\nprint(\"Causal Estimate is \" + str(causal_estimate_match.value))",
"INFO:dowhy.causal_estimator:INFO: Using Propensity Score Matching Estimator\nINFO:dowhy.causal_estimator:b: y~v+X3+Z1+X2+X0+X4+X1+Z0\n"
]
],
[
[
"## Method 4: Weighting\n\nWe will be using (inverse) propensity scores to assign weights to units in the data.",
"_____no_output_____"
]
],
[
[
"causal_estimate_ipw = model.estimate_effect(identified_estimand,\n method_name=\"backdoor.propensity_score_weighting\")\nprint(causal_estimate_ipw)\nprint(\"Causal Estimate is \" + str(causal_estimate_ipw.value))",
"INFO:dowhy.causal_estimator:INFO: Using Propensity Score Weighting Estimator\nINFO:dowhy.causal_estimator:b: y~v+X3+Z1+X2+X0+X4+X1+Z0\n"
]
],
[
[
"## Method 5: Instrumental Variable\n\nWe will be using Wald estimator for the provided instrumental variable.",
"_____no_output_____"
]
],
[
[
"causal_estimate_iv = model.estimate_effect(identified_estimand,\n method_name=\"iv.instrumental_variable\", method_params={'iv_instrument_name':'Z1'})\nprint(causal_estimate_iv)\nprint(\"Causal Estimate is \" + str(causal_estimate_iv.value))",
"INFO:dowhy.causal_estimator:INFO: Using Instrumental Variable Estimator\nINFO:dowhy.causal_estimator:Realized estimand: Wald Estimator\nRealized estimand type: ate\nEstimand expression:\n -1\nExpectation(Derivative(y, Z0))⋅Expectation(Derivative(v, Z0)) \nEstimand assumption 1, Exclusion: If we remove {Z0,Z1}→v, then ¬(Z0,Z1→y)\nEstimand assumption 2, outcome_effect_homogeneity: Each unit's outcome y isaffected in the same way by common causes of v and y\nEstimand assumption 3, As-if-random: If U→→y then ¬(U →→Z0,Z1)\nEstimand assumption 4, treatment_effect_homogeneity: Each unit's treatment v isaffected in the same way by common causes of v and y\n\n"
]
],
[
[
"## Method 6: Regression Discontinuity\n\nWe will be internally converting this to an equivalent instrumental variables problem.",
"_____no_output_____"
]
],
[
[
"causal_estimate_regdist = model.estimate_effect(identified_estimand,\n method_name=\"iv.regression_discontinuity\", \n method_params={'rd_variable_name':'Z1',\n 'rd_threshold_value':0.5,\n 'rd_bandwidth': 0.1})\nprint(causal_estimate_regdist)\nprint(\"Causal Estimate is \" + str(causal_estimate_regdist.value))",
"INFO:dowhy.causal_estimator:Using Regression Discontinuity Estimator\nINFO:dowhy.causal_estimator:\n"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ab68ae3908a236c8f280666aedf4b826bd0ad2c
| 1,608 |
ipynb
|
Jupyter Notebook
|
create_Threads.ipynb
|
PAIDI-RAMESH/Basic-python-threads
|
29e185978a360862576b2e007f2717cb0b8770de
|
[
"Unlicense"
] | null | null | null |
create_Threads.ipynb
|
PAIDI-RAMESH/Basic-python-threads
|
29e185978a360862576b2e007f2717cb0b8770de
|
[
"Unlicense"
] | null | null | null |
create_Threads.ipynb
|
PAIDI-RAMESH/Basic-python-threads
|
29e185978a360862576b2e007f2717cb0b8770de
|
[
"Unlicense"
] | null | null | null | 16.926316 | 58 | 0.468905 |
[
[
[
"# Importing Packages",
"_____no_output_____"
]
],
[
[
"import threading",
"_____no_output_____"
],
[
"def worker(num):\n # Threaded Worker function\n print(num)\n return",
"_____no_output_____"
]
],
[
[
"# Creating array and appending threads into array",
"_____no_output_____"
]
],
[
[
"threads = []",
"_____no_output_____"
],
[
"for i in range(5):\n t = threading.Thread(target=worker,args=(i,))\n threads.append(t)\n # Run the thread\n t.start()",
"01\n2\n\n3\n4\n"
]
]
] |
[
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4ab694425723e00e2e91ccfb540b28d29665ef43
| 34,056 |
ipynb
|
Jupyter Notebook
|
01_Pre_processing.ipynb
|
martinaviggiano/knowledge_extraction_project
|
7ddfa26f1f9387728d6d4a0bec7223bb0e4ce110
|
[
"BSD-3-Clause"
] | null | null | null |
01_Pre_processing.ipynb
|
martinaviggiano/knowledge_extraction_project
|
7ddfa26f1f9387728d6d4a0bec7223bb0e4ce110
|
[
"BSD-3-Clause"
] | null | null | null |
01_Pre_processing.ipynb
|
martinaviggiano/knowledge_extraction_project
|
7ddfa26f1f9387728d6d4a0bec7223bb0e4ce110
|
[
"BSD-3-Clause"
] | null | null | null | 34,056 | 34,056 | 0.644585 |
[
[
[
"# **01_PREPROCESSING**",
"_____no_output_____"
],
[
"Summary:\n\n\n1. Import and Normalization\n2. Split Opinions into Subjects of Interest\n3. Text Cleaning\n4. Split into Sentences\n\n\n\n",
"_____no_output_____"
],
[
"---",
"_____no_output_____"
]
],
[
[
"from google.colab import drive\ndrive.mount('/content/drive')",
"_____no_output_____"
],
[
"import sys\nsys.path.append('/content/drive/My Drive/Università/inforet_prj/')",
"_____no_output_____"
],
[
"!pip install -U spacy unidecode",
"_____no_output_____"
],
[
"!python -m spacy download en_core_web_sm",
"_____no_output_____"
],
[
"import lzma, json\nimport pandas as pd\nimport numpy as np\nimport pickle\nfrom tqdm import tqdm\nimport spacy\nimport string\nimport seaborn as sns\nimport matplotlib.pyplot as plt\nfrom nltk.corpus import stopwords\nfrom nltk.stem import PorterStemmer\nimport re\nfrom unidecode import unidecode\nimport nltk\nfrom nltk.tokenize import word_tokenize, sent_tokenize\nnltk.download('punkt')\nsns.set()\ntqdm.pandas()",
"_____no_output_____"
],
[
"nlp = spacy.load(\"en_core_web_sm\")",
"_____no_output_____"
]
],
[
[
"## **1. Import and Normalization**",
"_____no_output_____"
],
[
"### *1.1 Data Import*\n\n\n**NB**: run the 3 cells below only if on Google Colab. Otherwise skip them and download the compressed data manually from https://api.case.law/v1/bulk/22341/download/",
"_____no_output_____"
]
],
[
[
"!pip install selenium\n!apt-get update # to update ubuntu to correctly run apt install\n!apt install chromium-chromedriver\n!cp /usr/lib/chromium-browser/chromedriver /usr/bin\nimport sys\nsys.path.insert(0,'/usr/lib/chromium-browser/chromedriver')\nfrom selenium import webdriver\nchrome_options = webdriver.ChromeOptions()\nchrome_options.add_argument('--headless')\nchrome_options.add_argument('--no-sandbox')\nchrome_options.add_argument('--disable-dev-shm-usage')\nwd = webdriver.Chrome('chromedriver', options=chrome_options)\nwd.get(\"https://case.law/bulk/download/\")",
"_____no_output_____"
],
[
"wd.find_element_by_xpath(\"/html/body/div/main/div/div/div[2]/div/div[2]/div/div[2]/a\").click()",
"_____no_output_____"
],
[
"!unzip Illinois-20200302-text.zip\n!mv Illinois-20200302-text/data/data.jsonl.xz data.jsonl.xz\n!rm -r Illinois-20200302-text\n!rm Illinois-20200302-text.zip",
"_____no_output_____"
]
],
[
[
"### *1.2 Data Normalization*",
"_____no_output_____"
],
[
"Creation of opinions, citations and df",
"_____no_output_____"
]
],
[
[
"# We know that there will be 183146 items,\n# so we set this manually since tqdm will not\n# be able to display a progress bar when reading from\n# a file.\npbar = tqdm(total=183146)\n\n# Read directly from the compressed file.\n# We will create a list where each element is a line\n# of the file, which in turns is a json\n# (casted in python as a dict).\nwith lzma.open(\"data.jsonl.xz\") as f:\n cases = []\n\n for line in f:\n cases.append(json.loads(str(line, 'utf8')))\n pbar.update(1)\n\n pbar.close()",
"_____no_output_____"
],
[
"# https://pandas.pydata.org/docs/reference/api/pandas.json_normalize.html\ndf = pd.json_normalize(cases)",
"_____no_output_____"
],
[
"del cases",
"_____no_output_____"
],
[
"# Flattens the list of attorneys to a single string\n# with ; as separator\ndf[\"casebody.data.attorneys\"] = df.apply(lambda x: \"; \".join(x[\"casebody.data.attorneys\"]), axis=1)",
"_____no_output_____"
],
[
"\"\"\"\nEach element of the columns 'citations' and 'casebody.data.opinions' is\na list, and in turn each element of the list is a json object.\nThis means that we need to unravel those column to have a more \"flatten\"\nversion (like a simple table, eg. a DataFrame).\nThe approach shown here consists of creating two different DataFrames\nthat will contain data from the two columns. In order to preserve the\nassociation of each row of the new DataFrame with the corresponding data\nin the original DataFrame, we will add to each json a new key called \"id\"\nthat will have the original row number as value.\n\"\"\"\n\ndef add_id_todict(x, col):\n vals = x[col]\n\n for i, elem in enumerate(vals):\n d = elem\n d[\"id\"] = x.name\n vals[i] = d\n\n return vals",
"_____no_output_____"
],
[
"df[\"casebody.data.opinions\"] = df.apply(lambda x: add_id_todict(x, \"casebody.data.opinions\"), axis=1)\ndf[\"citations\"] = df.apply(lambda x: add_id_todict(x, \"citations\"), axis=1)",
"_____no_output_____"
],
[
"# For clarity, let's also add the \"id\" column to the original df\ndf[\"id\"] = df.index.values",
"_____no_output_____"
],
[
"# We merge each element in the \"citations\" column (which is a list)\n# to a single list called \"citations\".\n#\n# Using list comprehension instead of df[\"column\"].sum()\n# because the latter is slow for large df.\ncitations = [item for x in df[\"citations\"] for item in x]\ndf.drop(columns=[\"citations\"], inplace=True)",
"_____no_output_____"
],
[
"# Same for the opinions column\nopinions = [item for x in df[\"casebody.data.opinions\"] for item in x]\ndf.drop(columns=[\"casebody.data.opinions\"], inplace=True)",
"_____no_output_____"
],
[
"# Let's now get the flattened table from the citations\n# and from the opinions\ncitations_df = pd.json_normalize(citations)",
"_____no_output_____"
],
[
"opinions_df = pd.json_normalize(opinions)",
"_____no_output_____"
]
],
[
[
"We now have 3 dataframes that can be joined using the \"id\" column.",
"_____no_output_____"
]
],
[
[
"df['year'] = pd.to_datetime(df['decision_date']).apply(lambda x: x.year)\nopinions_df = pd.merge(opinions_df, df[['year','id']], on=\"id\", how=\"left\")",
"_____no_output_____"
]
],
[
[
"### *1.3 Serialize data*\n",
"_____no_output_____"
]
],
[
[
"with open(\"/content/drive/MyDrive/Università/inforet_prj/df.pkl\", \"wb\") as f:\n pickle.dump(df, f)",
"_____no_output_____"
],
[
"with open(\"/content/drive/MyDrive/Università/inforet_prj/citations.pkl\", \"wb\") as f:\n pickle.dump(citations_df, f)",
"_____no_output_____"
],
[
"with open(\"/content/drive/MyDrive/Università/inforet_prj/opinions.pkl\", \"wb\") as f:\n pickle.dump(opinions_df, f)",
"_____no_output_____"
],
[
"del df\ndel citations_df\ndel opinions\ndel citations\ndel opinions_df",
"_____no_output_____"
],
[
"import gc\ngc.collect()",
"_____no_output_____"
]
],
[
[
"---",
"_____no_output_____"
],
[
"## **2. Split Opinions into Subjects of Interest**\n\nWe divide into 3 groups rows based on the lists of terms provided for each subject of interest: narcotics, weapons and investigation.",
"_____no_output_____"
]
],
[
[
"with open(\"/content/drive/MyDrive/Università/inforet_prj/opinions.pkl\", \"rb\") as f:\n opinions_df = pickle.load(f)",
"_____no_output_____"
],
[
"opinions_df[\"text\"] = opinions_df[\"text\"].str.replace(\"|\", \" \")",
"_____no_output_____"
],
[
"opinions_df.author = opinions_df.author.fillna(\"\")\narray = opinions_df[\"author\"].progress_apply(lambda x: nltk.word_tokenize(x.lower()))\n\nauthors_judges = []\n\nfor op in array:\n for token in op:\n if token.isalpha() and len(token) > 1:\n authors_judges.append(token)\n\nauthors_judges = set(authors_judges)",
"_____no_output_____"
],
[
"with open(\"authors_judges.pkl\", \"wb\") as f:\n pickle.dump(authors_judges, f)",
"_____no_output_____"
],
[
"!cp authors_judges.pkl /content/drive/MyDrive/Università/inforet_prj",
"_____no_output_____"
],
[
"def typo(text):\n cleaned_text = (\n text.replace('cannabi ','cannabis ')\n .replace('lysergic acid diethylamide', 'lsd')\n .replace('methylenedioxymethamphetamine', 'mdma')\n .replace('MDMA', 'mdma')\n .replace('methylenedioxyamphetamine', 'mda')\n .replace('ciacetyl','diacetyl')\n .replace(' nar cotic', ' narcotic')\n .replace(' fi ','')\n )\n return cleaned_text",
"_____no_output_____"
],
[
"opinions_df['text'] = opinions_df.text.progress_apply(lambda x: typo(x))",
"_____no_output_____"
],
[
"narcotics = [\"cannabis\", \"marijuana\", \"lsd\", \"heroin\", 'methaqualone', \"ecstasy\", \"mdma\", \"cocaine\", \"cocaine\", \"methamphetamine\", \"hydromorphone\", \"dilaudid\", \"meperidine\", \"demerol\", \"oxycodone\", \"dexedrine\", \"fentanyl\", \"ritalin\", \"methadone\", \"amphetamine\", \"phencyclidine\", \"ephedrine\"]\nweapons = [\"gun\", \"knife\", \"weapon\", \"firearm\", \"rifle\", \"carabine\", \"shotgun\", \"assaults rifle\", \"sword\", \"blunt objects\"]\ninvestigations = [\"gang\", \"mafia\", \"serial killer\", \"rape\", \"thefts\", \"recidivism\", \"arrest\", \"ethnicity\", \"caucasian\", \"afroamerican\", \"native american\", \"hispanic\", \"gender\", \"male\", \"female\", \"man\", \"woman\", \"girl\", \"boy\", \"robbery\", \"cybercrime\"]",
"_____no_output_____"
],
[
"narco_df = opinions_df.loc[opinions_df['text'].str.contains(\"|\".join(narcotics)).any(level=0)] # 35410 rows / 6076 / 11038",
"_____no_output_____"
],
[
"narco_df.to_csv(\"narco_df.csv\", index=False, sep=\"|\")",
"_____no_output_____"
],
[
"!cp narco_df.csv /content/drive/MyDrive/Università/inforet_prj",
"_____no_output_____"
],
[
"del opinions_df\ndel authors_judges",
"_____no_output_____"
],
[
"import gc",
"_____no_output_____"
],
[
"gc.collect()",
"_____no_output_____"
]
],
[
[
"---",
"_____no_output_____"
],
[
"## **3. Text Cleaning**\nLoad Opinions from the previous step.",
"_____no_output_____"
]
],
[
[
"with open(\"/content/drive/MyDrive/Università/inforet_prj/authors_judges.pkl\", \"rb\") as f:\n authors_judges = pickle.load(f)",
"_____no_output_____"
],
[
"# Proper nouns found in the dataset\nnames = [\"Brinks\", \"Flores\", \"People v.\",\"Pinnix\", \"Garvey\", \"Steinbach\", \"Fowlar\", \"Mobil\", \"Milian\", \"TQ\", \"Yanez\", \"Tawanda\", \"Geder\", \"Mason\", \"Payne\", \"Bair\", \"ILCS\", \"tbe\", \"tbat\", \"Delores\",\"Stivers\", \"Spades\", \"Snyders\", \"Nally\", \"Budaj\", \"Yacoo\", \"Cosgrove\", \"Cos-grove\", \"Gayles\", \"Hodges\"]",
"_____no_output_____"
],
[
"def full_text_clean(text, is_sentence=False):\n if text == '' or pd.isna(text) or not isinstance(text, str):\n return ''\n\n bb = (\n text.replace(' U.S. ','US')\n .replace(' S.Ct. ','SCt')\n .replace(' f. supp. ', ' fsupp ')\n .replace(' cir.', ' cir ')\n .replace(\"[o]\", \"o\")\n .replace(\" CIR \", \" confidential source \")\n .replace(\"Reg.\", \" regulation \")\n .replace(\"miIe\", \" mile \")\n .replace(\" com mitted \", \" committed \")\n .replace(\"wtap\", \"tap\")\n )\n \n if bb.strip() == '' or pd.isna(bb):\n return ''\n\n if not ' ' in bb:\n return ''\n\n temp = bb.split()\n bb = \" \".join([ele for ele in temp if not ele[0].isupper()])\n \n if not is_sentence:\n bb = bb.split(\":\")\n if len(bb) > 1:\n bb.pop(0)\n bb = ' '.join(bb)\n\n if bb.strip() == '' or pd.isna(bb):\n return ''\n\n bb = unidecode(re.sub(' +', ' ', bb.strip())) #any additional whitespaces and foreign characters\n bb = bb.strip()\n bb = re.sub('[0-9]{1,2} [Uu]\\.[Ss]\\.[Cc]\\. §\\s?\\d+(\\w+)?( \\([0-9]{4}\\))?',' USCCITATION ', bb)\n bb = re.sub('[a-zA-Z]+ [vV]\\. [a-zA-Z]+',' CaseAvCaseB ', bb) #CaseA v. CaseB = CaseAvCaseB\n bb = re.sub('\\d+ (Ark|Ill)\\. \\d+',' StateCase ', bb) #300 Ark. 230 = 300Ark230\n bb = re.sub(' [Ss][Tt][Aa][Tt][Ss]\\.',' StateCase2 ',bb) #300 Ark. 230 = 300Ark230\n bb = re.sub('\\d+ [A-z]+\\.[ ]*[A-z]+\\.[ ]*\\d[A-z]+ \\d+',' CaseRef ',bb) #953 S.W.2d 559 or 87 L.Ed.2d 481\n bb = re.sub('[Jj][Rr]\\.', 'Jr ', bb)\n bb = re.sub('\\d+ (Ark|Ill)\\. App. \\d+',' StateAppCase ', bb)\n bb = re.sub('(Ark|Ill)\\. Code Ann\\. § ',' StateCodeSection ', bb)\n bb = re.sub(' [Ii][Dd]\\.',' Idem ', bb)\n bb = re.sub('§+',' Section ', bb)\n bb = re.sub('[Aa][Nn][Nn][Oo][:.]* \\d+ [Aa]\\.*[ ]*[Ll]\\.*[ ]*[Rr]\\.*[ ]*\\d+','anno', bb)\n bb = re.sub(' [Aa][Nn][Nn][Oo][:.]*',' anno', bb)\n bb = re.sub('[Cc][Ff]\\.','cf', bb)\n bb = re.sub(' [Rr][Ee][Vv]\\. [Ss][Tt][Aa][Tt]\\.',' revstat ', bb)\n bb = re.sub('[ \\d]+[Pp][Aa][Rr]\\.',' par ', bb)\n bb = re.sub('[ \\d]+[Ss][Tt][Aa][Tt]\\.',' stat ', bb)\n bb = re.sub(\"[\\(\\[].*?[\\)\\]]\", \"\", bb)\n\n \n bb = (\n bb.replace(\"USCCITATION\", \"\")\n .replace(\"CaseAvCaseB\", \"\")\n .replace(\"StateCase\", \"\")\n .replace(\"StateCase2\", \"\")\n .replace(\"CaseRef\", \"\")\n .replace(\"StateAppCase\", \"\")\n .replace(\"StateCodeSection\", \"\")\n .replace(\"anno\", \"\")\n )\n\n bb = unidecode(re.sub(' +', ' ', bb.strip()))\n bb = bb.strip()\n\n if bb.strip() == '' or pd.isna(bb):\n return '' \n\n doc = nlp(bb)\n persons = set([str(ent.text).lower() for ent in doc.ents if ent.label_ == \"PERSON\"])\n persons = [x.translate(str.maketrans('', '', string.punctuation)) for x in set(nltk.word_tokenize(\" \".join(persons)))]\n persons.extend(names)\n\n result = []\n for token in doc:\n if (len(token.text) > 1 \n and token.text.isalpha() # Token is word\n and token.pos_ not in ['NUM', 'PROPN'] # Token not NUM, PROPN nor ADV,, , 'ADV', 'PRON', 'CONJ'\n and not token.is_punct # Token not punctuation\n and not token.is_stop # Token not punctuation\n and token.text not in authors_judges # Token is not a judge\n and token.text not in persons # Token is not a persona name\n ):\n\n result.append(token.lemma_.lower())\n \n # Our result is a string of the form:\n # \"text lemma POS; text lemma POS; text lemma POS; ...\"\n result = \" \".join(result)\n \n return result",
"_____no_output_____"
],
[
"# 1 H\nwith open(\"narco_nlp_21set_nostop.csv\", \"w\") as my_empty_csv:\n pass\n\npbar = tqdm(total=6076 ) # narco_df total rows\nchunksize = 1\n\nfor chunk in pd.read_csv(\"narco_df.csv\", chunksize=chunksize, sep=\"|\", usecols=[\"text\"]):\n chunk['spacy_nlp'] = chunk.apply(lambda row: full_text_clean(row[\"text\"]), axis=1)\n chunk.drop(columns=[\"text\"], inplace=True)\n chunk.to_csv(\"narco_nlp_21set_nostop.csv\", index=False, sep=\"|\", mode=\"a\", header=False)\n\n pbar.update(1)\n\npbar.close()",
"_____no_output_____"
],
[
"!cp narco_nlp_21set_nostop.csv /content/drive/MyDrive/Università/inforet_prj",
"_____no_output_____"
]
],
[
[
"Check that the cleaning worked properly.",
"_____no_output_____"
]
],
[
[
"narco_nlp = pd.read_csv(\n \"/content/drive/MyDrive/Università/inforet_prj/narco_nlp_21set_nostop.csv\",\n sep=\"|\",\n names=['spacy_nlp'],\n header=None\n)",
"_____no_output_____"
],
[
"assert narco_nlp.shape[0] > 0",
"_____no_output_____"
],
[
"assert narco_nlp.loc[pd.isna(narco_nlp.spacy_nlp)].shape[0] == 0",
"_____no_output_____"
]
],
[
[
"## **4. Split into Sentences**",
"_____no_output_____"
]
],
[
[
"opinions_df = pd.read_csv(\"/content/drive/MyDrive/Università/inforet_prj/narco_df.csv\", sep=\"|\")",
"_____no_output_____"
],
[
"narco_pmi_ = opinions_df.loc[:, [ \"text\"]]",
"_____no_output_____"
],
[
"narco_pmi_[\"sentences\"] = narco_pmi_.text.progress_apply(lambda x: sent_tokenize(x))",
"100%|██████████| 6076/6076 [01:07<00:00, 89.42it/s]\n"
],
[
"narco_sentences = narco_pmi_.explode('sentences')",
"_____no_output_____"
],
[
"narco_sentences = narco_sentences.drop(columns=[\"text\"]).reset_index().rename(columns={\"index\": \"opinion_id\"})",
"_____no_output_____"
],
[
"narco_sentences.to_csv(\"narco_sentences.csv\", index=False, sep=\"|\")",
"_____no_output_____"
]
],
[
[
"---",
"_____no_output_____"
]
],
[
[
"opinions_df = pd.read_csv(\"/content/drive/MyDrive/Università/inforet_prj/narco_df.csv\", sep=\"|\")",
"_____no_output_____"
],
[
"schedule_1 = [\"cannabis\", \"marijuana\", \"lsd\", \"heroin\", 'methaqualone', \"ecstasy\", \"peyote\", \"mescaline\", \"mda\", \"mdma\"] #https://www.dea.gov/drug-information/drug-scheduling\nschedule_2 = [\"cocaine\", \"methamphetamine\", \"hydromorphone\", \"dilaudid\", \"meperidine\", \"demerol\", \"oxycodone\", \"dexedrine\", \"fentanyl\", \"ritalin\", \"methadone\", \"amphetamine\", \"phencyclidine\", \"pseudoephedrine\", \"ephedrine\", \"meth\", \"opium\", \"dilaudid\", \"preludin\"]\nschedule_3 = [\"ketamine\", \"anabolic\" , \"steroids\", \"testosterone\", \"ketamine\"]\nschedule_4 = [\"modafinil\", \"provigil\", \"adderall\", \"methylphenidate\", \"memantine\", \"axura\", \"soma\", \"xanax\", \"darvon\", \"darvocet\", \"valium\", \"ativan\", \"talwin\", \"ambien\", \"tramadol\", \"ethclorvynol\"]\nschedule_5 = [\"phenylpropanolamine\", \"lomotil\", \"motofen\", \"lyrica\", \"parepectolin\", \"tetracaine\"]",
"_____no_output_____"
],
[
"conditions = [\n (opinions_df['text'].str.contains(\"|\".join(schedule_1))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_3))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_2))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_4))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_5))),\n (opinions_df['text'].str.contains(\"|\".join(schedule_2))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_3))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_1))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_4))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_5))),\n (opinions_df['text'].str.contains(\"|\".join(schedule_3))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_1))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_2))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_4))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_5))),\n (opinions_df['text'].str.contains(\"|\".join(schedule_4))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_3))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_2))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_1))) & ~(opinions_df['text'].str.contains(\"|\".join(schedule_5)))\n ]\n\n# create a list of the values we want to assign for each condition\nvalues = ['narco_1', 'narco_2', 'narco_3', 'narco_4']\n\n# create a new column and use np.select to assign values to it using our lists as arguments\nopinions_df['schedule'] = np.select(conditions, values)\n\n# display updated DataFrame\nopinions_df.head()",
"_____no_output_____"
],
[
"narco_pmi = opinions_df.loc[:, [\"schedule\", \"text\"]]",
"_____no_output_____"
],
[
"narco_1_pmi = narco_pmi.loc[narco_pmi['schedule'] == 'narco_1'] # 1969\nnarco_2_pmi = narco_pmi.loc[narco_pmi['schedule'] == 'narco_2'] # 1782 ",
"_____no_output_____"
],
[
"narco_1_pmi[\"sentences\"] = narco_1_pmi.text.progress_apply(lambda x: sent_tokenize(x)) \nnarco_2_pmi[\"sentences\"] = narco_2_pmi.text.progress_apply(lambda x: sent_tokenize(x))",
"_____no_output_____"
],
[
"narco_sentences_1 = narco_1_pmi.explode('sentences')\nnarco_sentences_2 = narco_2_pmi.explode('sentences')",
"_____no_output_____"
],
[
"narco_sentences_1 = narco_sentences_1.drop(columns=[\"text\"]).reset_index().rename(columns={\"index\": \"opinion_id\"})\nnarco_sentences_2 = narco_sentences_2.drop(columns=[\"text\"]).reset_index().rename(columns={\"index\": \"opinion_id\"})",
"_____no_output_____"
],
[
"narco_sentences_1.to_csv(\"narco_sentences_1.csv\", index=False, sep=\"|\")\nnarco_sentences_2.to_csv(\"narco_sentences_2.csv\", index=False, sep=\"|\")",
"_____no_output_____"
],
[
"!cp narco_sentences_1.csv /content/drive/MyDrive/Università/inforet_prj # 239155",
"_____no_output_____"
],
[
"!cp narco_sentences_2.csv /content/drive/MyDrive/Università/inforet_prj # 325030",
"_____no_output_____"
]
],
[
[
"---",
"_____no_output_____"
]
],
[
[
"narco_sentences = pd.read_csv(\"/content/drive/MyDrive/Università/inforet_prj/narco_sentences.csv\", sep=\"|\")",
"_____no_output_____"
],
[
"narco_sentences = pd.read_csv(\"/content/drive/MyDrive/Università/inforet_prj/narco_sentences.csv\", sep=\"|\")\nnarco_sentences_1 = pd.read_csv(\"/content/drive/MyDrive/Università/inforet_prj/narco_sentences_1.csv\", sep=\"|\")\nnarco_sentences_2 = pd.read_csv(\"/content/drive/MyDrive/Università/inforet_prj/narco_sentences_2.csv\", sep=\"|\")",
"_____no_output_____"
],
[
"names = [\"Brinks\", \"Flores\", \"People v.\",\"Pinnix\", \"Garvey\", \"Steinbach\", \"Fowlar\", \"Mobil\", \"Milian\", \"TQ\", \"Yanez\", \"Tawanda\", \"Geder\", \"Mason\", \"Payne\", \"Bair\", \"ILCS\", \"tbe\", \"tbat\", \"Delores\",\"Stivers\", \"Spades\", \"Snyders\", \"Nally\", \"Budaj\", \"Yacoo\", \"Cosgrove\", \"Cos-grove\", \"Gayles\", \"Hodges\"]",
"_____no_output_____"
],
[
"# 5 H\nwith open(\"narco_pmi_nlp.csv\", \"w\") as my_empty_csv:\n pass\n\npbar = tqdm(total=1056006) # narco_sentences total rows 1056006\nchunksize = 1\n\nfor chunk in pd.read_csv(\"/content/drive/MyDrive/Università/inforet_prj/narco_sentences.csv\", chunksize=chunksize, sep=\"|\", usecols=[\"sentences\"]):\n chunk['sent_clean'] = chunk.apply(lambda row: full_text_clean(row[\"sentences\"], is_sentence=True), axis=1)\n chunk.drop(columns=[\"sentences\"], inplace=True)\n chunk.to_csv(\"narco_pmi_nlp.csv\", index=False, sep=\"|\", mode=\"a\", header=False)\n\n pbar.update(1)\n\npbar.close()",
"100%|██████████| 1056006/1056006 [5:06:51<00:00, 57.35it/s]\n"
],
[
"!cp narco_pmi_nlp.csv /content/drive/MyDrive/Università/inforet_prj",
"_____no_output_____"
],
[
"# 1 H\nwith open(\"narco_1_pmi_nlp.csv\", \"w\") as my_empty_csv:\n pass\n\npbar = tqdm(total=239155) # narco_sentences total rows 239155\nchunksize = 1\n\nfor chunk in pd.read_csv(\"/content/drive/MyDrive/Università/inforet_prj/narco_sentences_1.csv\", chunksize=chunksize, sep=\"|\", usecols=[\"sentences\"]):\n chunk['sent_clean'] = chunk.apply(lambda row: full_text_clean(row[\"sentences\"], is_sentence=True), axis=1)\n chunk.drop(columns=[\"sentences\"], inplace=True)\n chunk.to_csv(\"narco_1_pmi_nlp.csv\", index=False, sep=\"|\", mode=\"a\", header=False)\n\n pbar.update(1)\n\npbar.close()",
"_____no_output_____"
],
[
"!cp narco_1_pmi_nlp.csv /content/drive/MyDrive/Università/inforet_prj",
"_____no_output_____"
],
[
"# 1 H\nwith open(\"narco_2_pmi_nlp.csv\", \"w\") as my_empty_csv:\n pass\n\npbar = tqdm(total=325030) # narco_sentences total rows 325030\nchunksize = 1\n\nfor chunk in pd.read_csv(\"/content/drive/MyDrive/Università/inforet_prj/narco_sentences_2.csv\", chunksize=chunksize, sep=\"|\", usecols=[\"sentences\"]):\n chunk['sent_clean'] = chunk.apply(lambda row: full_text_clean(row[\"sentences\"], is_sentence=True), axis=1)\n chunk.drop(columns=[\"sentences\"], inplace=True)\n chunk.to_csv(\"narco_2_pmi_nlp.csv\", index=False, sep=\"|\", mode=\"a\", header=False)\n\n pbar.update(1)\n\npbar.close()",
"_____no_output_____"
],
[
"!cp narco_2_pmi_nlp.csv /content/drive/MyDrive/Università/inforet_prj",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab6990b4789e23a25b59815740f2226038a621b
| 39,052 |
ipynb
|
Jupyter Notebook
|
result/generate_img.ipynb
|
samcw/nanodet
|
dc7c4f6021199d6988221b516d49af392a52d748
|
[
"Apache-2.0"
] | null | null | null |
result/generate_img.ipynb
|
samcw/nanodet
|
dc7c4f6021199d6988221b516d49af392a52d748
|
[
"Apache-2.0"
] | null | null | null |
result/generate_img.ipynb
|
samcw/nanodet
|
dc7c4f6021199d6988221b516d49af392a52d748
|
[
"Apache-2.0"
] | null | null | null | 300.4 | 35,836 | 0.927993 |
[
[
[
"import pandas as pd\nimport matplotlib.pyplot as plt\nimport json\nimport os\nfrom matplotlib import rcParams\n\nconfig = {\n \"font.size\": 13,\n 'font.sans-serif': \"Times New Roman\"\n}\n\nrcParams.update(config)",
"_____no_output_____"
],
[
"def loadJSON(dir): \n with open(dir, 'r', encoding='utf8') as fp:\n json_data = json.load(fp)\n return json_data\n\ndef loadData():\n listdir = ['ap_l', 'ap_m', 'ap_s', 'mAP']\n content = []\n for item in listdir:\n _list = os.listdir(item)\n json_data = []\n for file in _list:\n data = loadJSON('./' + item + '/' + file)\n json_data.append(data)\n content.append(json_data)\n return content\n\ndef formateData(data):\n result = []\n for item in data:\n content = []\n for _item in item:\n content.append(_item[2])\n result.append(content)\n return result\n\ntableData = loadData()",
"_____no_output_____"
],
[
"def drawPic(data):\n epoch = []\n for i in range(20):\n epoch.append((i + 1) * 10)\n plt.plot(epoch, data[1], linewidth=0.5, color='b', label='pre', marker=\"v\")\n plt.plot(epoch, data[2], linewidth=0.5, color='y', label='post', marker=\"^\")\n plt.plot(epoch, data[3], linewidth=0.5, color='g', label='residual', marker=\"<\")\n plt.plot(epoch, data[4], linewidth=0.5, color='r', label='identity', marker=\">\")\n plt.xlabel('Epochs')\n plt.ylabel('Precision')\n plt.title('')\n plt.legend()\n \n plt.savefig('map.png', dpi=256)\n\nformatedData = formateData(tableData[3])\ndrawPic(formatedData)",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code"
]
] |
4ab6a45eb541f36a04669e17b3540e1fedf7dc76
| 148,522 |
ipynb
|
Jupyter Notebook
|
2. Wasserstein GAN-GP (Gradient Penalty)/walking in the latent space.ipynb
|
hee9joon/Face-Generation
|
caa9b4e0bb61e77ee6d32fc8687bad63f998ec9c
|
[
"MIT"
] | 14 |
2020-10-06T11:58:02.000Z
|
2022-02-21T13:20:32.000Z
|
2. Wasserstein GAN-GP (Gradient Penalty)/walking in the latent space.ipynb
|
ShambhaviCodes/Face-Generation
|
caa9b4e0bb61e77ee6d32fc8687bad63f998ec9c
|
[
"MIT"
] | null | null | null |
2. Wasserstein GAN-GP (Gradient Penalty)/walking in the latent space.ipynb
|
ShambhaviCodes/Face-Generation
|
caa9b4e0bb61e77ee6d32fc8687bad63f998ec9c
|
[
"MIT"
] | 4 |
2021-03-07T06:47:23.000Z
|
2021-11-27T06:01:58.000Z
| 707.247619 | 143,460 | 0.943544 |
[
[
[
"## Walking in the Latent Space\n\n### 1. Import Libraries",
"_____no_output_____"
]
],
[
[
"import torch\n\nimport os\nimport numpy as np\nfrom matplotlib import pyplot as plt\n\nfrom config import *\nfrom models import Generator",
"_____no_output_____"
]
],
[
[
"### 2. Make a Directory",
"_____no_output_____"
]
],
[
[
"interpolation_path = './results/interpolation'\n\nif not os.path.exists(interpolation_path):\n os.makedirs(interpolation_path)",
"_____no_output_____"
]
],
[
[
"### 3. Device Configuration",
"_____no_output_____"
]
],
[
[
"device = 'cuda' if torch.cuda.is_available() else 'cpu'",
"_____no_output_____"
]
],
[
[
"### 4. Prepare Generator",
"_____no_output_____"
]
],
[
[
"path = os.path.join(config.weights_path, 'Face_Generator_Epoch_{}.pkl'.format(config.num_epochs))",
"_____no_output_____"
],
[
"G = Generator().to(device)\nG.load_state_dict(torch.load(path))\nG.eval()",
"_____no_output_____"
]
],
[
[
"### 5. Walking in the Latent Space\n#### 5.1 Noise",
"_____no_output_____"
]
],
[
[
"z_1 = torch.empty(1, config.noise_dim, 1, 1).uniform_(-1, 1).to(device)\nz_2 = torch.empty(1, config.noise_dim, 1, 1).uniform_(-1, 1).to(device)\n\nlatent_space = np.linspace(0, 1, 10)",
"_____no_output_____"
]
],
[
[
"#### 5.2 Plot and Save",
"_____no_output_____"
]
],
[
[
"fig, axs = plt.subplots(nrows=1, ncols=10, figsize=(20, 2), constrained_layout=True)\n\nfor ax, alpha in zip(axs.flat, latent_space):\n ax.axis('off')\n \n z = alpha * z_1 + (1-alpha) * z_2\n \n image = G(z)\n image = image.detach().cpu().numpy().squeeze()\n image = image.transpose(1, 2, 0)\n image = (image+1)/2\n ax.imshow(image)\n \nfig.savefig(os.path.join(interpolation_path, './Generated_Face_Interpolation.png'))",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ab6ac6404ed0f3011cbf850ac45d0e9d074a894
| 31,974 |
ipynb
|
Jupyter Notebook
|
3_Training_a_Model.ipynb
|
6ones/plagiarism-detection
|
b92cb9852d2396337ef42b96b606af5288934381
|
[
"MIT"
] | null | null | null |
3_Training_a_Model.ipynb
|
6ones/plagiarism-detection
|
b92cb9852d2396337ef42b96b606af5288934381
|
[
"MIT"
] | null | null | null |
3_Training_a_Model.ipynb
|
6ones/plagiarism-detection
|
b92cb9852d2396337ef42b96b606af5288934381
|
[
"MIT"
] | null | null | null | 43.208108 | 1,211 | 0.622693 |
[
[
[
"# Plagiarism Detection Model\n\nNow that you've created training and test data, you are ready to define and train a model. Your goal in this notebook, will be to train a binary classification model that learns to label an answer file as either plagiarized or not, based on the features you provide the model.\n\nThis task will be broken down into a few discrete steps:\n\n* Upload your data to S3.\n* Define a binary classification model and a training script.\n* Train your model and deploy it.\n* Evaluate your deployed classifier and answer some questions about your approach.\n\nTo complete this notebook, you'll have to complete all given exercises and answer all the questions in this notebook.\n> All your tasks will be clearly labeled **EXERCISE** and questions as **QUESTION**.\n\nIt will be up to you to explore different classification models and decide on a model that gives you the best performance for this dataset.\n\n---",
"_____no_output_____"
],
[
"## Load Data to S3\n\nIn the last notebook, you should have created two files: a `training.csv` and `test.csv` file with the features and class labels for the given corpus of plagiarized/non-plagiarized text data. \n\n>The below cells load in some AWS SageMaker libraries and creates a default bucket. After creating this bucket, you can upload your locally stored data to S3.\n\nSave your train and test `.csv` feature files, locally. To do this you can run the second notebook \"2_Plagiarism_Feature_Engineering\" in SageMaker or you can manually upload your files to this notebook using the upload icon in Jupyter Lab. Then you can upload local files to S3 by using `sagemaker_session.upload_data` and pointing directly to where the training data is saved.",
"_____no_output_____"
]
],
[
[
"import pandas as pd\nimport boto3\nimport sagemaker",
"_____no_output_____"
],
[
"\"\"\"\nDON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE\n\"\"\"\n# session and role\nsagemaker_session = sagemaker.Session()\nrole = sagemaker.get_execution_role()\n\n# create an S3 bucket\nbucket = sagemaker_session.default_bucket()",
"_____no_output_____"
]
],
[
[
"## EXERCISE: Upload your training data to S3\n\nSpecify the `data_dir` where you've saved your `train.csv` file. Decide on a descriptive `prefix` that defines where your data will be uploaded in the default S3 bucket. Finally, create a pointer to your training data by calling `sagemaker_session.upload_data` and passing in the required parameters. It may help to look at the [Session documentation](https://sagemaker.readthedocs.io/en/stable/session.html#sagemaker.session.Session.upload_data) or previous SageMaker code examples.\n\nYou are expected to upload your entire directory. Later, the training script will only access the `train.csv` file.",
"_____no_output_____"
]
],
[
[
"import os\n# should be the name of directory you created to save your features data\ndata_dir = 'plagiarism_data'\n\n# set prefix, a descriptive name for a directory \nprefix = 'plagiarism'\n\n# upload all data to S3\ntest_location = sagemaker_session.upload_data(os.path.join(data_dir, 'test.csv'), key_prefix=prefix, bucket=bucket)\ntrain_location = sagemaker_session.upload_data(os.path.join(data_dir, 'train.csv'), key_prefix=prefix, bucket=bucket)",
"_____no_output_____"
]
],
[
[
"### Test cell\n\nTest that your data has been successfully uploaded. The below cell prints out the items in your S3 bucket and will throw an error if it is empty. You should see the contents of your `data_dir` and perhaps some checkpoints. If you see any other files listed, then you may have some old model files that you can delete via the S3 console (though, additional files shouldn't affect the performance of model developed in this notebook).",
"_____no_output_____"
]
],
[
[
"\"\"\"\nDON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE\n\"\"\"\n# confirm that data is in S3 bucket\nempty_check = []\nfor obj in boto3.resource('s3').Bucket(bucket).objects.all():\n empty_check.append(obj.key)\n print(obj.key)\n\nassert len(empty_check) !=0, 'S3 bucket is empty.'\nprint('Test passed!')",
"plagiarism/test.csv\nplagiarism/train.csv\nTest passed!\n"
]
],
[
[
"---\n\n# Modeling\n\nNow that you've uploaded your training data, it's time to define and train a model!\n\nThe type of model you create is up to you. For a binary classification task, you can choose to go one of three routes:\n* Use a built-in classification algorithm, like LinearLearner.\n* Define a custom Scikit-learn classifier, a comparison of models can be found [here](https://scikit-learn.org/stable/auto_examples/classification/plot_classifier_comparison.html).\n* Define a custom PyTorch neural network classifier. \n\nIt will be up to you to test out a variety of models and choose the best one. Your project will be graded on the accuracy of your final model. \n \n---\n\n## EXERCISE: Complete a training script \n\nTo implement a custom classifier, you'll need to complete a `train.py` script. You've been given the folders `source_sklearn` and `source_pytorch` which hold starting code for a custom Scikit-learn model and a PyTorch model, respectively. Each directory has a `train.py` training script. To complete this project **you only need to complete one of these scripts**; the script that is responsible for training your final model.\n\nA typical training script:\n* Loads training data from a specified directory\n* Parses any training & model hyperparameters (ex. nodes in a neural network, training epochs, etc.)\n* Instantiates a model of your design, with any specified hyperparams\n* Trains that model \n* Finally, saves the model so that it can be hosted/deployed, later\n\n### Defining and training a model\nMuch of the training script code is provided for you. Almost all of your work will be done in the `if __name__ == '__main__':` section. To complete a `train.py` file, you will:\n1. Import any extra libraries you need\n2. Define any additional model training hyperparameters using `parser.add_argument`\n2. Define a model in the `if __name__ == '__main__':` section\n3. Train the model in that same section\n\nBelow, you can use `!pygmentize` to display an existing `train.py` file. Read through the code; all of your tasks are marked with `TODO` comments. \n\n**Note: If you choose to create a custom PyTorch model, you will be responsible for defining the model in the `model.py` file,** and a `predict.py` file is provided. If you choose to use Scikit-learn, you only need a `train.py` file; you may import a classifier from the `sklearn` library.",
"_____no_output_____"
]
],
[
[
"# directory can be changed to: source_sklearn or source_pytorch\n!pygmentize source_sklearn/train.py",
"_____no_output_____"
]
],
[
[
"### Provided code\n\nIf you read the code above, you can see that the starter code includes a few things:\n* Model loading (`model_fn`) and saving code\n* Getting SageMaker's default hyperparameters\n* Loading the training data by name, `train.csv` and extracting the features and labels, `train_x`, and `train_y`\n\nIf you'd like to read more about model saving with [joblib for sklearn](https://scikit-learn.org/stable/modules/model_persistence.html) or with [torch.save](https://pytorch.org/tutorials/beginner/saving_loading_models.html), click on the provided links.",
"_____no_output_____"
],
[
"---\n# Create an Estimator\n\nWhen a custom model is constructed in SageMaker, an entry point must be specified. This is the Python file which will be executed when the model is trained; the `train.py` function you specified above. To run a custom training script in SageMaker, construct an estimator, and fill in the appropriate constructor arguments:\n\n* **entry_point**: The path to the Python script SageMaker runs for training and prediction.\n* **source_dir**: The path to the training script directory `source_sklearn` OR `source_pytorch`.\n* **entry_point**: The path to the Python script SageMaker runs for training and prediction.\n* **source_dir**: The path to the training script directory `train_sklearn` OR `train_pytorch`.\n* **entry_point**: The path to the Python script SageMaker runs for training.\n* **source_dir**: The path to the training script directory `train_sklearn` OR `train_pytorch`.\n* **role**: Role ARN, which was specified, above.\n* **train_instance_count**: The number of training instances (should be left at 1).\n* **train_instance_type**: The type of SageMaker instance for training. Note: Because Scikit-learn does not natively support GPU training, Sagemaker Scikit-learn does not currently support training on GPU instance types.\n* **sagemaker_session**: The session used to train on Sagemaker.\n* **hyperparameters** (optional): A dictionary `{'name':value, ..}` passed to the train function as hyperparameters.\n\nNote: For a PyTorch model, there is another optional argument **framework_version**, which you can set to the latest version of PyTorch, `1.0`.\n\n## EXERCISE: Define a Scikit-learn or PyTorch estimator\n\nTo import your desired estimator, use one of the following lines:\n```\nfrom sagemaker.sklearn.estimator import SKLearn\n```\n```\nfrom sagemaker.pytorch import PyTorch\n```",
"_____no_output_____"
]
],
[
[
"# your import and estimator code, here\n\nfrom sagemaker.sklearn.estimator import SKLearn\nestimator = SKLearn(entry_point='train.py',\n source_dir='source_sklearn',\n role=role,\n train_instance_count=1,\n train_instance_type='ml.c4.xlarge',\n framework_version='0.20.0', \n sagemaker_session=sagemaker_session,\n hyperparameters={\n 'max_leaf_nodes': 30,\n 'max_depth':2,\n }\n )",
"_____no_output_____"
]
],
[
[
"## EXERCISE: Train the estimator\n\nTrain your estimator on the training data stored in S3. This should create a training job that you can monitor in your SageMaker console.",
"_____no_output_____"
]
],
[
[
"%%time\n\n# Train your estimator on S3 training data\n\n# s3_input_train = sagemaker.s3_input(s3_data=train_location, content_type='text/csv')\n# linear_learner.fit({'train':s3_input_train})\nestimator.fit({'train': train_location})",
"2020-03-05 19:32:37 Starting - Starting the training job...\n2020-03-05 19:32:38 Starting - Launching requested ML instances...\n2020-03-05 19:33:37 Starting - Preparing the instances for training.........\n2020-03-05 19:35:06 Downloading - Downloading input data\n2020-03-05 19:35:06 Training - Downloading the training image...\n2020-03-05 19:35:33 Uploading - Uploading generated training model\n2020-03-05 19:35:33 Completed - Training job completed\n\u001b[34m2020-03-05 19:35:21,498 sagemaker-containers INFO Imported framework sagemaker_sklearn_container.training\u001b[0m\n\u001b[34m2020-03-05 19:35:21,500 sagemaker-containers INFO No GPUs detected (normal if no gpus installed)\u001b[0m\n\u001b[34m2020-03-05 19:35:21,510 sagemaker_sklearn_container.training INFO Invoking user training script.\u001b[0m\n\u001b[34m2020-03-05 19:35:21,786 sagemaker-containers INFO Module train does not provide a setup.py. \u001b[0m\n\u001b[34mGenerating setup.py\u001b[0m\n\u001b[34m2020-03-05 19:35:21,786 sagemaker-containers INFO Generating setup.cfg\u001b[0m\n\u001b[34m2020-03-05 19:35:21,786 sagemaker-containers INFO Generating MANIFEST.in\u001b[0m\n\u001b[34m2020-03-05 19:35:21,786 sagemaker-containers INFO Installing module with the following command:\u001b[0m\n\u001b[34m/miniconda3/bin/python -m pip install . \u001b[0m\n\u001b[34mProcessing /opt/ml/code\u001b[0m\n\u001b[34mBuilding wheels for collected packages: train\n Building wheel for train (setup.py): started\n Building wheel for train (setup.py): finished with status 'done'\n Created wheel for train: filename=train-1.0.0-py2.py3-none-any.whl size=7136 sha256=574938952486976c0956d67d5168387c7fc3e79ea3df77bceebda56d8e179f05\n Stored in directory: /tmp/pip-ephem-wheel-cache-lbjulvs7/wheels/35/24/16/37574d11bf9bde50616c67372a334f94fa8356bc7164af8ca3\u001b[0m\n\u001b[34mSuccessfully built train\u001b[0m\n\u001b[34mInstalling collected packages: train\u001b[0m\n\u001b[34mSuccessfully installed train-1.0.0\u001b[0m\n\u001b[34m2020-03-05 19:35:23,208 sagemaker-containers INFO No GPUs detected (normal if no gpus installed)\u001b[0m\n\u001b[34m2020-03-05 19:35:23,218 sagemaker-containers INFO Invoking user script\n\u001b[0m\n\u001b[34mTraining Env:\n\u001b[0m\n\u001b[34m{\n \"additional_framework_parameters\": {},\n \"channel_input_dirs\": {\n \"train\": \"/opt/ml/input/data/train\"\n },\n \"current_host\": \"algo-1\",\n \"framework_module\": \"sagemaker_sklearn_container.training:main\",\n \"hosts\": [\n \"algo-1\"\n ],\n \"hyperparameters\": {\n \"max_leaf_nodes\": 30,\n \"max_depth\": 2\n },\n \"input_config_dir\": \"/opt/ml/input/config\",\n \"input_data_config\": {\n \"train\": {\n \"TrainingInputMode\": \"File\",\n \"S3DistributionType\": \"FullyReplicated\",\n \"RecordWrapperType\": \"None\"\n }\n },\n \"input_dir\": \"/opt/ml/input\",\n \"is_master\": true,\n \"job_name\": \"sagemaker-scikit-learn-2020-03-05-19-32-36-719\",\n \"log_level\": 20,\n \"master_hostname\": \"algo-1\",\n \"model_dir\": \"/opt/ml/model\",\n \"module_dir\": \"s3://sagemaker-us-east-2-210288789627/sagemaker-scikit-learn-2020-03-05-19-32-36-719/source/sourcedir.tar.gz\",\n \"module_name\": \"train\",\n \"network_interface_name\": \"eth0\",\n \"num_cpus\": 4,\n \"num_gpus\": 0,\n \"output_data_dir\": \"/opt/ml/output/data\",\n \"output_dir\": \"/opt/ml/output\",\n \"output_intermediate_dir\": \"/opt/ml/output/intermediate\",\n \"resource_config\": {\n \"current_host\": \"algo-1\",\n \"hosts\": [\n \"algo-1\"\n ],\n \"network_interface_name\": \"eth0\"\n },\n \"user_entry_point\": \"train.py\"\u001b[0m\n\u001b[34m}\n\u001b[0m\n\u001b[34mEnvironment variables:\n\u001b[0m\n\u001b[34mSM_HOSTS=[\"algo-1\"]\u001b[0m\n\u001b[34mSM_NETWORK_INTERFACE_NAME=eth0\u001b[0m\n\u001b[34mSM_HPS={\"max_depth\":2,\"max_leaf_nodes\":30}\u001b[0m\n\u001b[34mSM_USER_ENTRY_POINT=train.py\u001b[0m\n\u001b[34mSM_FRAMEWORK_PARAMS={}\u001b[0m\n\u001b[34mSM_RESOURCE_CONFIG={\"current_host\":\"algo-1\",\"hosts\":[\"algo-1\"],\"network_interface_name\":\"eth0\"}\u001b[0m\n\u001b[34mSM_INPUT_DATA_CONFIG={\"train\":{\"RecordWrapperType\":\"None\",\"S3DistributionType\":\"FullyReplicated\",\"TrainingInputMode\":\"File\"}}\u001b[0m\n\u001b[34mSM_OUTPUT_DATA_DIR=/opt/ml/output/data\u001b[0m\n\u001b[34mSM_CHANNELS=[\"train\"]\u001b[0m\n\u001b[34mSM_CURRENT_HOST=algo-1\u001b[0m\n\u001b[34mSM_MODULE_NAME=train\u001b[0m\n\u001b[34mSM_LOG_LEVEL=20\u001b[0m\n\u001b[34mSM_FRAMEWORK_MODULE=sagemaker_sklearn_container.training:main\u001b[0m\n\u001b[34mSM_INPUT_DIR=/opt/ml/input\u001b[0m\n\u001b[34mSM_INPUT_CONFIG_DIR=/opt/ml/input/config\u001b[0m\n\u001b[34mSM_OUTPUT_DIR=/opt/ml/output\u001b[0m\n\u001b[34mSM_NUM_CPUS=4\u001b[0m\n\u001b[34mSM_NUM_GPUS=0\u001b[0m\n\u001b[34mSM_MODEL_DIR=/opt/ml/model\u001b[0m\n\u001b[34mSM_MODULE_DIR=s3://sagemaker-us-east-2-210288789627/sagemaker-scikit-learn-2020-03-05-19-32-36-719/source/sourcedir.tar.gz\u001b[0m\n\u001b[34mSM_TRAINING_ENV={\"additional_framework_parameters\":{},\"channel_input_dirs\":{\"train\":\"/opt/ml/input/data/train\"},\"current_host\":\"algo-1\",\"framework_module\":\"sagemaker_sklearn_container.training:main\",\"hosts\":[\"algo-1\"],\"hyperparameters\":{\"max_depth\":2,\"max_leaf_nodes\":30},\"input_config_dir\":\"/opt/ml/input/config\",\"input_data_config\":{\"train\":{\"RecordWrapperType\":\"None\",\"S3DistributionType\":\"FullyReplicated\",\"TrainingInputMode\":\"File\"}},\"input_dir\":\"/opt/ml/input\",\"is_master\":true,\"job_name\":\"sagemaker-scikit-learn-2020-03-05-19-32-36-719\",\"log_level\":20,\"master_hostname\":\"algo-1\",\"model_dir\":\"/opt/ml/model\",\"module_dir\":\"s3://sagemaker-us-east-2-210288789627/sagemaker-scikit-learn-2020-03-05-19-32-36-719/source/sourcedir.tar.gz\",\"module_name\":\"train\",\"network_interface_name\":\"eth0\",\"num_cpus\":4,\"num_gpus\":0,\"output_data_dir\":\"/opt/ml/output/data\",\"output_dir\":\"/opt/ml/output\",\"output_intermediate_dir\":\"/opt/ml/output/intermediate\",\"resource_config\":{\"current_host\":\"algo-1\",\"hosts\":[\"algo-1\"],\"network_interface_name\":\"eth0\"},\"user_entry_point\":\"train.py\"}\u001b[0m\n\u001b[34mSM_USER_ARGS=[\"--max_depth\",\"2\",\"--max_leaf_nodes\",\"30\"]\u001b[0m\n\u001b[34mSM_OUTPUT_INTERMEDIATE_DIR=/opt/ml/output/intermediate\u001b[0m\n\u001b[34mSM_CHANNEL_TRAIN=/opt/ml/input/data/train\u001b[0m\n\u001b[34mSM_HP_MAX_LEAF_NODES=30\u001b[0m\n\u001b[34mSM_HP_MAX_DEPTH=2\u001b[0m\n\u001b[34mPYTHONPATH=/miniconda3/bin:/miniconda3/lib/python37.zip:/miniconda3/lib/python3.7:/miniconda3/lib/python3.7/lib-dynload:/miniconda3/lib/python3.7/site-packages\n\u001b[0m\n\u001b[34mInvoking script with the following command:\n\u001b[0m\n\u001b[34m/miniconda3/bin/python -m train --max_depth 2 --max_leaf_nodes 30\n\n\u001b[0m\n\u001b[34m/miniconda3/lib/python3.7/site-packages/sklearn/externals/joblib/externals/cloudpickle/cloudpickle.py:47: DeprecationWarning: the imp module is deprecated in favour of importlib; see the module's documentation for alternative uses\n import imp\u001b[0m\n\u001b[34m2020-03-05 19:35:24,500 sagemaker-containers INFO Reporting training SUCCESS\u001b[0m\nTraining seconds: 50\nBillable seconds: 50\nCPU times: user 360 ms, sys: 33.3 ms, total: 393 ms\nWall time: 3min 11s\n"
]
],
[
[
"## EXERCISE: Deploy the trained model\n\nAfter training, deploy your model to create a `predictor`. If you're using a PyTorch model, you'll need to create a trained `PyTorchModel` that accepts the trained `<model>.model_data` as an input parameter and points to the provided `source_pytorch/predict.py` file as an entry point. \n\nTo deploy a trained model, you'll use `<model>.deploy`, which takes in two arguments:\n* **initial_instance_count**: The number of deployed instances (1).\n* **instance_type**: The type of SageMaker instance for deployment.\n\nNote: If you run into an instance error, it may be because you chose the wrong training or deployment instance_type. It may help to refer to your previous exercise code to see which types of instances we used.",
"_____no_output_____"
]
],
[
[
"%%time\n\n# uncomment, if needed\n# from sagemaker.pytorch import PyTorchModel\npredictor = estimator.deploy(initial_instance_count=1, instance_type='ml.t2.medium')",
"---------------!CPU times: user 257 ms, sys: 5.89 ms, total: 263 ms\nWall time: 7min 31s\n"
]
],
[
[
"---\n# Evaluating Your Model\n\nOnce your model is deployed, you can see how it performs when applied to our test data.\n\nThe provided cell below, reads in the test data, assuming it is stored locally in `data_dir` and named `test.csv`. The labels and features are extracted from the `.csv` file.",
"_____no_output_____"
]
],
[
[
"\"\"\"\nDON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE\n\"\"\"\nimport os\n\n# read in test data, assuming it is stored locally\ntest_data = pd.read_csv(os.path.join(data_dir, \"test.csv\"), header=None, names=None)\n\n# labels are in the first column\ntest_y = test_data.iloc[:,0]\ntest_x = test_data.iloc[:,1:]",
"_____no_output_____"
]
],
[
[
"## EXERCISE: Determine the accuracy of your model\n\nUse your deployed `predictor` to generate predicted, class labels for the test data. Compare those to the *true* labels, `test_y`, and calculate the accuracy as a value between 0 and 1.0 that indicates the fraction of test data that your model classified correctly. You may use [sklearn.metrics](https://scikit-learn.org/stable/modules/classes.html#module-sklearn.metrics) for this calculation.\n\n**To pass this project, your model should get at least 90% test accuracy.**",
"_____no_output_____"
]
],
[
[
"# First: generate predicted, class labels\nimport json \n\ntest_y_preds = predictor.predict(test_x.values)\n\n\n\"\"\"\nDON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE\n\"\"\"\n# test that your model generates the correct number of labels\nassert len(test_y_preds)==len(test_y), 'Unexpected number of predictions.'\nprint('Test passed!')",
"Test passed!\n"
],
[
"# Using Sklearn\n\nfrom sklearn.metrics import accuracy_score\naccuracy_score(y_true=test_y.values, y_pred=test_y_preds)",
"_____no_output_____"
],
[
"# Second: calculate the test accuracy\nimport numpy as np\n\ntest_labels = test_y.values\ntest_preds = np.asarray(test_y_preds)\n\ntp = np.logical_and(test_labels, test_preds).sum()\nfp = np.logical_and(1-test_labels, test_preds).sum()\nfn = np.logical_and(test_labels, 1-test_preds).sum()\ntn = np.logical_and(1-test_labels, 1-test_preds).sum()\n\naccuracy = (tp+tn)/(fp+fn+tn+tp)\n\nprint(accuracy)\n\n\n## print out the array of predicted and true labels, if you want\nprint('\\nPredicted class labels: ')\nprint(test_y_preds)\nprint('\\nTrue class labels: ')\nprint(test_y.values)",
"0.96\n\nPredicted class labels: \n[1 1 1 1 1 1 0 0 0 0 0 0 1 1 1 1 1 1 0 1 0 1 1 0 1]\n\nTrue class labels: \n[1 1 1 1 1 1 0 0 0 0 0 0 1 1 1 1 1 1 0 1 0 1 1 0 0]\n"
],
[
"print('True Positives: ', tp)\nprint('True Negatives: ', tn)\nprint('False Positives: ', fp)\nprint('False Negatives: ', fn)\n\nprint('Recall: ', tp / (tp + fn))\nprint('Precision: ', tp / (fp + tp))",
"True Positives: 15\nTrue Negatives: 9\nFalse Positives: 1\nFalse Negatives: 0\nRecall: 1.0\nPrecision: 0.9375\n"
]
],
[
[
"### Question 1: How many false positives and false negatives did your model produce, if any? And why do you think this is?",
"_____no_output_____"
],
[
"** Answer**: \n\nThe model produced 1 false positives and 0 false negatives\nIt produced this amount of false positives due to having a good accuracy score (96%)\n\nThis might be a reason to raise an alarm for overfitting, however with high scores in Recall (100%) and Precision (93%),\nI do not think that is the case.",
"_____no_output_____"
],
[
"### Question 2: How did you decide on the type of model to use? ",
"_____no_output_____"
],
[
"** Answer**:\n\nUsing an Decision Tree Classifier, I felt this would be a good decision as this algorithm can be used as a simple tree algorithm, and been able to set hyperparameters specific to this use case",
"_____no_output_____"
],
[
"----\n## EXERCISE: Clean up Resources\n\nAfter you're done evaluating your model, **delete your model endpoint**. You can do this with a call to `.delete_endpoint()`. You need to show, in this notebook, that the endpoint was deleted. Any other resources, you may delete from the AWS console, and you will find more instructions on cleaning up all your resources, below.",
"_____no_output_____"
]
],
[
[
"# uncomment and fill in the line below!\npredictor.delete_endpoint()",
"_____no_output_____"
]
],
[
[
"### Deleting S3 bucket\n\nWhen you are *completely* done with training and testing models, you can also delete your entire S3 bucket. If you do this before you are done training your model, you'll have to recreate your S3 bucket and upload your training data again.",
"_____no_output_____"
]
],
[
[
"# deleting bucket, uncomment lines below\n\nbucket_to_delete = boto3.resource('s3').Bucket(bucket)\nbucket_to_delete.objects.all().delete()",
"_____no_output_____"
]
],
[
[
"### Deleting all your models and instances\n\nWhen you are _completely_ done with this project and do **not** ever want to revisit this notebook, you can choose to delete all of your SageMaker notebook instances and models by following [these instructions](https://docs.aws.amazon.com/sagemaker/latest/dg/ex1-cleanup.html). Before you delete this notebook instance, I recommend at least downloading a copy and saving it, locally.",
"_____no_output_____"
],
[
"---\n## Further Directions\n\nThere are many ways to improve or add on to this project to expand your learning or make this more of a unique project for you. A few ideas are listed below:\n* Train a classifier to predict the *category* (1-3) of plagiarism and not just plagiarized (1) or not (0).\n* Utilize a different and larger dataset to see if this model can be extended to other types of plagiarism.\n* Use language or character-level analysis to find different (and more) similarity features.\n* Write a complete pipeline function that accepts a source text and submitted text file, and classifies the submitted text as plagiarized or not.\n* Use API Gateway and a lambda function to deploy your model to a web application.\n\nThese are all just options for extending your work. If you've completed all the exercises in this notebook, you've completed a real-world application, and can proceed to submit your project. Great job!",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"raw",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"raw"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
]
] |
4ab6af4d64a57da4096af4f1536018d6dac768a1
| 59,807 |
ipynb
|
Jupyter Notebook
|
code/dataset/generate_ReaSCAN.ipynb
|
frankaging/Reason-SCAN
|
1e6726aab70c915a3d3068bed77ae255feeaf6f1
|
[
"CC-BY-4.0"
] | 14 |
2021-07-05T19:05:53.000Z
|
2022-02-08T10:09:36.000Z
|
code/dataset/generate_ReaSCAN.ipynb
|
frankaging/Reason-SCAN
|
1e6726aab70c915a3d3068bed77ae255feeaf6f1
|
[
"CC-BY-4.0"
] | 1 |
2021-11-21T10:44:51.000Z
|
2021-12-02T15:11:51.000Z
|
code/dataset/generate_ReaSCAN.ipynb
|
frankaging/Reason-SCAN
|
1e6726aab70c915a3d3068bed77ae255feeaf6f1
|
[
"CC-BY-4.0"
] | 2 |
2021-06-17T08:16:36.000Z
|
2021-06-21T17:54:54.000Z
| 50.856293 | 152 | 0.498537 |
[
[
[
"import argparse\nfrom collections import namedtuple, OrderedDict\nimport itertools\nimport os\nimport numpy as np\nfrom typing import Tuple\nfrom typing import List\nfrom typing import Dict\nimport random\nfrom itertools import product\nimport copy\nimport re\nimport random\nimport hashlib\nimport pathlib\nimport json\nimport matplotlib as plt\nimport matplotlib.pyplot as plt\nimport matplotlib.image as mpimg\nos.environ['QT_QPA_PLATFORM']='offscreen'\nplt.rcParams[\"font.family\"] = \"DejaVu Serif\"\nfont = {'family' : 'DejaVu Serif',\n 'size' : 20}\nplt.rc('font', **font)\nimport plotly.tools as tls\n\nfrom utils import one_hot\nfrom utils import generate_possible_object_names\nfrom utils import numpy_array_to_image\n\nfrom vocabulary import *\nfrom object_vocabulary import *\nfrom world import *\nfrom grammer import *\nfrom simulator import *\nfrom relation_graph import *\n\nimport logging\n\nimport warnings\nwarnings.filterwarnings(\"ignore\")",
"_____no_output_____"
],
[
"# Helpers.\ndef get_relation_statistics(command_structs):\n \"\"\"\n Return a dictionary, (relation, position) with counts\n \"\"\"\n stats = {}\n for i in range(2): # at max 2!\n stats[f\"position-{i}\"] = {}\n for command in command_structs:\n pos_id = 0\n for k, v in command[\"rel_map\"].items():\n if v in stats[f\"position-{pos_id}\"].keys():\n stats[f\"position-{pos_id}\"][v] += 1\n else:\n stats[f\"position-{pos_id}\"][v] = 1\n pos_id += 1\n return stats\n\n\ndef get_attribute_statistics(command_structs, include_keywords=[\"circle\", \"cylinder\", \"square\", \"box\", \"object\"]):\n \n stats = {}\n # for k, v in command_structs[0][\"obj_map\"].items():\n # stats[k] = {} # we can do it in object level!\n for i in range(3): # at max 2!\n stats[f\"$OBJ_{i}\"] = {}\n \n for command in command_structs:\n for k, v in command[\"obj_map\"].items():\n for keyword in include_keywords:\n keyword_list = keyword.split(\" \") # in case there are a couple!\n match = True\n for sub_k in keyword_list:\n if sub_k not in v:\n match = False\n break\n if match:\n if keyword in stats[k].keys():\n stats[k][keyword] += 1\n else:\n stats[k][keyword] = 1\n return stats\n\n\ndef get_keyword_statistics(command_structs, include_keyword=\"adverb\"):\n stats = {}\n for command in command_structs:\n keyword = command[include_keyword]\n if keyword in stats.keys():\n stats[keyword] += 1\n else:\n stats[keyword] = 1\n return stats\n\ndef flatten_dictionary(\n dictionary_in\n):\n flat_dictionary = {}\n for k, v in dictionary_in.items():\n for kk, vv in v.items():\n if kk not in flat_dictionary:\n flat_dictionary[kk] = vv\n else:\n flat_dictionary[kk] += vv\n return flat_dictionary\n\ndef plot_dictionary(\n dictionary_in,\n y_label=\"Frequency\",\n x_label=\"Conditions\",\n title=\"Missing Title\",\n save_file=None,\n is_plot=False,\n wandb=None,\n):\n group_str = [k for k, _ in dictionary_in[0].items()]\n if len(group_str) > 8:\n rotate=90\n fontsize=10\n else:\n rotate=45\n fontsize=13\n all_stats = []\n for d in dictionary_in:\n group_stats = [d[k] for k in group_str]\n all_stats.append(group_stats)\n all_stats = np.array(all_stats)\n std = np.std(all_stats, axis=0)\n mean = np.mean(all_stats, axis=0)\n\n # input data\n mean_values = mean\n variance = std**2\n bar_labels = group_str\n \n # plot bars\n x_pos = list(range(len(bar_labels)))\n fig = plt.figure(figsize=(8,6))\n ax = fig.add_subplot(111)\n g = ax.bar(x_pos, mean_values, yerr=variance, align='center', alpha=0.5)\n\n plt.grid()\n\n # set height of the y-axis\n max_y = max(zip(mean_values, variance)) # returns a tuple, here: (3, 5)\n plt.ylim([0, (max_y[0] + max_y[1]) * 1.1])\n\n # set axes labels and title\n plt.ylabel(y_label)\n \n plt.xticks(x_pos, bar_labels)\n plt.xticks(rotation = rotate, fontsize=fontsize) \n plt.yticks(rotation = 45)\n plt.title(title, fontsize=10)\n if mean_values[0] > 10000:\n plt.ticklabel_format(axis='y', style='sci', scilimits=(4,4))\n \n if wandb != None:\n # Let us also try to log this plot to wandb!\n wandb.log({title: wandb.Image(fig)})\n\n if save_file != None:\n plt.savefig(save_file, dpi=100, bbox_inches='tight')\n plt.close(fig)\n else:\n if is_plot:\n plt.show()\n \ndef get_command_struct_statistics(\n command_structs, run_name=\"ReaSCAN-Awesome\", date=\"2021-05-06\", \n split=\"demo\",\n compositional_split=False,\n n_sample=-1, n_runs=10,\n output_dir=\"../../data-files/ReaSCAN-compositional_splits/\",\n save_to_disk=True,\n wandb=None\n):\n statistics = OrderedDict({\n \"run_name\": run_name,\n \"date\": date,\n \"splits\": split,\n \"number_of_these_examples_seen_in_training\": -1 if not compositional_split else 0,\n \"number_of_command_structs\": len(command_structs),\n })\n if n_sample == -1:\n n_sample = len(command_structs)\n # If we are downsampling, we need to do more runs as well!\n random.shuffle(command_structs)\n \n patterns = set([])\n for command_s in command_structs:\n patterns.add(command_s[\"grammer_pattern\"])\n statistics[\"command_patterns\"] = list(patterns)\n \n pattern_stats = get_keyword_statistics(command_structs, include_keyword=\"grammer_pattern\")\n statistics[\"pattern_stats\"] = pattern_stats\n \n # verb\n verb_stats = get_keyword_statistics(command_structs, include_keyword=\"verb\")\n statistics[\"verb_stats\"] = verb_stats\n plot_dictionary(\n [verb_stats],\n title=\"Verbs\",\n save_file=os.path.join(output_dir, f\"verb_stats-{split}.png\"),\n wandb=wandb,\n )\n \n # adverb\n adverb_stats = get_keyword_statistics(command_structs, include_keyword=\"adverb\")\n # special handling for adverb for better readabilities\n adverb_stats_rebuild = {}\n for k, v in adverb_stats.items():\n if k == \"\":\n adverb_stats_rebuild[\"EMPTY\"] = v\n else:\n adverb_stats_rebuild[k] = v\n statistics[\"adverb_stats\"] = adverb_stats_rebuild\n plot_dictionary(\n [adverb_stats_rebuild],\n title=\"Adverbs\",\n save_file=os.path.join(output_dir, f\"adverb_stats-{split}.png\"),\n wandb=wandb,\n )\n \n # relation\n relation_stats = get_relation_statistics(command_structs)\n if len(flatten_dictionary(relation_stats)) != 0:\n statistics[\"relation_stats\"] = relation_stats\n plot_dictionary(\n [flatten_dictionary(relation_stats)],\n title=\"Relation-Types\",\n save_file=os.path.join(output_dir, f\"relation_type_stats-{split}.png\"),\n wandb=wandb,\n )\n \n # attribute\n nouns = [\"circle\", \"cylinder\", \"square\", \"box\", \"object\"]\n n_stats = get_attribute_statistics(command_structs, include_keywords=nouns)\n statistics[\"shape_stats\"] = n_stats\n plot_dictionary(\n [flatten_dictionary(n_stats)],\n title=\"Shapes\",\n save_file=os.path.join(output_dir, f\"shape_stats-{split}.png\"),\n wandb=wandb,\n )\n \n color_adjectives = [\"red\", \"blue\", \"green\", \"yellow\"]\n c_stats = get_attribute_statistics(command_structs, include_keywords=color_adjectives)\n statistics[\"color_stats\"] = c_stats\n if len(flatten_dictionary(c_stats)) != 0:\n plot_dictionary(\n [flatten_dictionary(c_stats)],\n title=\"Colors\",\n save_file=os.path.join(output_dir, f\"color_stats-{split}.png\"),\n wandb=wandb,\n )\n\n size_adjectives = [\"big\", \"small\"]\n s_stats = get_attribute_statistics(command_structs, include_keywords=size_adjectives)\n if len(flatten_dictionary(s_stats)) != 0:\n statistics[\"size_stats\"] = s_stats\n plot_dictionary(\n [flatten_dictionary(s_stats)],\n title=\"Sizes\",\n save_file=os.path.join(output_dir, f\"size_stats-{split}.png\"),\n wandb=wandb,\n )\n \n # second order attribute\n color_adjectives = [\"red\", \"blue\", \"green\", \"yellow\"]\n nouns = [\"circle\", \"cylinder\", \"square\", \"box\", \"object\"]\n c_n_p = product(color_adjectives, nouns)\n include_keywords = [\" \".join(c_n) for c_n in c_n_p]\n c_n_stats = get_attribute_statistics(command_structs, include_keywords=include_keywords)\n statistics[\"color_and_shape_stats\"] = c_n_stats\n if len(flatten_dictionary(c_n_stats)) != 0:\n plot_dictionary(\n [flatten_dictionary(c_n_stats)],\n title=\"Colors-Shapes\",\n save_file=os.path.join(output_dir, f\"color+shape_stats-{split}.png\"),\n wandb=wandb,\n )\n\n size_adjectives = [\"big\", \"small\"]\n nouns = [\"circle\", \"cylinder\", \"square\", \"box\", \"object\"]\n s_n_p = product(size_adjectives, nouns)\n include_keywords = [\" \".join(s_n) for s_n in s_n_p]\n s_n_stats = get_attribute_statistics(command_structs, include_keywords=include_keywords)\n statistics[\"size_and_shape_stats\"] = s_n_stats\n if len(flatten_dictionary(s_n_stats)) != 0:\n plot_dictionary(\n [flatten_dictionary(s_n_stats)],\n title=\"Sizes-Shapes\",\n save_file=os.path.join(output_dir, f\"size+shape_stats-{split}.png\"),\n wandb=wandb,\n )\n \n # third order attribute\n size_adjectives = [\"big\", \"small\"]\n color_adjectives = [\"red\", \"blue\", \"green\", \"yellow\"]\n nouns = [\"circle\", \"cylinder\", \"square\", \"box\", \"object\"]\n all_p = product(size_adjectives, color_adjectives, nouns)\n include_keywords = [\" \".join(a) for a in all_p]\n all_stats = get_attribute_statistics(command_structs, include_keywords=include_keywords)\n statistics[\"size_and_color_and_shape_stats\"] = all_stats\n \n if save_to_disk:\n import yaml\n with open(os.path.join(output_dir, f\"command_struct_only_stats-{split}.yml\"), 'w') as yaml_file:\n yaml.dump(statistics, yaml_file, default_flow_style=False)\n \n return statistics\n\ndef arg_parse():\n \n # This is a single loop to generate the dataset.\n n_processes = 1\n mode = \"all\"\n n_command_struct = 10000\n grid_size = 6\n n_object_max = 10\n seed = 42\n date = \"2021-05-07\"\n per_command_world_retry_max = 200\n per_command_world_target_count = 10 # for each command, we target to have 50 shapeWorld!\n resumed_from_file_path = \"\"\n is_tensorboard = False\n \n parser = argparse.ArgumentParser(description='ReaSCAN argparse.')\n # Experiment management:\n parser.add_argument('--n_processes', type=int, default=1,\n help='Number of process used to generate the dataset.')\n parser.add_argument('--index_start', type=int, default=-1,\n help='Number of command sampled from the command population.')\n parser.add_argument('--index_end', type=int, default=-1,\n help='Number of command sampled from the command population.')\n\n parser.add_argument('--mode', type=str, default=\"all\",\n help='mode')\n parser.add_argument('--n_command_struct', type=int, default=10000,\n help='Number of command sampled from the command population.')\n parser.add_argument('--grid_size', type=int, default=6,\n help='Grid size of the world.')\n parser.add_argument('--n_object_max', type=int, default=10,\n help='Number of object at max in the shapeWorld (Note that you may still have more than this number!).')\n parser.add_argument('--seed', type=int, default=42,\n help='Random seed.')\n parser.add_argument('--date', type=str,\n help='date')\n parser.add_argument('--per_command_world_retry_max', type=int, default=200,\n help='How many times you can retry for each world generation.')\n parser.add_argument('--per_command_world_target_count', type=int, default=50,\n help='The targeted number of world to have per command.')\n parser.add_argument(\"--is_tensorboard\",\n default=False,\n action='store_true',\n help=\"Whether to use tensorboard.\")\n \n parser.add_argument(\"--include_relation_distractor\",\n default=False,\n action='store_true',\n help=\"Whether to use tensorboard.\")\n parser.add_argument(\"--include_attribute_distractor\",\n default=False,\n action='store_true',\n help=\"Whether to use tensorboard.\")\n parser.add_argument(\"--include_isomorphism_distractor\",\n default=False,\n action='store_true',\n help=\"Whether to use tensorboard.\")\n parser.add_argument(\"--include_random_distractor\",\n default=False,\n action='store_true',\n help=\"Whether to use tensorboard.\")\n parser.add_argument('--full_relation_probability', type=float, default=1.0,\n help='Probability of including full relation distractors.')\n \n parser.add_argument('--save_interal', type=int, default=200,\n help='Saving intervel in command count.')\n \n \n parser.add_argument('--command_pattern', type=str, default=\"p3\",\n help='What pattern to use, currently, we support p1-p4.')\n \n parser.add_argument('--resumed_from_file_path', type=str, default=\"\",\n help='Whether to resume for this file.')\n parser.add_argument('--output_dir', type=str, default=\"../../data-files/ReaSCAN-compositional_splits/\",\n help='Whether to resume for this file.')\n\n parser.set_defaults(\n # Exp management:\n n_processes=1,\n mode=\"all\",\n n_command_struct=10000,\n grid_size=6,\n n_object_max=10,\n seed=42,\n date=\"2021-05-07\",\n per_command_world_retry_max=200,\n per_command_world_target_count=50,\n resumed_from_file_path=\"\",\n is_tensorboard=False,\n output_dir=\"../../data-files/ReaSCAN-compositional_splits/\",\n )\n try:\n get_ipython().run_line_magic('matplotlib', 'inline')\n args = parser.parse_args([])\n except:\n args = parser.parse_args()\n return args\n\ndef example_classifier(\n task_info,\n mode=\"demo\",\n default_split_prob={\n \"train\": 0.9, \n \"dev\": 0.01,\n \"test\": 0.09,\n },\n):\n \"\"\"\n This will return the split this data belongs to.\n \"\"\"\n if mode == \"demo\" or mode == \"all\":\n if random.random() < default_split_prob[\"train\"]:\n return \"train\"\n else:\n if random.random() < 0.9:\n return \"test\"\n else:\n return \"dev\"\n else:\n # We need to add here logics to determine\n # compositional splits!\n pass",
"_____no_output_____"
],
[
"# Some tips:\n# Do not debug in this file, you can simply copy the questionable struct\n# to the lightweight demo file, and you can debug there!",
"_____no_output_____"
],
[
"if __name__ == \"__main__\":\n \n # Loading arguments\n args = arg_parse()\n try:\n# get_ipython().run_line_magic('matplotlib', 'inline')\n# # Experiment management:\n# args.n_processes=1\n# args.mode=\"demo\"\n# args.n_command_struct=20\n# args.grid_size=6\n# args.n_object_max=10\n# args.seed=42\n# args.date=\"2021-05-07\"\n# args.per_command_world_retry_max=20\n# args.per_command_world_target_count=3\n# args.resumed_from_file_path=\"\"\n# args.is_tensorboard=True # Let us try this!\n# args.output_dir=\"../../data-files/ReaSCAN-demo/\"\n# is_jupyter = True\n \n get_ipython().run_line_magic('matplotlib', 'inline')\n # Experiment management:\n args.n_processes=1\n args.mode=\"train\"\n args.n_command_struct=675*5\n args.grid_size=6\n args.n_object_max=10\n args.seed=42\n args.save_interal = 200\n args.date=\"2021-05-30\"\n args.per_command_world_retry_max=1000\n args.per_command_world_target_count=180\n args.resumed_from_file_path=\"\"\n args.is_tensorboard=True # Let us try this!\n args.output_dir=\"../../data-files/ReaSCAN-compositional-p3-full-relation/\"\n is_jupyter = True\n \n args.index_start = -1\n args.index_end = -1\n except:\n is_jupyter = False\n \n loading_p1 = True if args.command_pattern == \"p1\" else False\n p1_exhaustive_verb_adverb = False\n loading_p2 = True if args.command_pattern == \"p2\" else False\n loading_p3 = True if args.command_pattern == \"p3\" else False\n loading_p4 = True if args.command_pattern == \"p4\" else False\n\n save_command_stats = False\n save_at_interval = True\n save_interal = args.save_interal\n\n # TODO: add these to args.\n logging_interval = 1000\n \n # Create output directory if not exists.\n pathlib.Path(args.output_dir).mkdir(parents=True, exist_ok=True) \n \n logging.basicConfig(\n level=logging.INFO, \n format='%(asctime)s %(levelname)-8s %(message)s', \n datefmt='%a, %d %b %Y %H:%M:%S', \n filename=os.path.join(args.output_dir, \"generator.log\"),\n )\n logger = logging.getLogger(__name__)\n logging.getLogger().addHandler(logging.StreamHandler(os.sys.stdout))\n \n logger.info(\"Generating ReaSCAN with following parameters: \")\n logger.info(args)\n \n # This is a single loop to generate the dataset.\n n_processes = args.n_processes\n mode = args.mode\n n_command_struct = args.n_command_struct\n grid_size = args.grid_size\n n_object_max = args.n_object_max\n seed = args.seed\n date = args.date\n per_command_world_retry_max = args.per_command_world_retry_max\n per_command_world_target_count = args.per_command_world_target_count # for each command, we target to have 50 shapeWorld!\n resumed_from_file_path = args.resumed_from_file_path\n output_dir = args.output_dir\n is_tensorboard = args.is_tensorboard\n \n if is_tensorboard:\n logger.warning(\"Enabling wandb for tensorboard logging...\")\n import wandb\n run = wandb.init(project=\"ReaSCAN\", entity=\"wuzhengx\")\n run_name = wandb.run.name\n wandb.config.update(args)\n else:\n wandb = None\n\n random.seed(seed)\n np.random.seed(seed)\n \n # We also need something to generate generalization\n # splits!\n params = {\n \"n_processes\": n_processes,\n \"mode\": mode,\n \"n_command_struct\": n_command_struct,\n \"grid_size\": grid_size,\n \"n_object_max\": n_object_max,\n \"seed\": seed,\n \"per_command_world_retry_max\": per_command_world_retry_max,\n \"per_command_world_target_count\": per_command_world_target_count,\n }\n \n if mode == \"all\" or mode == \"demo\" or mode == \"train\":\n # Meaning we are generating the random ReaSCAN train + dev + test splits!\n logger.warning(f\"You are generating data for {mode} splits only!\")\n split_percentage = {\n \"train\": 0.9, \n }\n elif mode == \"all,noval_1,noval_2,noval_3,noval_4\":\n # here we need to define how to check for noval_*\n pass\n elif mode == \"compositional\":\n # Meaning we are generating the random ReaSCAN train + dev + test splits!\n logger.warning(\"You are generating data for all compositional splits!\")\n elif mode == \"\":\n pass # Not implemented!\n \n # Using the full vocabulary.\n intransitive_verbs = [\"walk\"]\n transitive_verbs = [\"push\", \"pull\"]\n adverbs = [\"while zigzagging\", \"while spinning\", \"cautiously\", \"hesitantly\"]\n nouns = [\"circle\", \"cylinder\", \"square\", \"box\"]\n color_adjectives = [\"red\", \"blue\", \"green\", \"yellow\"]\n size_adjectives = [\"big\", \"small\"]\n relative_pronouns = [\"that is\"]\n relation_clauses = [\"in the same row as\", \n \"in the same column as\", \n \"in the same color as\", \n \"in the same shape as\", \n \"in the same size as\",\n \"inside of\"]\n vocabulary = Vocabulary.initialize(intransitive_verbs=intransitive_verbs,\n transitive_verbs=transitive_verbs, adverbs=adverbs, nouns=nouns,\n color_adjectives=color_adjectives,\n size_adjectives=size_adjectives, \n relative_pronouns=relative_pronouns, \n relation_clauses=relation_clauses)\n \n # test out the object vocab\n min_object_size = 1\n max_object_size = 4\n object_vocabulary = ObjectVocabulary(shapes=vocabulary.get_semantic_shapes(),\n colors=vocabulary.get_semantic_colors(),\n min_size=min_object_size, max_size=max_object_size)\n \n # Generating all the core command structs.\n grammer = Grammer(vocabulary)\n \n # Bootup our simulator.\n simulator = Simulator(\n object_vocabulary, vocabulary, \n grid_size=grid_size, \n n_object_max=n_object_max,\n )\n \n command_structs = []\n logger.info(\"Finished loading required modules...\")\n # Sampling all the possible command score structs.\n \n if loading_p4:\n # Currently, we hard-code the pattern!\n grammer_pattern = '$OBJ_0 ^ $OBJ_1 & $OBJ_2 & $OBJ_3'\n logger.info(f\"Including pattern:= {grammer_pattern}...\")\n # Sampling relations\n relations = grammer.sample_object_relation_grammer(\n '$OBJ_0', \n grammer.build_dependency_graph(grammer_pattern))\n for relation in relations:\n obj_pattern_map = relation[0]\n rel_map = relation[1]\n grammer_bindings = grammer.grounding_grammer_with_vocabulary(grammer_pattern, obj_pattern_map, rel_map)\n for obj_map in grammer_bindings:\n # here, we also sample the verb and adverb bindings!\n adverb_enhance_list = vocabulary.get_adverbs()\n adverb_enhance_list += [\"\"]\n command_struct = {\n \"obj_pattern_map\" : obj_pattern_map,\n \"rel_map\" : rel_map,\n \"obj_map\" : obj_map,\n \"grammer_pattern\" : grammer_pattern,\n \"adverb\" : random.choice(adverb_enhance_list),\n \"verb\" : random.choice(vocabulary.get_transitive_verbs() + vocabulary.get_intransitive_verbs()),\n }\n command_structs += [command_struct]\n \n if loading_p3:\n # Currently, we hard-code the pattern!\n grammer_pattern = '$OBJ_0 ^ $OBJ_1 & $OBJ_2'\n logger.info(f\"Including pattern:= {grammer_pattern}...\")\n # Sampling relations\n relations = grammer.sample_object_relation_grammer(\n '$OBJ_0', \n grammer.build_dependency_graph(grammer_pattern))\n for relation in relations:\n obj_pattern_map = relation[0]\n rel_map = relation[1]\n grammer_bindings = grammer.grounding_grammer_with_vocabulary(grammer_pattern, obj_pattern_map, rel_map)\n for obj_map in grammer_bindings:\n # here, we also sample the verb and adverb bindings!\n adverb_enhance_list = vocabulary.get_adverbs()\n adverb_enhance_list += [\"\"]\n command_struct = {\n \"obj_pattern_map\" : obj_pattern_map,\n \"rel_map\" : rel_map,\n \"obj_map\" : obj_map,\n \"grammer_pattern\" : grammer_pattern,\n \"adverb\" : random.choice(adverb_enhance_list),\n \"verb\" : random.choice(vocabulary.get_transitive_verbs() + vocabulary.get_intransitive_verbs()),\n }\n command_structs += [command_struct]\n \n if loading_p2:\n grammer_pattern = '$OBJ_0 ^ $OBJ_1'\n logger.info(f\"Including pattern:= {grammer_pattern}...\")\n # Sampling relations\n relations = grammer.sample_object_relation_grammer(\n '$OBJ_0', \n grammer.build_dependency_graph(grammer_pattern))\n for relation in relations:\n obj_pattern_map = relation[0]\n rel_map = relation[1]\n grammer_bindings = grammer.grounding_grammer_with_vocabulary(grammer_pattern, obj_pattern_map, rel_map)\n for obj_map in grammer_bindings:\n # here, we also sample the verb and adverb bindings!\n adverb_enhance_list = vocabulary.get_adverbs()\n adverb_enhance_list += [\"\"]\n command_struct = {\n \"obj_pattern_map\" : obj_pattern_map,\n \"rel_map\" : rel_map,\n \"obj_map\" : obj_map,\n \"grammer_pattern\" : grammer_pattern,\n \"adverb\" : random.choice(adverb_enhance_list),\n \"verb\" : random.choice(vocabulary.get_transitive_verbs() + vocabulary.get_intransitive_verbs()),\n }\n command_structs += [command_struct]\n \n if loading_p1:\n p1_exhaustive_verb_adverb = True\n # for gSCAN command, we don't need to undersample, they are small!\n grammer_pattern = '$OBJ_0'\n logger.info(f\"Including pattern:= {grammer_pattern}...\")\n # Sampling relations\n relations = grammer.sample_object_relation_grammer(\n '$OBJ_0', \n grammer.build_dependency_graph(grammer_pattern))\n for relation in relations:\n obj_pattern_map = relation[0]\n rel_map = relation[1]\n grammer_bindings = grammer.grounding_grammer_with_vocabulary(grammer_pattern, obj_pattern_map, rel_map)\n for obj_map in grammer_bindings:\n if p1_exhaustive_verb_adverb:\n for adverb in vocabulary.get_adverbs() + [\"\"]:\n for verb in vocabulary.get_transitive_verbs() + vocabulary.get_intransitive_verbs():\n # here, we also sample the verb and adverb bindings!\n command_struct = {\n \"obj_pattern_map\" : obj_pattern_map,\n \"rel_map\" : rel_map,\n \"obj_map\" : obj_map,\n \"grammer_pattern\" : grammer_pattern,\n \"adverb\" : adverb,\n \"verb\" : verb,\n }\n command_structs += [command_struct]\n \n # We only sample these command!\n \"\"\"\n WARNING: beaware that not all command struct can\n be sampled for world-command pair! They may or\n may not fail.\n \"\"\"\n under_sample = True\n if under_sample:\n sampled_command_struct = []\n random.shuffle(command_structs)\n if n_command_struct != -1:\n sampled_command_struct = command_structs[:n_command_struct]\n if args.index_start == -1 or args.index_end == -1:\n pass\n else:\n # we only look at one shard! this is for multiprocess\n logger.info(f\"WARNING: contine with sharding: start at {args.index_start}; end at {args.index_end}\")\n sampled_command_struct = command_structs[args.index_start:args.index_end]\n logger.info(f\"Sampled {len(sampled_command_struct)} from {len(command_structs)} core command structs for pattern={grammer_pattern}.\")\n \n logger.info(f\"Finished sampling core command structs with total {len(sampled_command_struct)}...\")\n \n command_struct_file_path = os.path.join(args.output_dir, f\"command_struct-{args.mode}.txt\")\n formatted_sampled_command_struct = []\n for command_struct in sampled_command_struct:\n formatted_command_struct = {\n \"obj_pattern_map\" : command_struct[\"obj_pattern_map\"],\n \"rel_map\" : [(k, v) for k, v in command_struct[\"rel_map\"].items()],\n \"obj_map\" : command_struct[\"obj_map\"],\n \"grammer_pattern\" : command_struct[\"grammer_pattern\"],\n \"adverb\" : command_struct[\"adverb\"],\n \"verb\" : command_struct[\"verb\"],\n }\n formatted_sampled_command_struct += [formatted_command_struct]\n # dump to the disk.\n with open(command_struct_file_path, \"w\") as fd:\n json.dump(formatted_sampled_command_struct, fd, indent=4)\n logger.info(f\"Saved command struct to {command_struct_file_path} for later use...\")\n \n # print out quick stats on how many command per pattern!\n per_pattern_command_count = {}\n for command_struct in sampled_command_struct:\n grammer_pattern = command_struct[\"grammer_pattern\"]\n if grammer_pattern in per_pattern_command_count.keys():\n per_pattern_command_count[grammer_pattern] += 1\n else:\n per_pattern_command_count[grammer_pattern] = 1\n logger.info(f\"Counts per command pattern: \")\n logger.info(per_pattern_command_count)\n\n # From the struct, let us sample shape world.\n \"\"\"\n We just need a couple more steps beyond this point:\n (1) Sample a world\n (2) Making sure it is valid\n (3) Construct the command, providing determiners\n (4) Generate action sequences to the target\n (5) Get all the action related metadata as gSCAN\n (6) Save it to per command example\n \"\"\"\n \n # We need a way to index the sampled command.\n sampled_command_struct_indexed = OrderedDict({})\n global_command_struct_index = 0\n for command_struct in sampled_command_struct:\n sampled_command_struct_indexed[global_command_struct_index] = command_struct\n global_command_struct_index += 1\n \n root = \"$OBJ_0\"\n per_command_world_counts = OrderedDict({})\n if mode == \"demo\" or mode == \"all\" or mode == \"train\":\n created_examples_by_splits = OrderedDict({\n \"train\" : [],\n })\n else:\n pass\n shaperized_command_struct = []\n per_command_world_unique_check = OrderedDict({})\n \n # Some global control for data quality control.\n global_step = 0\n success_step = 0\n \n # Distractor info logs.\n d_full_relation_count = 0\n d_relation_count = 0\n d_attribute_count = 0\n d_iso_count = 0\n d_random_count = 0\n \n logger.info(f\"Started to generate the dataset...\")\n for command_struct_index, command_struct in sampled_command_struct_indexed.items():\n logger.info(f\"Generating for command struct (seed={seed}): {command_struct_index+1}/{len(sampled_command_struct_indexed)}...\")\n per_command_world_counts[command_struct_index] = 0 # 0 world for each command in the beginning!\n per_command_world_unique_check[command_struct_index] = set([])\n obj_pattern_map = command_struct[\"obj_pattern_map\"]\n rel_map = command_struct[\"rel_map\"]\n obj_map = command_struct[\"obj_map\"]\n grammer_pattern = command_struct[\"grammer_pattern\"]\n verb = command_struct[\"verb\"]\n adverb = command_struct[\"adverb\"]\n # This is the target world number generated for this command\n for n_world_try in range(per_command_world_target_count):\n # How many time we need to retry before we give up?\n at_least_success = False\n for n_retry in range(per_command_world_retry_max):\n global_step += 1\n if success_step == 0:\n denom = 1\n else:\n denom = success_step\n d_full_relation_ratio = 1.0*d_full_relation_count/denom\n d_relation_ratio = 1.0*d_relation_count/denom\n d_attribute_ratio = 1.0*d_attribute_count/denom\n d_iso_ratio = 1.0*d_iso_count/denom\n d_random_ratio = 1.0*d_random_count/denom\n global_success_ratio = 1.0*success_step/global_step\n # logging some very useful information to wandb if avaliable!\n if is_tensorboard:\n if (global_step%logging_interval) == 0:\n wandb.log({'global_success_ratio': global_success_ratio, 'global_step': global_step})\n wandb.log({'current_example_count': success_step, 'global_step': global_step})\n wandb.log({'d_full_relation_ratio': d_full_relation_ratio, 'global_step': global_step})\n wandb.log({'d_relation_ratio': d_relation_ratio, 'global_step': global_step})\n wandb.log({'d_attribute_ratio': d_attribute_ratio, 'global_step': global_step})\n wandb.log({'d_iso_ratio': d_iso_ratio, 'global_step': global_step})\n wandb.log({'d_random_ratio': d_random_ratio, 'global_step': global_step}) \n else:\n if (global_step%(logging_interval*10)) == 0:\n logger.info({'global_success_ratio': global_success_ratio, 'global_step': global_step})\n logger.info({'current_example_count': success_step, 'global_step': global_step})\n logger.info({'d_full_relation_ratio': d_full_relation_ratio, 'global_step': global_step})\n logger.info({'d_relation_ratio': d_relation_ratio, 'global_step': global_step})\n logger.info({'d_attribute_ratio': d_attribute_ratio, 'global_step': global_step})\n logger.info({'d_iso_ratio': d_iso_ratio, 'global_step': global_step})\n logger.info({'d_random_ratio': d_random_ratio, 'global_step': global_step})\n \n if mode == \"demo\":\n sampled_world = simulator.sample_situations_from_grounded_grammer(\n copy.deepcopy(grammer_pattern), \n copy.deepcopy(obj_pattern_map), \n copy.deepcopy(rel_map), \n copy.deepcopy(obj_map),\n is_plot=False,\n include_relation_distractor=args.include_relation_distractor, \n include_attribute_distractor=args.include_attribute_distractor, \n include_isomorphism_distractor=args.include_isomorphism_distractor, \n include_random_distractor=args.include_random_distractor,\n full_relation_probability=args.full_relation_probability,\n debug=False\n ) # This is the minimum settings! You need to turn on attribute always!\n else:\n # Sample a shapeWorld!\n sampled_world = simulator.sample_situations_from_grounded_grammer(\n copy.deepcopy(grammer_pattern), \n copy.deepcopy(obj_pattern_map), \n copy.deepcopy(rel_map), \n copy.deepcopy(obj_map),\n is_plot=False,\n include_relation_distractor=args.include_relation_distractor, \n include_attribute_distractor=args.include_attribute_distractor, \n include_isomorphism_distractor=args.include_isomorphism_distractor, \n include_random_distractor=args.include_random_distractor,\n full_relation_probability=args.full_relation_probability, # ReaSCAN Special: 15 distractors!\n debug=False\n )\n\n # Validate the world is valid!\n graph = ReaSCANGraph(\n objects=sampled_world[\"obj_map\"], \n object_patterns=sampled_world[\"obj_pattern_map\"], \n vocabulary=vocabulary,\n positions=sampled_world[\"pos_map\"], \n referred_object=sampled_world[\"referred_obj\"],\n debug=False\n )\n \n pattern_graph = ReaSCANGraph(\n objects=obj_map, \n object_patterns=None,\n vocabulary=vocabulary,\n relations=rel_map, \n referred_object='$OBJ_0', \n debug=False\n )\n \n potential_referent_target = graph.find_referred_object_super_fast(\n pattern_graph, referred_object='$OBJ_0', \n debug=False\n )\n\n # Save the result if the world is valid!\n \n # This may be to strict, but it ensures 100% correct!\n if len(potential_referent_target) == 1 and '$OBJ_0' in potential_referent_target:\n # A quick world repeat check!\n hash_world_str = hashlib.md5(str(sampled_world[\"situation\"].to_representation()).encode('utf-8')).hexdigest()\n if hash_world_str not in per_command_world_unique_check[command_struct_index]:\n per_command_world_unique_check[command_struct_index].add(hash_world_str)\n else:\n continue # This is highly unlikely, but just to prevent!\n \n # Form the command with grounded determiners!\n obj_determiner_map = graph.find_determiners(\n pattern_graph, \n referred_object='$OBJ_0', \n debug=False,\n )\n \n # we don't check this for P1 and P2?\n \n# valid_determiner = True\n# for k, v in obj_determiner_map.items():\n# if k != '$OBJ_0':\n# if v != \"a\":\n# valid_determiner = False\n# break\n# if not valid_determiner:\n# continue # we should abort and resample!\n \n at_least_success = True\n success_step += 1\n \n command_str = grammer.repre_str_command(\n grammer_pattern, rel_map, obj_map, \n obj_determiner_map, \n verb,\n adverb,\n )\n\n # Form the golden label for the action list!\n is_transitive = False\n if verb in simulator.vocabulary.get_transitive_verbs():\n is_transitive = True\n # Direct walk.\n action = \"walk\" # this is definit!\n primitive_command = simulator.vocabulary.translate_word(action)\n target_position = sampled_world[\"situation\"].target_object.position\n simulator._world.go_to_position(\n position=target_position, manner=adverb, \n primitive_command=primitive_command\n )\n # Object actions.\n if is_transitive:\n semantic_action = simulator.vocabulary.translate_word(verb)\n simulator._world.move_object_to_wall(action=semantic_action, manner=adverb)\n target_commands, _ = simulator._world.get_current_observations()\n \n has_relation_distractor = False\n full_relation_distractor = True\n for rel_bool in sampled_world[\"distractor_switch_map\"][\"relation\"]:\n if rel_bool:\n has_relation_distractor = True\n else:\n full_relation_distractor = False\n \n # Save all relevant information for a task.\n task_struct = OrderedDict({\n \"command\": \",\".join(command_str.split(\" \")),\n \"grammer_pattern\": grammer_pattern,\n \"meaning\": \",\".join(command_str.split(\" \")),\n \"derivation\": grammer_pattern,\n \"situation\": sampled_world[\"situation\"].to_representation(),\n \"target_commands\": \",\".join(target_commands),\n \"verb_in_command\": verb,\n \"adverb_in_command\": adverb,\n \"referred_target\": obj_map[\"$OBJ_0\"],\n \"object_pattern_map\": obj_pattern_map,\n \"relation_map\": [(k, v) for k, v in rel_map.items()],\n \"object_expression\": obj_map,\n \"n_object\": len(sampled_world[\"obj_map\"]),\n \"n_distractor\": len(sampled_world[\"obj_map\"])-len(obj_map),\n \"full_relation_distractor\": full_relation_distractor,\n \"has_relation_distractor\": has_relation_distractor,\n \"has_attribute_distractor\": sampled_world[\"distractor_switch_map\"][\"attribute\"],\n \"has_isomorphism_distractor\": sampled_world[\"distractor_switch_map\"][\"isomorphism\"],\n \"has_random_distractor\": True if sampled_world[\"n_random_distractor\"] != -1 else False,\n \"n_random_distractor\": sampled_world[\"n_random_distractor\"] if sampled_world[\"n_random_distractor\"] != -1 else 0,\n \"relation_distractor_metadata\": sampled_world[\"relation_distractor_metadata\"],\n \"attribute_distractor_metadata\": sampled_world[\"attribute_distractor_metadata\"],\n \"isomorphism_distractor_metadata\": sampled_world[\"isomorphism_distractor_metadata\"],\n \"random_distractor_metadata\": sampled_world[\"random_distractor_metadata\"],\n })\n \n # Record distractor related info\n if task_struct[\"full_relation_distractor\"]:\n d_full_relation_count += 1\n if task_struct[\"has_relation_distractor\"]:\n d_relation_count += 1\n if task_struct[\"has_attribute_distractor\"]:\n d_attribute_count += 1\n if task_struct[\"has_isomorphism_distractor\"]:\n d_iso_count += 1\n if task_struct[\"n_random_distractor\"]:\n d_random_count += 1\n \n # Here, we decide which split we put the example into!\n split = args.mode\n created_examples_by_splits[split].append(task_struct)\n per_command_world_counts[command_struct_index] += 1\n break # break the retry loop!\n if not at_least_success:\n logger.info(f\"WARNING: the success rate for this command is close to 0.0%, skipping...\")\n break # success rate for this comman is ~= 0.0%, let us directly skip\n if save_at_interval and (command_struct_index+1)% save_interal == 0:\n logger.info(f\"Saving data files and statistics to {args.output_dir} for checkpoints...\")\n # Now, we need to save data into the folder\n # along with possible statistics.\n to_save_command_struct = []\n per_command_count = []\n for command_struct_index, count in per_command_world_counts.items():\n per_command_count += [count]\n if count >= 1:\n to_save_command_struct.append(sampled_command_struct_indexed[command_struct_index])\n if save_command_stats:\n _ = get_command_struct_statistics(\n to_save_command_struct, run_name=f\"ReaSCAN-{mode}\", date=args.date, \n split=mode,\n compositional_split=False,\n n_sample=-1,\n output_dir=args.output_dir,\n save_to_disk=True if args.output_dir != \"\" else False,\n wandb=wandb\n )\n \n # wandb.log({\"per_command_world_count\": wandb.Histogram(per_command_count)})\n \n data_file_path = os.path.join(args.output_dir, f\"data-{args.mode}.txt\")\n \n if mode == \"demo\" or mode == \"all\" or mode == \"train\":\n logger.info(f\"total example count={success_step}...\")\n dataset_representation = {\n \"grid_size\": args.grid_size,\n \"type_grammar\": \"ReaSCAN-Grammer\",\n \"min_object_size\": 1,\n \"max_object_size\": 4,\n \"percentage_train\": split_percentage[\"train\"],\n \"examples\": created_examples_by_splits,\n \"intransitive_verbs\": intransitive_verbs,\n \"transitive_verbs\": transitive_verbs,\n \"adverbs\": adverbs,\n \"nouns\": nouns,\n \"color_adjectives\": color_adjectives,\n \"size_adjectives\": size_adjectives,\n \"relative_pronouns\": relative_pronouns,\n \"relation_clauses\": relation_clauses,\n }\n # dump to the disk.\n with open(data_file_path, \"w\") as fd:\n json.dump(dataset_representation, fd, indent=4)\n else:\n pass\n \n # Last round of saving!\n logger.info(f\"Saving FINAL data files and statistics to {args.output_dir}...\")\n # Now, we need to save data into the folder\n # along with possible statistics.\n to_save_command_struct = []\n per_command_count = []\n for command_struct_index, count in per_command_world_counts.items():\n per_command_count += [count]\n if count >= 1:\n to_save_command_struct.append(sampled_command_struct_indexed[command_struct_index])\n if save_command_stats:\n _ = get_command_struct_statistics(\n to_save_command_struct, run_name=f\"ReaSCAN-{mode}\", date=args.date, \n split=mode,\n compositional_split=False,\n n_sample=-1,\n output_dir=args.output_dir,\n save_to_disk=True if args.output_dir != \"\" else False,\n wandb=wandb\n )\n\n # wandb.log({\"per_command_world_count\": wandb.Histogram(per_command_count)})\n\n data_file_path = os.path.join(args.output_dir, f\"data-{args.mode}.txt\")\n\n if mode == \"demo\" or mode == \"all\" or mode == \"train\":\n logger.info(f\"total example count={success_step}...\")\n dataset_representation = {\n \"grid_size\": args.grid_size,\n \"type_grammar\": \"ReaSCAN-Grammer\",\n \"min_object_size\": 1,\n \"max_object_size\": 4,\n \"percentage_train\": split_percentage[\"train\"],\n \"examples\": created_examples_by_splits,\n \"intransitive_verbs\": intransitive_verbs,\n \"transitive_verbs\": transitive_verbs,\n \"adverbs\": adverbs,\n \"nouns\": nouns,\n \"color_adjectives\": color_adjectives,\n \"size_adjectives\": size_adjectives,\n \"relative_pronouns\": relative_pronouns,\n \"relation_clauses\": relation_clauses,\n }\n # dump to the disk.\n with open(data_file_path, \"w\") as fd:\n json.dump(dataset_representation, fd, indent=4)\n else:\n pass\n \n logger.info(\"==FINISH==\")\n \n if args.is_tensorboard:\n # end wandb\n wandb.finish()",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code"
]
] |
4ab6c78788ff3950881e7eb9e6aa18f400c75391
| 738,832 |
ipynb
|
Jupyter Notebook
|
data_analysis.ipynb
|
open-pisciculture/pisciculture-db
|
810175f6ced91090c4200da3cf982ede6af2026e
|
[
"MIT"
] | null | null | null |
data_analysis.ipynb
|
open-pisciculture/pisciculture-db
|
810175f6ced91090c4200da3cf982ede6af2026e
|
[
"MIT"
] | null | null | null |
data_analysis.ipynb
|
open-pisciculture/pisciculture-db
|
810175f6ced91090c4200da3cf982ede6af2026e
|
[
"MIT"
] | null | null | null | 522.142756 | 166,866 | 0.930708 |
[
[
[
"# Data Analysis\n\nHere we need a `.csv` file in order to do the desired analysis.",
"_____no_output_____"
]
],
[
[
"import json\nimport numpy as np\nimport matplotlib.pyplot as plt\nfrom datetime import datetime\nimport pandas as pd\nimport seaborn as sns",
"_____no_output_____"
],
[
"df = pd.read_csv('mergedDatalogger_formatted_data_nov1722.csv')\n\ndf.head()",
"_____no_output_____"
],
[
"df = pd.read_csv('mergedDatalogger_formatted_data_nov1722.csv')\ndf[\"datetime\"] = pd.to_datetime(df[\"datetime\"])\n# df = df.set_index('datetime')\ndel df[\"Datetime\"]\ndf[\"approx datetime\"] = df[\"datetime\"].round('1min') # redondeamos a los X mins más cercanos (X según frec. de los videos)\n\ndel df[\"Sigma T[sT]\"]\n# df = df.sort_index()\ndf.head(10)",
"_____no_output_____"
]
],
[
[
"## Gráficas Datos Datalogger",
"_____no_output_____"
]
],
[
[
"df.columns",
"_____no_output_____"
],
[
"df = pd.read_csv('mergedDatalogger_formatted_data_nov1722.csv')\ndf[\"datetime\"] = pd.to_datetime(df[\"Datetime\"])\nplt.figure(figsize=(20,15))\nplt.subplot(4,1,1)\nplt.scatter(df['datetime'], df['D.O.[mg/L]'], s=0.1)\nplt.subplot(4,1,2)\nplt.scatter(df['datetime'], df['pH '], s=0.1)\nplt.subplot(4,1,3)\nplt.scatter(df['datetime'], df['Temp'], s=0.1)\nplt.subplot(4,1,4)\nplt.scatter(df['datetime'], df['EC Abs'], s=0.1)\n\n# df = df.set_index('datetime')\n# df[['Temp', 'pH ', 'D.O.[mg/L]', 'EC Abs']].plot(subplots=True, figsize=(10,10))",
"_____no_output_____"
],
[
"df = pd.read_csv('datalogger_formatted_data_nov17.csv')\ndf[\"datetime\"] = pd.to_datetime(df[\"Datetime\"])\ndf = df.set_index('datetime')\ndf[['Temp', 'pH ', 'D.O.[mg/L]', 'EC Abs']].plot(subplots=True, figsize=(10,10))",
"_____no_output_____"
],
[
"df = pd.read_csv('datalogger_formatted_data_nov22.csv')\ndf[\"datetime\"] = pd.to_datetime(df[\"Datetime\"])\ndf = df.set_index('datetime')\ndf[['Temp', 'pH ', 'D.O.[mg/L]', 'EC Abs uS/cm']].plot(subplots=True, figsize=(10,10))",
"_____no_output_____"
],
[
"start_time = df.head(1)['datetime']\nend_time = df.tail(1)['datetime']\n\nprint(f\"Data starts at: {start_time}\")\nprint(f\"Data ends at: {end_time}\")",
"Data starts at: 0 2021-11-22 12:16:44\nName: datetime, dtype: datetime64[ns]\nData ends at: 19960 2021-11-23 16:00:04\nName: datetime, dtype: datetime64[ns]\n"
],
[
"#understanding correlation\nplt.figure(figsize = (15,9))\nsns.heatmap(df.corr(), annot = True)",
"_____no_output_____"
]
],
[
[
"## Strongly Correlated Variables",
"_____no_output_____"
]
],
[
[
"sns.scatterplot(x=df[\"Dissolved Oxygen\"], y=df[\"Ambient Temperature\"], data=df)",
"_____no_output_____"
],
[
"sns.scatterplot(x=df[\"Humidity\"], y=df[\"Ambient Temperature\"], data=df)",
"_____no_output_____"
],
[
"sns.scatterplot(x=df[\"Humidity\"], y=df[\"Dissolved Oxygen\"], data=df)",
"_____no_output_____"
],
[
"scatter = sns.scatterplot(x=df[\"Water Temperature\"], y=df[\"Total Dissolved Solids\"], data=df)\nscatter.set_ylim(top=400);",
"_____no_output_____"
]
],
[
[
"## Weakly/Not at all Corelated Variables",
"_____no_output_____"
]
],
[
[
"scatter = sns.scatterplot(x=df[\"Water Level\"], y=df[\"Rain\"], data=df)\nscatter.set_xlim(left=25, right=40)",
"_____no_output_____"
]
],
[
[
"# Merging Video Data with Physicochemical Vars",
"_____no_output_____"
]
],
[
[
"df = pd.read_csv('mergedDatalogger_formatted_data_nov1722.csv')\ndf[\"datetime\"] = pd.to_datetime(df[\"datetime\"])\n# df = df.set_index('datetime')\ndel df[\"Datetime\"]\ndf[\"approx datetime\"] = df[\"datetime\"].round('1min') # redondeamos a los X mins más cercanos (X según frec. de los videos)\n\ndel df[\"Sigma T[sT]\"]\ndf.head(10)",
"_____no_output_____"
],
[
"video_df = pd.read_csv('video_data_Nov1718_Nov2223.csv')\nvideo_df[\"datetime\"] = pd.to_datetime(video_df[\"timestamp\"])\ndel video_df[\"timestamp\"]\ndel video_df['Video Index']\nvideo_df[\"approx datetime\"] = video_df[\"datetime\"].round('1min') # redondeamos a los X mins más cercanos (X según frec. de los videos)\n# video_df = video_df.set_index('datetime')\n\nvideo_df = video_df.sort_values('datetime', ignore_index=True)\nvideo_df.head()",
"_____no_output_____"
],
[
"df.head()",
"_____no_output_____"
],
[
"merged_df = pd.merge(df, video_df, on=\"approx datetime\")\ndel merged_df['datetime_x'] # (Original formatted_data_ timestamp) Column with the same name\ndel merged_df['datetime_y'] # (Original video_data timestamp) Column with the same name\nmerged_df.head()",
"_____no_output_____"
],
[
"#understanding correlation\nplt.figure(figsize = (15,9))\nsns.heatmap(merged_df.corr(), annot = True)",
"_____no_output_____"
],
[
"scatter = sns.scatterplot(x=merged_df[\"Average Pairwise Distance\"], y=merged_df[\"Dissolved Oxygen\"], data=df)\n# scatter.set_ylim(top=400);",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab6d2a73439431a58b7371ff41e533fcc430a9c
| 20,188 |
ipynb
|
Jupyter Notebook
|
examples/01a_forecasting_sklearn.ipynb
|
0saurabh0/sktime
|
dddcd2c31f6400db1ca791c0d16d549d756ccab5
|
[
"BSD-3-Clause"
] | null | null | null |
examples/01a_forecasting_sklearn.ipynb
|
0saurabh0/sktime
|
dddcd2c31f6400db1ca791c0d16d549d756ccab5
|
[
"BSD-3-Clause"
] | null | null | null |
examples/01a_forecasting_sklearn.ipynb
|
0saurabh0/sktime
|
dddcd2c31f6400db1ca791c0d16d549d756ccab5
|
[
"BSD-3-Clause"
] | null | null | null | 34.806897 | 394 | 0.6277 |
[
[
[
"# Forecasting with sktime - appendix: forecasting, supervised regression, and pitfalls in confusing the two\n\nThis notebook provides some supplementary explanation about the relation between forecasting as implemented in `sktime`, and the very common supervised prediction tasks as supported by `scikit-learn` and similar toolboxes.\n\nKey points discussed in this notebook:\n\n* forecasting is not the same as supervised prediction\n* even though forecasting can be \"solved\" by algorithms for supervised prediction, this is indirect and requires careful composition\n* from an interface perspective, this is correctly formulated as \"reduction\", i.e., use of a supervised predictor as a component within a forecaster\n* there are a number of pitfalls if this is manually done - such as, over-optimistic performance evaluation, information leakage, or \"predicting the past\" type errors",
"_____no_output_____"
]
],
[
[
"# general imports\nimport numpy as np\nimport pandas as pd",
"_____no_output_____"
]
],
[
[
"## The pitfalls of mis-diagnosing forecasting as supervised regression\n\nA common mistake is to mis-identify a forecasting problem as supervised regression - after all, in both we predict numbers, so surely this must be the same thing?\n\nIndeed we predict numbers in both, but the set-up is different:\n\n* in supervised regression, we predict *label/target variables* from *feature variables*, in a cross-sectional set-up. This is after training on label/feature examples.\n* in forecasting, we predict *future values* from *past values*, of *the same variable*, in a temporal/sequential set-up. This is after training on the past.\n\nIn the common data frame representation:\n\n* in supervised regression, we predict entries in a column from other columns. For this, we mainly make use of the statistical relation between those columns, leart from examples of complete rows. The rows are all assumed exchangeable.\n* in forecasting, we predict new rows, assuming temporal ordering in the rows. For this, we mainly make use of the statistical relation between previous and subsequent rows, learnt from the example of the observed sequence of rows. The rows are not exchangeable, but in temporal sequence.\n\nTODO: add a nice picture on what is predicted from what, arrows and all. Contributions are welcome.",
"_____no_output_____"
],
[
"### Pitfall 1: over-optimism in performance evaluation, false confidence in \"broken\" forecasters\n\nConfusing the two tasks may lead to information leakage, and over-optimistic performance evaluation. This is because in supervised regression the ordering of rows does not matter, and train/test split is usually performed uniformly. In forecasting, the ordering does matter, both in training and in evaluation.\n\nAs subtle as it seems, this may have major practical consequences - since it can lead to the mistaken belief that a \"broken\" method is performant, which can cause damage to health, property, and other assets in real-life deployment.\n\nThe example below shows \"problematic\" performance estimation, when mistakenly using the regression evaluation workflow for forecasting.",
"_____no_output_____"
]
],
[
[
"from sklearn.model_selection import train_test_split\n\nfrom sktime.datasets import load_airline\nfrom sktime.forecasting.model_selection import temporal_train_test_split\nfrom sktime.utils.plotting import plot_series",
"_____no_output_____"
],
[
"y = load_airline()",
"_____no_output_____"
],
[
"y_train, y_test = train_test_split(y)\nplot_series(y_train.sort_index(), y_test.sort_index(), labels=[\"y_train\", \"y_test\"]);",
"_____no_output_____"
]
],
[
[
"This leads to leakage:\n\n> The data you are using to train a machine learning algorithm happens to have the information you are trying to predict.\n\nBut `train_test_split(y, shuffle=False)` works, which is what `temporal_train_test_split(y)` does in `sktime`:",
"_____no_output_____"
]
],
[
[
"y_train, y_test = temporal_train_test_split(y)\nplot_series(y_train, y_test, labels=[\"y_train\", \"y_test\"]);",
"_____no_output_____"
]
],
[
[
"### Pitfall 2: obscure data manipulations, brittle boilerplate code to apply regressors\n\nIt is common practice to apply supervised regressors after transforming the data for forecasting, through lagging - for example, in auto-regressive reduction strategies.\n\nTwo important pitfalls appear right at the start:\n\n* a lot of boilerplate code has to be written to transform the data to make it ready for fitting - this is highly error prone\n* there are a number of implicit hyper-parameters here, such as window and lag size. If done without caution, these are not explicit or tracked in the experiment, which can lead to \"p-value hacking\".\n\nBelow is an example of such boilerplate code to demonstrate this. The code is closely modelled on the R code used in the [M4 competition](https://github.com/Mcompetitions/M4-methods):",
"_____no_output_____"
]
],
[
[
"# suppose we want to predict 3 years ahead\nfh = np.arange(1, 37)",
"_____no_output_____"
],
[
"# slightly modified code from the M4 competition\ndef split_into_train_test(data, in_num, fh):\n \"\"\"\n Splits the series into train and test sets.\n\n Each step takes multiple points as inputs\n :param data: an individual TS\n :param fh: number of out of sample points\n :param in_num: number of input points for the forecast\n :return:\n \"\"\"\n train, test = data[:-fh], data[-(fh + in_num) :]\n x_train, y_train = train[:-1], np.roll(train, -in_num)[:-in_num]\n x_test, y_test = test[:-1], np.roll(test, -in_num)[:-in_num]\n # x_test, y_test = train[-in_num:], np.roll(test, -in_num)[:-in_num]\n\n # reshape input to be [samples, time steps, features]\n # (N-NF samples, 1 time step, 1 feature)\n x_train = np.reshape(x_train, (-1, 1))\n x_test = np.reshape(x_test, (-1, 1))\n temp_test = np.roll(x_test, -1)\n temp_train = np.roll(x_train, -1)\n for x in range(1, in_num):\n x_train = np.concatenate((x_train[:-1], temp_train[:-1]), 1)\n x_test = np.concatenate((x_test[:-1], temp_test[:-1]), 1)\n temp_test = np.roll(temp_test, -1)[:-1]\n temp_train = np.roll(temp_train, -1)[:-1]\n\n return x_train, y_train, x_test, y_test",
"_____no_output_____"
],
[
"# here we split the time index, rather than the actual values,\n# to show how we split the windows\nfeature_window, target_window, _, _ = split_into_train_test(\n np.arange(len(y)), 10, len(fh)\n)",
"_____no_output_____"
]
],
[
[
"To better understand the prior data transformation, we can look at how we can split the training series into windows. Here we show the generated windows expressed as integer indices:",
"_____no_output_____"
]
],
[
[
"feature_window[:5, :]",
"_____no_output_____"
],
[
"target_window[:5]",
"_____no_output_____"
],
[
"# now we can split the actual values of the time series\nx_train, y_train, x_test, y_test = split_into_train_test(y.values, 10, len(fh))\nprint(x_train.shape, y_train.shape)",
"_____no_output_____"
],
[
"from sklearn.ensemble import RandomForestRegressor\n\nmodel = RandomForestRegressor()\nmodel.fit(x_train, y_train)",
"_____no_output_____"
]
],
[
[
"To reiterate the potential pitfalls here:\n\n> The manual requires a lot of hand-written code which is often error-prone, not modular and not tuneable.\n\n> These steps involve a number of implicit hyper-parameters:\n> * the way you slice the time series into windows (e.g. the window length)\n> * the way you generate forecasts (recursive strategy, direct strategy, other hybrid strategies)",
"_____no_output_____"
],
[
"### Pitfall 3: Given a fitted regression algorithm, how can we generate forecasts?\n\nThe next important pitfall comes at the end:\n\nif making predictions along the \"manual route\" for supervised regressors, the supervised regressor's outputs have to be transformed back into forecasts. This is easily forgotten, and invites errors in forecasts and evaluation (see pitfall no.1) - especially, if one does not cleanly keep track of which data is known at what time, or how to invert the transformation made in fitting.\n\nA naive user might now proceed like this:",
"_____no_output_____"
]
],
[
[
"print(x_test.shape, y_test.shape)\n\n# add back time index to y_test\ny_test = pd.Series(y_test, index=y.index[-len(fh) :])",
"_____no_output_____"
],
[
"y_pred = model.predict(x_test)",
"_____no_output_____"
],
[
"from sktime.performance_metrics.forecasting import mean_absolute_percentage_error\n\nmean_absolute_percentage_error(\n y_test, pd.Series(y_pred, index=y_test.index), symmetric=False\n)",
"_____no_output_____"
]
],
[
[
"So easy, so wrong ... but what's the problem here? It's a bit subtle and not easy to spot:\n\n> We actually don't make a multi-step-ahead forecast up to the 36th step ahead. Instead, we make 36 single-step-ahead forecasts always using the most recent data. But that's a solution to a different learning task!\n\nTo fix this problem, we could write some code to do this recursively as in the M4 competition:",
"_____no_output_____"
]
],
[
[
"# slightly modified code from the M4 study\npredictions = []\nlast_window = x_train[-1, :].reshape(1, -1) # make it into 2d array\n\nlast_prediction = model.predict(last_window)[0] # take value from array\n\nfor i in range(len(fh)):\n # append prediction\n predictions.append(last_prediction)\n\n # update last window using previously predicted value\n last_window[0] = np.roll(last_window[0], -1)\n last_window[0, (len(last_window[0]) - 1)] = last_prediction\n\n # predict next step ahead\n last_prediction = model.predict(last_window)[0]\n\ny_pred_rec = pd.Series(predictions, index=y_test.index)",
"_____no_output_____"
],
[
"from sktime.performance_metrics.forecasting import mean_absolute_percentage_error\n\nmean_absolute_percentage_error(\n y_test, pd.Series(y_pred_rec, index=y_test.index), symmetric=False\n)",
"_____no_output_____"
]
],
[
[
"To summarize the potential pitfalls here:\n\n> Obtaining regressor predictions and converting them back into forecasts is non-trivial and error prone:\n> * some boilerplate code needs to be written, which just as in pitfall no.2 introduces potential for problems\n> * it isn't exactly obvious that this boilerplate code had to be written in the first place, creating a subtle failure point",
"_____no_output_____"
],
[
"### How does `sktime` help avoid the above pitfalls?\n\n`sktime` mitigates the above pitfalls by:\n\n* the unified interface for forecasters - any strategy to produce forecasts is a forecaster. Through the unified interface, forecasters are directly compatible with deployment and evaluation workflows appropriate for forecasters.\n* its declarative specification interface that minimizes boilerplate code - it's minimized to the bare necessities to tell `sktime` which forecaster you want to build\n\nNevertheless, `sktime` aims to be flexible, and tries to avoid to railroad the user into specific methodological choices.",
"_____no_output_____"
]
],
[
[
"from sklearn.neighbors import KNeighborsRegressor\n\nfrom sktime.forecasting.compose import make_reduction",
"_____no_output_____"
],
[
"# declarative forecaster specification - just two lines!\nregressor = KNeighborsRegressor(n_neighbors=1)\nforecaster = make_reduction(regressor, window_length=15, strategy=\"recursive\")",
"_____no_output_____"
],
[
"forecaster.fit(y_train)\ny_pred = forecaster.predict(fh)",
"_____no_output_____"
]
],
[
[
"... and that's it!\n\nNote that there is no `x_train` or other boilerplate artefacts, since construction of the lagged features and other boilerplate code are taken care of by the forecaster internally.\n\nFor more details on the `sktime` composition interface, refer to Section 3 of the main forecasting tutorial.",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
]
] |
4ab6d5d4ffc955ef1e6d361ccd1c516c341a03a8
| 966,530 |
ipynb
|
Jupyter Notebook
|
numpy/notebooks/01-Diabetes Data Analysis.ipynb
|
scicomforbio/ISCB20.06
|
399c776f56319f7b03ffd13750c3f7d7349c136d
|
[
"CC0-1.0"
] | 1 |
2020-12-09T19:51:53.000Z
|
2020-12-09T19:51:53.000Z
|
numpy/notebooks/01-Diabetes Data Analysis.ipynb
|
scicomforbio/ISCB20.06
|
399c776f56319f7b03ffd13750c3f7d7349c136d
|
[
"CC0-1.0"
] | null | null | null |
numpy/notebooks/01-Diabetes Data Analysis.ipynb
|
scicomforbio/ISCB20.06
|
399c776f56319f7b03ffd13750c3f7d7349c136d
|
[
"CC0-1.0"
] | 1 |
2020-12-09T19:56:51.000Z
|
2020-12-09T19:56:51.000Z
| 1,299.099462 | 860,432 | 0.951999 |
[
[
[
"# library \nimport numpy as np \nimport pandas as pd \nimport matplotlib.pyplot as plt \nimport seaborn as sns \n\n# options \npd.set_option('precision', 3)\npd.set_option('display.max_columns', 20)\npd.set_option('display.max_rows', 10000)\nsns.set_style('whitegrid')\n\n# visualization\nplt.rcParams['figure.figsize'] = (10,6) \nplt.rcParams['font.size'] = 14 \n%matplotlib inline ",
"_____no_output_____"
],
[
"df = pd.read_csv('../data/diabetes.csv')",
"_____no_output_____"
],
[
"df.head() ",
"_____no_output_____"
],
[
"df.shape ",
"_____no_output_____"
],
[
"df.info() ",
"<class 'pandas.core.frame.DataFrame'>\nRangeIndex: 768 entries, 0 to 767\nData columns (total 9 columns):\n # Column Non-Null Count Dtype \n--- ------ -------------- ----- \n 0 Pregnancies 768 non-null int64 \n 1 Glucose 768 non-null int64 \n 2 BloodPressure 768 non-null int64 \n 3 SkinThickness 768 non-null int64 \n 4 Insulin 768 non-null int64 \n 5 BMI 768 non-null float64\n 6 DiabetesPedigreeFunction 768 non-null float64\n 7 Age 768 non-null int64 \n 8 Outcome 768 non-null int64 \ndtypes: float64(2), int64(7)\nmemory usage: 54.1 KB\n"
],
[
"df.columns",
"_____no_output_____"
],
[
"df.describe() ",
"_____no_output_____"
],
[
"sns.distplot(df['BloodPressure'])",
"_____no_output_____"
],
[
"sns.pairplot(df)",
"_____no_output_____"
],
[
"df.corr() ",
"_____no_output_____"
],
[
"sns.heatmap(df.corr(), annot=True)",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab6e161ff639dc256e3226d10cb89ec1749fb81
| 24,119 |
ipynb
|
Jupyter Notebook
|
src/.ipynb_checkpoints/Taxi_Driver-checkpoint.ipynb
|
himanshu-02/rlagents
|
5acbacfd35e03c8d2111dd3d85bf593c1dd0a8fb
|
[
"MIT"
] | 5 |
2021-10-31T09:57:28.000Z
|
2022-01-16T17:07:57.000Z
|
src/.ipynb_checkpoints/Taxi_Driver-checkpoint.ipynb
|
himanshu-02/rlagents
|
5acbacfd35e03c8d2111dd3d85bf593c1dd0a8fb
|
[
"MIT"
] | null | null | null |
src/.ipynb_checkpoints/Taxi_Driver-checkpoint.ipynb
|
himanshu-02/rlagents
|
5acbacfd35e03c8d2111dd3d85bf593c1dd0a8fb
|
[
"MIT"
] | 1 |
2022-01-06T12:37:05.000Z
|
2022-01-06T12:37:05.000Z
| 28.242389 | 465 | 0.321448 |
[
[
[
"import gym\nimport numpy as np\nimport math",
"_____no_output_____"
]
],
[
[
"Description:\n There are four designated locations in the grid world indicated by R(ed), G(reen), Y(ellow), and B(lue). When the episode starts, the taxi starts off at a random square and the passenger is at a random location. The taxi drives to the passenger's location, picks up the passenger, drives to the passenger's destination (another one of the four specified locations), and then drops off the passenger. Once the passenger is dropped off, the episode ends.\n Observations:\n There are 500 discrete states since there are 25 taxi positions, 5 possible locations of the passenger (including the case when the passenger is in the taxi), and 4 destination locations.\n Note that there are 400 states that can actually be reached during an episode. The missing states correspond to situations in which the passenger is at the same location as their destination, as this typically signals the end of an episode.\n Four additional states can be observed right after a successful episodes, when both the passenger and the taxi are at the destination.\n This gives a total of 404 reachable discrete states.\n Passenger locations:\n - 0: R(ed)\n - 1: G(reen)\n - 2: Y(ellow)\n - 3: B(lue)\n - 4: in taxi\n Destinations:\n - 0: R(ed)\n - 1: G(reen)\n - 2: Y(ellow)\n - 3: B(lue)\n Actions:\n There are 6 discrete deterministic actions:\n - 0: move south\n - 1: move north\n - 2: move east\n - 3: move west\n - 4: pickup passenger\n - 5: drop off passenger\n Rewards:\n There is a default per-step reward of -1,\n except for delivering the passenger, which is +20,\n or executing \"pickup\" and \"drop-off\" actions illegally, which is -10.\n Rendering:\n - blue: passenger\n - magenta: destination\n - yellow: empty taxi\n - green: full taxi\n - other letters (R, G, Y and B): locations for passengers and destinations",
"_____no_output_____"
]
],
[
[
"env = gym.make(\"Taxi-v3\")\nq_table = np.zeros([env.observation_space.n, env.action_space.n])",
"_____no_output_____"
],
[
"env.render()",
"+---------+\n|R: | : :\u001b[34;1mG\u001b[0m|\n| : | : :\u001b[43m \u001b[0m|\n| : : : : |\n| | : | : |\n|Y| : |\u001b[35mB\u001b[0m: |\n+---------+\n\n"
],
[
"env.reset()",
"_____no_output_____"
],
[
"\"\"\"Training the agent\"\"\"\n\nimport random\nfrom IPython.display import clear_output\n\n# Hyperparameters\nalpha = 0.1\ngamma = 0.6\nepsilon = 0.1\n\n# For plotting metrics\nall_epochs = []\nall_penalties = []\n\nfor i in range(1, 100001):\n state = env.reset()\n\n epochs, penalties, reward, = 0, 0, 0\n done = False\n \n while not done:\n if random.uniform(0, 1) < epsilon:\n action = env.action_space.sample() # Explore action space\n else:\n action = np.argmax(q_table[state]) # Exploit learned values\n\n next_state, reward, done, info = env.step(action) \n \n old_value = q_table[state, action]\n next_max = np.max(q_table[next_state])\n \n new_value = (1 - alpha) * old_value + alpha * (reward + gamma * next_max)\n q_table[state, action] = new_value\n\n if reward == -10:\n penalties += 1\n\n state = next_state\n epochs += 1\n \n if i % 100 == 0:\n clear_output(wait=True)\n print(f\"Episode: {i}\")\n\nprint(\"Training finished.\\n\")\n",
"Episode: 100000\nTraining finished.\n\n"
],
[
"q_table[328]",
"_____no_output_____"
],
[
"\"\"\"Evaluate agent's performance after Q-learning\"\"\"\n\ntotal_epochs, total_penalties = 0, 0\nepisodes = 5\n\nfor _ in range(episodes):\n state = env.reset()\n epochs, penalties, reward = 0, 0, 0\n \n done = False\n \n while not done:\n action = np.argmax(q_table[state])\n state, reward, done, info = env.step(action)\n env.render()\n\n if reward == -10:\n penalties += 1\n\n epochs += 1\n\n total_penalties += penalties\n total_epochs += epochs\n\nprint(f\"Results after {episodes} episodes:\")\nprint(f\"Average timesteps per episode: {total_epochs / episodes}\")\nprint(f\"Average penalties per episode: {total_penalties / episodes}\")\n",
"+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : : : : |\n|\u001b[43m \u001b[0m| : | : |\n|Y| : |\u001b[34;1mB\u001b[0m: |\n+---------+\n (North)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n|\u001b[43m \u001b[0m: : : : |\n| | : | : |\n|Y| : |\u001b[34;1mB\u001b[0m: |\n+---------+\n (North)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| :\u001b[43m \u001b[0m: : : |\n| | : | : |\n|Y| : |\u001b[34;1mB\u001b[0m: |\n+---------+\n (East)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : :\u001b[43m \u001b[0m: : |\n| | : | : |\n|Y| : |\u001b[34;1mB\u001b[0m: |\n+---------+\n (East)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : : :\u001b[43m \u001b[0m: |\n| | : | : |\n|Y| : |\u001b[34;1mB\u001b[0m: |\n+---------+\n (East)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : : : : |\n| | : |\u001b[43m \u001b[0m: |\n|Y| : |\u001b[34;1mB\u001b[0m: |\n+---------+\n (South)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |\u001b[34;1m\u001b[43mB\u001b[0m\u001b[0m: |\n+---------+\n (South)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |\u001b[42mB\u001b[0m: |\n+---------+\n (Pickup)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : : : : |\n| | : |\u001b[42m_\u001b[0m: |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : : :\u001b[42m_\u001b[0m: |\n| | : | : |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | :\u001b[42m_\u001b[0m: |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|R: | :\u001b[42m_\u001b[0m:\u001b[35mG\u001b[0m|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|R: | : :\u001b[35m\u001b[42mG\u001b[0m\u001b[0m|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (East)\n+---------+\n|R: | : :\u001b[35m\u001b[34;1m\u001b[43mG\u001b[0m\u001b[0m\u001b[0m|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (Dropoff)\n+---------+\n|\u001b[34;1mR\u001b[0m: | : :G|\n| : | : : |\n| : :\u001b[43m \u001b[0m: : |\n| | : | : |\n|Y| : |\u001b[35mB\u001b[0m: |\n+---------+\n (West)\n+---------+\n|\u001b[34;1mR\u001b[0m: | : :G|\n| : | : : |\n| :\u001b[43m \u001b[0m: : : |\n| | : | : |\n|Y| : |\u001b[35mB\u001b[0m: |\n+---------+\n (West)\n+---------+\n|\u001b[34;1mR\u001b[0m: | : :G|\n| :\u001b[43m \u001b[0m| : : |\n| : : : : |\n| | : | : |\n|Y| : |\u001b[35mB\u001b[0m: |\n+---------+\n (North)\n+---------+\n|\u001b[34;1mR\u001b[0m: | : :G|\n|\u001b[43m \u001b[0m: | : : |\n| : : : : |\n| | : | : |\n|Y| : |\u001b[35mB\u001b[0m: |\n+---------+\n (West)\n+---------+\n|\u001b[34;1m\u001b[43mR\u001b[0m\u001b[0m: | : :G|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |\u001b[35mB\u001b[0m: |\n+---------+\n (North)\n+---------+\n|\u001b[42mR\u001b[0m: | : :G|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |\u001b[35mB\u001b[0m: |\n+---------+\n (Pickup)\n+---------+\n|R: | : :G|\n|\u001b[42m_\u001b[0m: | : : |\n| : : : : |\n| | : | : |\n|Y| : |\u001b[35mB\u001b[0m: |\n+---------+\n (South)\n+---------+\n|R: | : :G|\n| : | : : |\n|\u001b[42m_\u001b[0m: : : : |\n| | : | : |\n|Y| : |\u001b[35mB\u001b[0m: |\n+---------+\n (South)\n+---------+\n|R: | : :G|\n| : | : : |\n| :\u001b[42m_\u001b[0m: : : |\n| | : | : |\n|Y| : |\u001b[35mB\u001b[0m: |\n+---------+\n (East)\n+---------+\n|R: | : :G|\n| : | : : |\n| : :\u001b[42m_\u001b[0m: : |\n| | : | : |\n|Y| : |\u001b[35mB\u001b[0m: |\n+---------+\n (East)\n+---------+\n|R: | : :G|\n| : | : : |\n| : : :\u001b[42m_\u001b[0m: |\n| | : | : |\n|Y| : |\u001b[35mB\u001b[0m: |\n+---------+\n (East)\n+---------+\n|R: | : :G|\n| : | : : |\n| : : : : |\n| | : |\u001b[42m_\u001b[0m: |\n|Y| : |\u001b[35mB\u001b[0m: |\n+---------+\n (South)\n+---------+\n|R: | : :G|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |\u001b[35m\u001b[42mB\u001b[0m\u001b[0m: |\n+---------+\n (South)\n+---------+\n|R: | : :G|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |\u001b[35m\u001b[34;1m\u001b[43mB\u001b[0m\u001b[0m\u001b[0m: |\n+---------+\n (Dropoff)\n+---------+\n|\u001b[35mR\u001b[0m: | : :G|\n| :\u001b[43m \u001b[0m| : : |\n| : : : : |\n| | : | : |\n|Y| : |\u001b[34;1mB\u001b[0m: |\n+---------+\n (South)\n+---------+\n|\u001b[35mR\u001b[0m: | : :G|\n| : | : : |\n| :\u001b[43m \u001b[0m: : : |\n| | : | : |\n|Y| : |\u001b[34;1mB\u001b[0m: |\n+---------+\n (South)\n+---------+\n|\u001b[35mR\u001b[0m: | : :G|\n| : | : : |\n| : :\u001b[43m \u001b[0m: : |\n| | : | : |\n|Y| : |\u001b[34;1mB\u001b[0m: |\n+---------+\n (East)\n+---------+\n|\u001b[35mR\u001b[0m: | : :G|\n| : | : : |\n| : : :\u001b[43m \u001b[0m: |\n| | : | : |\n|Y| : |\u001b[34;1mB\u001b[0m: |\n+---------+\n (East)\n+---------+\n|\u001b[35mR\u001b[0m: | : :G|\n| : | : : |\n| : : : : |\n| | : |\u001b[43m \u001b[0m: |\n|Y| : |\u001b[34;1mB\u001b[0m: |\n+---------+\n (South)\n+---------+\n|\u001b[35mR\u001b[0m: | : :G|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |\u001b[34;1m\u001b[43mB\u001b[0m\u001b[0m: |\n+---------+\n (South)\n+---------+\n|\u001b[35mR\u001b[0m: | : :G|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |\u001b[42mB\u001b[0m: |\n+---------+\n (Pickup)\n+---------+\n|\u001b[35mR\u001b[0m: | : :G|\n| : | : : |\n| : : : : |\n| | : |\u001b[42m_\u001b[0m: |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|\u001b[35mR\u001b[0m: | : :G|\n| : | : : |\n| : : :\u001b[42m_\u001b[0m: |\n| | : | : |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|\u001b[35mR\u001b[0m: | : :G|\n| : | : : |\n| : :\u001b[42m_\u001b[0m: : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (West)\n+---------+\n|\u001b[35mR\u001b[0m: | : :G|\n| : | : : |\n| :\u001b[42m_\u001b[0m: : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (West)\n+---------+\n|\u001b[35mR\u001b[0m: | : :G|\n| : | : : |\n|\u001b[42m_\u001b[0m: : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (West)\n+---------+\n|\u001b[35mR\u001b[0m: | : :G|\n|\u001b[42m_\u001b[0m: | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|\u001b[35m\u001b[42mR\u001b[0m\u001b[0m: | : :G|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|\u001b[35m\u001b[34;1m\u001b[43mR\u001b[0m\u001b[0m\u001b[0m: | : :G|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (Dropoff)\n+---------+\n|\u001b[34;1mR\u001b[0m: | : :\u001b[35mG\u001b[0m|\n| :\u001b[43m \u001b[0m| : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|\u001b[34;1mR\u001b[0m: | : :\u001b[35mG\u001b[0m|\n|\u001b[43m \u001b[0m: | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (West)\n+---------+\n|\u001b[34;1m\u001b[43mR\u001b[0m\u001b[0m: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|\u001b[42mR\u001b[0m: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (Pickup)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n|\u001b[42m_\u001b[0m: | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (South)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| :\u001b[42m_\u001b[0m| : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (East)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| :\u001b[42m_\u001b[0m: : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (South)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : :\u001b[42m_\u001b[0m: : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (East)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : : :\u001b[42m_\u001b[0m: |\n| | : | : |\n|Y| : |B: |\n+---------+\n (East)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | :\u001b[42m_\u001b[0m: |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|R: | :\u001b[42m_\u001b[0m:\u001b[35mG\u001b[0m|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|R: | : :\u001b[35m\u001b[42mG\u001b[0m\u001b[0m|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (East)\n+---------+\n|R: | : :\u001b[35m\u001b[34;1m\u001b[43mG\u001b[0m\u001b[0m\u001b[0m|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (Dropoff)\n+---------+\n|\u001b[34;1mR\u001b[0m: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : : :\u001b[43m \u001b[0m: |\n| | : | : |\n|Y| : |B: |\n+---------+\n (South)\n+---------+\n|\u001b[34;1mR\u001b[0m: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : :\u001b[43m \u001b[0m: : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (West)\n+---------+\n|\u001b[34;1mR\u001b[0m: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| :\u001b[43m \u001b[0m: : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (West)\n+---------+\n|\u001b[34;1mR\u001b[0m: | : :\u001b[35mG\u001b[0m|\n| :\u001b[43m \u001b[0m| : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|\u001b[34;1mR\u001b[0m: | : :\u001b[35mG\u001b[0m|\n|\u001b[43m \u001b[0m: | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (West)\n+---------+\n|\u001b[34;1m\u001b[43mR\u001b[0m\u001b[0m: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|\u001b[42mR\u001b[0m: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (Pickup)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n|\u001b[42m_\u001b[0m: | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (South)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| :\u001b[42m_\u001b[0m| : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (East)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| :\u001b[42m_\u001b[0m: : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (South)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : :\u001b[42m_\u001b[0m: : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (East)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | : : |\n| : : :\u001b[42m_\u001b[0m: |\n| | : | : |\n|Y| : |B: |\n+---------+\n (East)\n+---------+\n|R: | : :\u001b[35mG\u001b[0m|\n| : | :\u001b[42m_\u001b[0m: |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|R: | :\u001b[42m_\u001b[0m:\u001b[35mG\u001b[0m|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (North)\n+---------+\n|R: | : :\u001b[35m\u001b[42mG\u001b[0m\u001b[0m|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (East)\n+---------+\n|R: | : :\u001b[35m\u001b[34;1m\u001b[43mG\u001b[0m\u001b[0m\u001b[0m|\n| : | : : |\n| : : : : |\n| | : | : |\n|Y| : |B: |\n+---------+\n (Dropoff)\nResults after 5 episodes:\nAverage timesteps per episode: 14.4\nAverage penalties per episode: 0.0\n"
]
]
] |
[
"code",
"markdown",
"code"
] |
[
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab6e34f21da24690bbafabc792af5ea59a2b851
| 23,704 |
ipynb
|
Jupyter Notebook
|
Lab8/034_K_Means.ipynb
|
Meet35/034_MeetDobariya
|
f5f0a32824609364d3e4c4332b489613e9407b03
|
[
"MIT"
] | null | null | null |
Lab8/034_K_Means.ipynb
|
Meet35/034_MeetDobariya
|
f5f0a32824609364d3e4c4332b489613e9407b03
|
[
"MIT"
] | null | null | null |
Lab8/034_K_Means.ipynb
|
Meet35/034_MeetDobariya
|
f5f0a32824609364d3e4c4332b489613e9407b03
|
[
"MIT"
] | null | null | null | 76.961039 | 7,882 | 0.727725 |
[
[
[
"import numpy as np \nimport pandas as pd \nfrom sklearn import datasets\nfrom sklearn.cluster import KMeans\nimport matplotlib.pyplot as plt",
"_____no_output_____"
],
[
"dataset=datasets.load_breast_cancer()\ndataset\n\nprint(dataset.data.shape)\nprint(dataset.target.shape)",
"(569, 30)\n(569,)\n"
],
[
"kmeans = KMeans(n_clusters=10, random_state=11)\nprediction = kmeans.fit_predict(dataset.data)\nprint(prediction)\n\nkmeans.cluster_centers_.shape\nprint(kmeans.cluster_centers_)",
"[9 9 1 0 1 5 1 5 5 5 2 8 8 2 5 2 2 8 9 5 0 7 2 6 9 8 2 8 8 8 1 5 8 1 8 8 5\n 0 5 5 5 0 1 5 5 1 7 5 0 5 0 5 0 8 2 0 9 2 0 7 7 7 2 7 5 2 7 0 7 0 1 7 1 5\n 0 8 0 1 1 0 0 0 6 8 0 1 5 1 0 5 5 5 5 5 2 1 0 7 0 5 5 7 0 7 7 5 0 0 6 0 7\n 0 5 7 7 0 7 2 8 8 0 1 6 5 0 5 5 1 5 1 0 2 8 2 1 0 0 0 2 7 7 8 0 0 7 0 0 5\n 5 5 0 7 7 7 5 0 8 2 0 0 0 1 9 0 6 5 7 8 1 5 0 2 2 7 7 7 7 2 0 0 4 9 8 0 2\n 7 8 0 0 0 5 0 7 5 5 0 5 8 1 2 0 8 9 2 0 2 7 8 0 2 1 0 6 2 5 5 0 7 9 9 5 5\n 7 2 5 5 7 5 0 5 8 0 0 9 7 5 4 1 5 8 5 0 0 5 1 7 0 0 7 0 9 0 9 8 9 5 9 2 8\n 2 9 8 8 2 8 4 7 5 0 7 5 0 6 7 8 0 0 8 5 5 1 0 1 2 0 0 0 0 7 0 5 5 0 0 0 5\n 7 0 5 7 9 0 1 7 0 0 0 7 5 5 0 5 0 0 7 0 0 8 7 0 7 1 0 9 0 0 5 0 2 2 2 0 7\n 0 0 8 0 1 7 4 5 7 7 1 0 7 0 2 0 0 0 2 4 2 0 0 0 5 7 7 0 0 0 2 5 1 1 0 4 9\n 8 5 1 9 5 2 7 5 5 0 0 7 0 0 0 5 0 5 0 8 7 7 8 9 0 5 5 0 0 0 8 0 0 0 0 0 2\n 0 8 0 0 0 7 2 2 0 7 1 0 0 0 5 0 5 7 7 7 0 7 0 5 0 1 1 5 5 0 5 5 5 0 8 5 7\n 8 0 8 5 5 9 0 1 0 5 0 5 0 0 0 7 1 3 5 0 5 5 5 7 8 0 7 0 2 0 7 0 5 5 0 2 0\n 5 5 5 2 0 5 1 0 2 0 8 8 0 5 5 0 0 1 1 5 5 0 6 7 7 0 7 2 2 0 5 5 5 2 0 1 1\n 0 0 7 6 7 5 7 7 5 0 5 0 0 0 5 1 7 1 5 0 7 7 0 5 5 0 5 5 7 7 7 0 7 7 0 7 0\n 7 7 7 5 0 5 7 2 9 9 1 2 1 7]\n[[1.21398295e+01 1.81243750e+01 7.80391477e+01 4.53930682e+02\n 9.30579545e-02 7.96476705e-02 4.52235324e-02 2.59481023e-02\n 1.74372727e-01 6.23869886e-02 2.76667614e-01 1.19831080e+00\n 1.96479489e+00 2.04931250e+01 6.79651136e-03 2.03912898e-02\n 2.40284352e-02 9.64013636e-03 2.03964205e-02 3.30967330e-03\n 1.34010227e+01 2.41963068e+01 8.71398295e+01 5.49269318e+02\n 1.26916705e-01 1.92223466e-01 1.75752653e-01 7.78560852e-02\n 2.78232386e-01 7.95895455e-02]\n [1.92993617e+01 2.17512766e+01 1.27459574e+02 1.15872340e+03\n 9.99821277e-02 1.46731915e-01 1.71502553e-01 9.74040426e-02\n 1.93663830e-01 6.02644681e-02 7.08710638e-01 1.24907660e+00\n 4.92457447e+00 8.57978723e+01 6.77202128e-03 3.29246809e-02\n 4.40670213e-02 1.61012979e-02 2.21217021e-02 3.89431915e-03\n 2.32834043e+01 2.88346809e+01 1.55308511e+02 1.64951064e+03\n 1.38865957e-01 3.60872340e-01 4.48112766e-01 1.86787234e-01\n 3.18602128e-01 8.55921277e-02]\n [1.54638000e+01 2.02392000e+01 1.01397600e+02 7.40524000e+02\n 1.00539400e-01 1.25822000e-01 1.23348200e-01 6.54630000e-02\n 1.85918000e-01 6.26102000e-02 3.90868000e-01 1.03997400e+00\n 2.75976000e+00 3.72140000e+01 6.65860000e-03 2.82472800e-02\n 3.78898000e-02 1.35351000e-02 1.91477200e-02 3.67792000e-03\n 1.80026000e+01 2.75786000e+01 1.19620000e+02 9.92134000e+02\n 1.41392800e-01 3.39973200e-01 3.95144000e-01 1.54146800e-01\n 3.16380000e-01 9.00534000e-02]\n [2.74200000e+01 2.62700000e+01 1.86900000e+02 2.50100000e+03\n 1.08400000e-01 1.98800000e-01 3.63500000e-01 1.68900000e-01\n 2.06100000e-01 5.62300000e-02 2.54700000e+00 1.30600000e+00\n 1.86500000e+01 5.42200000e+02 7.65000000e-03 5.37400000e-02\n 8.05500000e-02 2.59800000e-02 1.69700000e-02 4.55800000e-03\n 3.60400000e+01 3.13700000e+01 2.51200000e+02 4.25400000e+03\n 1.35700000e-01 4.25600000e-01 6.83300000e-01 2.62500000e-01\n 2.64100000e-01 7.42700000e-02]\n [2.36850000e+01 2.31566667e+01 1.56916667e+02 1.77366667e+03\n 1.02470000e-01 1.54020000e-01 2.17200000e-01 1.35818333e-01\n 1.82466667e-01 5.73900000e-02 1.04615000e+00 1.14798333e+00\n 7.03883333e+00 1.70916667e+02 5.63983333e-03 2.36733333e-02\n 3.28216667e-02 1.31516667e-02 1.60150000e-02 2.73416667e-03\n 3.18616667e+01 3.25450000e+01 2.12000000e+02 3.14583333e+03\n 1.46033333e-01 3.50783333e-01 4.78883333e-01 2.26750000e-01\n 2.93666667e-01 8.00400000e-02]\n [1.39387395e+01 1.90294958e+01 9.04831933e+01 6.00262185e+02\n 9.39314286e-02 1.00406387e-01 7.45220756e-02 4.14941345e-02\n 1.75155462e-01 6.16680672e-02 3.13209244e-01 1.08587899e+00\n 2.26420084e+00 2.71161345e+01 5.84906723e-03 2.42642689e-02\n 2.82606723e-02 1.11375714e-02 1.78284202e-02 3.49648571e-03\n 1.57033613e+01 2.55010924e+01 1.03702605e+02 7.57455462e+02\n 1.28325294e-01 2.62771765e-01 2.65218908e-01 1.12402101e-01\n 2.83356303e-01 8.43820168e-02]\n [2.37500000e+01 2.11944444e+01 1.59244444e+02 1.75011111e+03\n 1.07433333e-01 1.92500000e-01 2.63044444e-01 1.40782222e-01\n 1.94166667e-01 6.16344444e-02 1.25552222e+00 1.39952222e+00\n 9.29744444e+00 1.90815556e+02 8.61422222e-03 4.69666667e-02\n 5.96800000e-02 1.74600000e-02 2.50555556e-02 4.80877778e-03\n 2.87344444e+01 2.74311111e+01 1.97022222e+02 2.48000000e+03\n 1.39488889e-01 4.17266667e-01 5.16111111e-01 2.28388889e-01\n 2.95433333e-01 8.48355556e-02]\n [9.87515385e+00 1.78310989e+01 6.31275824e+01 2.99732967e+02\n 9.64635165e-02 8.09469231e-02 4.46973846e-02 1.89206154e-02\n 1.83812088e-01 6.83807692e-02 2.95191209e-01 1.51460549e+00\n 2.02103187e+00 1.79611868e+01 9.95848352e-03 2.56543407e-02\n 3.31990110e-02 1.02651538e-02 2.54141758e-02 4.97217582e-03\n 1.08238352e+01 2.29446154e+01 6.97957143e+01 3.58796703e+02\n 1.31465934e-01 1.64161758e-01 1.40192637e-01 5.34132967e-02\n 2.73173626e-01 8.52007692e-02]\n [1.72869565e+01 2.12302174e+01 1.13873913e+02 9.32139130e+02\n 9.95578261e-02 1.33787826e-01 1.40148913e-01 8.17313043e-02\n 1.90567391e-01 6.09047826e-02 5.72460870e-01 1.27289565e+00\n 4.09880435e+00 6.27041304e+01 6.37152174e-03 3.00010217e-02\n 3.73045652e-02 1.49175435e-02 1.98891304e-02 4.01458696e-03\n 2.05626087e+01 2.84769565e+01 1.36710870e+02 1.30030435e+03\n 1.38533478e-01 3.22255652e-01 3.89013043e-01 1.69940652e-01\n 3.12139130e-01 8.53423913e-02]\n [2.02387500e+01 2.21650000e+01 1.34295833e+02 1.28227500e+03\n 1.04119583e-01 1.60014167e-01 2.05120833e-01 1.14455000e-01\n 1.92020833e-01 6.12808333e-02 7.66895833e-01 1.01625417e+00\n 5.37841667e+00 1.00878750e+02 6.02283333e-03 3.03270833e-02\n 4.32558333e-02 1.55868750e-02 1.72483333e-02 3.85025000e-03\n 2.57820833e+01 2.96491667e+01 1.72391667e+02 2.04983333e+03\n 1.46254167e-01 3.98658333e-01 5.29562500e-01 2.20404167e-01\n 3.19550000e-01 9.13916667e-02]]\n"
],
[
"# Scatter plot of the data points\nimport matplotlib.pyplot as plt\nfig, ax = plt.subplots(2, 5, figsize=(8, 3))\ncenters = kmeans.cluster_centers_.reshape(10, 6, 5)\nfor axi, center in zip(ax.flat, centers):\n axi.set(xticks=[], yticks=[])\n axi.imshow(center, interpolation='nearest', cmap=plt.cm.binary)",
"_____no_output_____"
],
[
"import numpy as np\nfrom scipy.stats import mode\nlabels = np.zeros_like(prediction)\nfor i in range(10):\n mask = (prediction == i)\n labels[mask] = mode(dataset.target[mask])[0]",
"_____no_output_____"
],
[
"from sklearn.metrics import accuracy_score\naccuracy_score(dataset.target, labels)",
"_____no_output_____"
],
[
"from sklearn.metrics import confusion_matrix\nimport seaborn as sns\nmat = confusion_matrix(dataset.target, labels)\nax = sns.heatmap(mat.T, square=True, annot=True, fmt='d', cbar=False,xticklabels=dataset.target_names,yticklabels=dataset.target_names)\n\n#ax.set_ylim(10,10)\nplt.xlabel('true label')\nplt.ylabel('predicted label');",
"_____no_output_____"
],
[
"",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab6e7065bdda04cc8fcd97a4a55b4a4868b4f4c
| 58,290 |
ipynb
|
Jupyter Notebook
|
DataCamp DBSCAN blog/DBSCAN_A_macroscopic_investigation.ipynb
|
sayakpaul/DataCamp-blogs
|
902453e3f89485afb6a444db761e8209051bcd5f
|
[
"MIT"
] | 3 |
2019-07-27T04:37:02.000Z
|
2020-06-11T07:22:01.000Z
|
DataCamp DBSCAN blog/DBSCAN_A_macroscopic_investigation.ipynb
|
sayakpaul/DataCamp-blogs
|
902453e3f89485afb6a444db761e8209051bcd5f
|
[
"MIT"
] | null | null | null |
DataCamp DBSCAN blog/DBSCAN_A_macroscopic_investigation.ipynb
|
sayakpaul/DataCamp-blogs
|
902453e3f89485afb6a444db761e8209051bcd5f
|
[
"MIT"
] | 1 |
2019-07-07T18:48:22.000Z
|
2019-07-07T18:48:22.000Z
| 79.52251 | 23,032 | 0.768194 |
[
[
[
"<h1><center>DBSCAN: A macroscopic investigation in Python</center></h1><br>\nCluster analysis is an important problem in data analysis. Data scientists use clustering to identify malfunctioning servers, group genes with similar expression patterns, or various other applications.\n\nBriefly, clustering is the task of grouping together a set of objects in a way that objects in the same cluster are more similar to each other than to objects in other clusters. Similarity is an amount that reflects the strength of a relationship between two data objects. Clustering is mainly used for exploratory data mining. Clustering has manifold usage in many fields such as machine learning, pattern recognition, image analysis, information retrieval, bio-informatics, data compression, and computer graphics.\n\nThere are many families of clustering techniques, and you may be familiar with the most popular one: K-Means (which belongs to the *family of centroid-based clustering*). As a quick refresher, K-Means determines k centroids in the data and clusters points by assigning them to the nearest centroid.\n\nWhile K-Means is easy to understand and implement in practice, the algorithm does not take care of outliers, so all points are assigned to a cluster even if they do not belong in any. In the domain of anomaly detection, this causes problems as anomalous points will be assigned to the same cluster as “normal” data points. The anomalous points pull the cluster centroid towards them, making it harder to classify them as anomalous points.\n\nThis tutorial will cover another type of clustering technique known as density-based clustering specifically DBSCAN (a density-based based clustering technique). Compared to centroid-based clustering like K-Means, density-based clustering works by identifying “dense” clusters of points, allowing it to learn clusters of arbitrary shape and identify outliers in the data. \n\n<h2>In this post you will get to know about:</h2>\n* Disadvantage of centroid-based clustering technique\n* General introduction to density-based clustering technique\n* Inner workings of DBSCAN\n* A simple case study of DBSCAN in Python\n* Applications of DBSCAN\n\n\n<h3>Disadvantage of centroid-based clustering technique: </h3>\nBefore discussing the disadvantage of centroid-based clustering, let me give a brief introduction to it. A centroid is a data point (imaginary or real) at the center of a cluster. In centroid-based clustering, clusters are represented by a central vector or a centroid. This centroid might not necessarily be a member of the dataset. Centroid-based clustering is an iterative clustering algorithm in which the notion of similarity is derived by how close a data point is to the centroid of the cluster. <br><br>\nSometimes a dataset can contain extreme values that are outside the range of what is expected and unlike the other data. These are called outliers. More formally, an outlier is an observation that lies an abnormal distance from other values in a random sample from a population.\nThe main fundamental of centroid-based clustering techniques is driven by distance measurements between the data points and centroids. Therefore, centroid-based clustering techniques generally fail to identify the data points that deviate from the normal distribution of the data to a great extent. Even before predictive models are prepared on data, outliers can result in misleading representations and in turn misleading interpretations of collected data. This is essentially not desirable for building efficient predictive and analytical models from data. <br><br>\nYou can consider the following two taller bars (than rest of the bars) as outliers in that particular data:\n<center>\n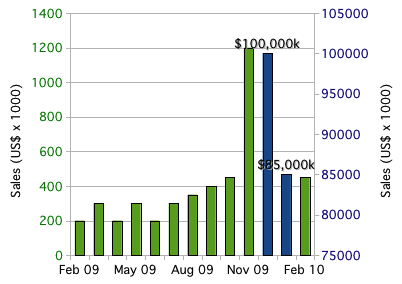 \n</center>\n\n<h3>General introduction to density-based clustering technique:</h3>\nBefore discussing density-based clustering, you first need to cover a topic : ɛ-neighborhoods.\n\nThe general idea behind ɛ-neighborhoods is given a data point, you want to be able to reason about the data points in the space around it. Formally, for some real-valued ɛ > 0 and some point p, the ɛ-neighborhood of p is defined as the set of points that are at most distance ɛ away from p.\n\nIf you think back to geometry, the shape in which all points are equidistant from the center is the circle. In 2D space, the ɛ-neighborhood of a point p is the set of points contained in a circle of radius ɛ, centered at p. In 3D space, the ɛ-neighborhood is a sphere of radius ɛ, centered at p, and in higher dimensional space, the ɛ-neighborhood is just the [N-sphere](https://en.wikipedia.org/wiki/N-sphere) of radius ɛ, centered at p.\n\nLet’s consider an example to make this idea more concrete.\nIn the image below 100 data points are scattered in the interval [1,3]X[2,4]. Let’s pick the point (3,2) to be our point p.\n\n<center>\n \n</center>\n\nFirst, let’s consider the neighborhood of p with radius 0.5 (ɛ = 0.5), the set of points that are distance 0.5 away from p.\n\n<center>\n \n</center>\n\nThe opaque green oval represents our neighborhood, and there are 31 data points in this neighborhood. Since 100 data points were scattered and 31 are in the neighborhood, this means that a little under one-third of the data points are contained within the neighborhood of p with radius 0.5.\n\nNow, let’s change our radius to 0.15 (ɛ = 0.15) and consider the resulting smaller neighborhood.\n\n<center>\n \n</center>\n\nNow the neighborhood is shrunk a bit, so now only 3 data points are contained within it. By decreasing ɛ from 0.5 to 0.15 (a 70% reduction), the number of points is decreased in our neighborhood from 31 to 3 (a 90% reduction).\n\nNow that you have a fair understanding of “neighborhood”, I will introduce the next important concept: the notion of a “density” for a neighborhood (You are proceeding towards learning “density-based clustering\", after all).\n\nIn a grade-school science class, children are taught that density = mass/volume. Let’s use this idea of mass divided by volume to define density at some point p. If you consider some point p and its neighborhood of radius ɛ, you can define the mass of the neighborhood as the number of data points (or alternatively, the fraction of data points) contained within the neighborhood, and the volume of the neighborhood is volume of the resulting shape of the neighborhood. In the 2D case, the neighborhood is a circle, so the volume of the neighborhood is just the area of the resulting circle. In the 3D and higher dimensional case, the neighborhood is a sphere or n-sphere, so you can calculate the volume of this shape.\n\nFor example, let’s consider our neighborhood of p = (3,2) of radius 0.5 again.\n\n<center>\n \n</center>\n\nThe mass is the number of data points in the neighborhood, so mass = 31. The volume is the area of the circle, so volume = π0.5<sup>2</sup> = π/4. Therefore, our local density approximation at * p = (3,2) is calculated as density = mass/volume = 31/(π/4) = 124/π ~= 39.5.\n\nThis value is meaningless by itself, but if you calculate the local density approximation for all points in our dataset, you could cluster our points by saying that points that are nearby (contained in the same neighborhood) and have similar local density approximations belong in the same cluster. If you decrease the value of ɛ, you can construct smaller neighborhoods (less volume) that would also contain fewer data points. Ideally, you want to identify highly dense neighborhoods where most of the data points are contained in these neighborhoods, but the volume of each of these neighborhoods is relatively small.<br>\nWhile this is not exactly what either DBSCAN or the Level Set Tree algorithm (another clustering technique belonging to the family of density-based clustering) does, it forms the general intuition behind density-based clustering.<br>\nTo recap, you covered the ɛ-neighborhoods and how they allow to reason about the space around a particular point. Then you learnt a notion of density at a particular point for a particular neighborhood. In the next section, you will get to know the DBSCAN algorithm where the ɛ-ball is a fundamental tool for defining clusters.\n\n\n<h3>Inner workings of DBSCAN:</h3>\nDBSCAN stands for Density-Based Spatial Clustering of Applications with Noise and it is hands down the most well-known density-based clustering algorithm. It was first introduced by first introduced in 1996 by [Ester et. al](http://citeseerx.ist.psu.edu/viewdoc/summary?doi=10.1.1.71.1980). Due to its importance in both theory and applications, this algorithm is one of three algorithms awarded the Test of Time Award at SIGKDD 2014.\n\nUnlike K-Means, DBSCAN does not require the number of clusters as a parameter. Rather it infers the number of clusters based on the data, and it can discover clusters of arbitrary shape (for comparison, K-Means usually discovers spherical clusters). As you saw earlier, the ɛ-neighborhood is fundamental to DBSCAN to approximate local density, so the algorithm has two parameters:\n\n* ɛ: The radius of our neighborhoods around a data point p.\n* minPts: The minimum number of data points you want in a neighborhood to define a cluster.\n\nUsing these two parameters, DBSCAN categories the data points into three categories:\n\n* Core Points: A data point p is a core point if Nbhd(p,ɛ) [ɛ-neighborhood of p] contains at least minPts ; |Nbhd(p,ɛ)| >= minPts.\n* Border Points: A data point *q is a border point if Nbhd(q, ɛ) contains less than minPts data points, but q is reachable from some core point p.\n* Outlier: A data point o is an outlier if it is neither a core point nor a border point. Essentially, this is the “other” class.\n\nThese definitions may seem abstract, so let’s cover what each one means in more detail.\n\n<b>Core Points: </b><br>\nCore Points are the foundations for our clusters are based on the density approximation I discussed in the previous section. You use the same ɛ to compute the neighborhood for each point, so the volume of all the neighborhoods is the same. However, the number of other points in each neighborhood is what differs. Recall that I said you can think of the number of data points in the neighborhood as its mass. The volume of each neighborhood is constant, and the mass of neighborhood is variable, so by putting a threshold on the minimum amount of mass needed to be core point, you are essentially setting a minimum density threshold. Therefore, core points are data points that satisfy a minimum density requirement. Our clusters are built around our core points (hence the core part), so by adjusting our minPts parameter, you can fine-tune how dense our clusters cores must be.\n\n<b>Border Points:</b><br>\nBorder Points are the points in our clusters that are not core points. In the definition above for border points, I used the term density-reachable. I have not defined this term yet, but the concept is simple. To explain this concept, let’s revisit our neighborhood example with epsilon = 0.15. Consider the point r (the black dot) that is outside of the point p‘s neighborhood.\n\n<center>\n \n</center>\n\nAll the points inside the point p‘s neighborhood are said to be directly reachable from p. Now, let’s explore the neighborhood of point q, a point directly reachable from p. The yellow circle represents q‘s neighborhood.\n\n<center>\n \n</center>\n\nNow while your target point r is not your starting point p‘s neighborhood, it is contained in the point q‘s neighborhood. This is the idea behind density-reachable: If you can get to the point r by jumping from neighborhood to neighborhood, starting at a point p, then the point r is density-reachable from the point p.\n\n<center>\n \n</center>\n\nAs an analogy, you can think of density-reachable points as being the “friends of a friend”. If the directly-reachable of a core point p are its “friends”, then the density-reachable points, points in neighborhood of the “friends” of p, are the “friends of its friends”. One thing that may not be clear is density-reachable points is not limited to just two adjacent neighborhood jumps. As long as you can reach the point doing “neighborhood jumps”, starting at a core point p, that point is density-reachable from p, so “friends of a friend of a friend … of a friend” are included as well. <br>\nIt is important to keep in mind that this idea of density-reachable is dependent on our value of ɛ. By picking larger values of ɛ, more points become density-reachable, and by choosing smaller values of ɛ, fewer points become density-reachable.\n\n\n<b>Outliers:</b><br> \nFinally, you get to the “other” class. Outliers are points that are neither core points nor are they close enough to a cluster to be density-reachable from a core point. Outliers are not assigned to any cluster and, depending on the context, may be considered anomalous points.\n\n<h3>Case study of DBSCAN in Python:</h3><br>\nDBSCAN is already beautifully implemented in the popular Python machine learning library *Scikit-Learn*, and because this implementation is scalable and well-tested, you will be using it to see how DBSCAN works in practice.\n\nThe steps to the DBSCAN algorithm are:\n\n* Pick a point at random that has not been assigned to a cluster or been designated as an outlier. Compute its neighborhood to determine if it’s a core point. If yes, start a cluster around this point. If no, label the point as an outlier.\n* Once we find a core point and thus a cluster, expand the cluster by adding all directly-reachable points to the cluster. Perform “neighborhood jumps” to find all density-reachable points and add them to the cluster. If an outlier is added, change that point’s status from outlier to border point.\n* Repeat these two steps until all points are either assigned to a cluster or designated as an outlier.\n\nFor this case study purpose you will be using [a dataset consisting of annual customer data for a wholesale distributor](https://archive.ics.uci.edu/ml/datasets/Wholesale+customers).\n\nSo, let's get started.\n\n\n\n\n",
"_____no_output_____"
]
],
[
[
"# Let's import all your dependencies first\n\nfrom sklearn.cluster import DBSCAN\nfrom sklearn.preprocessing import StandardScaler\nimport numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt",
"_____no_output_____"
]
],
[
[
"The dataset consists of 440 customers and has 8 attributes for each of these customers. You will use the Pandas library to import the .csv file and convert it into a DataFrame object.\n\nNow while importing your .csv file into the , make sure you supply the accurate path of that file.",
"_____no_output_____"
]
],
[
[
"# Import .csv file and convert it to a DataFrame object\ndf = pd.read_csv(\"C:/Users/Sayak/data/customers.csv\");\n\nprint(df.head())\n",
" Channel Region Fresh Milk Grocery Frozen Detergents_Paper \\\n0 2 3 12669 9656 7561 214 2674 \n1 2 3 7057 9810 9568 1762 3293 \n2 2 3 6353 8808 7684 2405 3516 \n3 1 3 13265 1196 4221 6404 507 \n4 2 3 22615 5410 7198 3915 1777 \n\n Delicatessen \n0 1338 \n1 1776 \n2 7844 \n3 1788 \n4 5185 \n"
]
],
[
[
"Now before proceeding any further with applying DBSCAN, it is very important that you know the data well so as know what kind of data is in the dataset, what distribution the data follows, and which features are numerical or not.\n\nAccording to the description given in the official [UCI machine learning repository of this dataset](https://archive.ics.uci.edu/ml/datasets/wholesale+customers), information about the features of the dataset is as follows:\n\n<li>\n FRESH: annual spending (m.u.) on fresh products (Continuous); </li>\n<li>MILK: annual spending (m.u.) on milk products (Continuous); </li>\n<li>GROCERY: annual spending (m.u.)on grocery products (Continuous); </li>\n<li>FROZEN: annual spending (m.u.)on frozen products (Continuous) </li>\n<li>DETERGENTS_PAPER: annual spending (m.u.) on detergents and paper products (Continuous) </li>\n<li>DELICATESSEN: annual spending (m.u.)on and delicatessen products (Continuous); </li>\n<li>CHANNEL: customers’ Channel - Horeca (Hotel/Restaurant/Café) or Retail channel (Nominal) \nREGION\n</li>\n\nNow that you know about the features about the dataset, let's display some stats of the data.",
"_____no_output_____"
]
],
[
[
"print(df.info())",
"<class 'pandas.core.frame.DataFrame'>\nRangeIndex: 440 entries, 0 to 439\nData columns (total 8 columns):\nChannel 440 non-null int64\nRegion 440 non-null int64\nFresh 440 non-null int64\nMilk 440 non-null int64\nGrocery 440 non-null int64\nFrozen 440 non-null int64\nDetergents_Paper 440 non-null int64\nDelicatessen 440 non-null int64\ndtypes: int64(8)\nmemory usage: 27.6 KB\nNone\n"
]
],
[
[
"As you can see from the above output, there is no missing value in the dataset and all the data is *integer* in type. This reduces the burden of further preprocessing the data. Let's dig a bit more.",
"_____no_output_____"
]
],
[
[
"print(df.describe())",
" Channel Region Fresh Milk Grocery \\\ncount 440.000000 440.000000 440.000000 440.000000 440.000000 \nmean 1.322727 2.543182 12000.297727 5796.265909 7951.277273 \nstd 0.468052 0.774272 12647.328865 7380.377175 9503.162829 \nmin 1.000000 1.000000 3.000000 55.000000 3.000000 \n25% 1.000000 2.000000 3127.750000 1533.000000 2153.000000 \n50% 1.000000 3.000000 8504.000000 3627.000000 4755.500000 \n75% 2.000000 3.000000 16933.750000 7190.250000 10655.750000 \nmax 2.000000 3.000000 112151.000000 73498.000000 92780.000000 \n\n Frozen Detergents_Paper Delicatessen \ncount 440.000000 440.000000 440.000000 \nmean 3071.931818 2881.493182 1524.870455 \nstd 4854.673333 4767.854448 2820.105937 \nmin 25.000000 3.000000 3.000000 \n25% 742.250000 256.750000 408.250000 \n50% 1526.000000 816.500000 965.500000 \n75% 3554.250000 3922.000000 1820.250000 \nmax 60869.000000 40827.000000 47943.000000 \n"
]
],
[
[
"From the above output, you can derive all the necessary statistical measures like standard deviation, mean, max of each and every feature present in the dataset. You can see most of the data in this dataset is *[continuous](https://stats.stackexchange.com/questions/206/what-is-the-difference-between-discrete-data-and-continuous-data)* in nature except for two features: Channel and Region. So for easing your computations, you will drop these two:",
"_____no_output_____"
]
],
[
[
"df.drop([\"Channel\", \"Region\"], axis = 1, inplace = True)",
"_____no_output_____"
],
[
"# Let's get a view of the data after the drop\n\nprint(df.head())",
" Fresh Milk Grocery Frozen Detergents_Paper Delicatessen\n0 12669 9656 7561 214 2674 1338\n1 7057 9810 9568 1762 3293 1776\n2 6353 8808 7684 2405 3516 7844\n3 13265 1196 4221 6404 507 1788\n4 22615 5410 7198 3915 1777 5185\n"
]
],
[
[
"So you can visualize the data, for that you are going to use two of the features:\n\n* Groceries: The customer’s annual spending (in some monetary unit) on grocery products.\n* Milk: The customer’s annual spending (in some monetary unit) on milk products.",
"_____no_output_____"
]
],
[
[
"# Let's plot the data now\nx = df['Grocery']\ny = df['Milk']\n\nplt.scatter(x,y)\nplt.xlabel(\"Groceries\")\nplt.ylabel(\"Milk\")\nplt.show()",
"_____no_output_____"
]
],
[
[
"Let's brief about the functions that you used for the plotting purpose:\nplt.scatter() : This function actually creates the scatter plot based on the data (as parameters that you supply [*x* and *y*]).\nplt.xlabel() : It helps you to put a label along the *X-axis*. (*Groceries* in this case)\nplt.ylabel() : It helps you to put a label along the *Y-axis*. (*Milk* in this case)\nplt.show() : After the plot is created, this function helps you to display it as the output.\n\nYou should really explore the beautiful world of *Matplotlib* for all your visualization purposes. Its [documentation](https://matplotlib.org/) is absolutely awesome. ",
"_____no_output_____"
],
[
"You can easily spot the data points that are far astray. Right? Well, those are your outliers. \n\nWith DBSCAN, we want to identify this main cluster of customers, but we also want to flag customers with more unusual annual purchasing habits as outliers.\n\nBecause the values of the data are in the thousands, you are going to normalize each attribute by scaling it to 0 mean and unit variance. What is does basically is it helps to keep the inter-relationships between the features intact so that a small change in one feature would reflect in the other.",
"_____no_output_____"
]
],
[
[
"df = df[[\"Grocery\", \"Milk\"]]\ndf = df.as_matrix().astype(\"float32\", copy = False)",
"_____no_output_____"
],
[
"stscaler = StandardScaler().fit(df)\ndf = stscaler.transform(df)\n",
"_____no_output_____"
]
],
[
[
"You will construct a DBSCAN object that requires a minimum of 15 data points in a neighborhood of radius 0.5 to be considered a core point.",
"_____no_output_____"
]
],
[
[
"dbsc = DBSCAN(eps = .5, min_samples = 15).fit(df)",
"_____no_output_____"
]
],
[
[
"Next, we can extract our cluster labels and outliers to plot our results.",
"_____no_output_____"
]
],
[
[
"labels = dbsc.labels_\ncore_samples = np.zeros_like(labels, dtype = bool)\ncore_samples[dbsc.core_sample_indices_] = True\n\n",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
],
[
"Lining up with the intuition, the DBSCAN algorithm was able to identify one cluster of customers who are around the mean grocery and mean milk product purchases. In addition, it was able to flag customers whose annual purchasing behavior deviated too heavily from other customers.\n\nBecause the outliers corresponded to customers with more extreme purchasing behavior, the wholesale distributor could specifically target these customers with exclusive discounts to encourage larger purchases.\n\n<h3>Real life applications of DBSCAN:</h3>\n\n* Suppose we have an e-commerce and we want to improve our sales by recommending relevant products to our customers. We don’t know exactly what our customers are looking for but based on a data set we can predict and recommend a relevant product to a specific customer. We can apply the DBSCAN to our data set (based on the e-commerce database) and find clusters based on the products that the users have bought. Using this clusters we can find similarities between customers, for example, if customer A has bought a pen, a book and one pair scissors, while customer B purchased a book and one pair of scissors, then you could recommend a pen to customer B.\n\n* Before the rise of deep learning based advanced methodologies, researchers used DBSCAN in order to segregate genes from a genes dataset that had the chance of mediating cancer. \n\n* Scientists have used DBSCAN in order to detect the stops in the trajectory data generated from mobile GPS devices. Stops represent the most meaningful and most important part of a trajectory.\n\n<h3>Conclusion:</h3>\nSo, in this blogpost you got to know about the prime disadvantages of centroid-based clustering and got familiar with another family of clustering techniques i.e. density-based clustering. You also saw how they overcome the shortcomings of centroid-based clustering. \n\nYou learnt how DBSCAN works and also did a case study of it. Besides, you got a fair overview of the real life problems where DBSCAN has been incorporated for solving them. As a further reading, I would really recommend you all go through the other density-based clustering methods like *Level Set Tree clustering* and how it is different from DBSCAN. \n\n<h4>References:</h4>\n* Martin Ester, Hans-Peter Kriegel, Jörg Sander, and Xiaowei Xu. 1996. A density-based algorithm for discovering clusters a density-based algorithm for discovering clusters in large spatial databases with noise. In Proceedings of the Second International Conference on Knowledge Discovery and Data Mining (KDD'96), Evangelos Simoudis, Jiawei Han, and Usama Fayyad (Eds.). AAAI Press 226-231.\n\n* https://towardsdatascience.com/how-dbscan-works-and-why-should-i-use-it-443b4a191c80\n\n* https://www.coursera.org/learn/predictive-analytics/lecture/EVHfy/dbscan\n",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
]
] |
4ab6ed0678b68a1f2cc812d2ea9cf056631a70ce
| 26,516 |
ipynb
|
Jupyter Notebook
|
Capstone/Untitled.ipynb
|
ClaudiaCCordeiro/Curso_IBM
|
2123ebf48e0657f61998f38fe803203be51877ff
|
[
"MIT"
] | null | null | null |
Capstone/Untitled.ipynb
|
ClaudiaCCordeiro/Curso_IBM
|
2123ebf48e0657f61998f38fe803203be51877ff
|
[
"MIT"
] | null | null | null |
Capstone/Untitled.ipynb
|
ClaudiaCCordeiro/Curso_IBM
|
2123ebf48e0657f61998f38fe803203be51877ff
|
[
"MIT"
] | null | null | null | 35.929539 | 138 | 0.363705 |
[
[
[
"# Segmenting and Clustering Neighborhoods in Toronto",
"_____no_output_____"
]
],
[
[
"import pandas as pd # importing library for data analsysis\n\n#I chose to use the pandas to extract the table of postal codes of the Wikpedia page.\nurl = \"https://en.wikipedia.org/wiki/List_of_postal_codes_of_Canada:_M\"\ndf_list = pd.read_html(url) #readind the html file.\ndf = df_list[0] # defining a dataframe\n\n#Ignoring cells with a borough that is Not assigned.\ndf = df[df.Borough != 'Not assigned'].reset_index(drop=True)\n\n#treating the cells that do not have a linked Neighbourhood to a Borough.\ndf[\"Neighbourhood\"] = df.apply(lambda row: row.Neighbourhood if row.Neighbourhood !=\"Not assigned\" else row.Borough, axis=1) \n\n#gathering the neighborhoods with the same postal code in a Borough.\ndf = df.groupby(['Postcode', 'Borough'])['Neighbourhood'].apply(', '.join).reset_index()\ndf",
"_____no_output_____"
],
[
"df.shape",
"_____no_output_____"
],
[
"df.groupby(['Borough'])['Neighbourhood'].apply(', '.join).reset_index()",
"_____no_output_____"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code"
]
] |
4ab70e64cc144a92fd677debfbb59e1271579c16
| 207,954 |
ipynb
|
Jupyter Notebook
|
floris/utils/run/demo/floris_demo/demo_1.ipynb
|
forestriveral/floris
|
02c31e121283ad6ccae987cfa3aa1bf1e4b43014
|
[
"Apache-2.0"
] | null | null | null |
floris/utils/run/demo/floris_demo/demo_1.ipynb
|
forestriveral/floris
|
02c31e121283ad6ccae987cfa3aa1bf1e4b43014
|
[
"Apache-2.0"
] | null | null | null |
floris/utils/run/demo/floris_demo/demo_1.ipynb
|
forestriveral/floris
|
02c31e121283ad6ccae987cfa3aa1bf1e4b43014
|
[
"Apache-2.0"
] | null | null | null | 256.733333 | 90,414 | 0.888081 |
[
[
[
"import os, sys\r\nsys.path.append(os.path.abspath('../..'))\r\nimport matplotlib.pyplot as plt\r\n\r\nimport floris.tools as wfct",
"_____no_output_____"
],
[
"# Initialize the FLORIS interface fi\r\n# For basic usage, the florice interface provides a simplified interface to\r\n# the underlying classes\r\nfi = wfct.floris_interface.FlorisInterface(\"../../examples/example_input.json\")\r\n\r\n# Calculate wake\r\nfi.calculate_wake()\r\n\r\n# Get horizontal plane at default height (hub-height)\r\nhor_plane = fi.get_hor_plane()\r\n\r\n# Plot and show\r\nfig, ax = plt.subplots()\r\nwfct.visualization.visualize_cut_plane(hor_plane, ax=ax)\r\nplt.show()",
"\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n"
],
[
"# Initialize the FLORIS interface fi\r\nfi = wfct.floris_interface.FlorisInterface(\"../../examples/example_input.json\")\r\n\r\n# Set to 2x2 farm\r\nfi.reinitialize_flow_field(layout_array=[[0, 0, 600, 600], [0, 300, 0, 300]])\r\n\r\n# Change turbine 0 and 3 to have a 35 m rotor diameter\r\nfi.change_turbine([0, 3], {\"rotor_diameter\": 35})\r\n\r\n# Calculate wake\r\nfi.calculate_wake()\r\n\r\n# Get horizontal plane at default height (hub-height)\r\nhor_plane = fi.get_hor_plane()\r\n\r\n# Plot and show\r\nfig, ax = plt.subplots()\r\nwfct.visualization.visualize_cut_plane(hor_plane, ax=ax)\r\nplt.show()",
"\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Updating turbine: 0\n\u001b[34mfloris.simulation.turbine.Turbine\u001b[0m \u001b[1;30mINFO\u001b[0m Setting rotor_diameter to 35\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Updating turbine: 3\n\u001b[34mfloris.simulation.turbine.Turbine\u001b[0m \u001b[1;30mINFO\u001b[0m Setting rotor_diameter to 35\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n"
],
[
"fi = wfct.floris_interface.FlorisInterface(\"../../examples/example_input.json\")\r\n\r\n# Declare a short-cut visualization function for brevity in this example\r\ndef plot_slice_shortcut(fi, ax, title):\r\n # Get horizontal plane at default height (hub-height)\r\n hor_plane = fi.get_hor_plane()\r\n ax.set_title(title)\r\n wfct.visualization.visualize_cut_plane(hor_plane, ax=ax, minSpeed=4.0, maxSpeed=8.0)\r\n\r\n\r\n# Define a plot\r\nfig, axarr = plt.subplots(3, 3, sharex=True, figsize=(12, 5), dpi=120)\r\naxarr = axarr.flatten()\r\n\r\n# Plot the initial setup\r\nfi.calculate_wake()\r\nplot_slice_shortcut(fi, axarr[0], \"Initial\")\r\n\r\n# Change the wind speed\r\nfi.reinitialize_flow_field(wind_speed=10.0)\r\nfi.calculate_wake()\r\nplot_slice_shortcut(fi, axarr[1], \"WS=10\")\r\n\r\n# Change the wind direction\r\nfi.reinitialize_flow_field(wind_direction=320.0)\r\nfi.calculate_wake()\r\nplot_slice_shortcut(fi, axarr[2], \"WD=320\")\r\n\r\n# Change the TI\r\nfi.reinitialize_flow_field(turbulence_intensity=0.15)\r\nfi.calculate_wake()\r\nplot_slice_shortcut(fi, axarr[3], \"TI=15%\")\r\n\r\n# Change the shear\r\nfi.reinitialize_flow_field(wind_shear=0.2)\r\nfi.calculate_wake()\r\nplot_slice_shortcut(fi, axarr[4], \"Shear=0.2\")\r\n\r\n# Change the veer\r\nfi.reinitialize_flow_field(wind_veer=5) # TODO IS THIS RIGHT?\r\nfi.calculate_wake()\r\nplot_slice_shortcut(fi, axarr[5], \"Veer=5\")\r\n\r\n# Change the air density\r\nfi.reinitialize_flow_field(air_density=1.0) # TODO IS THIS RIGHT?\r\nfi.calculate_wake()\r\nplot_slice_shortcut(fi, axarr[6], \"Air Density=1.0\")\r\n\r\n# Change the farm layout\r\nfi.reinitialize_flow_field(layout_array=[[0, 500, 1000], [0, 0, 0]]) # TODO IS THIS RIGHT?\r\nfi.calculate_wake()\r\nplot_slice_shortcut(fi, axarr[7], \"Change layout\")\r\nwfct.visualization.plot_turbines_with_fi(axarr[7], fi)\r\n\r\n# Changes the yaw angles\r\nfi.calculate_wake(yaw_angles=[25, 25, 25])\r\nplot_slice_shortcut(fi, axarr[8], \"Change yaw angles\")\r\nwfct.visualization.plot_turbines_with_fi(axarr[8], fi)\r\n\r\n\r\nplt.show()",
"\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n"
],
[
"# Initialize the FLORIS interface fi\r\nfi = wfct.floris_interface.FlorisInterface(\"../../examples/example_input.json\")\r\n\r\n# Show the current model parameters\r\nprint(\"All the model parameters and their current values:\\n\")\r\nfi.show_model_parameters()\r\nprint(\"\\n\")\r\n\r\n# Show the current model parameters with docstring info\r\nprint(\"All the model parameters, their current values, and docstrings:\\n\")\r\nfi.show_model_parameters(verbose=True)\r\nprint(\"\\n\")\r\n\r\n# Show a specific model parameter with its docstring\r\nprint(\"A specific model parameter, its current value, and its docstring:\\n\")\r\nfi.show_model_parameters(params=[\"ka\"], verbose=False)\r\nprint(\"\\n\")\r\n\r\n# Get the current model parameters\r\nmodel_params = fi.get_model_parameters()\r\nprint(\"The current model parameters:\\n\")\r\nprint(model_params)\r\nprint(\"\\n\")\r\n\r\n# Set parameters on the current model\r\nprint(\"Set specific model parameters on the current wake model:\\n\")\r\nparams = {\r\n \"Wake Velocity Parameters\": {\"alpha\": 0.2},\r\n \"Wake Deflection Parameters\": {\"alpha\": 0.2},\r\n \"Wake Turbulence Parameters\": {\"ti_constant\": 1.0},\r\n}\r\nfi.set_model_parameters(params)\r\nprint(\"\\n\")\r\n\r\n# Check that the parameters were changed\r\nprint(\"Observe that the requested paremeters changes have been made:\\n\")\r\nmodel_params = fi.get_model_parameters()\r\nprint(model_params)\r\nprint(\"\\n\")",
"\u001b[34mfloris.simulation.wake_velocity.gaussianModels.gauss_legacy.LegacyGauss\u001b[0m \u001b[1;30mINFO\u001b[0m Current value of alpha, 0.2, is not equal to tuned value of 0.58.\n\u001b[34mfloris.simulation.wake_deflection.gauss.Gauss\u001b[0m \u001b[1;30mINFO\u001b[0m Current value of alpha, 0.2, is not equal to tuned value of 0.58.\n\u001b[34mfloris.simulation.wake_turbulence.crespo_hernandez.CrespoHernandez\u001b[0m \u001b[1;30mINFO\u001b[0m Current value of constant, 1.0, is not equal to tuned value of 0.5.\n"
],
[
"# Initialize the FLORIS interface for 4 seperate models defined as JSONS\r\nfi_jensen = wfct.floris_interface.FlorisInterface(\"../../examples/other_jsons/jensen.json\")\r\nfi_turbopark = wfct.floris_interface.FlorisInterface(\"../../examples/other_jsons/turbopark.json\")\r\nfi_mz = wfct.floris_interface.FlorisInterface(\"../../examples/other_jsons/multizone.json\")\r\nfi_gauss = wfct.floris_interface.FlorisInterface(\"../../examples/other_jsons/input_legacy.json\")\r\nfi_gch = wfct.floris_interface.FlorisInterface(\"../../examples/example_input.json\")\r\n\r\nfig, axarr = plt.subplots(2, 5, figsize=(16, 4))\r\n\r\n\r\n# Use a python for loop to iterate over the models and plot a horizontal cut through\r\n# of the models for an aligned and yaw case to show some differences\r\nfor idx, (fi, name) in enumerate(\r\n zip(\r\n [fi_jensen,fi_turbopark, fi_mz, fi_gauss, fi_gch],\r\n [\"Jensen\", \"TurbOPark\", \"Multizone\", \"Gaussian\", \"GCH\"]\r\n )\r\n):\r\n\r\n # Aligned case\r\n fi.calculate_wake(yaw_angles=[0])\r\n ax = axarr[0, idx]\r\n hor_plane = fi.get_hor_plane()\r\n wfct.visualization.visualize_cut_plane(hor_plane, ax=ax, minSpeed=4, maxSpeed=8)\r\n ax.set_title(name)\r\n axarr[0, 0].set_ylabel(\"Aligned\")\r\n\r\n # Yawed case\r\n fi.calculate_wake(yaw_angles=[25])\r\n ax = axarr[1, idx]\r\n hor_plane = fi.get_hor_plane()\r\n wfct.visualization.visualize_cut_plane(hor_plane, ax=ax, minSpeed=4, maxSpeed=8)\r\n axarr[1, 0].set_ylabel(\"Yawed\")\r\n\r\n\r\n# Show the figure\r\nplt.show()",
"\u001b[34mfloris.simulation.wake_turbulence.crespo_hernandez.CrespoHernandez\u001b[0m \u001b[1;30mINFO\u001b[0m Current value of constant, 0.73, is not equal to tuned value of 0.5.\n\u001b[34mfloris.simulation.wake_turbulence.crespo_hernandez.CrespoHernandez\u001b[0m \u001b[1;30mINFO\u001b[0m Current value of downstream, -0.275, is not equal to tuned value of -0.32.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\nc:\\users\\li hang\\documents\\projects\\floris\\floris\\simulation\\wake_velocity\\turbopark.py:128: RuntimeWarning: invalid value encountered in log\n Dwx = D + ((A * I0 * D) / beta) * (term1 - term2 - np.log(term3 / term4))\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n\u001b[34mfloris.tools.floris_interface.FlorisInterface\u001b[0m \u001b[1;30mINFO\u001b[0m Default to hub height = 90.0 for horizontal plane.\n"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab7114f97cbbea02f300aab1d69452f9512cd7c
| 6,165 |
ipynb
|
Jupyter Notebook
|
pytorch/examples/SRNN/SRNN_Example.ipynb
|
krantikiran/EdgeML
|
e5c7bd7c56884ca61f6d54cedb0074553cfdc896
|
[
"MIT"
] | 1 |
2020-03-26T17:19:54.000Z
|
2020-03-26T17:19:54.000Z
|
pytorch/examples/SRNN/SRNN_Example.ipynb
|
krantikiran/EdgeML
|
e5c7bd7c56884ca61f6d54cedb0074553cfdc896
|
[
"MIT"
] | 2 |
2020-03-26T02:59:12.000Z
|
2020-04-23T19:09:00.000Z
|
pytorch/examples/SRNN/SRNN_Example.ipynb
|
krantikiran/EdgeML
|
e5c7bd7c56884ca61f6d54cedb0074553cfdc896
|
[
"MIT"
] | 3 |
2020-03-25T18:45:39.000Z
|
2020-12-17T19:09:54.000Z
| 27.64574 | 146 | 0.568045 |
[
[
[
"## SRNN on Speech Commands Dataset\n\n\nPlease use `fetch_google.sh` to download the Google Speech Commands Dataset and python `process_google.py` to create feature extracted data.",
"_____no_output_____"
]
],
[
[
"from __future__ import print_function\nimport sys\nimport os\nimport numpy as np\nimport torch",
"_____no_output_____"
],
[
"from pytorch_edgeml.graph.rnn import SRNN2\nfrom pytorch_edgeml.trainer.srnnTrainer import SRNNTrainer\nimport pytorch_edgeml.utils as utils",
"_____no_output_____"
],
[
"device = torch.device(\"cuda:0\" if torch.cuda.is_available() else \"cpu\")\nDATA_DIR = './GoogleSpeech/Extracted/'",
"_____no_output_____"
],
[
"x_train_, y_train = np.load(DATA_DIR + 'x_train.npy'), np.load(DATA_DIR + 'y_train.npy')\nx_val_, y_val = np.load(DATA_DIR + 'x_val.npy'), np.load(DATA_DIR + 'y_val.npy')\nx_test_, y_test = np.load(DATA_DIR + 'x_test.npy'), np.load(DATA_DIR + 'y_test.npy')\n# Mean-var normalize\nmean = np.mean(np.reshape(x_train_, [-1, x_train_.shape[-1]]), axis=0)\nstd = np.std(np.reshape(x_train_, [-1, x_train_.shape[-1]]), axis=0)\nstd[std[:] < 0.000001] = 1\nx_train_ = (x_train_ - mean) / std\nx_val_ = (x_val_ - mean) / std\nx_test_ = (x_test_ - mean) / std\n\nx_train = np.swapaxes(x_train_, 0, 1)\nx_val = np.swapaxes(x_val_, 0, 1)\nx_test = np.swapaxes(x_test_, 0, 1)\nprint(\"Train shape\", x_train.shape, y_train.shape)\nprint(\"Val shape\", x_val.shape, y_val.shape)\nprint(\"Test shape\", x_test.shape, y_test.shape)",
"Train shape (99, 51088, 32) (51088, 13)\nVal shape (99, 6798, 32) (6798, 13)\nTest shape (99, 6835, 32) (6835, 13)\n"
],
[
"numTimeSteps = x_train.shape[0]\nnumInput = x_train.shape[-1]\nnumClasses = y_train.shape[1]\n\n# Network Parameters\nbrickSize = 11\nhiddenDim0 = 64\nhiddenDim1 = 32\ncellType = 'LSTM'\nlearningRate = 0.01\nbatchSize = 128\nepochs = 10",
"_____no_output_____"
],
[
"srnn2 = SRNN2(numInput, numClasses, hiddenDim0, hiddenDim1, cellType).to(device) \ntrainer = SRNNTrainer(srnn2, learningRate, lossType='xentropy', device=device)",
"Using x-entropy loss\n"
],
[
"trainer.train(brickSize, batchSize, epochs, x_train, x_val, y_train, y_val, printStep=200, valStep=5)",
"Epoch 0 batch 0 loss 4.295151 acc 0.031250\nEpoch 0 batch 200 loss 1.002617 acc 0.718750\nEpoch 1 batch 0 loss 0.647069 acc 0.796875\nEpoch 1 batch 200 loss 0.469229 acc 0.835938\nEpoch 2 batch 0 loss 0.388671 acc 0.882812\nEpoch 2 batch 200 loss 0.396696 acc 0.859375\nEpoch 3 batch 0 loss 0.266433 acc 0.921875\nEpoch 3 batch 200 loss 0.281694 acc 0.867188\nEpoch 4 batch 0 loss 0.302240 acc 0.906250\nEpoch 4 batch 200 loss 0.245797 acc 0.929688\nValidation accuracy: 0.911003\nEpoch 5 batch 0 loss 0.202542 acc 0.945312\nEpoch 5 batch 200 loss 0.192004 acc 0.929688\nEpoch 6 batch 0 loss 0.256735 acc 0.921875\nEpoch 6 batch 200 loss 0.279066 acc 0.921875\nEpoch 7 batch 0 loss 0.228837 acc 0.945312\nEpoch 7 batch 200 loss 0.222357 acc 0.937500\nEpoch 8 batch 0 loss 0.164639 acc 0.960938\nEpoch 8 batch 200 loss 0.160117 acc 0.945312\nEpoch 9 batch 0 loss 0.173849 acc 0.953125\nEpoch 9 batch 200 loss 0.201694 acc 0.929688\nValidation accuracy: 0.912474\n"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab727083992dea382cc7ab9d71141d9c3843d12
| 928,386 |
ipynb
|
Jupyter Notebook
|
src/.ipynb_checkpoints/Timing Exploration-checkpoint.ipynb
|
danielle707/stockx_machine_learning_analysis
|
abbbabdce235ceaf73e37af7665182a35c2aa5af
|
[
"MIT"
] | 12 |
2020-04-07T15:32:57.000Z
|
2022-01-24T21:19:11.000Z
|
src/.ipynb_checkpoints/Timing Exploration-checkpoint.ipynb
|
danielle707/StockX-Predictive-Modeling
|
abbbabdce235ceaf73e37af7665182a35c2aa5af
|
[
"MIT"
] | null | null | null |
src/.ipynb_checkpoints/Timing Exploration-checkpoint.ipynb
|
danielle707/StockX-Predictive-Modeling
|
abbbabdce235ceaf73e37af7665182a35c2aa5af
|
[
"MIT"
] | null | null | null | 1,061.012571 | 110,420 | 0.955799 |
[
[
[
"import pandas as pd\nimport numpy as np\nimport seaborn as sns\nimport matplotlib.pyplot as plt\nfrom PIL import Image\nfrom IPython.display import Image as im\n%matplotlib inline",
"_____no_output_____"
],
[
"data = pd.read_csv(\"../data/StockX-Data-Consolidated.csv\")",
"_____no_output_____"
],
[
"data['week_since_release'] = (data['Days Since Release']/7).round(1)",
"_____no_output_____"
],
[
"data.columns[21:32]",
"_____no_output_____"
],
[
"## Get brands and regions",
"_____no_output_____"
],
[
"def get_brand(row):\n for brand in data.columns[4:14]:\n if row[brand] == 1:\n return brand\ndef get_region(row):\n for region in data.columns[14:20]:\n if row[region] == 1:\n return region\ndef get_col(row):\n for color in data.columns[21:32]:\n if row[color] == 1:\n return color\n \ndata['brand'] = data.apply(get_brand, axis=1)\ndata['region'] = data.apply(get_region, axis=1)\ndata['color'] = data.apply(get_col, axis=1)",
"_____no_output_____"
],
[
"timing = data[['Days Since Release',\"week_since_release\",'region', \"brand\",'color','Pct_change']]",
"_____no_output_____"
],
[
"timing = timing.rename(columns = {'Days Since Release':\"days_since_release\"})",
"_____no_output_____"
],
[
"np.random.seed(19680801)\nN = 99956\ncolors = np.random.rand(N)\narea = (50 * np.random.rand(N))**2\nplt.scatter(x = timing['week_since_release'], y = timing['Pct_change'], c=colors, alpha=0.5)\nplt.title('Price premium on Weeks since release')\nplt.xlabel('weeks since release')\nplt.ylabel('price premium')\nplt.show()",
"_____no_output_____"
],
[
"timing.drop_duplicates([\"week_since_release\",'region'], inplace=True)\npivot = timing.pivot(index='region', columns='week_since_release', values='Pct_change',)\nax = sns.heatmap(pivot,annot=True,cmap = 'YlGnBu')\nplt.show()",
"_____no_output_____"
],
[
"df1 = timing[[\"week_since_release\",'region','Pct_change']]\nheatmap1_data = pd.pivot_table(df1,values='Pct_change', index=['region'], columns='week_since_release')\nheatmap1_data.head(n=5)\nsns.heatmap(heatmap1_data, cmap=\"BuGn\")",
"_____no_output_____"
],
[
"fig, ax = plt.subplots()\nsc = ax.scatter(timing.region,timing.week_since_release, c=timing.Pct_change, cmap=\"YlGnBu\")\nfig.colorbar(sc, ax=ax)\nplt.show()\n",
"_____no_output_____"
],
[
"fig, ax = plt.subplots()\nsc = ax.scatter(timing.brand,timing.week_since_release, c=timing.Pct_change, cmap=\"YlGnBu\")\nfig.colorbar(sc, ax=ax)\nplt.figure(figsize=(20, 60))\nplt.show()",
"_____no_output_____"
]
],
[
[
"## Nike off-white days/weeks since release",
"_____no_output_____"
]
],
[
[
"offwhite= timing.loc[timing['brand'] != 'yeezy']",
"_____no_output_____"
],
[
"ow_nowhite = offwhite.loc[offwhite['color'] != 'White']\now_white = offwhite.loc[offwhite['color'] == 'White']",
"_____no_output_____"
],
[
"ow_color = ow_nowhite.groupby(['color'])",
"_____no_output_____"
],
[
"img = plt.imread('../data/media/nike.jpg')",
"_____no_output_____"
],
[
"# Plot\nfig, ax = plt.subplots()\nax.imshow(img, aspect='auto', extent=(-80, 800, 0, 8), zorder=-1,alpha = 0.5)\nax.yaxis.tick_left()\n#ax.margins(0.05) # Optional, just adds 5% padding to the autoscaling\ncmap1 = sns.color_palette(\"Paired\")\ncmap2 = sns.color_palette(\"Set2\")\ncolors = [cmap1[1],cmap2[-1],cmap1[7],cmap1[4],'brown']\nfor i, (name, group) in enumerate(ow_color):\n ax.plot(group.days_since_release, group.Pct_change, marker='o', linestyle='', \n c = colors[i], ms=4, label=name, alpha = 0.2)\n\nax.spines['bottom'].set_color('white')\nax.xaxis.label.set_color('white')\nax.tick_params(axis='x', colors='white')\nax.spines['left'].set_color('white')\nax.yaxis.label.set_color('white')\nax.tick_params(axis='y', colors='white')\n#ax.patch.set_visible(False) \n \n\nplt.title('Nike: Off-White', fontsize = 'large', color = 'white' )\nplt.xlabel('Days Since Release', color = 'white' )\nplt.ylabel('Price Premium', color = 'white')\nplt.legend()\nplt.show()",
"_____no_output_____"
],
[
"offwhite['brand'].value_counts(sort=True, ascending=False, bins=None, dropna=True)",
"_____no_output_____"
],
[
"## Nike Off white Blazer\naj = offwhite.loc[offwhite['brand'] == 'airjordan']\naj_color = aj.groupby(['color'])\npresto = offwhite.loc[offwhite['brand'] == 'presto']\npresto_color = presto.groupby(['color'])\nzoom = offwhite.loc[offwhite['brand'] == 'zoom']\nzoom_color = zoom.groupby(['color'])\nblazer = offwhite.loc[offwhite['brand'] == 'blazer']\nblazer_color = blazer.groupby(['color'])\naf = offwhite.loc[offwhite['brand'] == 'airforce']\naf_color = af.groupby(['color'])",
"_____no_output_____"
],
[
"# AJ Plot\nfig, ax = plt.subplots()\nax.imshow(img, aspect='auto', extent=(-20, 500, -2, 8), zorder=-1,alpha = 0.4)\nax.yaxis.tick_left()\n#ax.margins(0.05) # Optional, just adds 5% padding to the autoscaling\ncmap1 = sns.color_palette(\"Paired\")\ncmap2 = sns.color_palette(\"Set2\")\ncolors = [cmap1[0],cmap2[-1],cmap1[7],cmap1[4],'brown']\nfor i, (name, group) in enumerate(aj_color):\n ax.plot(group.days_since_release, group.Pct_change, marker='o', linestyle='', \n c = colors[i], ms=4, label=name, alpha = 0.4)\n\nplt.title('Nike: Off-White Air Jordan', fontsize = 'large', color = 'white')\nplt.xlabel('Days Since Release', color = 'white')\nplt.ylabel('Price Premium', color = 'white')\nax.spines['bottom'].set_color('white')\nax.xaxis.label.set_color('white')\nax.tick_params(axis='x', colors='white')\nax.spines['left'].set_color('white')\nax.yaxis.label.set_color('white')\nax.tick_params(axis='y', colors='white')\nplt.legend()\nplt.show()",
"_____no_output_____"
],
[
"# Zoom Plot\nfig, ax = plt.subplots()\nax.imshow(img, aspect='auto', extent=(-20, 500, -2, 8), zorder=-1,alpha = 0.4)\nax.yaxis.tick_left()\n#ax.margins(0.05) # Optional, just adds 5% padding to the autoscaling\ncmap1 = sns.color_palette(\"Paired\")\ncmap2 = sns.color_palette(\"Set2\")\ncolors = [cmap1[1],cmap1[7],cmap1[4],cmap1[0]]\nfor i, (name, group) in enumerate(zoom_color):\n ax.plot(group.days_since_release, group.Pct_change, marker='o', linestyle='', \n c = colors[i], ms=4, label=name, alpha = 0.3)\n\nplt.title('Nike: Off-White Zoom', fontsize = 'large', color = 'white')\nplt.xlabel('Days Since Release', color = 'white')\nplt.ylabel('Price Premium', color = 'white')\nax.spines['bottom'].set_color('white')\nax.xaxis.label.set_color('white')\nax.tick_params(axis='x', colors='white')\nax.spines['left'].set_color('white')\nax.yaxis.label.set_color('white')\nax.tick_params(axis='y', colors='white')\nplt.legend()\nplt.show()",
"_____no_output_____"
],
[
"# Presto Plot\nfig, ax = plt.subplots()\nax.imshow(img, aspect='auto', extent=(-20, 500, 0, 8), zorder=-1,alpha = 0.4)\nax.yaxis.tick_left()\n#ax.margins(0.05) # Optional, just adds 5% padding to the autoscaling\ncmap1 = sns.color_palette(\"Paired\")\ncmap2 = sns.color_palette(\"Set2\")\ncolors = [cmap1[1],cmap1[0],cmap1[7],cmap1[4],'brown']\nfor i, (name, group) in enumerate(presto_color):\n ax.plot(group.days_since_release, group.Pct_change, marker='o', linestyle='', \n c = colors[i], ms=4, label=name, alpha = 0.3)\n\nplt.title('Nike: Off-White Presto', fontsize = 'large', color = 'white')\nplt.xlabel('Days Since Release', color = 'white')\nplt.ylabel('Price Premium', color = 'white')\nax.spines['bottom'].set_color('white')\nax.xaxis.label.set_color('white')\nax.tick_params(axis='x', colors='white')\nax.spines['left'].set_color('white')\nax.yaxis.label.set_color('white')\nax.tick_params(axis='y', colors='white')\nplt.legend()\nplt.show()",
"_____no_output_____"
],
[
"# Blazer Plot\nfig, ax = plt.subplots()\nax.imshow(img, aspect='auto', extent=(-20, 500, 0, 8), zorder=-1,alpha = 0.4)\nax.yaxis.tick_left()\n#ax.margins(0.05) # Optional, just adds 5% padding to the autoscaling\ncmap1 = sns.color_palette(\"Paired\")\ncmap2 = sns.color_palette(\"Set2\")\ncolors = [cmap1[1],cmap1[0],cmap1[7],cmap1[0]]\nfor i, (name, group) in enumerate(blazer_color):\n ax.plot(group.days_since_release, group.Pct_change, marker='o', linestyle='', \n c = colors[i], ms=4, label=name, alpha = 0.3)\n\nplt.title('Nike: Off-White Blazer', fontsize = 'large', color = 'white')\nplt.xlabel('Days Since Release', color = 'white')\nplt.ylabel('Price Premium', color = 'white')\nax.spines['bottom'].set_color('white')\nax.xaxis.label.set_color('white')\nax.tick_params(axis='x', colors='white')\nax.spines['left'].set_color('white')\nax.yaxis.label.set_color('white')\nax.tick_params(axis='y', colors='white')\nplt.legend()\nplt.show()",
"_____no_output_____"
],
[
"aj = offwhite.loc[offwhite['brand'] == 'airjordan']\naj_color = aj.groupby(['color'])",
"_____no_output_____"
],
[
"# Presto Plot\nfig, ax = plt.subplots()\nax.imshow(img, aspect='auto', extent=(-20, 500, 0, 8), zorder=-1,alpha = 0.4)\nax.yaxis.tick_left()\n#ax.margins(0.05) # Optional, just adds 5% padding to the autoscaling\ncmap1 = sns.color_palette(\"Paired\")\ncmap2 = sns.color_palette(\"Set2\")\ncolors = [cmap1[1],cmap1[0],cmap1[7],cmap1[4],'brown']\nfor i, (name, group) in enumerate(presto_color):\n ax.plot(group.days_since_release, group.Pct_change, marker='o', linestyle='', \n c = colors[i], ms=4, label=name, alpha = 0.2)\n\nplt.title('Nike: Off-White Presto', fontsize = 'large')\nplt.xlabel('Days Since Release')\nplt.ylabel('Price Premium')\nax.spines['bottom'].set_color('white')\nax.xaxis.label.set_color('white')\nax.tick_params(axis='x', colors='white')\nax.spines['left'].set_color('white')\nax.yaxis.label.set_color('white')\nax.tick_params(axis='y', colors='white')\nplt.legend()\nplt.show()",
"_____no_output_____"
],
[
"aj.shape",
"_____no_output_____"
],
[
"np.random.seed(19680801)\nN = 5703\ncolors = np.random.rand(N)\narea = (50 * np.random.rand(N))**2\nplt.scatter(x = aj['week_since_release'], y = aj['Pct_change'], c=colors, alpha=0.5)\nplt.title('AJ: Price premium on Weeks since release')\nplt.xlabel('weeks since release')\nplt.ylabel('price premium')\nplt.show()",
"_____no_output_____"
],
[
"np.random.seed(19680801)\nN = 3622\ncolors = np.random.rand(N)\narea = (50 * np.random.rand(N))**2\nplt.scatter(x = blazer['week_since_release'], y = blazer['Pct_change'], c=colors, alpha=0.5)\nplt.title('Blazer: Price premium on Weeks since release')\nplt.xlabel('weeks since release')\nplt.ylabel('price premium')\nplt.show()",
"_____no_output_____"
]
],
[
[
"## Yeezy days/weeks since release",
"_____no_output_____"
]
],
[
[
"timing.brand.unique()\nyeezy= timing.loc[timing['brand'] == 'yeezy']\nimg2 = plt.imread('../data/media/yeezy.jpg')",
"_____no_output_____"
],
[
"yeezy.color.unique()",
"_____no_output_____"
],
[
"yeezy_color = yeezy.groupby(['color'])",
"_____no_output_____"
],
[
"# Plot\nfig, ax = plt.subplots()\nax.imshow(img2, aspect='auto', extent=(-5, 1500, -2, 12), zorder=-1,alpha = 0.5)\nax.yaxis.tick_left()\n#ax.margins(0.05) # Optional, just adds 5% padding to the autoscaling\ncmap1 = sns.color_palette(\"Paired\")\ncmap2 = sns.color_palette(\"Set2\")\ncolors = [cmap1[1],cmap2[-1],cmap1[-1],cmap1[4],cmap1[0]]\nfor i, (name, group) in enumerate(yeezy_color):\n ax.plot(group.days_since_release, group.Pct_change, marker='o', linestyle='', \n c = colors[i], ms=4, label=name, alpha = 0.3)\n\nplt.title('Adidas: Yeezy', fontsize = 'large')\nplt.xlabel('Days Since Release')\nplt.ylabel('Price Premium')\nplt.legend()\nplt.show()",
"_____no_output_____"
],
[
"yeezy.shape",
"_____no_output_____"
],
[
"np.random.seed(19680801)\nN = 72162\ncolors = np.random.rand(N)\narea = (50 * np.random.rand(N))**2\nplt.scatter(x = yeezy['week_since_release'], y = yeezy['Pct_change'], c=colors, alpha=0.5)\nplt.title('Yeezy: Price premium on Weeks since release')\nplt.xlabel('weeks since release')\nplt.ylabel('price premium')\nplt.show()",
"_____no_output_____"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab7371b0df657f5a71ed8bb72b5d571903089db
| 347,372 |
ipynb
|
Jupyter Notebook
|
contrib/UTS PEMROGRAMAN LANJUT 18117014_18117034.ipynb
|
renzydwis/pilrek
|
39f83d58f9a86d231db29a05e31cd83b7f395737
|
[
"MIT"
] | 2 |
2019-09-06T23:30:14.000Z
|
2019-10-15T13:45:04.000Z
|
contrib/UTS PEMROGRAMAN LANJUT 18117014_18117034.ipynb
|
renzydwis/pilrek
|
39f83d58f9a86d231db29a05e31cd83b7f395737
|
[
"MIT"
] | 4 |
2019-10-14T06:06:28.000Z
|
2019-10-16T01:25:41.000Z
|
contrib/UTS PEMROGRAMAN LANJUT 18117014_18117034.ipynb
|
renzydwis/pilrek
|
39f83d58f9a86d231db29a05e31cd83b7f395737
|
[
"MIT"
] | 44 |
2019-09-04T17:19:03.000Z
|
2019-10-16T01:28:44.000Z
| 77.382936 | 63,920 | 0.636142 |
[
[
[
"import pandas as pd\nimport matplotlib.pyplot as plt\nfrom math import pi\n%matplotlib inline\nimport seaborn as sns",
"_____no_output_____"
],
[
"url = 'https://github.com/eueung/pilrek/raw/master/pilrek.csv'\ndata = pd.read_csv(url)",
"_____no_output_____"
],
[
"data",
"_____no_output_____"
],
[
"data_rektor4 = data.groupby('Alasan Memilih CaRek').get_group('Sesuai dengan kriteria kekinian (Rektor 4.0)')",
"_____no_output_____"
],
[
"data_rektor42 = data_rektor4.groupby('Kategori Umur Anda').get_group('< 25 th')",
"_____no_output_____"
],
[
"data_rektor42",
"_____no_output_____"
],
[
"ax = data_rektor42['CaRek Pilihan'].value_counts().plot(kind='bar', color=['black', 'red', 'green', 'blue', 'cyan','yellow', 'orange','purple','magenta'])\nplt.title('Bakal Calon Rektor Pilihan Pemilih Kategori < 25 th Karena Dianggap Kekinian') ",
"_____no_output_____"
],
[
"data_rektorpilma = data.groupby('Kategori Anda').get_group('Mahasiswa')",
"_____no_output_____"
],
[
"data_rektorpilma",
"_____no_output_____"
],
[
"ax = data_rektorpilma['Alasan Memilih CaRek'].value_counts().plot(kind='bar',color=['black', 'red', 'green', 'blue', 'cyan','yellow', 'orange','purple','magenta'])\nplt.title('Alasan Mahasiswa Memilih Bakal Calon Rektor Tersebut') ",
"_____no_output_____"
],
[
"data_acep = data.groupby('CaRek Pilihan').get_group('Acep Purqon (FMIPA-FI)')\ndata_acep",
"_____no_output_____"
],
[
"data_acep.sort_values(by='Alasan Memilih CaRek')",
"_____no_output_____"
],
[
"data_acep.groupby('Alasan Memilih CaRek').count()",
"_____no_output_____"
],
[
"acep_skill = data_acep.groupby('Alasan Memilih CaRek').count()[['CaRek Pilihan']].transpose()\nacep_skill",
"_____no_output_____"
],
[
"import numpy as np",
"_____no_output_____"
],
[
"labels=np.array(['Keberhasilan dan prestasi', 'Mendatangkan dana riset dan kerjasama eksternal, dalam dan luar negeri', 'Muda dan berenergi', 'Sesuai dengan kriteria kekinian (Rektor 4.0)', 'Tipe yg futuristik dan outside-the-box yg bisa menangkap peluang masa depan'])\nstats= acep_skill.loc['CaRek Pilihan', labels].values\nstats",
"_____no_output_____"
],
[
"angles=np.linspace(0, 2*np.pi, len(labels), endpoint=False)\nstats=np.concatenate((stats,[stats[0]]))\nangles=np.concatenate((angles,[angles[0]]))",
"_____no_output_____"
],
[
"import seaborn as sns",
"_____no_output_____"
],
[
"fig=plt.figure()\nax = fig.add_subplot(111, polar=True)\nax.plot(angles, stats, 'o-', linewidth=2)\nax.fill(angles, stats, alpha=0.25)\nax.set_thetagrids(angles * 180/np.pi, labels)\nax.set_title('Acep Furqon Top 5 Skill')\nax.grid(True)",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab73866b46d4a4c538582b09e9cab4c5007afcc
| 17,966 |
ipynb
|
Jupyter Notebook
|
notebook/libtree.ipynb
|
Kerilk/libtree
|
7d75dff754ff90e14fe1031a7fa071067c54cf5b
|
[
"BSD-2-Clause"
] | null | null | null |
notebook/libtree.ipynb
|
Kerilk/libtree
|
7d75dff754ff90e14fe1031a7fa071067c54cf5b
|
[
"BSD-2-Clause"
] | null | null | null |
notebook/libtree.ipynb
|
Kerilk/libtree
|
7d75dff754ff90e14fe1031a7fa071067c54cf5b
|
[
"BSD-2-Clause"
] | null | null | null | 20.842227 | 96 | 0.47512 |
[
[
[
"empty"
]
]
] |
[
"empty"
] |
[
[
"empty"
]
] |
4ab73b65c245454113566101295cbcfb3b451b0e
| 31,277 |
ipynb
|
Jupyter Notebook
|
clases y trabajos/8 class.ipynb
|
SebastianArrobo/Robotica-Industrial
|
8317b98cce99b07a258abb78dbab8cbf678d67de
|
[
"MIT"
] | null | null | null |
clases y trabajos/8 class.ipynb
|
SebastianArrobo/Robotica-Industrial
|
8317b98cce99b07a258abb78dbab8cbf678d67de
|
[
"MIT"
] | null | null | null |
clases y trabajos/8 class.ipynb
|
SebastianArrobo/Robotica-Industrial
|
8317b98cce99b07a258abb78dbab8cbf678d67de
|
[
"MIT"
] | null | null | null | 27.268527 | 142 | 0.373597 |
[
[
[
"import os\nimport pandas as pd\nimport numpy as np",
"_____no_output_____"
],
[
"!dir",
" El volumen de la unidad C es Windows\n El número de serie del volumen es: 9059-79E2\n\n Directorio de C:\\Users\\USUARIO\\Robotica Industrial\n\n30/03/2020 05:16 p. m. <DIR> .\n30/03/2020 05:16 p. m. <DIR> ..\n30/03/2020 05:10 p. m. <DIR> .ipynb_checkpoints\n25/01/2020 08:12 p. m. 1,339 1 class.ipynb\n26/01/2020 10:21 p. m. 2,707 1 tarea.ipynb\n27/01/2020 07:27 p. m. 5,843 2 class.ipynb\n10/02/2020 07:19 p. m. 416,263 3 class.ipynb\n17/02/2020 08:07 p. m. 40,636 4 Class.ipynb\n24/02/2020 08:10 p. m. 105,338 5 Class.ipynb\n02/03/2020 06:09 p. m. 313,317 6 class.ipynb\n09/03/2020 06:53 p. m. 19,682 7 class.ipynb\n30/03/2020 03:40 p. m. <DIR> Datasets\n27/01/2020 06:23 p. m. 21,843 FuncionesMod.ipynb\n24/02/2020 07:03 p. m. 5,222 respuesta.xlsx\n27/01/2020 08:18 p. m. 9,210 Untitled.ipynb\n30/03/2020 05:16 p. m. 9,098 Untitled1.ipynb\n 12 archivos 950,498 bytes\n 4 dirs 148,719,321,088 bytes libres\n"
],
[
"dataset_path = os.path.join(\"Datasets\",\"titanic3.csv\")\ntitanic = pd.read_csv(dataset_path)\nprint(titanic.columns)",
"Index(['pclass', 'survived', 'name', 'sex', 'age', 'sibsp', 'parch', 'ticket',\n 'fare', 'cabin', 'embarked', 'boat', 'body', 'home.dest'],\n dtype='object')\n"
],
[
"titanic.head()",
"_____no_output_____"
],
[
"titanic.shape",
"_____no_output_____"
],
[
"titanic.columns",
"_____no_output_____"
],
[
"labels = titanic.survived.values\nlabels",
"_____no_output_____"
],
[
"features = titanic[['pclass','sex', 'age', 'sibsp', 'parch','fare','embarked']]",
"_____no_output_____"
],
[
"features.head()",
"_____no_output_____"
],
[
"pd.get_dummies(features).head()",
"_____no_output_____"
],
[
"features_dummies = pd.get_dummies(features, columns=['pclass','sex','embarked'])\nfeatures_dummies.head()",
"_____no_output_____"
],
[
"features_dummies.shape",
"_____no_output_____"
],
[
"features_dummies.isna().any()",
"_____no_output_____"
],
[
"features_dummies_train= features_dummies.dropna()\nfeatures_dummies_train.isna().any",
"_____no_output_____"
],
[
"features_dummies_train.shape",
"_____no_output_____"
],
[
"labels_train = labels[features_dummies_train.index]\nlabels_train",
"_____no_output_____"
]
],
[
[
"# Logistic Regresion",
"_____no_output_____"
]
],
[
[
"from sklearn.linear_model import LogisticRegression",
"_____no_output_____"
],
[
"LogisticRegression?",
"_____no_output_____"
],
[
"model= LogisticRegression(solver ='lbfgs', max_iter=500)",
"_____no_output_____"
],
[
"model.fit(features_dummies_train, labels_train)",
"_____no_output_____"
],
[
"labels_train #real",
"_____no_output_____"
],
[
"model.predict(features_dummies_train)#predict",
"_____no_output_____"
],
[
"sum(model.predict(features_dummies_train) == labels_train)",
"_____no_output_____"
],
[
"sum(model.predict(features_dummies_train) == labels_train)/(len(labels_train))",
"_____no_output_____"
]
],
[
[
" P(A)/P(n)",
"_____no_output_____"
]
],
[
[
"len(labels_train)",
"_____no_output_____"
],
[
"828/1045",
"_____no_output_____"
],
[
"print(\"Logistic regressio score: \", model.score(features_dummies_train, labels_train))",
"Logistic regressio score: 0.7923444976076555\n"
],
[
"from sklearn.model_selection import train_test_split",
"_____no_output_____"
],
[
"train_data, test_data, train_labels, test_labels = train_test_split(features_dummies_train, labels_train, random_state=0,test_size=0.33)",
"_____no_output_____"
],
[
"train_data.shape, test_data.shape",
"_____no_output_____"
],
[
"train_labels.shape,test_labels.shape",
"_____no_output_____"
],
[
"model = LogisticRegression(solver='lbfgs', max_iter=500).fit(train_data, train_labels)\nprint(\"Logistic regressio score: %f \"% model.score(test_data, test_labels))",
"Logistic regressio score: 0.788406 \n"
],
[
"[1, 'female', 29.0,0,0,211.3375]",
"_____no_output_____"
]
]
] |
[
"code",
"markdown",
"code",
"raw",
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"raw"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab73c6f3ba44e38d25bf150859b648571836f05
| 571,276 |
ipynb
|
Jupyter Notebook
|
course1: Neural Networks and Deep Learning/Planar_data_classification_with_onehidden_layer_v6c.ipynb
|
asenarmour/deep-learning-specialization-coursera
|
fb844de6d2f12343ce19eb1a9d356bdcfdcb8237
|
[
"MIT"
] | null | null | null |
course1: Neural Networks and Deep Learning/Planar_data_classification_with_onehidden_layer_v6c.ipynb
|
asenarmour/deep-learning-specialization-coursera
|
fb844de6d2f12343ce19eb1a9d356bdcfdcb8237
|
[
"MIT"
] | null | null | null |
course1: Neural Networks and Deep Learning/Planar_data_classification_with_onehidden_layer_v6c.ipynb
|
asenarmour/deep-learning-specialization-coursera
|
fb844de6d2f12343ce19eb1a9d356bdcfdcb8237
|
[
"MIT"
] | null | null | null | 345.599516 | 331,088 | 0.913938 |
[
[
[
"\n### <font color = \"darkblue\">Updates to Assignment</font>\n\n#### If you were working on the older version:\n* Please click on the \"Coursera\" icon in the top right to open up the folder directory. \n* Navigate to the folder: Week 3/ Planar data classification with one hidden layer. You can see your prior work in version 6b: \"Planar data classification with one hidden layer v6b.ipynb\"\n\n#### List of bug fixes and enhancements\n* Clarifies that the classifier will learn to classify regions as either red or blue.\n* compute_cost function fixes np.squeeze by casting it as a float.\n* compute_cost instructions clarify the purpose of np.squeeze.\n* compute_cost clarifies that \"parameters\" parameter is not needed, but is kept in the function definition until the auto-grader is also updated.\n* nn_model removes extraction of parameter values, as the entire parameter dictionary is passed to the invoked functions.",
"_____no_output_____"
],
[
"# Planar data classification with one hidden layer\n\nWelcome to your week 3 programming assignment. It's time to build your first neural network, which will have a hidden layer. You will see a big difference between this model and the one you implemented using logistic regression. \n\n**You will learn how to:**\n- Implement a 2-class classification neural network with a single hidden layer\n- Use units with a non-linear activation function, such as tanh \n- Compute the cross entropy loss \n- Implement forward and backward propagation\n",
"_____no_output_____"
],
[
"## 1 - Packages ##\n\nLet's first import all the packages that you will need during this assignment.\n- [numpy](https://www.numpy.org/) is the fundamental package for scientific computing with Python.\n- [sklearn](http://scikit-learn.org/stable/) provides simple and efficient tools for data mining and data analysis. \n- [matplotlib](http://matplotlib.org) is a library for plotting graphs in Python.\n- testCases provides some test examples to assess the correctness of your functions\n- planar_utils provide various useful functions used in this assignment",
"_____no_output_____"
]
],
[
[
"# Package imports\nimport numpy as np\nimport matplotlib.pyplot as plt\nfrom testCases_v2 import *\nimport sklearn\nimport sklearn.datasets\nimport sklearn.linear_model\nfrom planar_utils import plot_decision_boundary, sigmoid, load_planar_dataset, load_extra_datasets\n\n%matplotlib inline\n\nnp.random.seed(1) # set a seed so that the results are consistent",
"_____no_output_____"
]
],
[
[
"## 2 - Dataset ##\n\nFirst, let's get the dataset you will work on. The following code will load a \"flower\" 2-class dataset into variables `X` and `Y`.",
"_____no_output_____"
]
],
[
[
"X, Y = load_planar_dataset()",
"_____no_output_____"
]
],
[
[
"Visualize the dataset using matplotlib. The data looks like a \"flower\" with some red (label y=0) and some blue (y=1) points. Your goal is to build a model to fit this data. In other words, we want the classifier to define regions as either red or blue.",
"_____no_output_____"
]
],
[
[
"# Visualize the data:\nplt.scatter(X[0, :], X[1, :], c=Y, s=40, cmap=plt.cm.Spectral);",
"_____no_output_____"
],
[
"plt.figure(figsize=(35,30))\nplt.imshow(X[:1])",
"_____no_output_____"
]
],
[
[
"You have:\n - a numpy-array (matrix) X that contains your features (x1, x2)\n - a numpy-array (vector) Y that contains your labels (red:0, blue:1).\n\nLets first get a better sense of what our data is like. \n\n**Exercise**: How many training examples do you have? In addition, what is the `shape` of the variables `X` and `Y`? \n\n**Hint**: How do you get the shape of a numpy array? [(help)](https://docs.scipy.org/doc/numpy/reference/generated/numpy.ndarray.shape.html)",
"_____no_output_____"
]
],
[
[
"### START CODE HERE ### (≈ 3 lines of code)\nshape_X = X.shape\nshape_Y = Y.shape\nm = X.shape[1] # training set size\n### END CODE HERE ###\n\nprint ('The shape of X is: ' + str(shape_X))\nprint ('The shape of Y is: ' + str(shape_Y))\nprint ('I have m = %d training examples!' % (m))",
"The shape of X is: (2, 400)\nThe shape of Y is: (1, 400)\nI have m = 400 training examples!\n"
]
],
[
[
"**Expected Output**:\n \n<table style=\"width:20%\">\n \n <tr>\n <td>**shape of X**</td>\n <td> (2, 400) </td> \n </tr>\n \n <tr>\n <td>**shape of Y**</td>\n <td>(1, 400) </td> \n </tr>\n \n <tr>\n <td>**m**</td>\n <td> 400 </td> \n </tr>\n \n</table>",
"_____no_output_____"
],
[
"## 3 - Simple Logistic Regression\n\nBefore building a full neural network, lets first see how logistic regression performs on this problem. You can use sklearn's built-in functions to do that. Run the code below to train a logistic regression classifier on the dataset.",
"_____no_output_____"
]
],
[
[
"# Train the logistic regression classifier\nclf = sklearn.linear_model.LogisticRegressionCV();\nclf.fit(X.T, Y.T);",
"_____no_output_____"
]
],
[
[
"You can now plot the decision boundary of these models. Run the code below.",
"_____no_output_____"
]
],
[
[
"# Plot the decision boundary for logistic regression\nplot_decision_boundary(lambda x: clf.predict(x), X, Y)\nplt.title(\"Logistic Regression\")\n\n# Print accuracy\nLR_predictions = clf.predict(X.T)\nprint ('Accuracy of logistic regression: %d ' % float((np.dot(Y,LR_predictions) + np.dot(1-Y,1-LR_predictions))/float(Y.size)*100) +\n '% ' + \"(percentage of correctly labelled datapoints)\")",
"Accuracy of logistic regression: 47 % (percentage of correctly labelled datapoints)\n"
]
],
[
[
"**Expected Output**:\n\n<table style=\"width:20%\">\n <tr>\n <td>**Accuracy**</td>\n <td> 47% </td> \n </tr>\n \n</table>\n",
"_____no_output_____"
],
[
"**Interpretation**: The dataset is not linearly separable, so logistic regression doesn't perform well. Hopefully a neural network will do better. Let's try this now! ",
"_____no_output_____"
],
[
"## 4 - Neural Network model\n\nLogistic regression did not work well on the \"flower dataset\". You are going to train a Neural Network with a single hidden layer.\n\n**Here is our model**:\n<img src=\"images/classification_kiank.png\" style=\"width:600px;height:300px;\">\n\n**Mathematically**:\n\nFor one example $x^{(i)}$:\n$$z^{[1] (i)} = W^{[1]} x^{(i)} + b^{[1]}\\tag{1}$$ \n$$a^{[1] (i)} = \\tanh(z^{[1] (i)})\\tag{2}$$\n$$z^{[2] (i)} = W^{[2]} a^{[1] (i)} + b^{[2]}\\tag{3}$$\n$$\\hat{y}^{(i)} = a^{[2] (i)} = \\sigma(z^{ [2] (i)})\\tag{4}$$\n$$y^{(i)}_{prediction} = \\begin{cases} 1 & \\mbox{if } a^{[2](i)} > 0.5 \\\\ 0 & \\mbox{otherwise } \\end{cases}\\tag{5}$$\n\nGiven the predictions on all the examples, you can also compute the cost $J$ as follows: \n$$J = - \\frac{1}{m} \\sum\\limits_{i = 0}^{m} \\large\\left(\\small y^{(i)}\\log\\left(a^{[2] (i)}\\right) + (1-y^{(i)})\\log\\left(1- a^{[2] (i)}\\right) \\large \\right) \\small \\tag{6}$$\n\n**Reminder**: The general methodology to build a Neural Network is to:\n 1. Define the neural network structure ( # of input units, # of hidden units, etc). \n 2. Initialize the model's parameters\n 3. Loop:\n - Implement forward propagation\n - Compute loss\n - Implement backward propagation to get the gradients\n - Update parameters (gradient descent)\n\nYou often build helper functions to compute steps 1-3 and then merge them into one function we call `nn_model()`. Once you've built `nn_model()` and learnt the right parameters, you can make predictions on new data.",
"_____no_output_____"
],
[
"### 4.1 - Defining the neural network structure ####\n\n**Exercise**: Define three variables:\n - n_x: the size of the input layer\n - n_h: the size of the hidden layer (set this to 4) \n - n_y: the size of the output layer\n\n**Hint**: Use shapes of X and Y to find n_x and n_y. Also, hard code the hidden layer size to be 4.",
"_____no_output_____"
]
],
[
[
"# GRADED FUNCTION: layer_sizes\n\ndef layer_sizes(X, Y):\n \"\"\"\n Arguments:\n X -- input dataset of shape (input size, number of examples)\n Y -- labels of shape (output size, number of examples)\n \n Returns:\n n_x -- the size of the input layer\n n_h -- the size of the hidden layer\n n_y -- the size of the output layer\n \"\"\"\n ### START CODE HERE ### (≈ 3 lines of code)\n n_x = X.shape[0] # size of input layer\n n_h = 4\n n_y = Y.shape[0] # size of output layer\n ### END CODE HERE ###\n return (n_x, n_h, n_y)",
"_____no_output_____"
],
[
"X_assess, Y_assess = layer_sizes_test_case()\n(n_x, n_h, n_y) = layer_sizes(X_assess, Y_assess)\nprint(\"The size of the input layer is: n_x = \" + str(n_x))\nprint(\"The size of the hidden layer is: n_h = \" + str(n_h))\nprint(\"The size of the output layer is: n_y = \" + str(n_y))",
"The size of the input layer is: n_x = 5\nThe size of the hidden layer is: n_h = 4\nThe size of the output layer is: n_y = 2\n"
]
],
[
[
"**Expected Output** (these are not the sizes you will use for your network, they are just used to assess the function you've just coded).\n\n<table style=\"width:20%\">\n <tr>\n <td>**n_x**</td>\n <td> 5 </td> \n </tr>\n \n <tr>\n <td>**n_h**</td>\n <td> 4 </td> \n </tr>\n \n <tr>\n <td>**n_y**</td>\n <td> 2 </td> \n </tr>\n \n</table>",
"_____no_output_____"
],
[
"### 4.2 - Initialize the model's parameters ####\n\n**Exercise**: Implement the function `initialize_parameters()`.\n\n**Instructions**:\n- Make sure your parameters' sizes are right. Refer to the neural network figure above if needed.\n- You will initialize the weights matrices with random values. \n - Use: `np.random.randn(a,b) * 0.01` to randomly initialize a matrix of shape (a,b).\n- You will initialize the bias vectors as zeros. \n - Use: `np.zeros((a,b))` to initialize a matrix of shape (a,b) with zeros.",
"_____no_output_____"
]
],
[
[
"# GRADED FUNCTION: initialize_parameters\n\ndef initialize_parameters(n_x, n_h, n_y):\n \"\"\"\n Argument:\n n_x -- size of the input layer\n n_h -- size of the hidden layer\n n_y -- size of the output layer\n \n Returns:\n params -- python dictionary containing your parameters:\n W1 -- weight matrix of shape (n_h, n_x)\n b1 -- bias vector of shape (n_h, 1)\n W2 -- weight matrix of shape (n_y, n_h)\n b2 -- bias vector of shape (n_y, 1)\n \"\"\"\n \n np.random.seed(2) # we set up a seed so that your output matches ours although the initialization is random.\n \n ### START CODE HERE ### (≈ 4 lines of code)\n W1 = np.random.randn(n_h,n_x)*0.01\n b1 = np.zeros((n_h,1))\n W2 = np.random.randn(n_y,n_h)*0.01\n b2 = np.zeros((n_y,1))\n ### END CODE HERE ###\n \n assert (W1.shape == (n_h, n_x))\n assert (b1.shape == (n_h, 1))\n assert (W2.shape == (n_y, n_h))\n assert (b2.shape == (n_y, 1))\n \n parameters = {\"W1\": W1,\n \"b1\": b1,\n \"W2\": W2,\n \"b2\": b2}\n \n return parameters",
"_____no_output_____"
],
[
"n_x, n_h, n_y = initialize_parameters_test_case()\n\nparameters = initialize_parameters(n_x, n_h, n_y)\nprint(\"W1 = \" + str(parameters[\"W1\"]))\nprint(\"b1 = \" + str(parameters[\"b1\"]))\nprint(\"W2 = \" + str(parameters[\"W2\"]))\nprint(\"b2 = \" + str(parameters[\"b2\"]))",
"W1 = [[-0.00416758 -0.00056267]\n [-0.02136196 0.01640271]\n [-0.01793436 -0.00841747]\n [ 0.00502881 -0.01245288]]\nb1 = [[ 0.]\n [ 0.]\n [ 0.]\n [ 0.]]\nW2 = [[-0.01057952 -0.00909008 0.00551454 0.02292208]]\nb2 = [[ 0.]]\n"
]
],
[
[
"**Expected Output**:\n\n<table style=\"width:90%\">\n <tr>\n <td>**W1**</td>\n <td> [[-0.00416758 -0.00056267]\n [-0.02136196 0.01640271]\n [-0.01793436 -0.00841747]\n [ 0.00502881 -0.01245288]] </td> \n </tr>\n \n <tr>\n <td>**b1**</td>\n <td> [[ 0.]\n [ 0.]\n [ 0.]\n [ 0.]] </td> \n </tr>\n \n <tr>\n <td>**W2**</td>\n <td> [[-0.01057952 -0.00909008 0.00551454 0.02292208]]</td> \n </tr>\n \n\n <tr>\n <td>**b2**</td>\n <td> [[ 0.]] </td> \n </tr>\n \n</table>\n\n",
"_____no_output_____"
],
[
"### 4.3 - The Loop ####\n\n**Question**: Implement `forward_propagation()`.\n\n**Instructions**:\n- Look above at the mathematical representation of your classifier.\n- You can use the function `sigmoid()`. It is built-in (imported) in the notebook.\n- You can use the function `np.tanh()`. It is part of the numpy library.\n- The steps you have to implement are:\n 1. Retrieve each parameter from the dictionary \"parameters\" (which is the output of `initialize_parameters()`) by using `parameters[\"..\"]`.\n 2. Implement Forward Propagation. Compute $Z^{[1]}, A^{[1]}, Z^{[2]}$ and $A^{[2]}$ (the vector of all your predictions on all the examples in the training set).\n- Values needed in the backpropagation are stored in \"`cache`\". The `cache` will be given as an input to the backpropagation function.",
"_____no_output_____"
]
],
[
[
"X.shape",
"_____no_output_____"
],
[
"# GRADED FUNCTION: forward_propagation\n\ndef forward_propagation(X, parameters):\n \"\"\"\n Argument:\n X -- input data of size (n_x, m)\n parameters -- python dictionary containing your parameters (output of initialization function)\n \n Returns:\n A2 -- The sigmoid output of the second activation\n cache -- a dictionary containing \"Z1\", \"A1\", \"Z2\" and \"A2\"\n \"\"\"\n # Retrieve each parameter from the dictionary \"parameters\"\n ### START CODE HERE ### (≈ 4 lines of code)\n W1 = parameters['W1']\n b1 = parameters['b1']\n W2 = parameters['W2']\n b2 = parameters['b2']\n ### END CODE HERE ###\n \n # Implement Forward Propagation to calculate A2 (probabilities)\n ### START CODE HERE ### (≈ 4 lines of code)\n Z1 = np.dot(W1,X)+b1\n A1 = np.tanh(Z1)\n Z2 = np.dot(W2,A1)+b2\n A2 = sigmoid(Z2)\n ### END CODE HERE ###\n \n assert(A2.shape == (1, X.shape[1]))\n \n cache = {\"Z1\": Z1,\n \"A1\": A1,\n \"Z2\": Z2,\n \"A2\": A2}\n \n return A2, cache",
"_____no_output_____"
],
[
"X_assess, parameters = forward_propagation_test_case()\nA2, cache = forward_propagation(X_assess, parameters)\n\n# Note: we use the mean here just to make sure that your output matches ours. \nprint(np.mean(cache['Z1']) ,np.mean(cache['A1']),np.mean(cache['Z2']),np.mean(cache['A2']))",
"0.262818640198 0.091999045227 -1.30766601287 0.212877681719\n"
]
],
[
[
"**Expected Output**:\n<table style=\"width:50%\">\n <tr>\n <td> 0.262818640198 0.091999045227 -1.30766601287 0.212877681719 </td> \n </tr>\n</table>",
"_____no_output_____"
],
[
"Now that you have computed $A^{[2]}$ (in the Python variable \"`A2`\"), which contains $a^{[2](i)}$ for every example, you can compute the cost function as follows:\n\n$$J = - \\frac{1}{m} \\sum\\limits_{i = 1}^{m} \\large{(} \\small y^{(i)}\\log\\left(a^{[2] (i)}\\right) + (1-y^{(i)})\\log\\left(1- a^{[2] (i)}\\right) \\large{)} \\small\\tag{13}$$\n\n**Exercise**: Implement `compute_cost()` to compute the value of the cost $J$.\n\n**Instructions**:\n- There are many ways to implement the cross-entropy loss. To help you, we give you how we would have implemented\n$- \\sum\\limits_{i=0}^{m} y^{(i)}\\log(a^{[2](i)})$:\n```python\nlogprobs = np.multiply(np.log(A2),Y)\ncost = - np.sum(logprobs) # no need to use a for loop!\n```\n\n(you can use either `np.multiply()` and then `np.sum()` or directly `np.dot()`). \nNote that if you use `np.multiply` followed by `np.sum` the end result will be a type `float`, whereas if you use `np.dot`, the result will be a 2D numpy array. We can use `np.squeeze()` to remove redundant dimensions (in the case of single float, this will be reduced to a zero-dimension array). We can cast the array as a type `float` using `float()`.",
"_____no_output_____"
]
],
[
[
"# GRADED FUNCTION: compute_cost\n\ndef compute_cost(A2, Y, parameters):\n \"\"\"\n Computes the cross-entropy cost given in equation (13)\n \n Arguments:\n A2 -- The sigmoid output of the second activation, of shape (1, number of examples)\n Y -- \"true\" labels vector of shape (1, number of examples)\n parameters -- python dictionary containing your parameters W1, b1, W2 and b2\n [Note that the parameters argument is not used in this function, \n but the auto-grader currently expects this parameter.\n Future version of this notebook will fix both the notebook \n and the auto-grader so that `parameters` is not needed.\n For now, please include `parameters` in the function signature,\n and also when invoking this function.]\n \n Returns:\n cost -- cross-entropy cost given equation (13)\n \n \"\"\"\n \n m = Y.shape[1] # number of example\n\n # Compute the cross-entropy cost\n ### START CODE HERE ### (≈ 2 lines of code)\n logprobs = np.multiply(np.log(A2),Y)\n cost = -np.sum(logprobs)\n ### END CODE HERE ###\n \n cost = float(np.squeeze(cost)) # makes sure cost is the dimension we expect. \n # E.g., turns [[17]] into 17 \n assert(isinstance(cost, float))\n \n return cost",
"_____no_output_____"
],
[
"A2, Y_assess, parameters = compute_cost_test_case()\n\nprint(\"cost = \" + str(compute_cost(A2, Y_assess, parameters)))",
"cost = 0.6926858869721941\n"
]
],
[
[
"**Expected Output**:\n<table style=\"width:20%\">\n <tr>\n <td>**cost**</td>\n <td> 0.693058761... </td> \n </tr>\n \n</table>",
"_____no_output_____"
],
[
"Using the cache computed during forward propagation, you can now implement backward propagation.\n\n**Question**: Implement the function `backward_propagation()`.\n\n**Instructions**:\nBackpropagation is usually the hardest (most mathematical) part in deep learning. To help you, here again is the slide from the lecture on backpropagation. You'll want to use the six equations on the right of this slide, since you are building a vectorized implementation. \n\n<img src=\"images/grad_summary.png\" style=\"width:600px;height:300px;\">\n\n<!--\n$\\frac{\\partial \\mathcal{J} }{ \\partial z_{2}^{(i)} } = \\frac{1}{m} (a^{[2](i)} - y^{(i)})$\n\n$\\frac{\\partial \\mathcal{J} }{ \\partial W_2 } = \\frac{\\partial \\mathcal{J} }{ \\partial z_{2}^{(i)} } a^{[1] (i) T} $\n\n$\\frac{\\partial \\mathcal{J} }{ \\partial b_2 } = \\sum_i{\\frac{\\partial \\mathcal{J} }{ \\partial z_{2}^{(i)}}}$\n\n$\\frac{\\partial \\mathcal{J} }{ \\partial z_{1}^{(i)} } = W_2^T \\frac{\\partial \\mathcal{J} }{ \\partial z_{2}^{(i)} } * ( 1 - a^{[1] (i) 2}) $\n\n$\\frac{\\partial \\mathcal{J} }{ \\partial W_1 } = \\frac{\\partial \\mathcal{J} }{ \\partial z_{1}^{(i)} } X^T $\n\n$\\frac{\\partial \\mathcal{J} _i }{ \\partial b_1 } = \\sum_i{\\frac{\\partial \\mathcal{J} }{ \\partial z_{1}^{(i)}}}$\n\n- Note that $*$ denotes elementwise multiplication.\n- The notation you will use is common in deep learning coding:\n - dW1 = $\\frac{\\partial \\mathcal{J} }{ \\partial W_1 }$\n - db1 = $\\frac{\\partial \\mathcal{J} }{ \\partial b_1 }$\n - dW2 = $\\frac{\\partial \\mathcal{J} }{ \\partial W_2 }$\n - db2 = $\\frac{\\partial \\mathcal{J} }{ \\partial b_2 }$\n \n!-->\n\n- Tips:\n - To compute dZ1 you'll need to compute $g^{[1]'}(Z^{[1]})$. Since $g^{[1]}(.)$ is the tanh activation function, if $a = g^{[1]}(z)$ then $g^{[1]'}(z) = 1-a^2$. So you can compute \n $g^{[1]'}(Z^{[1]})$ using `(1 - np.power(A1, 2))`.",
"_____no_output_____"
]
],
[
[
"# GRADED FUNCTION: backward_propagation\n\ndef backward_propagation(parameters, cache, X, Y):\n \"\"\"\n Implement the backward propagation using the instructions above.\n \n Arguments:\n parameters -- python dictionary containing our parameters \n cache -- a dictionary containing \"Z1\", \"A1\", \"Z2\" and \"A2\".\n X -- input data of shape (2, number of examples)\n Y -- \"true\" labels vector of shape (1, number of examples)\n \n Returns:\n grads -- python dictionary containing your gradients with respect to different parameters\n \"\"\"\n m = X.shape[1]\n \n # First, retrieve W1 and W2 from the dictionary \"parameters\".\n ### START CODE HERE ### (≈ 2 lines of code)\n W1 = parameters['W1']\n W2 = parameters['W2']\n ### END CODE HERE ###\n \n # Retrieve also A1 and A2 from dictionary \"cache\".\n ### START CODE HERE ### (≈ 2 lines of code)\n A1 = cache['A1']\n A2 = cache['A2']\n ### END CODE HERE ###\n \n # Backward propagation: calculate dW1, db1, dW2, db2. \n ### START CODE HERE ### (≈ 6 lines of code, corresponding to 6 equations on slide above)\n dZ2 = A2-Y\n dW2 = (1/m)*np.dot(dZ2,A1.T)\n db2 = (1/m)*np.sum(dZ2,axis=1,keepdims=True)\n dZ1 = np.dot(W2.T,dZ2)*(1-np.power(A1,2))\n dW1 = (1/m)*np.dot(dZ1,X.T)\n db1 = (1/m)*np.sum(dZ1,axis=1,keepdims=True)\n ### END CODE HERE ###\n \n grads = {\"dW1\": dW1,\n \"db1\": db1,\n \"dW2\": dW2,\n \"db2\": db2}\n \n return grads",
"_____no_output_____"
],
[
"parameters, cache, X_assess, Y_assess = backward_propagation_test_case()\n\ngrads = backward_propagation(parameters, cache, X_assess, Y_assess)\nprint (\"dW1 = \"+ str(grads[\"dW1\"]))\nprint (\"db1 = \"+ str(grads[\"db1\"]))\nprint (\"dW2 = \"+ str(grads[\"dW2\"]))\nprint (\"db2 = \"+ str(grads[\"db2\"]))",
"dW1 = [[ 0.00301023 -0.00747267]\n [ 0.00257968 -0.00641288]\n [-0.00156892 0.003893 ]\n [-0.00652037 0.01618243]]\ndb1 = [[ 0.00176201]\n [ 0.00150995]\n [-0.00091736]\n [-0.00381422]]\ndW2 = [[ 0.00078841 0.01765429 -0.00084166 -0.01022527]]\ndb2 = [[-0.16655712]]\n"
]
],
[
[
"**Expected output**:\n\n\n\n<table style=\"width:80%\">\n <tr>\n <td>**dW1**</td>\n <td> [[ 0.00301023 -0.00747267]\n [ 0.00257968 -0.00641288]\n [-0.00156892 0.003893 ]\n [-0.00652037 0.01618243]] </td> \n </tr>\n \n <tr>\n <td>**db1**</td>\n <td> [[ 0.00176201]\n [ 0.00150995]\n [-0.00091736]\n [-0.00381422]] </td> \n </tr>\n \n <tr>\n <td>**dW2**</td>\n <td> [[ 0.00078841 0.01765429 -0.00084166 -0.01022527]] </td> \n </tr>\n \n\n <tr>\n <td>**db2**</td>\n <td> [[-0.16655712]] </td> \n </tr>\n \n</table> ",
"_____no_output_____"
],
[
"**Question**: Implement the update rule. Use gradient descent. You have to use (dW1, db1, dW2, db2) in order to update (W1, b1, W2, b2).\n\n**General gradient descent rule**: $ \\theta = \\theta - \\alpha \\frac{\\partial J }{ \\partial \\theta }$ where $\\alpha$ is the learning rate and $\\theta$ represents a parameter.\n\n**Illustration**: The gradient descent algorithm with a good learning rate (converging) and a bad learning rate (diverging). Images courtesy of Adam Harley.\n\n<img src=\"images/sgd.gif\" style=\"width:400;height:400;\"> <img src=\"images/sgd_bad.gif\" style=\"width:400;height:400;\">\n\n",
"_____no_output_____"
]
],
[
[
"# GRADED FUNCTION: update_parameters\n\ndef update_parameters(parameters, grads, learning_rate = 1.2):\n \"\"\"\n Updates parameters using the gradient descent update rule given above\n \n Arguments:\n parameters -- python dictionary containing your parameters \n grads -- python dictionary containing your gradients \n \n Returns:\n parameters -- python dictionary containing your updated parameters \n \"\"\"\n # Retrieve each parameter from the dictionary \"parameters\"\n ### START CODE HERE ### (≈ 4 lines of code)\n W1 = parameters['W1']\n b1 = parameters['b1']\n W2 = parameters['W2']\n b2 = parameters['b2']\n ### END CODE HERE ###\n \n # Retrieve each gradient from the dictionary \"grads\"\n ### START CODE HERE ### (≈ 4 lines of code)\n dW1 = grads['dW1']\n db1 = grads['db1']\n dW2 = grads['dW2']\n db2 = grads['db2']\n ## END CODE HERE ###\n \n # Update rule for each parameter\n ### START CODE HERE ### (≈ 4 lines of code)\n W1 = W1-learning_rate*dW1\n b1 = b1-learning_rate*db1\n W2 = W2-learning_rate*dW2\n b2 = b2-learning_rate*db2\n ### END CODE HERE ###\n \n parameters = {\"W1\": W1,\n \"b1\": b1,\n \"W2\": W2,\n \"b2\": b2}\n \n return parameters",
"_____no_output_____"
],
[
"parameters, grads = update_parameters_test_case()\nparameters = update_parameters(parameters, grads)\n\nprint(\"W1 = \" + str(parameters[\"W1\"]))\nprint(\"b1 = \" + str(parameters[\"b1\"]))\nprint(\"W2 = \" + str(parameters[\"W2\"]))\nprint(\"b2 = \" + str(parameters[\"b2\"]))",
"W1 = [[-0.00643025 0.01936718]\n [-0.02410458 0.03978052]\n [-0.01653973 -0.02096177]\n [ 0.01046864 -0.05990141]]\nb1 = [[ -1.02420756e-06]\n [ 1.27373948e-05]\n [ 8.32996807e-07]\n [ -3.20136836e-06]]\nW2 = [[-0.01041081 -0.04463285 0.01758031 0.04747113]]\nb2 = [[ 0.00010457]]\n"
]
],
[
[
"**Expected Output**:\n\n\n<table style=\"width:80%\">\n <tr>\n <td>**W1**</td>\n <td> [[-0.00643025 0.01936718]\n [-0.02410458 0.03978052]\n [-0.01653973 -0.02096177]\n [ 0.01046864 -0.05990141]]</td> \n </tr>\n \n <tr>\n <td>**b1**</td>\n <td> [[ -1.02420756e-06]\n [ 1.27373948e-05]\n [ 8.32996807e-07]\n [ -3.20136836e-06]]</td> \n </tr>\n \n <tr>\n <td>**W2**</td>\n <td> [[-0.01041081 -0.04463285 0.01758031 0.04747113]] </td> \n </tr>\n \n\n <tr>\n <td>**b2**</td>\n <td> [[ 0.00010457]] </td> \n </tr>\n \n</table> ",
"_____no_output_____"
],
[
"### 4.4 - Integrate parts 4.1, 4.2 and 4.3 in nn_model() ####\n\n**Question**: Build your neural network model in `nn_model()`.\n\n**Instructions**: The neural network model has to use the previous functions in the right order.",
"_____no_output_____"
]
],
[
[
"# GRADED FUNCTION: nn_model\n\ndef nn_model(X, Y, n_h, num_iterations = 10000, print_cost=False):\n \"\"\"\n Arguments:\n X -- dataset of shape (2, number of examples)\n Y -- labels of shape (1, number of examples)\n n_h -- size of the hidden layer\n num_iterations -- Number of iterations in gradient descent loop\n print_cost -- if True, print the cost every 1000 iterations\n \n Returns:\n parameters -- parameters learnt by the model. They can then be used to predict.\n \"\"\"\n \n np.random.seed(3)\n n_x = layer_sizes(X, Y)[0]\n n_y = layer_sizes(X, Y)[2]\n \n # Initialize parameters\n ### START CODE HERE ### (≈ 1 line of code)\n parameters = initialize_parameters(n_x, n_h, n_y)\n ### END CODE HERE ###\n \n # Loop (gradient descent)\n\n for i in range(0, num_iterations):\n \n ### START CODE HERE ### (≈ 4 lines of code)\n # Forward propagation. Inputs: \"X, parameters\". Outputs: \"A2, cache\".\n A2, cache = forward_propagation(X, parameters)\n \n # Cost function. Inputs: \"A2, Y, parameters\". Outputs: \"cost\".\n cost = compute_cost(A2, Y, parameters)\n \n # Backpropagation. Inputs: \"parameters, cache, X, Y\". Outputs: \"grads\".\n grads = backward_propagation(parameters, cache, X, Y)\n \n # Gradient descent parameter update. Inputs: \"parameters, grads\". Outputs: \"parameters\".\n parameters = update_parameters(parameters, grads, learning_rate = 1.2)\n \n ### END CODE HERE ###\n \n # Print the cost every 1000 iterations\n if print_cost and i % 1000 == 0:\n print (\"Cost after iteration %i: %f\" %(i, cost))\n\n return parameters",
"_____no_output_____"
],
[
"X_assess, Y_assess = nn_model_test_case()\nparameters = nn_model(X_assess, Y_assess, 4, num_iterations=10000, print_cost=True)\nprint(\"W1 = \" + str(parameters[\"W1\"]))\nprint(\"b1 = \" + str(parameters[\"b1\"]))\nprint(\"W2 = \" + str(parameters[\"W2\"]))\nprint(\"b2 = \" + str(parameters[\"b2\"]))",
"Cost after iteration 0: 1.385354\nCost after iteration 1000: 0.000317\nCost after iteration 2000: 0.000158\nCost after iteration 3000: 0.000105\nCost after iteration 4000: 0.000079\nCost after iteration 5000: 0.000063\nCost after iteration 6000: 0.000053\nCost after iteration 7000: 0.000045\nCost after iteration 8000: 0.000040\nCost after iteration 9000: 0.000035\nW1 = [[-0.65848169 1.21866811]\n [-0.76204273 1.39377573]\n [ 0.5792005 -1.10397703]\n [ 0.76773391 -1.41477129]]\nb1 = [[ 0.287592 ]\n [ 0.3511264 ]\n [-0.2431246 ]\n [-0.35772805]]\nW2 = [[-2.45566237 -3.27042274 2.00784958 3.36773273]]\nb2 = [[ 0.20459656]]\n"
]
],
[
[
"**Expected Output**:\n\n<table style=\"width:90%\">\n\n<tr> \n <td> \n **cost after iteration 0**\n </td>\n <td> \n 0.692739\n </td>\n</tr>\n\n<tr> \n <td> \n <center> $\\vdots$ </center>\n </td>\n <td> \n <center> $\\vdots$ </center>\n </td>\n</tr>\n\n <tr>\n <td>**W1**</td>\n <td> [[-0.65848169 1.21866811]\n [-0.76204273 1.39377573]\n [ 0.5792005 -1.10397703]\n [ 0.76773391 -1.41477129]]</td> \n </tr>\n \n <tr>\n <td>**b1**</td>\n <td> [[ 0.287592 ]\n [ 0.3511264 ]\n [-0.2431246 ]\n [-0.35772805]] </td> \n </tr>\n \n <tr>\n <td>**W2**</td>\n <td> [[-2.45566237 -3.27042274 2.00784958 3.36773273]] </td> \n </tr>\n \n\n <tr>\n <td>**b2**</td>\n <td> [[ 0.20459656]] </td> \n </tr>\n \n</table> ",
"_____no_output_____"
],
[
"### 4.5 Predictions\n\n**Question**: Use your model to predict by building predict().\nUse forward propagation to predict results.\n\n**Reminder**: predictions = $y_{prediction} = \\mathbb 1 \\text{{activation > 0.5}} = \\begin{cases}\n 1 & \\text{if}\\ activation > 0.5 \\\\\n 0 & \\text{otherwise}\n \\end{cases}$ \n \nAs an example, if you would like to set the entries of a matrix X to 0 and 1 based on a threshold you would do: ```X_new = (X > threshold)```",
"_____no_output_____"
]
],
[
[
"# GRADED FUNCTION: predict\n\ndef predict(parameters, X):\n \"\"\"\n Using the learned parameters, predicts a class for each example in X\n \n Arguments:\n parameters -- python dictionary containing your parameters \n X -- input data of size (n_x, m)\n \n Returns\n predictions -- vector of predictions of our model (red: 0 / blue: 1)\n \"\"\"\n \n # Computes probabilities using forward propagation, and classifies to 0/1 using 0.5 as the threshold.\n ### START CODE HERE ### (≈ 2 lines of code)\n A2, cache = forward_propagation(X, parameters)\n predictions=np.zeros(A2.shape)\n for i in range(A2.shape[1]):\n if A2[0,i]<0.5:\n predictions[0,i]=0\n else:\n predictions[0,i]=1\n ## END CODE HERE ###\n \n return predictions",
"_____no_output_____"
],
[
"parameters, X_assess = predict_test_case()\n\npredictions = predict(parameters, X_assess)\nprint(\"predictions mean = \" + str(np.mean(predictions)))",
"predictions mean = 0.666666666667\n"
]
],
[
[
"**Expected Output**: \n\n\n<table style=\"width:40%\">\n <tr>\n <td>**predictions mean**</td>\n <td> 0.666666666667 </td> \n </tr>\n \n</table>",
"_____no_output_____"
],
[
"It is time to run the model and see how it performs on a planar dataset. Run the following code to test your model with a single hidden layer of $n_h$ hidden units.",
"_____no_output_____"
]
],
[
[
"# Build a model with a n_h-dimensional hidden layer\nparameters = nn_model(X, Y, n_h = 4, num_iterations = 10000, print_cost=True)\n\n# Plot the decision boundary\nplot_decision_boundary(lambda x: predict(parameters, x.T), X, Y)\nplt.title(\"Decision Boundary for hidden layer size \" + str(4))",
"Cost after iteration 0: 138.613963\nCost after iteration 1000: 57.643326\nCost after iteration 2000: 50.677062\nCost after iteration 3000: 46.632785\nCost after iteration 4000: 45.276968\nCost after iteration 5000: 44.448380\nCost after iteration 6000: 43.847601\nCost after iteration 7000: 43.384818\nCost after iteration 8000: 39.839553\nCost after iteration 9000: 39.510576\n"
]
],
[
[
"**Expected Output**:\n\n<table style=\"width:40%\">\n <tr>\n <td>**Cost after iteration 9000**</td>\n <td> 0.218607 </td> \n </tr>\n \n</table>\n",
"_____no_output_____"
]
],
[
[
"# Print accuracy\npredictions = predict(parameters, X)\nprint ('Accuracy: %d' % float((np.dot(Y,predictions.T) + np.dot(1-Y,1-predictions.T))/float(Y.size)*100) + '%')",
"Accuracy: 90%\n"
]
],
[
[
"**Expected Output**: \n\n<table style=\"width:15%\">\n <tr>\n <td>**Accuracy**</td>\n <td> 90% </td> \n </tr>\n</table>",
"_____no_output_____"
],
[
"Accuracy is really high compared to Logistic Regression. The model has learnt the leaf patterns of the flower! Neural networks are able to learn even highly non-linear decision boundaries, unlike logistic regression. \n\nNow, let's try out several hidden layer sizes.",
"_____no_output_____"
],
[
"### 4.6 - Tuning hidden layer size (optional/ungraded exercise) ###\n\nRun the following code. It may take 1-2 minutes. You will observe different behaviors of the model for various hidden layer sizes.",
"_____no_output_____"
]
],
[
[
"# This may take about 2 minutes to run\n\nplt.figure(figsize=(16, 32))\nhidden_layer_sizes = [1, 2, 3, 4, 5, 20, 50]\nfor i, n_h in enumerate(hidden_layer_sizes):\n plt.subplot(5, 2, i+1)\n plt.title('Hidden Layer of size %d' % n_h)\n parameters = nn_model(X, Y, n_h, num_iterations = 5000)\n plot_decision_boundary(lambda x: predict(parameters, x.T), X, Y)\n predictions = predict(parameters, X)\n accuracy = float((np.dot(Y,predictions.T) + np.dot(1-Y,1-predictions.T))/float(Y.size)*100)\n print (\"Accuracy for {} hidden units: {} %\".format(n_h, accuracy))",
"Accuracy for 1 hidden units: 67.5 %\nAccuracy for 2 hidden units: 67.25 %\nAccuracy for 3 hidden units: 90.75 %\nAccuracy for 4 hidden units: 90.5 %\nAccuracy for 5 hidden units: 91.25 %\nAccuracy for 20 hidden units: 90.0 %\nAccuracy for 50 hidden units: 90.25 %\n"
]
],
[
[
"**Interpretation**:\n- The larger models (with more hidden units) are able to fit the training set better, until eventually the largest models overfit the data. \n- The best hidden layer size seems to be around n_h = 5. Indeed, a value around here seems to fits the data well without also incurring noticeable overfitting.\n- You will also learn later about regularization, which lets you use very large models (such as n_h = 50) without much overfitting. ",
"_____no_output_____"
],
[
"**Optional questions**:\n\n**Note**: Remember to submit the assignment by clicking the blue \"Submit Assignment\" button at the upper-right. \n\nSome optional/ungraded questions that you can explore if you wish: \n- What happens when you change the tanh activation for a sigmoid activation or a ReLU activation?\n- Play with the learning_rate. What happens?\n- What if we change the dataset? (See part 5 below!)",
"_____no_output_____"
],
[
"<font color='blue'>\n**You've learnt to:**\n- Build a complete neural network with a hidden layer\n- Make a good use of a non-linear unit\n- Implemented forward propagation and backpropagation, and trained a neural network\n- See the impact of varying the hidden layer size, including overfitting.",
"_____no_output_____"
],
[
"Nice work! ",
"_____no_output_____"
],
[
"## 5) Performance on other datasets",
"_____no_output_____"
],
[
"If you want, you can rerun the whole notebook (minus the dataset part) for each of the following datasets.",
"_____no_output_____"
]
],
[
[
"# Datasets\nnoisy_circles, noisy_moons, blobs, gaussian_quantiles, no_structure = load_extra_datasets()\n\ndatasets = {\"noisy_circles\": noisy_circles,\n \"noisy_moons\": noisy_moons,\n \"blobs\": blobs,\n \"gaussian_quantiles\": gaussian_quantiles}\n\n### START CODE HERE ### (choose your dataset)\ndataset = \"noisy_moons\"\n### END CODE HERE ###\n\nX, Y = datasets[dataset]\nX, Y = X.T, Y.reshape(1, Y.shape[0])\n\n# make blobs binary\nif dataset == \"blobs\":\n Y = Y%2\n\n# Visualize the data\nplt.scatter(X[0, :], X[1, :], c=Y, s=40, cmap=plt.cm.Spectral);",
"_____no_output_____"
]
],
[
[
"Congrats on finishing this Programming Assignment!\n\nReference:\n- http://scs.ryerson.ca/~aharley/neural-networks/\n- http://cs231n.github.io/neural-networks-case-study/",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
4ab743b06320f0e27413ea462714ec00b6f7e527
| 200,782 |
ipynb
|
Jupyter Notebook
|
EDA_Martina.ipynb
|
olehansen538/MPO-Kickstarter
|
678bca98d62e86382d2da474a4920f589c90e915
|
[
"MIT"
] | 1 |
2020-12-15T11:30:15.000Z
|
2020-12-15T11:30:15.000Z
|
EDA_Martina.ipynb
|
olehansen538/MPO-Kickstarter
|
678bca98d62e86382d2da474a4920f589c90e915
|
[
"MIT"
] | 1 |
2020-11-05T09:54:15.000Z
|
2020-11-05T09:54:15.000Z
|
EDA_Martina.ipynb
|
olehansen538/MPO-Kickstarter
|
678bca98d62e86382d2da474a4920f589c90e915
|
[
"MIT"
] | 2 |
2020-10-30T09:33:03.000Z
|
2020-10-30T09:34:21.000Z
| 111.113448 | 22,556 | 0.830189 |
[
[
[
"# EDA classification\n\naim: When will a project succeed?\nWhich features influence the success of a project?\n\n\n#### assumptions\n * the higher the goal, the lower the probability for success\n * the longer the duration the higher the probability for success\n * the longer the preparation time the higher the probability for success\n * the month of launch influences the probability for success\n * the country influences the probability for success\n * pledged amount per backer influences the probability for success",
"_____no_output_____"
]
],
[
[
"# import packages\nimport pandas as pd\nimport numpy as np\nimport seaborn as sns\nimport matplotlib.pyplot as plt",
"_____no_output_____"
],
[
"# read dataframe in\ndf = pd.read_csv('data/kickstarter_preprocess.csv')",
"_____no_output_____"
],
[
"# first summary\ndf.shape",
"_____no_output_____"
],
[
"df.info()",
"<class 'pandas.core.frame.DataFrame'>\nRangeIndex: 177594 entries, 0 to 177593\nData columns (total 26 columns):\n # Column Non-Null Count Dtype \n--- ------ -------------- ----- \n 0 backers_count 177594 non-null int64 \n 1 country 177594 non-null object \n 2 goal 177594 non-null float64\n 3 staff_pick 177594 non-null bool \n 4 state 177594 non-null object \n 5 usd_pledged 177594 non-null int64 \n 6 blurb_len_c 177594 non-null int64 \n 7 blurb_len_w 177594 non-null int64 \n 8 slug_len_c 177594 non-null int64 \n 9 slug_len_w 177594 non-null int64 \n 10 cat_in_slug 177594 non-null int64 \n 11 category_parent_id 177594 non-null int64 \n 12 category_id 177594 non-null int64 \n 13 category_name 177594 non-null object \n 14 created_year 177594 non-null int64 \n 15 created_month 177594 non-null int64 \n 16 deadline_year 177594 non-null int64 \n 17 deadline_month 177594 non-null int64 \n 18 launched_year 177594 non-null int64 \n 19 launched_month 177594 non-null int64 \n 20 duration_days 177594 non-null int64 \n 21 preparation 177594 non-null int64 \n 22 pledged_per_backer 177594 non-null int64 \n 23 rel_pledged_goal 177594 non-null float64\n 24 filled_parent 177594 non-null int64 \n 25 parent_name 177594 non-null object \ndtypes: bool(1), float64(2), int64(19), object(4)\nmemory usage: 34.0+ MB\n"
],
[
"df.describe()",
"_____no_output_____"
],
[
"df.head()",
"_____no_output_____"
],
[
"# overview how many projects were successful, failed, canceled\ndf['state'].hist();",
"_____no_output_____"
],
[
"# make three new dataframes: one for success, one for failed and the last for canceled\ndf_suc = df.query('state == \"successful\"')\ndf_fai = df.query('state == \"failed\"')\ndf_can = df.query('state == \"canceled\"')",
"_____no_output_____"
]
],
[
[
"### assumption 1: the higher the goal, the lower the probability for success",
"_____no_output_____"
]
],
[
[
"plt.boxplot(df_suc['goal'])\nplt.yscale('log');",
"_____no_output_____"
],
[
"df_suc.query('goal < 1').shape",
"_____no_output_____"
],
[
"df_suc.query('goal >= 500000').shape",
"_____no_output_____"
],
[
"# remove outlier\n#df_suc.drop(df_suc[df_suc['goal'] > 100000].index, inplace=True)",
"_____no_output_____"
],
[
"df_suc1 = df_suc.query('goal <= 1500')\n#plt.boxplot(df_suc1['goal'])\ndf_suc1.shape",
"_____no_output_____"
],
[
"df_suc2 = df_suc.query('1500 < goal < 7000')\n#plt.boxplot(df_suc2['goal'])\ndf_suc2.shape",
"_____no_output_____"
],
[
"df_suc3 = df_suc.query('goal >= 7000')\n#plt.boxplot(df_suc3['goal'])\ndf_suc3.shape",
"_____no_output_____"
],
[
"df_fai1 = df_fai.query('goal <= 1500')\ndf_fai1.shape",
"_____no_output_____"
],
[
"df_fai2 = df_fai.query('1500 < goal < 7000')\ndf_fai2.shape",
"_____no_output_____"
],
[
"df_fai3 = df_fai.query('goal >= 7000')\ndf_fai3.shape",
"_____no_output_____"
],
[
"df_can1 = df_can.query('goal <= 1500')\ndf_can1.shape",
"_____no_output_____"
],
[
"df_can2 = df_can.query('1500 < goal < 7000')\ndf_can2.shape",
"_____no_output_____"
],
[
"df_can3 = df_can.query('goal >= 7000')\ndf_can3.shape",
"_____no_output_____"
],
[
"# making a categorical variable for goal 0='goal <= 1500' 1='1500 < goal < 7000', 2='goal >= 7000'\n#df.loc[df['goal'] <= 1500, 'goal_split'] = 0\n#df.loc[(df['goal'] > 1500) & (df['goal'] < 7000), 'goal_split'] = 1\n#df.loc[df['goal'] >= 7000, 'goal_split'] = 2\n#sns.barplot(x='goal_split', y=None, hue=\"state\", data=df)\n\n# set width of bar\nbarWidth = 0.25\nfig = plt.subplots(figsize =(12, 8)) \n\n# set height of bar \nsuc = [29467, 35129, 29650] \nfai = [13763, 22526, 37909] \ncan = [1656, 2502, 4460]\n\n# Set position of bar on X axis \nbr1 = np.arange(3) \nbr2 = [x + barWidth for x in br1] \nbr3 = [x + barWidth for x in br2]\n\np1 = plt.bar(br1, suc, color ='g', width = barWidth, \n edgecolor ='grey', tick_label ='success') \np2 = plt.bar(br2, fai, color ='r', width = barWidth, \n edgecolor ='grey', tick_label ='failed') \np3 = plt.bar(br3, can, color ='b', width = barWidth, \n edgecolor ='grey', tick_label ='canceled')\n\n# Adding Xticks \nplt.xlabel('goal_split', fontweight ='bold') \nplt.ylabel('count', fontweight ='bold') \nplt.xticks([r + barWidth for r in range(3)], \n ['goal <= 1500', '1500 < goal < 7000', 'goal >= 7000'])\nplt.legend((p1[0], p2[0], p3[0]), ('success', 'failed', 'canceled'))\n\nplt.show()\n\n#df1 = df.query('goal <= 1500')\n#df2 = df.query('1500 < goal < 7000')\n#df3 = df.query('goal >= 7000')",
"_____no_output_____"
]
],
[
[
"## conclucsion\nthe lower the goal the higher the probability for success",
"_____no_output_____"
]
],
[
[
"sns.violinplot(x =\"state\", y =\"goal\", data = df_suc);",
"_____no_output_____"
],
[
"var1 = 'state'\ndata1 = pd.concat([df_suc['goal'], df_suc[var1]], axis=1)\nf, ax = plt.subplots(figsize=(8, 6))\nfig1 = sns.violinplot(x=var1, y=\"goal\", data=data1, scale=\"count\")\nfig1.axis(ymin=0, ymax=100000);\n#plt.yscale('log')",
"_____no_output_____"
],
[
"df.query('goal <= 1').shape",
"_____no_output_____"
],
[
"df.query('goal >= 1000000').shape",
"_____no_output_____"
],
[
"df.query('goal >= 1000000 and state == \"successful\"').shape",
"_____no_output_____"
],
[
"sta_dur = df.plot(x='state', \n y='duration_days', \n kind='scatter')",
"_____no_output_____"
]
],
[
[
"### assumption 2: the longer the duration the higher the probability for success",
"_____no_output_____"
]
],
[
[
"plt.boxplot(df['duration_days']);",
"_____no_output_____"
],
[
"dur = [df.query('duration_days <= 20').shape, df.query('20 < duration_days <= 30').shape, \n df.query('30 < duration_days <= 40').shape, df.query('duration_days > 40').shape]\ndur",
"_____no_output_____"
],
[
"dur1 = [df_suc.query('duration_days <= 20').shape, df_suc.query('20 < duration_days <= 30').shape,\n df_suc.query('30 < duration_days <= 40').shape, df_suc.query('duration_days > 40').shape]\ndur1",
"_____no_output_____"
],
[
"dur2 = [df_fai.query('duration_days <= 20').shape, df_fai.query('20 < duration_days <= 30').shape,\n df_fai.query('30 < duration_days <= 40').shape, df_fai.query('duration_days > 40').shape]\ndur2",
"_____no_output_____"
],
[
"dur3 = [df_can.query('duration_days <= 20').shape, df_can.query('20 < duration_days <= 30').shape,\n df_can.query('30 < duration_days <= 40').shape, df_can.query('duration_days > 40').shape]\ndur3",
"_____no_output_____"
],
[
"# set width of bar\nbarWidth = 0.25\nfig = plt.subplots(figsize =(12, 8)) \n\n# set height of bar \nsuc = [12052, 55097, 15513, 12116] \nfai = [6150, 43036, 8119, 16893] \ncan = [653, 4661, 1163, 2141]\n\n# Set position of bar on X axis \nbr1 = np.arange(4) \nbr2 = [x + barWidth for x in br1] \nbr3 = [x + barWidth for x in br2]\n\np1 = plt.bar(br1, suc, color ='g', width = barWidth, \n edgecolor ='grey', tick_label ='success') \np2 = plt.bar(br2, fai, color ='r', width = barWidth, \n edgecolor ='grey', tick_label ='failed') \np3 = plt.bar(br3, can, color ='b', width = barWidth, \n edgecolor ='grey', tick_label ='canceled')\n\n# Adding Xticks \nplt.xlabel('goal_split', fontweight ='bold') \nplt.ylabel('count', fontweight ='bold') \nplt.xticks([r + barWidth for r in range(4)], \n ['duration_days <= 20', '20 < duration_days <= 30', '30 < duration_days <= 40', 'duration_days > 40'])\nplt.legend((p1[0], p2[0], p3[0]), ('success', 'failed', 'canceled'))\n\nplt.show()",
"_____no_output_____"
]
],
[
[
"### assumption 3: the longer the preparation time the higher the probability for success",
"_____no_output_____"
],
[
"### assumption 4: the month of launch influences the probability for success",
"_____no_output_____"
]
],
[
[
"sta_month = df.plot(x='launched_month', \n y='state', \n kind='scatter')",
"_____no_output_____"
],
[
"# boxplot sqft_living (small houses, big houses, bad neighborhood)\nfig, axes = plt.subplots(ncols=3,sharex=True,sharey=True,figsize=(9,6))\n\nax1 = df_suc.boxplot(column=['duration_days'],ax=axes[0])\nax1.set_title(\"duration_days successful\", fontsize = 10)\nax1.set_ylabel(\"count\");\nax2 = df_fai.boxplot(column=['duration_days'],ax=axes[1])\nax2.set_title(\"duration_days failed\", fontsize = 10);\nax3 = df_can.boxplot(column=['duration_days'],ax=axes[2]);\nax3.set_title(\"duration_days canceled\", fontsize = 10);",
"_____no_output_____"
],
[
"df_suc['duration_days'].mean(), df_suc['duration_days'].median()",
"_____no_output_____"
],
[
"df_fai['duration_days'].mean(), df_fai['duration_days'].median()",
"_____no_output_____"
],
[
"df_can['duration_days'].mean(), df_can['duration_days'].median()",
"_____no_output_____"
],
[
"df.groupby('state').count()['successful']",
"_____no_output_____"
]
],
[
[
"### assumption 5: the country influences the probability for success",
"_____no_output_____"
]
],
[
[
"cou_suc = df_suc.groupby(['country'])['country'].count()\ncou_fai = df_fai.groupby(['country'])['country'].count()\ncou_can = df_can.groupby(['country'])['country'].count()\npd.merge(cou_suc, cou_fai, cou_can, on=['country'],suffixes=[' successful', ' failed', ' canceled'])\n#df['country'].unique()",
"_____no_output_____"
],
[
"cou_can",
"_____no_output_____"
],
[
"fig, axes = plt.subplots(ncols=3,sharex=True,sharey=True,figsize=(15,6))\n\nax1 = cou_suc.hist(column=['country'],ax=axes[0])\naxes[0].set_title('country successful')\naxes[0].set_xlabel('country')\naxes[0].set_ylabel('count')\n\n\nax2 = cou_fai.hist(column=['country'],ax=axes[1])\naxes[1].set_title('country failed')\naxes[1].set_xlabel('country')\naxes[1].set_ylabel('count')\n\n\nax3 = cou_can.hist(column=['country'],ax=axes[2])\naxes[2].set_title('country canceled')\naxes[2].set_xlabel('country')\naxes[2].set_ylabel('count');",
"_____no_output_____"
],
[
"cou_suc.plot(kind='bar');",
"_____no_output_____"
],
[
"cou_fai.plot(kind='bar');",
"_____no_output_____"
],
[
"cou_can.plot(kind='bar');",
"_____no_output_____"
],
[
"cou = df_suc.groupby('country')['country'].count()\ncou = list(cou)\n\ncou1 = df['country'].unique()\n\n# Creating plot \nfig = plt.figure(figsize =(10, 7)) \nplt.pie(cou, labels = cou1) \n \n# show plot \nplt.show() ",
"_____no_output_____"
],
[
"\n\ncountry_suc = df_suc.groupby(df_suc['country'])\n#pledged = amt_pledged.sum().sort_values(ascending=0)[0:10]\n\n\nax = country_suc.plot(kind=\"bar\")\nax.set_title(\"Amount by Country\")\nax.set_ylabel(\"Amount\")\nax.set_xlabel(\"Country\")\nvals = ax.get_yticks()",
"_____no_output_____"
]
],
[
[
"### assumption 6: pledged amount per backer influences the probability for success",
"_____no_output_____"
]
],
[
[
"df.groupby('state').pledged_per_backer.mean()",
"_____no_output_____"
],
[
"df_suc.groupby('staff_pick').count()",
"_____no_output_____"
],
[
"df_fai.groupby('staff_pick').count()",
"_____no_output_____"
],
[
"df_can.groupby('staff_pick').count()",
"_____no_output_____"
],
[
"df_suc['cat_in_slug'].hist()",
"_____no_output_____"
],
[
"df_fai['cat_in_slug'].hist()",
"_____no_output_____"
],
[
"df_can['cat_in_slug'].hist()",
"_____no_output_____"
],
[
"sns.catplot(x = \"cat_in_slug\", kind = 'count', hue=\"state\", data=df);\n#sns.barplot(x='cat_in_slug', hue='state', data=df)\n#df.groupby('cat_in_slug').plot(x='state', kind='bar')",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab74e35386115ea4071d3f5f2422037c86c3ef9
| 544,816 |
ipynb
|
Jupyter Notebook
|
playground/AK/initial_test/wkw_test_mSEM_v2.ipynb
|
generem/generem
|
b77387d33d177a3cc79742f873a73b636024fe38
|
[
"MIT"
] | null | null | null |
playground/AK/initial_test/wkw_test_mSEM_v2.ipynb
|
generem/generem
|
b77387d33d177a3cc79742f873a73b636024fe38
|
[
"MIT"
] | null | null | null |
playground/AK/initial_test/wkw_test_mSEM_v2.ipynb
|
generem/generem
|
b77387d33d177a3cc79742f873a73b636024fe38
|
[
"MIT"
] | null | null | null | 317.676968 | 460,508 | 0.90111 |
[
[
[
"%load_ext autoreload\n%autoreload 2\n%matplotlib inline\nimport os\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport wkw\n\nimport torch\nfrom torch.utils import data\nfrom torch.utils.tensorboard import SummaryWriter\n# imports from genem package\nfrom genem.util import viewData,gpu\n# Get the empty gpu\ngpu.get_empty_gpu()",
"The autoreload extension is already loaded. To reload it, use:\n %reload_ext autoreload\nFound 1 process(es) running on GPU\nFound the following processes: \nGPU ID: 0, PID: 257592, Type: b'C', Pname: b'python', Mem: b'15511MiB'\nEmpty GPUs: [1]\nAssigned GPU 1\n"
],
[
"# Log directory\nlog_root = './__logs__/'",
"_____no_output_____"
],
[
"# The mag 1 of this dataset points to mag 8-8-1 in the original dataset\npath_wkw = '/tmpscratch/webknossos/Connectomics_Department/2018-11-13_scMS109_1to7199_v01_l4_06_24_fixed_mag8/color/1/'\n# Open the dataset for reading\ndataset = wkw.Dataset.open(path_wkw)\n# Bounding box taken out of the datasource.json\ndataset_bbox = 19500, 15250, 9, 11250, 11250,7168\n# Read a single plane\nemData = dataset.read([19500, 15250, 9], [302, 302, 7000])",
"_____no_output_____"
],
[
"# View the data\nviewData.ImageSliceViewer3D(emData.squeeze(0))",
"_____no_output_____"
],
[
"# Transform data so that the third dimension would be the first so that each slice would be an individual sample\ncorrectedDimforPytorch = emData.squeeze(0).transpose((2,0,1))\nviewData.ImageSliceViewer3D(correctedDimforPytorch)",
"_____no_output_____"
],
[
"# Normalize data\ndata_floatType = correctedDimforPytorch.astype(np.single)\ndata_normalized = np.asarray((data_floatType - data_floatType.mean())/data_floatType.std())",
"_____no_output_____"
],
[
"data_expanded = np.expand_dims(data_normalized,1)\ndata_expanded.shape,data_expanded.mean(),data_expanded.std()",
"_____no_output_____"
],
[
"viewData.ImageSliceViewer3D(data_normalized)",
"_____no_output_____"
],
[
"torch.from_numpy(data_expanded).shape",
"_____no_output_____"
],
[
"# create pytorch data loader from the numpy array\ndataSet = data.TensorDataset(torch.from_numpy(data_expanded))\ntrainloader = data.DataLoader(dataSet, batch_size=32, shuffle=True, num_workers=2)",
"_____no_output_____"
],
[
"for i,curTensor in enumerate(trainloader):\n print(curTensor[0].shape)",
"torch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([32, 1, 302, 302])\ntorch.Size([24, 1, 302, 302])\n"
],
[
"# Code copied from the autoencoder script\ndef get_conv_pad(input_size, kernel_size, stride):\n padding = np.ceil(((stride-1)*input_size-stride+kernel_size)/2).astype(int)\n return padding\n\ndef data2fig_subplot(inputs, outputs, idx):\n fig, axs = plt.subplots(1, 2, figsize=(16,12))\n input_cpu = inputs[idx].data.cpu()\n img_input = input_cpu.numpy().squeeze()\n axs[0].imshow(img_input, cmap='gray')\n output_cpu = outputs[idx].data.cpu()\n img_output = output_cpu.numpy().squeeze()\n axs[1].imshow(img_output, cmap='gray')\n return fig\n\nclass Encoder_4_sampling_bn(torch.nn.Module):\n \n def __init__(self, input_size, kernel_size, stride, n_fmaps, n_latent):\n super().__init__()\n \n self.input_size = input_size\n self.kernel_size = kernel_size\n self.stride = stride\n self.n_fmaps = n_fmaps\n self.n_latent = n_latent\n \n self.encoding_conv1 = torch.nn.Sequential(\n torch.nn.Conv2d(1, n_fmaps, kernel_size, stride),\n torch.nn.ReLU(),\n torch.nn.MaxPool2d(2))\n \n self.encoding_conv2 = torch.nn.Sequential(\n torch.nn.Conv2d(n_fmaps, n_fmaps*2, kernel_size, stride),\n torch.nn.BatchNorm2d(n_fmaps*2),\n torch.nn.ReLU(),\n torch.nn.MaxPool2d(2))\n \n self.encoding_conv3 = torch.nn.Sequential(\n torch.nn.Conv2d(n_fmaps*2, n_fmaps*4, kernel_size, stride),\n torch.nn.BatchNorm2d(n_fmaps*4),\n torch.nn.ReLU(),\n torch.nn.MaxPool2d(2))\n \n self.encoding_conv4 = torch.nn.Sequential(\n torch.nn.Conv2d(n_fmaps*4, n_fmaps*8, kernel_size, stride),\n torch.nn.BatchNorm2d(n_fmaps*8),\n torch.nn.ReLU(),\n torch.nn.MaxPool2d(2))\n \n self.encoding_fc = torch.nn.Sequential(\n torch.nn.Linear(17**2*n_fmaps*8, n_latent),\n torch.nn.ReLU())\n \n def forward(self, x):\n \n x = self.encoding_conv1(x)\n x = self.encoding_conv2(x)\n x = self.encoding_conv3(x)\n x = self.encoding_conv4(x)\n x = self.encoding_fc(x.reshape((-1, 1, 17**2*n_fmaps*8)))\n \n return x\n\nclass Decoder_4_sampling_bn(torch.nn.Module):\n \n def __init__(self, output_size, kernel_size, stride, n_fmaps, n_latent):\n super().__init__()\n \n self.output_size = output_size\n self.kernel_size = kernel_size\n self.stride = stride\n self.n_fmaps = n_fmaps\n self.n_latent = n_latent\n \n self.decoding_fc = torch.nn.Sequential(\n torch.nn.Linear(n_latent, 17**2*n_fmaps*8),\n torch.nn.ReLU())\n \n self.decoding_convt1 = torch.nn.Sequential(\n torch.nn.Upsample(scale_factor=2, mode='nearest'),\n torch.nn.ConvTranspose2d(n_fmaps*8, n_fmaps*4, kernel_size, stride),\n torch.nn.BatchNorm2d(n_fmaps*4),\n torch.nn.ReLU())\n \n self.decoding_convt2 = torch.nn.Sequential(\n torch.nn.Upsample(scale_factor=2, mode='nearest'),\n torch.nn.ConvTranspose2d(n_fmaps*4, n_fmaps*2, kernel_size, stride),\n torch.nn.BatchNorm2d(n_fmaps*2),\n torch.nn.ReLU())\n\n self.decoding_convt3 = torch.nn.Sequential(\n torch.nn.Upsample(scale_factor=2, mode='nearest'),\n torch.nn.ConvTranspose2d(n_fmaps*2, n_fmaps, kernel_size, stride),\n torch.nn.BatchNorm2d(n_fmaps),\n torch.nn.ReLU())\n \n self.decoding_convt4 = torch.nn.Sequential(\n torch.nn.Upsample(scale_factor=2, mode='nearest'),\n torch.nn.ConvTranspose2d(n_fmaps, 1, kernel_size, stride))\n \n def forward(self, x):\n \n x = self.decoding_fc(x)\n x = self.decoding_convt1(x.reshape((-1, n_fmaps*8, 17, 17)))\n x = self.decoding_convt2(x)\n x = self.decoding_convt3(x)\n x = self.decoding_convt4(x)\n \n return x\n\nclass AE(torch.nn.Module):\n \n def __init__(self, encoder, decoder):\n super().__init__()\n \n self.encoder = encoder\n self.decoder = decoder\n \n def forward(self, x):\n \n x = self.encoder(x)\n x = self.decoder(x)\n \n return x",
"_____no_output_____"
],
[
"input_size=302\noutput_size=input_size\nkernel_size=3\nstride=1\nn_fmaps=8\nn_latent=5000\nnet = AE(\n Encoder_4_sampling_bn(input_size, kernel_size, stride, n_fmaps, n_latent), \n Decoder_4_sampling_bn(output_size, kernel_size, stride, n_fmaps, n_latent))\ncriterion = torch.nn.MSELoss()\noptimizer = torch.optim.SGD(net.parameters(), lr=0.02, momentum=0.9)\n# Copy network to cuda\n#net = net.cuda();",
"_____no_output_____"
],
[
"writer = SummaryWriter(log_root)\nwriter_int = 5\n#epoch_writer_int = 5\nn_epoch = 50000\nit = 0\nfor epoch in range(n_epoch): # loop over the dataset multiple times\n \n running_loss = 0.0\n for i, curData in enumerate(trainloader):\n it += 1\n\n inputs = curData[0]\n #inputs = inputs.cuda();\n labels = inputs\n\n # zero the parameter gradients\n optimizer.zero_grad()\n\n # forward + backward + optimize\n outputs = net(inputs)\n loss = criterion(outputs, labels)\n loss.backward()\n optimizer.step()\n\n # print statistics \n running_loss += loss.item()\n \n if (i+1) % writer_int == 0:\n avg_loss = running_loss / writer_int\n #if (epoch+1) % epoch_writer_int == 0:\n print('it: {} (epoch: {}, batch: {}), running loss: {:0.3f}'.format(it, epoch, i+1, avg_loss))\n\n writer.add_scalar('loss', loss.item(), it)\n writer.add_scalar('avg_loss', avg_loss, it)\n writer.add_figure('inputs', data2fig_subplot(inputs, outputs, 0), it)\n\n running_loss = 0.0",
"it: 5 (epoch: 0, batch: 5), running loss: 0.710\nit: 10 (epoch: 0, batch: 10), running loss: 0.745\nit: 15 (epoch: 0, batch: 15), running loss: 0.726\nit: 20 (epoch: 0, batch: 20), running loss: 0.730\nit: 25 (epoch: 0, batch: 25), running loss: 0.712\nit: 30 (epoch: 0, batch: 30), running loss: 0.752\nit: 35 (epoch: 0, batch: 35), running loss: 0.759\nit: 40 (epoch: 0, batch: 40), running loss: 0.702\nit: 45 (epoch: 0, batch: 45), running loss: 0.740\nit: 50 (epoch: 0, batch: 50), running loss: 0.736\nit: 55 (epoch: 0, batch: 55), running loss: 0.763\nit: 60 (epoch: 0, batch: 60), running loss: 0.750\nit: 65 (epoch: 0, batch: 65), running loss: 0.754\nit: 70 (epoch: 0, batch: 70), running loss: 0.715\nit: 75 (epoch: 0, batch: 75), running loss: 0.738\nit: 80 (epoch: 0, batch: 80), running loss: 0.720\nit: 85 (epoch: 0, batch: 85), running loss: 0.734\nit: 90 (epoch: 0, batch: 90), running loss: 0.745\nit: 95 (epoch: 0, batch: 95), running loss: 0.724\nit: 100 (epoch: 0, batch: 100), running loss: 0.762\nit: 105 (epoch: 0, batch: 105), running loss: 0.724\nit: 110 (epoch: 0, batch: 110), running loss: 0.712\nit: 115 (epoch: 0, batch: 115), running loss: 0.745\nit: 120 (epoch: 0, batch: 120), running loss: 0.717\nit: 125 (epoch: 0, batch: 125), running loss: 0.743\nit: 130 (epoch: 0, batch: 130), running loss: 0.733\nit: 135 (epoch: 0, batch: 135), running loss: 0.685\nit: 140 (epoch: 0, batch: 140), running loss: 0.714\nit: 145 (epoch: 0, batch: 145), running loss: 0.744\nit: 150 (epoch: 0, batch: 150), running loss: 0.728\nit: 155 (epoch: 0, batch: 155), running loss: 0.712\nit: 160 (epoch: 0, batch: 160), running loss: 0.746\nit: 165 (epoch: 0, batch: 165), running loss: 0.715\nit: 170 (epoch: 0, batch: 170), running loss: 0.744\nit: 175 (epoch: 0, batch: 175), running loss: 0.744\nit: 180 (epoch: 0, batch: 180), running loss: 0.722\nit: 185 (epoch: 0, batch: 185), running loss: 0.740\nit: 190 (epoch: 0, batch: 190), running loss: 0.749\nit: 195 (epoch: 0, batch: 195), running loss: 0.747\nit: 200 (epoch: 0, batch: 200), running loss: 0.732\nit: 205 (epoch: 0, batch: 205), running loss: 0.745\nit: 210 (epoch: 0, batch: 210), running loss: 0.708\nit: 215 (epoch: 0, batch: 215), running loss: 0.727\nit: 224 (epoch: 1, batch: 5), running loss: 0.705\nit: 229 (epoch: 1, batch: 10), running loss: 0.716\nit: 234 (epoch: 1, batch: 15), running loss: 0.719\nit: 239 (epoch: 1, batch: 20), running loss: 0.744\nit: 244 (epoch: 1, batch: 25), running loss: 0.733\nit: 249 (epoch: 1, batch: 30), running loss: 0.756\nit: 254 (epoch: 1, batch: 35), running loss: 0.729\nit: 259 (epoch: 1, batch: 40), running loss: 0.746\nit: 264 (epoch: 1, batch: 45), running loss: 0.724\nit: 269 (epoch: 1, batch: 50), running loss: 0.714\nit: 274 (epoch: 1, batch: 55), running loss: 0.726\nit: 279 (epoch: 1, batch: 60), running loss: 0.718\nit: 284 (epoch: 1, batch: 65), running loss: 0.737\nit: 289 (epoch: 1, batch: 70), running loss: 0.733\nit: 294 (epoch: 1, batch: 75), running loss: 0.738\nit: 299 (epoch: 1, batch: 80), running loss: 0.727\nit: 304 (epoch: 1, batch: 85), running loss: 0.710\nit: 309 (epoch: 1, batch: 90), running loss: 0.744\nit: 314 (epoch: 1, batch: 95), running loss: 0.727\nit: 319 (epoch: 1, batch: 100), running loss: 0.745\nit: 324 (epoch: 1, batch: 105), running loss: 0.736\nit: 329 (epoch: 1, batch: 110), running loss: 0.733\nit: 334 (epoch: 1, batch: 115), running loss: 0.761\nit: 339 (epoch: 1, batch: 120), running loss: 0.737\nit: 344 (epoch: 1, batch: 125), running loss: 0.745\nit: 349 (epoch: 1, batch: 130), running loss: 0.723\nit: 354 (epoch: 1, batch: 135), running loss: 0.726\nit: 359 (epoch: 1, batch: 140), running loss: 0.719\nit: 364 (epoch: 1, batch: 145), running loss: 0.710\nit: 369 (epoch: 1, batch: 150), running loss: 0.712\nit: 374 (epoch: 1, batch: 155), running loss: 0.715\nit: 379 (epoch: 1, batch: 160), running loss: 0.740\nit: 384 (epoch: 1, batch: 165), running loss: 0.697\nit: 389 (epoch: 1, batch: 170), running loss: 0.732\nit: 394 (epoch: 1, batch: 175), running loss: 0.713\nit: 399 (epoch: 1, batch: 180), running loss: 0.737\nit: 404 (epoch: 1, batch: 185), running loss: 0.727\nit: 409 (epoch: 1, batch: 190), running loss: 0.749\nit: 414 (epoch: 1, batch: 195), running loss: 0.727\nit: 419 (epoch: 1, batch: 200), running loss: 0.754\nit: 424 (epoch: 1, batch: 205), running loss: 0.724\nit: 429 (epoch: 1, batch: 210), running loss: 0.725\nit: 434 (epoch: 1, batch: 215), running loss: 0.712\nit: 443 (epoch: 2, batch: 5), running loss: 0.731\nit: 448 (epoch: 2, batch: 10), running loss: 0.715\nit: 453 (epoch: 2, batch: 15), running loss: 0.745\nit: 458 (epoch: 2, batch: 20), running loss: 0.743\nit: 463 (epoch: 2, batch: 25), running loss: 0.749\nit: 468 (epoch: 2, batch: 30), running loss: 0.727\nit: 473 (epoch: 2, batch: 35), running loss: 0.728\nit: 478 (epoch: 2, batch: 40), running loss: 0.693\nit: 483 (epoch: 2, batch: 45), running loss: 0.730\nit: 488 (epoch: 2, batch: 50), running loss: 0.733\nit: 493 (epoch: 2, batch: 55), running loss: 0.699\nit: 498 (epoch: 2, batch: 60), running loss: 0.717\nit: 503 (epoch: 2, batch: 65), running loss: 0.721\nit: 508 (epoch: 2, batch: 70), running loss: 0.728\nit: 513 (epoch: 2, batch: 75), running loss: 0.712\nit: 518 (epoch: 2, batch: 80), running loss: 0.761\nit: 523 (epoch: 2, batch: 85), running loss: 0.736\nit: 528 (epoch: 2, batch: 90), running loss: 0.698\nit: 533 (epoch: 2, batch: 95), running loss: 0.728\nit: 538 (epoch: 2, batch: 100), running loss: 0.732\nit: 543 (epoch: 2, batch: 105), running loss: 0.732\nit: 548 (epoch: 2, batch: 110), running loss: 0.749\nit: 553 (epoch: 2, batch: 115), running loss: 0.719\nit: 558 (epoch: 2, batch: 120), running loss: 0.725\nit: 563 (epoch: 2, batch: 125), running loss: 0.721\nit: 568 (epoch: 2, batch: 130), running loss: 0.710\nit: 573 (epoch: 2, batch: 135), running loss: 0.745\nit: 578 (epoch: 2, batch: 140), running loss: 0.712\nit: 583 (epoch: 2, batch: 145), running loss: 0.742\nit: 588 (epoch: 2, batch: 150), running loss: 0.731\nit: 593 (epoch: 2, batch: 155), running loss: 0.731\nit: 598 (epoch: 2, batch: 160), running loss: 0.709\nit: 603 (epoch: 2, batch: 165), running loss: 0.719\nit: 608 (epoch: 2, batch: 170), running loss: 0.738\nit: 613 (epoch: 2, batch: 175), running loss: 0.694\nit: 618 (epoch: 2, batch: 180), running loss: 0.712\nit: 623 (epoch: 2, batch: 185), running loss: 0.699\nit: 628 (epoch: 2, batch: 190), running loss: 0.731\nit: 633 (epoch: 2, batch: 195), running loss: 0.728\nit: 638 (epoch: 2, batch: 200), running loss: 0.742\nit: 643 (epoch: 2, batch: 205), running loss: 0.725\nit: 648 (epoch: 2, batch: 210), running loss: 0.732\nit: 653 (epoch: 2, batch: 215), running loss: 0.753\nit: 662 (epoch: 3, batch: 5), running loss: 0.725\nit: 667 (epoch: 3, batch: 10), running loss: 0.737\nit: 672 (epoch: 3, batch: 15), running loss: 0.729\nit: 677 (epoch: 3, batch: 20), running loss: 0.722\nit: 682 (epoch: 3, batch: 25), running loss: 0.742\nit: 687 (epoch: 3, batch: 30), running loss: 0.707\nit: 692 (epoch: 3, batch: 35), running loss: 0.708\nit: 697 (epoch: 3, batch: 40), running loss: 0.708\nit: 702 (epoch: 3, batch: 45), running loss: 0.730\nit: 707 (epoch: 3, batch: 50), running loss: 0.715\nit: 712 (epoch: 3, batch: 55), running loss: 0.742\nit: 717 (epoch: 3, batch: 60), running loss: 0.723\nit: 722 (epoch: 3, batch: 65), running loss: 0.728\nit: 727 (epoch: 3, batch: 70), running loss: 0.721\nit: 732 (epoch: 3, batch: 75), running loss: 0.734\nit: 737 (epoch: 3, batch: 80), running loss: 0.735\nit: 742 (epoch: 3, batch: 85), running loss: 0.740\nit: 747 (epoch: 3, batch: 90), running loss: 0.715\nit: 752 (epoch: 3, batch: 95), running loss: 0.714\nit: 757 (epoch: 3, batch: 100), running loss: 0.735\nit: 762 (epoch: 3, batch: 105), running loss: 0.708\nit: 767 (epoch: 3, batch: 110), running loss: 0.736\nit: 772 (epoch: 3, batch: 115), running loss: 0.711\nit: 777 (epoch: 3, batch: 120), running loss: 0.766\nit: 782 (epoch: 3, batch: 125), running loss: 0.700\nit: 787 (epoch: 3, batch: 130), running loss: 0.737\nit: 792 (epoch: 3, batch: 135), running loss: 0.718\nit: 797 (epoch: 3, batch: 140), running loss: 0.755\nit: 802 (epoch: 3, batch: 145), running loss: 0.715\nit: 807 (epoch: 3, batch: 150), running loss: 0.728\nit: 812 (epoch: 3, batch: 155), running loss: 0.701\n"
],
[
"print(len(trainloader))",
"_____no_output_____"
],
[
"enumerate(trainloader)",
"_____no_output_____"
],
[
"data2fig_subplot(inputs, outputs, 11);",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab752a203450c0a23b203686ae7b556f9651e39
| 29,309 |
ipynb
|
Jupyter Notebook
|
1_introduction_to_data_science/Assignment 2.ipynb
|
necromuralist/data_scienc
|
5106152e7cbd0f6f5a3c82579477417c146f75b5
|
[
"MIT"
] | null | null | null |
1_introduction_to_data_science/Assignment 2.ipynb
|
necromuralist/data_scienc
|
5106152e7cbd0f6f5a3c82579477417c146f75b5
|
[
"MIT"
] | null | null | null |
1_introduction_to_data_science/Assignment 2.ipynb
|
necromuralist/data_scienc
|
5106152e7cbd0f6f5a3c82579477417c146f75b5
|
[
"MIT"
] | null | null | null | 28.847441 | 508 | 0.474666 |
[
[
[
"---\n\n_You are currently looking at **version 1.2** of this notebook. To download notebooks and datafiles, as well as get help on Jupyter notebooks in the Coursera platform, visit the [Jupyter Notebook FAQ](https://www.coursera.org/learn/python-data-analysis/resources/0dhYG) course resource._\n\n---",
"_____no_output_____"
],
[
"# Assignment 2 - Pandas Introduction",
"_____no_output_____"
],
[
"All questions are weighted the same in this assignment.",
"_____no_output_____"
],
[
"## Part 1",
"_____no_output_____"
],
[
"The following code loads the olympics dataset (olympics.csv), which was derrived from the Wikipedia entry on [All Time Olympic Games Medals](https://en.wikipedia.org/wiki/All-time_Olympic_Games_medal_table), and does some basic data cleaning. \n\nThe columns are organized as # of Summer games, Summer medals, # of Winter games, Winter medals, total # number of games, total # of medals. Use this dataset to answer the questions below.",
"_____no_output_____"
]
],
[
[
"import numpy\nimport pandas as pd\nimport pandas",
"_____no_output_____"
],
[
"df = pd.read_csv('olympics.csv', index_col=0, skiprows=1)\ndata = df",
"_____no_output_____"
],
[
"data.describe()",
"_____no_output_____"
],
[
"data.head()",
"_____no_output_____"
]
],
[
[
"First the unicode is removed from the column names and the numbered column names are changed to their meaning.",
"_____no_output_____"
]
],
[
[
"for col in df.columns:\n if col[:2]=='01':\n df.rename(columns={col:'Gold'+col[4:]}, inplace=True)\n if col[:2]=='02':\n df.rename(columns={col:'Silver'+col[4:]}, inplace=True)\n if col[:2]=='03':\n df.rename(columns={col:'Bronze'+col[4:]}, inplace=True)\n if col[:1]=='№':\n df.rename(columns={col:'#'+col[1:]}, inplace=True)\ndata.head()",
"_____no_output_____"
],
[
"names_ids = df.index.str.split('\\s\\(') # split the index by '('\n\ndf.index = names_ids.str[0] # the [0] element is the country name (new index) \ndf['ID'] = names_ids.str[1].str[:3] # the [1] element is the abbreviation or ID (take first 3 characters from that)\n\ndf = df.drop('Totals')\ndf.head()",
"_____no_output_____"
]
],
[
[
"### Question 0 (Example)\n\nWhat is the first country in df?\n\n*This function should return a Series.*",
"_____no_output_____"
]
],
[
[
"# You should write your whole answer within the function provided. The autograder will call\n# this function and compare the return value against the correct solution value\ndef answer_zero():\n # This function returns the row for Afghanistan, which is a Series object. The assignment\n # question description will tell you the general format the autograder is expecting\n return df.iloc[0]\n\n# You can examine what your function returns by calling it in the cell. If you have questions\n# about the assignment formats, check out the discussion forums for any FAQs\nanswer_zero() ",
"_____no_output_____"
],
[
"data = df\nsummer_gold = \"Gold\"\nwinter_gold = \"Gold.1\"\ntotal_gold = \"Gold.2\"\ntotal_silver = \"Silver.2\"\ntotal_bronze = \"Bronze.2\"",
"_____no_output_____"
]
],
[
[
"### Question 1\nWhich country has won the most gold medals in summer games?\n\n*This function should return a single string value.*",
"_____no_output_____"
]
],
[
[
"def answer_one():\n return data[summer_gold].argmax()\nanswer_one()",
"_____no_output_____"
]
],
[
[
"### Question 2\nWhich country had the biggest difference between their summer and winter gold medal counts?\n\n*This function should return a single string value.*",
"_____no_output_____"
]
],
[
[
"def answer_two():\n return (data[summer_gold] - data[winter_gold]).abs().argmax() \nanswer_two()",
"_____no_output_____"
]
],
[
[
"### Question 3\nWhich country has the biggest difference between their summer gold medal counts and winter gold medal counts relative to their total gold medal count? \n\n$$\\frac{Summer~Gold - Winter~Gold}{Total~Gold}$$\n\nOnly include countries that have won at least 1 gold in both summer and winter.\n\n*This function should return a single string value.*",
"_____no_output_____"
]
],
[
[
"def answer_three():\n elegible = data[(data[summer_gold]>=1) & (data[winter_gold]>=1)]\n ratios = (elegible[summer_gold] - elegible[winter_gold]).abs()/elegible[total_gold]\n return ratios.argmax()\n\nanswer_three()",
"_____no_output_____"
]
],
[
[
"### Question 4\nWrite a function that creates a Series called \"Points\" which is a weighted value where each gold medal (`Gold.2`) counts for 3 points, silver medals (`Silver.2`) for 2 points, and bronze medals (`Bronze.2`) for 1 point. The function should return only the column (a Series object) which you created.\n\n*This function should return a Series named `Points` of length 146*",
"_____no_output_____"
]
],
[
[
"def answer_four():\n \"\"\"\n Creates weighted points based on medals\n * Gold: 3 points\n * Silver: 2 points\n * Bronze: 1 point\n\n Returns\n -------\n\n Series: column of points for each NOC\n \"\"\"\n points = numpy.zeros(len(data))\n points += data[total_gold] * 3\n points += data[total_silver] * 2\n points += data[total_bronze]\n return pandas.Series(points, index=data.index)\n\npoints = answer_four()\nassert points.loc[\"United States\"] == 5684",
"_____no_output_____"
]
],
[
[
"## Part 2\nFor the next set of questions, we will be using census data from the [United States Census Bureau](http://www.census.gov/popest/data/counties/totals/2015/CO-EST2015-alldata.html). Counties are political and geographic subdivisions of states in the United States. This dataset contains population data for counties and states in the US from 2010 to 2015. [See this document](http://www.census.gov/popest/data/counties/totals/2015/files/CO-EST2015-alldata.pdf) for a description of the variable names.\n\nThe census dataset (census.csv) should be loaded as census_df. Answer questions using this as appropriate.\n\n### Question 5\nWhich state has the most counties in it? (hint: consider the sumlevel key carefully! You will need this for future questions too...)\n\n*This function should return a single string value.*",
"_____no_output_____"
]
],
[
[
"census_df = pd.read_csv('census.csv')\ncensus_df.head()\ncensus_data = census_df",
"_____no_output_____"
],
[
"class CensusVariables:\n state_name = \"STNAME\"\n county_name = \"CTYNAME\"\n census_population = \"CENSUS2010POP\"\n region = \"REGION\"\n population_2014 = \"POPESTIMATE2014\"\n population_2015 = \"POPESTIMATE2015\"\n population_estimates = [\"POPESTIMATE2010\",\n \"POPESTIMATE2011\",\n \"POPESTIMATE2012\",\n \"POPESTIMATE2013\",\n population_2014,\n population_2015]\n county_level = 50\n summary_level = \"SUMLEV\"",
"_____no_output_____"
],
[
"counties = census_data[census_data[\n CensusVariables.summary_level]==CensusVariables.county_level]\n# this throws off the numeric index for the argmax method so reset it\ncounties = counties.reset_index()\n\n# but the last question wants the original index\ncounties_original_index = census_data[census_data[\n CensusVariables.summary_level]==CensusVariables.county_level]\n",
"_____no_output_____"
],
[
"def answer_five():\n \"\"\"finds state with the most counties\n\n Returns\n -------\n\n str: name of state with the most counties\n \"\"\"\n return counties.groupby(\n CensusVariables.state_name).count().COUNTY.argmax()\n\nanswer_five()",
"_____no_output_____"
]
],
[
[
"### Question 6\nOnly looking at the three most populous counties for each state, what are the three most populous states (in order of highest population to lowest population)? Use `CENSUS2010POP`.\n\n*This function should return a list of string values.*",
"_____no_output_____"
]
],
[
[
"def answer_six():\n \"\"\"finds three most populous states based on top three counties in each\n\n Returns\n -------\n\n List: top three state-names (highest to lowest)\n \"\"\"\n top_threes = counties.groupby(\n CensusVariables.state_name\n )[CensusVariables.census_population].nlargest(3)\n states = top_threes.groupby(level=0).sum()\n return list(states.nlargest(3).index)\n\nanswer_six()",
"_____no_output_____"
]
],
[
[
"### Question 7\nWhich county has had the largest absolute change in population within the period 2010-2015? (Hint: population values are stored in columns POPESTIMATE2010 through POPESTIMATE2015, you need to consider all six columns.)\n\ne.g. If County Population in the 5 year period is 100, 120, 80, 105, 100, 130, then its largest change in the period would be |130-80| = 50.\n\n*This function should return a single string value.*",
"_____no_output_____"
]
],
[
[
"def answer_seven():\n \"\"\"Find county with largest absolute population variance\n\n Returns\n -------\n\n str: name of the county\n \"\"\"\n return counties.iloc[\n (counties[\n CensusVariables.population_estimates].max(axis=1) -\n counties[\n CensusVariables.population_estimates].min(axis=1)\n ).argmax()][CensusVariables.county_name]\n\nanswer_seven()",
"_____no_output_____"
]
],
[
[
"### Question 8\nIn this datafile, the United States is broken up into four regions using the \"REGION\" column. \n\nCreate a query that finds the counties that belong to regions 1 or 2, whose name starts with 'Washington', and whose POPESTIMATE2015 was greater than their POPESTIMATE 2014.\n\n*This function should return a 5x2 DataFrame with the columns = ['STNAME', 'CTYNAME'] and the same index ID as the census_df (sorted ascending by index).*",
"_____no_output_____"
]
],
[
[
"def answer_eight():\n \"\"\"find region 1 or 2 counties:\n\n ,* with names that start with Washington\n ,* whose population grew from 2014 to 2015\n\n .. note:: the index in the final data-frame has to match the original\n census data\n\n Returns\n -------\n\n DataFrame: with the county and state-name columns\n \"\"\"\n regions = counties_original_index[\n (counties_original_index[CensusVariables.region]==1) |\n (counties_original_index[CensusVariables.region]==2)]\n washingtons = regions[\n regions[CensusVariables.county_name].str.startswith(\"Washington\")]\n grew = washingtons[washingtons[CensusVariables.population_2015] >\n washingtons[CensusVariables.population_2014]]\n return grew[[CensusVariables.state_name,\n CensusVariables.county_name]]\n\noutcome = answer_eight()\nassert outcome.shape == (5,2)\nprint(outcome)",
" STNAME CTYNAME\n896 Iowa Washington County\n1419 Minnesota Washington County\n2345 Pennsylvania Washington County\n2355 Rhode Island Washington County\n3163 Wisconsin Washington County\n"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ab765845918da0ddb802944bd2048f6f5b33fe7
| 12,579 |
ipynb
|
Jupyter Notebook
|
Neural Networks/3_Backprop.ipynb
|
sjchiass/ml_guides
|
2f437ce6e0117a140f4a5e43bb10b51ba9e7cb63
|
[
"BSD-3-Clause"
] | null | null | null |
Neural Networks/3_Backprop.ipynb
|
sjchiass/ml_guides
|
2f437ce6e0117a140f4a5e43bb10b51ba9e7cb63
|
[
"BSD-3-Clause"
] | null | null | null |
Neural Networks/3_Backprop.ipynb
|
sjchiass/ml_guides
|
2f437ce6e0117a140f4a5e43bb10b51ba9e7cb63
|
[
"BSD-3-Clause"
] | null | null | null | 74.875 | 625 | 0.675332 |
[
[
[
"# Neural Networks: Back-propagation\n\nBackprop is how the network evaluates its performance during feed forward. Combined with gradient descent, back-propagation helps train the network.\n\n\n# Further reading\n\n* [Peter's Notes](http://peterroelants.github.io/posts/neural_network_implementation_part01/) are a bit mathy and specific, but I've found them helpful when confused\n* [Deep Learning Basics](http://alexminnaar.com/deep-learning-basics-neural-networks-backpropagation-and-stochastic-gradient-descent.html), a guide that covers about the same ground as this one\n* [A Step by Step Backpropagation Example](https://mattmazur.com/2015/03/17/a-step-by-step-backpropagation-example/) using actual numbers\n* [3Blue1Brown's calculus videos.](https://www.youtube.com/playlist?list=PLZHQObOWTQDMsr9K-rj53DwVRMYO3t5Yr) If you want to go deeper into calculus, these are good to get you motivated.\n* [Again, 3Blue2Brown's neural network videos might be useful.](https://www.youtube.com/playlist?list=PLZHQObOWTQDNU6R1_67000Dx_ZCJB-3pi)\n\n## Intuition\n\nIf the feed-forward is data pushed all the way forward to the outputs, then back-propagation is the trickling back down of errors from the outputs all the way back to the earliest neurons.\n\nBack-propagation is necessary if you want to use gradient descent on neural networks of multiple layers. Single-layer models can calculate their errors in one step, but multi-layer networks require a multi-step process to get it done.\n\n\n\nThe back-propagation journey all starts at the output. Here there is a clear link between the choice of parameters (weights and biases) and the output error. The approach here is the same as simple gradient descent.\n\nAt the layer preceeding the output, we'll call it $l_2$, there is an extra step. What is the link between $l_2$ weights and biases and the output error? It has multiple steps: $l_2$ has a direct effect on the output layer's data, and the output layer's data has a direct effect on what the model decides to output. It takes two steps to get back to the end.\n\nIn other words, the output layer is the boss and it is directly responsible for the model's error. If the output layer changes its behaviour, it can directly improve its accuracy. It's the easiest to train.\n\nThe hidden layers are not directly responsible for the model's error; however, they are responsible for providing the output layer accurate analyses of the model's input data. Knowing their boss, they have an idea of how to change their computations so that the big cheese makes more informed decisions. Their gradient formulas in fact depend on the output layer's weights (the boss's personality, you might say).\n\n## Detour: gradient checking\n\nBackprop takes snapshots of errors everywhere in the NN and uses these to adjust parameters. Normally this is done with calculus and repeated applications of the chain rule of differentiation.\n\nBackprop can also be done by more primitive methods, albeit much more slowly. Numerical differentiation is used to teach students calculus, so it makes sense to show it here first before breaking out the chain rule.\n\nThe idea behind numerical differentiation is this: \n\n1. Take your NN as is\n2. Adjust a parameter slightly and see the effect on output error\n3. You now know the effect of that parameter on error\n\n\n\nGiven $J(\\theta_{i,j})$ your cost function, $\\theta$ any parameter anywhere in the neural network, and $\\epsilon$ a small value as a \"nudge\", \n\n$$\\frac{\\partial J(\\theta)}{\\partial \\theta_{i,j}} \\approx \\frac{J(\\theta_{i,j} + \\epsilon) - J(\\theta_{i,j} - \\epsilon)}{2\\epsilon}$$\n\nBackprop does this for every parameter in the NN. If it sees that the output error increases when a parameter is increased, it will decrease the parameter. If output error increases when the parameter is decreased, backprop increases the parameter instead.\n\n(This is the gradient descent algorithm: it sees error and rolls down the slope in the opposite direction.)\n\nUltimately, this makes a neural network more complicated than any collection of corporate committees. Except in rare prophetic instances, an office worker will not know how many dollars their actions win/lose their company. With neural networks, a single neuron will know how much error it is causing its network. And yet, it's never guaranteed that the neuron can do something useful with this information!\n\nIt's a good idea to use gradient checking. It's a good backup.\n\n## Back propagation II\n\nI hope the above has made back propagation make some sense. It's now time for some light mathematics. Don't worry I'll just paste the answers and skip the algebraic Tetris.\n\nHere are the gradients with respect to error for parameters in the NN model.\n\nGlossary:\n* An error gradient: the \"slope\" of the error. All the model needs to know is which direction this is.\n* The Jacobian: a matrix full of gradients. Since all of our parameters are stored in matrices, it makes sense that we'd store all of our gradients in matrices too.\n\n**Note 1:** With these equations, the most important part is whether they're positive or negative, so you can look at them to see what affects their sign. Gradient decsent will generally work alright as long as it's heading in the right direction (has the right sign).\n\n**Note 2:** When a gradient is positive (error is increasing with parameter) you want to decrease the parameter. When the gradient is negative, you want to increase the parameter.\n\nThe gradients for the output layer weights and biases are\n\n$$\\frac{\\partial J(\\theta)}{\\partial B_{out}} = 1 \\cdot (a_{out} - y), \\frac{\\partial J(\\theta)}{\\partial W_{out}} = a_2^T \\cdot (a_{out} - y)$$\n\nThe above equations make some sense. If the output neuron is overshooting the target, reduce the bias. It's a similar idea with the weights: if the weights cause the neuron to overshoot when they are given a positive input, they need to be reduced.\n\n(You need the 1 in the bias gradient. It represents the intercept but is also necessary to get the right dimension.)\n\nTo proceed lower into the previous layer, we have to do some backprop. Here it is:\n\n$$\\delta_{out} = (a_{out} - y) \\cdot W_{out}^T$$\n\nWe also need the derivative of the sigmoid function. We'll just call it $\\sigma^\\prime$.\n\nWe just have to include that in our equations and we'll be fine. The gradients for the second hidden layer are:\n\n$$\\frac{\\partial J(\\theta)}{\\partial W_2} = a_1^T \\cdot (\\sigma^\\prime(a_2) \\circ \\delta_{out}), \\frac{\\partial J(\\theta)}{\\partial B_2} = 1 \\cdot \\sigma^\\prime(a_2) \\circ \\delta_{out}$$\n\nThe sigmoid derivative $\\sigma^\\prime$ is a newcomer, but otherwise these are similar to before. The weight gradients depend on layer 2's input, which comes from layer 1. The bias gradient is simpler, but it still has to pass through the $\\sigma^\\prime$ and the $\\delta_{out}$.\n\nFor layer 1 we need a new delta.\n\n$$\\delta_2 = (\\delta_{out} \\circ \\sigma^\\prime(a_2)) \\cdot W_2^T$$\n\nFinally, the last backprop step.\n\n$$\\frac{\\partial J(\\theta)}{\\partial W_1} = x^T \\cdot (\\sigma^\\prime(a_1) \\circ \\delta_2), \\frac{\\partial J(\\theta)}{\\partial B_1} = 1 \\cdot \\sigma^\\prime(a_1) \\circ \\delta_2$$\n\nThat's all there is to it.\n\n## Gradient interpretation\n\nIn this section, I'll do my best to narrate what back propagation is doing. Feel free to skip this section.\n\nBack propagation is a repeated application of the chain-rule of differentiation, and its purpose is to determine the effect of a parameter on model error, which is the error gradient with respect to the parameter ($W_1$, $B_{out}$, etc).\n\nLet's take the weight update below as an example, starting with the $a_1^T$ term.\n\n$$\\frac{\\partial J(\\theta)}{\\partial W_2} = a_1^T \\cdot (\\sigma^\\prime(a_2) \\circ \\delta_{out})$$\n\nRecall that $W_2$'s role in the neural network is to do the following:\n\n$$a_2 = \\sigma(a_1 W_2 + B_2), z_2 = a_1 W_2 + B_2$$\n\nLet's put everything together by answering the question: *how does $W_2$ affect $z_2$?*\n\nThe answer: *$W_2$ affects $z_2$ through its interaction with $a_1$.*\n\nThe derivative $\\frac{\\partial z_2}{\\partial W_2} = \\frac{\\partial a_1 W_2 + B_2}{\\partial W_2} = a_1$ signifies that $z_2$ increases by $a_1$ when $W_2$ increases by 1. This works out nicely here since $a_1 W_2$ is linear; normally though the derivative \"slope\" only holds in a very small area around the current point.\n\nThis is where the $a_1^T$ in the gradient comes from. What does it mean? It means that $W_2$'s job is to multiply $a_1$, so its contribution to the model output is $a_1$. Since model error is closely related to model output (through the cost function), $a_1$ is also $W_2$'s contribution to model error.\n\nThat covers $a_1^T$.\n\nLet's now look at that $\\sigma^\\prime(a_2)$ term.\n\n$\\sigma^\\prime(a_2)$ is the sigmoid's contribution to error. $W_2$'s contribution to error, seen above, passes through the simgoid prime. The derivative of the sigmoid function is $\\sigma(a_2)^\\prime = \\sigma(a_2) (1 - \\sigma(a_2))$. Looking at it a bit, it becomes apparent that the derivative reaches its maximum value when $\\sigma(a_2) = 1 - \\sigma(a_2)$, or when $\\sigma(a_2) = 0.5$ or $a_2 = 0$. The slope of the derivative vanishes when $a_2$ approaches 0 or 1. Thus the sigmoid's contribution to error: it restricts the flow of error depending on the value of $a_2$ fed to it, $\\sigma^\\prime(a_2)$.\n\nStory so far: $\\frac{\\partial J(\\theta)}{\\partial W_2}$ is $a_1$ passed through $\\sigma^\\prime$, the latter at most being 0.5 but possibly 0.0.\n\nThe next component of $W_2$'s gradient is $\\delta_{out} = (a_{out} - y) \\cdot W_{out}^T$.\n\nWe are seeing $W_{out}$ here because $W_2$'s effect must pass through it to reach the model error. The idea here is that stronger $W_{out}$ values mean that whatever $x W$ outputs will be mangnified, while weaker $W_{out}$ will attenuate $W_2$'s influence. Therefore $W_{out}$ is a part of $W_2$'s effect on error.\n\n$(a_{out} - y)$ is more difficult to explain because it is an algebraic simplication. Its full form is $\\frac{\\partial NLL_{cost} \\sigma(a_{out})}{\\partial a_{out}}$. But that isn't too important. The first role of $a_{out} - y$ is to keep the gradient positive if $a_{out} > y$ but turn it negative if $a_{out} < y$: this makes sense since you want to increase/lower $a_{out}$ if it has undershot/overshot y. The second role of $a_{out} - y$ is to return a higher error value the wider a gap there is between model output and true output: this gap resides in $[-1, 1]$ since all outputs belong in $[0, 1]$.\n\nThere you have it. I hope this has helped you understand gradients a little bit better.",
"_____no_output_____"
]
]
] |
[
"markdown"
] |
[
[
"markdown"
]
] |
4ab76d6e5fb624dde14dc4870c6990a5aa8b4a91
| 15,082 |
ipynb
|
Jupyter Notebook
|
automl/06.auto-ml-sparse-data-custom-cv-split.ipynb
|
msdavx/MachineLearningNotebooks
|
cff9606bf9bb1ac877f63d4f9318c814478ebb20
|
[
"MIT"
] | 1 |
2018-10-12T01:38:23.000Z
|
2018-10-12T01:38:23.000Z
|
automl/06.auto-ml-sparse-data-custom-cv-split.ipynb
|
msdavx/MachineLearningNotebooks
|
cff9606bf9bb1ac877f63d4f9318c814478ebb20
|
[
"MIT"
] | null | null | null |
automl/06.auto-ml-sparse-data-custom-cv-split.ipynb
|
msdavx/MachineLearningNotebooks
|
cff9606bf9bb1ac877f63d4f9318c814478ebb20
|
[
"MIT"
] | 1 |
2018-10-11T21:44:00.000Z
|
2018-10-11T21:44:00.000Z
| 36.08134 | 313 | 0.532688 |
[
[
[
"Copyright (c) Microsoft Corporation. All rights reserved.\n\nLicensed under the MIT License.",
"_____no_output_____"
],
[
"# AutoML 06: Custom CV Splits and Handling Sparse Data\n\nIn this example we use the scikit-learn's [20newsgroup](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.fetch_20newsgroups.html) to showcase how you can use AutoML for handling sparse data and how to specify custom cross validations splits.\n\nMake sure you have executed the [00.configuration](00.configuration.ipynb) before running this notebook.\n\nIn this notebook you will learn how to:\n1. Create an `Experiment` in an existing `Workspace`.\n2. Configure AutoML using `AutoMLConfig`.\n4. Train the model.\n5. Explore the results.\n6. Test the best fitted model.\n\nIn addition this notebook showcases the following features\n- **Custom CV** splits \n- Handling **sparse data** in the input",
"_____no_output_____"
],
[
"## Create an Experiment\n\nAs part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments.",
"_____no_output_____"
]
],
[
[
"import logging\nimport os\nimport random\n\nfrom matplotlib import pyplot as plt\nfrom matplotlib.pyplot import imshow\nimport numpy as np\nimport pandas as pd\nfrom sklearn import datasets\n\nimport azureml.core\nfrom azureml.core.experiment import Experiment\nfrom azureml.core.workspace import Workspace\nfrom azureml.train.automl import AutoMLConfig\nfrom azureml.train.automl.run import AutoMLRun",
"_____no_output_____"
],
[
"ws = Workspace.from_config()\n\n# choose a name for the experiment\nexperiment_name = 'automl-local-missing-data'\n# project folder\nproject_folder = './sample_projects/automl-local-missing-data'\n\nexperiment = Experiment(ws, experiment_name)\n\noutput = {}\noutput['SDK version'] = azureml.core.VERSION\noutput['Subscription ID'] = ws.subscription_id\noutput['Workspace'] = ws.name\noutput['Resource Group'] = ws.resource_group\noutput['Location'] = ws.location\noutput['Project Directory'] = project_folder\noutput['Experiment Name'] = experiment.name\npd.set_option('display.max_colwidth', -1)\npd.DataFrame(data=output, index=['']).T",
"_____no_output_____"
]
],
[
[
"## Diagnostics\n\nOpt-in diagnostics for better experience, quality, and security of future releases.",
"_____no_output_____"
]
],
[
[
"from azureml.telemetry import set_diagnostics_collection\nset_diagnostics_collection(send_diagnostics = True)",
"_____no_output_____"
]
],
[
[
"## Creating Sparse Data",
"_____no_output_____"
]
],
[
[
"from sklearn.datasets import fetch_20newsgroups\nfrom sklearn.feature_extraction.text import HashingVectorizer\nfrom sklearn.model_selection import train_test_split\n\nremove = ('headers', 'footers', 'quotes')\ncategories = [\n 'alt.atheism',\n 'talk.religion.misc',\n 'comp.graphics',\n 'sci.space',\n]\ndata_train = fetch_20newsgroups(subset = 'train', categories = categories,\n shuffle = True, random_state = 42,\n remove = remove)\n\nX_train, X_validation, y_train, y_validation = train_test_split(data_train.data, data_train.target, test_size = 0.33, random_state = 42)\n\n\nvectorizer = HashingVectorizer(stop_words = 'english', alternate_sign = False,\n n_features = 2**16)\nX_train = vectorizer.transform(X_train)\nX_validation = vectorizer.transform(X_validation)\n\nsummary_df = pd.DataFrame(index = ['No of Samples', 'No of Features'])\nsummary_df['Train Set'] = [X_train.shape[0], X_train.shape[1]]\nsummary_df['Validation Set'] = [X_validation.shape[0], X_validation.shape[1]]\nsummary_df",
"_____no_output_____"
]
],
[
[
"## Configure AutoML\n\nInstantiate an `AutoMLConfig` object to specify the settings and data used to run the experiment.\n\n|Property|Description|\n|-|-|\n|**task**|classification or regression|\n|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics: <br><i>accuracy</i><br><i>AUC_weighted</i><br><i>balanced_accuracy</i><br><i>average_precision_score_weighted</i><br><i>precision_score_weighted</i>|\n|**max_time_sec**|Time limit in seconds for each iteration.|\n|**iterations**|Number of iterations. In each iteration AutoML trains a specific pipeline with the data.|\n|**preprocess**|Setting this to *True* enables AutoML to perform preprocessing on the input to handle *missing data*, and to perform some common *feature extraction*.<br>**Note:** If input data is sparse, you cannot use *True*.|\n|**X**|(sparse) array-like, shape = [n_samples, n_features]|\n|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]<br>Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers.|\n|**X_valid**|(sparse) array-like, shape = [n_samples, n_features] for the custom validation set.|\n|**y_valid**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]<br>Multi-class targets. An indicator matrix turns on multilabel classification for the custom validation set.|\n|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder.|",
"_____no_output_____"
]
],
[
[
"automl_config = AutoMLConfig(task = 'classification',\n debug_log = 'automl_errors.log',\n primary_metric = 'AUC_weighted',\n max_time_sec = 3600,\n iterations = 5,\n preprocess = False,\n verbosity = logging.INFO,\n X = X_train, \n y = y_train,\n X_valid = X_validation, \n y_valid = y_validation, \n path = project_folder)",
"_____no_output_____"
]
],
[
[
"## Train the Model\n\nCall the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.\nIn this example, we specify `show_output = True` to print currently running iterations to the console.",
"_____no_output_____"
]
],
[
[
"local_run = experiment.submit(automl_config, show_output=True)",
"_____no_output_____"
]
],
[
[
"## Explore the Results",
"_____no_output_____"
],
[
"#### Widget for Monitoring Runs\n\nThe widget will first report a \"loading\" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.\n\n**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details.",
"_____no_output_____"
]
],
[
[
"from azureml.train.widgets import RunDetails\nRunDetails(local_run).show() ",
"_____no_output_____"
]
],
[
[
"\n#### Retrieve All Child Runs\nYou can also use SDK methods to fetch all the child runs and see individual metrics that we log.",
"_____no_output_____"
]
],
[
[
"children = list(local_run.get_children())\nmetricslist = {}\nfor run in children:\n properties = run.get_properties()\n metrics = {k: v for k, v in run.get_metrics().items() if isinstance(v, float)}\n metricslist[int(properties['iteration'])] = metrics\n \nrundata = pd.DataFrame(metricslist).sort_index(1)\nrundata",
"_____no_output_____"
]
],
[
[
"### Retrieve the Best Model\n\nBelow we select the best pipeline from our iterations. The `get_output` method on `automl_classifier` returns the best run and the fitted model for the last invocation. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*.",
"_____no_output_____"
]
],
[
[
"best_run, fitted_model = local_run.get_output()",
"_____no_output_____"
]
],
[
[
"#### Best Model Based on Any Other Metric\nShow the run and the model which has the smallest `accuracy` value:",
"_____no_output_____"
]
],
[
[
"# lookup_metric = \"accuracy\"\n# best_run, fitted_model = local_run.get_output(metric = lookup_metric)",
"_____no_output_____"
]
],
[
[
"#### Model from a Specific Iteration\nShow the run and the model from the third iteration:",
"_____no_output_____"
]
],
[
[
"# iteration = 3\n# best_run, fitted_model = local_run.get_output(iteration = iteration)",
"_____no_output_____"
]
],
[
[
"### Register the Fitted Model for Deployment",
"_____no_output_____"
]
],
[
[
"description = 'AutoML Model'\ntags = None\nlocal_run.register_model(description = description, tags = tags)\nlocal_run.model_id # Use this id to deploy the model as a web service in Azure.",
"_____no_output_____"
]
],
[
[
"### Testing the Fitted Model",
"_____no_output_____"
]
],
[
[
"# Load test data.\nimport sklearn\nfrom pandas_ml import ConfusionMatrix\n\nremove = ('headers', 'footers', 'quotes')\ncategories = [\n 'alt.atheism',\n 'talk.religion.misc',\n 'comp.graphics',\n 'sci.space',\n]\n\n\ndata_test = fetch_20newsgroups(subset = 'test', categories = categories,\n shuffle = True, random_state = 42,\n remove = remove)\n\nvectorizer = HashingVectorizer(stop_words = 'english', alternate_sign = False,\n n_features = 2**16)\n\nX_test = vectorizer.transform(data_test.data)\ny_test = data_test.target\n\n# Test our best pipeline.\n\ny_pred = fitted_model.predict(X_test)\ny_pred_strings = [data_test.target_names[i] for i in y_pred]\ny_test_strings = [data_test.target_names[i] for i in y_test]\n\ncm = ConfusionMatrix(y_test_strings, y_pred_strings)\nprint(cm)\ncm.plot()",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ab76e2a228c1260a7aba8f7b6ab2a0fd02bb1ce
| 2,146 |
ipynb
|
Jupyter Notebook
|
Projects/Calculator.ipynb
|
NsikakabasiUmoh/FreeCodeCamp
|
368b078c4229dfb7694f49b70bbae959ab2b10c9
|
[
"MIT"
] | null | null | null |
Projects/Calculator.ipynb
|
NsikakabasiUmoh/FreeCodeCamp
|
368b078c4229dfb7694f49b70bbae959ab2b10c9
|
[
"MIT"
] | null | null | null |
Projects/Calculator.ipynb
|
NsikakabasiUmoh/FreeCodeCamp
|
368b078c4229dfb7694f49b70bbae959ab2b10c9
|
[
"MIT"
] | null | null | null | 20.634615 | 59 | 0.491146 |
[
[
[
"# Basic Calculator",
"_____no_output_____"
]
],
[
[
"# get information\nnumber_1 = input(\"Enter a number: \")\nnumber_2 = input(\"Enter another number: \")\n\n# covert to integer and print\nprint(float(number_1) + float(number_2))",
"Enter a number: 4.5\nEnter another number: 3\n7.5\n"
]
],
[
[
"# Advanced Calculator",
"_____no_output_____"
]
],
[
[
"# get input\n\nnumber_1 = float(input(\"Enter first number: \"))\nnumber_2 = float(input(\"Enter second number: \"))\noperator = input(\"Enter operator: \")\n\n# process\nif operator == \"+\":\n print(number_1 + number_2)\nelif operator == \"-\":\n print(number_1 - number_2)\nelif operator == \"*\":\n print(number_1 * number_2)\nelif operator == \"/\":\n print(number_1 / number_2)\nelse:\n print(\"Invalid\")",
"Enter first number: 5\nEnter second number: 6\nEnter operator: /\n0.8333333333333334\n"
]
]
] |
[
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ab7708597c96939a0fd8656ea1957b500f3f4be
| 21,282 |
ipynb
|
Jupyter Notebook
|
Lab6_Classification_service.ipynb
|
YeonKang/Python-for-Machine-Learning
|
84e8ff72155ddef4bf82521e9301c1ca75c213f5
|
[
"Apache-2.0"
] | null | null | null |
Lab6_Classification_service.ipynb
|
YeonKang/Python-for-Machine-Learning
|
84e8ff72155ddef4bf82521e9301c1ca75c213f5
|
[
"Apache-2.0"
] | null | null | null |
Lab6_Classification_service.ipynb
|
YeonKang/Python-for-Machine-Learning
|
84e8ff72155ddef4bf82521e9301c1ca75c213f5
|
[
"Apache-2.0"
] | null | null | null | 40.306818 | 6,937 | 0.574523 |
[
[
[
"<a href=\"https://colab.research.google.com/github/YeonKang/Python-for-Machine-Learning/blob/main/Lab6_Classification_service.ipynb\" target=\"_parent\"><img src=\"https://colab.research.google.com/assets/colab-badge.svg\" alt=\"Open In Colab\"/></a>",
"_____no_output_____"
]
],
[
[
"import pandas as pd\r\nimport numpy as np",
"_____no_output_____"
],
[
"from google.colab import files\r\nmyfile = files.upload()",
"_____no_output_____"
],
[
"import io\r\n\r\ndf = pd.read_csv(io.BytesIO(myfile['binary.csv']))\r\n\r\ndf[:5]",
"_____no_output_____"
],
[
"y_data = df[\"admit\"].values.reshape(-1,1)\r\nx_data = df.iloc[:,1:].values\r\n\r\ny_data[:5], x_data[:5]",
"_____no_output_____"
],
[
"from sklearn import preprocessing #Min-Max Standardzation\r\n\r\nmin_max_scaler = preprocessing.MinMaxScaler()\r\nx_data = min_max_scaler.fit_transform(x_data)\r\n\r\nx_data[:5]",
"_____no_output_____"
],
[
"from sklearn import linear_model, datasets\r\n\r\nlogreg = linear_model.LogisticRegression(fit_intercept=True)\r\nlogreg.fit(x_data, y_data.ravel())\r\n\r\nsum(logreg.predict(x_data) == y_data.ravel()) / len(y_data.ravel())",
"_____no_output_____"
],
[
"theta = np.append(logreg.intercept_, logreg.coef_.ravel())\r\ntheta",
"_____no_output_____"
],
[
"theta_file_name = \"theta_bin.npy\"\r\nnp.save(theta_file_name, theta)",
"_____no_output_____"
],
[
"np.load(theta_file_name)",
"_____no_output_____"
],
[
"min_max = np.vstack( (df.iloc[:,1:].values.min(axis=0), df.iloc[:,1:].values.max(axis=0)) )\r\nmin_max",
"_____no_output_____"
],
[
"min_max_file_name = \"min_max.npy\"\r\nnp.save(min_max_file_name, min_max)",
"_____no_output_____"
],
[
"np.load(min_max_file_name)",
"_____no_output_____"
],
[
"np.load(theta_file_name)",
"_____no_output_____"
],
[
"import pickle\r\n\r\nf = open('logicmodel.pkl', 'wb')\r\nf.seek(0)\r\n\r\npickle.dump(logreg,f )\r\nf.close()",
"_____no_output_____"
],
[
"fr = open('./logicmodel.pkl', 'rb')\r\nlogreg_pickle = pickle.load(fr)\r\n\r\nsum(logreg.predict(x_data) == logreg_pickle.predict(x_data)) / len(y_data.ravel())",
"_____no_output_____"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab772ae42f3654c789b8de44514693e57932e89
| 9,853 |
ipynb
|
Jupyter Notebook
|
math/Math32_Tensor_Product_Solutions.ipynb
|
kanishkmittal/qbronze
|
5b1bcf4e4f48ab35f2e47753e303af2aef24b9b8
|
[
"Apache-2.0",
"CC-BY-4.0"
] | null | null | null |
math/Math32_Tensor_Product_Solutions.ipynb
|
kanishkmittal/qbronze
|
5b1bcf4e4f48ab35f2e47753e303af2aef24b9b8
|
[
"Apache-2.0",
"CC-BY-4.0"
] | null | null | null |
math/Math32_Tensor_Product_Solutions.ipynb
|
kanishkmittal/qbronze
|
5b1bcf4e4f48ab35f2e47753e303af2aef24b9b8
|
[
"Apache-2.0",
"CC-BY-4.0"
] | 1 |
2022-03-05T21:01:01.000Z
|
2022-03-05T21:01:01.000Z
| 29.237389 | 309 | 0.415914 |
[
[
[
"<table> <tr>\n <td style=\"background-color:#ffffff;\">\n <a href=\"http://qworld.lu.lv\" target=\"_blank\"><img src=\"../images/qworld.jpg\" width=\"25%\" align=\"left\"> </a></td>\n <td style=\"background-color:#ffffff;vertical-align:bottom;text-align:right;\">\n prepared by <a href=\"http://abu.lu.lv\" target=\"_blank\">Abuzer Yakaryilmaz</a> (<a href=\"http://qworld.lu.lv/index.php/qlatvia/\" target=\"_blank\">QLatvia</a>)\n </td> \n</tr></table>",
"_____no_output_____"
],
[
"<table width=\"100%\"><tr><td style=\"color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;text-align:right;\">This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros. </td></tr></table>\n$ \\newcommand{\\bra}[1]{\\langle #1|} $\n$ \\newcommand{\\ket}[1]{|#1\\rangle} $\n$ \\newcommand{\\braket}[2]{\\langle #1|#2\\rangle} $\n$ \\newcommand{\\dot}[2]{ #1 \\cdot #2} $\n$ \\newcommand{\\biginner}[2]{\\left\\langle #1,#2\\right\\rangle} $\n$ \\newcommand{\\mymatrix}[2]{\\left( \\begin{array}{#1} #2\\end{array} \\right)} $\n$ \\newcommand{\\myvector}[1]{\\mymatrix{c}{#1}} $\n$ \\newcommand{\\myrvector}[1]{\\mymatrix{r}{#1}} $\n$ \\newcommand{\\mypar}[1]{\\left( #1 \\right)} $\n$ \\newcommand{\\mybigpar}[1]{ \\Big( #1 \\Big)} $\n$ \\newcommand{\\sqrttwo}{\\frac{1}{\\sqrt{2}}} $\n$ \\newcommand{\\dsqrttwo}{\\dfrac{1}{\\sqrt{2}}} $\n$ \\newcommand{\\onehalf}{\\frac{1}{2}} $\n$ \\newcommand{\\donehalf}{\\dfrac{1}{2}} $\n$ \\newcommand{\\hadamard}{ \\mymatrix{rr}{ \\sqrttwo & \\sqrttwo \\\\ \\sqrttwo & -\\sqrttwo }} $\n$ \\newcommand{\\vzero}{\\myvector{1\\\\0}} $\n$ \\newcommand{\\vone}{\\myvector{0\\\\1}} $\n$ \\newcommand{\\vhadamardzero}{\\myvector{ \\sqrttwo \\\\ \\sqrttwo } } $\n$ \\newcommand{\\vhadamardone}{ \\myrvector{ \\sqrttwo \\\\ -\\sqrttwo } } $\n$ \\newcommand{\\myarray}[2]{ \\begin{array}{#1}#2\\end{array}} $\n$ \\newcommand{\\X}{ \\mymatrix{cc}{0 & 1 \\\\ 1 & 0} } $\n$ \\newcommand{\\Z}{ \\mymatrix{rr}{1 & 0 \\\\ 0 & -1} } $\n$ \\newcommand{\\Htwo}{ \\mymatrix{rrrr}{ \\frac{1}{2} & \\frac{1}{2} & \\frac{1}{2} & \\frac{1}{2} \\\\ \\frac{1}{2} & -\\frac{1}{2} & \\frac{1}{2} & -\\frac{1}{2} \\\\ \\frac{1}{2} & \\frac{1}{2} & -\\frac{1}{2} & -\\frac{1}{2} \\\\ \\frac{1}{2} & -\\frac{1}{2} & -\\frac{1}{2} & \\frac{1}{2} } } $\n$ \\newcommand{\\CNOT}{ \\mymatrix{cccc}{1 & 0 & 0 & 0 \\\\ 0 & 1 & 0 & 0 \\\\ 0 & 0 & 0 & 1 \\\\ 0 & 0 & 1 & 0} } $\n$ \\newcommand{\\norm}[1]{ \\left\\lVert #1 \\right\\rVert } $",
"_____no_output_____"
],
[
"<h2> <font color=\"blue\"> Solutions for </font> Matrices: Tensor Product </h2>",
"_____no_output_____"
],
[
"<a id=\"task1\"></a>\n<h3> Task 1 </h3>\n\nFind $ u \\otimes v $ and $ v \\otimes u $ for the given vectors $ u = \\myrvector{-2 \\\\ -1 \\\\ 0 \\\\ 1} $ and $ v = \\myrvector{ 1 \\\\ 2 \\\\ 3 } $.",
"_____no_output_____"
],
[
"<h3>Solution</h3>",
"_____no_output_____"
]
],
[
[
"u = [-2,-1,0,1]\nv = [1,2,3]\n\nuv = []\nvu = []\n\n\nfor i in range(len(u)): # one element of u is picked\n for j in range(len(v)): # now we iteratively select every element of v\n uv.append(u[i]*v[j]) # this one element of u is iteratively multiplied with every element of v \n \nprint(\"u-tensor-v is\",uv) \n\nfor i in range(len(v)): # one element of v is picked\n for j in range(len(u)): # now we iteratively select every element of u\n vu.append(v[i]*u[j]) # this one element of v is iteratively multiplied with every element of u \n \nprint(\"v-tensor-u is\",vu) ",
"u-tensor-v is [-2, -4, -6, -1, -2, -3, 0, 0, 0, 1, 2, 3]\nv-tensor-u is [-2, -1, 0, 1, -4, -2, 0, 2, -6, -3, 0, 3]\n"
]
],
[
[
"<a id=\"task2\"></a>\n<h3> Task 2 </h3>\n\nFind $ A \\otimes B $ for the given matrices\n$\n A = \\mymatrix{rrr}{-1 & 0 & 1 \\\\ -2 & -1 & 2} ~~\\mbox{and}~~ \n B = \\mymatrix{rr}{0 & 2 \\\\ 3 & -1 \\\\ -1 & 1 }.\n$",
"_____no_output_____"
],
[
"<h3>Solution</h3>",
"_____no_output_____"
]
],
[
[
"A = [\n [-1,0,1],\n [-2,-1,2]\n]\n\nB = [\n [0,2],\n [3,-1],\n [-1,1]\n]\n\nprint(\"A =\")\nfor i in range(len(A)):\n print(A[i])\n\nprint() # print a line\nprint(\"B =\")\nfor i in range(len(B)):\n print(B[i])\n\n# let's define A-tensor-B as a (6x6)-dimensional zero matrix\nAB = []\nfor i in range(6):\n AB.append([])\n for j in range(6):\n AB[i].append(0)\n\n \n \n# let's find A-tensor-B\nfor i in range(2):\n for j in range(3):\n # for each A(i,j) we execute the following codes\n a = A[i][j]\n # we access each element of B\n for m in range(3):\n for n in range(2):\n b = B[m][n]\n # now we put (a*b) in the appropriate index of AB\n AB[3*i+m][2*j+n] = a * b\n \n \n\nprint() # print a line\nprint(\"A-tensor-B =\") \nprint() # print a line\nfor i in range(6):\n print(AB[i])",
"A =\n[-1, 0, 1]\n[-2, -1, 2]\n\nB =\n[0, 2]\n[3, -1]\n[-1, 1]\n\nA-tensor-B =\n\n[0, -2, 0, 0, 0, 2]\n[-3, 1, 0, 0, 3, -1]\n[1, -1, 0, 0, -1, 1]\n[0, -4, 0, -2, 0, 4]\n[-6, 2, -3, 1, 6, -2]\n[2, -2, 1, -1, -2, 2]\n"
]
],
[
[
"<a id=\"task3\"></a>\n<h3> Task 3 </h3>\n\nFind $ B \\otimes A $ for the given matrices\n$\n A = \\mymatrix{rrr}{-1 & 0 & 1 \\\\ -2 & -1 & 2} ~~\\mbox{and}~~ \n B = \\mymatrix{rr}{0 & 2 \\\\ 3 & -1 \\\\ -1 & 1 }.\n$",
"_____no_output_____"
],
[
"<h3>Solution</h3>",
"_____no_output_____"
]
],
[
[
"A = [\n [-1,0,1],\n [-2,-1,2]\n]\n\nB = [\n [0,2],\n [3,-1],\n [-1,1]\n]\n\nprint() # print a line\nprint(\"B =\")\nfor i in range(len(B)):\n print(B[i])\n \nprint(\"A =\")\nfor i in range(len(A)):\n print(A[i])\n\n# let's define B-tensor-A as a (6x6)-dimensional zero matrix\nBA = []\nfor i in range(6):\n BA.append([])\n for j in range(6):\n BA[i].append(0)\n \n# let's find B-tensor-A\nfor i in range(3):\n for j in range(2):\n # for each B(i,j) we execute the following codes\n b = B[i][j]\n # we access each element of A\n for m in range(2):\n for n in range(3):\n a = A[m][n]\n # now we put (a*b) in the appropriate index of AB\n BA[2*i+m][3*j+n] = b * a\n \n \n\nprint() # print a line\nprint(\"B-tensor-A =\") \nprint() # print a line\nfor i in range(6):\n print(BA[i])",
"\nB =\n[0, 2]\n[3, -1]\n[-1, 1]\nA =\n[-1, 0, 1]\n[-2, -1, 2]\n\nB-tensor-A =\n\n[0, 0, 0, -2, 0, 2]\n[0, 0, 0, -4, -2, 4]\n[-3, 0, 3, 1, 0, -1]\n[-6, -3, 6, 2, 1, -2]\n[1, 0, -1, -1, 0, 1]\n[2, 1, -2, -2, -1, 2]\n"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
]
] |
4ab776941b5e8338119d5a631bc55bc6ce57b66e
| 66,828 |
ipynb
|
Jupyter Notebook
|
_build/jupyter_execute/curriculum-notebooks/Mathematics/TellingTime/telling-time.ipynb
|
BryceHaley/curriculum-jbook
|
d1246799ddfe62b0cf5c389394a18c2904383437
|
[
"CC-BY-4.0"
] | 1 |
2022-03-18T18:19:40.000Z
|
2022-03-18T18:19:40.000Z
|
_build/jupyter_execute/curriculum-notebooks/Mathematics/TellingTime/telling-time.ipynb
|
callysto/curriculum-jbook
|
ffb685901e266b0ae91d1250bf63e05a87c456d9
|
[
"CC-BY-4.0"
] | null | null | null |
_build/jupyter_execute/curriculum-notebooks/Mathematics/TellingTime/telling-time.ipynb
|
callysto/curriculum-jbook
|
ffb685901e266b0ae91d1250bf63e05a87c456d9
|
[
"CC-BY-4.0"
] | null | null | null | 60.097122 | 19,225 | 0.639717 |
[
[
[
"\n\n<a href=\"https://hub.callysto.ca/jupyter/hub/user-redirect/git-pull?repo=https%3A%2F%2Fgithub.com%2Fcallysto%2Fcurriculum-notebooks&branch=master&subPath=Mathematics/TellingTime/telling-time.ipynb&depth=1\" target=\"_parent\"><img src=\"https://raw.githubusercontent.com/callysto/curriculum-notebooks/master/open-in-callysto-button.svg?sanitize=true\" width=\"123\" height=\"24\" alt=\"Open in Callysto\"/></a>",
"_____no_output_____"
]
],
[
[
"from IPython.display import HTML\nfrom IPython.display import YouTubeVideo\nimport myMagics\n%uiButtons",
"_____no_output_____"
]
],
[
[
"*Note: Run the cell above then click on the \"Initialize\" button to get notebook ready*",
"_____no_output_____"
],
[
"# Telling Time\nTime is a concept we are all very familiar with. Seconds, minutes, hours, and days are simply part of our everyday life. Have you ever wondered about how all these units of time relate or why we use \"am\" and \"pm\" when talking about the hours of the day? \n\nThere are two important distinctions to make when thinking of time:\n 1. Telling the time - looking at a clock and knowing what time it is.\n 2. Measuring the time - using a clock or other tools to measure how long something takes.\n\nIn this notebook we will explore the relationships between all the different units of time and how to use different tools to read and keep track of time. We will also learn how to easily convert between hours, minutes and seconds.",
"_____no_output_____"
],
[
"## A Little History\nWhen thinking of the different times of the day one thing stands out: there are two of each hour in the day; for example, 2 AM and 2 PM. Why is that?\n\nThis dates back to the Roman times. It was decided a day should be split up into two parts, one part for day time and the other for night time.\n\nEventually this split was made to use noon and midnight as the points where it changes from one part of the day to the other\n\n- AM means \"ante meridiem\" which means \"before midday\"\n\n- PM means \"post meridiem\" which means \"after midday\"\n\nClick on the \"More\" button for more details",
"_____no_output_____"
],
[
"<div class=\"hideMe\">\nInitially to do this split the Romans decided to break the day up into two 12 hour blocks. As we can imagine from back in those days it would only be logical to make the 12 hours in the \"day\" start at sunrise and the 12 hours of the \"night\" start at dusk. But since the day/night cycle changes over the year (shorter days in winter for example), this caused problems.<br>\n\nEventually it was decided to change from sunset/dusk to midnight/midday, this is where AM and PM where born. AM means \"ante meridiem\" which stands for \"before midday\" and PM means \"post meridiem\" meaning, you guessed it, \"after midday\". When you think about it this makes sense: 3PM is 3 hours past the midday. Eventually it was decided that keeping one day split up into 24 hours instead of two blocks of 12 hours made more sense. The 24 hour clock was then introduced. The hours on this clock range from 0 to 23 (totalling 24 hours). We in North America still frequently use the AM/PM 12 hour day format but many parts of the world use the 24 hour clock.",
"_____no_output_____"
]
],
[
[
"%toggleMore",
"_____no_output_____"
]
],
[
[
"## How Well do you Know Time?\nOk so now that we have a little background on how our measurements of time came about, let's play a little game. Do you think you can guess exactly 15 seconds without using a clock? If you think you can do it, click the button below. When you think it’s been 15 seconds click it again and see how close you are. (Click again if you want to retry)",
"_____no_output_____"
]
],
[
[
"HTML(filename='TimeGuessWidget.html')",
"_____no_output_____"
]
],
[
[
"So how did you do? Not as easy as it seems eh. Most people, when trying this the first time, end up clicking much too early. You may have counted up to 15 \"Mississippi’s\" in your head which can help get closer to 15 seconds, but what if I asked you to guess 3 minutes? Rhythmically counting to 180 \"Mississippi’s\" is not particularly fun . This example shows the importance of using tools to more accurately measure time.",
"_____no_output_____"
],
[
"## Reading Time\n\nLong ago before electricity was invented, ancient civilizations had to find other cleaver ways of reading time. Two common techniques used where:\n1. Using the position of the sun to know the time of day\n<img src=\"https://upload.wikimedia.org/wikipedia/commons/6/66/Sundial_-_Canonical_Hour.jpg\" width=\"400\" style=\"box-shadow: 4px 4px 12px gray;margin: 5px;\">\n2. Studying the position of the stars was used to know time in the night\n<img src=\"https://c.tadst.com/gfx/750x500/tell-time-with-stars.png?1\" width=\"400\" style=\"box-shadow: 4px 4px 12px gray;margin: 5px;\">\n\nNow days the time is everywhere and is easily accessible. The main two ways of displaying time are using a digital clock or analog. Digital represents the time as numbers, an analog clock represents the time using clock hands going in circle.",
"_____no_output_____"
],
[
"<div class=\"hideMe\">\n\nThese days the time is everywhere. We simply look for a clock on a wall, appliance, a watch, or our phones. This was not always the case though. Does this mean time did not exist long ago? Egyptians 1500 BC (roughly 3500 years ago) were very much aware of time and found very clever ways for measuring it. For these ancient civilizations, as for us, knowing the time and months of the year was crucial to their survival. It turns out the most important clocks of all where not found on earth but in the sky! The stars and sun have been used to measure time for thousands of years and by many civilizations. Using the position of the sun casting shadows was used during the day and the position of known constellations were used at night. \n\nLuckily for us we have evolved far beyond using the sun and stars to tell time. Imagine trying to get to school on time on a cloudy day! Now when we get a new watch or clock we simply synchronize it to match another time piece showing the correct time. More and more, as devices are connected to the internet we don’t even have to set the time, it is done automatically!",
"_____no_output_____"
]
],
[
[
"%toggleMore",
"_____no_output_____"
]
],
[
[
"## Units of Time\nSo if I ask you what time it is right now you could easily look at a clock and tell me right? For example you could go as far as saying \"it is 11:37:12 on Monday December 17, 2018\". Now, that is probably a lot more information then was asked when asking for the time but bear with me. Let's break down all the components of that sentence: \n\n- 1 year is made up of 365 days (366 if the year is leap)\n- 1 day is made up of 24 hours\n- 1 hour is made up of 60 minutes\n- 1 minutes is made up of 60 seconds\n- 1 second is made up of 1000 milliseconds\n\nWe could keep going but already 1 millisecond happens so fast that they are rarely used in everyday life.\n\nLet's visualize this by using an analog clock. If you count all the ticks around the clock you will find out that there are 60 of them. This makes sense, as 1 one hour is made up of 60 minutes and 1 minute is made up of 60 seconds. In everyday life we know a clock only ever goes forward, and some might say it moves relatively slow. This can make it hard to fully understand its pattern. This example breaks these rules and allows you to manipulate the clock forward and backwards, fast or slow. \n\nIf you adjust the slider below the clock you will see that the hands will begin to move. Each tick on the slider represents a second. Try adjusting the time to see how each hand behaves. (You can also use your keyboard's side arrows to tick through 1 second at a time)",
"_____no_output_____"
]
],
[
[
"from IPython.display import HTML\nHTML(filename='ClockWidget.html')",
"_____no_output_____"
]
],
[
[
"What have you noticed about the relationships between the hands as you slide back and forth? \nTwo important things to notice about the clock:\n1. In order for the minute hand (blue) to move one full tick the seconds hand (red) must do one full rotation\n2. When the minute hand does a full rotation the hour hand will have moved 5 ticks\n\nWhy does the hour hand move 5 ticks per minute rotation? That is because a day has 24 hours, not 60 hours. Remember earlier when we talked about AM and PM, the 24 hour day was broken down into two 12 hour sections. So we can see that if we divide a full rotation (60 minutes) into 12 hours we get $$60\\div12=5$$ This means in the time the minute hand does a full rotation, meaning 60 minutes, the hour hand will advance 5 ticks on the clock. You will see this happening if you slide the slider from the far left all the way to the far right, the minute hand will have done a full rotation and the hour hand will have moved 5 ticks. \n\nNow that we have a better understanding of the relationships between the units, can we figure out how many seconds are in 1 hour? Sure we can! Let's think about this. In 1 hour the minute hand goes around 60 times and for each one of these minutes the seconds hand goes around 60 times, this must mean $$60_\\frac{min}{hr} \\times60_\\frac{sec}{min}=3600_\\frac{sec}{hr}$$ So 1 hour has 3600 seconds. This means if you use your keyboard arrows on the slider from left to right you will need to push it 3600 times!! (Don't do that.)\n\nBased on this math can you figure out how many seconds are in a day? or how many minutes are in a week?",
"_____no_output_____"
],
[
"## Measuring Time\nSo as we all know being able to tell the time is a crucial part of our everyday lives. It helps us know when we have appointments, when we should eat or when we should go to sleep. \n\nTime also has many other great uses like keeping time for a hockey game or measuring how long it takes to drive from one city to another. \n\nLet's take travelling from city to another as an example. Say you are going from Calgary to Edmonton and you want to calculate how long the trip takes. The simplest way of doing this without extra tools is to write down the time when you leave and then check the time when you arrive. Now all we do is take the difference between these two times.\n\nLet's say you leave at 1:32 and arrive at 4:47. You can probably estimate in your head that the trip took a little over 3 hours, but we can calculate it exactly. To make this simpler we will convert the format from hours:minutes to just have minutes. Let's recall how many minutes are in 1 hour to get the following: \n$$(1_{hr}\\times60_\\frac{min}{hr})+32_{min}=60_{min} + 32_{min}=92_{min}$$ \n$$(4_{hr}\\times60_\\frac{min}{hr})+47_{min}=240_{min} + 47_{min}=287_{min}$$ \n\n*Notice these times in minutes actually mean 92 min and 287 min past noon respectively*\n\nAnd now we get the difference: \n$$287_{min}-92_{min}=195_{min}$$ \n\nSo the trip took 195 minutes to get from Calgary to Edmonton. To get it back into the hours:minutes format we need to figure out how many times $195$ can be divided by $60$. We can see that $60$ will fit $3$ times inside $195$ and we will be left with a remainder of $15$, so the trip took $3$ hours and $15$ minutes.\n\nOk so that wasn't too bad. It took some work to get an exact value but it is definitely doable. Now let's take our hockey example and look at how we could keep track of the time for the game. A few things to consider before attempting this \n1. The time goes backwards from 20 minutes down to 0 milliseconds\n2. The time has to stop every time the whistle is blown\n3. The time has to be accurate to the 100th millisecond\n\nAnalyzing this problem we can quickly see that if all we have is a regular clock then a hockey game would take a very long time between each whistle blown as someone would have to calculate the differences in time between each stop and start. Thankfully we have many different types of tools to measure times like this. In this case a scoreboard with a timeclock built in does the trick. \n\nNow all the time keeper has to do is stop and start time as the whistle is blown and the play starts again.",
"_____no_output_____"
],
[
"<div class=\"hideMe\">\nIn other sports sometimes a fraction of a second makes the difference between first and second place. Precise measurements of time may also be needed during a critical science experiment. With examples like these we can see that a person's reflex to start/stop a clock is probably not going to cut it. Many other techniques have been developed to overcome these challenges. Laser sensors are far more accurate then the human hand-to-eye coordination. \n<img src=\"https://upload.wikimedia.org/wikipedia/commons/thumb/c/c0/LightBeamTiming.jpg/220px-LightBeamTiming.jpg\" style=\"box-shadow: 4px 4px 12px gray;\">\nThe image above is of a light beam sensor used on the Olympic track.",
"_____no_output_____"
]
],
[
[
"%toggleMore",
"_____no_output_____"
]
],
[
[
"## 24-Hour Clock\nAs we now know a day has 24 hours. We also know why AM and PM were introduced and still used today. Another wildly popular way of representing time is to use the 24-hour clock. The 24-hour clock eliminates the need to us AM and PM. Using the 24-hour clock simply means we don't go back to one after we pass noon, we keep going from 12 to 13. This may seem odd at first since saying it is 13 o'clock is not something we are use to. \n\nOne major benefit to using this format is that you will never set your alarm wrong by putting 8PM instead of 8AM; 8 just means 8 in the morning and 20 means 8 at night. \n\nIf you use this format enough, knowing that 16:03 simply means 4:03pm becomes second nature, but you're probably wondering how to quickly get this answer when you are not used to it yet. \n\nAll you have to do is take the hour $16$ and subtract $12$ from it so $$16-12=4\\text{ o'clock PM.}$$ A good way to quickly do this in your head is to first take away $10$ which is easy to do then remove the last $2$, so $$16-10=6,$$ and then $$6-2=4\\text{ o'clock.}$$ Give this a try: what time is 18:39? How about 22:18?\n\nMany modern watches, smartphones, alarm clocks, etc. allow you to use the 24 hour clock. Try it out for a week or two and see how fast you adjust to this format.",
"_____no_output_____"
],
[
"## Different Ways to Express Time\nNow that we have a much better understanding of how time works and the relationships between the different units we can start getting creative and come up with other ways to express time. Check out this video for an abstract wooden pendulum clock",
"_____no_output_____"
]
],
[
[
"YouTubeVideo('9ZzkMIrWdPE', width=800, height=550)",
"_____no_output_____"
],
[
"%%html\n<iframe width=\"560\" height=\"315\" src=\"https://www.youtube.com/embed/9ZzkMIrWdPE\" frameborder=\"0\" allow=\"accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture\" allowfullscreen></iframe>",
"_____no_output_____"
]
],
[
[
"Here is another abstract way of telling time, can you decipher what each colour represent?\n\nYou can speed up the clock to see how each ring behaves over time, when you think you have figured it out check your answers below.",
"_____no_output_____"
]
],
[
[
"from IPython.display import HTML\nHTML(filename='AbsClockWidget.html')",
"_____no_output_____"
],
[
"from IPython.display import HTML\nHTML(filename='questions.html')",
"_____no_output_____"
]
],
[
[
"## Conclusion\nIn this notebook we explored:\n1. Some history of time and where AM and PM comes from\n2. The relationships between the different units of time and how the behave together\n3. Examples of tools to use the time in different ways (Time for a sports game)\n4. How to use the 24-hour clock",
"_____no_output_____"
],
[
"[](https://github.com/callysto/curriculum-notebooks/blob/master/LICENSE.md)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
]
] |
4ab77843814907d3fd188edab839e87d4ed9298c
| 434,002 |
ipynb
|
Jupyter Notebook
|
examples/navier-stokes/single_instance/q2_ns_fps_resmin.ipynb
|
adityabalu/DiffNet
|
a21e024ad9948fa76fe73796e216a0a6601f2c7c
|
[
"MIT"
] | 1 |
2021-12-02T06:42:38.000Z
|
2021-12-02T06:42:38.000Z
|
examples/navier-stokes/single_instance/q2_ns_fps_resmin.ipynb
|
adityabalu/DiffNet
|
a21e024ad9948fa76fe73796e216a0a6601f2c7c
|
[
"MIT"
] | null | null | null |
examples/navier-stokes/single_instance/q2_ns_fps_resmin.ipynb
|
adityabalu/DiffNet
|
a21e024ad9948fa76fe73796e216a0a6601f2c7c
|
[
"MIT"
] | 2 |
2021-12-01T20:53:24.000Z
|
2021-12-02T06:42:39.000Z
| 707.996737 | 156,676 | 0.947309 |
[
[
[
"import os\nimport sys\nimport math\nimport json\nimport torch\nimport numpy as np\n\nimport scipy.io\nfrom scipy import ndimage\nimport matplotlib\n# from skimage import io\n# matplotlib.use(\"pgf\")\nmatplotlib.rcParams.update({\n # 'font.family': 'serif',\n 'font.size':10,\n})\nfrom matplotlib import pyplot as plt\n\nimport pytorch_lightning as pl\nfrom pytorch_lightning import Trainer, seed_everything\nfrom pytorch_lightning.loggers import TensorBoardLogger\nseed_everything(42)\n\nimport DiffNet\nfrom DiffNet.DiffNetFEM import DiffNet2DFEM\nfrom torch.utils import data\n# from e1_stokes_base_resmin import Stokes2D\n\nfrom pytorch_lightning.callbacks.base import Callback\nfrom e2_ns_fps_resmin import OptimSwitchLBFGS, NS_FPS_Dataset, NS_FPS",
"/work/baskarg/bkhara/python_virtual_envs/lightning/lib/python3.6/site-packages/pandas/compat/__init__.py:120: UserWarning: Could not import the lzma module. Your installed Python is incomplete. Attempting to use lzma compression will result in a RuntimeError.\n warnings.warn(msg)\nGlobal seed set to 42\nGlobal seed set to 42\n"
],
[
"def plot_contours(module, u, v, p, u_x_gp, v_y_gp, path=None):\n self = module\n fig, axs = plt.subplots(3, 3, figsize=(6*3,3*3),\n subplot_kw={'aspect': 'auto'}, squeeze=True)\n \n for i in range(axs.shape[0]-1):\n for j in range(axs.shape[1]):\n axs[i,j].set_xticks([])\n axs[i,j].set_yticks([])\n\n div_gp = u_x_gp + v_y_gp\n div_elmwise = torch.sum(div_gp, 0)\n div_total = torch.sum(div_elmwise)\n \n interp_method = 'bilinear'\n im0 = axs[0,0].imshow(u,cmap='jet', origin='lower', interpolation=interp_method)\n fig.colorbar(im0, ax=axs[0,0]); axs[0,0].set_title(r'$u_x$')\n im1 = axs[0,1].imshow(v,cmap='jet',origin='lower', interpolation=interp_method)\n fig.colorbar(im1, ax=axs[0,1]); axs[0,1].set_title(r'$u_y$')\n im2 = axs[0,2].imshow(p,cmap='jet',origin='lower', interpolation=interp_method)\n fig.colorbar(im2, ax=axs[0,2]); axs[0,2].set_title(r'$p$')\n\n im3 = axs[1,0].imshow(div_elmwise,cmap='jet',origin='lower', interpolation=interp_method)\n fig.colorbar(im3, ax=axs[1,0]); axs[1,0].set_title(r'$\\int(\\nabla\\cdot u) d\\Omega = $' + '{:.3e}'.format(div_total.item()))\n im4 = axs[1,1].imshow((u**2 + v**2)**0.5,cmap='jet',origin='lower', interpolation=interp_method)\n fig.colorbar(im4, ax=axs[1,1]); axs[1,1].set_title(r'$\\sqrt{u_x^2+u_y^2}$')\n x = np.linspace(0, 1, u.shape[1])\n y = np.linspace(0, 1, u.shape[0])\n xx , yy = np.meshgrid(x, y)\n print(x.shape)\n print(y.shape)\n print(xx.shape)\n print(yy.shape)\n print(u.shape)\n print(v.shape)\n im5 = axs[1,2].streamplot(xx, yy, u, v, color='k', cmap='jet'); axs[1,2].set_title(\"Streamlines\")\n\n mid_idxX = int(self.domain_sizeX/2)\n mid_idxY = int(self.domain_sizeY/2)\n# im = axs[2,0].plot(self.dataset.y[:,0], u[:,0],label='u_inlet')\n im = axs[2,0].plot(self.dataset.x[mid_idxY,:], u[mid_idxY,:],label='u_mid')\n im = axs[2,1].plot(self.dataset.x[mid_idxY,:], v[mid_idxY,:],label='v_mid')\n im = axs[2,2].plot(self.dataset.x[mid_idxY,:], p[mid_idxY,:],label='p_mid')\n \n if not path == None:\n plt.savefig(path)\n# im = axs[2,0].plot(self.dataset.y[:,mid_idx], u[:,mid_idx],label='DiffNet')\n# im = axs[2,0].plot(self.midline_Y,self.midline_U,label='Numerical')\n# axs[2,0].set_xlabel('y'); axs[2,0].legend(); axs[2,0].set_title(r'$u_x @ x=0.5$')\n# im = axs[2,1].plot(self.dataset.x[mid_idx,:], v[mid_idx,:],label='DiffNet')\n# im = axs[2,1].plot(self.midline_X,self.midline_V,label='Numerical')\n# axs[2,1].set_xlabel('x'); axs[2,1].legend(); axs[2,1].set_title(r'$u_y @ y=0.5$')\n# im = axs[2,2].plot(self.dataset.x[-1,:], p[-1,:],label='DiffNet')\n# im = axs[2,2].plot(self.midline_X,self.topline_P,label='Numerical')\n# axs[2,2].set_xlabel('x'); axs[2,2].legend(); axs[2,2].set_title(r'$p @ y=1.0$')\n\n# fig.suptitle(\"Re = {:.1f}, N = {}, LR = {:.1e}\".format(self.Re, self.domain_size, self.learning_rate), fontsize=12)\n\n# plt.savefig(os.path.join(self.logger[0].log_dir, 'contour_' + str(self.current_epoch) + '.png'))\n# self.logger[0].experiment.add_figure('Contour Plots', fig, self.current_epoch)\n# plt.close('all')",
"_____no_output_____"
],
[
"lx = 12.\nly = 6.\nNx = 128\nNy = 64\ndomain_size = 32\nRe = 1.\ndir_string = \"ns_fps\"\nmax_epochs = 50001\nplot_frequency = 100\nLR = 5e-3\nopt_switch_epochs = max_epochs\nload_from_prev = False\nload_version_id = 25\n\nx = np.linspace(0, lx, Nx)\ny = np.linspace(0, ly, Ny)\nxx , yy = np.meshgrid(x, y)",
"_____no_output_____"
],
[
"dataset = NS_FPS_Dataset(domain_lengths=(lx,ly), domain_sizes=(Nx,Ny), Re=Re)\n\nif load_from_prev:\n print(\"LOADING FROM PREVIOUS VERSION: \", load_version_id)\n case_dir = './ns_fps/version_'+str(load_version_id)\n net_u = torch.load(os.path.join(case_dir, 'net_u.pt'))\n net_v = torch.load(os.path.join(case_dir, 'net_v.pt'))\n net_p = torch.load(os.path.join(case_dir, 'net_p.pt')) \nelse:\n print(\"INITIALIZING PARAMETERS TO ZERO\")\n v1 = np.zeros_like(dataset.x)\n v2 = np.zeros_like(dataset.x)\n p = np.zeros_like(dataset.x)\n u_tensor = np.expand_dims(np.array([v1,v2,p]),0)\n\n # network = torch.nn.ParameterList([torch.nn.Parameter(torch.FloatTensor(u_tensor), requires_grad=True)])\n net_u = torch.nn.ParameterList([torch.nn.Parameter(torch.FloatTensor(u_tensor[:,0:1,:,:]), requires_grad=True)])\n net_v = torch.nn.ParameterList([torch.nn.Parameter(torch.FloatTensor(u_tensor[:,1:2,:,:]), requires_grad=True)])\n net_p = torch.nn.ParameterList([torch.nn.Parameter(torch.FloatTensor(u_tensor[:,2:3,:,:]), requires_grad=True)])\n# print(\"net_u = \\n\", net_u[0])\n# print(\"net_v = \\n\", net_v[0])\n# print(\"net_p = \\n\", net_p[0])\nnetwork = (net_u, net_v, net_p)\nbasecase = NS_FPS(network, dataset, domain_lengths=(lx,ly), domain_sizes=(Nx,Ny), batch_size=1, fem_basis_deg=1, learning_rate=LR, plot_frequency=plot_frequency)\n\n# Initialize trainer\nlogger = pl.loggers.TensorBoardLogger('.', name=dir_string)\ncsv_logger = pl.loggers.CSVLogger(logger.save_dir, name=logger.name, version=logger.version)\n\nearly_stopping = pl.callbacks.early_stopping.EarlyStopping('loss',\n min_delta=1e-8, patience=10, verbose=False, mode='max', strict=True)\ncheckpoint = pl.callbacks.model_checkpoint.ModelCheckpoint(monitor='loss',\n dirpath=logger.log_dir, filename='{epoch}-{step}',\n mode='min', save_last=True)\n\nlbfgs_switch = OptimSwitchLBFGS(epochs=opt_switch_epochs)\n\ntrainer = Trainer(gpus=[0],callbacks=[early_stopping,lbfgs_switch],\n checkpoint_callback=checkpoint, logger=[logger,csv_logger],\n max_epochs=max_epochs, deterministic=True, profiler=\"simple\")\n\n# Training\ntrainer.fit(basecase)\n# Save network\ntorch.save(basecase.net_u, os.path.join(logger.log_dir, 'net_u.pt'))\ntorch.save(basecase.net_v, os.path.join(logger.log_dir, 'net_v.pt'))\ntorch.save(basecase.net_p, os.path.join(logger.log_dir, 'net_p.pt'))",
"GPU available: True, used: True\nTPU available: False, using: 0 TPU cores\nLOCAL_RANK: 0 - CUDA_VISIBLE_DEVICES: [1]\n"
],
[
"# Query\nbasecase.dataset[0]\ninputs, forcing = basecase.dataset[0]\nu, v, p, u_x, v_y = basecase.do_query(inputs, forcing)\nu = u.squeeze().detach().cpu()\nv = v.squeeze().detach().cpu()\np = p.squeeze().detach().cpu()\nu_x = u_x.squeeze().detach().cpu()\nv_y = v_y.squeeze().detach().cpu()\n\n# plot\nplot_contours(basecase, u, v, p, u_x, v_y)",
"(128,)\n(64,)\n(64, 128)\n(64, 128)\ntorch.Size([64, 128])\ntorch.Size([64, 128])\n"
],
[
"# separate query\nversion_id = 81\ncase_dir = './ns_fps/version_'+str(version_id)\ndataset = NS_FPS_Dataset(domain_lengths=(lx,ly), domain_sizes=(Nx,Ny), Re=Re)\nnet_u = torch.load(os.path.join(case_dir, 'net_u.pt'))\nnet_v = torch.load(os.path.join(case_dir, 'net_v.pt'))\nnet_p = torch.load(os.path.join(case_dir, 'net_p.pt'))\n# network = (net_u, net_v, net_p)\nnetwork = (net_u.cpu(), net_v.cpu(), net_p.cpu())\nequation = NS_FPS(network, dataset, domain_lengths=(lx,ly), domain_sizes=(Nx,Ny), batch_size=1, fem_basis_deg=1, learning_rate=LR, plot_frequency=plot_frequency)\n# Query\ninputs, forcing = equation.dataset[0]\nu, v, p, u_x, v_y = equation.do_query(inputs, forcing)\nu = u.squeeze().detach().cpu()\nv = v.squeeze().detach().cpu()\np = p.squeeze().detach().cpu()\nu_x = u_x.squeeze().detach().cpu()\nv_y = v_y.squeeze().detach().cpu()",
"hx = 0.09448818897637795 , hy = 0.09523809523809523\nnelmX = 127 , nelmY = 63\nexact_solution -- LDC class called\nforcing -- LDC class called\n"
],
[
"obj_left_idx = dataset.obj_left_idx\nobj_rght_idx = dataset.obj_rght_idx\nobj_bttm_idx = dataset.obj_bttm_idx\nobj_top__idx = dataset.obj_top__idx\nu[obj_bttm_idx:obj_top__idx, obj_left_idx:obj_rght_idx] = float('inf')\nv[obj_bttm_idx:obj_top__idx, obj_left_idx:obj_rght_idx] = float('inf')\np[obj_bttm_idx:obj_top__idx, obj_left_idx:obj_rght_idx] = float('inf')\n# plot\nfilepath = os.path.join(case_dir,'query_ns_fps.png')\nplot_contours(equation, u, v, p, u_x, v_y, filepath)",
"(128,)\n(64,)\n(64, 128)\n(64, 128)\ntorch.Size([64, 128])\ntorch.Size([64, 128])\n"
],
[
"net_u.cpu()",
"_____no_output_____"
],
[
"net_u",
"_____no_output_____"
],
[
"simdata = np.loadtxt('ns-ldc-numerical-results/re-30-ns-L12-H6-midlineX.csv', skiprows=1,delimiter=',')",
"_____no_output_____"
],
[
"fig, axs = plt.subplots(3, 3, figsize=(6*3,3.6*3), subplot_kw={'aspect': 'auto'}, squeeze=True)\naxs[0,0].plot(simdata[:,0], simdata[:,2],label='num')\naxs[0,1].plot(simdata[:,0], simdata[:,3],label='num')\naxs[0,2].plot(simdata[:,0], simdata[:,1],label='num')\nmid_idxX = int(Nx/2)\nmid_idxY = int(Ny/2)\naxs[0,0].plot(equation.dataset.x[mid_idxY,:], u[mid_idxY,:],label='u_mid'); axs[0,0].legend()\naxs[0,1].plot(equation.dataset.x[mid_idxY,:], v[mid_idxY,:],label='v_mid'); axs[0,1].legend()\naxs[0,2].plot(equation.dataset.x[mid_idxY,:], p[mid_idxY,:],label='p_mid'); axs[0,2].legend()",
"_____no_output_____"
],
[
"simdataY = np.loadtxt('ns-ldc-numerical-results/re-30-ns-L12-H6-midlineY.csv', skiprows=1,delimiter=',')",
"_____no_output_____"
],
[
"fig, axs = plt.subplots(3, 3, figsize=(6*3,3.6*3), subplot_kw={'aspect': 'auto'}, squeeze=True)\naxs[0,0].plot(simdataY[:,0], simdataY[:,2],label='num')\naxs[0,1].plot(simdataY[:,0], simdataY[:,3],label='num')\naxs[0,2].plot(simdataY[:,0], simdataY[:,1],label='num')\nmid_idxX = int(Nx/2)\nmid_idxY = int(Ny/2)\naxs[0,0].plot(equation.dataset.y[:,mid_idxY], u[:,mid_idxY],label='u_mid'); axs[0,0].legend()\naxs[0,1].plot(equation.dataset.y[:,mid_idxY], v[:,mid_idxY],label='v_mid'); axs[0,1].legend()\naxs[0,2].plot(equation.dataset.y[:,mid_idxY], p[:,mid_idxY],label='p_mid'); axs[0,2].legend()",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab780c091ff06ce2b2c754628900721edc8468c
| 43,169 |
ipynb
|
Jupyter Notebook
|
ONNX/ONNX-Python-Inference-Perf-Issue/TestPerf.ipynb
|
CESARDELATORRE/AML-Issues
|
3aa16de0ff3536e5416bed673a0c9e8c4d91d9d8
|
[
"MIT"
] | null | null | null |
ONNX/ONNX-Python-Inference-Perf-Issue/TestPerf.ipynb
|
CESARDELATORRE/AML-Issues
|
3aa16de0ff3536e5416bed673a0c9e8c4d91d9d8
|
[
"MIT"
] | null | null | null |
ONNX/ONNX-Python-Inference-Perf-Issue/TestPerf.ipynb
|
CESARDELATORRE/AML-Issues
|
3aa16de0ff3536e5416bed673a0c9e8c4d91d9d8
|
[
"MIT"
] | null | null | null | 38.92606 | 102 | 0.312724 |
[
[
[
"import numpy as np\nimport pandas as pd\n\npd.set_option('display.max_columns', None)\n\npd.set_option('display.max_rows', 10)",
"_____no_output_____"
],
[
"import warnings\nimport logging\nimport os\nimport onnxruntime\nfrom azureml.automl.runtime.onnx_convert import OnnxInferenceHelper\n# from azureml.automl.core.onnx_convert import OnnxInferenceHelper\nimport json\nimport time",
"_____no_output_____"
],
[
"from typing import Any, Tuple\nfrom numpy import ndarray\n\n\nclass OnnxModelWrapper:\n \"\"\"\n helper class for prediction when using onnx model\n \"\"\"\n def __init__(self, onnx_model_bytes: bytes, onnx_input_map: dict):\n \"\"\"\n :param onnx_model_bytes: the onnx model in bytes\n :param onnx_input_map: the onnx_resource dictionary\n \"\"\"\n self.onnx_model_bytes = onnx_model_bytes\n self.onnx_input_map = onnx_input_map\n self.wrapper_model = OnnxInferenceHelper(self.onnx_model_bytes, self.onnx_input_map)\n\n def predict(self, X) -> Tuple[Any, Any]:\n \"\"\"\n predict by using OnnxInferenceHelper\n :param X: features to predict\n :returns tuple of <label, prob>\n \"\"\"\n return self.wrapper_model.predict(X)\n\n def predict_proba(self, X) -> ndarray:\n \"\"\"\n predict proba by using OnnxInferenceHelper\n :param X: features to predict\n :returns ndarray of prob\n \"\"\"\n _, y_prob = self.wrapper_model.predict(X, with_prob=True)\n return y_prob\n",
"_____no_output_____"
],
[
"onnx_model_file = open('onnx.model', 'rb')\nonnx_res_file = open('onnx.res', 'r')\nonnx_model_data = onnx_model_file.read()\nonnx_res_data = onnx_res_file.read()",
"_____no_output_____"
],
[
"onnxrt_wrapper = OnnxModelWrapper(onnx_model_data, json.loads(onnx_res_data))",
"_____no_output_____"
],
[
"filepath = 'invoice.csv'\ndata_df = pd.read_csv(filepath)",
"_____no_output_____"
],
[
"data_df.head()",
"_____no_output_____"
],
[
"data_df.shape",
"_____no_output_____"
],
[
"start = time.time()\npredictions = onnxrt_wrapper.predict_proba(data_df)\nend = time.time()\nprint(end - start)",
"31.1851806640625\n"
],
[
"from azureml.explain.model.mimic.mimic_explainer import MimicExplainer\nfrom azureml.explain.model.mimic.models.lightgbm_model import LGBMExplainableModel\nstart = time.time()\nexplainer = MimicExplainer(onnxrt_wrapper, data_df, LGBMExplainableModel, augment_data=False)\nend = time.time()\nprint(end - start)",
"32.04781436920166\n"
],
[
"def explain(train_data):\n explanation = explainer.explain_global(train_data, include_local=False)",
"_____no_output_____"
],
[
"sample = data_df.sample(1000)\nstart = time.time()\nexplain(sample)\nend = time.time()\nprint(end - start)",
"47.51412320137024\n"
],
[
"sample = data_df.sample(2000)\nstart = time.time()\nexplain(sample)\nend = time.time()\nprint(end - start)",
"91.24858832359314\n"
],
[
"sample = data_df.sample(5000)\nstart = time.time()\nexplain(sample)\nend = time.time()\nprint(end - start)",
"218.98222756385803\n"
],
[
"start = time.time()\nexplain(data_df)\nend = time.time()\nprint(end - start)",
"639.5890746116638\n"
],
[
"data_df.shape",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab7866f3308954abb2f501cbaf18ab75a0649a1
| 29,151 |
ipynb
|
Jupyter Notebook
|
notebooks/hpc_project_bechmarks.ipynb
|
qualiphal/parallel-phal
|
a6bbfdb104d13c4c45914e02d53f32e1b134ca3c
|
[
"MIT"
] | null | null | null |
notebooks/hpc_project_bechmarks.ipynb
|
qualiphal/parallel-phal
|
a6bbfdb104d13c4c45914e02d53f32e1b134ca3c
|
[
"MIT"
] | null | null | null |
notebooks/hpc_project_bechmarks.ipynb
|
qualiphal/parallel-phal
|
a6bbfdb104d13c4c45914e02d53f32e1b134ca3c
|
[
"MIT"
] | null | null | null | 44.847692 | 11,290 | 0.638537 |
[
[
[
"from random import randint\nfrom timeit import default_timer\nsize = 100\nmat_1 = [[randint(0, size) for _ in range(size)] for _ in range(size)]\nmat_2 = [[randint(0, size) for _ in range(size)] for _ in range(size)]\nresult = [[0 for _ in range(size)] for _ in range(size)]",
"_____no_output_____"
]
],
[
[
"### 1. Serial Implementation",
"_____no_output_____"
]
],
[
[
"starttime = default_timer()\n\nfor i in range(size):\n for j in range(size):\n for k in range(size):\n result[i][j] += mat_1[i][k] * mat_2[k][j]\n\nt1 = default_timer() - starttime\nprint(\"Serial Time Taken :\", t1)\nresult = [[0 for _ in range(size)] for _ in range(size)]",
"Serial Time Taken : 0.4240078129998892\n"
]
],
[
[
"### 2. Data Parallel Implementation",
"_____no_output_____"
]
],
[
[
"!pip install -q pymp-pypi\nimport pymp\n\nstarttime = default_timer()\nres_arr = pymp.shared.array((size, size), dtype='uint8')\nwith pymp.Parallel(2) as p:\n for i in p.range(size):\n for j in range(size):\n for k in range(size):\n res_arr[i][j] += mat_1[i][k] * mat_2[k][j]\n\nt2 = default_timer() - starttime\nprint(\"Parallel Time Taken :\", t2)",
"Parallel Time Taken : 3.070237331000044\n"
]
],
[
[
"### 3. Numpy",
"_____no_output_____"
]
],
[
[
"import numpy as np \n\nstarttime = default_timer()\n\nres = np.dot(mat_1,mat_2) \n\nt3 = default_timer() - starttime\nprint(\"Numpy Time Taken :\", t3)",
"Numpy Time Taken : 0.004947687999901973\n"
]
],
[
[
"### 4. Scipy",
"_____no_output_____"
]
],
[
[
"from scipy import sparse\n\nm1 = sparse.csr_matrix(mat_1)\nm2 = sparse.csr_matrix(mat_2)\nstarttime = default_timer()\n\nres = m1.multiply(m2)\n\nt4 = default_timer() - starttime\nprint(\"Scipy Time Taken :\", t4)",
"Scipy Time Taken : 0.0029525400000238733\n"
]
],
[
[
"### 5. Pandas",
"_____no_output_____"
]
],
[
[
"import numpy as np\nimport pandas as pd\n\n\ndf_1 = pd.DataFrame(mat_1)\ndf_2 = pd.DataFrame(mat_2)\n\nstarttime = default_timer()\ndf_1.dot(df_2)\n\nt5 = default_timer() - starttime\nprint(\"Pandas Time Taken :\", t5)",
"Pandas Time Taken : 0.0017128819999925327\n"
]
],
[
[
"### 6. Sympy",
"_____no_output_____"
]
],
[
[
"from sympy import Matrix\nm1 = Matrix(mat_1)\nm2 = Matrix(mat_2)\n\nstarttime = default_timer()\nr = m1*m2\n\nt6 = default_timer() - starttime\nprint(\"Sympy Time Taken :\", t6)",
"Sympy Time Taken : 9.038939131000006\n"
]
],
[
[
"### 7. Numba",
"_____no_output_____"
]
],
[
[
"import numpy as np\nimport timeit\nfrom numba import jit, float64, prange\n\n@jit('float64[:,:](float64[:,:],float64[:,:])', parallel=True, nopython=True)\ndef matmul(A, B):\n C = np.zeros((A.shape[0], B.shape[1]))\n for i in prange(A.shape[0]):\n for j in prange(B.shape[1]):\n for k in range(A.shape[0]):\n C[i,j] = C[i,j] + A[i,k]*B[k,j]\n return C\n\nA = np.random.rand(size, size)\nB = np.random.rand(size, size)\n\nstart = default_timer()\n\nmatmul(A, B)\n\nt7 = default_timer() - start\n\nprint(\"Numba Time Taken :\", t7) ",
"/usr/local/lib/python3.7/dist-packages/numba/np/ufunc/parallel.py:363: NumbaWarning: The TBB threading layer requires TBB version 2019.5 or later i.e., TBB_INTERFACE_VERSION >= 11005. Found TBB_INTERFACE_VERSION = 9107. The TBB threading layer is disabled.\n warnings.warn(problem)\n"
]
],
[
[
"### 8. Linalg",
"_____no_output_____"
]
],
[
[
"from numpy.linalg import multi_dot\n\nstart = default_timer()\n\n_ = multi_dot([mat_1, mat_2])\n\nt8 = default_timer() - start\n\nprint(\"linalg Time Taken :\", t8) ",
"linalg Time Taken : 0.004973874999905092\n"
]
],
[
[
"### 9. Pymatrix",
"_____no_output_____"
]
],
[
[
"!wget https://raw.githubusercontent.com/dthul/pymatrix/master/matrix.py\nimport sys\nsys.path.append('/content/')\n\nfrom matrix import Matrix\nm1 = Matrix(mat_1)\nm2 = Matrix(mat_2)\n\nstart = default_timer()\n\nres = m1 * m2\n\nt9 = default_timer() - start\n\nprint(\"pymatrix Time Taken :\", t9) ",
"--2021-04-27 14:43:17-- https://raw.githubusercontent.com/dthul/pymatrix/master/matrix.py\nResolving raw.githubusercontent.com (raw.githubusercontent.com)... 185.199.111.133, 185.199.109.133, 185.199.108.133, ...\nConnecting to raw.githubusercontent.com (raw.githubusercontent.com)|185.199.111.133|:443... connected.\nHTTP request sent, awaiting response... 200 OK\nLength: 16087 (16K) [text/plain]\nSaving to: ‘matrix.py’\n\nmatrix.py 100%[===================>] 15.71K --.-KB/s in 0s \n\n2021-04-27 14:43:17 (111 MB/s) - ‘matrix.py’ saved [16087/16087]\n\npymatrix Time Taken : 0.34896002799996495\n"
]
],
[
[
"### 10. Tensorflow",
"_____no_output_____"
]
],
[
[
"from tensorflow.linalg import matmul\n\nstart = default_timer()\n\n_ = matmul(mat_1, mat_2)\n\nt10 = default_timer() - start\n\nprint(\"tensorflow Time Taken :\", t10) ",
"tensorflow Time Taken : 5.036223599000095\n"
]
],
[
[
"### 12. Pymc",
"_____no_output_____"
]
],
[
[
"!pip install -q pymc3\nimport pymc3 as pm\n\nstart = default_timer()\n\nres = pm.math.dot(mat_1, mat_2)\nt12 = default_timer() - start\n\nprint(\"pymc Time Taken :\", t12) ",
"pymc Time Taken : 0.006076633000020593\n"
],
[
"import numpy as np\nfrom functools import lru_cache, wraps\nimport numpy as np\n\ndef np_cache(*args, **kwargs):\n \"\"\"LRU cache implementation for functions whose FIRST parameter is a numpy array\n >>> array = np.array([[1, 2, 3], [4, 5, 6]])\n >>> @np_cache(maxsize=256)\n ... def multiply(array, factor):\n ... print(\"Calculating...\")\n ... return factor*array\n >>> multiply(array, 2)\n Calculating...\n array([[ 2, 4, 6],\n [ 8, 10, 12]])\n >>> multiply(array, 2)\n array([[ 2, 4, 6],\n [ 8, 10, 12]])\n >>> multiply.cache_info()\n CacheInfo(hits=1, misses=1, maxsize=256, currsize=1)\n \n \"\"\"\n def decorator(function):\n @wraps(function)\n def wrapper(np_array, *args, **kwargs):\n hashable_array = array_to_tuple(np_array)\n return cached_wrapper(hashable_array, *args, **kwargs)\n\n @lru_cache(*args, **kwargs)\n def cached_wrapper(hashable_array, *args, **kwargs):\n array = np.array(hashable_array)\n return function(array, *args, **kwargs)\n\n def array_to_tuple(np_array):\n \"\"\"Iterates recursivelly.\"\"\"\n try:\n return tuple(array_to_tuple(_) for _ in np_array)\n except TypeError:\n return np_array\n\n # copy lru_cache attributes over too\n wrapper.cache_info = cached_wrapper.cache_info\n wrapper.cache_clear = cached_wrapper.cache_clear\n\n return wrapper\n\n return decorator\n\n@np_cache(maxsize=256)\ndef sq_cache(array):\n return array*array\n \nstarttime = default_timer()\n\nl1 = np.array(mat_1)\n\nsq_cache(l1)\n\nt13 = default_timer() - starttime\nprint(\"Custom Time Taken :\", t13)",
"Custom Time Taken : 0.009742387999722268\n"
],
[
"import matplotlib.pyplot as plt\nfig, ax = plt.subplots()\n\n\n# methods = ['Serial','Parallel','Numpy','Scipy','Pandas','Sympy','Numba','Linalg','Pymatrix','TF','Pymc','Custom']\n# times = [t1, t2, t3, t4, t5, t6, t7, t8, t9, t10, t12, t13]\n\nmethods = ['Serial','Parallel','Sympy','Pymatrix','TF','Pymc','Custom']\ntimes = [t1, t2, t6, t9, t10, t12, t13]\nax.bar(methods,times)\nax.set_ylabel('Time in Seconds')\nax.set_title(f'Speedup Matrix Multiplication ({size}*{size})')\n\nfig.set_size_inches(10, 3)\n\nplt.yscale('log')\nplt.show()",
"_____no_output_____"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
]
] |
4ab7a1cacc4f2ee01340a77f3503f18fdce4dffa
| 223,806 |
ipynb
|
Jupyter Notebook
|
MetENP_Appyter.ipynb
|
metabolomicsworkbench/MetENPAppyter
|
e3bfdddd6694d739be8aeb63efbe35069ebb3e34
|
[
"MIT"
] | null | null | null |
MetENP_Appyter.ipynb
|
metabolomicsworkbench/MetENPAppyter
|
e3bfdddd6694d739be8aeb63efbe35069ebb3e34
|
[
"MIT"
] | null | null | null |
MetENP_Appyter.ipynb
|
metabolomicsworkbench/MetENPAppyter
|
e3bfdddd6694d739be8aeb63efbe35069ebb3e34
|
[
"MIT"
] | null | null | null | 88.425919 | 36,344 | 0.791802 |
[
[
[
"#%%appyter init\nfrom appyter import magic\nmagic.init(lambda _=globals: _())",
"_____no_output_____"
]
],
[
[
"### Load all necessary packages",
"_____no_output_____"
]
],
[
[
"import rpy2\nimport rpy2.robjects as robjects\nimport rpy2.robjects.packages as rpackages\n\nfrom rpy2.robjects import numpy2ri, pandas2ri\n\nimport rpy2.ipython.html\nrpy2.ipython.html.init_printing()\n\nimport rpy2.robjects.lib.ggplot2 as gp\nfrom rpy2.ipython.ggplot import image_png\n\nfrom rpy2.rinterface_lib import openrlib\n\nrobjects.r('''\n\n## add the full path to folder inside which MetENP R package is installed; relative path may not work\n\n.libPaths( c( .libPaths(), \"/home/mano/.local/R\" ))\n\n##.libPaths( c( .libPaths(), \"/var/www/html/appyters/MetENP/R\" ))\n\nprint(.libPaths())\n''')\n;",
"[1] \"/usr/lib64/R/library\" \"/usr/share/R/library\" \"/home/mano/.local/R\" \n"
],
[
"%%appyter hide_code \n\nimport pandas as pd\nfrom io import StringIO\n\n\n_formatter = robjects.r('''\ninstall.packages(\"dplyr\")\nfunction (X) {\n return (\n format_tsv(\n as.data.frame(apply(X, c(1,2), function(x) {\n unlist(x, TRUE, TRUE)\n }))\n )\n )\n}\n''')\ndef as_df(data, index_col=None):\n return pd.read_csv(StringIO(_formatter(data)[0]), sep='\\t', index_col=index_col)",
"_____no_output_____"
],
[
"from rpy2.robjects.packages import importr\nmetenp = importr('MetENP')",
"_____no_output_____"
],
[
"%%appyter hide_code\n{% do SectionField(\n name='INPUT',\n title='Input the Study, Factors and Analysis type Information',\n subtitle='Specify your study here',\n img='upload_icon.png',\n)%}\n{% do SectionField(\n name='SIGM',\n title='Find Significant Metabolites',\n subtitle='Run significance of all the analysis summary together',\n img='vol.png',\n)%}\n{% do SectionField(\n name='COUNTCLASS',\n title='Count Metabolites by Class',\n subtitle='Count the metabolites of each class',\n img='bar.png',\n)%}\n{% do SectionField(\n name='ENRICH',\n title='Enrichment Score by Class',\n subtitle='Calculate the enrichment score of each metabolite class',\n img='enrich.png',\n)%}\n{% do SectionField(\n name='HEAT',\n title='Heatmap',\n subtitle='Heatmap of metabolites x enriched pathways',\n img='heat.png',\n)%}\n{% do SectionField(\n name='DOT',\n title='Dotplot',\n subtitle='Plot a dotplot of enrichment by class',\n img='dot.png',\n)%}\n",
"_____no_output_____"
],
[
"%%appyter code_exec\n\n{% set study = MetabolomicsStudyField(\n name='study',\n label='Study Information',\n description = \"Resolve the factors and analysis type for the study\",\n default={'study_id': 'ST000915', 'fac1': 'Cirrhosis', 'fac2': 'Normal', 'anal': 'Core G Fatty acids/Eicosanoids', 'faccol': 'Diagnosis' },\n section='INPUT',\n) \n\n%}\n\nprint(\"Study ID:\", {{study.value.study_id|jsonify}})\n",
"_____no_output_____"
],
[
"%%appyter hide_code \n\n{% set studyOrg = StringField(name='studyOrg', label='KEGG Organism Code', description = \"hsa for human, mmu for mouse, eco for E. coli, rno for rat. See https://www.genome.jp/kegg/catalog/org_list.html for all organisms\", default='hsa', section='INPUT') %}",
"_____no_output_____"
],
[
"%%appyter hide_code\n\n{% set classtype = MultiChoiceField(\n name='classtype',\n label='Choose a Metabolite Class',\n description='Choose between sub_class, main_class and super_class',\n default=[\n 'sub_class',\n ],\n choices=[\n 'sub_class',\n 'main_class',\n 'super_class',\n ],\n section='INPUT'\n) %}",
"_____no_output_____"
]
],
[
[
"### Start data processing\n\n### Gets metabolomics data, metadata and metabolite info from Metabolomics Workbench using REST service",
"_____no_output_____"
]
],
[
[
"%%appyter code_exec\n\ndata_1 = robjects.r['getmwstudies']({{study.value.study_id|jsonify}}, \n 'data'\n)",
"_____no_output_____"
],
[
"%%appyter hide_code\n\ndata_1 = metenp.getmwstudies({{study.value.study_id|jsonify}}, 'data')\nas_df(data_1, index_col=[0, 1, 2, 3, 4])",
"_____no_output_____"
],
[
"with openrlib.rlock:\n \n robjects.r.assign('data_1', data_1)\n \n robjects.r('''\n\n data_1_print = data.frame(apply(data_1, c(1,2), function(x){unlist(x, TRUE, TRUE)}))\n head(data_1_print)\n\n ''')\n \n pass",
"_____no_output_____"
]
],
[
[
"### Gets metabolomics data, metadata and metabolite info from Metabolomics Workbench using REST service",
"_____no_output_____"
]
],
[
[
"%%appyter code_exec\n\nmetadata = robjects.r['getmwstudies']({{study.value.study_id|jsonify}}, \n 'factors'\n)\n",
"_____no_output_____"
],
[
"robjects.r.assign('metadata', metadata)\n\nrobjects.r('''\nmetadata_print = data.frame(apply(metadata, c(1,2), function(x){unlist(x, TRUE, TRUE)}))\nhead(metadata_print)\n''')",
"_____no_output_____"
]
],
[
[
"### Associate metabolomics data to the refmet class",
"_____no_output_____"
]
],
[
[
"refmet_class = robjects.r['convert_refmet'](data_1)\n\nrobjects.r.assign('refmet_class', refmet_class)\n\nrobjects.r('''\n\nrefmet_class_print = data.frame(apply(refmet_class, c(1,2), function(x){unlist(x, TRUE, TRUE)}))\nhead(refmet_class_print)\n\n''')",
"_____no_output_____"
]
],
[
[
"### This lists each unique metadata factor/independent variable",
"_____no_output_____"
]
],
[
[
"robjects.r('''\nstudy_facs = data.frame(unique(metadata_print$factors))\n''')",
"_____no_output_____"
]
],
[
[
"### This lists each different analysis terms",
"_____no_output_____"
]
],
[
[
"robjects.r('''\nstudy_anals = data.frame(unique(data_1_print$analysis_summary))\n''')",
"_____no_output_____"
]
],
[
[
"### Here, find significant metabolites and the significance of all the analysis summary together. The analysis summary/modes you got in the previous section.",
"_____no_output_____"
]
],
[
[
"%%appyter hide_code\n\n \n \n\n\n\n\n",
"_____no_output_____"
],
[
"%%appyter hide_code\n\n{% set normtype = MultiChoiceField(\n name='normtype',\n label='Select Type of Normalization. How would you like to handle missing data?',\n description='Select 1. half_of_min: where the NAs are replaced by half of min values in the data, 2. remove_NAs: where Cols with NAs values are removed, or 3. 50percent: where cols with more than 50% NAs values are removed. half_of_min is ideal when you wish to see which metabolites were present in either group. Very high fold change would mean it was present in either group.',\n default=[\n '50percent',\n ],\n choices=[\n 'half_of_min',\n 'remove_NAs',\n '50percent',\n ],\n section='SIGM'\n) %}\n",
"_____no_output_____"
],
[
"%%appyter hide_code\n\n{% set pthres = MultiChoiceField(\n name='pthres',\n label='P-Value Threshhold',\n description='P-Value Threshhold for Volcano Plot. Float between 0 and 1.',\n default=[\n '0.05',\n ],\n choices=[\n '0.001',\n '0.002',\n '0.005',\n '0.01',\n '0.02',\n '0.05',\n '0.10'\n ],\n section='SIGM'\n) %}\n",
"_____no_output_____"
],
[
"%%appyter hide_code\n\n#0.0, 0.5, 0.6, 0.8, 1, 1.2, 1.5, 2, 2.5, 3\n#TODO - don't do yet \n\n{% set thres_log2foldchange = StringField(\n name = 'tres_log2foldchange',\n label = 'Log2FoldChange Threshhold',\n description= \"Log2FoldChange Threshhold for Volcano Plot.\",\n default='0.5',\n section='SIGM'\n) %}\n",
"_____no_output_____"
],
[
"%%appyter hide_code\n\n{% set padj = MultiChoiceField(\n name='padj',\n label='Select P Adjust Method',\n description='P adjust method for significance analysis of metabolites',\n default=[\n 'fdr',\n ],\n choices=[\n 'fdr',\n 'BH',\n 'holm',\n 'bonferroni',\n 'hochberg',\n 'hommel',\n ],\n section='SIGM'\n) %}",
"_____no_output_____"
],
[
"%%appyter code_eval\n\nrobjects.r.assign('refmet_class', refmet_class)\nrobjects.r.assign('metcol', 'metabolite_name')\nrobjects.r.assign('fac1', {{study.value.fac1|jsonify}})\nrobjects.r.assign('fac2', {{study.value.fac2|jsonify}})\nrobjects.r.assign('faccol', {{study.value.faccol|jsonify}})\nrobjects.r.assign('samcol', 'local_sample_id')\nrobjects.r.assign('normtype', {{normtype}})\nrobjects.r.assign('pthres', {{pthres}})\nrobjects.r.assign('thres_log2foldchangevar', {{thres_log2foldchange}})\nrobjects.r.assign('padj', {{padj}})",
"_____no_output_____"
],
[
"robjects.r('''\nptreshnum<-as.numeric(pthres)\nthres_log2foldchangenum<-as.numeric(thres_log2foldchangevar)\npadjchar<-as.character(padj[1])\n''')\n;",
"_____no_output_____"
],
[
"%%appyter code_exec\n\nrobjects.r.assign('anal',{{study.value.anal|jsonify}} )\n\n",
"_____no_output_____"
]
],
[
[
"### Find the log2fold change and p value on metabolomics data using a t-test. ",
"_____no_output_____"
]
],
[
[
"robjects.r('''\nanalysis_type_sep = \"___\";\nanal_vec = trimws(unlist(strsplit(anal, analysis_type_sep)))\nprint(anal_vec)\nstats_metabolites = significant_met(metabolomics_data=refmet_class_print, met_col='metabolite_name',analysis_type=anal_vec, metadata=metadata, factor1=fac1, factor2=fac2, factor_col=faccol,sample_col='local_sample_id', p_adjust=padjchar,normalization=normtype)\n''')\n;",
"[1] \"Core G Fatty acids/Eicosanoids\"\n"
],
[
"robjects.r('''\nsig_metabolites = stats_metabolites[which(stats_metabolites[,\"pval\"] <= ptreshnum&abs(stats_metabolites[,\"log2Fold_change\"])>thres_log2foldchangenum),]\n''')\n;",
"_____no_output_____"
],
[
"image_png(robjects.r('''\nplot_volcano(stats_metabolites, thres_pval= ptreshnum, thres_log2foldchange = thres_log2foldchangenum, TRUE)\n'''))",
"_____no_output_____"
]
],
[
[
"Each color describes a different metabolite group.",
"_____no_output_____"
],
[
"### Map metabolite class of the significant metabolites utilzing refmet classification in Metabolomics Workbench\nThis function not only maps metabolite to metabolite class but also to external databases such as pubchem, inchi key, smiles, and KEGG (if you chose to add in KEGG).",
"_____no_output_____"
]
],
[
[
"with openrlib.rlock:\n robjects.r('''\n sig_metabolites_kegg_id= map_keggid(sig_metabolites)\n ''')\n pass",
"_____no_output_____"
]
],
[
[
"#### Check all your significant metabolites have not been assigned metabolite class\nOutcome should be 0 elements.",
"_____no_output_____"
]
],
[
[
"robjects.r('''\nsetdiff(sig_metabolites$refmet_name, sig_metabolites_kegg_id$refmet_name)\n''')",
"_____no_output_____"
]
],
[
[
"### Count the number of metabolites in each of the metabolite classes and plot them with a bar graph",
"_____no_output_____"
]
],
[
[
"%%appyter hide_code\n\n{% set thres_log2foldchange_countclass = StringField(\n name = 'thres_log2foldchange_countclass',\n label = 'Log2FoldChange Threshhold for counting',\n description= \"Log2FoldChange Threshhold for counting a metabolite in a class. Float between 0 and 1.\",\n default='0.5',\n section='COUNTCLASS'\n) %}",
"_____no_output_____"
],
[
"%%appyter code_exec\n\nrobjects.r.assign('classtype', {{classtype}})\n\nrobjects.r.assign('thres_log2foldchange_countclass', {{thres_log2foldchange_countclass}})\n\nrobjects.r('''\nmetclassvar <- as.character(classtype[1])\ncount_changes = metcountplot(df_metclass=sig_metabolites_kegg_id, metclass=metclassvar, plotting=TRUE, thres_logfC = thres_log2foldchange_countclass)\n''')",
"_____no_output_____"
],
[
"image_png(robjects.r('''\ncount_changes$plotimg\n'''))",
"_____no_output_____"
]
],
[
[
"### Enrichment score (p-value) for metabolite classes\n\nCalculate the enrichment score (p-value) of each metabolite class. \n\nEnrichment score (p-value) is calculated through the hypergeometric method as described by Choudhary et al. [https://www.biorxiv.org/content/10.1101/2020.11.20.391912v1].",
"_____no_output_____"
]
],
[
[
"%%appyter hide_code\n\n{% set sigmets = IntField(\n name = 'sigmets',\n label = 'Minimum # of (most significant) metabolites needed in a class to include the class in enrichment.',\n description= \"We advise to use the number of metabolites in each class as 3 or more. However, if you want to know the enrichment score for all the metabolites, choose 1.\",\n default=3,\n section = 'ENRICH'\n) %}",
"_____no_output_____"
],
[
"%%appyter code_eval\nrobjects.r.assign('sigmets', {{sigmets}})",
"_____no_output_____"
],
[
"robjects.r('''\nmetenrichment = metclassenrichment(df_metclass=sig_metabolites_kegg_id,refmet_class, metclass=metclassvar,enrich_stats=\"HG\",no=sigmets)\n''')\n;",
"_____no_output_____"
]
],
[
[
"## Plot the enrichment score (p-value) via function plot_met_enrichment\n\n### If no figure shows here, or an error returns, this means no classes had enough significant metabolites. Decrease the minimum number of significant metabolites per class needed and reload/rerun the appyter program.",
"_____no_output_____"
]
],
[
[
"image_png(robjects.r('''\nplot_met_enrichment(metenrichment, metclassvar,\"HG\", no=sigmets)\n'''))",
"_____no_output_____"
]
],
[
[
"### Check the pathways with reactions involving the significant metabolites",
"_____no_output_____"
]
],
[
[
"%%appyter code_exec\n\nrobjects.r.assign('studyOrg', {{studyOrg}})\n\nrobjects.r('''\nmet_path = met_pathways(df_metenrichment = metenrichment, studyOrg)\nhead(met_path)\n''')",
"_____no_output_____"
]
],
[
[
"### Get pathway enrichment score (p-value).\n\nOnce we have the pathway information, we can calculate enrichment score (p-value) of pathways using the hypergeometric scores for each pathway.\n\nFor hypergeometric score, we need to know the total number of KEGG compounds that are linked to KEGG pathways. \n\nN = Total no. of compounds linked to KEGG pathway, \nL = No. of compounds in a pathway, \nM = No. of altered (significant) compounds in a pathway, \nK = Total no. of altered (significant) compounds. \nThen, \nscore or p-value = phyper(M-1, L, N-L, K)\n\nThis function also utilizes korg dataset from pathview package.",
"_____no_output_____"
]
],
[
[
"%%appyter hide_code\n\n{% set keggTF = MultiChoiceField(\n name='keggTF',\n label='Analysis with KEGG',\n description='Would you like the enrichment analysis to include all KEGG pathways?',\n default=[\n 'FALSE',\n ],\n choices=[\n 'TRUE',\n 'FALSE'\n ],\n section='ENRICH'\n) %}",
"_____no_output_____"
],
[
"%%appyter code_exec\n\nrobjects.r.assign('keggTF', {{keggTF}})\n\nrobjects.r('''\nkeggTFchar<-as.character(keggTF[1])\n''')",
"_____no_output_____"
],
[
"%%appyter code_exec\n\n# relative paths do not work, exclude the starting / as it gets added automatically when appyter runs\n#korgvar = \"{{ url_for('home/sumana/appyters/MetENP/static', filename='korg.RData') }}\"\n#ls_pathvar = \"{{ url_for('home/sumana/appyters/MetENP/static', filename='ls_path.RData') }}\"\n\nkorgvar = \"{{ url_for('var/www/html/appyters/MetENP/static', filename='korg.RData') }}\"\nls_pathvar = \"{{ url_for('var/www/html/appyters/MetENP/static', filename='ls_path.RData') }}\"\n\nrobjects.r.assign('korgvar', korgvar)\nrobjects.r.assign('ls_pathvar', ls_pathvar)",
"_____no_output_____"
],
[
"robjects.r('''\nload(korgvar)\nload(ls_pathvar)\n''')",
"R[write to console]: Error in readChar(con, 5L, useBytes = TRUE) : cannot open the connection\n\nR[write to console]: In addition: \nR[write to console]: There were 11 warnings (use warnings() to see them)\nR[write to console]: \n\n"
],
[
"robjects.r('''\nkegg_es = path_enrichmentscore(met_path,sig_metabolite_kegg_id=sig_metabolite_kegg_id,ls_path=ls_path,refmet_class=refmet_class,sps=studyOrg,padj=padjchar, kegg_comp_path=keggTFchar)\nhead(kegg_es)\n''')",
"_____no_output_____"
]
],
[
[
"### Plot pathway network\nOnly plotting significant pathways (pathways with >= the minimum number significant metabolites). There is no p-value or adjusted p-value threshold applied for the pathways.\n\nThe pathway network is such that it shows metabolites that are connected to different pathways and same metabolite in different pathway. Color of nodes of metabolites are according to the fold change of metabolites (low expression in green and high in red) and size of pathway nodes (square nodes) are according to the number of branches (meaning no of metabolites). All metabolite are written in blue.",
"_____no_output_____"
]
],
[
[
"image_png(robjects.r('''\nplot_pathway_networks (met_path,kegg_es, TRUE)\n'''))",
"_____no_output_____"
]
],
[
[
"### Heatmap",
"_____no_output_____"
]
],
[
[
"%%appyter hide_code\n\n{% set x_size_heat = IntField(\n name = 'x_size_heat',\n label = 'Font Size for X',\n description= \"To change the size of the heatmap\",\n default=8,\n section = 'HEAT'\n) %}",
"_____no_output_____"
],
[
"%%appyter hide_code\n\n{% set y_size_heat = IntField(\n name = 'y_size_heat',\n label = 'Font Size for Y',\n description= \"To change the size of the heatmap\",\n default=6,\n section = 'HEAT'\n) %}",
"_____no_output_____"
],
[
"%%appyter code_exec\n\nrobjects.r.assign('x_size_heat', {{x_size_heat}})\nrobjects.r.assign('y_size_heat', {{y_size_heat}})",
"_____no_output_____"
],
[
"image_png(robjects.r('''\nplot_heatmap(met_path, shorten_name=TRUE,refmet_name=FALSE, xaxis=x_size_heat, yaxis=y_size_heat)\n'''))",
"_____no_output_____"
]
],
[
[
"### Dotplot",
"_____no_output_____"
]
],
[
[
"%%appyter hide_code\n\n{% set x_size_dot = IntField(\n name = 'x_size_dot',\n label = 'Font Size for X',\n description= \"To change the size of the dotplot\",\n default=8,\n section = 'DOT'\n) %}",
"_____no_output_____"
],
[
"%%appyter hide_code\n\n{% set y_size_dot = IntField(\n name = 'y_size_dot',\n label = 'Font Size for Y',\n description= \"To change the size of the dotplot\",\n default=6,\n section = 'DOT'\n) %}",
"_____no_output_____"
],
[
"%%appyter code_exec\n\nrobjects.r.assign('x_size_dot', {{x_size_dot}})\nrobjects.r.assign('y_size_dot', {{y_size_dot}})",
"_____no_output_____"
],
[
"image_png(robjects.r('''\ndotplot_met_class_path (met_path, kegg_es, metclassvar,xaxis=x_size_dot,yaxis=y_size_dot)\n'''))",
"_____no_output_____"
]
],
[
[
"### Get the gene and enzyme info\n\nHere we get the information of genes involved in enriched pathways for specified organism using KEGG database.",
"_____no_output_____"
]
],
[
[
"robjects.r('''\nmet_gene_info = enzyme_gene_info (metenrichment, studyOrg, metclassvar)\nhead(met_gene_info)\n''')",
"_____no_output_____"
]
],
[
[
"#### Get the information if metabolite is a reactant or substrate",
"_____no_output_____"
]
],
[
[
"robjects.r('''\nrclass_info = react_substrate(met_gene_info)\nhead(rclass_info)\n''')",
"_____no_output_____"
]
],
[
[
"#### Get gene info in short form",
"_____no_output_____"
]
],
[
[
"robjects.r('''\nmet_gene_info2=data.table::data.table(rclass_info)[,lapply(.SD, function(x) toString(unique(x))), by = 'Metabolite']\n''')",
"_____no_output_____"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ab7b4e8dc78c7ee89c8dbe6f80157a7c72b3a34
| 547,517 |
ipynb
|
Jupyter Notebook
|
50_modules/01_dust/04_assignments/solutions/04_assignment_solution.ipynb
|
trivedi-c/atm_Practical3
|
1cd1e0bd263d274cada781d871ad37314ebe139e
|
[
"MIT"
] | null | null | null |
50_modules/01_dust/04_assignments/solutions/04_assignment_solution.ipynb
|
trivedi-c/atm_Practical3
|
1cd1e0bd263d274cada781d871ad37314ebe139e
|
[
"MIT"
] | null | null | null |
50_modules/01_dust/04_assignments/solutions/04_assignment_solution.ipynb
|
trivedi-c/atm_Practical3
|
1cd1e0bd263d274cada781d871ad37314ebe139e
|
[
"MIT"
] | null | null | null | 119.336748 | 153,808 | 0.783342 |
[
[
[
"<img src='../../../../img/EU-Copernicus-EUM_3Logos.png' alt='Logo EU Copernicus EUMETSAT' align='right' width='50%'></img>\n<br>",
"_____no_output_____"
],
[
"# 04 - Assignment - Solution",
"_____no_output_____"
],
[
"### About",
"_____no_output_____"
],
[
"> So far, we analysed Aerosol Optical Depth from different types of data (satellite, model-based and observations) for a single dust event. Let us now broaden our view and analyse the annual cycle in 2020 of Aerosol Optical Depth from AERONET and compare it with the CAMS global reanalysis data.",
"_____no_output_____"
],
[
"### Tasks",
"_____no_output_____"
],
[
"#### 1. Download and plot time-series of AERONET data for Santa Cruz, Tenerife in 2020\n * **Hint** \n * [AERONET - Example notebook](../../02_ground-based_observations/21_AERONET.ipynb)\n * you can select daily aggregates of the station observations with setting `AVG=20`\n * **Interpret the results:**\n * Have there been other times in 2020 with increased AOD values?\n * If yes, how could you find out if the increase in AOD is caused by dust? Try to find out by visualizing the AOD time-series together with another parameter from the AERONET data.\n * [MSG SEVIRI Dust RGB](https://sds-was.aemet.es/forecast-products/dust-observations/msg-2013-eumetsat) and [MODIS RGB](https://worldview.earthdata.nasa.gov/) quick looks might be helpful to get a more complete picture of other events that might have happened in 2020.\n\n\n#### 2. Download CAMS global reanalysis (EAC4) and select 2020 time-series for *Santa Cruz, Tenerife*\n * **Hint**\n * [CAMS global forecast - Example notebook](../../03_model-based_data/32_CAMS_global_forecast_duaod_load_browse.ipynb) (**Note:** the notebook works with CAMS forecast data, but they have a similar data structure to the CAMS global reanalysis data)\n * [Data access](https://ads.atmosphere.copernicus.eu/cdsapp#!/dataset/cams-global-reanalysis-eac4?tab=form) with the following specifications:\n > Variable on single levels: `Dust aerosol optical depth at 550 nm` <br>\n > Date: `Start=2020-01-01`, `End=2020-12-31` <br>\n > Time: `[00:00, 03:00, 06:00, 09:00, 12:00, 15:00, 18:00, 21:00]` <br>\n > Restricted area: `N: 30., W: -20, E: 14, S: 20.` <br>\n >Format: `netCDF` <br>\n * With the xarray function `sel()` and keyword argument `method='nearest'` you can select data based on coordinate information\n * We also recommend you to transform your xarray.DataArray into a pandas.DataFrame with the function `to_dataframe()`\n \n\n#### 3. Visualize both time-series of CAMS reanalysis and AERONET daily aggregates in one plot\n * **Interpret the results:** What can you say about the annual cycle in 2020 of AOD in Santa Cruz, Tenerife?",
"_____no_output_____"
],
[
"### Module outline\n* [1 - Select latitude / longitude values for Santa Cruz, Tenerife](#select_lat_lon)\n* [2 - Download and plot time-series of AERONET data](#aeronet)\n* [3 - Download CAMS global reanalysis (EAC4) and select 2020 time-series for Santa Cruz, Tenerife](#cams_reanalysis)\n* [4 - Combine both annual time-series and visualize both in one plot](#visualize_annual_ts)\n",
"_____no_output_____"
],
[
"<hr>",
"_____no_output_____"
],
[
"##### Load required libraries",
"_____no_output_____"
]
],
[
[
"%matplotlib inline\nimport os\nimport xarray as xr\nimport numpy as np\nimport netCDF4 as nc\nimport pandas as pd\n\nfrom IPython.display import HTML\n\nimport matplotlib.pyplot as plt\nimport matplotlib.colors\nfrom matplotlib.cm import get_cmap\nfrom matplotlib import animation\nimport cartopy.crs as ccrs\nfrom cartopy.mpl.gridliner import LONGITUDE_FORMATTER, LATITUDE_FORMATTER\nimport cartopy.feature as cfeature\n\nfrom matplotlib.axes import Axes\nfrom cartopy.mpl.geoaxes import GeoAxes\nGeoAxes._pcolormesh_patched = Axes.pcolormesh\n\nimport wget\n\nimport warnings\nwarnings.simplefilter(action = \"ignore\", category = RuntimeWarning)",
"_____no_output_____"
]
],
[
[
"##### Load helper functions",
"_____no_output_____"
]
],
[
[
"%run ../functions.ipynb",
"_____no_output_____"
]
],
[
[
"<hr>",
"_____no_output_____"
],
[
"### <a id='select_lat_lon'></a>1. Select latitude / longitude values for Santa Cruz, Tenerife",
"_____no_output_____"
],
[
"You can see an overview of all available AERONET Site Names [here](https://aeronet.gsfc.nasa.gov/cgi-bin/draw_map_display_aod_v3?long1=-180&long2=180&lat1=-90&lat2=90&multiplier=2&what_map=4&nachal=1&formatter=0&level=3&place_code=10&place_limit=0).",
"_____no_output_____"
]
],
[
[
"lat = 28.473\nlon = -16.247",
"_____no_output_____"
]
],
[
[
"<br>",
"_____no_output_____"
],
[
"### <a id='aeronet'></a>2. Download and plot time-series of AERONET data",
"_____no_output_____"
],
[
"As a first step, let us create a Python dictionary in which we store all the parameters we would like to use for the request as dictionary keys. You can initiate a dictionary with curled brackets `{}`. Below, we specify the following parameters:\n* `endpoint`: Endpoint of the AERONET web service\n* `station`: Name of the AERONET station\n* `year`: year 1 of interest\n* `month`: month 1 of interest\n* `day`: day 1 of interest\n* `year2`: year 2 of interest\n* `month2`: month 2 of interest\n* `day2`: day 2 of interest\n* `AOD15`: data type, other options include `AOD10`, `AOD20`, etc.\n* `AVG`: data format, `AVG=10` - all points, `AVG=20` - daily averages",
"_____no_output_____"
],
[
"The keywords below are those we will need for requesting daily averaged observations of Aerosol Optical Depth Level 1.5 data for the station Santa Cruz, Tenerife from 1 January to 31 December 2020.",
"_____no_output_____"
]
],
[
[
"data_dict = {\n 'endpoint': 'https://aeronet.gsfc.nasa.gov/cgi-bin/print_web_data_v3',\n 'station':'Santa_Cruz_Tenerife',\n 'year': 2020,\n 'month': 1,\n 'day': 1,\n 'year2': 2020,\n 'month2': 12,\n 'day2': 31,\n 'AOD15': 1,\n 'AVG': 20\n}\n",
"_____no_output_____"
]
],
[
[
"In a next step, we construct the final string for the wget request with the `format` function. You construct a string by adding the dictionary keys in curled brackets. At the end of the string, you provide the dictionary key informatoin to the string with the `format()` function. A print of the resulting url shows, that the format function replaced the information in the curled brackets with the data in the dictionary.",
"_____no_output_____"
]
],
[
[
"url = '{endpoint}?site={station}&year={year}&month={month}&day={day}&year2={year2}&month2={month2}&day2={day2}&AOD15={AOD15}&AVG={AVG}'.format(**data_dict)\nurl",
"_____no_output_____"
]
],
[
[
"<br>",
"_____no_output_____"
],
[
"Now we are ready to request the data with the function `download()` from the wget Python library. You have to pass to the function the constructed url above together with a file path of where the downloaded that shall be stored. Let us store the data as `txt` file in the folder `../data/2_observations/aeronet/`.",
"_____no_output_____"
]
],
[
[
"wget.download(url, '../../../../eodata/50_modules/01_dust/04_assignment/aeronet/2020_santa_cruz_tenerife_20.txt')",
"_____no_output_____"
]
],
[
[
"<br>",
"_____no_output_____"
],
[
"After we downloaded the station observations as `txt` file, we can open it with the pandas function `read_table()`. We additonally set specific keyword arguments that allow us to specify the columns and rows of interest:\n* `delimiter`: specify the delimiter in the text file, e.g. comma\n* `header`: specify the index of the row that shall be set as header.\n* `index_col`: specify the index of the column that shall be set as index\n\nYou see below that the resulting dataframe has 296 rows and 81 columns.",
"_____no_output_____"
]
],
[
[
"df = pd.read_table('../../../../eodata/50_modules/01_dust/04_assignment/aeronet/2020_santa_cruz_tenerife_20.txt', delimiter=',', header=[7], index_col=1)\ndf",
"_____no_output_____"
]
],
[
[
"Now, we can inspect the entries in the loaded data frame a bit more. Above you see that the last entry is a NaN entry, which is best to drop with the function `dropna()`.\n\nThe next step is then to replace the entries with -999.0 and set them as NaN. We can use the function `replace()` to do so. ",
"_____no_output_____"
]
],
[
[
"df = df.dropna()\ndf = df.replace(-999.0, np.nan)\ndf",
"_____no_output_____"
]
],
[
[
"Let us now convert the index entry to a `DateTimeIndex` format with the function `to_datetime()`. Important here, you have to specify the format of the index string: `%d:%m:%Y`.\n\nYou see below that we do not have observations for every day. E.g on 2 January 2020, the data frame does not list any entry.",
"_____no_output_____"
]
],
[
[
"df.index = pd.to_datetime(df.index, format = '%d:%m:%Y')\ndf",
"_____no_output_____"
]
],
[
[
"<br>",
"_____no_output_____"
],
[
"We can now plot the column `AOD_500nm` as annual time-series. You see that the station `Santa Cruz, Tenerife` was affected by other dust events later in 2020.",
"_____no_output_____"
]
],
[
[
"# Initiate a figure\nfig = plt.figure(figsize=(20,8))\nax = plt.subplot()\n\n# Define the plotting function\nax.plot(df.AOD_500nm, 'o-', color='green', label='AERONET observations')\n\n# Customize the title and axes lables\nax.set_title('\\nAerosol Optical Depth at 500 nm - Santa Cruz Tenerife\\n', fontsize=20)\nax.set_ylabel('~', fontsize=14)\nax.set_xlabel('\\nDay', fontsize=14)\n\n# Customize the fontsize of the axes tickes\nplt.xticks(fontsize=14)\nplt.yticks(fontsize=14)\n\n# Add a gridline to the plot\nax.grid(linestyle='--')\n\nplt.legend(fontsize=16, loc=2)\n",
"_____no_output_____"
]
],
[
[
"<br>",
"_____no_output_____"
],
[
"The next question is now, how you can find out, if the strong increase of AOD at the end of August 2020 was because of dust? For this to find out, you can use the `Angstrom Exponent`, which gives us an indication of the particle size. If the `Angstrom Exponent` is below 0.6, then it is an indication that the increase of AOD is caused by coarser dust particles.\n\nLet us visualize the AOD at 500nm for 2020 together with the `Angstrom Exponent 440-675nm`.",
"_____no_output_____"
]
],
[
[
"# Initiate a figure\nfig = plt.figure(figsize=(20,8))\nax = plt.subplot()\n\n# Define the plotting function\nax.plot(df.AOD_500nm, 'o-', color='green', label='AERONET observations')\nax.plot(df['440-675_Angstrom_Exponent'], '-', color='lightgrey', label='Angstrom Exponent - 440-675nm')\n\nplt.axhline(y=0.6, color='r', linestyle='dotted', label='Angstrom Exponent <0.6 is dust')\n\n# Customize the title and axes lables\nax.set_title('\\nAerosol Optical Depth at 500 nm - Santa Cruz Tenerife\\n', fontsize=20)\nax.set_ylabel('~', fontsize=14)\nax.set_xlabel('\\nDay', fontsize=14)\n\n# Customize the fontsize of the axes tickes\nplt.xticks(fontsize=14)\nplt.yticks(fontsize=14)\n\n# Add a gridline to the plot\nax.grid(linestyle='--')\n\nplt.legend(fontsize=16, loc=2)",
"_____no_output_____"
]
],
[
[
"Above, you see that the Angstrom Exponent during the high AOD values at the end of August is very low. Hence, we could interpret this event as a strong dust intrusion. But is this really the case? You can also check [here](https://sds-was.aemet.es/forecast-products/dust-observations/msg-2013-eumetsat) the MSG SEVIRI Dust RGB for e.g. 26 August 2020 and [here](https://worldview.earthdata.nasa.gov/?v=-39.451155087380556,13.025874527486357,5.067364489712844,36.350274677008436&l=Reference_Labels_15m(hidden),Reference_Features_15m(hidden),Coastlines_15m,MODIS_Aqua_CorrectedReflectance_TrueColor(hidden),MODIS_Terra_CorrectedReflectance_TrueColor&lg=false&t=2020-08-26-T00%3A00%3A00Z) the MODIS RGB to better understand the event and what could have caused the high AOD values.",
"_____no_output_____"
],
[
"<br>",
"_____no_output_____"
],
[
"### <a id='cams_reanalysis'></a> 3. Download CAMS global reanalysis (EAC4) and select 2020 time-series for Santa Cruz, Tenerife",
"_____no_output_____"
],
[
"First, we have to download the `CAMS global reanalysis (EAC4)` from the [Copernicus Atmosphere Data Store](https://ads.atmosphere.copernicus.eu/cdsapp#!/dataset/cams-global-reanalysis-eac4?tab=form) with the following specifications:\n* Variable on single levels: `Dust aerosol optical depth at 550 nm`\n* Date: `Start=2020-01-01`, `End=2020-12-31`\n* Time: `[00:00, 03:00, 06:00, 09:00, 12:00, 15:00, 18:00, 21:00]`\n* Restricted area: `N: 30., W: -20, E: 14, S: 20.`\n* Format: `netCDF`\n\nSee `CDSAPI` request below.",
"_____no_output_____"
]
],
[
[
"URL = 'https://ads.atmosphere.copernicus.eu/api/v2'\nKEY = '######################'",
"_____no_output_____"
],
[
"import cdsapi\n\nc = cdsapi.Client(url=URL, key=KEY)\n\nc.retrieve(\n 'cams-global-reanalysis-eac4',\n {\n 'variable': 'dust_aerosol_optical_depth_550nm',\n 'date': '2020-01-01/2020-12-31',\n 'time': [\n '00:00', '03:00', '06:00',\n '09:00', '12:00', '15:00',\n '18:00', '21:00',\n ],\n 'area': [\n 30, -20, 20,\n 15,\n ],\n 'format': 'netcdf',\n },\n '../../../../eodata/50_modules/01_dust/04_assignment/cams/2020_dustAOD_cams_eac4.nc'}",
"_____no_output_____"
]
],
[
[
"<br>",
"_____no_output_____"
],
[
"The data is in netCDF, so we can open the netCDF file with the xarray function `open_dataset()`. We see that the data has three dimensions (`latitude`, `longitude`, `time`) and one data variable:\n* `duaod550`: Dust Aerosol Optical Depth at 550nm",
"_____no_output_____"
]
],
[
[
"file = xr.open_dataset('../../../../eodata/50_modules/01_dust/04_assignment/cams/2020_dustAOD_cams_eac4.nc')\nfile",
"_____no_output_____"
]
],
[
[
"<br>",
"_____no_output_____"
],
[
"Let us now store the data variable `Dust Aerosol Optical Depth (AOD) at 550nm` as `xarray.DataArray` with the name `duaod_cams`.",
"_____no_output_____"
]
],
[
[
"duaod_cams = file['duaod550']\nduaod_cams",
"_____no_output_____"
]
],
[
[
"<br>",
"_____no_output_____"
],
[
"Now, we can select the time-series of the grid point nearest to the station in Santa Cruz, Tenerife. We can use the function `sel()` to select data based on the longitude and latitude dimensions. The keyword argument `method='nearest'` selects the grid point entry closest to the station coordinates.",
"_____no_output_____"
]
],
[
[
"cams_ts = duaod_cams.sel(longitude=lon, latitude=lat, method='nearest')\ncams_ts",
"_____no_output_____"
]
],
[
[
"<br>",
"_____no_output_____"
],
[
"The next step is now to resample the 3-hourly time entries and aggregate it to daily averages. We can use a combination of the functions `resample()` and `mean()` to create daily averages.",
"_____no_output_____"
]
],
[
[
"cams_ts_resample = cams_ts.resample(time='1D').mean()\ncams_ts_resample",
"_____no_output_____"
]
],
[
[
"<br>",
"_____no_output_____"
],
[
"A closer look at the `time` dimension shows us that we now have an entry for each day in 2020.",
"_____no_output_____"
]
],
[
[
"cams_ts_resample.time",
"_____no_output_____"
]
],
[
[
"<br>",
"_____no_output_____"
],
[
"Now, we can convert the `xarray.DataArray` to a `pandas.DataFrame`, as pandas is more efficient to handle time-series data. The function `to_dataframe()` easily converts a data array to a dataframe. The resulting dataframe has 366 rows and 3 columns.",
"_____no_output_____"
]
],
[
[
"cams_ts_df = cams_ts_resample.to_dataframe()\ncams_ts_df",
"_____no_output_____"
]
],
[
[
"<br>",
"_____no_output_____"
],
[
"### <a id='visualize_annual_ts'></a>4. Combine both annual time-series and visualize both in one plot",
"_____no_output_____"
],
[
"Let us now use the function `join()` and combine the two time-series `cams_ts_df` and `df['AOD_500nm]`. The resulting dataframe has 366 rows and 4 columns.",
"_____no_output_____"
]
],
[
[
"df_combined = cams_ts_df.join(df['AOD_500nm'])\ndf_combined",
"_____no_output_____"
]
],
[
[
"Let us safe the pandas dataframe as csv file. This allows us to easily load the time-series again at a later stage. You can use the function `to_csv()` to save a pandas.DataFrame as csv.",
"_____no_output_____"
]
],
[
[
"df_combined.to_csv(\"../../../../eodata/50_modules/01_dust/04_assignment/2020_ts_cams_aeronet.csv\", index_label='time')",
"_____no_output_____"
]
],
[
[
"<br>",
"_____no_output_____"
],
[
"The last step is now to plot the two columns of the pandas.DataFrame `df_combined` as two individual line plots.",
"_____no_output_____"
]
],
[
[
"# Initiate a figure\nfig = plt.figure(figsize=(20,8))\nax = plt.subplot()\n\n# Define the plotting function\nax.plot(df_combined.duaod550, '-', color='blue', label='CAMS global reanalysis (EAC4) - 550 nm')\nax.plot(df_combined.AOD_500nm, '-', color='green', label='AERONET observations - 500 nm')\nplt.axhline(y=0.6, color='r', linestyle='dotted', label='PM10 daily limit')\n\n# Customize the title and axes lables\nax.set_title('\\nAerosol Optical Depth at 500 / 550 nm - Santa Cruz Tenerife\\n', fontsize=20)\nax.set_ylabel(cams_ts.units, fontsize=14)\nax.set_xlabel('\\nDay', fontsize=14)\n\n# Customize the fontsize of the axes tickes\nplt.xticks(fontsize=14)\nplt.yticks(fontsize=14)\n\n# Add a gridline to the plot\nax.grid(linestyle='--')\n\nplt.legend(fontsize=14, loc=2)\n",
"_____no_output_____"
]
],
[
[
"<br>",
"_____no_output_____"
],
[
"You see in the plot above that the model and the AERONET observations follow a similar annual cycle of AOD in 2020 for the Santa Cruz station in Tenerife. You also see that for higher AOD values measured by AERONET, the CAMS model mostly underpredicts the AOD intensity.",
"_____no_output_____"
],
[
"<hr>",
"_____no_output_____"
],
[
"<img src='../../img/copernicus_logo.png' alt='Logo EU Copernicus' align='right' width='20%'><br><br><br><br>\n<p style=\"text-align:right;\">This project is licensed under the <a href=\"./LICENSE\">MIT License</a> and is developed under a Copernicus contract.",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
4ab7b58a16aae22c6edf8e3639702c45a765c651
| 13,760 |
ipynb
|
Jupyter Notebook
|
notebooks/artwork/setup.ipynb
|
tomwhite/shiny-train
|
5366760067fc98a8008a195dbb28475b5aa9f3af
|
[
"MIT"
] | 1 |
2020-10-27T23:30:59.000Z
|
2020-10-27T23:30:59.000Z
|
notebooks/artwork/setup.ipynb
|
tomwhite/shiny-train
|
5366760067fc98a8008a195dbb28475b5aa9f3af
|
[
"MIT"
] | null | null | null |
notebooks/artwork/setup.ipynb
|
tomwhite/shiny-train
|
5366760067fc98a8008a195dbb28475b5aa9f3af
|
[
"MIT"
] | 1 |
2020-10-27T13:53:24.000Z
|
2020-10-27T13:53:24.000Z
| 32.761905 | 121 | 0.522747 |
[
[
[
"import zarr\nfrom pyprojroot import here\nimport pandas as pd\nimport numpy as np\nimport allel\nimport yaml\nimport matplotlib.pyplot as plt\nimport functools\nimport seaborn as sns\nsns.set_context('paper')\nsns.set_style('darkgrid')\nimport dask.array as da\nimport scipy.interpolate\nimport scipy.stats\nimport petl as etl\nimport pyfasta\nfrom matplotlib.ticker import MultipleLocator, FormatStrFormatter\n%matplotlib inline\n%config InlineBackend.figure_format = 'retina'",
"_____no_output_____"
],
[
"# OLD VECTORBASE - gone\n\n# genome_path = here() / 'data/external/vectorbase/Anopheles-gambiae-PEST_CHROMOSOMES_AgamP4.fa'\n# genome = pyfasta.Fasta(str(genome_path), key_fn=lambda x: x.split()[0])\n\n# NEW VECTORBASE\n\ndef _genome_key_fn(s):\n k = s.split()[0]\n if k.startswith('AgamP4'):\n k = k.split('_')[1]\n return k\n\ngenome_path = here() / 'data/external/vectorbase/VectorBase-48_AgambiaePEST_Genome.fasta'\ngenome = pyfasta.Fasta(str(genome_path), key_fn=_genome_key_fn)",
"_____no_output_____"
],
[
"chromosomes = '2', '3', 'X'\nchromosome_plen = {\n '2': len(genome['2R']) + len(genome['2L']), \n '3': len(genome['3R']) + len(genome['3L']), \n 'X': len(genome['X'])\n}",
"_____no_output_____"
],
[
"pop_defs_path = here() / 'notebooks/gwss/pop_defs.yml'\nwith open(pop_defs_path, 'rt') as f:\n pop_defs = yaml.safe_load(f)\npops = list(pop_defs)",
"_____no_output_____"
],
[
"h12_root_path = here() / 'data/gwss/h12/h12.zarr'\nh12_root = zarr.open_consolidated(str(h12_root_path))\n\n \ndef load_h12_gwss(pop, chromosome):\n window_size = pop_defs[pop]['h12_window_size']\n window_step = 200\n grp = h12_root[f'{pop}/{window_size}/{window_step}/{chromosome}']\n return (\n grp['windows'][:],\n grp['gwindows'][:],\n grp['h1'][:],\n grp['h12'][:],\n grp['h123'][:],\n grp['h2_h1'][:]\n )",
"_____no_output_____"
],
[
"ihs_root_path = here() / 'data/gwss/ihs/ihs.zarr'\nihs_root = zarr.open_consolidated(str(ihs_root_path))\n\n\[email protected]_cache(maxsize=None)\ndef load_ihs_gwss(pop, chromosome, window_size=200, window_step=100):\n grp = ihs_root[f'{pop}/{chromosome}']\n pos = grp['pos'][:]\n gpos = grp['gpos'][:]\n ihs_std = np.fabs(grp['ihs_std'][:])\n x = allel.moving_statistic(pos, np.mean, size=window_size, step=window_step)\n gx = allel.moving_statistic(gpos, np.mean, size=window_size, step=window_step)\n y_max = allel.moving_statistic(ihs_std, np.max, size=window_size, step=window_step)\n y_pc95 = allel.moving_statistic(ihs_std, lambda v: np.percentile(v, 95), size=window_size, step=window_step)\n y_pc75 = allel.moving_statistic(ihs_std, lambda v: np.percentile(v, 75), size=window_size, step=window_step)\n y_pc50 = allel.moving_statistic(ihs_std, np.median, size=window_size, step=window_step)\n return x, gx, y_max, y_pc95, y_pc75, y_pc50\n",
"_____no_output_____"
],
[
"xpehh_root_path = here() / 'data/gwss/xpehh/xpehh.zarr'\nxpehh_root = zarr.open_consolidated(str(xpehh_root_path))\n\n\[email protected]_cache(maxsize=None)\ndef load_xpehh_gwss(pop1, pop2, chromosome, window_size=500, window_step=250):\n\n # avoid running the same scan twice\n orig_pop1, orig_pop2 = pop1, pop2\n pop1, pop2 = sorted([pop1, pop2])\n \n grp = xpehh_root[f'{pop1}_{pop2}/{chromosome}']\n pos = grp['pos'][:]\n gpos = grp['gpos'][:]\n xpehh = grp['xpehh'][:]\n\n if pop1 == orig_pop2:\n # flip back\n xpehh = -xpehh\n pop1, pop2 = pop2, pop1\n\n # centre\n xpehh = xpehh - np.median(xpehh)\n \n # clip at zero to focus on selection in pop1\n xpehh1 = np.clip(xpehh, a_min=0, a_max=None)\n \n x = allel.moving_statistic(pos, np.mean, size=window_size, step=window_step)\n gx = allel.moving_statistic(gpos, np.mean, size=window_size, step=window_step)\n y_max = allel.moving_statistic(xpehh1, np.max, size=window_size, step=window_step)\n y_pc95 = allel.moving_statistic(xpehh1, lambda v: np.percentile(v, 95), size=window_size, step=window_step)\n y_pc75 = allel.moving_statistic(xpehh1, lambda v: np.percentile(v, 75), size=window_size, step=window_step)\n y_pc50 = allel.moving_statistic(xpehh1, np.median, size=window_size, step=window_step)\n \n return x, gx, y_max, y_pc95, y_pc75, y_pc50\n",
"_____no_output_____"
],
[
"pbs_root_path = here() / 'data/gwss/pbs/pbs.zarr'\npbs_root = zarr.open_consolidated(str(pbs_root_path))\n\n\ndef load_pbs_gwss(pop1, pop2, pop3, chromosome, window_size=500, window_step=250):\n\n grp_path = f'/{pop1}_{pop2}_{pop3}/{window_size}/{window_step}/{chromosome}'\n grp = pbs_root[grp_path]\n windows = grp['windows'][:]\n gwindows = grp['gwindows'][:]\n pbs = grp['pbs'][:]\n pbs_scaled = grp['pbs_scaled'][:]\n \n return windows, gwindows, pbs, pbs_scaled\n",
"_____no_output_____"
],
[
"def load_genes():\n\n # OLD VECTORBASE\n# features_path = here() / 'data/external/vectorbase/Anopheles-gambiae-PEST_BASEFEATURES_AgamP4.12.gff3'\n# df_genes = (\n# allel.gff3_to_dataframe(\n# str(features_path),\n# attributes=['ID', 'Name', 'biotype']\n# )\n# .set_index('ID')\n# .query(\"type == 'gene' and biotype == 'protein_coding'\")\n# )\n\n # NEW VECTORBASE\n features_path = here() / 'data/external/vectorbase/VectorBase-48_AgambiaePEST.gff'\n df_genes = (\n allel.gff3_to_dataframe(\n str(features_path),\n attributes=['ID', 'description']\n )\n .sort_values(['seqid', 'start'])\n .set_index('ID')\n .query(\"type == 'gene'\")\n )\n # fix chromosome IDs\n df_genes['seqid'] = df_genes['seqid'].str.split('_', expand=True).loc[:, 1]\n\n # convert to chromosomal coordinates\n df_genes['chromosome'] = df_genes['seqid'].copy()\n df_genes['chromosome_start'] = df_genes['start'].copy()\n df_genes['chromosome_end'] = df_genes['end'].copy()\n loc_2R = df_genes.seqid == '2R'\n df_genes.loc[loc_2R, 'chromosome'] = '2'\n loc_2L = df_genes.seqid == '2L'\n df_genes.loc[loc_2L, 'chromosome'] = '2'\n df_genes.loc[loc_2L, 'chromosome_start'] = df_genes.loc[loc_2L, 'start'] + len(genome['2R'])\n df_genes.loc[loc_2L, 'chromosome_end'] = df_genes.loc[loc_2L, 'end'] + len(genome['2R'])\n loc_3R = df_genes.seqid == '3R'\n df_genes.loc[loc_3R, 'chromosome'] = '3'\n loc_3L = df_genes.seqid == '3L'\n df_genes.loc[loc_3L, 'chromosome'] = '3'\n df_genes.loc[loc_3L, 'chromosome_start'] = df_genes.loc[loc_3L, 'start'] + len(genome['3R'])\n df_genes.loc[loc_3L, 'chromosome_end'] = df_genes.loc[loc_3L, 'end'] + len(genome['3R'])\n df_genes['chromosome_center'] = (df_genes['chromosome_start'] + df_genes['chromosome_end']) / 2\n \n return df_genes\n\n\ndf_genes = load_genes()",
"_____no_output_____"
],
[
"import warnings",
"_____no_output_____"
],
[
"with warnings.catch_warnings():\n warnings.simplefilter('ignore')\n ace1 = df_genes.loc['AGAP001356']\n ace1['Name'] = 'Ace1'\n cyp6p3 = df_genes.loc['AGAP002865']\n cyp6p3['Name'] = 'Cyp6p3'\n vgsc = df_genes.loc['AGAP004707']\n vgsc['Name'] = 'Vgsc'\n gaba = df_genes.loc['AGAP006028']\n gaba['Name'] = 'Gaba'\n gste2 = df_genes.loc['AGAP009194']\n gste2['Name'] = 'Gste2'\n cyp9k1 = df_genes.loc['AGAP000818']\n cyp9k1['Name'] = 'Cyp9k1'\nir_genes = [ace1, cyp6p3, vgsc, gaba, gste2, cyp9k1]",
"_____no_output_____"
],
[
"novel_loci = {\n 'A': ('2', 24_860_000),\n 'B': ('2', 40_940_000),\n 'C': ('2', 28_549_590 + len(genome['2R'])),\n 'D': ('2', 34_050_000 + len(genome['2R'])),\n 'E': ('X', 4_360_000),\n 'F': ('X', 9_220_000),\n}",
"_____no_output_____"
],
[
"tbl_chromatin = [\n ('name', 'chrom', 'start', 'end'),\n ('CHX', 'X', 20009764, 24393108),\n ('CH2R', '2R', 58984778, 61545105),\n ('CH2L', '2L', 1, 2431617),\n ('PEU2L', '2L', 2487770, 5042389),\n ('IH2L', '2L', 5078962, 5788875),\n ('IH3R', '3R', 38988757, 41860198),\n ('CH3R', '3R', 52161877, 53200684),\n ('CH3L', '3L', 1, 1815119),\n ('PEU3L', '3L', 1896830, 4235209),\n ('IH3L', '3L', 4264713, 5031692)\n]",
"_____no_output_____"
],
[
"seq_ids = '2R', '2L', '3R', '3L', 'X'",
"_____no_output_____"
],
[
"def build_gmap():\n \n # crude recombination rate lookup, keyed off chromatin state\n # use units of cM / bp, assume 2 cM / Mbp == 2x10^-6 cM / bp\n tbl_rr = (\n etl.wrap(tbl_chromatin)\n # extend heterochromatin on 2L - this is empirical, based on making vgsc peaks symmetrical\n .update('end', 2840000, where=lambda r: r.name == 'CH2L')\n .update('start', 2840001, where=lambda r: r.name == 'PEU2L')\n .addfield('rr', lambda r: .5e-6 if 'H' in r.name else 2e-6)\n )\n \n # per-base map of recombination rates\n rr_map = {seq_id: np.full(len(genome[seq_id]), fill_value=2e-6, dtype='f8')\n for seq_id in seq_ids}\n for row in tbl_rr.records():\n rr_map[row.chrom][row.start - 1:row.end] = row.rr\n \n # genetic map\n gmap = {seq_id: np.cumsum(rr_map[seq_id]) for seq_id in seq_ids}\n gmap['2'] = np.concatenate([gmap['2R'], gmap['2L'] + gmap['2R'][-1]])\n gmap['3'] = np.concatenate([gmap['3R'], gmap['3L'] + gmap['3R'][-1]])\n \n return gmap\n\ngmap = build_gmap()",
"_____no_output_____"
],
[
"def tex_italicize_species(s):\n return (\n s\n .replace('An. gambiae', '\\textit{An. gambiae}')\n .replace('An. coluzzii', '\\textit{An. coluzzii}')\n )\n",
"_____no_output_____"
],
[
"def root_mean_square(s):\n return np.sqrt(np.mean(s**2))\n\n\ndef mean_absolute(s):\n return np.mean(np.fabs(s))\n",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab7c797d02bf798860772bbd59b18552872f383
| 4,717 |
ipynb
|
Jupyter Notebook
|
gs_quant/tutorials/0_price_and_risk_basics.ipynb
|
jeamick/gs-quant
|
a61dd2866dafc8453949391e900f9bf1ce2ad52e
|
[
"Apache-2.0"
] | null | null | null |
gs_quant/tutorials/0_price_and_risk_basics.ipynb
|
jeamick/gs-quant
|
a61dd2866dafc8453949391e900f9bf1ce2ad52e
|
[
"Apache-2.0"
] | 1 |
2020-02-15T11:58:29.000Z
|
2020-02-15T11:58:29.000Z
|
gs_quant/tutorials/0_price_and_risk_basics.ipynb
|
jeamick/gs-quant
|
a61dd2866dafc8453949391e900f9bf1ce2ad52e
|
[
"Apache-2.0"
] | null | null | null | 24.567708 | 112 | 0.569642 |
[
[
[
"# Pricing and Risk Calculation - The Basics\nIn this tutorial we construct IR Swaps and IR Swaptions and calculate price and risk for these instruments",
"_____no_output_____"
]
],
[
[
"from gs_quant.common import PayReceive\nfrom gs_quant.instrument import IRSwap, IRSwaption\nfrom gs_quant.markets import PricingContext\nfrom gs_quant.session import GsSession\nfrom gs_quant import risk\n\nclient_id = None\nclient_secret = None\n\nGsSession.use(client_id=client_id, client_secret=client_secret,\n scopes=['run_analytics', 'read_financial_data'])",
"_____no_output_____"
]
],
[
[
"### Pricing Context\nThe current PricingContext shows current pricing environment and market data paramaterization",
"_____no_output_____"
]
],
[
[
"print(PricingContext.current.market_data_location)\nprint(PricingContext.current.market_data_as_of)\nprint(PricingContext.current.pricing_date)",
"_____no_output_____"
]
],
[
[
"### IR Swaps",
"_____no_output_____"
]
],
[
[
"# Creating Swaps - spot starting, forward starting\nswap = IRSwap(PayReceive.Receive, '10y', 'GBP',\n fixed_rate='atm+50', notional_amount=1e8) # running\nswap_fwd_start = IRSwap(PayReceive.Pay, '5y', 'EUR',\n fixed_rate='atm+20', effective_date='3y') # fwd starting",
"_____no_output_____"
],
[
"# View Instrument w/ specified relative parameters\nswap_fwd_start.as_dict()",
"_____no_output_____"
],
[
"# Resolve Instrument, View fixed parameters\nswap_fwd_start.resolve()\nswap_fwd_start.as_dict()",
"_____no_output_____"
],
[
"# Calculate Price - Local Ccy and USD\nprint('swap price (local): {price}'.format(price=swap.price()))\nprint('swap price (usd): {price}'.format(price=swap.dollar_price()))",
"_____no_output_____"
],
[
"# Calculate IR Delta (Parallel and Ladder)\nir_risk = swap.calc(\n (risk.IRDeltaParallel, risk.IRDeltaParallelLocalCcy, risk.IRDelta))\n\n# Print Risks\nprint(ir_risk[risk.IRDeltaParallel])\nprint(ir_risk[risk.IRDeltaParallelLocalCcy])\nprint(ir_risk[risk.IRDelta])",
"_____no_output_____"
]
],
[
[
"### IR Swaptions",
"_____no_output_____"
]
],
[
[
"# Swaption Definition - 13m5y USD Swaption struck atm+50\nswaption = IRSwaption(PayReceive.Receive, '10y', 'GBP',\n strike='atm+50', expiration_date='13m')",
"_____no_output_____"
],
[
"# View Swaption Strike\nprint('Unresolved strike: {strike}'.format(strike=swaption.strike))\nswaption.resolve()\nprint('Resolved strike: {strike}'.format(strike=swaption.strike))",
"_____no_output_____"
],
[
"# Swaption Risk - IRDelta, IRGamma, IRVega, Theta\nres = swaption.calc(\n (risk.DollarPrice, risk.IRDeltaParallel, risk.IRVega, risk.Theta))\n\nprint(res[risk.Theta])",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
]
] |
4ab7f7cf0c36daed082caf4851169a126f7842c3
| 243,326 |
ipynb
|
Jupyter Notebook
|
notebooks/models/2_modelling_svr_linear_msp.ipynb
|
JeromeSauer/Capstone_WindPowerPredicting
|
469e362853990cc78338933e1bf5ce76dd41b78d
|
[
"MIT"
] | null | null | null |
notebooks/models/2_modelling_svr_linear_msp.ipynb
|
JeromeSauer/Capstone_WindPowerPredicting
|
469e362853990cc78338933e1bf5ce76dd41b78d
|
[
"MIT"
] | 28 |
2021-11-22T12:22:37.000Z
|
2021-12-21T14:14:10.000Z
|
notebooks/models/2_modelling_svr_linear_msp.ipynb
|
JeromeSauer/Capstone_WindPowerPredicting
|
469e362853990cc78338933e1bf5ce76dd41b78d
|
[
"MIT"
] | 2 |
2021-12-02T14:33:11.000Z
|
2021-12-23T17:30:09.000Z
| 65.49825 | 277 | 0.302808 |
[
[
[
"# 2. Modelling SVR Linear\n\n---",
"_____no_output_____"
]
],
[
[
"## load modules and run mlflow_logging.ipynb to get function to track model information on MLFLow\nimport sys\nsys.path.append(\"..\")\nimport numpy as np\nimport pandas as pd\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.metrics import mean_squared_error, r2_score\nfrom sklearn.svm import LinearSVR\nimport seaborn as sns\nfrom sklearn.preprocessing import MinMaxScaler, StandardScaler\n\nfrom sklearn.model_selection import GridSearchCV\n\nfrom modeling.functions import modelling, log_to_mlflow, get_features, save_models, save_results",
"_____no_output_____"
],
[
"data = pd.read_csv('../data/GEFCom2014Data/Wind/raw_data_incl_features.csv', parse_dates=['TIMESTAMP'])\ndata.head()\ndata.dropna(inplace=True)\ndata.info()\nRSEED = 42",
"<class 'pandas.core.frame.DataFrame'>\nInt64Index: 175265 entries, 0 to 175433\nData columns (total 19 columns):\n # Column Non-Null Count Dtype \n--- ------ -------------- ----- \n 0 ZONEID 175265 non-null int64 \n 1 TIMESTAMP 175265 non-null datetime64[ns]\n 2 TARGETVAR 175265 non-null float64 \n 3 U10 175265 non-null float64 \n 4 V10 175265 non-null float64 \n 5 U100 175265 non-null float64 \n 6 V100 175265 non-null float64 \n 7 HOUR 175265 non-null int64 \n 8 MONTH 175265 non-null int64 \n 9 WEEKDAY 175265 non-null int64 \n 10 IS_HOLIDAY 175265 non-null int64 \n 11 WS10 175265 non-null float64 \n 12 WS100 175265 non-null float64 \n 13 WD10 175265 non-null float64 \n 14 WD100 175265 non-null float64 \n 15 WD100CARD 175265 non-null object \n 16 WD10CARD 175265 non-null object \n 17 U100NORM 175265 non-null float64 \n 18 V100NORM 175265 non-null float64 \ndtypes: datetime64[ns](1), float64(11), int64(5), object(2)\nmemory usage: 26.7+ MB\n"
],
[
"data = pd.get_dummies(data, columns = ['WD100CARD','WD10CARD'])\ndata.head()",
"_____no_output_____"
],
[
"## train-test-split\n#data_train, data_test = train_test_split(data, test_size=0.25, random_state=RSEED, stratify=data.ZONEID)\ndata_train = data[data.TIMESTAMP <= '2013-07-01 00:00:00']\ndata_test = data[data.TIMESTAMP > '2013-07-01 00:00:00']",
"_____no_output_____"
],
[
"# define features and feature dict\nfeature_dict = get_features(data)\n\nfeatures = feature_dict['all']",
"_____no_output_____"
],
[
"# define zone\nzone = 1",
"_____no_output_____"
],
[
"# split train and test data in feature and TARGETVAR parts and cut data to desired zones\nX_train = data_train[data_train.ZONEID == zone][features]\ny_train = data_train[data_train.ZONEID == zone].TARGETVAR\n\nX_test = data_test[data_test.ZONEID == zone][features]\ny_test = data_test[data_test.ZONEID == zone].TARGETVAR",
"_____no_output_____"
],
[
"# Scale data\nscaler = MinMaxScaler()\nX_train = scaler.fit_transform(X_train)\nX_test = scaler.transform(X_test)",
"_____no_output_____"
],
[
"model = LinearSVR()\n\nparam_grid = {'C': [0.01, 0.1, 1, 10, 100]}\n\n# cv = GridSearchCV(model, param_grid=param_grid)\n# cv.fit(X_train, y_train)",
"_____no_output_____"
],
[
"#y_pred = cv.predict(X_test)\n\n#print(mean_squared_error(y_test, y_pred, squared=False))",
"_____no_output_____"
],
[
"model = LinearSVR(max_iter=100000)\nscaler = MinMaxScaler()\n\nmodel_dict = {}\nresults_train = {}\nresults_test = {}\nparam_grid = {'C': [0.1, 1, 10]}\n\nfor key in feature_dict.keys():\n print(f'Features: {key}')\n results_train[key],results_test[key], model_dict[key] = modelling(data_train, data_test, feature_dict[key], model = model, scaler=scaler, print_scores=True, log=None, infotext_mlflow=None, save_model= True, perform_gridCV= True, param_grid= param_grid, n_jobs = 3)\n results_test[key] = {k : np.round(value,5) for k,value in results_test[key].items()}",
"Features: all\nScaler: MinMaxScaler\nScaled X_train min/max: 0.0, 1.0\nScaled X_test min/max: -0.0, 1.0\n\nZONEID 1\nFitting 5 folds for each of 3 candidates, totalling 15 fits\n"
],
[
"results_test\n#save_models(model_dict)\nsave_results(results_train, results_test, '../saved_models/211203_1152_LinearSVR')",
"_____no_output_____"
],
[
"def save_scores(results_dict):\n pass\n\n\n\n\nfeatures = []\nzones = []\ntrain_score = []\ntest_score = []\n\nfor key in results_train.keys():\n for zone in results_train[key].keys():\n features.append(key)\n zones.append(zone)\n train_score.append(results_train[key][zone])\n test_score.append(results_test[key][zone])\n\ndf = pd.DataFrame({'features':features,'zone': zones,'train_score': train_score,'test_score': test_score})\n\ndf.to_csv(path + '/' + file_name, index=False)\n\n",
"_____no_output_____"
],
[
"df",
"_____no_output_____"
],
[
"df.zone.unique()",
"_____no_output_____"
],
[
"path = '../saved_models/211201_0915_GridSearchCV'\n\nfile_name = path.split('/')[-1] + '.csv'\n\nfile_name",
"_____no_output_____"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab7f8db29ae64faf5351b206b7e9e97e2e5679d
| 57,943 |
ipynb
|
Jupyter Notebook
|
KNN Classifier On Iris Dataset.ipynb
|
PrasoonThakur5/KNN-Classifier-On-Iris-Dataset
|
8aec5b2d6155ae5edc107b85bc9f4e8af0c75093
|
[
"Apache-2.0"
] | null | null | null |
KNN Classifier On Iris Dataset.ipynb
|
PrasoonThakur5/KNN-Classifier-On-Iris-Dataset
|
8aec5b2d6155ae5edc107b85bc9f4e8af0c75093
|
[
"Apache-2.0"
] | null | null | null |
KNN Classifier On Iris Dataset.ipynb
|
PrasoonThakur5/KNN-Classifier-On-Iris-Dataset
|
8aec5b2d6155ae5edc107b85bc9f4e8af0c75093
|
[
"Apache-2.0"
] | null | null | null | 55.341929 | 15,368 | 0.707247 |
[
[
[
"import numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport seaborn as sns",
"_____no_output_____"
],
[
"data=sns.load_dataset(\"iris\")",
"_____no_output_____"
],
[
"data.head()",
"_____no_output_____"
],
[
"data.drop(columns=['sepal_width'],inplace=True)",
"_____no_output_____"
],
[
"data.head()",
"_____no_output_____"
],
[
"data.iloc[:,0:2].values",
"_____no_output_____"
],
[
"data.iloc[:,-1].values",
"_____no_output_____"
],
[
"species=data['species'].replace({'setosa':1,'versicolor':2,'virginica':3},inplace=True)",
"_____no_output_____"
],
[
"data.iloc[:,-1].values",
"_____no_output_____"
],
[
"x=data.iloc[:,0:2].values",
"_____no_output_____"
],
[
"y=data.iloc[:,-1].values",
"_____no_output_____"
],
[
"x.shape",
"_____no_output_____"
],
[
"y.shape",
"_____no_output_____"
],
[
"from sklearn.model_selection import train_test_split\nx_train,x_test,y_train,y_test=train_test_split(x,y,test_size=0.20)",
"_____no_output_____"
],
[
"x_train.shape",
"_____no_output_____"
],
[
"x_test.shape",
"_____no_output_____"
],
[
"np.sqrt(x_train.shape[0])",
"_____no_output_____"
],
[
"k=25",
"_____no_output_____"
],
[
"from sklearn.neighbors import KNeighborsClassifier\nknn=KNeighborsClassifier(n_neighbors=k)",
"_____no_output_____"
],
[
"knn.fit(x_train,y_train)",
"_____no_output_____"
],
[
"y_pred=knn.predict(x_test)",
"_____no_output_____"
],
[
"y_pred",
"_____no_output_____"
],
[
"y_test",
"_____no_output_____"
],
[
"y_test.shape",
"_____no_output_____"
],
[
"from sklearn.metrics import accuracy_score\naccuracy_score(y_test,y_pred)",
"_____no_output_____"
],
[
"from sklearn.metrics import confusion_matrix\nconfusion_matrix(y_test,y_pred)",
"_____no_output_____"
],
[
"accuracy=[]\nfor i in range(1,100):\n knn=KNeighborsClassifier(n_neighbors=i)\n knn.fit(x_train,y_train)\n accuracy.append(accuracy_score(y_test,knn.predict(x_test)))",
"_____no_output_____"
],
[
"len(accuracy)",
"_____no_output_____"
],
[
"plt.plot(range(1,100),accuracy)",
"_____no_output_____"
],
[
"knn=KNeighborsClassifier(n_neighbors=40)\nknn.fit(x_train,y_train)",
"_____no_output_____"
],
[
"y_pred=knn.predict(x_test)",
"_____no_output_____"
],
[
"accuracy_score(y_test,y_pred)",
"_____no_output_____"
],
[
"def predict_species():\n sepal=float(input(\"Enter The Sepal Length\"))\n petal=float(input(\"Enter The Petal Length\"))\n \n x_new=np.array([[sepal],[petal]]).reshape(1,2)\n \n if knn.predict(x_new)[0]==1:\n return \"The Species Is Setosa\"\n elif knn.predict(x_new)[0]==2:\n return \"The Species Is Versicolor\"\n \n else:\n return \"The Species Is Virginica\"",
"_____no_output_____"
],
[
"predict_species()",
"Enter The Sepal Length5.1\nEnter The Petal Length1.4\n"
]
],
[
[
"# Decision Boundary",
"_____no_output_____"
]
],
[
[
"a=np.arange(start=x_train[:,0].min()-1,stop=x_train[:,0].max()+1,step=0.01)\nb=np.arange(start=x_train[:,1].min()-1,stop=x_train[:,1].max()+1,step=0.01)\n\nxx,yy=np.meshgrid(a,b)",
"_____no_output_____"
],
[
"input_array=np.array([xx.ravel(),yy.ravel()]).T\nlabels=knn.predict(input_array)",
"_____no_output_____"
],
[
"plt.contourf(xx,yy,labels.reshape(xx.shape))",
"_____no_output_____"
],
[
"plt.contourf(xx,yy,labels.reshape(xx.shape))\nplt.scatter(x_train[:,0],x_train[:,1], c=y_train)",
"_____no_output_____"
]
]
] |
[
"code",
"markdown",
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
4ab7fbebc5a53f14e86dcbc0a4c552c70689d604
| 45,525 |
ipynb
|
Jupyter Notebook
|
hw1/.ipynb_checkpoints/ MIDS-W261-2015-HWK-Week01-AlejandroRojas-checkpoint.ipynb
|
venamax/ucb-261
|
7df292668a971f49f8a099d38f899e9961432e7c
|
[
"MIT-0",
"Apache-2.0",
"BSD-2-Clause",
"BSD-3-Clause"
] | 1 |
2022-01-05T14:27:20.000Z
|
2022-01-05T14:27:20.000Z
|
hw1/.ipynb_checkpoints/ MIDS-W261-2015-HWK-Week01-AlejandroRojas-checkpoint.ipynb
|
venamax/ucb-261
|
7df292668a971f49f8a099d38f899e9961432e7c
|
[
"MIT-0",
"Apache-2.0",
"BSD-2-Clause",
"BSD-3-Clause"
] | null | null | null |
hw1/.ipynb_checkpoints/ MIDS-W261-2015-HWK-Week01-AlejandroRojas-checkpoint.ipynb
|
venamax/ucb-261
|
7df292668a971f49f8a099d38f899e9961432e7c
|
[
"MIT-0",
"Apache-2.0",
"BSD-2-Clause",
"BSD-3-Clause"
] | null | null | null | 45.298507 | 883 | 0.436068 |
[
[
[
"<h1> <b>Homework 1</b></h1>\n<i>Alejandro J. Rojas<br>\[email protected]<br>\nW261: Machine Learning at Scale<br>\nWeek: 01<br>\nJan 21, 2016</i></li>",
"_____no_output_____"
],
[
"<h2>HW1.0.0.</h2> Define big data. Provide an example of a big data problem in your domain of expertise. ",
"_____no_output_____"
],
[
"The term big data is asoociated to datasets that cannot be processed, stored and transformed using traditional applications and tools because of high volume, high velocity and high variety. By high volume, we mean datasets that not only require high storage capacity, usually beyond 1T, but also datasets that are too big to have a decent processing and thoughput time. By high velocity, we mean data that requires real-time processing with throughput speeds that can be bursty. High variety includes data that comes from different formats, some structured some that are not, that all need to be ingested and transformed to be processed. Big data is simply changing the way we used to collect and analyze data at the time that it opens opportunity to increase the scale, scope and intimacy of the analyses that we are now able to do.\n\nThe social web is a leading source of big data applications given our ability to log almost anything that the user does when interacting to an application. In my field, I've seen how online videos are increasingly the way users consume media. A video, per se, is an unstructured data item and its interactions are usually captured by leading social media platforms like Facebook, Twitter and Youtube in the form of JSON, a semi unstructured format that can capture user interactions such as likes, shares and comments. Across the internet, the amount of videos being upload and downstream is exploding making it a challenge to measure real-time, the media consuming habits of our target users. Big data can help in providing insights from all of this information so that we can better predict the taste of users visiting our site properties to serve them content they like. \n",
"_____no_output_____"
],
[
"<h2>HW1.0.1.</h2>In 500 words (English or pseudo code or a combination) describe how to estimate the bias, the variance, the irreduciable error for a test dataset T when using polynomial regression models of degree 1, 2,3, 4,5 are considered. How would you select a model?",
"_____no_output_____"
],
[
"For any dataset T that contains n independent variables (x1, x2, ..xn) and one dependent variable y_true, we can observe the following:\n\nIf we try to estimate y as a function of x:\n \n y_pred = f(x)\n\nThe estimate of our function will produce an error shown as:\n\n <img src=\"error.png\">\n\n\nThis error varies as we increase the complexity of our models as the following chart shows:\n\n<img src=\"Prediction-Error.png\">\n\n\nThe source of this error can be divided into three types:\n \n bias \n variance\n irreducible error\n\nand can be derived mathematically the following way\n\n<img src=\"mathematicalerrors.jpg\">\n\nBias error is introduced by us when we try to simplify the dynamics that we observe in the data, for instace by using a linear function to estimate y.\n\nAs we try to better fit the underlying data, we can try implementing nonlinear functions.\n\nAs the order of the polynomial regression increases, our function f(x) will more closely match the underlying portion of the dataset T and consequently we reduced our bias error.\n\nHowever, if we randomly applied our high-ordered polynomial f(x) to another portion of dataset T, we will find that our error will increase because we introduced variance error by overfitting the prior dataset.\n\nSo as a rule of thumb, we can say that\n\nas the degree of the predictive polynomial function f(x) increases:\n\n bias error is reduced\n variance error is increased\n \nthe trick is to find the optimal point where the sum of these two errors are at the minimum.Even at that point, our function(x) will still show some error that will be irreducible because it comes from imprecisions in the way data was collected or other type of noise present in the dataset T.\n\n\nIn this chart you can see how each of these errors varies as we bootstrap 50 samples of dataset T:\n\n<img src=\"bootstrapping.jpg\">\n\n\n\n\n\n\n",
"_____no_output_____"
],
[
"<h2> HW1.1.</h2> Read through the provided control script (pNaiveBayes.sh)\n and all of its comments. When you are comfortable with their\n purpose and function, respond to the remaining homework questions below. \n A simple cell in the notebook with a print statmement with a \"done\" string will suffice here. (dont forget to include the Question Number and the quesition in the cell as a multiline comment!)",
"_____no_output_____"
],
[
"# <----------------------------------End of HW1.1------------------------------------->",
"_____no_output_____"
],
[
"<h2>HW1.2.</h2>Provide a mapper/reducer pair that, when executed by pNaiveBayes.sh\n will determine the number of occurrences of a single, user-specified word. Examine the word “assistance” and report your results.",
"_____no_output_____"
],
[
"# Map",
"_____no_output_____"
]
],
[
[
"%%writefile mapper.py\n#!/usr/bin/python\n## mapper.py\n## Author: Alejandro J. Rojas\n## Description: mapper code for HW1.2-1.5\n\nimport sys\nimport re\ncount = 0\nrecords = 0\nwords = 0\n\n## collect user input\nfilename = sys.argv[1]\nfindwords = re.split(\" \",sys.argv[2].lower())\n\nwith open (filename, \"r\") as myfile:\n for line in myfile.readlines():\n record = re.split(r'\\t+', line)\n records = records + 1\n for i in range (len(record)):\n bagofwords = re.split(\" \",record[i]) ### Break each email records into words\n for word in bagofwords:\n words = words + 1\n for keyword in findwords:\n if keyword in word:\n count = count + 1 ### Add one the count of found words\n \n\n##print '# of Records analized',records\n##print '# of Words analized', words\n##print '# of Ocurrences', count\nprint count",
"Overwriting mapper.py\n"
],
[
"!chmod +x mapper.py",
"_____no_output_____"
]
],
[
[
"# Reduce",
"_____no_output_____"
]
],
[
[
"%%writefile reducer.py\n#!/usr/bin/python\n## reducer.py\n## Author: Alejandro J. Rojas\n## Description: reducer code for HW1.2\n\nimport sys\nimport re\nsum = 0\n\n## collect user input\nfilenames = sys.argv[1:]\nfor file in filenames:\n with open (file, \"r\") as myfile:\n for line in myfile.readlines():\n if line.strip():\n sum = sum + int(line) ### Add counts present on all mapper produced files\n \nprint sum",
"Overwriting reducer.py\n"
],
[
"!chmod +x reducer.py",
"_____no_output_____"
]
],
[
[
"# Write script to file",
"_____no_output_____"
]
],
[
[
"%%writefile pNaiveBayes.sh\n## pNaiveBayes.sh\n## Author: Jake Ryland Williams\n## Usage: pNaiveBayes.sh m wordlist\n## Input:\n## m = number of processes (maps), e.g., 4\n## wordlist = a space-separated list of words in quotes, e.g., \"the and of\"\n##\n## Instructions: Read this script and its comments closely.\n## Do your best to understand the purpose of each command,\n## and focus on how arguments are supplied to mapper.py/reducer.py,\n## as this will determine how the python scripts take input.\n## When you are comfortable with the unix code below,\n## answer the questions on the LMS for HW1 about the starter code.\n\n## collect user input\nm=$1 ## the number of parallel processes (maps) to run\nwordlist=$2 ## if set to \"*\", then all words are used\n\n## a test set data of 100 messages\ndata=\"enronemail_1h.txt\" \n\n## the full set of data (33746 messages)\n# data=\"enronemail.txt\" \n\n## 'wc' determines the number of lines in the data\n## 'perl -pe' regex strips the piped wc output to a number\nlinesindata=`wc -l $data | perl -pe 's/^.*?(\\d+).*?$/$1/'`\n\n## determine the lines per chunk for the desired number of processes\nlinesinchunk=`echo \"$linesindata/$m+1\" | bc`\n\n## split the original file into chunks by line\nsplit -l $linesinchunk $data $data.chunk.\n\n## assign python mappers (mapper.py) to the chunks of data\n## and emit their output to temporary files\nfor datachunk in $data.chunk.*; do\n ## feed word list to the python mapper here and redirect STDOUT to a temporary file on disk\n ####\n ####\n ./mapper.py $datachunk \"$wordlist\" > $datachunk.counts &\n ####\n ####\ndone\n## wait for the mappers to finish their work\nwait\n\n## 'ls' makes a list of the temporary count files\n## 'perl -pe' regex replaces line breaks with spaces\ncountfiles=`\\ls $data.chunk.*.counts | perl -pe 's/\\n/ /'`\n\n## feed the list of countfiles to the python reducer and redirect STDOUT to disk\n####\n####\n./reducer.py $countfiles > $data.output\n####\n####\nnumOfInstances=$(cat $data.output)\necho \"found [$numOfInstances] [$wordlist]\" ## Report how many were found\n## clean up the data chunks and temporary count files\n\\rm $data.chunk.*",
"Overwriting pNaiveBayes.sh\n"
],
[
"!chmod a+x pNaiveBayes.sh",
"_____no_output_____"
]
],
[
[
"# Run file",
"_____no_output_____"
],
[
"Usage: usage: pNaiveBayes.sh m wordlist",
"_____no_output_____"
]
],
[
[
"!./pNaiveBayes.sh 5 \"assistance\"",
"found [10] [assistance]\r\n"
]
],
[
[
"# <----------------------------------End of HW1.2------------------------------------->",
"_____no_output_____"
],
[
"<h2>HW1.3.</h2> Provide a mapper/reducer pair that, when executed by pNaiveBayes.sh\n will classify the email messages by a single, user-specified word using the multinomial Naive Bayes Formulation. Examine the word “assistance” and report your results. ",
"_____no_output_____"
],
[
"# Map",
"_____no_output_____"
]
],
[
[
"%%writefile mapper.py\n#!/usr/bin/python\n## mapper.py\n## Author: Alejandro J. Rojas\n## Description: mapper code for HW1.3\n\nimport sys\nimport re\n\n\n\n########## Collect user input ###############\nfilename = sys.argv[1]\nfindwords = re.split(\" \",sys.argv[2].lower())\n\n\n\n\nwith open (filename, \"r\") as myfile:\n\n for line in myfile.readlines():\n record = re.split(r'\\t+', line) ### Each email is a record with 4 components\n ### 1) ID 2) Spam Truth 3) Subject 4) Content\n if len(record)==4: ### Take only complete records\n\n ########## Variables to collect and measure #########\n records = 0 ### Each record corresponds to a unique email\n words = 0 ### Words written in all emails incluidng Subject \n spam_records, spam_words, spam_count = 0,0,0 ### Spam email count, words in spam email, user-specified word count\n ham_records, ham_words, ham_count = 0, 0, 0 ### Same as above but for not spam emails\n\n\n records += 1 ### add one the the total sum of emails\n if int(record[1]) == 1: ### If the email is labeled as spam\n spam_records += 1 ### add one to the email spam count\n for i in range (2,len(record)): ### Starting from Subject to the Content \n bagofwords = re.split(\" \",record[i]) ### Collect all words present on each email \n for word in bagofwords: ### For each word\n words += 1 ### add one to the total sum of words\n spam_words += 1 ### add one to the total sum of spam words \n for keyword in findwords: ### for each word specified by user\n if keyword in word: ### If there's a match then\n spam_count += 1 ### add one to the user specified word count as spam\n \n else: ### If email is not labeled as spam\n ham_records +=1 ### add one to the email ham count\n for i in range (2,len(record)): ### Starting from Subject to the Content \n bagofwords = re.split(\" \",record[i]) ### Collect all words present on each email \n for word in bagofwords: ### For each word\n words += 1 ### add one to the total sum of words\n ham_words += 1 ### add one to the total sum of ham words \n for keyword in findwords: ### for each word specified by user\n if keyword in word: ### If there's a match then\n ham_count += 1 ### add one to the user specified word count as ham\n \n record_id = record[0]\n truth = record[1]\n print spam_count, \" \", spam_words, \" \", spam_records, \" \", \\\n ham_count, \" \", ham_words, \" \", ham_records, \" \", \\\n words, \" \", records, \" \", record_id, \" \", truth ",
"Overwriting mapper.py\n"
],
[
"!chmod +x mapper.py",
"_____no_output_____"
]
],
[
[
"# Reduce",
"_____no_output_____"
]
],
[
[
"%%writefile reducer.py\n#!/usr/bin/python\n## reducer.py\n## Author: Alejandro J. Rojas\n## Description: reducer code for HW1.3-1.4\n\nimport sys\nimport re\nsum_spam_records, sum_spam_words, sum_spam_count = 0,0,0\nsum_ham_records, sum_ham_words, sum_ham_count = 0,0,0\nsum_records,sum_words = 0,0\n\n## collect user input\nfilenames = sys.argv[1:]\nfor file in filenames:\n with open (file, \"r\") as myfile:\n for line in myfile.readlines():\n if line.strip():\n factors = re.split(\" \", line)\n sum_spam_count += int(factors[0]) ## sum up every time the word was found in a spam\n sum_spam_words += int(factors[3]) ## sum up all words from spams\n sum_spam_records+= int(factors[6]) ## sum up all emails labeled as spam\n sum_ham_count += int(factors[9]) ## sum up every time the word was found in a ham\n sum_ham_words += int(factors[12]) ## sum up all words from hams\n sum_ham_records += int(factors[15]) ## sum up all emails labeled as ham\n sum_words += int(factors[18]) ## sum all words from all emails\n sum_records += int(factors[21]) ## sum all emails\n \n\nprior_spam = float(sum_spam_records)/float(sum_records) ## prior prob of a spam email\nprior_ham = float(sum_ham_records)/float(sum_records) ## prior prob of a ham email\nprob_word_spam = float(sum_spam_count)/float(sum_spam_words)## prob of word given that email is spam\nprob_word_ham = float(sum_ham_count)/float(sum_ham_words) ## prob of word given that email is ham\n\n##check_prior = prior_spam + prior_ham ## check priors -> sum to 1\n##check_words = float(sum_words)/float(sum_spam_words+sum_ham_words) ## check probabilities of a word -> sum to 1\n##check_spam = prob_word_spam*float(sum_spam_words)/float(sum_spam_count) ## check spam counts -> sum to 1\n##check_ham = prob_word_ham*float(sum_ham_words)/float(sum_ham_count) ## check ham count -> sum to 1\nsum_count = sum_spam_count+sum_ham_count\n\nprint \"Summary of Data\"\nprint '%4s'%sum_records ,'emails examined, containing %6s'%sum_words, 'words, we found %3s'%sum_count ,'matches.'\n\nprint '%30s' %'ID', '%10s' %'TRUTH', '%10s' %'CLASS', '%20s' %'CUMULATIVE ACCURACY'\nmiss, sample_size = 0,0 \nfor file in filenames: \n with open (file, \"r\") as myfile:\n for line in myfile.readlines():\n if line.strip():\n data = re.split(\" \", line)\n record_id = data[24]\n y_true = int(data[27][0])\n count = int(data[0]) + int(data[9])\n p_spam = prior_spam*prob_word_spam**count\n p_ham = prior_ham*prob_word_ham**count\n if p_spam > p_ham:\n y_pred = 1\n else:\n y_pred = 0\n \n if y_pred != y_true:\n miss+= 1.0\n sample_size += 1.0\n accuracy = ((sample_size-miss)/sample_size)*100\n \n print '%30s' %record_id, '%10s' %y_true, '%10s' %y_pred, '%18.2f %%' % accuracy\n \n ",
"Overwriting reducer.py\n"
],
[
"!chmod +x reducer.py",
"_____no_output_____"
]
],
[
[
"# Write script to file",
"_____no_output_____"
]
],
[
[
"%%writefile pNaiveBayes.sh\n## pNaiveBayes.sh\n## Author: Jake Ryland Williams\n## Usage: pNaiveBayes.sh m wordlist\n## Input:\n## m = number of processes (maps), e.g., 4\n## wordlist = a space-separated list of words in quotes, e.g., \"the and of\"\n##\n## Instructions: Read this script and its comments closely.\n## Do your best to understand the purpose of each command,\n## and focus on how arguments are supplied to mapper.py/reducer.py,\n## as this will determine how the python scripts take input.\n## When you are comfortable with the unix code below,\n## answer the questions on the LMS for HW1 about the starter code.\n\n## collect user input\nm=$1 ## the number of parallel processes (maps) to run\nwordlist=$2 ## if set to \"*\", then all words are used\n\n## a test set data of 100 messages\ndata=\"enronemail_1h.txt\" \n\n## the full set of data (33746 messages)\n# data=\"enronemail.txt\" \n\n## 'wc' determines the number of lines in the data\n## 'perl -pe' regex strips the piped wc output to a number\nlinesindata=`wc -l $data | perl -pe 's/^.*?(\\d+).*?$/$1/'`\n\n## determine the lines per chunk for the desired number of processes\nlinesinchunk=`echo \"$linesindata/$m+1\" | bc`\n\n## split the original file into chunks by line\nsplit -l $linesinchunk $data $data.chunk.\n\n## assign python mappers (mapper.py) to the chunks of data\n## and emit their output to temporary files\nfor datachunk in $data.chunk.*; do\n ## feed word list to the python mapper here and redirect STDOUT to a temporary file on disk\n ####\n ####\n ./mapper.py $datachunk \"$wordlist\" > $datachunk.counts &\n ####\n ####\ndone\n## wait for the mappers to finish their work\nwait\n\n## 'ls' makes a list of the temporary count files\n## 'perl -pe' regex replaces line breaks with spaces\ncountfiles=`\\ls $data.chunk.*.counts | perl -pe 's/\\n/ /'`\n\n## feed the list of countfiles to the python reducer and redirect STDOUT to disk\n####\n####\n./reducer.py $countfiles > $data.output\n####\n####\nnumOfInstances=$(cat $data.output)\necho \"NB Classifier based on word(s): $wordlist\" ## Print out words \necho \"$numOfInstances\" ## Print out output data\n## clean up the data chunks and temporary count files\n\\rm $data.chunk.*",
"Overwriting pNaiveBayes.sh\n"
]
],
[
[
"# Run file",
"_____no_output_____"
]
],
[
[
"!./pNaiveBayes.sh 5 \"assistance\"",
"NB Classifier based on word(s): assistance\r\nSummary of Data\r\n 98 emails examined, containing 35352 words, we found 9 matches.\r\n ID TRUTH CLASS CUMULATIVE ACCURACY\r\n 0001.1999-12-10.farmer 0 0 100.00 %\r\n 0001.1999-12-10.kaminski 0 0 100.00 %\r\n 0001.2000-01-17.beck 0 0 100.00 %\r\n 0001.2001-02-07.kitchen 0 0 100.00 %\r\n 0001.2001-04-02.williams 0 0 100.00 %\r\n 0002.1999-12-13.farmer 0 0 100.00 %\r\n 0002.2001-02-07.kitchen 0 0 100.00 %\r\n 0002.2001-05-25.SA_and_HP 1 0 87.50 %\r\n 0002.2003-12-18.GP 1 0 77.78 %\r\n 0002.2004-08-01.BG 1 1 80.00 %\r\n 0003.1999-12-10.kaminski 0 0 81.82 %\r\n 0003.1999-12-14.farmer 0 0 83.33 %\r\n 0003.2000-01-17.beck 0 0 84.62 %\r\n 0003.2001-02-08.kitchen 0 0 85.71 %\r\n 0003.2003-12-18.GP 1 0 80.00 %\r\n 0003.2004-08-01.BG 1 0 75.00 %\r\n 0004.1999-12-10.kaminski 0 1 70.59 %\r\n 0004.1999-12-14.farmer 0 0 72.22 %\r\n 0004.2001-04-02.williams 0 0 73.68 %\r\n 0004.2001-06-12.SA_and_HP 1 0 70.00 %\r\n 0004.2004-08-01.BG 1 0 66.67 %\r\n 0005.1999-12-12.kaminski 0 1 63.64 %\r\n 0005.1999-12-14.farmer 0 0 65.22 %\r\n 0005.2000-06-06.lokay 0 0 66.67 %\r\n 0005.2001-02-08.kitchen 0 0 68.00 %\r\n 0005.2001-06-23.SA_and_HP 1 0 65.38 %\r\n 0005.2003-12-18.GP 1 0 62.96 %\r\n 0006.1999-12-13.kaminski 0 0 64.29 %\r\n 0006.2001-02-08.kitchen 0 0 65.52 %\r\n 0006.2001-04-03.williams 0 0 66.67 %\r\n 0006.2001-06-25.SA_and_HP 1 0 64.52 %\r\n 0006.2003-12-18.GP 1 0 62.50 %\r\n 0006.2004-08-01.BG 1 0 60.61 %\r\n 0007.1999-12-13.kaminski 0 0 61.76 %\r\n 0007.1999-12-14.farmer 0 0 62.86 %\r\n 0007.2000-01-17.beck 0 0 63.89 %\r\n 0007.2001-02-09.kitchen 0 0 64.86 %\r\n 0007.2003-12-18.GP 1 0 63.16 %\r\n 0007.2004-08-01.BG 1 0 61.54 %\r\n 0008.2001-02-09.kitchen 0 0 62.50 %\r\n 0008.2001-06-12.SA_and_HP 1 0 60.98 %\r\n 0008.2001-06-25.SA_and_HP 1 0 59.52 %\r\n 0008.2003-12-18.GP 1 0 58.14 %\r\n 0008.2004-08-01.BG 1 0 56.82 %\r\n 0009.1999-12-13.kaminski 0 0 57.78 %\r\n 0009.1999-12-14.farmer 0 0 58.70 %\r\n 0009.2000-06-07.lokay 0 0 59.57 %\r\n 0009.2001-02-09.kitchen 0 0 60.42 %\r\n 0009.2003-12-18.GP 1 0 59.18 %\r\n 0010.1999-12-14.farmer 0 0 60.00 %\r\n 0010.1999-12-14.kaminski 0 0 60.78 %\r\n 0010.2001-02-09.kitchen 0 0 61.54 %\r\n 0010.2001-06-28.SA_and_HP 1 1 62.26 %\r\n 0010.2003-12-18.GP 1 0 61.11 %\r\n 0010.2004-08-01.BG 1 0 60.00 %\r\n 0011.1999-12-14.farmer 0 0 60.71 %\r\n 0011.2001-06-28.SA_and_HP 1 0 59.65 %\r\n 0011.2001-06-29.SA_and_HP 1 0 58.62 %\r\n 0011.2003-12-18.GP 1 0 57.63 %\r\n 0011.2004-08-01.BG 1 0 56.67 %\r\n 0012.1999-12-14.farmer 0 0 57.38 %\r\n 0012.1999-12-14.kaminski 0 0 58.06 %\r\n 0012.2000-01-17.beck 0 0 58.73 %\r\n 0012.2000-06-08.lokay 0 0 59.38 %\r\n 0012.2001-02-09.kitchen 0 0 60.00 %\r\n 0012.2003-12-19.GP 1 0 59.09 %\r\n 0013.1999-12-14.farmer 0 0 59.70 %\r\n 0013.1999-12-14.kaminski 0 0 60.29 %\r\n 0013.2001-04-03.williams 0 0 60.87 %\r\n 0013.2001-06-30.SA_and_HP 1 0 60.00 %\r\n 0013.2004-08-01.BG 1 1 60.56 %\r\n 0014.1999-12-14.kaminski 0 0 61.11 %\r\n 0014.1999-12-15.farmer 0 0 61.64 %\r\n 0014.2001-02-12.kitchen 0 0 62.16 %\r\n 0014.2001-07-04.SA_and_HP 1 0 61.33 %\r\n 0014.2003-12-19.GP 1 0 60.53 %\r\n 0014.2004-08-01.BG 1 0 59.74 %\r\n 0015.1999-12-14.kaminski 0 0 60.26 %\r\n 0015.1999-12-15.farmer 0 0 60.76 %\r\n 0015.2000-06-09.lokay 0 0 61.25 %\r\n 0015.2001-02-12.kitchen 0 0 61.73 %\r\n 0015.2001-07-05.SA_and_HP 1 0 60.98 %\r\n 0015.2003-12-19.GP 1 0 60.24 %\r\n 0016.1999-12-15.farmer 0 0 60.71 %\r\n 0016.2001-02-12.kitchen 0 0 61.18 %\r\n 0016.2001-07-05.SA_and_HP 1 0 60.47 %\r\n 0016.2001-07-06.SA_and_HP 1 0 59.77 %\r\n 0016.2003-12-19.GP 1 0 59.09 %\r\n 0016.2004-08-01.BG 1 0 58.43 %\r\n 0017.1999-12-14.kaminski 0 0 58.89 %\r\n 0017.2000-01-17.beck 0 0 59.34 %\r\n 0017.2001-04-03.williams 0 0 59.78 %\r\n 0017.2003-12-18.GP 1 0 59.14 %\r\n 0017.2004-08-01.BG 1 0 58.51 %\r\n 0017.2004-08-02.BG 1 0 57.89 %\r\n 0018.1999-12-14.kaminski 0 0 58.33 %\r\n 0018.2001-07-13.SA_and_HP 1 1 58.76 %\r\n 0018.2003-12-18.GP 1 1 59.18 %\r\n"
]
],
[
[
"# <----------------------------------End of HW1.3------------------------------------->",
"_____no_output_____"
],
[
"<h2>HW1.4.</h2> Provide a mapper/reducer pair that, when executed by pNaiveBayes.sh\n will classify the email messages by a list of one or more user-specified words. Examine the words “assistance”, “valium”, and “enlargementWithATypo” and report your results",
"_____no_output_____"
],
[
"# Run file",
"_____no_output_____"
]
],
[
[
"!./pNaiveBayes.sh 5 \"assistance valium enlargementWithATypo\"",
"NB Classifier based on word(s): assistance valium enlargementWithATypo\r\nSummary of Data\r\n 98 emails examined, containing 35352 words, we found 12 matches.\r\n ID TRUTH CLASS CUMULATIVE ACCURACY\r\n 0001.1999-12-10.farmer 0 0 100.00 %\r\n 0001.1999-12-10.kaminski 0 0 100.00 %\r\n 0001.2000-01-17.beck 0 0 100.00 %\r\n 0001.2001-02-07.kitchen 0 0 100.00 %\r\n 0001.2001-04-02.williams 0 0 100.00 %\r\n 0002.1999-12-13.farmer 0 0 100.00 %\r\n 0002.2001-02-07.kitchen 0 0 100.00 %\r\n 0002.2001-05-25.SA_and_HP 1 0 87.50 %\r\n 0002.2003-12-18.GP 1 0 77.78 %\r\n 0002.2004-08-01.BG 1 1 80.00 %\r\n 0003.1999-12-10.kaminski 0 0 81.82 %\r\n 0003.1999-12-14.farmer 0 0 83.33 %\r\n 0003.2000-01-17.beck 0 0 84.62 %\r\n 0003.2001-02-08.kitchen 0 0 85.71 %\r\n 0003.2003-12-18.GP 1 0 80.00 %\r\n 0003.2004-08-01.BG 1 0 75.00 %\r\n 0004.1999-12-10.kaminski 0 1 70.59 %\r\n 0004.1999-12-14.farmer 0 0 72.22 %\r\n 0004.2001-04-02.williams 0 0 73.68 %\r\n 0004.2001-06-12.SA_and_HP 1 0 70.00 %\r\n 0004.2004-08-01.BG 1 0 66.67 %\r\n 0005.1999-12-12.kaminski 0 1 63.64 %\r\n 0005.1999-12-14.farmer 0 0 65.22 %\r\n 0005.2000-06-06.lokay 0 0 66.67 %\r\n 0005.2001-02-08.kitchen 0 0 68.00 %\r\n 0005.2001-06-23.SA_and_HP 1 0 65.38 %\r\n 0005.2003-12-18.GP 1 0 62.96 %\r\n 0006.1999-12-13.kaminski 0 0 64.29 %\r\n 0006.2001-02-08.kitchen 0 0 65.52 %\r\n 0006.2001-04-03.williams 0 0 66.67 %\r\n 0006.2001-06-25.SA_and_HP 1 0 64.52 %\r\n 0006.2003-12-18.GP 1 0 62.50 %\r\n 0006.2004-08-01.BG 1 0 60.61 %\r\n 0007.1999-12-13.kaminski 0 0 61.76 %\r\n 0007.1999-12-14.farmer 0 0 62.86 %\r\n 0007.2000-01-17.beck 0 0 63.89 %\r\n 0007.2001-02-09.kitchen 0 0 64.86 %\r\n 0007.2003-12-18.GP 1 0 63.16 %\r\n 0007.2004-08-01.BG 1 0 61.54 %\r\n 0008.2001-02-09.kitchen 0 0 62.50 %\r\n 0008.2001-06-12.SA_and_HP 1 0 60.98 %\r\n 0008.2001-06-25.SA_and_HP 1 0 59.52 %\r\n 0008.2003-12-18.GP 1 0 58.14 %\r\n 0008.2004-08-01.BG 1 0 56.82 %\r\n 0009.1999-12-13.kaminski 0 0 57.78 %\r\n 0009.1999-12-14.farmer 0 0 58.70 %\r\n 0009.2000-06-07.lokay 0 0 59.57 %\r\n 0009.2001-02-09.kitchen 0 0 60.42 %\r\n 0009.2003-12-18.GP 1 1 61.22 %\r\n 0010.1999-12-14.farmer 0 0 62.00 %\r\n 0010.1999-12-14.kaminski 0 0 62.75 %\r\n 0010.2001-02-09.kitchen 0 0 63.46 %\r\n 0010.2001-06-28.SA_and_HP 1 1 64.15 %\r\n 0010.2003-12-18.GP 1 0 62.96 %\r\n 0010.2004-08-01.BG 1 0 61.82 %\r\n 0011.1999-12-14.farmer 0 0 62.50 %\r\n 0011.2001-06-28.SA_and_HP 1 0 61.40 %\r\n 0011.2001-06-29.SA_and_HP 1 0 60.34 %\r\n 0011.2003-12-18.GP 1 0 59.32 %\r\n 0011.2004-08-01.BG 1 0 58.33 %\r\n 0012.1999-12-14.farmer 0 0 59.02 %\r\n 0012.1999-12-14.kaminski 0 0 59.68 %\r\n 0012.2000-01-17.beck 0 0 60.32 %\r\n 0012.2000-06-08.lokay 0 0 60.94 %\r\n 0012.2001-02-09.kitchen 0 0 61.54 %\r\n 0012.2003-12-19.GP 1 0 60.61 %\r\n 0013.1999-12-14.farmer 0 0 61.19 %\r\n 0013.1999-12-14.kaminski 0 0 61.76 %\r\n 0013.2001-04-03.williams 0 0 62.32 %\r\n 0013.2001-06-30.SA_and_HP 1 0 61.43 %\r\n 0013.2004-08-01.BG 1 1 61.97 %\r\n 0014.1999-12-14.kaminski 0 0 62.50 %\r\n 0014.1999-12-15.farmer 0 0 63.01 %\r\n 0014.2001-02-12.kitchen 0 0 63.51 %\r\n 0014.2001-07-04.SA_and_HP 1 0 62.67 %\r\n 0014.2003-12-19.GP 1 0 61.84 %\r\n 0014.2004-08-01.BG 1 0 61.04 %\r\n 0015.1999-12-14.kaminski 0 0 61.54 %\r\n 0015.1999-12-15.farmer 0 0 62.03 %\r\n 0015.2000-06-09.lokay 0 0 62.50 %\r\n 0015.2001-02-12.kitchen 0 0 62.96 %\r\n 0015.2001-07-05.SA_and_HP 1 0 62.20 %\r\n 0015.2003-12-19.GP 1 0 61.45 %\r\n 0016.1999-12-15.farmer 0 0 61.90 %\r\n 0016.2001-02-12.kitchen 0 0 62.35 %\r\n 0016.2001-07-05.SA_and_HP 1 0 61.63 %\r\n 0016.2001-07-06.SA_and_HP 1 0 60.92 %\r\n 0016.2003-12-19.GP 1 1 61.36 %\r\n 0016.2004-08-01.BG 1 0 60.67 %\r\n 0017.1999-12-14.kaminski 0 0 61.11 %\r\n 0017.2000-01-17.beck 0 0 61.54 %\r\n 0017.2001-04-03.williams 0 0 61.96 %\r\n 0017.2003-12-18.GP 1 0 61.29 %\r\n 0017.2004-08-01.BG 1 1 61.70 %\r\n 0017.2004-08-02.BG 1 0 61.05 %\r\n 0018.1999-12-14.kaminski 0 0 61.46 %\r\n 0018.2001-07-13.SA_and_HP 1 1 61.86 %\r\n 0018.2003-12-18.GP 1 1 62.24 %\r\n"
]
],
[
[
"# <----------------------------------End of HW1.4------------------------------------->",
"_____no_output_____"
],
[
"# <----------------------------------End of HW1------------------------------------->",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
]
] |
4ab7fd83fd12972bd5f7e869c90fc47aa4fede2a
| 278,025 |
ipynb
|
Jupyter Notebook
|
pytorch_example.ipynb
|
eladhoffer/colab-notebooks
|
c586853baadfd768182923d858edb3cae3ac134a
|
[
"MIT"
] | 1 |
2018-10-04T11:46:25.000Z
|
2018-10-04T11:46:25.000Z
|
pytorch_example.ipynb
|
eladhoffer/colab-notebooks
|
c586853baadfd768182923d858edb3cae3ac134a
|
[
"MIT"
] | null | null | null |
pytorch_example.ipynb
|
eladhoffer/colab-notebooks
|
c586853baadfd768182923d858edb3cae3ac134a
|
[
"MIT"
] | null | null | null | 122.531952 | 40,251 | 0.586192 |
[
[
[
"[View in Colaboratory](https://colab.research.google.com/github/eladhoffer/colab-notebooks/blob/master/pytorch_example.ipynb)",
"_____no_output_____"
],
[
"# Train using Pytorch with a free K80 GPU from Google\nThis is an example how to use one of repos to train Cifar10\n## using a K80 gpu\nchange \"Runtime->change runtime type->Hardware accelerator->GPU\" before running",
"_____no_output_____"
],
[
"# Install dependencies",
"_____no_output_____"
]
],
[
[
"import os\naccelerator = 'cu80' if os.path.exists('/opt/bin/nvidia-smi') else 'cpu'\nassert accelerator == 'cu80'\ntry:\n import torch\nexcept:\n !pip3 install http://download.pytorch.org/whl/cu80/torch-0.4.1-cp36-cp36m-linux_x86_64.whl\n\n!pip3 install torchvision\n!pip3 install bokeh",
"Collecting torch==0.4.1 from http://download.pytorch.org/whl/cu80/torch-0.4.1-cp36-cp36m-linux_x86_64.whl\n\u001b[?25l Downloading http://download.pytorch.org/whl/cu80/torch-0.4.1-cp36-cp36m-linux_x86_64.whl (483.0MB)\n\u001b[K 100% |████████████████████████████████| 483.0MB 2.3MB/s \ntcmalloc: large alloc 1073750016 bytes == 0x561b6000 @ 0x7fdc180dc2a4 0x594e17 0x626104 0x51190a 0x4f5277 0x510c78 0x5119bd 0x4f5277 0x4f3338 0x510fb0 0x5119bd 0x4f5277 0x4f3338 0x510fb0 0x5119bd 0x4f5277 0x4f3338 0x510fb0 0x5119bd 0x4f6070 0x510c78 0x5119bd 0x4f5277 0x4f3338 0x510fb0 0x5119bd 0x4f6070 0x4f3338 0x510fb0 0x5119bd 0x4f6070\n\u001b[?25hInstalling collected packages: torch\nSuccessfully installed torch-0.4.1\nCollecting torchvision\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/ca/0d/f00b2885711e08bd71242ebe7b96561e6f6d01fdb4b9dcf4d37e2e13c5e1/torchvision-0.2.1-py2.py3-none-any.whl (54kB)\n\u001b[K 100% |████████████████████████████████| 61kB 3.3MB/s \n\u001b[?25hRequirement already satisfied: six in /usr/local/lib/python3.6/dist-packages (from torchvision) (1.11.0)\nRequirement already satisfied: numpy in /usr/local/lib/python3.6/dist-packages (from torchvision) (1.14.6)\nRequirement already satisfied: torch in /usr/local/lib/python3.6/dist-packages (from torchvision) (0.4.1)\nCollecting pillow>=4.1.1 (from torchvision)\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/62/94/5430ebaa83f91cc7a9f687ff5238e26164a779cca2ef9903232268b0a318/Pillow-5.3.0-cp36-cp36m-manylinux1_x86_64.whl (2.0MB)\n\u001b[K 100% |████████████████████████████████| 2.0MB 9.8MB/s \n\u001b[?25hInstalling collected packages: pillow, torchvision\n Found existing installation: Pillow 4.0.0\n Uninstalling Pillow-4.0.0:\n Successfully uninstalled Pillow-4.0.0\nSuccessfully installed pillow-5.3.0 torchvision-0.2.1\nCollecting bokeh\n\u001b[?25l Downloading https://files.pythonhosted.org/packages/07/1b/1bb751797f0bbbafc2642c629656ce158e7e7b7fb1110f449f7c320fb819/bokeh-0.13.0.tar.gz (16.0MB)\n\u001b[K 100% |████████████████████████████████| 16.0MB 919kB/s \n\u001b[?25hRequirement already satisfied: six>=1.5.2 in /usr/local/lib/python3.6/dist-packages (from bokeh) (1.11.0)\nRequirement already satisfied: PyYAML>=3.10 in /usr/local/lib/python3.6/dist-packages (from bokeh) (3.13)\nRequirement already satisfied: python-dateutil>=2.1 in /usr/local/lib/python3.6/dist-packages (from bokeh) (2.5.3)\nRequirement already satisfied: Jinja2>=2.7 in /usr/local/lib/python3.6/dist-packages (from bokeh) (2.10)\nRequirement already satisfied: numpy>=1.7.1 in /usr/local/lib/python3.6/dist-packages (from bokeh) (1.14.6)\nCollecting packaging>=16.8 (from bokeh)\n Downloading https://files.pythonhosted.org/packages/89/d1/92e6df2e503a69df9faab187c684585f0136662c12bb1f36901d426f3fab/packaging-18.0-py2.py3-none-any.whl\nRequirement already satisfied: tornado>=4.3 in /usr/local/lib/python3.6/dist-packages (from bokeh) (4.5.3)\nRequirement already satisfied: MarkupSafe>=0.23 in /usr/local/lib/python3.6/dist-packages (from Jinja2>=2.7->bokeh) (1.0)\nRequirement already satisfied: pyparsing>=2.0.2 in /usr/local/lib/python3.6/dist-packages (from packaging>=16.8->bokeh) (2.2.2)\nBuilding wheels for collected packages: bokeh\n Running setup.py bdist_wheel for bokeh ... \u001b[?25l-\b \b\\\b \b|\b \b/\b \b-\b \b\\\b \b|\b \b/\b \b-\b \b\\\b \b|\b \b/\b \b-\b \bdone\n\u001b[?25h Stored in directory: /root/.cache/pip/wheels/05/3e/43/95ff0bde940a0a5d86ec13c22d2a4bddc97271cd788f441a63\nSuccessfully built bokeh\nInstalling collected packages: packaging, bokeh\nSuccessfully installed bokeh-0.13.0 packaging-18.0\n"
]
],
[
[
"# Create a Google Drive folder to save results",
"_____no_output_____"
]
],
[
[
"GOOGLE_DRIVE_PATH = 'colab'\n\nfrom google.colab import drive\ndrive.mount('/content/drive/')\npath = os.path.join(\"/content/drive/My Drive/\", GOOGLE_DRIVE_PATH)\nif not os.path.exists(path):\n os.makedirs(path)\n\nos.chdir(path)",
"_____no_output_____"
]
],
[
[
"# Use a specific github repo",
"_____no_output_____"
]
],
[
[
"if not os.path.exists('convNet.pytorch'):\n !git clone https://github.com/eladhoffer/convNet.pytorch --recursive\nos.chdir(os.path.join(path, 'convNet.pytorch'))",
"_____no_output_____"
]
],
[
[
"## Run experiment",
"_____no_output_____"
]
],
[
[
"!python main.py --dataset cifar10 --model resnet --model-config \"{'depth': 18}\" -b 512 --epochs 100 --save resnet18_test ",
"saving to ./results/resnet18_test\ncreating model resnet\ncreated model with configuration: {'dataset': 'cifar10', 'depth': 18}\nnumber of parameters: 175258\nFiles already downloaded and verified\nFiles already downloaded and verified\noptimization regime: [{'epoch': 0, 'optimizer': 'SGD', 'lr': 0.1, 'weight_decay': 0, 'momentum': 0.9}, {'epoch': 81, 'lr': 0.01}, {'epoch': 122, 'lr': 0.001, 'weight_decay': 0}, {'epoch': 164, 'lr': 0.0001}]\n\nStarting Epoch: 1\n\nTRAINING - Epoch: [0][0/97]\tTime 2.424 (2.424)\tData 0.998 (0.998)\tLoss 2.3051 (2.3051)\tPrec@1 8.594 (8.594)\tPrec@5 46.875 (46.875)\t\nTRAINING - Epoch: [0][10/97]\tTime 0.149 (0.361)\tData 0.000 (0.092)\tLoss 2.1968 (2.2665)\tPrec@1 19.336 (13.459)\tPrec@5 72.461 (60.831)\t\nTRAINING - Epoch: [0][20/97]\tTime 0.152 (0.268)\tData 0.000 (0.049)\tLoss 2.0724 (2.1926)\tPrec@1 22.070 (17.866)\tPrec@5 75.391 (67.522)\t\nTRAINING - Epoch: [0][30/97]\tTime 0.155 (0.232)\tData 0.005 (0.034)\tLoss 2.0517 (2.1419)\tPrec@1 27.148 (19.890)\tPrec@5 78.906 (70.974)\t\nTRAINING - Epoch: [0][40/97]\tTime 0.152 (0.214)\tData 0.000 (0.026)\tLoss 1.9294 (2.1056)\tPrec@1 27.539 (21.151)\tPrec@5 80.664 (73.042)\t\nTRAINING - Epoch: [0][50/97]\tTime 0.172 (0.204)\tData 0.000 (0.021)\tLoss 1.8897 (2.0754)\tPrec@1 29.297 (22.300)\tPrec@5 83.789 (74.740)\t\nTRAINING - Epoch: [0][60/97]\tTime 0.153 (0.196)\tData 0.000 (0.018)\tLoss 1.9542 (2.0503)\tPrec@1 27.734 (23.121)\tPrec@5 81.055 (76.002)\t\nTRAINING - Epoch: [0][70/97]\tTime 0.151 (0.191)\tData 0.000 (0.016)\tLoss 1.8472 (2.0208)\tPrec@1 31.641 (24.122)\tPrec@5 84.570 (77.195)\t\nTRAINING - Epoch: [0][80/97]\tTime 0.159 (0.188)\tData 0.000 (0.014)\tLoss 1.7615 (1.9925)\tPrec@1 34.570 (25.096)\tPrec@5 87.891 (78.376)\t\nTRAINING - Epoch: [0][90/97]\tTime 0.149 (0.184)\tData 0.000 (0.012)\tLoss 1.7687 (1.9670)\tPrec@1 36.133 (26.080)\tPrec@5 86.719 (79.320)\t\nEVALUATING - Epoch: [0][0/20]\tTime 1.117 (1.117)\tData 1.056 (1.056)\tLoss 1.7817 (1.7817)\tPrec@1 33.398 (33.398)\tPrec@5 87.695 (87.695)\t\nEVALUATING - Epoch: [0][10/20]\tTime 0.050 (0.189)\tData 0.000 (0.129)\tLoss 1.8167 (1.7600)\tPrec@1 34.180 (34.126)\tPrec@5 86.523 (86.754)\t\n\nResults - Epoch: 1\nTraining Loss 1.9527 \tTraining Prec@1 26.560 \tTraining Prec@5 79.820 \tValidation Loss 1.7631 \tValidation Prec@1 33.890 \tValidation Prec@5 86.460 \t\n\nPlot file saved at: /content/drive/My Drive/colab/convNet.pytorch/results/resnet18_test/results.html\n\nStarting Epoch: 2\n\nTRAINING - Epoch: [1][0/97]\tTime 1.338 (1.338)\tData 1.014 (1.014)\tLoss 1.7351 (1.7351)\tPrec@1 34.961 (34.961)\tPrec@5 86.914 (86.914)\t\nTRAINING - Epoch: [1][10/97]\tTime 0.176 (0.325)\tData 0.000 (0.098)\tLoss 1.7274 (1.7138)\tPrec@1 34.961 (34.055)\tPrec@5 88.477 (88.228)\t\nTRAINING - Epoch: [1][20/97]\tTime 0.157 (0.248)\tData 0.001 (0.052)\tLoss 1.7589 (1.7056)\tPrec@1 34.961 (34.626)\tPrec@5 87.695 (88.597)\t\nTRAINING - Epoch: [1][30/97]\tTime 0.160 (0.220)\tData 0.000 (0.035)\tLoss 1.7013 (1.7011)\tPrec@1 36.914 (34.690)\tPrec@5 87.891 (88.577)\t\nTRAINING - Epoch: [1][40/97]\tTime 0.164 (0.205)\tData 0.000 (0.027)\tLoss 1.5834 (1.6884)\tPrec@1 37.891 (35.175)\tPrec@5 91.211 (88.839)\t\nTRAINING - Epoch: [1][50/97]\tTime 0.174 (0.197)\tData 0.000 (0.022)\tLoss 1.6043 (1.6801)\tPrec@1 39.453 (35.616)\tPrec@5 90.625 (88.917)\t\nTRAINING - Epoch: [1][60/97]\tTime 0.157 (0.191)\tData 0.000 (0.018)\tLoss 1.6374 (1.6704)\tPrec@1 38.867 (36.014)\tPrec@5 89.648 (89.168)\t\nTRAINING - Epoch: [1][70/97]\tTime 0.156 (0.186)\tData 0.000 (0.016)\tLoss 1.5859 (1.6626)\tPrec@1 36.719 (36.257)\tPrec@5 90.820 (89.277)\t\nTRAINING - Epoch: [1][80/97]\tTime 0.147 (0.183)\tData 0.000 (0.014)\tLoss 1.5735 (1.6529)\tPrec@1 41.016 (36.605)\tPrec@5 91.211 (89.475)\t\nTRAINING - Epoch: [1][90/97]\tTime 0.150 (0.180)\tData 0.000 (0.013)\tLoss 1.5651 (1.6422)\tPrec@1 40.820 (37.090)\tPrec@5 91.406 (89.668)\t\nEVALUATING - Epoch: [1][0/20]\tTime 1.267 (1.267)\tData 1.220 (1.220)\tLoss 1.5646 (1.5646)\tPrec@1 43.555 (43.555)\tPrec@5 91.797 (91.797)\t\nEVALUATING - Epoch: [1][10/20]\tTime 0.043 (0.191)\tData 0.000 (0.138)\tLoss 1.6707 (1.5776)\tPrec@1 41.211 (40.785)\tPrec@5 88.477 (90.891)\t\n\nResults - Epoch: 2\nTraining Loss 1.6352 \tTraining Prec@1 37.371 \tTraining Prec@5 89.812 \tValidation Loss 1.5663 \tValidation Prec@1 40.940 \tValidation Prec@5 91.070 \t\n\n\nStarting Epoch: 3\n\nTRAINING - Epoch: [2][0/97]\tTime 1.359 (1.359)\tData 1.085 (1.085)\tLoss 1.5318 (1.5318)\tPrec@1 42.578 (42.578)\tPrec@5 92.773 (92.773)\t\nTRAINING - Epoch: [2][10/97]\tTime 0.180 (0.334)\tData 0.004 (0.104)\tLoss 1.4891 (1.5167)\tPrec@1 44.922 (42.436)\tPrec@5 90.625 (91.460)\t\nTRAINING - Epoch: [2][20/97]\tTime 0.168 (0.251)\tData 0.000 (0.055)\tLoss 1.5377 (1.5122)\tPrec@1 43.164 (43.331)\tPrec@5 92.188 (91.546)\t\nTRAINING - Epoch: [2][30/97]\tTime 0.175 (0.222)\tData 0.000 (0.037)\tLoss 1.4374 (1.5026)\tPrec@1 49.414 (43.845)\tPrec@5 92.773 (91.734)\t\nTRAINING - Epoch: [2][40/97]\tTime 0.161 (0.207)\tData 0.000 (0.029)\tLoss 1.4675 (1.4924)\tPrec@1 46.094 (44.336)\tPrec@5 91.211 (91.825)\t\nTRAINING - Epoch: [2][50/97]\tTime 0.186 (0.198)\tData 0.000 (0.023)\tLoss 1.4264 (1.4826)\tPrec@1 48.438 (44.684)\tPrec@5 92.188 (91.981)\t\nTRAINING - Epoch: [2][60/97]\tTime 0.159 (0.192)\tData 0.007 (0.020)\tLoss 1.4103 (1.4725)\tPrec@1 47.852 (45.060)\tPrec@5 93.945 (92.143)\t\nTRAINING - Epoch: [2][70/97]\tTime 0.162 (0.188)\tData 0.000 (0.017)\tLoss 1.4074 (1.4570)\tPrec@1 46.094 (45.698)\tPrec@5 92.578 (92.344)\t\nTRAINING - Epoch: [2][80/97]\tTime 0.155 (0.184)\tData 0.000 (0.015)\tLoss 1.4573 (1.4467)\tPrec@1 46.875 (46.094)\tPrec@5 92.773 (92.527)\t\nTRAINING - Epoch: [2][90/97]\tTime 0.150 (0.181)\tData 0.000 (0.014)\tLoss 1.3143 (1.4358)\tPrec@1 52.148 (46.596)\tPrec@5 93.945 (92.700)\t\nEVALUATING - Epoch: [2][0/20]\tTime 0.989 (0.989)\tData 0.945 (0.945)\tLoss 1.5360 (1.5360)\tPrec@1 43.750 (43.750)\tPrec@5 94.336 (94.336)\t\nEVALUATING - Epoch: [2][10/20]\tTime 0.051 (0.196)\tData 0.000 (0.144)\tLoss 1.7384 (1.5593)\tPrec@1 43.555 (46.982)\tPrec@5 90.234 (92.791)\t\n\nResults - Epoch: 3\nTraining Loss 1.4283 \tTraining Prec@1 46.952 \tTraining Prec@5 92.808 \tValidation Loss 1.5452 \tValidation Prec@1 46.540 \tValidation Prec@5 92.890 \t\n\n\nStarting Epoch: 4\n\nTRAINING - Epoch: [3][0/97]\tTime 1.618 (1.618)\tData 1.317 (1.317)\tLoss 1.3083 (1.3083)\tPrec@1 52.930 (52.930)\tPrec@5 92.188 (92.188)\t\nTRAINING - Epoch: [3][10/97]\tTime 0.156 (0.334)\tData 0.000 (0.122)\tLoss 1.4001 (1.3278)\tPrec@1 47.656 (50.373)\tPrec@5 92.188 (94.460)\t\nTRAINING - Epoch: [3][20/97]\tTime 0.147 (0.252)\tData 0.000 (0.064)\tLoss 1.3078 (1.3136)\tPrec@1 55.273 (51.618)\tPrec@5 94.336 (94.522)\t\nTRAINING - Epoch: [3][30/97]\tTime 0.155 (0.222)\tData 0.005 (0.044)\tLoss 1.3143 (1.3124)\tPrec@1 52.539 (51.670)\tPrec@5 92.969 (94.361)\t\nTRAINING - Epoch: [3][40/97]\tTime 0.171 (0.207)\tData 0.000 (0.033)\tLoss 1.2481 (1.3039)\tPrec@1 55.273 (51.944)\tPrec@5 95.508 (94.393)\t\nTRAINING - Epoch: [3][50/97]\tTime 0.164 (0.198)\tData 0.000 (0.027)\tLoss 1.2716 (1.3016)\tPrec@1 53.711 (52.106)\tPrec@5 93.750 (94.328)\t\nTRAINING - Epoch: [3][60/97]\tTime 0.174 (0.192)\tData 0.000 (0.023)\tLoss 1.2269 (1.2989)\tPrec@1 53.125 (52.251)\tPrec@5 95.703 (94.349)\t\nTRAINING - Epoch: [3][70/97]\tTime 0.148 (0.187)\tData 0.000 (0.020)\tLoss 1.2283 (1.2907)\tPrec@1 56.641 (52.616)\tPrec@5 94.141 (94.460)\t\nTRAINING - Epoch: [3][80/97]\tTime 0.153 (0.184)\tData 0.001 (0.018)\tLoss 1.1743 (1.2824)\tPrec@1 54.102 (52.889)\tPrec@5 96.289 (94.507)\t\nTRAINING - Epoch: [3][90/97]\tTime 0.150 (0.180)\tData 0.000 (0.016)\tLoss 1.1634 (1.2745)\tPrec@1 60.156 (53.207)\tPrec@5 94.727 (94.602)\t\nEVALUATING - Epoch: [3][0/20]\tTime 1.251 (1.251)\tData 1.196 (1.196)\tLoss 1.2980 (1.2980)\tPrec@1 55.469 (55.469)\tPrec@5 96.094 (96.094)\t\nEVALUATING - Epoch: [3][10/20]\tTime 0.051 (0.200)\tData 0.000 (0.143)\tLoss 1.3498 (1.2883)\tPrec@1 51.562 (55.433)\tPrec@5 94.141 (95.348)\t\n\nResults - Epoch: 4\nTraining Loss 1.2684 \tTraining Prec@1 53.455 \tTraining Prec@5 94.648 \tValidation Loss 1.2883 \tValidation Prec@1 55.010 \tValidation Prec@5 95.420 \t\n\n\nStarting Epoch: 5\n\nTRAINING - Epoch: [4][0/97]\tTime 1.097 (1.097)\tData 0.834 (0.834)\tLoss 1.1543 (1.1543)\tPrec@1 56.445 (56.445)\tPrec@5 96.680 (96.680)\t\nTRAINING - Epoch: [4][10/97]\tTime 0.196 (0.330)\tData 0.000 (0.096)\tLoss 1.1648 (1.1551)\tPrec@1 58.008 (58.416)\tPrec@5 96.289 (95.774)\t\nTRAINING - Epoch: [4][20/97]\tTime 0.170 (0.249)\tData 0.000 (0.051)\tLoss 1.1032 (1.1645)\tPrec@1 61.719 (58.119)\tPrec@5 96.875 (95.685)\t\nTRAINING - Epoch: [4][30/97]\tTime 0.147 (0.221)\tData 0.000 (0.034)\tLoss 1.0998 (1.1533)\tPrec@1 61.523 (58.531)\tPrec@5 96.289 (95.785)\t\nTRAINING - Epoch: [4][40/97]\tTime 0.161 (0.205)\tData 0.000 (0.026)\tLoss 1.1098 (1.1455)\tPrec@1 60.547 (58.851)\tPrec@5 96.094 (95.875)\t\nTRAINING - Epoch: [4][50/97]\tTime 0.152 (0.196)\tData 0.000 (0.021)\tLoss 1.1601 (1.1418)\tPrec@1 55.859 (58.950)\tPrec@5 96.289 (95.837)\t\nTRAINING - Epoch: [4][60/97]\tTime 0.163 (0.191)\tData 0.005 (0.018)\tLoss 1.1030 (1.1357)\tPrec@1 58.594 (59.138)\tPrec@5 95.898 (95.950)\t\nTRAINING - Epoch: [4][70/97]\tTime 0.152 (0.187)\tData 0.005 (0.016)\tLoss 0.9912 (1.1283)\tPrec@1 63.086 (59.433)\tPrec@5 97.070 (95.995)\t\nTRAINING - Epoch: [4][80/97]\tTime 0.145 (0.183)\tData 0.000 (0.014)\tLoss 1.0663 (1.1206)\tPrec@1 62.695 (59.698)\tPrec@5 96.484 (96.050)\t\nTRAINING - Epoch: [4][90/97]\tTime 0.150 (0.180)\tData 0.000 (0.013)\tLoss 1.0019 (1.1150)\tPrec@1 65.039 (59.944)\tPrec@5 95.508 (96.051)\t\nEVALUATING - Epoch: [4][0/20]\tTime 1.251 (1.251)\tData 1.193 (1.193)\tLoss 1.5090 (1.5090)\tPrec@1 49.609 (49.609)\tPrec@5 92.578 (92.578)\t\nEVALUATING - Epoch: [4][10/20]\tTime 0.044 (0.204)\tData 0.000 (0.151)\tLoss 1.6838 (1.5325)\tPrec@1 49.609 (52.557)\tPrec@5 92.969 (92.738)\t\n\nResults - Epoch: 5\nTraining Loss 1.1115 \tTraining Prec@1 60.027 \tTraining Prec@5 96.035 \tValidation Loss 1.5081 \tValidation Prec@1 52.400 \tValidation Prec@5 92.920 \t\n\n\nStarting Epoch: 6\n\nTRAINING - Epoch: [5][0/97]\tTime 0.835 (0.835)\tData 0.578 (0.578)\tLoss 1.0855 (1.0855)\tPrec@1 62.305 (62.305)\tPrec@5 95.312 (95.312)\t\nTRAINING - Epoch: [5][10/97]\tTime 0.196 (0.329)\tData 0.003 (0.097)\tLoss 1.0500 (1.0637)\tPrec@1 64.062 (61.896)\tPrec@5 94.922 (96.662)\t\nTRAINING - Epoch: [5][20/97]\tTime 0.162 (0.250)\tData 0.005 (0.052)\tLoss 1.0470 (1.0544)\tPrec@1 62.305 (62.174)\tPrec@5 96.484 (96.652)\t\nTRAINING - Epoch: [5][30/97]\tTime 0.153 (0.221)\tData 0.000 (0.035)\tLoss 1.0823 (1.0520)\tPrec@1 62.305 (62.279)\tPrec@5 96.680 (96.705)\t\nTRAINING - Epoch: [5][40/97]\tTime 0.154 (0.206)\tData 0.000 (0.027)\tLoss 0.9990 (1.0454)\tPrec@1 62.891 (62.400)\tPrec@5 97.266 (96.789)\t\nTRAINING - Epoch: [5][50/97]\tTime 0.173 (0.197)\tData 0.000 (0.022)\tLoss 1.0229 (1.0394)\tPrec@1 62.500 (62.542)\tPrec@5 97.070 (96.791)\t\nTRAINING - Epoch: [5][60/97]\tTime 0.156 (0.191)\tData 0.000 (0.018)\tLoss 0.9604 (1.0290)\tPrec@1 64.258 (62.820)\tPrec@5 96.875 (96.843)\t\nTRAINING - Epoch: [5][70/97]\tTime 0.162 (0.187)\tData 0.000 (0.016)\tLoss 0.8739 (1.0193)\tPrec@1 66.797 (63.254)\tPrec@5 97.852 (96.911)\t\nTRAINING - Epoch: [5][80/97]\tTime 0.154 (0.184)\tData 0.000 (0.014)\tLoss 1.0185 (1.0174)\tPrec@1 61.914 (63.320)\tPrec@5 97.266 (96.882)\t\nTRAINING - Epoch: [5][90/97]\tTime 0.150 (0.180)\tData 0.000 (0.013)\tLoss 0.9708 (1.0155)\tPrec@1 66.797 (63.447)\tPrec@5 96.875 (96.884)\t\nEVALUATING - Epoch: [5][0/20]\tTime 1.029 (1.029)\tData 0.977 (0.977)\tLoss 1.1466 (1.1466)\tPrec@1 60.742 (60.742)\tPrec@5 95.508 (95.508)\t\nEVALUATING - Epoch: [5][10/20]\tTime 0.045 (0.192)\tData 0.000 (0.136)\tLoss 1.2510 (1.1842)\tPrec@1 55.664 (58.629)\tPrec@5 95.312 (96.023)\t\n\nResults - Epoch: 6\nTraining Loss 1.0152 \tTraining Prec@1 63.531 \tTraining Prec@5 96.875 \tValidation Loss 1.1851 \tValidation Prec@1 58.320 \tValidation Prec@5 96.000 \t\n\n\nStarting Epoch: 7\n\nTRAINING - Epoch: [6][0/97]\tTime 1.301 (1.301)\tData 1.030 (1.030)\tLoss 0.9252 (0.9252)\tPrec@1 68.164 (68.164)\tPrec@5 97.266 (97.266)\t\nTRAINING - Epoch: [6][10/97]\tTime 0.210 (0.335)\tData 0.012 (0.097)\tLoss 0.9724 (0.9638)\tPrec@1 64.648 (65.909)\tPrec@5 97.266 (96.928)\t\nTRAINING - Epoch: [6][20/97]\tTime 0.150 (0.253)\tData 0.000 (0.051)\tLoss 0.9216 (0.9621)\tPrec@1 67.773 (65.904)\tPrec@5 98.242 (97.024)\t\nTRAINING - Epoch: [6][30/97]\tTime 0.149 (0.221)\tData 0.001 (0.035)\tLoss 1.0226 (0.9606)\tPrec@1 64.453 (65.770)\tPrec@5 96.680 (96.925)\t\nTRAINING - Epoch: [6][40/97]\tTime 0.157 (0.207)\tData 0.000 (0.027)\tLoss 0.9550 (0.9562)\tPrec@1 65.234 (65.844)\tPrec@5 96.875 (97.004)\t\nTRAINING - Epoch: [6][50/97]\tTime 0.156 (0.198)\tData 0.000 (0.022)\tLoss 0.9495 (0.9519)\tPrec@1 66.211 (65.939)\tPrec@5 96.875 (97.032)\t\nTRAINING - Epoch: [6][60/97]\tTime 0.157 (0.192)\tData 0.000 (0.018)\tLoss 0.8893 (0.9508)\tPrec@1 68.555 (65.968)\tPrec@5 96.875 (97.090)\t\nTRAINING - Epoch: [6][70/97]\tTime 0.161 (0.187)\tData 0.000 (0.016)\tLoss 0.9984 (0.9489)\tPrec@1 64.258 (66.010)\tPrec@5 97.070 (97.120)\t\nTRAINING - Epoch: [6][80/97]\tTime 0.168 (0.184)\tData 0.003 (0.014)\tLoss 0.9349 (0.9471)\tPrec@1 64.062 (66.023)\tPrec@5 97.266 (97.145)\t\nTRAINING - Epoch: [6][90/97]\tTime 0.149 (0.180)\tData 0.000 (0.013)\tLoss 0.9366 (0.9445)\tPrec@1 68.555 (66.054)\tPrec@5 97.070 (97.184)\t\nEVALUATING - Epoch: [6][0/20]\tTime 1.023 (1.023)\tData 0.966 (0.966)\tLoss 1.0044 (1.0044)\tPrec@1 63.867 (63.867)\tPrec@5 97.461 (97.461)\t\nEVALUATING - Epoch: [6][10/20]\tTime 0.052 (0.191)\tData 0.000 (0.138)\tLoss 1.0503 (1.0261)\tPrec@1 62.695 (64.702)\tPrec@5 97.070 (97.159)\t\n\nResults - Epoch: 7\nTraining Loss 0.9424 \tTraining Prec@1 66.159 \tTraining Prec@5 97.219 \tValidation Loss 1.0193 \tValidation Prec@1 64.590 \tValidation Prec@5 97.090 \t\n\n\nStarting Epoch: 8\n\nTRAINING - Epoch: [7][0/97]\tTime 1.358 (1.358)\tData 1.072 (1.072)\tLoss 0.9595 (0.9595)\tPrec@1 64.844 (64.844)\tPrec@5 97.266 (97.266)\t\nTRAINING - Epoch: [7][10/97]\tTime 0.203 (0.338)\tData 0.010 (0.102)\tLoss 0.8368 (0.8881)\tPrec@1 70.703 (68.164)\tPrec@5 98.047 (97.638)\t\nTRAINING - Epoch: [7][20/97]\tTime 0.152 (0.252)\tData 0.005 (0.054)\tLoss 0.7772 (0.8916)\tPrec@1 70.898 (68.034)\tPrec@5 98.438 (97.684)\t\nTRAINING - Epoch: [7][30/97]\tTime 0.167 (0.222)\tData 0.000 (0.037)\tLoss 0.8358 (0.8899)\tPrec@1 70.117 (68.145)\tPrec@5 98.047 (97.612)\t\nTRAINING - Epoch: [7][40/97]\tTime 0.156 (0.208)\tData 0.000 (0.028)\tLoss 0.8501 (0.8856)\tPrec@1 68.359 (68.345)\tPrec@5 98.438 (97.642)\t\nTRAINING - Epoch: [7][50/97]\tTime 0.158 (0.198)\tData 0.000 (0.023)\tLoss 0.8423 (0.8804)\tPrec@1 70.312 (68.413)\tPrec@5 97.070 (97.664)\t\nTRAINING - Epoch: [7][60/97]\tTime 0.148 (0.192)\tData 0.000 (0.019)\tLoss 0.8959 (0.8756)\tPrec@1 68.359 (68.564)\tPrec@5 97.070 (97.704)\t\nTRAINING - Epoch: [7][70/97]\tTime 0.150 (0.187)\tData 0.000 (0.017)\tLoss 0.9092 (0.8755)\tPrec@1 65.625 (68.491)\tPrec@5 97.266 (97.706)\t\nTRAINING - Epoch: [7][80/97]\tTime 0.150 (0.184)\tData 0.000 (0.015)\tLoss 0.8593 (0.8708)\tPrec@1 67.773 (68.680)\tPrec@5 97.852 (97.741)\t\nTRAINING - Epoch: [7][90/97]\tTime 0.149 (0.180)\tData 0.000 (0.013)\tLoss 0.8497 (0.8684)\tPrec@1 67.578 (68.814)\tPrec@5 97.852 (97.742)\t\nEVALUATING - Epoch: [7][0/20]\tTime 1.028 (1.028)\tData 0.975 (0.975)\tLoss 1.2370 (1.2370)\tPrec@1 61.523 (61.523)\tPrec@5 96.289 (96.289)\t\nEVALUATING - Epoch: [7][10/20]\tTime 0.063 (0.189)\tData 0.000 (0.134)\tLoss 1.2470 (1.2720)\tPrec@1 59.961 (59.712)\tPrec@5 96.289 (95.188)\t\n\nResults - Epoch: 8\nTraining Loss 0.8663 \tTraining Prec@1 68.861 \tTraining Prec@5 97.761 \tValidation Loss 1.2764 \tValidation Prec@1 59.790 \tValidation Prec@5 95.090 \t\n\n\nStarting Epoch: 9\n\nTRAINING - Epoch: [8][0/97]\tTime 1.452 (1.452)\tData 1.172 (1.172)\tLoss 0.9020 (0.9020)\tPrec@1 69.727 (69.727)\tPrec@5 97.656 (97.656)\t\nTRAINING - Epoch: [8][10/97]\tTime 0.163 (0.337)\tData 0.000 (0.110)\tLoss 0.8453 (0.8469)\tPrec@1 70.312 (69.549)\tPrec@5 97.852 (97.727)\t\nTRAINING - Epoch: [8][20/97]\tTime 0.166 (0.254)\tData 0.000 (0.058)\tLoss 0.7194 (0.8314)\tPrec@1 73.047 (70.136)\tPrec@5 98.633 (97.805)\t\nTRAINING - Epoch: [8][30/97]\tTime 0.150 (0.224)\tData 0.000 (0.039)\tLoss 0.8919 (0.8274)\tPrec@1 68.555 (70.237)\tPrec@5 97.852 (97.921)\t\nTRAINING - Epoch: [8][40/97]\tTime 0.191 (0.208)\tData 0.000 (0.030)\tLoss 0.8080 (0.8210)\tPrec@1 68.945 (70.251)\tPrec@5 98.633 (97.990)\t\nTRAINING - Epoch: [8][50/97]\tTime 0.166 (0.199)\tData 0.000 (0.024)\tLoss 0.8117 (0.8179)\tPrec@1 72.852 (70.508)\tPrec@5 97.461 (98.024)\t\nTRAINING - Epoch: [8][60/97]\tTime 0.161 (0.194)\tData 0.000 (0.020)\tLoss 0.9292 (0.8169)\tPrec@1 68.164 (70.694)\tPrec@5 96.484 (97.960)\t\nTRAINING - Epoch: [8][70/97]\tTime 0.155 (0.189)\tData 0.000 (0.018)\tLoss 0.8907 (0.8206)\tPrec@1 69.141 (70.555)\tPrec@5 97.461 (97.931)\t\nTRAINING - Epoch: [8][80/97]\tTime 0.154 (0.185)\tData 0.000 (0.016)\tLoss 0.8297 (0.8186)\tPrec@1 70.312 (70.616)\tPrec@5 98.047 (97.967)\t\nTRAINING - Epoch: [8][90/97]\tTime 0.149 (0.181)\tData 0.000 (0.014)\tLoss 0.6810 (0.8161)\tPrec@1 74.609 (70.737)\tPrec@5 99.219 (97.995)\t\nEVALUATING - Epoch: [8][0/20]\tTime 1.001 (1.001)\tData 0.950 (0.950)\tLoss 0.9103 (0.9103)\tPrec@1 68.164 (68.164)\tPrec@5 98.242 (98.242)\t\nEVALUATING - Epoch: [8][10/20]\tTime 0.060 (0.194)\tData 0.000 (0.138)\tLoss 0.9905 (0.9874)\tPrec@1 69.336 (67.010)\tPrec@5 98.047 (97.798)\t\n\nResults - Epoch: 9\nTraining Loss 0.8129 \tTraining Prec@1 70.852 \tTraining Prec@5 98.001 \tValidation Loss 0.9813 \tValidation Prec@1 66.870 \tValidation Prec@5 97.920 \t\n\n\nStarting Epoch: 10\n\nTRAINING - Epoch: [9][0/97]\tTime 1.376 (1.376)\tData 1.103 (1.103)\tLoss 0.7292 (0.7292)\tPrec@1 74.023 (74.023)\tPrec@5 98.242 (98.242)\t\nTRAINING - Epoch: [9][10/97]\tTime 0.203 (0.337)\tData 0.000 (0.103)\tLoss 0.7811 (0.7801)\tPrec@1 73.047 (72.230)\tPrec@5 97.852 (98.331)\t\nTRAINING - Epoch: [9][20/97]\tTime 0.157 (0.255)\tData 0.000 (0.055)\tLoss 0.8006 (0.7804)\tPrec@1 70.508 (72.405)\tPrec@5 98.242 (98.298)\t\nTRAINING - Epoch: [9][30/97]\tTime 0.181 (0.226)\tData 0.000 (0.037)\tLoss 0.8448 (0.7791)\tPrec@1 68.945 (72.360)\tPrec@5 98.047 (98.230)\t\nTRAINING - Epoch: [9][40/97]\tTime 0.148 (0.210)\tData 0.000 (0.029)\tLoss 0.8310 (0.7758)\tPrec@1 70.898 (72.447)\tPrec@5 98.047 (98.247)\t\nTRAINING - Epoch: [9][50/97]\tTime 0.151 (0.200)\tData 0.000 (0.023)\tLoss 0.8339 (0.7751)\tPrec@1 70.703 (72.403)\tPrec@5 98.828 (98.235)\t\nTRAINING - Epoch: [9][60/97]\tTime 0.154 (0.193)\tData 0.000 (0.020)\tLoss 0.7483 (0.7718)\tPrec@1 73.047 (72.541)\tPrec@5 98.438 (98.236)\t\nTRAINING - Epoch: [9][70/97]\tTime 0.159 (0.189)\tData 0.000 (0.017)\tLoss 0.7302 (0.7701)\tPrec@1 74.805 (72.585)\tPrec@5 98.242 (98.253)\t\nTRAINING - Epoch: [9][80/97]\tTime 0.151 (0.185)\tData 0.000 (0.015)\tLoss 0.7995 (0.7742)\tPrec@1 72.656 (72.468)\tPrec@5 98.438 (98.225)\t\nTRAINING - Epoch: [9][90/97]\tTime 0.149 (0.181)\tData 0.000 (0.013)\tLoss 0.7868 (0.7734)\tPrec@1 73.242 (72.540)\tPrec@5 97.852 (98.234)\t\nEVALUATING - Epoch: [9][0/20]\tTime 1.003 (1.003)\tData 0.930 (0.930)\tLoss 0.8861 (0.8861)\tPrec@1 70.117 (70.117)\tPrec@5 99.023 (99.023)\t\nEVALUATING - Epoch: [9][10/20]\tTime 0.051 (0.190)\tData 0.000 (0.138)\tLoss 0.8534 (0.8989)\tPrec@1 71.289 (69.300)\tPrec@5 97.656 (97.976)\t\n\nResults - Epoch: 10\nTraining Loss 0.7700 \tTraining Prec@1 72.682 \tTraining Prec@5 98.262 \tValidation Loss 0.8970 \tValidation Prec@1 69.420 \tValidation Prec@5 98.010 \t\n\n\nStarting Epoch: 11\n\nTRAINING - Epoch: [10][0/97]\tTime 1.289 (1.289)\tData 1.008 (1.008)\tLoss 0.7349 (0.7349)\tPrec@1 71.680 (71.680)\tPrec@5 98.242 (98.242)\t\nTRAINING - Epoch: [10][10/97]\tTime 0.175 (0.327)\tData 0.000 (0.095)\tLoss 0.7151 (0.7182)\tPrec@1 73.242 (74.148)\tPrec@5 98.828 (98.526)\t\nTRAINING - Epoch: [10][20/97]\tTime 0.151 (0.247)\tData 0.000 (0.050)\tLoss 0.7631 (0.7186)\tPrec@1 71.484 (73.856)\tPrec@5 98.828 (98.428)\t\nTRAINING - Epoch: [10][30/97]\tTime 0.157 (0.218)\tData 0.000 (0.034)\tLoss 0.6582 (0.7214)\tPrec@1 77.539 (73.771)\tPrec@5 97.852 (98.482)\t\nTRAINING - Epoch: [10][40/97]\tTime 0.164 (0.204)\tData 0.001 (0.026)\tLoss 0.7692 (0.7233)\tPrec@1 74.219 (73.823)\tPrec@5 98.047 (98.447)\t\nTRAINING - Epoch: [10][50/97]\tTime 0.157 (0.195)\tData 0.000 (0.021)\tLoss 0.7170 (0.7260)\tPrec@1 74.219 (73.790)\tPrec@5 98.242 (98.430)\t\nTRAINING - Epoch: [10][60/97]\tTime 0.155 (0.189)\tData 0.000 (0.018)\tLoss 0.7524 (0.7281)\tPrec@1 75.586 (73.851)\tPrec@5 97.852 (98.447)\t\nTRAINING - Epoch: [10][70/97]\tTime 0.150 (0.184)\tData 0.000 (0.016)\tLoss 0.7266 (0.7264)\tPrec@1 75.195 (74.026)\tPrec@5 97.852 (98.415)\t\nTRAINING - Epoch: [10][80/97]\tTime 0.168 (0.181)\tData 0.000 (0.014)\tLoss 0.7751 (0.7292)\tPrec@1 75.391 (73.978)\tPrec@5 97.656 (98.423)\t\nTRAINING - Epoch: [10][90/97]\tTime 0.149 (0.178)\tData 0.000 (0.012)\tLoss 0.7150 (0.7265)\tPrec@1 71.289 (74.062)\tPrec@5 99.414 (98.483)\t\nEVALUATING - Epoch: [10][0/20]\tTime 1.107 (1.107)\tData 1.051 (1.051)\tLoss 0.8944 (0.8944)\tPrec@1 69.141 (69.141)\tPrec@5 96.875 (96.875)\t\nEVALUATING - Epoch: [10][10/20]\tTime 0.116 (0.190)\tData 0.072 (0.132)\tLoss 0.9578 (0.9712)\tPrec@1 70.312 (69.283)\tPrec@5 97.461 (96.378)\t\n\nResults - Epoch: 11\nTraining Loss 0.7277 \tTraining Prec@1 74.096 \tTraining Prec@5 98.452 \tValidation Loss 0.9770 \tValidation Prec@1 68.720 \tValidation Prec@5 96.620 \t\n\n\nStarting Epoch: 12\n\nTRAINING - Epoch: [11][0/97]\tTime 1.409 (1.409)\tData 1.135 (1.135)\tLoss 0.6537 (0.6537)\tPrec@1 75.391 (75.391)\tPrec@5 98.828 (98.828)\t\nTRAINING - Epoch: [11][10/97]\tTime 0.160 (0.333)\tData 0.000 (0.109)\tLoss 0.7393 (0.6995)\tPrec@1 75.000 (75.036)\tPrec@5 98.047 (98.526)\t\nTRAINING - Epoch: [11][20/97]\tTime 0.152 (0.251)\tData 0.000 (0.058)\tLoss 0.7330 (0.6958)\tPrec@1 72.266 (75.009)\tPrec@5 99.414 (98.689)\t\nTRAINING - Epoch: [11][30/97]\tTime 0.174 (0.222)\tData 0.000 (0.040)\tLoss 0.6346 (0.6886)\tPrec@1 76.172 (75.315)\tPrec@5 98.438 (98.677)\t\nTRAINING - Epoch: [11][40/97]\tTime 0.170 (0.207)\tData 0.000 (0.031)\tLoss 0.7012 (0.6886)\tPrec@1 75.000 (75.476)\tPrec@5 98.633 (98.714)\t\nTRAINING - Epoch: [11][50/97]\tTime 0.173 (0.198)\tData 0.000 (0.025)\tLoss 0.6991 (0.6915)\tPrec@1 75.391 (75.452)\tPrec@5 98.438 (98.671)\t\nTRAINING - Epoch: [11][60/97]\tTime 0.156 (0.192)\tData 0.000 (0.021)\tLoss 0.5976 (0.6972)\tPrec@1 82.617 (75.227)\tPrec@5 98.633 (98.620)\t\nTRAINING - Epoch: [11][70/97]\tTime 0.161 (0.187)\tData 0.000 (0.018)\tLoss 0.6919 (0.6970)\tPrec@1 75.391 (75.162)\tPrec@5 97.852 (98.605)\t\nTRAINING - Epoch: [11][80/97]\tTime 0.159 (0.184)\tData 0.000 (0.016)\tLoss 0.7154 (0.6952)\tPrec@1 77.539 (75.229)\tPrec@5 98.242 (98.592)\t\nTRAINING - Epoch: [11][90/97]\tTime 0.149 (0.180)\tData 0.000 (0.014)\tLoss 0.5731 (0.6929)\tPrec@1 81.055 (75.376)\tPrec@5 99.414 (98.592)\t\nEVALUATING - Epoch: [11][0/20]\tTime 1.113 (1.113)\tData 1.049 (1.049)\tLoss 0.7345 (0.7345)\tPrec@1 73.633 (73.633)\tPrec@5 99.023 (99.023)\t\nEVALUATING - Epoch: [11][10/20]\tTime 0.048 (0.188)\tData 0.000 (0.134)\tLoss 0.8193 (0.8157)\tPrec@1 73.438 (72.532)\tPrec@5 98.047 (98.207)\t\n\nResults - Epoch: 12\nTraining Loss 0.6927 \tTraining Prec@1 75.415 \tTraining Prec@5 98.587 \tValidation Loss 0.8176 \tValidation Prec@1 72.510 \tValidation Prec@5 98.220 \t\n\n\nStarting Epoch: 13\n\nTRAINING - Epoch: [12][0/97]\tTime 1.370 (1.370)\tData 1.114 (1.114)\tLoss 0.7758 (0.7758)\tPrec@1 73.047 (73.047)\tPrec@5 97.656 (97.656)\t\nTRAINING - Epoch: [12][10/97]\tTime 0.154 (0.333)\tData 0.000 (0.104)\tLoss 0.6978 (0.6755)\tPrec@1 72.852 (75.763)\tPrec@5 98.828 (98.739)\t\nTRAINING - Epoch: [12][20/97]\tTime 0.153 (0.252)\tData 0.000 (0.055)\tLoss 0.6267 (0.6740)\tPrec@1 78.125 (76.144)\tPrec@5 98.438 (98.865)\t\nTRAINING - Epoch: [12][30/97]\tTime 0.163 (0.222)\tData 0.009 (0.038)\tLoss 0.7076 (0.6665)\tPrec@1 73.828 (76.481)\tPrec@5 99.414 (98.872)\t\nTRAINING - Epoch: [12][40/97]\tTime 0.162 (0.208)\tData 0.000 (0.029)\tLoss 0.6959 (0.6659)\tPrec@1 73.828 (76.577)\tPrec@5 98.828 (98.795)\t\nTRAINING - Epoch: [12][50/97]\tTime 0.173 (0.199)\tData 0.000 (0.024)\tLoss 0.6375 (0.6636)\tPrec@1 79.102 (76.727)\tPrec@5 98.828 (98.805)\t\nTRAINING - Epoch: [12][60/97]\tTime 0.168 (0.193)\tData 0.000 (0.020)\tLoss 0.6544 (0.6581)\tPrec@1 77.930 (76.873)\tPrec@5 98.828 (98.812)\t\nTRAINING - Epoch: [12][70/97]\tTime 0.148 (0.188)\tData 0.000 (0.018)\tLoss 0.5998 (0.6541)\tPrec@1 77.930 (76.997)\tPrec@5 98.438 (98.809)\t\nTRAINING - Epoch: [12][80/97]\tTime 0.159 (0.185)\tData 0.005 (0.016)\tLoss 0.6916 (0.6567)\tPrec@1 77.148 (76.893)\tPrec@5 98.242 (98.787)\t\nTRAINING - Epoch: [12][90/97]\tTime 0.149 (0.181)\tData 0.000 (0.014)\tLoss 0.6968 (0.6596)\tPrec@1 75.391 (76.814)\tPrec@5 98.242 (98.740)\t\nEVALUATING - Epoch: [12][0/20]\tTime 1.031 (1.031)\tData 0.979 (0.979)\tLoss 0.7283 (0.7283)\tPrec@1 72.852 (72.852)\tPrec@5 98.633 (98.633)\t\nEVALUATING - Epoch: [12][10/20]\tTime 0.047 (0.192)\tData 0.000 (0.137)\tLoss 0.7600 (0.7460)\tPrec@1 73.633 (74.165)\tPrec@5 98.828 (98.136)\t\n\nResults - Epoch: 13\nTraining Loss 0.6593 \tTraining Prec@1 76.834 \tTraining Prec@5 98.740 \tValidation Loss 0.7543 \tValidation Prec@1 73.910 \tValidation Prec@5 98.220 \t\n\n\nStarting Epoch: 14\n\nTRAINING - Epoch: [13][0/97]\tTime 0.925 (0.925)\tData 0.668 (0.668)\tLoss 0.5987 (0.5987)\tPrec@1 78.711 (78.711)\tPrec@5 99.219 (99.219)\t\nTRAINING - Epoch: [13][10/97]\tTime 0.165 (0.326)\tData 0.000 (0.096)\tLoss 0.6147 (0.6284)\tPrec@1 79.102 (78.107)\tPrec@5 99.219 (99.006)\t\nTRAINING - Epoch: [13][20/97]\tTime 0.175 (0.250)\tData 0.011 (0.051)\tLoss 0.6706 (0.6265)\tPrec@1 76.367 (77.883)\tPrec@5 98.828 (98.903)\t\nTRAINING - Epoch: [13][30/97]\tTime 0.156 (0.221)\tData 0.000 (0.035)\tLoss 0.6013 (0.6354)\tPrec@1 77.344 (77.545)\tPrec@5 98.047 (98.803)\t\nTRAINING - Epoch: [13][40/97]\tTime 0.150 (0.207)\tData 0.000 (0.027)\tLoss 0.5827 (0.6376)\tPrec@1 80.859 (77.711)\tPrec@5 99.609 (98.866)\t\nTRAINING - Epoch: [13][50/97]\tTime 0.151 (0.197)\tData 0.000 (0.022)\tLoss 0.5834 (0.6317)\tPrec@1 79.297 (77.930)\tPrec@5 99.414 (98.832)\t\nTRAINING - Epoch: [13][60/97]\tTime 0.160 (0.192)\tData 0.000 (0.018)\tLoss 0.6882 (0.6295)\tPrec@1 77.148 (78.106)\tPrec@5 98.047 (98.822)\t\nTRAINING - Epoch: [13][70/97]\tTime 0.153 (0.188)\tData 0.000 (0.016)\tLoss 0.5542 (0.6280)\tPrec@1 80.273 (78.037)\tPrec@5 99.219 (98.858)\t\nTRAINING - Epoch: [13][80/97]\tTime 0.152 (0.184)\tData 0.000 (0.014)\tLoss 0.6250 (0.6259)\tPrec@1 80.078 (78.103)\tPrec@5 98.438 (98.840)\t\nTRAINING - Epoch: [13][90/97]\tTime 0.151 (0.181)\tData 0.000 (0.013)\tLoss 0.6270 (0.6275)\tPrec@1 78.906 (78.037)\tPrec@5 99.414 (98.852)\t\nEVALUATING - Epoch: [13][0/20]\tTime 0.984 (0.984)\tData 0.926 (0.926)\tLoss 0.7417 (0.7417)\tPrec@1 74.414 (74.414)\tPrec@5 99.023 (99.023)\t\nEVALUATING - Epoch: [13][10/20]\tTime 0.043 (0.198)\tData 0.000 (0.143)\tLoss 0.8301 (0.7733)\tPrec@1 74.219 (75.710)\tPrec@5 98.047 (98.189)\t\n\nResults - Epoch: 14\nTraining Loss 0.6275 \tTraining Prec@1 78.089 \tTraining Prec@5 98.858 \tValidation Loss 0.7674 \tValidation Prec@1 75.340 \tValidation Prec@5 98.420 \t\n\n\nStarting Epoch: 15\n\nTRAINING - Epoch: [14][0/97]\tTime 1.121 (1.121)\tData 0.874 (0.874)\tLoss 0.6056 (0.6056)\tPrec@1 78.711 (78.711)\tPrec@5 99.219 (99.219)\t\nTRAINING - Epoch: [14][10/97]\tTime 0.177 (0.334)\tData 0.000 (0.101)\tLoss 0.6199 (0.6193)\tPrec@1 78.125 (78.214)\tPrec@5 99.219 (99.023)\t\nTRAINING - Epoch: [14][20/97]\tTime 0.163 (0.252)\tData 0.000 (0.054)\tLoss 0.6455 (0.6124)\tPrec@1 78.906 (78.646)\tPrec@5 99.023 (99.014)\t\nTRAINING - Epoch: [14][30/97]\tTime 0.158 (0.223)\tData 0.000 (0.036)\tLoss 0.7027 (0.6089)\tPrec@1 74.414 (78.604)\tPrec@5 98.438 (98.929)\t\nTRAINING - Epoch: [14][40/97]\tTime 0.164 (0.208)\tData 0.000 (0.028)\tLoss 0.6439 (0.6070)\tPrec@1 77.148 (78.563)\tPrec@5 99.219 (98.952)\t\nTRAINING - Epoch: [14][50/97]\tTime 0.173 (0.198)\tData 0.000 (0.023)\tLoss 0.5371 (0.6040)\tPrec@1 79.492 (78.588)\tPrec@5 99.219 (98.985)\t\nTRAINING - Epoch: [14][60/97]\tTime 0.167 (0.192)\tData 0.000 (0.019)\tLoss 0.5558 (0.6014)\tPrec@1 81.055 (78.749)\tPrec@5 99.023 (98.950)\t\nTRAINING - Epoch: [14][70/97]\tTime 0.168 (0.187)\tData 0.000 (0.017)\tLoss 0.6243 (0.6016)\tPrec@1 78.906 (78.804)\tPrec@5 98.438 (98.963)\t\nTRAINING - Epoch: [14][80/97]\tTime 0.161 (0.184)\tData 0.000 (0.015)\tLoss 0.5349 (0.6021)\tPrec@1 81.836 (78.872)\tPrec@5 99.023 (98.937)\t\nTRAINING - Epoch: [14][90/97]\tTime 0.150 (0.180)\tData 0.000 (0.013)\tLoss 0.5714 (0.6053)\tPrec@1 80.078 (78.735)\tPrec@5 98.828 (98.950)\t\nEVALUATING - Epoch: [14][0/20]\tTime 1.078 (1.078)\tData 1.027 (1.027)\tLoss 0.7717 (0.7717)\tPrec@1 74.609 (74.609)\tPrec@5 98.828 (98.828)\t\nEVALUATING - Epoch: [14][10/20]\tTime 0.062 (0.202)\tData 0.000 (0.146)\tLoss 0.8157 (0.8385)\tPrec@1 73.828 (72.994)\tPrec@5 98.242 (98.065)\t\n\nResults - Epoch: 15\nTraining Loss 0.6062 \tTraining Prec@1 78.661 \tTraining Prec@5 98.951 \tValidation Loss 0.8457 \tValidation Prec@1 72.350 \tValidation Prec@5 98.200 \t\n\n\nStarting Epoch: 16\n\nTRAINING - Epoch: [15][0/97]\tTime 1.103 (1.103)\tData 0.943 (0.943)\tLoss 0.6289 (0.6289)\tPrec@1 78.516 (78.516)\tPrec@5 99.023 (99.023)\t\nTRAINING - Epoch: [15][10/97]\tTime 0.157 (0.330)\tData 0.000 (0.104)\tLoss 0.6272 (0.6113)\tPrec@1 76.953 (78.587)\tPrec@5 99.219 (98.970)\t\nTRAINING - Epoch: [15][20/97]\tTime 0.172 (0.253)\tData 0.000 (0.055)\tLoss 0.5787 (0.5941)\tPrec@1 77.539 (78.943)\tPrec@5 98.828 (98.986)\t\nTRAINING - Epoch: [15][30/97]\tTime 0.170 (0.223)\tData 0.000 (0.038)\tLoss 0.6217 (0.5964)\tPrec@1 77.539 (78.931)\tPrec@5 99.023 (98.942)\t\nTRAINING - Epoch: [15][40/97]\tTime 0.150 (0.207)\tData 0.000 (0.029)\tLoss 0.6192 (0.5931)\tPrec@1 78.125 (79.140)\tPrec@5 97.852 (98.876)\t\nTRAINING - Epoch: [15][50/97]\tTime 0.172 (0.199)\tData 0.000 (0.024)\tLoss 0.5516 (0.5884)\tPrec@1 82.031 (79.362)\tPrec@5 99.219 (98.939)\t\nTRAINING - Epoch: [15][60/97]\tTime 0.168 (0.192)\tData 0.005 (0.020)\tLoss 0.5365 (0.5835)\tPrec@1 80.469 (79.454)\tPrec@5 99.219 (98.979)\t\nTRAINING - Epoch: [15][70/97]\tTime 0.171 (0.187)\tData 0.000 (0.017)\tLoss 0.5565 (0.5832)\tPrec@1 80.273 (79.478)\tPrec@5 99.414 (98.993)\t\nTRAINING - Epoch: [15][80/97]\tTime 0.171 (0.184)\tData 0.004 (0.015)\tLoss 0.5891 (0.5831)\tPrec@1 80.078 (79.519)\tPrec@5 98.828 (99.007)\t\nTRAINING - Epoch: [15][90/97]\tTime 0.149 (0.180)\tData 0.000 (0.014)\tLoss 0.5453 (0.5803)\tPrec@1 82.617 (79.636)\tPrec@5 98.438 (99.017)\t\nEVALUATING - Epoch: [15][0/20]\tTime 1.013 (1.013)\tData 0.960 (0.960)\tLoss 0.8046 (0.8046)\tPrec@1 73.047 (73.047)\tPrec@5 98.633 (98.633)\t\nEVALUATING - Epoch: [15][10/20]\tTime 0.054 (0.191)\tData 0.000 (0.138)\tLoss 0.8960 (0.8681)\tPrec@1 73.242 (73.118)\tPrec@5 97.656 (98.100)\t\n\nResults - Epoch: 16\nTraining Loss 0.5794 \tTraining Prec@1 79.671 \tTraining Prec@5 99.027 \tValidation Loss 0.8586 \tValidation Prec@1 73.180 \tValidation Prec@5 98.310 \t\n\n\nStarting Epoch: 17\n\nTRAINING - Epoch: [16][0/97]\tTime 1.221 (1.221)\tData 0.963 (0.963)\tLoss 0.6031 (0.6031)\tPrec@1 78.906 (78.906)\tPrec@5 99.414 (99.414)\t\nTRAINING - Epoch: [16][10/97]\tTime 0.154 (0.333)\tData 0.000 (0.091)\tLoss 0.5738 (0.5553)\tPrec@1 79.883 (80.362)\tPrec@5 100.000 (99.201)\t\nTRAINING - Epoch: [16][20/97]\tTime 0.158 (0.252)\tData 0.000 (0.048)\tLoss 0.4950 (0.5558)\tPrec@1 83.984 (80.506)\tPrec@5 99.219 (99.116)\t\nTRAINING - Epoch: [16][30/97]\tTime 0.161 (0.223)\tData 0.000 (0.033)\tLoss 0.5387 (0.5623)\tPrec@1 81.445 (80.393)\tPrec@5 98.633 (98.986)\t\nTRAINING - Epoch: [16][40/97]\tTime 0.173 (0.208)\tData 0.005 (0.026)\tLoss 0.5556 (0.5609)\tPrec@1 79.297 (80.426)\tPrec@5 99.219 (99.057)\t\nTRAINING - Epoch: [16][50/97]\tTime 0.151 (0.198)\tData 0.000 (0.021)\tLoss 0.5524 (0.5577)\tPrec@1 80.078 (80.522)\tPrec@5 99.414 (99.073)\t\nTRAINING - Epoch: [16][60/97]\tTime 0.150 (0.192)\tData 0.000 (0.018)\tLoss 0.6007 (0.5587)\tPrec@1 79.688 (80.459)\tPrec@5 99.219 (99.068)\t\nTRAINING - Epoch: [16][70/97]\tTime 0.172 (0.189)\tData 0.000 (0.015)\tLoss 0.5795 (0.5552)\tPrec@1 80.273 (80.628)\tPrec@5 99.023 (99.067)\t\nTRAINING - Epoch: [16][80/97]\tTime 0.156 (0.185)\tData 0.000 (0.014)\tLoss 0.5556 (0.5562)\tPrec@1 81.250 (80.534)\tPrec@5 98.047 (99.062)\t\nTRAINING - Epoch: [16][90/97]\tTime 0.149 (0.181)\tData 0.000 (0.012)\tLoss 0.5909 (0.5568)\tPrec@1 77.344 (80.499)\tPrec@5 99.023 (99.066)\t\nEVALUATING - Epoch: [16][0/20]\tTime 1.015 (1.015)\tData 0.954 (0.954)\tLoss 0.8124 (0.8124)\tPrec@1 72.852 (72.852)\tPrec@5 96.289 (96.289)\t\nEVALUATING - Epoch: [16][10/20]\tTime 0.061 (0.192)\tData 0.000 (0.135)\tLoss 0.8328 (0.8037)\tPrec@1 75.586 (74.716)\tPrec@5 97.461 (97.656)\t\n\nResults - Epoch: 17\nTraining Loss 0.5568 \tTraining Prec@1 80.515 \tTraining Prec@5 99.076 \tValidation Loss 0.8090 \tValidation Prec@1 74.340 \tValidation Prec@5 97.790 \t\n\n\nStarting Epoch: 18\n\nTRAINING - Epoch: [17][0/97]\tTime 1.002 (1.002)\tData 0.725 (0.725)\tLoss 0.5302 (0.5302)\tPrec@1 81.641 (81.641)\tPrec@5 98.828 (98.828)\t\nTRAINING - Epoch: [17][10/97]\tTime 0.178 (0.335)\tData 0.000 (0.098)\tLoss 0.5623 (0.5329)\tPrec@1 80.273 (81.197)\tPrec@5 99.219 (99.219)\t\nTRAINING - Epoch: [17][20/97]\tTime 0.176 (0.255)\tData 0.005 (0.053)\tLoss 0.5829 (0.5356)\tPrec@1 82.227 (81.185)\tPrec@5 98.828 (99.163)\t\nTRAINING - Epoch: [17][30/97]\tTime 0.148 (0.224)\tData 0.000 (0.037)\tLoss 0.5647 (0.5423)\tPrec@1 81.055 (81.074)\tPrec@5 99.219 (99.149)\t\nTRAINING - Epoch: [17][40/97]\tTime 0.174 (0.210)\tData 0.000 (0.028)\tLoss 0.5723 (0.5452)\tPrec@1 81.641 (81.036)\tPrec@5 99.609 (99.128)\t\nTRAINING - Epoch: [17][50/97]\tTime 0.168 (0.200)\tData 0.000 (0.023)\tLoss 0.5352 (0.5476)\tPrec@1 81.641 (80.940)\tPrec@5 98.828 (99.085)\t\nTRAINING - Epoch: [17][60/97]\tTime 0.147 (0.194)\tData 0.000 (0.019)\tLoss 0.5110 (0.5511)\tPrec@1 82.812 (80.866)\tPrec@5 99.609 (99.091)\t\nTRAINING - Epoch: [17][70/97]\tTime 0.178 (0.189)\tData 0.000 (0.017)\tLoss 0.5347 (0.5504)\tPrec@1 81.641 (80.898)\tPrec@5 99.805 (99.103)\t\nTRAINING - Epoch: [17][80/97]\tTime 0.156 (0.186)\tData 0.000 (0.015)\tLoss 0.4938 (0.5489)\tPrec@1 82.422 (80.876)\tPrec@5 99.414 (99.108)\t\nTRAINING - Epoch: [17][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.013)\tLoss 0.5351 (0.5500)\tPrec@1 79.688 (80.789)\tPrec@5 99.609 (99.105)\t\nEVALUATING - Epoch: [17][0/20]\tTime 1.130 (1.130)\tData 1.076 (1.076)\tLoss 0.6355 (0.6355)\tPrec@1 76.562 (76.562)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [17][10/20]\tTime 0.053 (0.194)\tData 0.000 (0.141)\tLoss 0.6818 (0.6951)\tPrec@1 78.711 (76.918)\tPrec@5 98.438 (98.668)\t\n\nResults - Epoch: 18\nTraining Loss 0.5511 \tTraining Prec@1 80.765 \tTraining Prec@5 99.120 \tValidation Loss 0.7013 \tValidation Prec@1 76.530 \tValidation Prec@5 98.710 \t\n\n\nStarting Epoch: 19\n\nTRAINING - Epoch: [18][0/97]\tTime 1.436 (1.436)\tData 1.169 (1.169)\tLoss 0.5829 (0.5829)\tPrec@1 79.297 (79.297)\tPrec@5 99.023 (99.023)\t\nTRAINING - Epoch: [18][10/97]\tTime 0.168 (0.332)\tData 0.000 (0.110)\tLoss 0.4312 (0.5163)\tPrec@1 86.914 (81.676)\tPrec@5 99.414 (99.290)\t\nTRAINING - Epoch: [18][20/97]\tTime 0.177 (0.252)\tData 0.000 (0.059)\tLoss 0.4959 (0.5189)\tPrec@1 83.789 (81.780)\tPrec@5 99.414 (99.191)\t\nTRAINING - Epoch: [18][30/97]\tTime 0.149 (0.223)\tData 0.000 (0.040)\tLoss 0.5250 (0.5106)\tPrec@1 79.492 (81.937)\tPrec@5 99.414 (99.168)\t\nTRAINING - Epoch: [18][40/97]\tTime 0.175 (0.207)\tData 0.000 (0.031)\tLoss 0.5434 (0.5120)\tPrec@1 82.422 (81.960)\tPrec@5 99.023 (99.157)\t\nTRAINING - Epoch: [18][50/97]\tTime 0.176 (0.198)\tData 0.005 (0.025)\tLoss 0.5370 (0.5141)\tPrec@1 80.469 (81.916)\tPrec@5 98.633 (99.154)\t\nTRAINING - Epoch: [18][60/97]\tTime 0.161 (0.192)\tData 0.000 (0.021)\tLoss 0.6024 (0.5169)\tPrec@1 79.297 (81.839)\tPrec@5 99.023 (99.148)\t\nTRAINING - Epoch: [18][70/97]\tTime 0.166 (0.188)\tData 0.000 (0.019)\tLoss 0.4198 (0.5164)\tPrec@1 85.352 (81.778)\tPrec@5 99.805 (99.180)\t\nTRAINING - Epoch: [18][80/97]\tTime 0.168 (0.184)\tData 0.000 (0.016)\tLoss 0.5464 (0.5198)\tPrec@1 80.664 (81.662)\tPrec@5 99.805 (99.192)\t\nTRAINING - Epoch: [18][90/97]\tTime 0.150 (0.180)\tData 0.000 (0.015)\tLoss 0.5338 (0.5203)\tPrec@1 82.422 (81.696)\tPrec@5 98.438 (99.189)\t\nEVALUATING - Epoch: [18][0/20]\tTime 1.015 (1.015)\tData 0.960 (0.960)\tLoss 0.5785 (0.5785)\tPrec@1 81.250 (81.250)\tPrec@5 99.219 (99.219)\t\nEVALUATING - Epoch: [18][10/20]\tTime 0.066 (0.191)\tData 0.000 (0.133)\tLoss 0.5853 (0.6469)\tPrec@1 80.469 (78.942)\tPrec@5 98.633 (98.668)\t\n\nResults - Epoch: 19\nTraining Loss 0.5203 \tTraining Prec@1 81.729 \tTraining Prec@5 99.185 \tValidation Loss 0.6505 \tValidation Prec@1 78.430 \tValidation Prec@5 98.840 \t\n\n\nStarting Epoch: 20\n\nTRAINING - Epoch: [19][0/97]\tTime 1.072 (1.072)\tData 0.799 (0.799)\tLoss 0.4498 (0.4498)\tPrec@1 83.398 (83.398)\tPrec@5 99.805 (99.805)\t\nTRAINING - Epoch: [19][10/97]\tTime 0.153 (0.337)\tData 0.008 (0.098)\tLoss 0.4913 (0.4874)\tPrec@1 83.008 (83.274)\tPrec@5 99.023 (99.343)\t\nTRAINING - Epoch: [19][20/97]\tTime 0.157 (0.254)\tData 0.000 (0.052)\tLoss 0.5327 (0.4934)\tPrec@1 82.227 (83.073)\tPrec@5 99.023 (99.302)\t\nTRAINING - Epoch: [19][30/97]\tTime 0.152 (0.224)\tData 0.000 (0.036)\tLoss 0.5125 (0.4986)\tPrec@1 83.203 (82.850)\tPrec@5 99.023 (99.263)\t\nTRAINING - Epoch: [19][40/97]\tTime 0.148 (0.209)\tData 0.000 (0.027)\tLoss 0.4207 (0.5034)\tPrec@1 84.570 (82.512)\tPrec@5 99.414 (99.281)\t\nTRAINING - Epoch: [19][50/97]\tTime 0.169 (0.200)\tData 0.000 (0.022)\tLoss 0.5199 (0.5082)\tPrec@1 81.836 (82.426)\tPrec@5 99.219 (99.234)\t\nTRAINING - Epoch: [19][60/97]\tTime 0.188 (0.194)\tData 0.000 (0.019)\tLoss 0.5278 (0.5086)\tPrec@1 80.273 (82.284)\tPrec@5 99.609 (99.264)\t\nTRAINING - Epoch: [19][70/97]\tTime 0.183 (0.193)\tData 0.005 (0.017)\tLoss 0.5081 (0.5063)\tPrec@1 82.031 (82.326)\tPrec@5 99.805 (99.263)\t\nTRAINING - Epoch: [19][80/97]\tTime 0.157 (0.190)\tData 0.000 (0.015)\tLoss 0.4453 (0.5096)\tPrec@1 85.156 (82.222)\tPrec@5 100.000 (99.248)\t\nTRAINING - Epoch: [19][90/97]\tTime 0.150 (0.185)\tData 0.000 (0.013)\tLoss 0.5619 (0.5071)\tPrec@1 80.859 (82.293)\tPrec@5 99.023 (99.270)\t\nEVALUATING - Epoch: [19][0/20]\tTime 1.011 (1.011)\tData 0.962 (0.962)\tLoss 0.5751 (0.5751)\tPrec@1 79.102 (79.102)\tPrec@5 99.414 (99.414)\t\nEVALUATING - Epoch: [19][10/20]\tTime 0.045 (0.197)\tData 0.000 (0.145)\tLoss 0.6033 (0.5877)\tPrec@1 79.102 (79.883)\tPrec@5 98.828 (98.864)\t\n\nResults - Epoch: 20\nTraining Loss 0.5064 \tTraining Prec@1 82.287 \tTraining Prec@5 99.269 \tValidation Loss 0.5980 \tValidation Prec@1 79.290 \tValidation Prec@5 98.970 \t\n\n\nStarting Epoch: 21\n\nTRAINING - Epoch: [20][0/97]\tTime 1.435 (1.435)\tData 1.127 (1.127)\tLoss 0.4494 (0.4494)\tPrec@1 85.156 (85.156)\tPrec@5 99.414 (99.414)\t\nTRAINING - Epoch: [20][10/97]\tTime 0.175 (0.336)\tData 0.005 (0.107)\tLoss 0.5937 (0.4959)\tPrec@1 81.250 (83.097)\tPrec@5 98.438 (99.201)\t\nTRAINING - Epoch: [20][20/97]\tTime 0.154 (0.253)\tData 0.000 (0.057)\tLoss 0.4385 (0.4940)\tPrec@1 85.352 (83.175)\tPrec@5 99.414 (99.293)\t\nTRAINING - Epoch: [20][30/97]\tTime 0.158 (0.224)\tData 0.000 (0.039)\tLoss 0.4923 (0.4922)\tPrec@1 83.203 (83.172)\tPrec@5 99.023 (99.275)\t\nTRAINING - Epoch: [20][40/97]\tTime 0.170 (0.216)\tData 0.000 (0.030)\tLoss 0.4847 (0.4912)\tPrec@1 84.375 (83.117)\tPrec@5 99.414 (99.281)\t\nTRAINING - Epoch: [20][50/97]\tTime 0.157 (0.205)\tData 0.000 (0.024)\tLoss 0.4656 (0.4926)\tPrec@1 82.617 (83.046)\tPrec@5 99.414 (99.280)\t\nTRAINING - Epoch: [20][60/97]\tTime 0.163 (0.198)\tData 0.000 (0.020)\tLoss 0.4707 (0.4933)\tPrec@1 83.984 (82.928)\tPrec@5 99.219 (99.292)\t\nTRAINING - Epoch: [20][70/97]\tTime 0.148 (0.193)\tData 0.000 (0.018)\tLoss 0.5224 (0.4919)\tPrec@1 81.836 (82.972)\tPrec@5 99.219 (99.304)\t\nTRAINING - Epoch: [20][80/97]\tTime 0.170 (0.189)\tData 0.002 (0.016)\tLoss 0.5325 (0.4932)\tPrec@1 81.445 (82.931)\tPrec@5 99.609 (99.296)\t\nTRAINING - Epoch: [20][90/97]\tTime 0.149 (0.185)\tData 0.000 (0.014)\tLoss 0.4585 (0.4922)\tPrec@1 84.375 (82.931)\tPrec@5 99.219 (99.294)\t\nEVALUATING - Epoch: [20][0/20]\tTime 1.128 (1.128)\tData 1.075 (1.075)\tLoss 0.7080 (0.7080)\tPrec@1 73.633 (73.633)\tPrec@5 99.414 (99.414)\t\nEVALUATING - Epoch: [20][10/20]\tTime 0.044 (0.194)\tData 0.000 (0.144)\tLoss 0.7402 (0.7456)\tPrec@1 75.391 (76.278)\tPrec@5 99.023 (98.704)\t\n\nResults - Epoch: 21\nTraining Loss 0.4923 \tTraining Prec@1 82.917 \tTraining Prec@5 99.285 \tValidation Loss 0.7660 \tValidation Prec@1 75.640 \tValidation Prec@5 98.780 \t\n\n\nStarting Epoch: 22\n\nTRAINING - Epoch: [21][0/97]\tTime 1.374 (1.374)\tData 1.101 (1.101)\tLoss 0.4778 (0.4778)\tPrec@1 83.594 (83.594)\tPrec@5 98.633 (98.633)\t\nTRAINING - Epoch: [21][10/97]\tTime 0.263 (0.366)\tData 0.004 (0.106)\tLoss 0.5170 (0.4538)\tPrec@1 81.641 (84.126)\tPrec@5 99.414 (99.272)\t\nTRAINING - Epoch: [21][20/97]\tTime 0.186 (0.273)\tData 0.005 (0.058)\tLoss 0.4162 (0.4654)\tPrec@1 84.961 (84.022)\tPrec@5 99.805 (99.135)\t\nTRAINING - Epoch: [21][30/97]\tTime 0.161 (0.236)\tData 0.000 (0.039)\tLoss 0.5443 (0.4706)\tPrec@1 80.859 (83.965)\tPrec@5 99.219 (99.149)\t\nTRAINING - Epoch: [21][40/97]\tTime 0.150 (0.218)\tData 0.000 (0.030)\tLoss 0.4968 (0.4720)\tPrec@1 83.398 (83.884)\tPrec@5 98.047 (99.209)\t\nTRAINING - Epoch: [21][50/97]\tTime 0.173 (0.207)\tData 0.000 (0.024)\tLoss 0.4909 (0.4750)\tPrec@1 83.398 (83.716)\tPrec@5 99.414 (99.203)\t\nTRAINING - Epoch: [21][60/97]\tTime 0.159 (0.199)\tData 0.006 (0.020)\tLoss 0.4543 (0.4759)\tPrec@1 84.961 (83.568)\tPrec@5 99.414 (99.241)\t\nTRAINING - Epoch: [21][70/97]\tTime 0.177 (0.194)\tData 0.000 (0.018)\tLoss 0.4930 (0.4774)\tPrec@1 82.227 (83.519)\tPrec@5 98.828 (99.257)\t\nTRAINING - Epoch: [21][80/97]\tTime 0.187 (0.190)\tData 0.005 (0.016)\tLoss 0.4786 (0.4784)\tPrec@1 83.398 (83.471)\tPrec@5 99.414 (99.272)\t\nTRAINING - Epoch: [21][90/97]\tTime 0.150 (0.186)\tData 0.000 (0.014)\tLoss 0.4640 (0.4791)\tPrec@1 84.180 (83.506)\tPrec@5 99.023 (99.251)\t\nEVALUATING - Epoch: [21][0/20]\tTime 1.064 (1.064)\tData 1.017 (1.017)\tLoss 0.9279 (0.9279)\tPrec@1 69.727 (69.727)\tPrec@5 99.023 (99.023)\t\nEVALUATING - Epoch: [21][10/20]\tTime 0.048 (0.229)\tData 0.000 (0.175)\tLoss 0.8393 (0.8951)\tPrec@1 75.586 (72.976)\tPrec@5 98.633 (98.597)\t\n\nResults - Epoch: 22\nTraining Loss 0.4795 \tTraining Prec@1 83.425 \tTraining Prec@5 99.261 \tValidation Loss 0.8921 \tValidation Prec@1 72.820 \tValidation Prec@5 98.620 \t\n\n\nStarting Epoch: 23\n\nTRAINING - Epoch: [22][0/97]\tTime 1.413 (1.413)\tData 1.115 (1.115)\tLoss 0.4931 (0.4931)\tPrec@1 83.789 (83.789)\tPrec@5 99.219 (99.219)\t\nTRAINING - Epoch: [22][10/97]\tTime 0.163 (0.336)\tData 0.000 (0.105)\tLoss 0.4394 (0.4537)\tPrec@1 84.375 (84.180)\tPrec@5 99.805 (99.467)\t\nTRAINING - Epoch: [22][20/97]\tTime 0.155 (0.255)\tData 0.000 (0.057)\tLoss 0.4152 (0.4544)\tPrec@1 86.328 (84.096)\tPrec@5 99.414 (99.516)\t\nTRAINING - Epoch: [22][30/97]\tTime 0.155 (0.224)\tData 0.000 (0.039)\tLoss 0.4279 (0.4651)\tPrec@1 83.789 (83.833)\tPrec@5 99.805 (99.502)\t\nTRAINING - Epoch: [22][40/97]\tTime 0.176 (0.210)\tData 0.000 (0.029)\tLoss 0.5230 (0.4661)\tPrec@1 82.422 (83.803)\tPrec@5 98.242 (99.438)\t\nTRAINING - Epoch: [22][50/97]\tTime 0.154 (0.200)\tData 0.000 (0.024)\tLoss 0.4387 (0.4633)\tPrec@1 84.766 (83.824)\tPrec@5 99.219 (99.437)\t\nTRAINING - Epoch: [22][60/97]\tTime 0.154 (0.194)\tData 0.000 (0.020)\tLoss 0.4456 (0.4646)\tPrec@1 84.766 (83.786)\tPrec@5 99.414 (99.411)\t\nTRAINING - Epoch: [22][70/97]\tTime 0.152 (0.189)\tData 0.000 (0.018)\tLoss 0.4387 (0.4633)\tPrec@1 84.570 (83.850)\tPrec@5 99.805 (99.395)\t\nTRAINING - Epoch: [22][80/97]\tTime 0.160 (0.186)\tData 0.001 (0.016)\tLoss 0.4490 (0.4615)\tPrec@1 84.570 (83.893)\tPrec@5 99.219 (99.397)\t\nTRAINING - Epoch: [22][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.014)\tLoss 0.4742 (0.4606)\tPrec@1 83.984 (83.937)\tPrec@5 99.414 (99.382)\t\nEVALUATING - Epoch: [22][0/20]\tTime 1.071 (1.071)\tData 1.016 (1.016)\tLoss 0.6533 (0.6533)\tPrec@1 77.734 (77.734)\tPrec@5 99.023 (99.023)\t\nEVALUATING - Epoch: [22][10/20]\tTime 0.058 (0.197)\tData 0.000 (0.139)\tLoss 0.6574 (0.6810)\tPrec@1 79.883 (78.089)\tPrec@5 98.633 (98.828)\t\n\nResults - Epoch: 23\nTraining Loss 0.4599 \tTraining Prec@1 83.976 \tTraining Prec@5 99.376 \tValidation Loss 0.6772 \tValidation Prec@1 78.160 \tValidation Prec@5 98.840 \t\n\n\nStarting Epoch: 24\n\nTRAINING - Epoch: [23][0/97]\tTime 1.560 (1.560)\tData 1.303 (1.303)\tLoss 0.4261 (0.4261)\tPrec@1 83.789 (83.789)\tPrec@5 99.609 (99.609)\t\nTRAINING - Epoch: [23][10/97]\tTime 0.150 (0.333)\tData 0.000 (0.123)\tLoss 0.4817 (0.4444)\tPrec@1 82.812 (84.482)\tPrec@5 99.219 (99.485)\t\nTRAINING - Epoch: [23][20/97]\tTime 0.181 (0.256)\tData 0.000 (0.065)\tLoss 0.4688 (0.4472)\tPrec@1 83.398 (84.301)\tPrec@5 99.609 (99.442)\t\nTRAINING - Epoch: [23][30/97]\tTime 0.161 (0.225)\tData 0.000 (0.044)\tLoss 0.4939 (0.4501)\tPrec@1 83.594 (84.268)\tPrec@5 99.219 (99.383)\t\nTRAINING - Epoch: [23][40/97]\tTime 0.162 (0.210)\tData 0.004 (0.034)\tLoss 0.4448 (0.4475)\tPrec@1 84.375 (84.361)\tPrec@5 99.219 (99.371)\t\nTRAINING - Epoch: [23][50/97]\tTime 0.164 (0.200)\tData 0.001 (0.027)\tLoss 0.5215 (0.4479)\tPrec@1 80.273 (84.375)\tPrec@5 99.219 (99.391)\t\nTRAINING - Epoch: [23][60/97]\tTime 0.169 (0.194)\tData 0.000 (0.023)\tLoss 0.4194 (0.4475)\tPrec@1 87.695 (84.410)\tPrec@5 99.414 (99.395)\t\nTRAINING - Epoch: [23][70/97]\tTime 0.147 (0.189)\tData 0.000 (0.020)\tLoss 0.4687 (0.4509)\tPrec@1 82.812 (84.336)\tPrec@5 99.023 (99.359)\t\nTRAINING - Epoch: [23][80/97]\tTime 0.153 (0.185)\tData 0.000 (0.018)\tLoss 0.4529 (0.4488)\tPrec@1 85.352 (84.447)\tPrec@5 99.219 (99.383)\t\nTRAINING - Epoch: [23][90/97]\tTime 0.149 (0.181)\tData 0.000 (0.016)\tLoss 0.4574 (0.4501)\tPrec@1 84.766 (84.422)\tPrec@5 98.828 (99.365)\t\nEVALUATING - Epoch: [23][0/20]\tTime 1.004 (1.004)\tData 0.958 (0.958)\tLoss 0.5353 (0.5353)\tPrec@1 82.422 (82.422)\tPrec@5 99.219 (99.219)\t\nEVALUATING - Epoch: [23][10/20]\tTime 0.053 (0.200)\tData 0.000 (0.147)\tLoss 0.5529 (0.5893)\tPrec@1 82.617 (80.078)\tPrec@5 98.828 (98.881)\t\n\nResults - Epoch: 24\nTraining Loss 0.4511 \tTraining Prec@1 84.375 \tTraining Prec@5 99.374 \tValidation Loss 0.5917 \tValidation Prec@1 80.000 \tValidation Prec@5 99.050 \t\n\n\nStarting Epoch: 25\n\nTRAINING - Epoch: [24][0/97]\tTime 1.214 (1.214)\tData 0.943 (0.943)\tLoss 0.3863 (0.3863)\tPrec@1 88.281 (88.281)\tPrec@5 99.219 (99.219)\t\nTRAINING - Epoch: [24][10/97]\tTime 0.161 (0.335)\tData 0.000 (0.092)\tLoss 0.4669 (0.4324)\tPrec@1 83.203 (85.014)\tPrec@5 98.633 (99.414)\t\nTRAINING - Epoch: [24][20/97]\tTime 0.177 (0.256)\tData 0.005 (0.049)\tLoss 0.4911 (0.4353)\tPrec@1 81.445 (84.859)\tPrec@5 99.219 (99.442)\t\nTRAINING - Epoch: [24][30/97]\tTime 0.174 (0.225)\tData 0.000 (0.033)\tLoss 0.4599 (0.4384)\tPrec@1 83.789 (84.892)\tPrec@5 99.414 (99.420)\t\nTRAINING - Epoch: [24][40/97]\tTime 0.190 (0.209)\tData 0.000 (0.026)\tLoss 0.4639 (0.4378)\tPrec@1 83.984 (84.880)\tPrec@5 99.219 (99.443)\t\nTRAINING - Epoch: [24][50/97]\tTime 0.156 (0.200)\tData 0.000 (0.021)\tLoss 0.4488 (0.4355)\tPrec@1 83.398 (85.007)\tPrec@5 99.805 (99.429)\t\nTRAINING - Epoch: [24][60/97]\tTime 0.156 (0.193)\tData 0.000 (0.017)\tLoss 0.5100 (0.4381)\tPrec@1 83.008 (84.942)\tPrec@5 99.414 (99.417)\t\nTRAINING - Epoch: [24][70/97]\tTime 0.174 (0.189)\tData 0.005 (0.015)\tLoss 0.4288 (0.4404)\tPrec@1 85.938 (84.870)\tPrec@5 99.805 (99.403)\t\nTRAINING - Epoch: [24][80/97]\tTime 0.159 (0.185)\tData 0.000 (0.013)\tLoss 0.4600 (0.4408)\tPrec@1 83.789 (84.886)\tPrec@5 99.609 (99.409)\t\nTRAINING - Epoch: [24][90/97]\tTime 0.151 (0.181)\tData 0.000 (0.012)\tLoss 0.4023 (0.4401)\tPrec@1 86.523 (84.849)\tPrec@5 99.609 (99.416)\t\nEVALUATING - Epoch: [24][0/20]\tTime 1.052 (1.052)\tData 1.002 (1.002)\tLoss 0.5570 (0.5570)\tPrec@1 82.031 (82.031)\tPrec@5 98.828 (98.828)\t\nEVALUATING - Epoch: [24][10/20]\tTime 0.059 (0.193)\tData 0.000 (0.138)\tLoss 0.5607 (0.6007)\tPrec@1 81.445 (80.771)\tPrec@5 99.023 (98.580)\t\n\nResults - Epoch: 25\nTraining Loss 0.4405 \tTraining Prec@1 84.838 \tTraining Prec@5 99.420 \tValidation Loss 0.6008 \tValidation Prec@1 80.750 \tValidation Prec@5 98.620 \t\n\n\nStarting Epoch: 26\n\nTRAINING - Epoch: [25][0/97]\tTime 1.472 (1.472)\tData 1.196 (1.196)\tLoss 0.3694 (0.3694)\tPrec@1 87.305 (87.305)\tPrec@5 99.805 (99.805)\t\nTRAINING - Epoch: [25][10/97]\tTime 0.180 (0.329)\tData 0.000 (0.111)\tLoss 0.3459 (0.4044)\tPrec@1 88.086 (85.653)\tPrec@5 100.000 (99.538)\t\nTRAINING - Epoch: [25][20/97]\tTime 0.149 (0.248)\tData 0.000 (0.059)\tLoss 0.4003 (0.4128)\tPrec@1 85.742 (85.398)\tPrec@5 99.023 (99.535)\t\nTRAINING - Epoch: [25][30/97]\tTime 0.158 (0.219)\tData 0.000 (0.041)\tLoss 0.3231 (0.4097)\tPrec@1 88.672 (85.761)\tPrec@5 100.000 (99.521)\t\nTRAINING - Epoch: [25][40/97]\tTime 0.165 (0.204)\tData 0.000 (0.031)\tLoss 0.4115 (0.4118)\tPrec@1 85.156 (85.737)\tPrec@5 99.805 (99.543)\t\nTRAINING - Epoch: [25][50/97]\tTime 0.160 (0.195)\tData 0.000 (0.025)\tLoss 0.4154 (0.4150)\tPrec@1 85.156 (85.585)\tPrec@5 99.805 (99.525)\t\nTRAINING - Epoch: [25][60/97]\tTime 0.153 (0.189)\tData 0.000 (0.021)\tLoss 0.4272 (0.4169)\tPrec@1 84.375 (85.537)\tPrec@5 99.219 (99.497)\t\nTRAINING - Epoch: [25][70/97]\tTime 0.165 (0.185)\tData 0.012 (0.018)\tLoss 0.4262 (0.4175)\tPrec@1 85.156 (85.550)\tPrec@5 99.414 (99.491)\t\nTRAINING - Epoch: [25][80/97]\tTime 0.148 (0.182)\tData 0.001 (0.016)\tLoss 0.4537 (0.4194)\tPrec@1 84.180 (85.528)\tPrec@5 99.023 (99.474)\t\nTRAINING - Epoch: [25][90/97]\tTime 0.149 (0.179)\tData 0.000 (0.015)\tLoss 0.4861 (0.4232)\tPrec@1 82.422 (85.349)\tPrec@5 99.219 (99.476)\t\nEVALUATING - Epoch: [25][0/20]\tTime 1.007 (1.007)\tData 0.956 (0.956)\tLoss 0.5916 (0.5916)\tPrec@1 80.859 (80.859)\tPrec@5 98.828 (98.828)\t\nEVALUATING - Epoch: [25][10/20]\tTime 0.080 (0.191)\tData 0.000 (0.133)\tLoss 0.6068 (0.6438)\tPrec@1 80.664 (78.942)\tPrec@5 98.242 (98.757)\t\n\nResults - Epoch: 26\nTraining Loss 0.4261 \tTraining Prec@1 85.311 \tTraining Prec@5 99.462 \tValidation Loss 0.6399 \tValidation Prec@1 78.820 \tValidation Prec@5 98.930 \t\n\n\nStarting Epoch: 27\n\nTRAINING - Epoch: [26][0/97]\tTime 1.606 (1.606)\tData 1.350 (1.350)\tLoss 0.3967 (0.3967)\tPrec@1 85.742 (85.742)\tPrec@5 98.828 (98.828)\t\nTRAINING - Epoch: [26][10/97]\tTime 0.146 (0.336)\tData 0.000 (0.126)\tLoss 0.3750 (0.4038)\tPrec@1 87.500 (85.866)\tPrec@5 99.414 (99.272)\t\nTRAINING - Epoch: [26][20/97]\tTime 0.158 (0.253)\tData 0.000 (0.066)\tLoss 0.4126 (0.4169)\tPrec@1 85.742 (85.724)\tPrec@5 99.219 (99.405)\t\nTRAINING - Epoch: [26][30/97]\tTime 0.157 (0.225)\tData 0.000 (0.045)\tLoss 0.4247 (0.4117)\tPrec@1 85.742 (85.843)\tPrec@5 99.219 (99.420)\t\nTRAINING - Epoch: [26][40/97]\tTime 0.145 (0.210)\tData 0.000 (0.035)\tLoss 0.4660 (0.4200)\tPrec@1 83.984 (85.595)\tPrec@5 99.023 (99.433)\t\nTRAINING - Epoch: [26][50/97]\tTime 0.165 (0.201)\tData 0.000 (0.028)\tLoss 0.4749 (0.4238)\tPrec@1 83.984 (85.451)\tPrec@5 99.023 (99.441)\t\nTRAINING - Epoch: [26][60/97]\tTime 0.158 (0.195)\tData 0.000 (0.024)\tLoss 0.3831 (0.4221)\tPrec@1 86.914 (85.508)\tPrec@5 100.000 (99.446)\t\nTRAINING - Epoch: [26][70/97]\tTime 0.155 (0.191)\tData 0.000 (0.021)\tLoss 0.3626 (0.4198)\tPrec@1 87.891 (85.572)\tPrec@5 99.414 (99.455)\t\nTRAINING - Epoch: [26][80/97]\tTime 0.150 (0.187)\tData 0.005 (0.018)\tLoss 0.3498 (0.4207)\tPrec@1 88.281 (85.523)\tPrec@5 99.414 (99.438)\t\nTRAINING - Epoch: [26][90/97]\tTime 0.150 (0.183)\tData 0.000 (0.016)\tLoss 0.4278 (0.4194)\tPrec@1 84.375 (85.532)\tPrec@5 100.000 (99.466)\t\nEVALUATING - Epoch: [26][0/20]\tTime 1.145 (1.145)\tData 1.092 (1.092)\tLoss 0.5603 (0.5603)\tPrec@1 80.078 (80.078)\tPrec@5 99.023 (99.023)\t\nEVALUATING - Epoch: [26][10/20]\tTime 0.061 (0.189)\tData 0.017 (0.136)\tLoss 0.5859 (0.6122)\tPrec@1 81.055 (80.966)\tPrec@5 98.828 (98.935)\t\n\nResults - Epoch: 27\nTraining Loss 0.4186 \tTraining Prec@1 85.521 \tTraining Prec@5 99.466 \tValidation Loss 0.6091 \tValidation Prec@1 80.630 \tValidation Prec@5 99.050 \t\n\n\nStarting Epoch: 28\n\nTRAINING - Epoch: [27][0/97]\tTime 1.323 (1.323)\tData 1.040 (1.040)\tLoss 0.4259 (0.4259)\tPrec@1 85.156 (85.156)\tPrec@5 99.219 (99.219)\t\nTRAINING - Epoch: [27][10/97]\tTime 0.171 (0.336)\tData 0.000 (0.097)\tLoss 0.4090 (0.3964)\tPrec@1 86.328 (86.204)\tPrec@5 100.000 (99.485)\t\nTRAINING - Epoch: [27][20/97]\tTime 0.147 (0.254)\tData 0.000 (0.052)\tLoss 0.5369 (0.4028)\tPrec@1 81.445 (85.965)\tPrec@5 99.609 (99.507)\t\nTRAINING - Epoch: [27][30/97]\tTime 0.148 (0.223)\tData 0.000 (0.036)\tLoss 0.4281 (0.4083)\tPrec@1 85.742 (85.685)\tPrec@5 99.414 (99.502)\t\nTRAINING - Epoch: [27][40/97]\tTime 0.150 (0.207)\tData 0.000 (0.027)\tLoss 0.3467 (0.4096)\tPrec@1 87.305 (85.685)\tPrec@5 99.805 (99.509)\t\nTRAINING - Epoch: [27][50/97]\tTime 0.179 (0.199)\tData 0.000 (0.022)\tLoss 0.4479 (0.4089)\tPrec@1 84.570 (85.681)\tPrec@5 99.219 (99.506)\t\nTRAINING - Epoch: [27][60/97]\tTime 0.172 (0.193)\tData 0.000 (0.018)\tLoss 0.3920 (0.4109)\tPrec@1 87.109 (85.611)\tPrec@5 99.414 (99.494)\t\nTRAINING - Epoch: [27][70/97]\tTime 0.148 (0.188)\tData 0.000 (0.016)\tLoss 0.4105 (0.4107)\tPrec@1 85.938 (85.605)\tPrec@5 99.219 (99.494)\t\nTRAINING - Epoch: [27][80/97]\tTime 0.149 (0.185)\tData 0.000 (0.014)\tLoss 0.3881 (0.4114)\tPrec@1 87.891 (85.610)\tPrec@5 99.219 (99.501)\t\nTRAINING - Epoch: [27][90/97]\tTime 0.149 (0.181)\tData 0.000 (0.013)\tLoss 0.3437 (0.4097)\tPrec@1 88.477 (85.667)\tPrec@5 100.000 (99.500)\t\nEVALUATING - Epoch: [27][0/20]\tTime 1.002 (1.002)\tData 0.950 (0.950)\tLoss 0.5007 (0.5007)\tPrec@1 82.617 (82.617)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [27][10/20]\tTime 0.055 (0.194)\tData 0.000 (0.137)\tLoss 0.5331 (0.5536)\tPrec@1 81.836 (82.085)\tPrec@5 98.828 (99.006)\t\n\nResults - Epoch: 28\nTraining Loss 0.4094 \tTraining Prec@1 85.692 \tTraining Prec@5 99.507 \tValidation Loss 0.5581 \tValidation Prec@1 82.160 \tValidation Prec@5 98.980 \t\n\n\nStarting Epoch: 29\n\nTRAINING - Epoch: [28][0/97]\tTime 1.485 (1.485)\tData 1.215 (1.215)\tLoss 0.3506 (0.3506)\tPrec@1 88.281 (88.281)\tPrec@5 99.609 (99.609)\t\nTRAINING - Epoch: [28][10/97]\tTime 0.157 (0.335)\tData 0.000 (0.113)\tLoss 0.3569 (0.3833)\tPrec@1 86.719 (86.435)\tPrec@5 100.000 (99.592)\t\nTRAINING - Epoch: [28][20/97]\tTime 0.169 (0.254)\tData 0.000 (0.060)\tLoss 0.3878 (0.3917)\tPrec@1 87.109 (86.412)\tPrec@5 99.609 (99.581)\t\nTRAINING - Epoch: [28][30/97]\tTime 0.175 (0.225)\tData 0.000 (0.041)\tLoss 0.3858 (0.3933)\tPrec@1 87.695 (86.297)\tPrec@5 99.023 (99.572)\t\nTRAINING - Epoch: [28][40/97]\tTime 0.148 (0.210)\tData 0.000 (0.031)\tLoss 0.3922 (0.3971)\tPrec@1 85.156 (86.176)\tPrec@5 99.414 (99.581)\t\nTRAINING - Epoch: [28][50/97]\tTime 0.162 (0.201)\tData 0.005 (0.026)\tLoss 0.4191 (0.4006)\tPrec@1 83.398 (86.037)\tPrec@5 99.609 (99.560)\t\nTRAINING - Epoch: [28][60/97]\tTime 0.165 (0.194)\tData 0.000 (0.022)\tLoss 0.3565 (0.3969)\tPrec@1 88.672 (86.216)\tPrec@5 99.609 (99.561)\t\nTRAINING - Epoch: [28][70/97]\tTime 0.150 (0.189)\tData 0.000 (0.019)\tLoss 0.3761 (0.3949)\tPrec@1 87.695 (86.224)\tPrec@5 99.805 (99.571)\t\nTRAINING - Epoch: [28][80/97]\tTime 0.159 (0.187)\tData 0.000 (0.017)\tLoss 0.4379 (0.3968)\tPrec@1 85.938 (86.164)\tPrec@5 99.219 (99.564)\t\nTRAINING - Epoch: [28][90/97]\tTime 0.151 (0.183)\tData 0.000 (0.015)\tLoss 0.3787 (0.3971)\tPrec@1 85.938 (86.176)\tPrec@5 99.609 (99.556)\t\nEVALUATING - Epoch: [28][0/20]\tTime 1.223 (1.223)\tData 1.171 (1.171)\tLoss 0.6773 (0.6773)\tPrec@1 78.906 (78.906)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [28][10/20]\tTime 0.049 (0.197)\tData 0.000 (0.144)\tLoss 0.6653 (0.6843)\tPrec@1 78.320 (79.261)\tPrec@5 98.828 (98.899)\t\n\nResults - Epoch: 29\nTraining Loss 0.3960 \tTraining Prec@1 86.238 \tTraining Prec@5 99.553 \tValidation Loss 0.6928 \tValidation Prec@1 78.370 \tValidation Prec@5 98.900 \t\n\n\nStarting Epoch: 30\n\nTRAINING - Epoch: [29][0/97]\tTime 1.461 (1.461)\tData 1.155 (1.155)\tLoss 0.3621 (0.3621)\tPrec@1 87.695 (87.695)\tPrec@5 99.609 (99.609)\t\nTRAINING - Epoch: [29][10/97]\tTime 0.152 (0.337)\tData 0.000 (0.106)\tLoss 0.4040 (0.3780)\tPrec@1 85.352 (86.737)\tPrec@5 99.609 (99.663)\t\nTRAINING - Epoch: [29][20/97]\tTime 0.158 (0.253)\tData 0.000 (0.057)\tLoss 0.4405 (0.3843)\tPrec@1 83.789 (86.579)\tPrec@5 99.609 (99.581)\t\nTRAINING - Epoch: [29][30/97]\tTime 0.169 (0.223)\tData 0.000 (0.039)\tLoss 0.3502 (0.3858)\tPrec@1 88.281 (86.618)\tPrec@5 99.414 (99.622)\t\nTRAINING - Epoch: [29][40/97]\tTime 0.164 (0.208)\tData 0.000 (0.030)\tLoss 0.4403 (0.3952)\tPrec@1 85.352 (86.381)\tPrec@5 98.828 (99.562)\t\nTRAINING - Epoch: [29][50/97]\tTime 0.171 (0.199)\tData 0.000 (0.024)\tLoss 0.3703 (0.3938)\tPrec@1 86.719 (86.405)\tPrec@5 99.609 (99.533)\t\nTRAINING - Epoch: [29][60/97]\tTime 0.165 (0.193)\tData 0.000 (0.020)\tLoss 0.3492 (0.3942)\tPrec@1 88.086 (86.405)\tPrec@5 99.805 (99.539)\t\nTRAINING - Epoch: [29][70/97]\tTime 0.155 (0.188)\tData 0.000 (0.018)\tLoss 0.3830 (0.3910)\tPrec@1 86.523 (86.397)\tPrec@5 98.438 (99.524)\t\nTRAINING - Epoch: [29][80/97]\tTime 0.171 (0.185)\tData 0.001 (0.015)\tLoss 0.4033 (0.3912)\tPrec@1 86.328 (86.432)\tPrec@5 98.828 (99.520)\t\nTRAINING - Epoch: [29][90/97]\tTime 0.150 (0.181)\tData 0.000 (0.014)\tLoss 0.3837 (0.3915)\tPrec@1 86.914 (86.384)\tPrec@5 99.414 (99.508)\t\nEVALUATING - Epoch: [29][0/20]\tTime 1.030 (1.030)\tData 0.983 (0.983)\tLoss 0.4667 (0.4667)\tPrec@1 84.180 (84.180)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [29][10/20]\tTime 0.046 (0.201)\tData 0.000 (0.144)\tLoss 0.4499 (0.4910)\tPrec@1 86.328 (84.038)\tPrec@5 99.609 (99.272)\t\n\nResults - Epoch: 30\nTraining Loss 0.3916 \tTraining Prec@1 86.364 \tTraining Prec@5 99.519 \tValidation Loss 0.4904 \tValidation Prec@1 83.910 \tValidation Prec@5 99.380 \t\n\n\nStarting Epoch: 31\n\nTRAINING - Epoch: [30][0/97]\tTime 1.339 (1.339)\tData 1.039 (1.039)\tLoss 0.3194 (0.3194)\tPrec@1 89.648 (89.648)\tPrec@5 99.609 (99.609)\t\nTRAINING - Epoch: [30][10/97]\tTime 0.165 (0.329)\tData 0.000 (0.100)\tLoss 0.3619 (0.3684)\tPrec@1 87.109 (87.216)\tPrec@5 99.609 (99.574)\t\nTRAINING - Epoch: [30][20/97]\tTime 0.157 (0.252)\tData 0.000 (0.053)\tLoss 0.3698 (0.3652)\tPrec@1 87.891 (87.435)\tPrec@5 99.805 (99.600)\t\nTRAINING - Epoch: [30][30/97]\tTime 0.147 (0.222)\tData 0.000 (0.037)\tLoss 0.3518 (0.3670)\tPrec@1 87.500 (87.279)\tPrec@5 99.805 (99.565)\t\nTRAINING - Epoch: [30][40/97]\tTime 0.169 (0.207)\tData 0.000 (0.028)\tLoss 0.3295 (0.3702)\tPrec@1 88.086 (87.062)\tPrec@5 99.609 (99.562)\t\nTRAINING - Epoch: [30][50/97]\tTime 0.162 (0.198)\tData 0.007 (0.023)\tLoss 0.3888 (0.3719)\tPrec@1 86.914 (87.002)\tPrec@5 99.805 (99.556)\t\nTRAINING - Epoch: [30][60/97]\tTime 0.159 (0.192)\tData 0.000 (0.019)\tLoss 0.3763 (0.3712)\tPrec@1 86.523 (87.058)\tPrec@5 99.219 (99.561)\t\nTRAINING - Epoch: [30][70/97]\tTime 0.147 (0.188)\tData 0.000 (0.017)\tLoss 0.3811 (0.3742)\tPrec@1 88.086 (86.964)\tPrec@5 99.609 (99.563)\t\nTRAINING - Epoch: [30][80/97]\tTime 0.154 (0.184)\tData 0.000 (0.015)\tLoss 0.3541 (0.3751)\tPrec@1 88.672 (86.926)\tPrec@5 99.805 (99.544)\t\nTRAINING - Epoch: [30][90/97]\tTime 0.150 (0.180)\tData 0.000 (0.013)\tLoss 0.3246 (0.3772)\tPrec@1 88.867 (86.854)\tPrec@5 99.805 (99.541)\t\nEVALUATING - Epoch: [30][0/20]\tTime 1.265 (1.265)\tData 1.204 (1.204)\tLoss 0.6887 (0.6887)\tPrec@1 77.539 (77.539)\tPrec@5 98.633 (98.633)\t\nEVALUATING - Epoch: [30][10/20]\tTime 0.043 (0.199)\tData 0.000 (0.141)\tLoss 0.7820 (0.7433)\tPrec@1 77.344 (76.687)\tPrec@5 98.047 (98.544)\t\n\nResults - Epoch: 31\nTraining Loss 0.3796 \tTraining Prec@1 86.789 \tTraining Prec@5 99.545 \tValidation Loss 0.7327 \tValidation Prec@1 76.560 \tValidation Prec@5 98.740 \t\n\n\nStarting Epoch: 32\n\nTRAINING - Epoch: [31][0/97]\tTime 1.457 (1.457)\tData 1.216 (1.216)\tLoss 0.3315 (0.3315)\tPrec@1 87.305 (87.305)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [31][10/97]\tTime 0.149 (0.332)\tData 0.000 (0.115)\tLoss 0.2992 (0.3600)\tPrec@1 89.453 (87.340)\tPrec@5 99.414 (99.556)\t\nTRAINING - Epoch: [31][20/97]\tTime 0.174 (0.252)\tData 0.005 (0.061)\tLoss 0.3858 (0.3666)\tPrec@1 86.914 (87.156)\tPrec@5 99.609 (99.609)\t\nTRAINING - Epoch: [31][30/97]\tTime 0.156 (0.223)\tData 0.000 (0.042)\tLoss 0.4031 (0.3717)\tPrec@1 85.547 (87.009)\tPrec@5 99.609 (99.590)\t\nTRAINING - Epoch: [31][40/97]\tTime 0.174 (0.208)\tData 0.000 (0.032)\tLoss 0.3673 (0.3719)\tPrec@1 87.695 (87.009)\tPrec@5 99.609 (99.609)\t\nTRAINING - Epoch: [31][50/97]\tTime 0.156 (0.198)\tData 0.000 (0.026)\tLoss 0.3633 (0.3735)\tPrec@1 86.914 (86.972)\tPrec@5 99.609 (99.613)\t\nTRAINING - Epoch: [31][60/97]\tTime 0.158 (0.192)\tData 0.000 (0.022)\tLoss 0.3469 (0.3746)\tPrec@1 87.305 (87.026)\tPrec@5 99.023 (99.587)\t\nTRAINING - Epoch: [31][70/97]\tTime 0.153 (0.188)\tData 0.000 (0.019)\tLoss 0.3159 (0.3736)\tPrec@1 89.844 (87.065)\tPrec@5 99.414 (99.582)\t\nTRAINING - Epoch: [31][80/97]\tTime 0.168 (0.185)\tData 0.000 (0.017)\tLoss 0.3921 (0.3758)\tPrec@1 85.352 (86.986)\tPrec@5 99.609 (99.564)\t\nTRAINING - Epoch: [31][90/97]\tTime 0.150 (0.181)\tData 0.000 (0.015)\tLoss 0.3786 (0.3766)\tPrec@1 87.305 (86.912)\tPrec@5 99.609 (99.560)\t\nEVALUATING - Epoch: [31][0/20]\tTime 1.020 (1.020)\tData 0.971 (0.971)\tLoss 0.6753 (0.6753)\tPrec@1 79.102 (79.102)\tPrec@5 99.219 (99.219)\t\nEVALUATING - Epoch: [31][10/20]\tTime 0.053 (0.188)\tData 0.000 (0.134)\tLoss 0.6753 (0.6978)\tPrec@1 77.930 (78.214)\tPrec@5 99.023 (99.094)\t\n\nResults - Epoch: 32\nTraining Loss 0.3763 \tTraining Prec@1 86.934 \tTraining Prec@5 99.561 \tValidation Loss 0.6871 \tValidation Prec@1 78.410 \tValidation Prec@5 99.120 \t\n\n\nStarting Epoch: 33\n\nTRAINING - Epoch: [32][0/97]\tTime 1.265 (1.265)\tData 0.985 (0.985)\tLoss 0.3289 (0.3289)\tPrec@1 88.672 (88.672)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [32][10/97]\tTime 0.183 (0.332)\tData 0.000 (0.095)\tLoss 0.3784 (0.3443)\tPrec@1 87.109 (87.962)\tPrec@5 99.219 (99.698)\t\nTRAINING - Epoch: [32][20/97]\tTime 0.181 (0.254)\tData 0.000 (0.050)\tLoss 0.2827 (0.3546)\tPrec@1 91.211 (87.844)\tPrec@5 100.000 (99.647)\t\nTRAINING - Epoch: [32][30/97]\tTime 0.175 (0.225)\tData 0.000 (0.034)\tLoss 0.3450 (0.3531)\tPrec@1 89.062 (87.897)\tPrec@5 99.609 (99.647)\t\nTRAINING - Epoch: [32][40/97]\tTime 0.159 (0.210)\tData 0.000 (0.026)\tLoss 0.3811 (0.3541)\tPrec@1 85.156 (87.657)\tPrec@5 99.805 (99.662)\t\nTRAINING - Epoch: [32][50/97]\tTime 0.157 (0.200)\tData 0.000 (0.021)\tLoss 0.4108 (0.3595)\tPrec@1 85.352 (87.531)\tPrec@5 99.414 (99.648)\t\nTRAINING - Epoch: [32][60/97]\tTime 0.165 (0.193)\tData 0.000 (0.018)\tLoss 0.4176 (0.3651)\tPrec@1 84.375 (87.369)\tPrec@5 99.805 (99.603)\t\nTRAINING - Epoch: [32][70/97]\tTime 0.167 (0.189)\tData 0.000 (0.015)\tLoss 0.3627 (0.3657)\tPrec@1 87.500 (87.332)\tPrec@5 99.805 (99.609)\t\nTRAINING - Epoch: [32][80/97]\tTime 0.160 (0.185)\tData 0.000 (0.014)\tLoss 0.3319 (0.3659)\tPrec@1 87.305 (87.285)\tPrec@5 99.805 (99.583)\t\nTRAINING - Epoch: [32][90/97]\tTime 0.150 (0.181)\tData 0.000 (0.012)\tLoss 0.3989 (0.3649)\tPrec@1 87.695 (87.324)\tPrec@5 99.219 (99.562)\t\nEVALUATING - Epoch: [32][0/20]\tTime 1.130 (1.130)\tData 1.084 (1.084)\tLoss 0.5199 (0.5199)\tPrec@1 82.227 (82.227)\tPrec@5 98.828 (98.828)\t\nEVALUATING - Epoch: [32][10/20]\tTime 0.049 (0.192)\tData 0.000 (0.138)\tLoss 0.5509 (0.5207)\tPrec@1 82.227 (82.422)\tPrec@5 99.023 (99.130)\t\n\nResults - Epoch: 33\nTraining Loss 0.3655 \tTraining Prec@1 87.303 \tTraining Prec@5 99.559 \tValidation Loss 0.5141 \tValidation Prec@1 82.830 \tValidation Prec@5 99.260 \t\n\n\nStarting Epoch: 34\n\nTRAINING - Epoch: [33][0/97]\tTime 1.491 (1.491)\tData 1.244 (1.244)\tLoss 0.3433 (0.3433)\tPrec@1 87.500 (87.500)\tPrec@5 99.805 (99.805)\t\nTRAINING - Epoch: [33][10/97]\tTime 0.159 (0.334)\tData 0.001 (0.116)\tLoss 0.3701 (0.3619)\tPrec@1 88.281 (87.660)\tPrec@5 99.414 (99.592)\t\nTRAINING - Epoch: [33][20/97]\tTime 0.153 (0.251)\tData 0.000 (0.061)\tLoss 0.3502 (0.3558)\tPrec@1 88.477 (87.993)\tPrec@5 99.805 (99.684)\t\nTRAINING - Epoch: [33][30/97]\tTime 0.180 (0.223)\tData 0.005 (0.042)\tLoss 0.3546 (0.3496)\tPrec@1 88.281 (88.143)\tPrec@5 99.609 (99.723)\t\nTRAINING - Epoch: [33][40/97]\tTime 0.168 (0.206)\tData 0.005 (0.032)\tLoss 0.3360 (0.3546)\tPrec@1 89.062 (87.881)\tPrec@5 99.805 (99.662)\t\nTRAINING - Epoch: [33][50/97]\tTime 0.173 (0.198)\tData 0.010 (0.026)\tLoss 0.4116 (0.3578)\tPrec@1 84.961 (87.711)\tPrec@5 99.805 (99.648)\t\nTRAINING - Epoch: [33][60/97]\tTime 0.162 (0.192)\tData 0.000 (0.022)\tLoss 0.3711 (0.3598)\tPrec@1 85.547 (87.545)\tPrec@5 100.000 (99.645)\t\nTRAINING - Epoch: [33][70/97]\tTime 0.151 (0.188)\tData 0.000 (0.019)\tLoss 0.3753 (0.3597)\tPrec@1 87.305 (87.599)\tPrec@5 99.609 (99.645)\t\nTRAINING - Epoch: [33][80/97]\tTime 0.162 (0.185)\tData 0.000 (0.017)\tLoss 0.3774 (0.3578)\tPrec@1 88.477 (87.666)\tPrec@5 99.414 (99.650)\t\nTRAINING - Epoch: [33][90/97]\tTime 0.150 (0.181)\tData 0.000 (0.015)\tLoss 0.4301 (0.3595)\tPrec@1 85.156 (87.564)\tPrec@5 99.805 (99.637)\t\nEVALUATING - Epoch: [33][0/20]\tTime 1.005 (1.005)\tData 0.952 (0.952)\tLoss 0.4994 (0.4994)\tPrec@1 82.031 (82.031)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [33][10/20]\tTime 0.054 (0.195)\tData 0.000 (0.141)\tLoss 0.4255 (0.5140)\tPrec@1 84.375 (82.990)\tPrec@5 99.023 (99.183)\t\n\nResults - Epoch: 34\nTraining Loss 0.3591 \tTraining Prec@1 87.572 \tTraining Prec@5 99.642 \tValidation Loss 0.5143 \tValidation Prec@1 83.000 \tValidation Prec@5 99.290 \t\n\n\nStarting Epoch: 35\n\nTRAINING - Epoch: [34][0/97]\tTime 1.276 (1.276)\tData 1.010 (1.010)\tLoss 0.3069 (0.3069)\tPrec@1 88.477 (88.477)\tPrec@5 99.805 (99.805)\t\nTRAINING - Epoch: [34][10/97]\tTime 0.192 (0.319)\tData 0.000 (0.096)\tLoss 0.3040 (0.3346)\tPrec@1 89.844 (88.281)\tPrec@5 99.805 (99.716)\t\nTRAINING - Epoch: [34][20/97]\tTime 0.158 (0.248)\tData 0.005 (0.052)\tLoss 0.2658 (0.3301)\tPrec@1 92.383 (88.486)\tPrec@5 99.414 (99.702)\t\nTRAINING - Epoch: [34][30/97]\tTime 0.165 (0.220)\tData 0.000 (0.035)\tLoss 0.3416 (0.3379)\tPrec@1 87.891 (88.225)\tPrec@5 99.609 (99.691)\t\nTRAINING - Epoch: [34][40/97]\tTime 0.149 (0.207)\tData 0.000 (0.027)\tLoss 0.3581 (0.3471)\tPrec@1 88.867 (87.853)\tPrec@5 99.805 (99.638)\t\nTRAINING - Epoch: [34][50/97]\tTime 0.169 (0.198)\tData 0.000 (0.022)\tLoss 0.3307 (0.3503)\tPrec@1 89.258 (87.818)\tPrec@5 99.805 (99.636)\t\nTRAINING - Epoch: [34][60/97]\tTime 0.161 (0.192)\tData 0.000 (0.018)\tLoss 0.3824 (0.3529)\tPrec@1 87.500 (87.737)\tPrec@5 99.805 (99.629)\t\nTRAINING - Epoch: [34][70/97]\tTime 0.159 (0.187)\tData 0.000 (0.016)\tLoss 0.3368 (0.3492)\tPrec@1 88.867 (87.841)\tPrec@5 99.805 (99.640)\t\nTRAINING - Epoch: [34][80/97]\tTime 0.170 (0.184)\tData 0.004 (0.014)\tLoss 0.3222 (0.3478)\tPrec@1 88.086 (87.847)\tPrec@5 99.609 (99.650)\t\nTRAINING - Epoch: [34][90/97]\tTime 0.150 (0.180)\tData 0.000 (0.013)\tLoss 0.3682 (0.3469)\tPrec@1 87.109 (87.831)\tPrec@5 99.805 (99.637)\t\nEVALUATING - Epoch: [34][0/20]\tTime 0.962 (0.962)\tData 0.913 (0.913)\tLoss 0.6524 (0.6524)\tPrec@1 80.273 (80.273)\tPrec@5 99.414 (99.414)\t\nEVALUATING - Epoch: [34][10/20]\tTime 0.069 (0.190)\tData 0.000 (0.136)\tLoss 0.7071 (0.6807)\tPrec@1 79.102 (79.190)\tPrec@5 98.828 (98.846)\t\n\nResults - Epoch: 35\nTraining Loss 0.3478 \tTraining Prec@1 87.786 \tTraining Prec@5 99.638 \tValidation Loss 0.6864 \tValidation Prec@1 78.560 \tValidation Prec@5 98.900 \t\n\n\nStarting Epoch: 36\n\nTRAINING - Epoch: [35][0/97]\tTime 1.272 (1.272)\tData 0.987 (0.987)\tLoss 0.3604 (0.3604)\tPrec@1 87.695 (87.695)\tPrec@5 99.414 (99.414)\t\nTRAINING - Epoch: [35][10/97]\tTime 0.185 (0.330)\tData 0.000 (0.091)\tLoss 0.3339 (0.3690)\tPrec@1 87.891 (87.411)\tPrec@5 100.000 (99.485)\t\nTRAINING - Epoch: [35][20/97]\tTime 0.150 (0.249)\tData 0.000 (0.049)\tLoss 0.3202 (0.3495)\tPrec@1 88.672 (87.891)\tPrec@5 99.609 (99.554)\t\nTRAINING - Epoch: [35][30/97]\tTime 0.154 (0.222)\tData 0.000 (0.033)\tLoss 0.3455 (0.3556)\tPrec@1 88.477 (87.525)\tPrec@5 99.023 (99.540)\t\nTRAINING - Epoch: [35][40/97]\tTime 0.155 (0.207)\tData 0.000 (0.026)\tLoss 0.2891 (0.3528)\tPrec@1 89.844 (87.624)\tPrec@5 99.023 (99.538)\t\nTRAINING - Epoch: [35][50/97]\tTime 0.180 (0.198)\tData 0.000 (0.021)\tLoss 0.3796 (0.3497)\tPrec@1 87.305 (87.741)\tPrec@5 99.609 (99.583)\t\nTRAINING - Epoch: [35][60/97]\tTime 0.157 (0.191)\tData 0.000 (0.017)\tLoss 0.2660 (0.3492)\tPrec@1 90.234 (87.779)\tPrec@5 99.805 (99.577)\t\nTRAINING - Epoch: [35][70/97]\tTime 0.159 (0.187)\tData 0.005 (0.015)\tLoss 0.2836 (0.3463)\tPrec@1 89.453 (87.880)\tPrec@5 100.000 (99.590)\t\nTRAINING - Epoch: [35][80/97]\tTime 0.157 (0.184)\tData 0.000 (0.014)\tLoss 0.3641 (0.3469)\tPrec@1 87.500 (87.927)\tPrec@5 99.609 (99.590)\t\nTRAINING - Epoch: [35][90/97]\tTime 0.149 (0.180)\tData 0.000 (0.012)\tLoss 0.3352 (0.3477)\tPrec@1 88.867 (87.953)\tPrec@5 99.609 (99.601)\t\nEVALUATING - Epoch: [35][0/20]\tTime 1.081 (1.081)\tData 1.029 (1.029)\tLoss 0.4928 (0.4928)\tPrec@1 82.031 (82.031)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [35][10/20]\tTime 0.048 (0.190)\tData 0.000 (0.136)\tLoss 0.4651 (0.5621)\tPrec@1 84.766 (82.156)\tPrec@5 98.828 (99.112)\t\n\nResults - Epoch: 36\nTraining Loss 0.3483 \tTraining Prec@1 87.925 \tTraining Prec@5 99.603 \tValidation Loss 0.5672 \tValidation Prec@1 81.810 \tValidation Prec@5 99.090 \t\n\n\nStarting Epoch: 37\n\nTRAINING - Epoch: [36][0/97]\tTime 1.402 (1.402)\tData 1.132 (1.132)\tLoss 0.2987 (0.2987)\tPrec@1 89.258 (89.258)\tPrec@5 99.609 (99.609)\t\nTRAINING - Epoch: [36][10/97]\tTime 0.175 (0.336)\tData 0.000 (0.107)\tLoss 0.3244 (0.3272)\tPrec@1 88.867 (88.849)\tPrec@5 99.805 (99.751)\t\nTRAINING - Epoch: [36][20/97]\tTime 0.158 (0.253)\tData 0.000 (0.057)\tLoss 0.3379 (0.3311)\tPrec@1 88.281 (88.616)\tPrec@5 99.609 (99.693)\t\nTRAINING - Epoch: [36][30/97]\tTime 0.157 (0.223)\tData 0.000 (0.039)\tLoss 0.3289 (0.3274)\tPrec@1 88.672 (88.653)\tPrec@5 99.805 (99.704)\t\nTRAINING - Epoch: [36][40/97]\tTime 0.165 (0.209)\tData 0.000 (0.030)\tLoss 0.3742 (0.3309)\tPrec@1 86.523 (88.458)\tPrec@5 99.609 (99.709)\t\nTRAINING - Epoch: [36][50/97]\tTime 0.160 (0.200)\tData 0.005 (0.024)\tLoss 0.3771 (0.3330)\tPrec@1 87.305 (88.392)\tPrec@5 99.805 (99.717)\t\nTRAINING - Epoch: [36][60/97]\tTime 0.152 (0.194)\tData 0.000 (0.020)\tLoss 0.3048 (0.3333)\tPrec@1 89.258 (88.323)\tPrec@5 99.609 (99.709)\t\nTRAINING - Epoch: [36][70/97]\tTime 0.161 (0.190)\tData 0.000 (0.018)\tLoss 0.4087 (0.3374)\tPrec@1 86.523 (88.207)\tPrec@5 99.609 (99.684)\t\nTRAINING - Epoch: [36][80/97]\tTime 0.150 (0.186)\tData 0.000 (0.016)\tLoss 0.3151 (0.3392)\tPrec@1 88.867 (88.175)\tPrec@5 99.805 (99.670)\t\nTRAINING - Epoch: [36][90/97]\tTime 0.149 (0.182)\tData 0.000 (0.014)\tLoss 0.3625 (0.3397)\tPrec@1 87.695 (88.172)\tPrec@5 99.609 (99.667)\t\nEVALUATING - Epoch: [36][0/20]\tTime 0.986 (0.986)\tData 0.934 (0.934)\tLoss 0.5563 (0.5563)\tPrec@1 79.688 (79.688)\tPrec@5 99.414 (99.414)\t\nEVALUATING - Epoch: [36][10/20]\tTime 0.060 (0.188)\tData 0.000 (0.127)\tLoss 0.5411 (0.5658)\tPrec@1 84.766 (82.528)\tPrec@5 99.023 (99.077)\t\n\nResults - Epoch: 37\nTraining Loss 0.3406 \tTraining Prec@1 88.130 \tTraining Prec@5 99.668 \tValidation Loss 0.5668 \tValidation Prec@1 82.250 \tValidation Prec@5 99.140 \t\n\n\nStarting Epoch: 38\n\nTRAINING - Epoch: [37][0/97]\tTime 1.326 (1.326)\tData 1.047 (1.047)\tLoss 0.3201 (0.3201)\tPrec@1 88.672 (88.672)\tPrec@5 99.609 (99.609)\t\nTRAINING - Epoch: [37][10/97]\tTime 0.197 (0.339)\tData 0.000 (0.099)\tLoss 0.3422 (0.3285)\tPrec@1 87.500 (88.317)\tPrec@5 99.609 (99.734)\t\nTRAINING - Epoch: [37][20/97]\tTime 0.200 (0.256)\tData 0.010 (0.054)\tLoss 0.3692 (0.3412)\tPrec@1 86.328 (87.993)\tPrec@5 100.000 (99.674)\t\nTRAINING - Epoch: [37][30/97]\tTime 0.165 (0.226)\tData 0.000 (0.037)\tLoss 0.4104 (0.3486)\tPrec@1 85.156 (87.739)\tPrec@5 99.414 (99.647)\t\nTRAINING - Epoch: [37][40/97]\tTime 0.170 (0.211)\tData 0.000 (0.028)\tLoss 0.3291 (0.3458)\tPrec@1 86.133 (87.905)\tPrec@5 100.000 (99.662)\t\nTRAINING - Epoch: [37][50/97]\tTime 0.159 (0.201)\tData 0.000 (0.023)\tLoss 0.3480 (0.3424)\tPrec@1 87.695 (87.944)\tPrec@5 99.805 (99.671)\t\nTRAINING - Epoch: [37][60/97]\tTime 0.166 (0.195)\tData 0.000 (0.020)\tLoss 0.3337 (0.3442)\tPrec@1 87.695 (87.887)\tPrec@5 100.000 (99.670)\t\nTRAINING - Epoch: [37][70/97]\tTime 0.192 (0.191)\tData 0.005 (0.017)\tLoss 0.3459 (0.3432)\tPrec@1 87.500 (87.979)\tPrec@5 99.219 (99.662)\t\nTRAINING - Epoch: [37][80/97]\tTime 0.154 (0.187)\tData 0.000 (0.015)\tLoss 0.2865 (0.3391)\tPrec@1 88.867 (88.074)\tPrec@5 99.609 (99.667)\t\nTRAINING - Epoch: [37][90/97]\tTime 0.150 (0.183)\tData 0.000 (0.014)\tLoss 0.3110 (0.3391)\tPrec@1 88.867 (88.062)\tPrec@5 99.414 (99.663)\t\nEVALUATING - Epoch: [37][0/20]\tTime 1.007 (1.007)\tData 0.960 (0.960)\tLoss 0.5374 (0.5374)\tPrec@1 79.883 (79.883)\tPrec@5 99.219 (99.219)\t\nEVALUATING - Epoch: [37][10/20]\tTime 0.068 (0.192)\tData 0.000 (0.135)\tLoss 0.5747 (0.5804)\tPrec@1 82.031 (81.072)\tPrec@5 98.828 (99.023)\t\n\nResults - Epoch: 38\nTraining Loss 0.3381 \tTraining Prec@1 88.108 \tTraining Prec@5 99.664 \tValidation Loss 0.5784 \tValidation Prec@1 81.070 \tValidation Prec@5 99.180 \t\n\n\nStarting Epoch: 39\n\nTRAINING - Epoch: [38][0/97]\tTime 1.494 (1.494)\tData 1.258 (1.258)\tLoss 0.3037 (0.3037)\tPrec@1 89.453 (89.453)\tPrec@5 99.805 (99.805)\t\nTRAINING - Epoch: [38][10/97]\tTime 0.195 (0.333)\tData 0.000 (0.116)\tLoss 0.3089 (0.3226)\tPrec@1 90.234 (88.601)\tPrec@5 99.805 (99.734)\t\nTRAINING - Epoch: [38][20/97]\tTime 0.151 (0.254)\tData 0.000 (0.061)\tLoss 0.3548 (0.3225)\tPrec@1 87.305 (88.672)\tPrec@5 99.609 (99.767)\t\nTRAINING - Epoch: [38][30/97]\tTime 0.172 (0.224)\tData 0.000 (0.042)\tLoss 0.3377 (0.3244)\tPrec@1 87.109 (88.697)\tPrec@5 99.805 (99.698)\t\nTRAINING - Epoch: [38][40/97]\tTime 0.171 (0.209)\tData 0.000 (0.032)\tLoss 0.3341 (0.3241)\tPrec@1 87.305 (88.643)\tPrec@5 100.000 (99.709)\t\nTRAINING - Epoch: [38][50/97]\tTime 0.168 (0.199)\tData 0.000 (0.026)\tLoss 0.2712 (0.3231)\tPrec@1 90.234 (88.680)\tPrec@5 99.805 (99.713)\t\nTRAINING - Epoch: [38][60/97]\tTime 0.150 (0.193)\tData 0.000 (0.022)\tLoss 0.3747 (0.3219)\tPrec@1 88.672 (88.765)\tPrec@5 99.609 (99.712)\t\nTRAINING - Epoch: [38][70/97]\tTime 0.157 (0.189)\tData 0.000 (0.019)\tLoss 0.3592 (0.3242)\tPrec@1 86.328 (88.622)\tPrec@5 99.609 (99.722)\t\nTRAINING - Epoch: [38][80/97]\tTime 0.168 (0.186)\tData 0.000 (0.017)\tLoss 0.3128 (0.3247)\tPrec@1 90.039 (88.604)\tPrec@5 99.609 (99.715)\t\nTRAINING - Epoch: [38][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.015)\tLoss 0.2854 (0.3256)\tPrec@1 91.211 (88.562)\tPrec@5 99.414 (99.719)\t\nEVALUATING - Epoch: [38][0/20]\tTime 0.981 (0.981)\tData 0.930 (0.930)\tLoss 0.4083 (0.4083)\tPrec@1 85.352 (85.352)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [38][10/20]\tTime 0.057 (0.189)\tData 0.000 (0.135)\tLoss 0.4326 (0.4786)\tPrec@1 85.352 (84.517)\tPrec@5 99.023 (99.290)\t\n\nResults - Epoch: 39\nTraining Loss 0.3262 \tTraining Prec@1 88.547 \tTraining Prec@5 99.712 \tValidation Loss 0.4715 \tValidation Prec@1 84.620 \tValidation Prec@5 99.300 \t\n\n\nStarting Epoch: 40\n\nTRAINING - Epoch: [39][0/97]\tTime 1.027 (1.027)\tData 0.768 (0.768)\tLoss 0.2805 (0.2805)\tPrec@1 89.844 (89.844)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [39][10/97]\tTime 0.188 (0.334)\tData 0.000 (0.088)\tLoss 0.2938 (0.3034)\tPrec@1 90.039 (89.631)\tPrec@5 99.609 (99.716)\t\nTRAINING - Epoch: [39][20/97]\tTime 0.167 (0.253)\tData 0.000 (0.046)\tLoss 0.2480 (0.3003)\tPrec@1 92.188 (89.509)\tPrec@5 99.805 (99.712)\t\nTRAINING - Epoch: [39][30/97]\tTime 0.156 (0.221)\tData 0.007 (0.032)\tLoss 0.3237 (0.3025)\tPrec@1 88.867 (89.352)\tPrec@5 100.000 (99.742)\t\nTRAINING - Epoch: [39][40/97]\tTime 0.183 (0.207)\tData 0.000 (0.024)\tLoss 0.2918 (0.3033)\tPrec@1 90.430 (89.353)\tPrec@5 99.609 (99.757)\t\nTRAINING - Epoch: [39][50/97]\tTime 0.152 (0.198)\tData 0.005 (0.020)\tLoss 0.3295 (0.3141)\tPrec@1 89.258 (89.020)\tPrec@5 99.219 (99.717)\t\nTRAINING - Epoch: [39][60/97]\tTime 0.162 (0.192)\tData 0.000 (0.017)\tLoss 0.3776 (0.3163)\tPrec@1 87.109 (88.922)\tPrec@5 99.805 (99.725)\t\nTRAINING - Epoch: [39][70/97]\tTime 0.178 (0.188)\tData 0.008 (0.015)\tLoss 0.3098 (0.3152)\tPrec@1 90.625 (88.983)\tPrec@5 100.000 (99.728)\t\nTRAINING - Epoch: [39][80/97]\tTime 0.160 (0.185)\tData 0.000 (0.013)\tLoss 0.2918 (0.3161)\tPrec@1 89.453 (88.935)\tPrec@5 99.219 (99.725)\t\nTRAINING - Epoch: [39][90/97]\tTime 0.150 (0.181)\tData 0.000 (0.012)\tLoss 0.3547 (0.3177)\tPrec@1 88.867 (88.891)\tPrec@5 99.805 (99.740)\t\nEVALUATING - Epoch: [39][0/20]\tTime 1.087 (1.087)\tData 1.034 (1.034)\tLoss 0.6338 (0.6338)\tPrec@1 79.688 (79.688)\tPrec@5 99.414 (99.414)\t\nEVALUATING - Epoch: [39][10/20]\tTime 0.058 (0.194)\tData 0.000 (0.138)\tLoss 0.6461 (0.6751)\tPrec@1 77.734 (78.942)\tPrec@5 98.828 (99.023)\t\n\nResults - Epoch: 40\nTraining Loss 0.3199 \tTraining Prec@1 88.831 \tTraining Prec@5 99.728 \tValidation Loss 0.6837 \tValidation Prec@1 78.830 \tValidation Prec@5 98.860 \t\n\n\nStarting Epoch: 41\n\nTRAINING - Epoch: [40][0/97]\tTime 1.258 (1.258)\tData 0.979 (0.979)\tLoss 0.2921 (0.2921)\tPrec@1 89.453 (89.453)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [40][10/97]\tTime 0.147 (0.333)\tData 0.000 (0.091)\tLoss 0.3404 (0.2981)\tPrec@1 89.844 (89.648)\tPrec@5 99.609 (99.805)\t\nTRAINING - Epoch: [40][20/97]\tTime 0.149 (0.251)\tData 0.000 (0.048)\tLoss 0.3452 (0.3105)\tPrec@1 86.328 (89.165)\tPrec@5 100.000 (99.749)\t\nTRAINING - Epoch: [40][30/97]\tTime 0.180 (0.223)\tData 0.005 (0.033)\tLoss 0.2965 (0.3092)\tPrec@1 88.477 (89.233)\tPrec@5 99.609 (99.773)\t\nTRAINING - Epoch: [40][40/97]\tTime 0.154 (0.207)\tData 0.000 (0.025)\tLoss 0.3616 (0.3065)\tPrec@1 88.672 (89.448)\tPrec@5 99.219 (99.743)\t\nTRAINING - Epoch: [40][50/97]\tTime 0.154 (0.197)\tData 0.000 (0.021)\tLoss 0.3393 (0.3081)\tPrec@1 89.258 (89.403)\tPrec@5 100.000 (99.751)\t\nTRAINING - Epoch: [40][60/97]\tTime 0.150 (0.191)\tData 0.000 (0.018)\tLoss 0.2500 (0.3087)\tPrec@1 90.430 (89.322)\tPrec@5 100.000 (99.757)\t\nTRAINING - Epoch: [40][70/97]\tTime 0.162 (0.187)\tData 0.000 (0.015)\tLoss 0.3080 (0.3115)\tPrec@1 89.844 (89.252)\tPrec@5 99.805 (99.725)\t\nTRAINING - Epoch: [40][80/97]\tTime 0.153 (0.184)\tData 0.000 (0.014)\tLoss 0.2519 (0.3108)\tPrec@1 92.578 (89.280)\tPrec@5 99.609 (99.708)\t\nTRAINING - Epoch: [40][90/97]\tTime 0.149 (0.180)\tData 0.000 (0.012)\tLoss 0.2973 (0.3120)\tPrec@1 88.281 (89.249)\tPrec@5 99.805 (99.712)\t\nEVALUATING - Epoch: [40][0/20]\tTime 1.089 (1.089)\tData 1.028 (1.028)\tLoss 0.5318 (0.5318)\tPrec@1 83.008 (83.008)\tPrec@5 99.219 (99.219)\t\nEVALUATING - Epoch: [40][10/20]\tTime 0.045 (0.195)\tData 0.000 (0.143)\tLoss 0.5164 (0.5605)\tPrec@1 84.180 (82.901)\tPrec@5 98.438 (99.094)\t\n\nResults - Epoch: 41\nTraining Loss 0.3132 \tTraining Prec@1 89.207 \tTraining Prec@5 99.710 \tValidation Loss 0.5614 \tValidation Prec@1 82.600 \tValidation Prec@5 99.210 \t\n\n\nStarting Epoch: 42\n\nTRAINING - Epoch: [41][0/97]\tTime 1.363 (1.363)\tData 1.087 (1.087)\tLoss 0.3426 (0.3426)\tPrec@1 89.648 (89.648)\tPrec@5 99.219 (99.219)\t\nTRAINING - Epoch: [41][10/97]\tTime 0.161 (0.332)\tData 0.000 (0.101)\tLoss 0.2963 (0.3034)\tPrec@1 89.258 (89.382)\tPrec@5 99.805 (99.663)\t\nTRAINING - Epoch: [41][20/97]\tTime 0.155 (0.251)\tData 0.000 (0.053)\tLoss 0.3090 (0.3083)\tPrec@1 88.672 (89.183)\tPrec@5 99.805 (99.628)\t\nTRAINING - Epoch: [41][30/97]\tTime 0.173 (0.224)\tData 0.000 (0.036)\tLoss 0.2897 (0.3115)\tPrec@1 90.234 (89.132)\tPrec@5 100.000 (99.716)\t\nTRAINING - Epoch: [41][40/97]\tTime 0.171 (0.208)\tData 0.000 (0.028)\tLoss 0.2248 (0.3131)\tPrec@1 92.773 (89.120)\tPrec@5 100.000 (99.700)\t\nTRAINING - Epoch: [41][50/97]\tTime 0.153 (0.199)\tData 0.000 (0.023)\tLoss 0.3603 (0.3159)\tPrec@1 86.914 (89.024)\tPrec@5 99.609 (99.701)\t\nTRAINING - Epoch: [41][60/97]\tTime 0.154 (0.193)\tData 0.000 (0.019)\tLoss 0.3409 (0.3184)\tPrec@1 87.500 (88.947)\tPrec@5 99.805 (99.696)\t\nTRAINING - Epoch: [41][70/97]\tTime 0.160 (0.188)\tData 0.000 (0.017)\tLoss 0.3349 (0.3183)\tPrec@1 87.891 (88.958)\tPrec@5 99.609 (99.692)\t\nTRAINING - Epoch: [41][80/97]\tTime 0.160 (0.185)\tData 0.000 (0.015)\tLoss 0.3857 (0.3193)\tPrec@1 87.305 (88.850)\tPrec@5 100.000 (99.694)\t\nTRAINING - Epoch: [41][90/97]\tTime 0.150 (0.181)\tData 0.000 (0.013)\tLoss 0.3003 (0.3180)\tPrec@1 90.039 (88.893)\tPrec@5 99.805 (99.695)\t\nEVALUATING - Epoch: [41][0/20]\tTime 1.011 (1.011)\tData 0.960 (0.960)\tLoss 0.5795 (0.5795)\tPrec@1 80.859 (80.859)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [41][10/20]\tTime 0.059 (0.196)\tData 0.000 (0.139)\tLoss 0.5922 (0.5510)\tPrec@1 82.227 (82.653)\tPrec@5 99.219 (99.254)\t\n\nResults - Epoch: 42\nTraining Loss 0.3179 \tTraining Prec@1 88.865 \tTraining Prec@5 99.704 \tValidation Loss 0.5514 \tValidation Prec@1 82.910 \tValidation Prec@5 99.280 \t\n\n\nStarting Epoch: 43\n\nTRAINING - Epoch: [42][0/97]\tTime 1.536 (1.536)\tData 1.238 (1.238)\tLoss 0.3152 (0.3152)\tPrec@1 88.086 (88.086)\tPrec@5 99.805 (99.805)\t\nTRAINING - Epoch: [42][10/97]\tTime 0.157 (0.339)\tData 0.000 (0.115)\tLoss 0.2749 (0.2938)\tPrec@1 90.234 (89.382)\tPrec@5 99.219 (99.698)\t\nTRAINING - Epoch: [42][20/97]\tTime 0.154 (0.255)\tData 0.000 (0.061)\tLoss 0.2695 (0.2857)\tPrec@1 90.039 (89.723)\tPrec@5 100.000 (99.767)\t\nTRAINING - Epoch: [42][30/97]\tTime 0.155 (0.225)\tData 0.005 (0.042)\tLoss 0.2972 (0.2890)\tPrec@1 89.844 (89.617)\tPrec@5 99.805 (99.735)\t\nTRAINING - Epoch: [42][40/97]\tTime 0.150 (0.210)\tData 0.000 (0.032)\tLoss 0.3237 (0.2974)\tPrec@1 89.258 (89.420)\tPrec@5 99.609 (99.719)\t\nTRAINING - Epoch: [42][50/97]\tTime 0.148 (0.200)\tData 0.000 (0.026)\tLoss 0.3276 (0.2962)\tPrec@1 88.477 (89.396)\tPrec@5 99.609 (99.736)\t\nTRAINING - Epoch: [42][60/97]\tTime 0.185 (0.194)\tData 0.005 (0.022)\tLoss 0.3469 (0.2995)\tPrec@1 87.891 (89.293)\tPrec@5 100.000 (99.741)\t\nTRAINING - Epoch: [42][70/97]\tTime 0.159 (0.189)\tData 0.005 (0.019)\tLoss 0.3681 (0.3003)\tPrec@1 86.523 (89.338)\tPrec@5 99.805 (99.736)\t\nTRAINING - Epoch: [42][80/97]\tTime 0.157 (0.186)\tData 0.000 (0.017)\tLoss 0.3054 (0.3040)\tPrec@1 90.430 (89.243)\tPrec@5 100.000 (99.735)\t\nTRAINING - Epoch: [42][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.015)\tLoss 0.3496 (0.3059)\tPrec@1 87.500 (89.204)\tPrec@5 99.805 (99.727)\t\nEVALUATING - Epoch: [42][0/20]\tTime 1.093 (1.093)\tData 1.039 (1.039)\tLoss 0.4657 (0.4657)\tPrec@1 84.766 (84.766)\tPrec@5 100.000 (100.000)\t\nEVALUATING - Epoch: [42][10/20]\tTime 0.049 (0.199)\tData 0.000 (0.142)\tLoss 0.4935 (0.4899)\tPrec@1 85.547 (84.677)\tPrec@5 99.023 (99.450)\t\n\nResults - Epoch: 43\nTraining Loss 0.3056 \tTraining Prec@1 89.258 \tTraining Prec@5 99.722 \tValidation Loss 0.4942 \tValidation Prec@1 84.550 \tValidation Prec@5 99.420 \t\n\n\nStarting Epoch: 44\n\nTRAINING - Epoch: [43][0/97]\tTime 1.513 (1.513)\tData 1.227 (1.227)\tLoss 0.2871 (0.2871)\tPrec@1 91.211 (91.211)\tPrec@5 99.414 (99.414)\t\nTRAINING - Epoch: [43][10/97]\tTime 0.156 (0.339)\tData 0.001 (0.117)\tLoss 0.3066 (0.2777)\tPrec@1 88.477 (90.643)\tPrec@5 100.000 (99.769)\t\nTRAINING - Epoch: [43][20/97]\tTime 0.151 (0.253)\tData 0.000 (0.062)\tLoss 0.3108 (0.2822)\tPrec@1 87.891 (90.030)\tPrec@5 100.000 (99.777)\t\nTRAINING - Epoch: [43][30/97]\tTime 0.177 (0.224)\tData 0.000 (0.042)\tLoss 0.2310 (0.2766)\tPrec@1 90.820 (90.272)\tPrec@5 99.805 (99.792)\t\nTRAINING - Epoch: [43][40/97]\tTime 0.153 (0.209)\tData 0.000 (0.032)\tLoss 0.3154 (0.2869)\tPrec@1 88.281 (89.853)\tPrec@5 99.805 (99.786)\t\nTRAINING - Epoch: [43][50/97]\tTime 0.150 (0.200)\tData 0.000 (0.026)\tLoss 0.2988 (0.2884)\tPrec@1 90.430 (89.828)\tPrec@5 99.609 (99.797)\t\nTRAINING - Epoch: [43][60/97]\tTime 0.154 (0.194)\tData 0.000 (0.022)\tLoss 0.3473 (0.2919)\tPrec@1 86.719 (89.693)\tPrec@5 99.609 (99.776)\t\nTRAINING - Epoch: [43][70/97]\tTime 0.152 (0.190)\tData 0.000 (0.019)\tLoss 0.3285 (0.2928)\tPrec@1 88.477 (89.668)\tPrec@5 99.414 (99.766)\t\nTRAINING - Epoch: [43][80/97]\tTime 0.181 (0.186)\tData 0.000 (0.017)\tLoss 0.3471 (0.2944)\tPrec@1 88.477 (89.624)\tPrec@5 99.609 (99.769)\t\nTRAINING - Epoch: [43][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.015)\tLoss 0.3444 (0.2952)\tPrec@1 87.891 (89.601)\tPrec@5 99.609 (99.768)\t\nEVALUATING - Epoch: [43][0/20]\tTime 0.927 (0.927)\tData 0.876 (0.876)\tLoss 0.5761 (0.5761)\tPrec@1 81.445 (81.445)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [43][10/20]\tTime 0.057 (0.197)\tData 0.000 (0.145)\tLoss 0.4837 (0.5557)\tPrec@1 84.961 (83.203)\tPrec@5 99.414 (99.254)\t\n\nResults - Epoch: 44\nTraining Loss 0.2963 \tTraining Prec@1 89.562 \tTraining Prec@5 99.760 \tValidation Loss 0.5460 \tValidation Prec@1 83.290 \tValidation Prec@5 99.280 \t\n\n\nStarting Epoch: 45\n\nTRAINING - Epoch: [44][0/97]\tTime 1.331 (1.331)\tData 1.099 (1.099)\tLoss 0.2342 (0.2342)\tPrec@1 90.820 (90.820)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [44][10/97]\tTime 0.151 (0.337)\tData 0.000 (0.102)\tLoss 0.2547 (0.2622)\tPrec@1 91.992 (91.087)\tPrec@5 100.000 (99.840)\t\nTRAINING - Epoch: [44][20/97]\tTime 0.154 (0.255)\tData 0.000 (0.054)\tLoss 0.2770 (0.2727)\tPrec@1 90.430 (90.467)\tPrec@5 99.805 (99.814)\t\nTRAINING - Epoch: [44][30/97]\tTime 0.163 (0.225)\tData 0.000 (0.037)\tLoss 0.2879 (0.2807)\tPrec@1 89.062 (90.241)\tPrec@5 100.000 (99.786)\t\nTRAINING - Epoch: [44][40/97]\tTime 0.161 (0.211)\tData 0.000 (0.028)\tLoss 0.3149 (0.2806)\tPrec@1 88.672 (90.201)\tPrec@5 99.414 (99.757)\t\nTRAINING - Epoch: [44][50/97]\tTime 0.155 (0.202)\tData 0.000 (0.023)\tLoss 0.3024 (0.2826)\tPrec@1 88.477 (90.047)\tPrec@5 100.000 (99.778)\t\nTRAINING - Epoch: [44][60/97]\tTime 0.161 (0.195)\tData 0.000 (0.019)\tLoss 0.3109 (0.2858)\tPrec@1 88.281 (89.943)\tPrec@5 100.000 (99.782)\t\nTRAINING - Epoch: [44][70/97]\tTime 0.157 (0.190)\tData 0.000 (0.017)\tLoss 0.2668 (0.2910)\tPrec@1 90.430 (89.805)\tPrec@5 99.805 (99.769)\t\nTRAINING - Epoch: [44][80/97]\tTime 0.152 (0.186)\tData 0.000 (0.015)\tLoss 0.2758 (0.2904)\tPrec@1 90.234 (89.841)\tPrec@5 99.805 (99.769)\t\nTRAINING - Epoch: [44][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.013)\tLoss 0.3079 (0.2926)\tPrec@1 88.867 (89.730)\tPrec@5 99.414 (99.760)\t\nEVALUATING - Epoch: [44][0/20]\tTime 0.972 (0.972)\tData 0.925 (0.925)\tLoss 0.4755 (0.4755)\tPrec@1 82.617 (82.617)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [44][10/20]\tTime 0.048 (0.198)\tData 0.000 (0.144)\tLoss 0.4931 (0.5590)\tPrec@1 84.570 (83.612)\tPrec@5 99.219 (99.077)\t\n\nResults - Epoch: 45\nTraining Loss 0.2925 \tTraining Prec@1 89.719 \tTraining Prec@5 99.766 \tValidation Loss 0.5588 \tValidation Prec@1 83.450 \tValidation Prec@5 99.040 \t\n\n\nStarting Epoch: 46\n\nTRAINING - Epoch: [45][0/97]\tTime 1.519 (1.519)\tData 1.224 (1.224)\tLoss 0.2966 (0.2966)\tPrec@1 87.500 (87.500)\tPrec@5 99.609 (99.609)\t\nTRAINING - Epoch: [45][10/97]\tTime 0.184 (0.339)\tData 0.000 (0.116)\tLoss 0.3137 (0.2882)\tPrec@1 91.016 (90.004)\tPrec@5 99.609 (99.751)\t\nTRAINING - Epoch: [45][20/97]\tTime 0.172 (0.253)\tData 0.005 (0.062)\tLoss 0.2835 (0.2823)\tPrec@1 89.258 (90.179)\tPrec@5 99.805 (99.795)\t\nTRAINING - Epoch: [45][30/97]\tTime 0.153 (0.224)\tData 0.000 (0.042)\tLoss 0.2587 (0.2900)\tPrec@1 90.625 (89.737)\tPrec@5 99.414 (99.786)\t\nTRAINING - Epoch: [45][40/97]\tTime 0.163 (0.210)\tData 0.000 (0.032)\tLoss 0.2604 (0.2897)\tPrec@1 91.992 (89.791)\tPrec@5 99.805 (99.781)\t\nTRAINING - Epoch: [45][50/97]\tTime 0.176 (0.201)\tData 0.000 (0.026)\tLoss 0.2968 (0.2890)\tPrec@1 89.258 (89.817)\tPrec@5 99.414 (99.789)\t\nTRAINING - Epoch: [45][60/97]\tTime 0.155 (0.194)\tData 0.000 (0.022)\tLoss 0.2836 (0.2923)\tPrec@1 90.039 (89.677)\tPrec@5 99.805 (99.776)\t\nTRAINING - Epoch: [45][70/97]\tTime 0.149 (0.189)\tData 0.000 (0.019)\tLoss 0.3458 (0.2938)\tPrec@1 88.867 (89.643)\tPrec@5 99.609 (99.772)\t\nTRAINING - Epoch: [45][80/97]\tTime 0.153 (0.186)\tData 0.000 (0.017)\tLoss 0.3466 (0.2940)\tPrec@1 86.719 (89.581)\tPrec@5 99.414 (99.771)\t\nTRAINING - Epoch: [45][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.015)\tLoss 0.2684 (0.2937)\tPrec@1 91.797 (89.548)\tPrec@5 99.414 (99.772)\t\nEVALUATING - Epoch: [45][0/20]\tTime 1.079 (1.079)\tData 1.012 (1.012)\tLoss 0.5177 (0.5177)\tPrec@1 82.227 (82.227)\tPrec@5 99.219 (99.219)\t\nEVALUATING - Epoch: [45][10/20]\tTime 0.059 (0.187)\tData 0.000 (0.131)\tLoss 0.4774 (0.5493)\tPrec@1 84.375 (82.599)\tPrec@5 99.023 (99.059)\t\n\nResults - Epoch: 46\nTraining Loss 0.2960 \tTraining Prec@1 89.479 \tTraining Prec@5 99.764 \tValidation Loss 0.5472 \tValidation Prec@1 82.550 \tValidation Prec@5 99.220 \t\n\n\nStarting Epoch: 47\n\nTRAINING - Epoch: [46][0/97]\tTime 1.387 (1.387)\tData 1.101 (1.101)\tLoss 0.2795 (0.2795)\tPrec@1 89.062 (89.062)\tPrec@5 99.609 (99.609)\t\nTRAINING - Epoch: [46][10/97]\tTime 0.150 (0.334)\tData 0.000 (0.104)\tLoss 0.3049 (0.2711)\tPrec@1 89.258 (90.572)\tPrec@5 99.805 (99.645)\t\nTRAINING - Epoch: [46][20/97]\tTime 0.151 (0.254)\tData 0.000 (0.055)\tLoss 0.2744 (0.2808)\tPrec@1 88.672 (90.104)\tPrec@5 99.805 (99.712)\t\nTRAINING - Epoch: [46][30/97]\tTime 0.148 (0.224)\tData 0.000 (0.038)\tLoss 0.2798 (0.2763)\tPrec@1 90.039 (90.228)\tPrec@5 99.805 (99.742)\t\nTRAINING - Epoch: [46][40/97]\tTime 0.149 (0.210)\tData 0.000 (0.029)\tLoss 0.2731 (0.2755)\tPrec@1 90.430 (90.215)\tPrec@5 100.000 (99.786)\t\nTRAINING - Epoch: [46][50/97]\tTime 0.171 (0.200)\tData 0.000 (0.024)\tLoss 0.3150 (0.2827)\tPrec@1 88.672 (90.012)\tPrec@5 99.219 (99.774)\t\nTRAINING - Epoch: [46][60/97]\tTime 0.157 (0.194)\tData 0.000 (0.020)\tLoss 0.2614 (0.2855)\tPrec@1 89.453 (89.853)\tPrec@5 100.000 (99.760)\t\nTRAINING - Epoch: [46][70/97]\tTime 0.158 (0.190)\tData 0.000 (0.017)\tLoss 0.3402 (0.2880)\tPrec@1 87.500 (89.747)\tPrec@5 99.805 (99.763)\t\nTRAINING - Epoch: [46][80/97]\tTime 0.161 (0.186)\tData 0.007 (0.015)\tLoss 0.2660 (0.2896)\tPrec@1 90.820 (89.689)\tPrec@5 99.805 (99.761)\t\nTRAINING - Epoch: [46][90/97]\tTime 0.151 (0.182)\tData 0.000 (0.014)\tLoss 0.3098 (0.2943)\tPrec@1 88.281 (89.582)\tPrec@5 100.000 (99.753)\t\nEVALUATING - Epoch: [46][0/20]\tTime 1.377 (1.377)\tData 1.326 (1.326)\tLoss 0.5247 (0.5247)\tPrec@1 82.812 (82.812)\tPrec@5 99.414 (99.414)\t\nEVALUATING - Epoch: [46][10/20]\tTime 0.044 (0.204)\tData 0.000 (0.152)\tLoss 0.4944 (0.5292)\tPrec@1 84.766 (83.345)\tPrec@5 99.414 (99.308)\t\n\nResults - Epoch: 47\nTraining Loss 0.2944 \tTraining Prec@1 89.576 \tTraining Prec@5 99.764 \tValidation Loss 0.5251 \tValidation Prec@1 83.240 \tValidation Prec@5 99.350 \t\n\n\nStarting Epoch: 48\n\nTRAINING - Epoch: [47][0/97]\tTime 1.448 (1.448)\tData 1.166 (1.166)\tLoss 0.3609 (0.3609)\tPrec@1 87.500 (87.500)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [47][10/97]\tTime 0.183 (0.339)\tData 0.000 (0.108)\tLoss 0.3300 (0.2985)\tPrec@1 88.672 (89.577)\tPrec@5 100.000 (99.858)\t\nTRAINING - Epoch: [47][20/97]\tTime 0.168 (0.256)\tData 0.000 (0.057)\tLoss 0.3295 (0.2948)\tPrec@1 89.062 (89.872)\tPrec@5 100.000 (99.833)\t\nTRAINING - Epoch: [47][30/97]\tTime 0.149 (0.226)\tData 0.001 (0.039)\tLoss 0.2673 (0.2886)\tPrec@1 90.820 (89.945)\tPrec@5 99.805 (99.805)\t\nTRAINING - Epoch: [47][40/97]\tTime 0.152 (0.210)\tData 0.000 (0.030)\tLoss 0.2454 (0.2841)\tPrec@1 91.211 (90.134)\tPrec@5 99.805 (99.814)\t\nTRAINING - Epoch: [47][50/97]\tTime 0.155 (0.201)\tData 0.000 (0.024)\tLoss 0.3313 (0.2859)\tPrec@1 90.039 (90.028)\tPrec@5 99.414 (99.816)\t\nTRAINING - Epoch: [47][60/97]\tTime 0.150 (0.195)\tData 0.000 (0.020)\tLoss 0.2923 (0.2886)\tPrec@1 90.430 (89.949)\tPrec@5 99.805 (99.821)\t\nTRAINING - Epoch: [47][70/97]\tTime 0.150 (0.190)\tData 0.000 (0.018)\tLoss 0.2548 (0.2876)\tPrec@1 90.430 (89.987)\tPrec@5 99.805 (99.802)\t\nTRAINING - Epoch: [47][80/97]\tTime 0.158 (0.186)\tData 0.000 (0.016)\tLoss 0.3096 (0.2857)\tPrec@1 89.844 (90.044)\tPrec@5 99.609 (99.797)\t\nTRAINING - Epoch: [47][90/97]\tTime 0.150 (0.183)\tData 0.000 (0.014)\tLoss 0.2893 (0.2861)\tPrec@1 90.039 (90.046)\tPrec@5 99.805 (99.798)\t\nEVALUATING - Epoch: [47][0/20]\tTime 1.113 (1.113)\tData 1.065 (1.065)\tLoss 0.5412 (0.5412)\tPrec@1 81.445 (81.445)\tPrec@5 99.219 (99.219)\t\nEVALUATING - Epoch: [47][10/20]\tTime 0.045 (0.196)\tData 0.000 (0.141)\tLoss 0.6485 (0.5972)\tPrec@1 81.445 (82.102)\tPrec@5 99.023 (98.810)\t\n\nResults - Epoch: 48\nTraining Loss 0.2866 \tTraining Prec@1 89.993 \tTraining Prec@5 99.795 \tValidation Loss 0.6043 \tValidation Prec@1 82.220 \tValidation Prec@5 98.740 \t\n\n\nStarting Epoch: 49\n\nTRAINING - Epoch: [48][0/97]\tTime 1.320 (1.320)\tData 1.042 (1.042)\tLoss 0.2853 (0.2853)\tPrec@1 89.648 (89.648)\tPrec@5 99.609 (99.609)\t\nTRAINING - Epoch: [48][10/97]\tTime 0.151 (0.333)\tData 0.000 (0.098)\tLoss 0.2637 (0.2747)\tPrec@1 91.016 (90.163)\tPrec@5 99.805 (99.822)\t\nTRAINING - Epoch: [48][20/97]\tTime 0.169 (0.252)\tData 0.000 (0.052)\tLoss 0.3247 (0.2725)\tPrec@1 89.844 (90.281)\tPrec@5 99.414 (99.786)\t\nTRAINING - Epoch: [48][30/97]\tTime 0.150 (0.223)\tData 0.000 (0.036)\tLoss 0.2647 (0.2780)\tPrec@1 89.844 (90.077)\tPrec@5 100.000 (99.779)\t\nTRAINING - Epoch: [48][40/97]\tTime 0.185 (0.209)\tData 0.000 (0.028)\tLoss 0.2820 (0.2787)\tPrec@1 90.820 (90.144)\tPrec@5 99.805 (99.786)\t\nTRAINING - Epoch: [48][50/97]\tTime 0.153 (0.200)\tData 0.000 (0.022)\tLoss 0.2801 (0.2780)\tPrec@1 90.234 (90.169)\tPrec@5 99.805 (99.797)\t\nTRAINING - Epoch: [48][60/97]\tTime 0.162 (0.193)\tData 0.007 (0.019)\tLoss 0.2600 (0.2790)\tPrec@1 89.453 (90.180)\tPrec@5 99.805 (99.795)\t\nTRAINING - Epoch: [48][70/97]\tTime 0.166 (0.188)\tData 0.000 (0.017)\tLoss 0.2576 (0.2773)\tPrec@1 90.625 (90.276)\tPrec@5 99.805 (99.783)\t\nTRAINING - Epoch: [48][80/97]\tTime 0.152 (0.185)\tData 0.000 (0.015)\tLoss 0.3207 (0.2787)\tPrec@1 88.086 (90.254)\tPrec@5 99.805 (99.785)\t\nTRAINING - Epoch: [48][90/97]\tTime 0.150 (0.181)\tData 0.000 (0.013)\tLoss 0.2319 (0.2801)\tPrec@1 90.430 (90.191)\tPrec@5 100.000 (99.779)\t\nEVALUATING - Epoch: [48][0/20]\tTime 1.400 (1.400)\tData 1.344 (1.344)\tLoss 0.4657 (0.4657)\tPrec@1 83.398 (83.398)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [48][10/20]\tTime 0.044 (0.205)\tData 0.000 (0.152)\tLoss 0.4743 (0.4926)\tPrec@1 86.523 (84.979)\tPrec@5 99.219 (99.308)\t\n\nResults - Epoch: 49\nTraining Loss 0.2820 \tTraining Prec@1 90.124 \tTraining Prec@5 99.781 \tValidation Loss 0.4932 \tValidation Prec@1 85.030 \tValidation Prec@5 99.370 \t\n\n\nStarting Epoch: 50\n\nTRAINING - Epoch: [49][0/97]\tTime 1.527 (1.527)\tData 1.246 (1.246)\tLoss 0.2889 (0.2889)\tPrec@1 89.648 (89.648)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [49][10/97]\tTime 0.152 (0.337)\tData 0.000 (0.117)\tLoss 0.2559 (0.2690)\tPrec@1 90.430 (90.661)\tPrec@5 100.000 (99.787)\t\nTRAINING - Epoch: [49][20/97]\tTime 0.185 (0.254)\tData 0.000 (0.062)\tLoss 0.2738 (0.2634)\tPrec@1 89.062 (90.606)\tPrec@5 99.609 (99.777)\t\nTRAINING - Epoch: [49][30/97]\tTime 0.148 (0.224)\tData 0.000 (0.043)\tLoss 0.2899 (0.2660)\tPrec@1 89.648 (90.669)\tPrec@5 99.609 (99.742)\t\nTRAINING - Epoch: [49][40/97]\tTime 0.148 (0.209)\tData 0.000 (0.032)\tLoss 0.3027 (0.2678)\tPrec@1 88.086 (90.525)\tPrec@5 99.805 (99.738)\t\nTRAINING - Epoch: [49][50/97]\tTime 0.171 (0.200)\tData 0.005 (0.026)\tLoss 0.2440 (0.2699)\tPrec@1 91.992 (90.518)\tPrec@5 99.805 (99.755)\t\nTRAINING - Epoch: [49][60/97]\tTime 0.153 (0.194)\tData 0.000 (0.022)\tLoss 0.2127 (0.2720)\tPrec@1 92.969 (90.452)\tPrec@5 99.805 (99.757)\t\nTRAINING - Epoch: [49][70/97]\tTime 0.162 (0.189)\tData 0.000 (0.019)\tLoss 0.2717 (0.2741)\tPrec@1 91.211 (90.375)\tPrec@5 100.000 (99.763)\t\nTRAINING - Epoch: [49][80/97]\tTime 0.163 (0.186)\tData 0.000 (0.017)\tLoss 0.3066 (0.2755)\tPrec@1 89.062 (90.295)\tPrec@5 100.000 (99.771)\t\nTRAINING - Epoch: [49][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.015)\tLoss 0.3274 (0.2794)\tPrec@1 88.867 (90.133)\tPrec@5 99.805 (99.764)\t\nEVALUATING - Epoch: [49][0/20]\tTime 0.993 (0.993)\tData 0.947 (0.947)\tLoss 0.6519 (0.6519)\tPrec@1 80.469 (80.469)\tPrec@5 99.414 (99.414)\t\nEVALUATING - Epoch: [49][10/20]\tTime 0.057 (0.197)\tData 0.000 (0.141)\tLoss 0.6405 (0.6842)\tPrec@1 81.445 (80.717)\tPrec@5 99.023 (98.846)\t\n\nResults - Epoch: 50\nTraining Loss 0.2806 \tTraining Prec@1 90.140 \tTraining Prec@5 99.758 \tValidation Loss 0.6839 \tValidation Prec@1 80.170 \tValidation Prec@5 99.020 \t\n\n\nStarting Epoch: 51\n\nTRAINING - Epoch: [50][0/97]\tTime 1.467 (1.467)\tData 1.161 (1.161)\tLoss 0.2536 (0.2536)\tPrec@1 90.820 (90.820)\tPrec@5 99.805 (99.805)\t\nTRAINING - Epoch: [50][10/97]\tTime 0.165 (0.341)\tData 0.000 (0.110)\tLoss 0.2321 (0.2689)\tPrec@1 90.625 (90.252)\tPrec@5 99.805 (99.840)\t\nTRAINING - Epoch: [50][20/97]\tTime 0.171 (0.255)\tData 0.006 (0.058)\tLoss 0.2900 (0.2727)\tPrec@1 89.258 (90.355)\tPrec@5 100.000 (99.851)\t\nTRAINING - Epoch: [50][30/97]\tTime 0.154 (0.224)\tData 0.000 (0.040)\tLoss 0.2397 (0.2659)\tPrec@1 92.578 (90.568)\tPrec@5 99.805 (99.842)\t\nTRAINING - Epoch: [50][40/97]\tTime 0.156 (0.208)\tData 0.005 (0.031)\tLoss 0.2873 (0.2665)\tPrec@1 88.281 (90.454)\tPrec@5 99.805 (99.824)\t\nTRAINING - Epoch: [50][50/97]\tTime 0.157 (0.199)\tData 0.000 (0.025)\tLoss 0.2710 (0.2703)\tPrec@1 90.234 (90.349)\tPrec@5 100.000 (99.820)\t\nTRAINING - Epoch: [50][60/97]\tTime 0.152 (0.193)\tData 0.000 (0.021)\tLoss 0.2683 (0.2721)\tPrec@1 90.234 (90.398)\tPrec@5 99.609 (99.805)\t\nTRAINING - Epoch: [50][70/97]\tTime 0.149 (0.188)\tData 0.000 (0.018)\tLoss 0.2709 (0.2707)\tPrec@1 90.039 (90.372)\tPrec@5 99.805 (99.813)\t\nTRAINING - Epoch: [50][80/97]\tTime 0.159 (0.185)\tData 0.000 (0.016)\tLoss 0.2411 (0.2717)\tPrec@1 91.211 (90.357)\tPrec@5 100.000 (99.817)\t\nTRAINING - Epoch: [50][90/97]\tTime 0.149 (0.181)\tData 0.000 (0.014)\tLoss 0.3029 (0.2736)\tPrec@1 88.281 (90.292)\tPrec@5 100.000 (99.809)\t\nEVALUATING - Epoch: [50][0/20]\tTime 1.210 (1.210)\tData 1.153 (1.153)\tLoss 0.4516 (0.4516)\tPrec@1 83.398 (83.398)\tPrec@5 99.414 (99.414)\t\nEVALUATING - Epoch: [50][10/20]\tTime 0.047 (0.196)\tData 0.000 (0.139)\tLoss 0.4632 (0.5030)\tPrec@1 86.914 (84.624)\tPrec@5 99.414 (99.272)\t\n\nResults - Epoch: 51\nTraining Loss 0.2744 \tTraining Prec@1 90.259 \tTraining Prec@5 99.811 \tValidation Loss 0.4914 \tValidation Prec@1 84.520 \tValidation Prec@5 99.370 \t\n\n\nStarting Epoch: 52\n\nTRAINING - Epoch: [51][0/97]\tTime 1.526 (1.526)\tData 1.265 (1.265)\tLoss 0.2186 (0.2186)\tPrec@1 92.578 (92.578)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [51][10/97]\tTime 0.214 (0.338)\tData 0.000 (0.119)\tLoss 0.2167 (0.2510)\tPrec@1 92.188 (91.264)\tPrec@5 100.000 (99.822)\t\nTRAINING - Epoch: [51][20/97]\tTime 0.147 (0.253)\tData 0.000 (0.063)\tLoss 0.3045 (0.2582)\tPrec@1 89.844 (90.969)\tPrec@5 99.414 (99.758)\t\nTRAINING - Epoch: [51][30/97]\tTime 0.183 (0.224)\tData 0.000 (0.043)\tLoss 0.2695 (0.2575)\tPrec@1 90.430 (91.066)\tPrec@5 100.000 (99.767)\t\nTRAINING - Epoch: [51][40/97]\tTime 0.160 (0.209)\tData 0.000 (0.033)\tLoss 0.2775 (0.2621)\tPrec@1 90.820 (90.882)\tPrec@5 99.609 (99.757)\t\nTRAINING - Epoch: [51][50/97]\tTime 0.170 (0.199)\tData 0.000 (0.026)\tLoss 0.2991 (0.2675)\tPrec@1 90.625 (90.702)\tPrec@5 99.414 (99.759)\t\nTRAINING - Epoch: [51][60/97]\tTime 0.164 (0.193)\tData 0.000 (0.023)\tLoss 0.2522 (0.2696)\tPrec@1 92.578 (90.689)\tPrec@5 99.805 (99.773)\t\nTRAINING - Epoch: [51][70/97]\tTime 0.167 (0.189)\tData 0.005 (0.020)\tLoss 0.2727 (0.2699)\tPrec@1 89.648 (90.589)\tPrec@5 100.000 (99.791)\t\nTRAINING - Epoch: [51][80/97]\tTime 0.150 (0.185)\tData 0.000 (0.017)\tLoss 0.3063 (0.2707)\tPrec@1 89.844 (90.536)\tPrec@5 99.805 (99.795)\t\nTRAINING - Epoch: [51][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.015)\tLoss 0.2997 (0.2715)\tPrec@1 89.453 (90.475)\tPrec@5 99.805 (99.794)\t\nEVALUATING - Epoch: [51][0/20]\tTime 1.025 (1.025)\tData 0.966 (0.966)\tLoss 0.4995 (0.4995)\tPrec@1 83.398 (83.398)\tPrec@5 99.414 (99.414)\t\nEVALUATING - Epoch: [51][10/20]\tTime 0.062 (0.190)\tData 0.000 (0.134)\tLoss 0.5612 (0.5216)\tPrec@1 81.641 (83.434)\tPrec@5 99.219 (99.219)\t\n\nResults - Epoch: 52\nTraining Loss 0.2725 \tTraining Prec@1 90.432 \tTraining Prec@5 99.789 \tValidation Loss 0.5218 \tValidation Prec@1 83.380 \tValidation Prec@5 99.350 \t\n\n\nStarting Epoch: 53\n\nTRAINING - Epoch: [52][0/97]\tTime 1.487 (1.487)\tData 1.194 (1.194)\tLoss 0.2158 (0.2158)\tPrec@1 91.406 (91.406)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [52][10/97]\tTime 0.157 (0.334)\tData 0.000 (0.114)\tLoss 0.2403 (0.2498)\tPrec@1 90.625 (90.962)\tPrec@5 100.000 (99.876)\t\nTRAINING - Epoch: [52][20/97]\tTime 0.181 (0.255)\tData 0.000 (0.061)\tLoss 0.2647 (0.2625)\tPrec@1 89.648 (90.699)\tPrec@5 99.414 (99.823)\t\nTRAINING - Epoch: [52][30/97]\tTime 0.158 (0.225)\tData 0.000 (0.041)\tLoss 0.2754 (0.2615)\tPrec@1 90.234 (90.789)\tPrec@5 99.609 (99.792)\t\nTRAINING - Epoch: [52][40/97]\tTime 0.172 (0.210)\tData 0.000 (0.032)\tLoss 0.2461 (0.2625)\tPrec@1 91.211 (90.749)\tPrec@5 100.000 (99.814)\t\nTRAINING - Epoch: [52][50/97]\tTime 0.157 (0.201)\tData 0.000 (0.026)\tLoss 0.3261 (0.2637)\tPrec@1 89.062 (90.671)\tPrec@5 99.805 (99.820)\t\nTRAINING - Epoch: [52][60/97]\tTime 0.153 (0.195)\tData 0.000 (0.022)\tLoss 0.2796 (0.2627)\tPrec@1 91.016 (90.663)\tPrec@5 99.414 (99.814)\t\nTRAINING - Epoch: [52][70/97]\tTime 0.156 (0.190)\tData 0.000 (0.019)\tLoss 0.3283 (0.2628)\tPrec@1 89.258 (90.691)\tPrec@5 99.805 (99.824)\t\nTRAINING - Epoch: [52][80/97]\tTime 0.156 (0.186)\tData 0.000 (0.017)\tLoss 0.2385 (0.2619)\tPrec@1 90.820 (90.731)\tPrec@5 100.000 (99.826)\t\nTRAINING - Epoch: [52][90/97]\tTime 0.149 (0.182)\tData 0.000 (0.015)\tLoss 0.2734 (0.2626)\tPrec@1 90.234 (90.719)\tPrec@5 99.609 (99.818)\t\nEVALUATING - Epoch: [52][0/20]\tTime 1.004 (1.004)\tData 0.954 (0.954)\tLoss 0.5493 (0.5493)\tPrec@1 83.203 (83.203)\tPrec@5 100.000 (100.000)\t\nEVALUATING - Epoch: [52][10/20]\tTime 0.064 (0.192)\tData 0.000 (0.138)\tLoss 0.6350 (0.6006)\tPrec@1 81.445 (82.457)\tPrec@5 99.023 (99.059)\t\n\nResults - Epoch: 53\nTraining Loss 0.2622 \tTraining Prec@1 90.732 \tTraining Prec@5 99.813 \tValidation Loss 0.6038 \tValidation Prec@1 82.400 \tValidation Prec@5 99.140 \t\n\n\nStarting Epoch: 54\n\nTRAINING - Epoch: [53][0/97]\tTime 1.441 (1.441)\tData 1.163 (1.163)\tLoss 0.2264 (0.2264)\tPrec@1 92.383 (92.383)\tPrec@5 99.609 (99.609)\t\nTRAINING - Epoch: [53][10/97]\tTime 0.155 (0.340)\tData 0.000 (0.110)\tLoss 0.2585 (0.2513)\tPrec@1 90.234 (91.317)\tPrec@5 99.805 (99.787)\t\nTRAINING - Epoch: [53][20/97]\tTime 0.164 (0.256)\tData 0.000 (0.058)\tLoss 0.2429 (0.2612)\tPrec@1 91.016 (90.913)\tPrec@5 100.000 (99.814)\t\nTRAINING - Epoch: [53][30/97]\tTime 0.160 (0.224)\tData 0.000 (0.040)\tLoss 0.2579 (0.2635)\tPrec@1 90.234 (90.915)\tPrec@5 100.000 (99.811)\t\nTRAINING - Epoch: [53][40/97]\tTime 0.170 (0.209)\tData 0.000 (0.030)\tLoss 0.2356 (0.2621)\tPrec@1 91.406 (90.916)\tPrec@5 99.805 (99.805)\t\nTRAINING - Epoch: [53][50/97]\tTime 0.156 (0.200)\tData 0.000 (0.025)\tLoss 0.2904 (0.2625)\tPrec@1 90.430 (90.855)\tPrec@5 100.000 (99.824)\t\nTRAINING - Epoch: [53][60/97]\tTime 0.172 (0.195)\tData 0.000 (0.021)\tLoss 0.1889 (0.2609)\tPrec@1 92.969 (90.862)\tPrec@5 99.805 (99.834)\t\nTRAINING - Epoch: [53][70/97]\tTime 0.160 (0.190)\tData 0.000 (0.018)\tLoss 0.2436 (0.2619)\tPrec@1 91.992 (90.804)\tPrec@5 99.609 (99.824)\t\nTRAINING - Epoch: [53][80/97]\tTime 0.154 (0.187)\tData 0.000 (0.016)\tLoss 0.2644 (0.2629)\tPrec@1 89.844 (90.772)\tPrec@5 99.805 (99.819)\t\nTRAINING - Epoch: [53][90/97]\tTime 0.152 (0.183)\tData 0.000 (0.014)\tLoss 0.2837 (0.2643)\tPrec@1 90.039 (90.685)\tPrec@5 99.805 (99.813)\t\nEVALUATING - Epoch: [53][0/20]\tTime 1.243 (1.243)\tData 1.194 (1.194)\tLoss 0.6212 (0.6212)\tPrec@1 81.055 (81.055)\tPrec@5 99.414 (99.414)\t\nEVALUATING - Epoch: [53][10/20]\tTime 0.052 (0.199)\tData 0.000 (0.144)\tLoss 0.6225 (0.6562)\tPrec@1 82.617 (81.197)\tPrec@5 99.023 (99.148)\t\n\nResults - Epoch: 54\nTraining Loss 0.2644 \tTraining Prec@1 90.677 \tTraining Prec@5 99.815 \tValidation Loss 0.6550 \tValidation Prec@1 81.090 \tValidation Prec@5 99.190 \t\n\n\nStarting Epoch: 55\n\nTRAINING - Epoch: [54][0/97]\tTime 1.408 (1.408)\tData 1.130 (1.130)\tLoss 0.2405 (0.2405)\tPrec@1 91.016 (91.016)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [54][10/97]\tTime 0.152 (0.340)\tData 0.000 (0.106)\tLoss 0.2670 (0.2646)\tPrec@1 90.430 (90.589)\tPrec@5 99.805 (99.840)\t\nTRAINING - Epoch: [54][20/97]\tTime 0.150 (0.255)\tData 0.000 (0.056)\tLoss 0.2069 (0.2526)\tPrec@1 93.164 (91.034)\tPrec@5 99.805 (99.823)\t\nTRAINING - Epoch: [54][30/97]\tTime 0.158 (0.225)\tData 0.000 (0.038)\tLoss 0.2717 (0.2488)\tPrec@1 90.430 (91.224)\tPrec@5 99.805 (99.842)\t\nTRAINING - Epoch: [54][40/97]\tTime 0.180 (0.210)\tData 0.008 (0.030)\tLoss 0.2601 (0.2504)\tPrec@1 90.820 (91.278)\tPrec@5 100.000 (99.829)\t\nTRAINING - Epoch: [54][50/97]\tTime 0.156 (0.200)\tData 0.000 (0.024)\tLoss 0.2852 (0.2544)\tPrec@1 88.672 (91.085)\tPrec@5 99.805 (99.831)\t\nTRAINING - Epoch: [54][60/97]\tTime 0.150 (0.194)\tData 0.000 (0.020)\tLoss 0.2208 (0.2543)\tPrec@1 92.773 (91.131)\tPrec@5 99.805 (99.827)\t\nTRAINING - Epoch: [54][70/97]\tTime 0.162 (0.189)\tData 0.000 (0.018)\tLoss 0.2844 (0.2569)\tPrec@1 89.453 (90.983)\tPrec@5 99.609 (99.827)\t\nTRAINING - Epoch: [54][80/97]\tTime 0.161 (0.186)\tData 0.005 (0.016)\tLoss 0.3061 (0.2587)\tPrec@1 89.258 (90.934)\tPrec@5 99.609 (99.819)\t\nTRAINING - Epoch: [54][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.014)\tLoss 0.2513 (0.2595)\tPrec@1 90.625 (90.878)\tPrec@5 99.805 (99.811)\t\nEVALUATING - Epoch: [54][0/20]\tTime 1.058 (1.058)\tData 0.984 (0.984)\tLoss 0.4649 (0.4649)\tPrec@1 83.789 (83.789)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [54][10/20]\tTime 0.044 (0.199)\tData 0.000 (0.143)\tLoss 0.4912 (0.4943)\tPrec@1 85.742 (84.801)\tPrec@5 99.219 (99.272)\t\n\nResults - Epoch: 55\nTraining Loss 0.2597 \tTraining Prec@1 90.857 \tTraining Prec@5 99.811 \tValidation Loss 0.5002 \tValidation Prec@1 84.640 \tValidation Prec@5 99.350 \t\n\n\nStarting Epoch: 56\n\nTRAINING - Epoch: [55][0/97]\tTime 1.404 (1.404)\tData 1.086 (1.086)\tLoss 0.2130 (0.2130)\tPrec@1 93.164 (93.164)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [55][10/97]\tTime 0.182 (0.338)\tData 0.000 (0.101)\tLoss 0.2924 (0.2621)\tPrec@1 89.844 (90.980)\tPrec@5 99.805 (99.858)\t\nTRAINING - Epoch: [55][20/97]\tTime 0.148 (0.253)\tData 0.000 (0.055)\tLoss 0.2935 (0.2583)\tPrec@1 90.820 (90.997)\tPrec@5 99.805 (99.833)\t\nTRAINING - Epoch: [55][30/97]\tTime 0.198 (0.224)\tData 0.004 (0.037)\tLoss 0.2663 (0.2567)\tPrec@1 90.234 (90.896)\tPrec@5 100.000 (99.842)\t\nTRAINING - Epoch: [55][40/97]\tTime 0.158 (0.210)\tData 0.005 (0.029)\tLoss 0.2284 (0.2534)\tPrec@1 91.602 (90.982)\tPrec@5 100.000 (99.848)\t\nTRAINING - Epoch: [55][50/97]\tTime 0.185 (0.200)\tData 0.000 (0.023)\tLoss 0.2663 (0.2533)\tPrec@1 90.820 (90.920)\tPrec@5 99.805 (99.847)\t\nTRAINING - Epoch: [55][60/97]\tTime 0.162 (0.194)\tData 0.000 (0.020)\tLoss 0.2378 (0.2529)\tPrec@1 92.188 (91.009)\tPrec@5 100.000 (99.840)\t\nTRAINING - Epoch: [55][70/97]\tTime 0.149 (0.188)\tData 0.000 (0.017)\tLoss 0.2847 (0.2530)\tPrec@1 90.039 (91.032)\tPrec@5 99.805 (99.838)\t\nTRAINING - Epoch: [55][80/97]\tTime 0.167 (0.185)\tData 0.000 (0.015)\tLoss 0.2678 (0.2545)\tPrec@1 89.844 (90.967)\tPrec@5 100.000 (99.836)\t\nTRAINING - Epoch: [55][90/97]\tTime 0.149 (0.181)\tData 0.000 (0.014)\tLoss 0.3001 (0.2559)\tPrec@1 87.500 (90.906)\tPrec@5 100.000 (99.839)\t\nEVALUATING - Epoch: [55][0/20]\tTime 0.977 (0.977)\tData 0.933 (0.933)\tLoss 0.5131 (0.5131)\tPrec@1 83.594 (83.594)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [55][10/20]\tTime 0.057 (0.180)\tData 0.000 (0.129)\tLoss 0.4910 (0.5497)\tPrec@1 86.719 (84.375)\tPrec@5 99.219 (99.254)\t\n\nResults - Epoch: 56\nTraining Loss 0.2568 \tTraining Prec@1 90.893 \tTraining Prec@5 99.833 \tValidation Loss 0.5460 \tValidation Prec@1 84.350 \tValidation Prec@5 99.280 \t\n\n\nStarting Epoch: 57\n\nTRAINING - Epoch: [56][0/97]\tTime 1.325 (1.325)\tData 1.040 (1.040)\tLoss 0.2408 (0.2408)\tPrec@1 91.211 (91.211)\tPrec@5 99.805 (99.805)\t\nTRAINING - Epoch: [56][10/97]\tTime 0.168 (0.328)\tData 0.000 (0.099)\tLoss 0.2065 (0.2413)\tPrec@1 92.969 (91.193)\tPrec@5 100.000 (99.787)\t\nTRAINING - Epoch: [56][20/97]\tTime 0.167 (0.251)\tData 0.000 (0.052)\tLoss 0.2427 (0.2353)\tPrec@1 89.844 (91.397)\tPrec@5 99.805 (99.823)\t\nTRAINING - Epoch: [56][30/97]\tTime 0.147 (0.221)\tData 0.000 (0.036)\tLoss 0.2263 (0.2401)\tPrec@1 92.383 (91.501)\tPrec@5 100.000 (99.855)\t\nTRAINING - Epoch: [56][40/97]\tTime 0.150 (0.206)\tData 0.000 (0.027)\tLoss 0.2527 (0.2429)\tPrec@1 90.820 (91.502)\tPrec@5 100.000 (99.824)\t\nTRAINING - Epoch: [56][50/97]\tTime 0.156 (0.197)\tData 0.000 (0.022)\tLoss 0.2226 (0.2473)\tPrec@1 92.578 (91.337)\tPrec@5 99.805 (99.831)\t\nTRAINING - Epoch: [56][60/97]\tTime 0.162 (0.191)\tData 0.000 (0.018)\tLoss 0.2558 (0.2494)\tPrec@1 90.625 (91.233)\tPrec@5 99.805 (99.834)\t\nTRAINING - Epoch: [56][70/97]\tTime 0.148 (0.186)\tData 0.000 (0.016)\tLoss 0.2612 (0.2489)\tPrec@1 90.039 (91.238)\tPrec@5 99.805 (99.835)\t\nTRAINING - Epoch: [56][80/97]\tTime 0.159 (0.183)\tData 0.000 (0.014)\tLoss 0.2524 (0.2516)\tPrec@1 90.820 (91.119)\tPrec@5 100.000 (99.838)\t\nTRAINING - Epoch: [56][90/97]\tTime 0.149 (0.179)\tData 0.000 (0.013)\tLoss 0.2239 (0.2524)\tPrec@1 92.188 (91.071)\tPrec@5 100.000 (99.843)\t\nEVALUATING - Epoch: [56][0/20]\tTime 1.019 (1.019)\tData 0.972 (0.972)\tLoss 0.4733 (0.4733)\tPrec@1 84.766 (84.766)\tPrec@5 100.000 (100.000)\t\nEVALUATING - Epoch: [56][10/20]\tTime 0.054 (0.189)\tData 0.000 (0.136)\tLoss 0.4805 (0.5362)\tPrec@1 85.938 (84.766)\tPrec@5 99.023 (99.254)\t\n\nResults - Epoch: 57\nTraining Loss 0.2535 \tTraining Prec@1 91.044 \tTraining Prec@5 99.839 \tValidation Loss 0.5307 \tValidation Prec@1 84.580 \tValidation Prec@5 99.410 \t\n\n\nStarting Epoch: 58\n\nTRAINING - Epoch: [57][0/97]\tTime 1.467 (1.467)\tData 1.132 (1.132)\tLoss 0.2228 (0.2228)\tPrec@1 91.992 (91.992)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [57][10/97]\tTime 0.152 (0.339)\tData 0.000 (0.107)\tLoss 0.2705 (0.2333)\tPrec@1 91.406 (91.921)\tPrec@5 99.805 (99.929)\t\nTRAINING - Epoch: [57][20/97]\tTime 0.145 (0.256)\tData 0.000 (0.057)\tLoss 0.3051 (0.2348)\tPrec@1 90.039 (91.955)\tPrec@5 99.805 (99.879)\t\nTRAINING - Epoch: [57][30/97]\tTime 0.150 (0.226)\tData 0.000 (0.039)\tLoss 0.2019 (0.2386)\tPrec@1 92.188 (91.816)\tPrec@5 99.805 (99.868)\t\nTRAINING - Epoch: [57][40/97]\tTime 0.172 (0.210)\tData 0.000 (0.030)\tLoss 0.2875 (0.2417)\tPrec@1 89.258 (91.597)\tPrec@5 100.000 (99.838)\t\nTRAINING - Epoch: [57][50/97]\tTime 0.182 (0.202)\tData 0.000 (0.025)\tLoss 0.2060 (0.2446)\tPrec@1 92.383 (91.483)\tPrec@5 100.000 (99.839)\t\nTRAINING - Epoch: [57][60/97]\tTime 0.153 (0.195)\tData 0.000 (0.021)\tLoss 0.2544 (0.2486)\tPrec@1 91.602 (91.301)\tPrec@5 99.805 (99.834)\t\nTRAINING - Epoch: [57][70/97]\tTime 0.149 (0.190)\tData 0.000 (0.018)\tLoss 0.2319 (0.2504)\tPrec@1 91.016 (91.214)\tPrec@5 99.609 (99.835)\t\nTRAINING - Epoch: [57][80/97]\tTime 0.160 (0.186)\tData 0.000 (0.016)\tLoss 0.3066 (0.2509)\tPrec@1 87.891 (91.151)\tPrec@5 100.000 (99.843)\t\nTRAINING - Epoch: [57][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.014)\tLoss 0.2657 (0.2527)\tPrec@1 89.453 (91.046)\tPrec@5 100.000 (99.845)\t\nEVALUATING - Epoch: [57][0/20]\tTime 1.115 (1.115)\tData 1.068 (1.068)\tLoss 0.4834 (0.4834)\tPrec@1 83.203 (83.203)\tPrec@5 100.000 (100.000)\t\nEVALUATING - Epoch: [57][10/20]\tTime 0.045 (0.194)\tData 0.000 (0.141)\tLoss 0.4654 (0.5171)\tPrec@1 85.156 (84.730)\tPrec@5 99.609 (99.361)\t\n\nResults - Epoch: 58\nTraining Loss 0.2530 \tTraining Prec@1 91.032 \tTraining Prec@5 99.849 \tValidation Loss 0.5196 \tValidation Prec@1 84.650 \tValidation Prec@5 99.340 \t\n\n\nStarting Epoch: 59\n\nTRAINING - Epoch: [58][0/97]\tTime 1.261 (1.261)\tData 0.988 (0.988)\tLoss 0.2543 (0.2543)\tPrec@1 90.820 (90.820)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [58][10/97]\tTime 0.159 (0.334)\tData 0.000 (0.093)\tLoss 0.2300 (0.2327)\tPrec@1 91.602 (91.744)\tPrec@5 99.805 (99.858)\t\nTRAINING - Epoch: [58][20/97]\tTime 0.152 (0.254)\tData 0.000 (0.050)\tLoss 0.2675 (0.2343)\tPrec@1 90.430 (91.927)\tPrec@5 99.805 (99.860)\t\nTRAINING - Epoch: [58][30/97]\tTime 0.157 (0.224)\tData 0.000 (0.034)\tLoss 0.2170 (0.2338)\tPrec@1 92.383 (91.791)\tPrec@5 100.000 (99.887)\t\nTRAINING - Epoch: [58][40/97]\tTime 0.164 (0.209)\tData 0.000 (0.026)\tLoss 0.2350 (0.2337)\tPrec@1 91.602 (91.683)\tPrec@5 100.000 (99.881)\t\nTRAINING - Epoch: [58][50/97]\tTime 0.148 (0.199)\tData 0.000 (0.021)\tLoss 0.2579 (0.2367)\tPrec@1 90.039 (91.613)\tPrec@5 99.609 (99.851)\t\nTRAINING - Epoch: [58][60/97]\tTime 0.149 (0.192)\tData 0.000 (0.018)\tLoss 0.2675 (0.2377)\tPrec@1 90.234 (91.528)\tPrec@5 100.000 (99.846)\t\nTRAINING - Epoch: [58][70/97]\tTime 0.159 (0.189)\tData 0.000 (0.016)\tLoss 0.2211 (0.2390)\tPrec@1 91.406 (91.472)\tPrec@5 100.000 (99.851)\t\nTRAINING - Epoch: [58][80/97]\tTime 0.149 (0.185)\tData 0.000 (0.014)\tLoss 0.2543 (0.2411)\tPrec@1 90.039 (91.442)\tPrec@5 100.000 (99.863)\t\nTRAINING - Epoch: [58][90/97]\tTime 0.149 (0.181)\tData 0.000 (0.013)\tLoss 0.2332 (0.2444)\tPrec@1 91.602 (91.340)\tPrec@5 99.805 (99.848)\t\nEVALUATING - Epoch: [58][0/20]\tTime 1.038 (1.038)\tData 0.984 (0.984)\tLoss 0.5082 (0.5082)\tPrec@1 82.422 (82.422)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [58][10/20]\tTime 0.061 (0.189)\tData 0.000 (0.132)\tLoss 0.5563 (0.5273)\tPrec@1 85.156 (84.162)\tPrec@5 99.414 (99.343)\t\n\nResults - Epoch: 59\nTraining Loss 0.2449 \tTraining Prec@1 91.318 \tTraining Prec@5 99.847 \tValidation Loss 0.5218 \tValidation Prec@1 84.600 \tValidation Prec@5 99.380 \t\n\n\nStarting Epoch: 60\n\nTRAINING - Epoch: [59][0/97]\tTime 1.418 (1.418)\tData 1.115 (1.115)\tLoss 0.2424 (0.2424)\tPrec@1 93.359 (93.359)\tPrec@5 99.414 (99.414)\t\nTRAINING - Epoch: [59][10/97]\tTime 0.179 (0.339)\tData 0.000 (0.103)\tLoss 0.2256 (0.2508)\tPrec@1 91.797 (91.264)\tPrec@5 100.000 (99.876)\t\nTRAINING - Epoch: [59][20/97]\tTime 0.163 (0.253)\tData 0.000 (0.054)\tLoss 0.2457 (0.2417)\tPrec@1 91.602 (91.518)\tPrec@5 100.000 (99.870)\t\nTRAINING - Epoch: [59][30/97]\tTime 0.154 (0.224)\tData 0.000 (0.037)\tLoss 0.2146 (0.2392)\tPrec@1 92.188 (91.570)\tPrec@5 99.609 (99.861)\t\nTRAINING - Epoch: [59][40/97]\tTime 0.156 (0.209)\tData 0.007 (0.028)\tLoss 0.2767 (0.2421)\tPrec@1 89.648 (91.340)\tPrec@5 100.000 (99.871)\t\nTRAINING - Epoch: [59][50/97]\tTime 0.149 (0.201)\tData 0.000 (0.023)\tLoss 0.2437 (0.2425)\tPrec@1 90.430 (91.291)\tPrec@5 100.000 (99.858)\t\nTRAINING - Epoch: [59][60/97]\tTime 0.153 (0.194)\tData 0.000 (0.020)\tLoss 0.3038 (0.2462)\tPrec@1 90.234 (91.198)\tPrec@5 99.609 (99.850)\t\nTRAINING - Epoch: [59][70/97]\tTime 0.158 (0.190)\tData 0.000 (0.017)\tLoss 0.2077 (0.2446)\tPrec@1 93.359 (91.255)\tPrec@5 99.805 (99.838)\t\nTRAINING - Epoch: [59][80/97]\tTime 0.148 (0.186)\tData 0.000 (0.015)\tLoss 0.2295 (0.2449)\tPrec@1 91.406 (91.254)\tPrec@5 100.000 (99.846)\t\nTRAINING - Epoch: [59][90/97]\tTime 0.149 (0.182)\tData 0.000 (0.013)\tLoss 0.2146 (0.2461)\tPrec@1 92.773 (91.220)\tPrec@5 99.805 (99.839)\t\nEVALUATING - Epoch: [59][0/20]\tTime 1.113 (1.113)\tData 1.066 (1.066)\tLoss 0.4573 (0.4573)\tPrec@1 85.742 (85.742)\tPrec@5 99.219 (99.219)\t\nEVALUATING - Epoch: [59][10/20]\tTime 0.047 (0.193)\tData 0.000 (0.135)\tLoss 0.5627 (0.5658)\tPrec@1 84.375 (84.038)\tPrec@5 99.219 (99.148)\t\n\nResults - Epoch: 60\nTraining Loss 0.2459 \tTraining Prec@1 91.227 \tTraining Prec@5 99.841 \tValidation Loss 0.5626 \tValidation Prec@1 83.760 \tValidation Prec@5 99.220 \t\n\n\nStarting Epoch: 61\n\nTRAINING - Epoch: [60][0/97]\tTime 1.595 (1.595)\tData 1.337 (1.337)\tLoss 0.2250 (0.2250)\tPrec@1 92.773 (92.773)\tPrec@5 99.609 (99.609)\t\nTRAINING - Epoch: [60][10/97]\tTime 0.146 (0.343)\tData 0.000 (0.124)\tLoss 0.2775 (0.2256)\tPrec@1 90.234 (91.992)\tPrec@5 100.000 (99.840)\t\nTRAINING - Epoch: [60][20/97]\tTime 0.152 (0.259)\tData 0.000 (0.066)\tLoss 0.2528 (0.2342)\tPrec@1 90.234 (91.611)\tPrec@5 99.609 (99.805)\t\nTRAINING - Epoch: [60][30/97]\tTime 0.169 (0.227)\tData 0.000 (0.045)\tLoss 0.2016 (0.2345)\tPrec@1 91.602 (91.513)\tPrec@5 100.000 (99.824)\t\nTRAINING - Epoch: [60][40/97]\tTime 0.150 (0.212)\tData 0.000 (0.034)\tLoss 0.2049 (0.2360)\tPrec@1 93.164 (91.559)\tPrec@5 100.000 (99.843)\t\nTRAINING - Epoch: [60][50/97]\tTime 0.190 (0.202)\tData 0.000 (0.027)\tLoss 0.2535 (0.2393)\tPrec@1 91.016 (91.429)\tPrec@5 100.000 (99.851)\t\nTRAINING - Epoch: [60][60/97]\tTime 0.153 (0.196)\tData 0.000 (0.023)\tLoss 0.1673 (0.2405)\tPrec@1 94.727 (91.384)\tPrec@5 100.000 (99.837)\t\nTRAINING - Epoch: [60][70/97]\tTime 0.159 (0.191)\tData 0.000 (0.020)\tLoss 0.2629 (0.2405)\tPrec@1 91.016 (91.392)\tPrec@5 99.805 (99.838)\t\nTRAINING - Epoch: [60][80/97]\tTime 0.167 (0.187)\tData 0.000 (0.018)\tLoss 0.2789 (0.2418)\tPrec@1 91.016 (91.339)\tPrec@5 99.609 (99.838)\t\nTRAINING - Epoch: [60][90/97]\tTime 0.150 (0.183)\tData 0.000 (0.016)\tLoss 0.1674 (0.2432)\tPrec@1 94.336 (91.338)\tPrec@5 100.000 (99.835)\t\nEVALUATING - Epoch: [60][0/20]\tTime 1.003 (1.003)\tData 0.957 (0.957)\tLoss 0.5001 (0.5001)\tPrec@1 83.594 (83.594)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [60][10/20]\tTime 0.045 (0.197)\tData 0.000 (0.147)\tLoss 0.5587 (0.5692)\tPrec@1 84.766 (83.487)\tPrec@5 99.219 (99.361)\t\n\nResults - Epoch: 61\nTraining Loss 0.2442 \tTraining Prec@1 91.277 \tTraining Prec@5 99.831 \tValidation Loss 0.5741 \tValidation Prec@1 83.410 \tValidation Prec@5 99.400 \t\n\n\nStarting Epoch: 62\n\nTRAINING - Epoch: [61][0/97]\tTime 1.190 (1.190)\tData 0.937 (0.937)\tLoss 0.2130 (0.2130)\tPrec@1 92.578 (92.578)\tPrec@5 99.805 (99.805)\t\nTRAINING - Epoch: [61][10/97]\tTime 0.162 (0.341)\tData 0.000 (0.094)\tLoss 0.2237 (0.2130)\tPrec@1 92.773 (92.667)\tPrec@5 99.805 (99.840)\t\nTRAINING - Epoch: [61][20/97]\tTime 0.155 (0.254)\tData 0.000 (0.050)\tLoss 0.2374 (0.2235)\tPrec@1 90.820 (92.215)\tPrec@5 100.000 (99.823)\t\nTRAINING - Epoch: [61][30/97]\tTime 0.204 (0.226)\tData 0.002 (0.034)\tLoss 0.2374 (0.2315)\tPrec@1 91.992 (91.872)\tPrec@5 99.609 (99.811)\t\nTRAINING - Epoch: [61][40/97]\tTime 0.157 (0.211)\tData 0.000 (0.027)\tLoss 0.2461 (0.2355)\tPrec@1 91.797 (91.721)\tPrec@5 99.609 (99.776)\t\nTRAINING - Epoch: [61][50/97]\tTime 0.153 (0.201)\tData 0.000 (0.022)\tLoss 0.1836 (0.2365)\tPrec@1 93.750 (91.678)\tPrec@5 99.805 (99.801)\t\nTRAINING - Epoch: [61][60/97]\tTime 0.154 (0.195)\tData 0.000 (0.018)\tLoss 0.2417 (0.2397)\tPrec@1 91.406 (91.489)\tPrec@5 99.609 (99.808)\t\nTRAINING - Epoch: [61][70/97]\tTime 0.151 (0.190)\tData 0.000 (0.016)\tLoss 0.2714 (0.2384)\tPrec@1 91.211 (91.566)\tPrec@5 99.609 (99.824)\t\nTRAINING - Epoch: [61][80/97]\tTime 0.163 (0.187)\tData 0.000 (0.014)\tLoss 0.2631 (0.2396)\tPrec@1 90.430 (91.532)\tPrec@5 100.000 (99.819)\t\nTRAINING - Epoch: [61][90/97]\tTime 0.149 (0.183)\tData 0.000 (0.013)\tLoss 0.2565 (0.2416)\tPrec@1 91.992 (91.458)\tPrec@5 99.609 (99.822)\t\nEVALUATING - Epoch: [61][0/20]\tTime 1.037 (1.037)\tData 0.985 (0.985)\tLoss 0.5306 (0.5306)\tPrec@1 81.250 (81.250)\tPrec@5 100.000 (100.000)\t\nEVALUATING - Epoch: [61][10/20]\tTime 0.060 (0.195)\tData 0.000 (0.140)\tLoss 0.5601 (0.5287)\tPrec@1 83.984 (83.825)\tPrec@5 99.219 (99.450)\t\n\nResults - Epoch: 62\nTraining Loss 0.2428 \tTraining Prec@1 91.410 \tTraining Prec@5 99.831 \tValidation Loss 0.5343 \tValidation Prec@1 83.830 \tValidation Prec@5 99.450 \t\n\n\nStarting Epoch: 63\n\nTRAINING - Epoch: [62][0/97]\tTime 1.502 (1.502)\tData 1.201 (1.201)\tLoss 0.2397 (0.2397)\tPrec@1 91.211 (91.211)\tPrec@5 99.609 (99.609)\t\nTRAINING - Epoch: [62][10/97]\tTime 0.191 (0.338)\tData 0.000 (0.112)\tLoss 0.2045 (0.2250)\tPrec@1 93.555 (92.383)\tPrec@5 99.805 (99.876)\t\nTRAINING - Epoch: [62][20/97]\tTime 0.165 (0.256)\tData 0.000 (0.060)\tLoss 0.2311 (0.2277)\tPrec@1 91.602 (92.206)\tPrec@5 100.000 (99.898)\t\nTRAINING - Epoch: [62][30/97]\tTime 0.164 (0.227)\tData 0.004 (0.041)\tLoss 0.2196 (0.2321)\tPrec@1 91.602 (91.891)\tPrec@5 100.000 (99.899)\t\nTRAINING - Epoch: [62][40/97]\tTime 0.159 (0.211)\tData 0.000 (0.031)\tLoss 0.2327 (0.2342)\tPrec@1 92.969 (91.778)\tPrec@5 99.609 (99.871)\t\nTRAINING - Epoch: [62][50/97]\tTime 0.177 (0.201)\tData 0.000 (0.025)\tLoss 0.2621 (0.2339)\tPrec@1 90.234 (91.793)\tPrec@5 99.609 (99.862)\t\nTRAINING - Epoch: [62][60/97]\tTime 0.155 (0.195)\tData 0.005 (0.021)\tLoss 0.2277 (0.2360)\tPrec@1 92.188 (91.666)\tPrec@5 100.000 (99.872)\t\nTRAINING - Epoch: [62][70/97]\tTime 0.156 (0.190)\tData 0.000 (0.019)\tLoss 0.2194 (0.2373)\tPrec@1 92.383 (91.643)\tPrec@5 99.805 (99.876)\t\nTRAINING - Epoch: [62][80/97]\tTime 0.153 (0.186)\tData 0.000 (0.017)\tLoss 0.2766 (0.2364)\tPrec@1 90.234 (91.664)\tPrec@5 100.000 (99.875)\t\nTRAINING - Epoch: [62][90/97]\tTime 0.149 (0.182)\tData 0.000 (0.015)\tLoss 0.2400 (0.2391)\tPrec@1 93.164 (91.569)\tPrec@5 99.805 (99.876)\t\nEVALUATING - Epoch: [62][0/20]\tTime 1.183 (1.183)\tData 1.131 (1.131)\tLoss 0.3947 (0.3947)\tPrec@1 84.375 (84.375)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [62][10/20]\tTime 0.052 (0.194)\tData 0.000 (0.134)\tLoss 0.4316 (0.4570)\tPrec@1 88.086 (85.707)\tPrec@5 98.828 (99.379)\t\n\nResults - Epoch: 63\nTraining Loss 0.2378 \tTraining Prec@1 91.630 \tTraining Prec@5 99.875 \tValidation Loss 0.4546 \tValidation Prec@1 85.870 \tValidation Prec@5 99.450 \t\n\n\nStarting Epoch: 64\n\nTRAINING - Epoch: [63][0/97]\tTime 1.507 (1.507)\tData 1.259 (1.259)\tLoss 0.2445 (0.2445)\tPrec@1 92.383 (92.383)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [63][10/97]\tTime 0.184 (0.333)\tData 0.000 (0.118)\tLoss 0.2043 (0.2229)\tPrec@1 92.969 (92.134)\tPrec@5 100.000 (99.929)\t\nTRAINING - Epoch: [63][20/97]\tTime 0.176 (0.255)\tData 0.000 (0.063)\tLoss 0.1733 (0.2177)\tPrec@1 93.750 (92.374)\tPrec@5 100.000 (99.907)\t\nTRAINING - Epoch: [63][30/97]\tTime 0.159 (0.225)\tData 0.000 (0.043)\tLoss 0.2112 (0.2232)\tPrec@1 93.945 (92.074)\tPrec@5 100.000 (99.899)\t\nTRAINING - Epoch: [63][40/97]\tTime 0.171 (0.209)\tData 0.000 (0.033)\tLoss 0.2658 (0.2253)\tPrec@1 91.211 (92.006)\tPrec@5 100.000 (99.881)\t\nTRAINING - Epoch: [63][50/97]\tTime 0.147 (0.200)\tData 0.000 (0.026)\tLoss 0.2085 (0.2262)\tPrec@1 92.969 (91.931)\tPrec@5 100.000 (99.889)\t\nTRAINING - Epoch: [63][60/97]\tTime 0.160 (0.193)\tData 0.000 (0.022)\tLoss 0.3052 (0.2297)\tPrec@1 91.016 (91.842)\tPrec@5 99.219 (99.875)\t\nTRAINING - Epoch: [63][70/97]\tTime 0.156 (0.189)\tData 0.000 (0.019)\tLoss 0.2613 (0.2307)\tPrec@1 90.820 (91.783)\tPrec@5 99.805 (99.873)\t\nTRAINING - Epoch: [63][80/97]\tTime 0.189 (0.186)\tData 0.005 (0.017)\tLoss 0.2313 (0.2328)\tPrec@1 92.383 (91.717)\tPrec@5 99.805 (99.863)\t\nTRAINING - Epoch: [63][90/97]\tTime 0.149 (0.182)\tData 0.000 (0.015)\tLoss 0.2152 (0.2308)\tPrec@1 92.383 (91.801)\tPrec@5 100.000 (99.869)\t\nEVALUATING - Epoch: [63][0/20]\tTime 1.049 (1.049)\tData 0.977 (0.977)\tLoss 0.4539 (0.4539)\tPrec@1 83.984 (83.984)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [63][10/20]\tTime 0.047 (0.196)\tData 0.000 (0.140)\tLoss 0.5627 (0.5309)\tPrec@1 82.422 (84.109)\tPrec@5 99.219 (99.165)\t\n\nResults - Epoch: 64\nTraining Loss 0.2309 \tTraining Prec@1 91.785 \tTraining Prec@5 99.873 \tValidation Loss 0.5378 \tValidation Prec@1 83.960 \tValidation Prec@5 99.170 \t\n\n\nStarting Epoch: 65\n\nTRAINING - Epoch: [64][0/97]\tTime 1.375 (1.375)\tData 1.097 (1.097)\tLoss 0.2132 (0.2132)\tPrec@1 92.969 (92.969)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [64][10/97]\tTime 0.147 (0.337)\tData 0.000 (0.105)\tLoss 0.2080 (0.2015)\tPrec@1 92.578 (92.809)\tPrec@5 100.000 (99.947)\t\nTRAINING - Epoch: [64][20/97]\tTime 0.168 (0.256)\tData 0.000 (0.056)\tLoss 0.1905 (0.2126)\tPrec@1 93.750 (92.615)\tPrec@5 99.805 (99.888)\t\nTRAINING - Epoch: [64][30/97]\tTime 0.189 (0.226)\tData 0.000 (0.038)\tLoss 0.2318 (0.2196)\tPrec@1 91.016 (92.156)\tPrec@5 100.000 (99.880)\t\nTRAINING - Epoch: [64][40/97]\tTime 0.148 (0.209)\tData 0.000 (0.029)\tLoss 0.2278 (0.2229)\tPrec@1 91.797 (92.097)\tPrec@5 100.000 (99.871)\t\nTRAINING - Epoch: [64][50/97]\tTime 0.150 (0.200)\tData 0.000 (0.024)\tLoss 0.2259 (0.2249)\tPrec@1 92.188 (91.977)\tPrec@5 100.000 (99.858)\t\nTRAINING - Epoch: [64][60/97]\tTime 0.150 (0.193)\tData 0.000 (0.020)\tLoss 0.1774 (0.2253)\tPrec@1 92.773 (91.973)\tPrec@5 100.000 (99.856)\t\nTRAINING - Epoch: [64][70/97]\tTime 0.150 (0.189)\tData 0.000 (0.017)\tLoss 0.2135 (0.2246)\tPrec@1 92.383 (92.017)\tPrec@5 100.000 (99.862)\t\nTRAINING - Epoch: [64][80/97]\tTime 0.155 (0.185)\tData 0.000 (0.015)\tLoss 0.1422 (0.2263)\tPrec@1 96.484 (91.932)\tPrec@5 99.805 (99.853)\t\nTRAINING - Epoch: [64][90/97]\tTime 0.149 (0.182)\tData 0.000 (0.014)\tLoss 0.2409 (0.2270)\tPrec@1 92.383 (91.913)\tPrec@5 100.000 (99.854)\t\nEVALUATING - Epoch: [64][0/20]\tTime 1.046 (1.046)\tData 0.985 (0.985)\tLoss 0.6193 (0.6193)\tPrec@1 81.836 (81.836)\tPrec@5 99.023 (99.023)\t\nEVALUATING - Epoch: [64][10/20]\tTime 0.050 (0.194)\tData 0.000 (0.141)\tLoss 0.5939 (0.6347)\tPrec@1 84.766 (82.546)\tPrec@5 99.023 (98.828)\t\n\nResults - Epoch: 65\nTraining Loss 0.2273 \tTraining Prec@1 91.877 \tTraining Prec@5 99.859 \tValidation Loss 0.6296 \tValidation Prec@1 82.610 \tValidation Prec@5 98.840 \t\n\n\nStarting Epoch: 66\n\nTRAINING - Epoch: [65][0/97]\tTime 1.412 (1.412)\tData 1.122 (1.122)\tLoss 0.2300 (0.2300)\tPrec@1 91.406 (91.406)\tPrec@5 99.805 (99.805)\t\nTRAINING - Epoch: [65][10/97]\tTime 0.198 (0.340)\tData 0.005 (0.108)\tLoss 0.2476 (0.2349)\tPrec@1 91.016 (91.602)\tPrec@5 99.805 (99.893)\t\nTRAINING - Epoch: [65][20/97]\tTime 0.165 (0.255)\tData 0.003 (0.057)\tLoss 0.2285 (0.2242)\tPrec@1 91.602 (92.076)\tPrec@5 100.000 (99.898)\t\nTRAINING - Epoch: [65][30/97]\tTime 0.155 (0.224)\tData 0.000 (0.039)\tLoss 0.2181 (0.2241)\tPrec@1 94.727 (92.112)\tPrec@5 99.805 (99.899)\t\nTRAINING - Epoch: [65][40/97]\tTime 0.154 (0.209)\tData 0.000 (0.029)\tLoss 0.2481 (0.2227)\tPrec@1 90.430 (92.178)\tPrec@5 99.805 (99.890)\t\nTRAINING - Epoch: [65][50/97]\tTime 0.155 (0.201)\tData 0.000 (0.024)\tLoss 0.2470 (0.2211)\tPrec@1 90.430 (92.260)\tPrec@5 99.805 (99.881)\t\nTRAINING - Epoch: [65][60/97]\tTime 0.161 (0.194)\tData 0.000 (0.020)\tLoss 0.2508 (0.2237)\tPrec@1 91.992 (92.123)\tPrec@5 100.000 (99.891)\t\nTRAINING - Epoch: [65][70/97]\tTime 0.151 (0.190)\tData 0.000 (0.018)\tLoss 0.1738 (0.2237)\tPrec@1 93.945 (92.091)\tPrec@5 99.805 (99.884)\t\nTRAINING - Epoch: [65][80/97]\tTime 0.155 (0.186)\tData 0.000 (0.016)\tLoss 0.2259 (0.2273)\tPrec@1 91.211 (91.944)\tPrec@5 100.000 (99.889)\t\nTRAINING - Epoch: [65][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.014)\tLoss 0.2653 (0.2318)\tPrec@1 90.820 (91.793)\tPrec@5 99.805 (99.888)\t\nEVALUATING - Epoch: [65][0/20]\tTime 1.180 (1.180)\tData 1.130 (1.130)\tLoss 0.4897 (0.4897)\tPrec@1 82.812 (82.812)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [65][10/20]\tTime 0.053 (0.198)\tData 0.000 (0.142)\tLoss 0.4837 (0.5091)\tPrec@1 86.328 (85.281)\tPrec@5 99.219 (99.432)\t\n\nResults - Epoch: 66\nTraining Loss 0.2333 \tTraining Prec@1 91.747 \tTraining Prec@5 99.879 \tValidation Loss 0.5168 \tValidation Prec@1 84.830 \tValidation Prec@5 99.420 \t\n\n\nStarting Epoch: 67\n\nTRAINING - Epoch: [66][0/97]\tTime 1.242 (1.242)\tData 0.990 (0.990)\tLoss 0.2691 (0.2691)\tPrec@1 91.016 (91.016)\tPrec@5 99.805 (99.805)\t\nTRAINING - Epoch: [66][10/97]\tTime 0.170 (0.338)\tData 0.004 (0.093)\tLoss 0.2441 (0.2193)\tPrec@1 91.406 (92.383)\tPrec@5 99.805 (99.858)\t\nTRAINING - Epoch: [66][20/97]\tTime 0.154 (0.255)\tData 0.000 (0.049)\tLoss 0.2208 (0.2153)\tPrec@1 91.992 (92.448)\tPrec@5 100.000 (99.860)\t\nTRAINING - Epoch: [66][30/97]\tTime 0.156 (0.225)\tData 0.000 (0.034)\tLoss 0.2289 (0.2178)\tPrec@1 92.969 (92.295)\tPrec@5 100.000 (99.874)\t\nTRAINING - Epoch: [66][40/97]\tTime 0.152 (0.209)\tData 0.005 (0.026)\tLoss 0.2130 (0.2186)\tPrec@1 91.992 (92.283)\tPrec@5 100.000 (99.871)\t\nTRAINING - Epoch: [66][50/97]\tTime 0.156 (0.200)\tData 0.000 (0.021)\tLoss 0.2257 (0.2214)\tPrec@1 91.797 (92.168)\tPrec@5 99.805 (99.877)\t\nTRAINING - Epoch: [66][60/97]\tTime 0.148 (0.194)\tData 0.000 (0.018)\tLoss 0.1910 (0.2212)\tPrec@1 93.359 (92.255)\tPrec@5 100.000 (99.872)\t\nTRAINING - Epoch: [66][70/97]\tTime 0.192 (0.190)\tData 0.000 (0.016)\tLoss 0.2315 (0.2235)\tPrec@1 92.578 (92.149)\tPrec@5 100.000 (99.871)\t\nTRAINING - Epoch: [66][80/97]\tTime 0.167 (0.186)\tData 0.000 (0.014)\tLoss 0.2114 (0.2253)\tPrec@1 93.359 (92.089)\tPrec@5 99.609 (99.877)\t\nTRAINING - Epoch: [66][90/97]\tTime 0.151 (0.182)\tData 0.000 (0.012)\tLoss 0.1645 (0.2258)\tPrec@1 92.969 (92.046)\tPrec@5 100.000 (99.871)\t\nEVALUATING - Epoch: [66][0/20]\tTime 1.117 (1.117)\tData 1.071 (1.071)\tLoss 0.4763 (0.4763)\tPrec@1 82.422 (82.422)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [66][10/20]\tTime 0.055 (0.192)\tData 0.000 (0.136)\tLoss 0.4726 (0.5346)\tPrec@1 85.938 (84.499)\tPrec@5 99.609 (99.450)\t\n\nResults - Epoch: 67\nTraining Loss 0.2264 \tTraining Prec@1 92.028 \tTraining Prec@5 99.871 \tValidation Loss 0.5342 \tValidation Prec@1 84.540 \tValidation Prec@5 99.470 \t\n\n\nStarting Epoch: 68\n\nTRAINING - Epoch: [67][0/97]\tTime 1.505 (1.505)\tData 1.240 (1.240)\tLoss 0.2132 (0.2132)\tPrec@1 93.359 (93.359)\tPrec@5 99.414 (99.414)\t\nTRAINING - Epoch: [67][10/97]\tTime 0.154 (0.340)\tData 0.000 (0.117)\tLoss 0.2688 (0.2119)\tPrec@1 89.648 (92.507)\tPrec@5 99.609 (99.840)\t\nTRAINING - Epoch: [67][20/97]\tTime 0.151 (0.254)\tData 0.000 (0.062)\tLoss 0.2436 (0.2186)\tPrec@1 91.211 (92.215)\tPrec@5 99.805 (99.833)\t\nTRAINING - Epoch: [67][30/97]\tTime 0.160 (0.224)\tData 0.000 (0.042)\tLoss 0.1884 (0.2150)\tPrec@1 94.141 (92.251)\tPrec@5 99.805 (99.861)\t\nTRAINING - Epoch: [67][40/97]\tTime 0.159 (0.209)\tData 0.000 (0.032)\tLoss 0.3038 (0.2214)\tPrec@1 89.062 (91.964)\tPrec@5 99.805 (99.862)\t\nTRAINING - Epoch: [67][50/97]\tTime 0.160 (0.199)\tData 0.000 (0.026)\tLoss 0.2308 (0.2207)\tPrec@1 91.797 (92.000)\tPrec@5 99.805 (99.866)\t\nTRAINING - Epoch: [67][60/97]\tTime 0.165 (0.193)\tData 0.000 (0.022)\tLoss 0.1707 (0.2220)\tPrec@1 92.969 (91.970)\tPrec@5 100.000 (99.878)\t\nTRAINING - Epoch: [67][70/97]\tTime 0.152 (0.188)\tData 0.000 (0.019)\tLoss 0.2933 (0.2246)\tPrec@1 88.672 (91.868)\tPrec@5 100.000 (99.873)\t\nTRAINING - Epoch: [67][80/97]\tTime 0.150 (0.185)\tData 0.000 (0.017)\tLoss 0.2224 (0.2248)\tPrec@1 91.602 (91.879)\tPrec@5 99.805 (99.877)\t\nTRAINING - Epoch: [67][90/97]\tTime 0.149 (0.181)\tData 0.000 (0.015)\tLoss 0.2091 (0.2251)\tPrec@1 92.188 (91.908)\tPrec@5 99.805 (99.876)\t\nEVALUATING - Epoch: [67][0/20]\tTime 1.352 (1.352)\tData 1.290 (1.290)\tLoss 0.4701 (0.4701)\tPrec@1 83.789 (83.789)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [67][10/20]\tTime 0.051 (0.206)\tData 0.000 (0.151)\tLoss 0.5034 (0.5207)\tPrec@1 83.594 (84.109)\tPrec@5 99.414 (99.379)\t\n\nResults - Epoch: 68\nTraining Loss 0.2256 \tTraining Prec@1 91.954 \tTraining Prec@5 99.875 \tValidation Loss 0.5185 \tValidation Prec@1 84.460 \tValidation Prec@5 99.440 \t\n\n\nStarting Epoch: 69\n\nTRAINING - Epoch: [68][0/97]\tTime 1.522 (1.522)\tData 1.209 (1.209)\tLoss 0.1792 (0.1792)\tPrec@1 93.164 (93.164)\tPrec@5 99.609 (99.609)\t\nTRAINING - Epoch: [68][10/97]\tTime 0.148 (0.343)\tData 0.000 (0.112)\tLoss 0.2264 (0.2178)\tPrec@1 92.578 (92.383)\tPrec@5 99.805 (99.769)\t\nTRAINING - Epoch: [68][20/97]\tTime 0.165 (0.259)\tData 0.000 (0.060)\tLoss 0.2774 (0.2276)\tPrec@1 90.430 (91.983)\tPrec@5 100.000 (99.814)\t\nTRAINING - Epoch: [68][30/97]\tTime 0.183 (0.229)\tData 0.000 (0.041)\tLoss 0.2488 (0.2257)\tPrec@1 91.016 (92.017)\tPrec@5 100.000 (99.830)\t\nTRAINING - Epoch: [68][40/97]\tTime 0.156 (0.213)\tData 0.000 (0.031)\tLoss 0.2850 (0.2266)\tPrec@1 90.430 (91.983)\tPrec@5 99.805 (99.838)\t\nTRAINING - Epoch: [68][50/97]\tTime 0.174 (0.203)\tData 0.000 (0.025)\tLoss 0.2490 (0.2267)\tPrec@1 91.406 (91.954)\tPrec@5 99.805 (99.847)\t\nTRAINING - Epoch: [68][60/97]\tTime 0.161 (0.197)\tData 0.000 (0.022)\tLoss 0.2374 (0.2258)\tPrec@1 90.430 (91.979)\tPrec@5 99.609 (99.853)\t\nTRAINING - Epoch: [68][70/97]\tTime 0.157 (0.192)\tData 0.000 (0.019)\tLoss 0.1606 (0.2237)\tPrec@1 94.531 (92.009)\tPrec@5 99.805 (99.854)\t\nTRAINING - Epoch: [68][80/97]\tTime 0.169 (0.188)\tData 0.000 (0.016)\tLoss 0.1946 (0.2234)\tPrec@1 93.555 (92.036)\tPrec@5 100.000 (99.863)\t\nTRAINING - Epoch: [68][90/97]\tTime 0.150 (0.184)\tData 0.000 (0.015)\tLoss 0.2442 (0.2239)\tPrec@1 91.406 (91.988)\tPrec@5 99.805 (99.863)\t\nEVALUATING - Epoch: [68][0/20]\tTime 1.122 (1.122)\tData 1.078 (1.078)\tLoss 0.3316 (0.3316)\tPrec@1 88.477 (88.477)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [68][10/20]\tTime 0.056 (0.198)\tData 0.000 (0.144)\tLoss 0.4669 (0.4437)\tPrec@1 85.352 (86.541)\tPrec@5 99.414 (99.485)\t\n\nResults - Epoch: 69\nTraining Loss 0.2247 \tTraining Prec@1 91.974 \tTraining Prec@5 99.857 \tValidation Loss 0.4491 \tValidation Prec@1 86.280 \tValidation Prec@5 99.430 \t\n\n\nStarting Epoch: 70\n\nTRAINING - Epoch: [69][0/97]\tTime 1.428 (1.428)\tData 1.145 (1.145)\tLoss 0.2140 (0.2140)\tPrec@1 92.383 (92.383)\tPrec@5 99.805 (99.805)\t\nTRAINING - Epoch: [69][10/97]\tTime 0.176 (0.338)\tData 0.005 (0.110)\tLoss 0.1945 (0.2022)\tPrec@1 93.750 (92.898)\tPrec@5 99.805 (99.858)\t\nTRAINING - Epoch: [69][20/97]\tTime 0.160 (0.252)\tData 0.000 (0.058)\tLoss 0.2523 (0.2038)\tPrec@1 90.820 (92.671)\tPrec@5 99.609 (99.879)\t\nTRAINING - Epoch: [69][30/97]\tTime 0.174 (0.224)\tData 0.000 (0.039)\tLoss 0.2263 (0.2040)\tPrec@1 90.625 (92.773)\tPrec@5 99.805 (99.861)\t\nTRAINING - Epoch: [69][40/97]\tTime 0.168 (0.208)\tData 0.000 (0.030)\tLoss 0.2530 (0.2081)\tPrec@1 91.211 (92.564)\tPrec@5 99.805 (99.867)\t\nTRAINING - Epoch: [69][50/97]\tTime 0.180 (0.200)\tData 0.000 (0.024)\tLoss 0.2020 (0.2094)\tPrec@1 94.727 (92.605)\tPrec@5 99.805 (99.877)\t\nTRAINING - Epoch: [69][60/97]\tTime 0.184 (0.193)\tData 0.000 (0.021)\tLoss 0.2225 (0.2102)\tPrec@1 91.602 (92.565)\tPrec@5 100.000 (99.866)\t\nTRAINING - Epoch: [69][70/97]\tTime 0.161 (0.189)\tData 0.000 (0.018)\tLoss 0.2690 (0.2128)\tPrec@1 91.016 (92.432)\tPrec@5 100.000 (99.873)\t\nTRAINING - Epoch: [69][80/97]\tTime 0.150 (0.185)\tData 0.000 (0.016)\tLoss 0.2532 (0.2137)\tPrec@1 91.211 (92.426)\tPrec@5 99.414 (99.870)\t\nTRAINING - Epoch: [69][90/97]\tTime 0.149 (0.182)\tData 0.000 (0.014)\tLoss 0.2231 (0.2143)\tPrec@1 92.578 (92.404)\tPrec@5 99.609 (99.865)\t\nEVALUATING - Epoch: [69][0/20]\tTime 1.006 (1.006)\tData 0.958 (0.958)\tLoss 0.5168 (0.5168)\tPrec@1 84.180 (84.180)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [69][10/20]\tTime 0.043 (0.194)\tData 0.000 (0.142)\tLoss 0.6146 (0.5896)\tPrec@1 83.008 (83.487)\tPrec@5 99.023 (99.112)\t\n\nResults - Epoch: 70\nTraining Loss 0.2154 \tTraining Prec@1 92.371 \tTraining Prec@5 99.867 \tValidation Loss 0.5874 \tValidation Prec@1 83.610 \tValidation Prec@5 99.090 \t\n\n\nStarting Epoch: 71\n\nTRAINING - Epoch: [70][0/97]\tTime 1.350 (1.350)\tData 1.073 (1.073)\tLoss 0.2304 (0.2304)\tPrec@1 91.211 (91.211)\tPrec@5 99.805 (99.805)\t\nTRAINING - Epoch: [70][10/97]\tTime 0.164 (0.339)\tData 0.001 (0.102)\tLoss 0.2582 (0.2308)\tPrec@1 91.797 (92.099)\tPrec@5 100.000 (99.893)\t\nTRAINING - Epoch: [70][20/97]\tTime 0.179 (0.255)\tData 0.005 (0.054)\tLoss 0.1445 (0.2159)\tPrec@1 95.312 (92.587)\tPrec@5 100.000 (99.879)\t\nTRAINING - Epoch: [70][30/97]\tTime 0.157 (0.225)\tData 0.000 (0.037)\tLoss 0.2091 (0.2128)\tPrec@1 92.383 (92.647)\tPrec@5 100.000 (99.880)\t\nTRAINING - Epoch: [70][40/97]\tTime 0.193 (0.211)\tData 0.000 (0.028)\tLoss 0.2044 (0.2090)\tPrec@1 91.602 (92.697)\tPrec@5 100.000 (99.895)\t\nTRAINING - Epoch: [70][50/97]\tTime 0.160 (0.201)\tData 0.000 (0.023)\tLoss 0.2168 (0.2097)\tPrec@1 92.969 (92.662)\tPrec@5 99.805 (99.885)\t\nTRAINING - Epoch: [70][60/97]\tTime 0.150 (0.194)\tData 0.000 (0.019)\tLoss 0.1913 (0.2122)\tPrec@1 92.773 (92.591)\tPrec@5 99.805 (99.888)\t\nTRAINING - Epoch: [70][70/97]\tTime 0.155 (0.190)\tData 0.000 (0.017)\tLoss 0.2196 (0.2125)\tPrec@1 91.992 (92.570)\tPrec@5 99.609 (99.890)\t\nTRAINING - Epoch: [70][80/97]\tTime 0.147 (0.186)\tData 0.000 (0.015)\tLoss 0.2253 (0.2135)\tPrec@1 92.578 (92.513)\tPrec@5 99.805 (99.887)\t\nTRAINING - Epoch: [70][90/97]\tTime 0.149 (0.182)\tData 0.000 (0.013)\tLoss 0.2039 (0.2134)\tPrec@1 91.992 (92.449)\tPrec@5 100.000 (99.888)\t\nEVALUATING - Epoch: [70][0/20]\tTime 0.937 (0.937)\tData 0.890 (0.890)\tLoss 0.5709 (0.5709)\tPrec@1 83.008 (83.008)\tPrec@5 99.414 (99.414)\t\nEVALUATING - Epoch: [70][10/20]\tTime 0.047 (0.177)\tData 0.000 (0.122)\tLoss 0.5103 (0.5440)\tPrec@1 84.570 (84.322)\tPrec@5 99.414 (99.450)\t\n\nResults - Epoch: 71\nTraining Loss 0.2148 \tTraining Prec@1 92.403 \tTraining Prec@5 99.889 \tValidation Loss 0.5354 \tValidation Prec@1 84.390 \tValidation Prec@5 99.440 \t\n\n\nStarting Epoch: 72\n\nTRAINING - Epoch: [71][0/97]\tTime 1.212 (1.212)\tData 0.944 (0.944)\tLoss 0.1772 (0.1772)\tPrec@1 94.336 (94.336)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [71][10/97]\tTime 0.179 (0.333)\tData 0.007 (0.089)\tLoss 0.2263 (0.2034)\tPrec@1 93.359 (92.933)\tPrec@5 100.000 (99.947)\t\nTRAINING - Epoch: [71][20/97]\tTime 0.147 (0.250)\tData 0.000 (0.047)\tLoss 0.1756 (0.2035)\tPrec@1 93.359 (92.857)\tPrec@5 100.000 (99.944)\t\nTRAINING - Epoch: [71][30/97]\tTime 0.169 (0.221)\tData 0.000 (0.032)\tLoss 0.2138 (0.2050)\tPrec@1 91.992 (92.666)\tPrec@5 100.000 (99.905)\t\nTRAINING - Epoch: [71][40/97]\tTime 0.161 (0.206)\tData 0.000 (0.024)\tLoss 0.2299 (0.2111)\tPrec@1 90.430 (92.321)\tPrec@5 99.805 (99.871)\t\nTRAINING - Epoch: [71][50/97]\tTime 0.170 (0.197)\tData 0.000 (0.020)\tLoss 0.2210 (0.2144)\tPrec@1 93.359 (92.310)\tPrec@5 99.805 (99.858)\t\nTRAINING - Epoch: [71][60/97]\tTime 0.153 (0.191)\tData 0.000 (0.017)\tLoss 0.2094 (0.2153)\tPrec@1 92.773 (92.328)\tPrec@5 100.000 (99.872)\t\nTRAINING - Epoch: [71][70/97]\tTime 0.157 (0.186)\tData 0.000 (0.014)\tLoss 0.2290 (0.2158)\tPrec@1 92.969 (92.325)\tPrec@5 100.000 (99.871)\t\nTRAINING - Epoch: [71][80/97]\tTime 0.168 (0.183)\tData 0.000 (0.013)\tLoss 0.2596 (0.2164)\tPrec@1 91.211 (92.298)\tPrec@5 100.000 (99.867)\t\nTRAINING - Epoch: [71][90/97]\tTime 0.150 (0.180)\tData 0.000 (0.012)\tLoss 0.2510 (0.2179)\tPrec@1 91.992 (92.260)\tPrec@5 100.000 (99.873)\t\nEVALUATING - Epoch: [71][0/20]\tTime 1.109 (1.109)\tData 1.058 (1.058)\tLoss 0.4460 (0.4460)\tPrec@1 86.328 (86.328)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [71][10/20]\tTime 0.053 (0.195)\tData 0.000 (0.141)\tLoss 0.5609 (0.5388)\tPrec@1 84.766 (84.712)\tPrec@5 99.414 (99.201)\t\n\nResults - Epoch: 72\nTraining Loss 0.2180 \tTraining Prec@1 92.244 \tTraining Prec@5 99.877 \tValidation Loss 0.5223 \tValidation Prec@1 84.940 \tValidation Prec@5 99.320 \t\n\n\nStarting Epoch: 73\n\nTRAINING - Epoch: [72][0/97]\tTime 0.939 (0.939)\tData 0.677 (0.677)\tLoss 0.2268 (0.2268)\tPrec@1 91.602 (91.602)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [72][10/97]\tTime 0.193 (0.336)\tData 0.000 (0.094)\tLoss 0.1904 (0.2097)\tPrec@1 92.969 (92.649)\tPrec@5 100.000 (99.947)\t\nTRAINING - Epoch: [72][20/97]\tTime 0.160 (0.253)\tData 0.000 (0.050)\tLoss 0.1910 (0.2030)\tPrec@1 92.969 (92.727)\tPrec@5 100.000 (99.926)\t\nTRAINING - Epoch: [72][30/97]\tTime 0.158 (0.224)\tData 0.000 (0.034)\tLoss 0.2487 (0.2041)\tPrec@1 91.016 (92.654)\tPrec@5 99.805 (99.912)\t\nTRAINING - Epoch: [72][40/97]\tTime 0.156 (0.208)\tData 0.000 (0.026)\tLoss 0.1916 (0.2075)\tPrec@1 92.773 (92.530)\tPrec@5 100.000 (99.905)\t\nTRAINING - Epoch: [72][50/97]\tTime 0.160 (0.199)\tData 0.000 (0.021)\tLoss 0.2325 (0.2090)\tPrec@1 91.992 (92.440)\tPrec@5 100.000 (99.885)\t\nTRAINING - Epoch: [72][60/97]\tTime 0.161 (0.193)\tData 0.000 (0.018)\tLoss 0.2199 (0.2129)\tPrec@1 92.578 (92.332)\tPrec@5 99.609 (99.872)\t\nTRAINING - Epoch: [72][70/97]\tTime 0.150 (0.188)\tData 0.000 (0.015)\tLoss 0.2324 (0.2160)\tPrec@1 92.383 (92.229)\tPrec@5 99.609 (99.862)\t\nTRAINING - Epoch: [72][80/97]\tTime 0.163 (0.186)\tData 0.000 (0.014)\tLoss 0.1950 (0.2166)\tPrec@1 92.773 (92.224)\tPrec@5 100.000 (99.851)\t\nTRAINING - Epoch: [72][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.012)\tLoss 0.2743 (0.2199)\tPrec@1 90.625 (92.106)\tPrec@5 99.805 (99.850)\t\nEVALUATING - Epoch: [72][0/20]\tTime 1.319 (1.319)\tData 1.268 (1.268)\tLoss 0.5735 (0.5735)\tPrec@1 81.250 (81.250)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [72][10/20]\tTime 0.063 (0.198)\tData 0.019 (0.144)\tLoss 0.5485 (0.6254)\tPrec@1 84.961 (82.440)\tPrec@5 99.414 (99.201)\t\n\nResults - Epoch: 73\nTraining Loss 0.2199 \tTraining Prec@1 92.155 \tTraining Prec@5 99.849 \tValidation Loss 0.6082 \tValidation Prec@1 82.420 \tValidation Prec@5 99.270 \t\n\n\nStarting Epoch: 74\n\nTRAINING - Epoch: [73][0/97]\tTime 1.445 (1.445)\tData 1.140 (1.140)\tLoss 0.2082 (0.2082)\tPrec@1 93.359 (93.359)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [73][10/97]\tTime 0.178 (0.338)\tData 0.000 (0.107)\tLoss 0.2418 (0.1935)\tPrec@1 91.992 (93.235)\tPrec@5 99.609 (99.876)\t\nTRAINING - Epoch: [73][20/97]\tTime 0.168 (0.255)\tData 0.000 (0.056)\tLoss 0.1630 (0.1912)\tPrec@1 94.336 (93.341)\tPrec@5 99.805 (99.888)\t\nTRAINING - Epoch: [73][30/97]\tTime 0.165 (0.228)\tData 0.000 (0.038)\tLoss 0.2369 (0.1978)\tPrec@1 92.969 (93.076)\tPrec@5 99.805 (99.880)\t\nTRAINING - Epoch: [73][40/97]\tTime 0.179 (0.212)\tData 0.000 (0.029)\tLoss 0.2003 (0.1962)\tPrec@1 93.359 (93.059)\tPrec@5 100.000 (99.895)\t\nTRAINING - Epoch: [73][50/97]\tTime 0.154 (0.202)\tData 0.000 (0.024)\tLoss 0.2366 (0.1976)\tPrec@1 91.992 (93.049)\tPrec@5 100.000 (99.904)\t\nTRAINING - Epoch: [73][60/97]\tTime 0.180 (0.196)\tData 0.000 (0.020)\tLoss 0.1840 (0.2006)\tPrec@1 93.945 (92.930)\tPrec@5 99.805 (99.901)\t\nTRAINING - Epoch: [73][70/97]\tTime 0.174 (0.192)\tData 0.000 (0.018)\tLoss 0.2341 (0.2017)\tPrec@1 91.211 (92.922)\tPrec@5 100.000 (99.890)\t\nTRAINING - Epoch: [73][80/97]\tTime 0.161 (0.188)\tData 0.000 (0.016)\tLoss 0.2743 (0.2038)\tPrec@1 89.648 (92.814)\tPrec@5 99.609 (99.887)\t\nTRAINING - Epoch: [73][90/97]\tTime 0.150 (0.184)\tData 0.000 (0.014)\tLoss 0.2146 (0.2045)\tPrec@1 92.969 (92.812)\tPrec@5 100.000 (99.882)\t\nEVALUATING - Epoch: [73][0/20]\tTime 1.021 (1.021)\tData 0.975 (0.975)\tLoss 0.4840 (0.4840)\tPrec@1 83.594 (83.594)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [73][10/20]\tTime 0.064 (0.195)\tData 0.000 (0.143)\tLoss 0.4718 (0.5136)\tPrec@1 85.938 (85.227)\tPrec@5 99.609 (99.308)\t\n\nResults - Epoch: 74\nTraining Loss 0.2060 \tTraining Prec@1 92.753 \tTraining Prec@5 99.879 \tValidation Loss 0.5046 \tValidation Prec@1 85.520 \tValidation Prec@5 99.380 \t\n\n\nStarting Epoch: 75\n\nTRAINING - Epoch: [74][0/97]\tTime 1.394 (1.394)\tData 1.080 (1.080)\tLoss 0.1961 (0.1961)\tPrec@1 93.359 (93.359)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [74][10/97]\tTime 0.212 (0.338)\tData 0.005 (0.101)\tLoss 0.2025 (0.1862)\tPrec@1 92.188 (93.501)\tPrec@5 99.805 (99.893)\t\nTRAINING - Epoch: [74][20/97]\tTime 0.160 (0.254)\tData 0.000 (0.054)\tLoss 0.1988 (0.1914)\tPrec@1 93.555 (93.313)\tPrec@5 99.805 (99.907)\t\nTRAINING - Epoch: [74][30/97]\tTime 0.154 (0.225)\tData 0.000 (0.037)\tLoss 0.1589 (0.1924)\tPrec@1 93.164 (93.145)\tPrec@5 100.000 (99.912)\t\nTRAINING - Epoch: [74][40/97]\tTime 0.182 (0.210)\tData 0.000 (0.028)\tLoss 0.2049 (0.1995)\tPrec@1 93.945 (92.926)\tPrec@5 99.805 (99.905)\t\nTRAINING - Epoch: [74][50/97]\tTime 0.147 (0.201)\tData 0.000 (0.023)\tLoss 0.2472 (0.2014)\tPrec@1 90.625 (92.785)\tPrec@5 100.000 (99.904)\t\nTRAINING - Epoch: [74][60/97]\tTime 0.192 (0.194)\tData 0.000 (0.019)\tLoss 0.2152 (0.2047)\tPrec@1 93.945 (92.706)\tPrec@5 100.000 (99.914)\t\nTRAINING - Epoch: [74][70/97]\tTime 0.162 (0.190)\tData 0.000 (0.016)\tLoss 0.2422 (0.2074)\tPrec@1 91.992 (92.586)\tPrec@5 99.609 (99.912)\t\nTRAINING - Epoch: [74][80/97]\tTime 0.181 (0.186)\tData 0.005 (0.015)\tLoss 0.2314 (0.2090)\tPrec@1 91.406 (92.549)\tPrec@5 100.000 (99.913)\t\nTRAINING - Epoch: [74][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.013)\tLoss 0.2168 (0.2119)\tPrec@1 92.188 (92.484)\tPrec@5 100.000 (99.908)\t\nEVALUATING - Epoch: [74][0/20]\tTime 1.066 (1.066)\tData 1.011 (1.011)\tLoss 0.5636 (0.5636)\tPrec@1 83.594 (83.594)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [74][10/20]\tTime 0.046 (0.187)\tData 0.000 (0.137)\tLoss 0.5091 (0.5691)\tPrec@1 83.008 (83.842)\tPrec@5 99.609 (99.254)\t\n\nResults - Epoch: 75\nTraining Loss 0.2121 \tTraining Prec@1 92.508 \tTraining Prec@5 99.905 \tValidation Loss 0.5624 \tValidation Prec@1 83.880 \tValidation Prec@5 99.340 \t\n\n\nStarting Epoch: 76\n\nTRAINING - Epoch: [75][0/97]\tTime 1.565 (1.565)\tData 1.259 (1.259)\tLoss 0.2142 (0.2142)\tPrec@1 92.188 (92.188)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [75][10/97]\tTime 0.149 (0.345)\tData 0.000 (0.118)\tLoss 0.2087 (0.2112)\tPrec@1 92.188 (92.347)\tPrec@5 99.609 (99.947)\t\nTRAINING - Epoch: [75][20/97]\tTime 0.173 (0.258)\tData 0.000 (0.062)\tLoss 0.1893 (0.2004)\tPrec@1 93.945 (93.015)\tPrec@5 99.805 (99.944)\t\nTRAINING - Epoch: [75][30/97]\tTime 0.159 (0.227)\tData 0.000 (0.043)\tLoss 0.2155 (0.2052)\tPrec@1 92.969 (92.786)\tPrec@5 100.000 (99.931)\t\nTRAINING - Epoch: [75][40/97]\tTime 0.155 (0.211)\tData 0.005 (0.033)\tLoss 0.2290 (0.2029)\tPrec@1 92.578 (92.854)\tPrec@5 99.609 (99.919)\t\nTRAINING - Epoch: [75][50/97]\tTime 0.150 (0.201)\tData 0.000 (0.027)\tLoss 0.1619 (0.2031)\tPrec@1 94.141 (92.800)\tPrec@5 100.000 (99.897)\t\nTRAINING - Epoch: [75][60/97]\tTime 0.149 (0.194)\tData 0.000 (0.023)\tLoss 0.2538 (0.2028)\tPrec@1 91.406 (92.725)\tPrec@5 100.000 (99.901)\t\nTRAINING - Epoch: [75][70/97]\tTime 0.168 (0.190)\tData 0.000 (0.020)\tLoss 0.2242 (0.2025)\tPrec@1 92.383 (92.749)\tPrec@5 99.609 (99.895)\t\nTRAINING - Epoch: [75][80/97]\tTime 0.149 (0.187)\tData 0.000 (0.018)\tLoss 0.2253 (0.2033)\tPrec@1 91.211 (92.737)\tPrec@5 100.000 (99.896)\t\nTRAINING - Epoch: [75][90/97]\tTime 0.152 (0.183)\tData 0.000 (0.016)\tLoss 0.2032 (0.2052)\tPrec@1 92.969 (92.660)\tPrec@5 100.000 (99.901)\t\nEVALUATING - Epoch: [75][0/20]\tTime 1.045 (1.045)\tData 0.987 (0.987)\tLoss 0.5903 (0.5903)\tPrec@1 82.617 (82.617)\tPrec@5 99.219 (99.219)\t\nEVALUATING - Epoch: [75][10/20]\tTime 0.062 (0.194)\tData 0.000 (0.139)\tLoss 0.5416 (0.5501)\tPrec@1 81.836 (84.606)\tPrec@5 98.828 (99.272)\t\n\nResults - Epoch: 76\nTraining Loss 0.2068 \tTraining Prec@1 92.578 \tTraining Prec@5 99.893 \tValidation Loss 0.5485 \tValidation Prec@1 84.450 \tValidation Prec@5 99.420 \t\n\n\nStarting Epoch: 77\n\nTRAINING - Epoch: [76][0/97]\tTime 1.504 (1.504)\tData 1.213 (1.213)\tLoss 0.2129 (0.2129)\tPrec@1 92.578 (92.578)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [76][10/97]\tTime 0.148 (0.337)\tData 0.000 (0.113)\tLoss 0.1973 (0.1913)\tPrec@1 93.555 (93.288)\tPrec@5 99.805 (99.840)\t\nTRAINING - Epoch: [76][20/97]\tTime 0.182 (0.258)\tData 0.007 (0.060)\tLoss 0.1977 (0.2042)\tPrec@1 92.188 (92.801)\tPrec@5 100.000 (99.860)\t\nTRAINING - Epoch: [76][30/97]\tTime 0.158 (0.227)\tData 0.000 (0.041)\tLoss 0.2143 (0.2003)\tPrec@1 93.555 (93.051)\tPrec@5 99.609 (99.880)\t\nTRAINING - Epoch: [76][40/97]\tTime 0.165 (0.212)\tData 0.005 (0.032)\tLoss 0.2110 (0.2018)\tPrec@1 92.383 (93.031)\tPrec@5 100.000 (99.895)\t\nTRAINING - Epoch: [76][50/97]\tTime 0.150 (0.201)\tData 0.000 (0.026)\tLoss 0.1732 (0.2002)\tPrec@1 94.141 (93.064)\tPrec@5 99.805 (99.893)\t\nTRAINING - Epoch: [76][60/97]\tTime 0.146 (0.195)\tData 0.000 (0.022)\tLoss 0.1962 (0.2010)\tPrec@1 92.773 (92.982)\tPrec@5 100.000 (99.901)\t\nTRAINING - Epoch: [76][70/97]\tTime 0.180 (0.191)\tData 0.005 (0.019)\tLoss 0.2239 (0.2015)\tPrec@1 93.359 (92.927)\tPrec@5 99.805 (99.901)\t\nTRAINING - Epoch: [76][80/97]\tTime 0.151 (0.188)\tData 0.000 (0.017)\tLoss 0.2033 (0.2019)\tPrec@1 93.359 (92.901)\tPrec@5 100.000 (99.908)\t\nTRAINING - Epoch: [76][90/97]\tTime 0.150 (0.184)\tData 0.000 (0.015)\tLoss 0.1815 (0.2016)\tPrec@1 93.945 (92.870)\tPrec@5 99.805 (99.914)\t\nEVALUATING - Epoch: [76][0/20]\tTime 1.024 (1.024)\tData 0.974 (0.974)\tLoss 0.5748 (0.5748)\tPrec@1 83.398 (83.398)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [76][10/20]\tTime 0.068 (0.195)\tData 0.000 (0.137)\tLoss 0.5051 (0.5914)\tPrec@1 84.766 (83.612)\tPrec@5 99.219 (99.094)\t\n\nResults - Epoch: 77\nTraining Loss 0.2014 \tTraining Prec@1 92.870 \tTraining Prec@5 99.911 \tValidation Loss 0.5945 \tValidation Prec@1 83.160 \tValidation Prec@5 99.170 \t\n\n\nStarting Epoch: 78\n\nTRAINING - Epoch: [77][0/97]\tTime 1.171 (1.171)\tData 0.910 (0.910)\tLoss 0.1810 (0.1810)\tPrec@1 91.797 (91.797)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [77][10/97]\tTime 0.174 (0.335)\tData 0.000 (0.092)\tLoss 0.2566 (0.1950)\tPrec@1 90.625 (92.898)\tPrec@5 100.000 (99.893)\t\nTRAINING - Epoch: [77][20/97]\tTime 0.156 (0.252)\tData 0.000 (0.049)\tLoss 0.1719 (0.1990)\tPrec@1 93.750 (92.755)\tPrec@5 99.805 (99.916)\t\nTRAINING - Epoch: [77][30/97]\tTime 0.154 (0.223)\tData 0.000 (0.033)\tLoss 0.2107 (0.1962)\tPrec@1 92.383 (92.868)\tPrec@5 100.000 (99.931)\t\nTRAINING - Epoch: [77][40/97]\tTime 0.163 (0.209)\tData 0.000 (0.025)\tLoss 0.2143 (0.1977)\tPrec@1 92.773 (92.802)\tPrec@5 99.414 (99.900)\t\nTRAINING - Epoch: [77][50/97]\tTime 0.171 (0.200)\tData 0.000 (0.021)\tLoss 0.1600 (0.1991)\tPrec@1 94.531 (92.793)\tPrec@5 100.000 (99.897)\t\nTRAINING - Epoch: [77][60/97]\tTime 0.195 (0.200)\tData 0.000 (0.018)\tLoss 0.1883 (0.1998)\tPrec@1 92.773 (92.754)\tPrec@5 100.000 (99.901)\t\nTRAINING - Epoch: [77][70/97]\tTime 0.148 (0.195)\tData 0.000 (0.016)\tLoss 0.2356 (0.2035)\tPrec@1 91.016 (92.633)\tPrec@5 100.000 (99.909)\t\nTRAINING - Epoch: [77][80/97]\tTime 0.149 (0.191)\tData 0.000 (0.014)\tLoss 0.2353 (0.2051)\tPrec@1 91.406 (92.561)\tPrec@5 99.414 (99.894)\t\nTRAINING - Epoch: [77][90/97]\tTime 0.150 (0.187)\tData 0.000 (0.012)\tLoss 0.2198 (0.2072)\tPrec@1 91.602 (92.507)\tPrec@5 100.000 (99.893)\t\nEVALUATING - Epoch: [77][0/20]\tTime 1.084 (1.084)\tData 1.031 (1.031)\tLoss 0.4322 (0.4322)\tPrec@1 86.133 (86.133)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [77][10/20]\tTime 0.062 (0.195)\tData 0.000 (0.142)\tLoss 0.4214 (0.4870)\tPrec@1 87.305 (86.346)\tPrec@5 99.414 (99.290)\t\n\nResults - Epoch: 78\nTraining Loss 0.2089 \tTraining Prec@1 92.455 \tTraining Prec@5 99.891 \tValidation Loss 0.4878 \tValidation Prec@1 86.100 \tValidation Prec@5 99.340 \t\n\n\nStarting Epoch: 79\n\nTRAINING - Epoch: [78][0/97]\tTime 1.150 (1.150)\tData 0.878 (0.878)\tLoss 0.1617 (0.1617)\tPrec@1 93.945 (93.945)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [78][10/97]\tTime 0.183 (0.334)\tData 0.000 (0.091)\tLoss 0.2168 (0.1893)\tPrec@1 91.602 (93.395)\tPrec@5 99.805 (99.947)\t\nTRAINING - Epoch: [78][20/97]\tTime 0.180 (0.255)\tData 0.000 (0.049)\tLoss 0.1782 (0.1915)\tPrec@1 94.727 (93.313)\tPrec@5 100.000 (99.935)\t\nTRAINING - Epoch: [78][30/97]\tTime 0.181 (0.235)\tData 0.000 (0.034)\tLoss 0.1860 (0.1964)\tPrec@1 93.164 (93.126)\tPrec@5 100.000 (99.943)\t\nTRAINING - Epoch: [78][40/97]\tTime 0.175 (0.218)\tData 0.000 (0.026)\tLoss 0.2163 (0.1977)\tPrec@1 91.797 (93.097)\tPrec@5 99.805 (99.948)\t\nTRAINING - Epoch: [78][50/97]\tTime 0.153 (0.206)\tData 0.005 (0.021)\tLoss 0.2390 (0.1993)\tPrec@1 91.211 (93.015)\tPrec@5 99.805 (99.943)\t\nTRAINING - Epoch: [78][60/97]\tTime 0.151 (0.199)\tData 0.000 (0.018)\tLoss 0.2476 (0.2044)\tPrec@1 91.992 (92.821)\tPrec@5 99.609 (99.926)\t\nTRAINING - Epoch: [78][70/97]\tTime 0.210 (0.195)\tData 0.002 (0.015)\tLoss 0.1876 (0.2046)\tPrec@1 94.336 (92.831)\tPrec@5 99.805 (99.915)\t\nTRAINING - Epoch: [78][80/97]\tTime 0.160 (0.190)\tData 0.000 (0.013)\tLoss 0.2158 (0.2069)\tPrec@1 91.602 (92.740)\tPrec@5 99.805 (99.913)\t\nTRAINING - Epoch: [78][90/97]\tTime 0.150 (0.186)\tData 0.000 (0.012)\tLoss 0.2028 (0.2072)\tPrec@1 93.359 (92.658)\tPrec@5 99.805 (99.912)\t\nEVALUATING - Epoch: [78][0/20]\tTime 0.923 (0.923)\tData 0.870 (0.870)\tLoss 0.5548 (0.5548)\tPrec@1 83.203 (83.203)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [78][10/20]\tTime 0.068 (0.194)\tData 0.000 (0.139)\tLoss 0.6108 (0.6133)\tPrec@1 83.984 (83.043)\tPrec@5 99.219 (99.343)\t\n\nResults - Epoch: 79\nTraining Loss 0.2079 \tTraining Prec@1 92.610 \tTraining Prec@5 99.915 \tValidation Loss 0.6134 \tValidation Prec@1 82.870 \tValidation Prec@5 99.440 \t\n\n\nStarting Epoch: 80\n\nTRAINING - Epoch: [79][0/97]\tTime 1.449 (1.449)\tData 1.141 (1.141)\tLoss 0.1927 (0.1927)\tPrec@1 92.773 (92.773)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [79][10/97]\tTime 0.179 (0.343)\tData 0.000 (0.109)\tLoss 0.2082 (0.2006)\tPrec@1 92.773 (92.685)\tPrec@5 100.000 (99.929)\t\nTRAINING - Epoch: [79][20/97]\tTime 0.167 (0.257)\tData 0.000 (0.057)\tLoss 0.1756 (0.1911)\tPrec@1 93.750 (93.118)\tPrec@5 100.000 (99.916)\t\nTRAINING - Epoch: [79][30/97]\tTime 0.160 (0.226)\tData 0.000 (0.039)\tLoss 0.2199 (0.1960)\tPrec@1 91.992 (92.918)\tPrec@5 99.609 (99.905)\t\nTRAINING - Epoch: [79][40/97]\tTime 0.155 (0.210)\tData 0.000 (0.030)\tLoss 0.1621 (0.1952)\tPrec@1 94.727 (93.031)\tPrec@5 100.000 (99.905)\t\nTRAINING - Epoch: [79][50/97]\tTime 0.176 (0.201)\tData 0.006 (0.024)\tLoss 0.1797 (0.1980)\tPrec@1 93.555 (92.862)\tPrec@5 100.000 (99.904)\t\nTRAINING - Epoch: [79][60/97]\tTime 0.147 (0.195)\tData 0.000 (0.020)\tLoss 0.2200 (0.1975)\tPrec@1 91.602 (92.886)\tPrec@5 99.609 (99.894)\t\nTRAINING - Epoch: [79][70/97]\tTime 0.180 (0.191)\tData 0.000 (0.018)\tLoss 0.1381 (0.1968)\tPrec@1 95.117 (92.905)\tPrec@5 100.000 (99.887)\t\nTRAINING - Epoch: [79][80/97]\tTime 0.165 (0.187)\tData 0.000 (0.016)\tLoss 0.2042 (0.1980)\tPrec@1 91.992 (92.851)\tPrec@5 99.805 (99.889)\t\nTRAINING - Epoch: [79][90/97]\tTime 0.149 (0.183)\tData 0.000 (0.014)\tLoss 0.2293 (0.2007)\tPrec@1 91.797 (92.763)\tPrec@5 99.414 (99.888)\t\nEVALUATING - Epoch: [79][0/20]\tTime 0.726 (0.726)\tData 0.672 (0.672)\tLoss 0.4336 (0.4336)\tPrec@1 84.961 (84.961)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [79][10/20]\tTime 0.085 (0.207)\tData 0.019 (0.150)\tLoss 0.5140 (0.5222)\tPrec@1 85.156 (84.304)\tPrec@5 99.805 (99.379)\t\n\nResults - Epoch: 80\nTraining Loss 0.2023 \tTraining Prec@1 92.705 \tTraining Prec@5 99.889 \tValidation Loss 0.5262 \tValidation Prec@1 84.330 \tValidation Prec@5 99.340 \t\n\n\nStarting Epoch: 81\n\nTRAINING - Epoch: [80][0/97]\tTime 1.485 (1.485)\tData 1.218 (1.218)\tLoss 0.2069 (0.2069)\tPrec@1 92.578 (92.578)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [80][10/97]\tTime 0.217 (0.337)\tData 0.000 (0.113)\tLoss 0.2096 (0.2058)\tPrec@1 92.773 (92.472)\tPrec@5 99.805 (99.964)\t\nTRAINING - Epoch: [80][20/97]\tTime 0.160 (0.254)\tData 0.000 (0.060)\tLoss 0.1989 (0.2002)\tPrec@1 92.578 (92.885)\tPrec@5 99.805 (99.963)\t\nTRAINING - Epoch: [80][30/97]\tTime 0.176 (0.225)\tData 0.000 (0.041)\tLoss 0.1971 (0.2025)\tPrec@1 91.992 (92.698)\tPrec@5 100.000 (99.943)\t\nTRAINING - Epoch: [80][40/97]\tTime 0.155 (0.210)\tData 0.000 (0.031)\tLoss 0.1697 (0.1982)\tPrec@1 93.945 (92.816)\tPrec@5 100.000 (99.938)\t\nTRAINING - Epoch: [80][50/97]\tTime 0.168 (0.201)\tData 0.000 (0.026)\tLoss 0.2357 (0.1998)\tPrec@1 92.773 (92.743)\tPrec@5 99.805 (99.912)\t\nTRAINING - Epoch: [80][60/97]\tTime 0.154 (0.194)\tData 0.000 (0.022)\tLoss 0.2358 (0.2020)\tPrec@1 91.406 (92.709)\tPrec@5 99.805 (99.901)\t\nTRAINING - Epoch: [80][70/97]\tTime 0.177 (0.190)\tData 0.000 (0.019)\tLoss 0.2309 (0.2021)\tPrec@1 92.578 (92.718)\tPrec@5 99.805 (99.906)\t\nTRAINING - Epoch: [80][80/97]\tTime 0.168 (0.191)\tData 0.000 (0.017)\tLoss 0.1756 (0.2021)\tPrec@1 93.945 (92.766)\tPrec@5 99.805 (99.908)\t\nTRAINING - Epoch: [80][90/97]\tTime 0.150 (0.186)\tData 0.000 (0.015)\tLoss 0.1915 (0.2029)\tPrec@1 93.750 (92.748)\tPrec@5 100.000 (99.897)\t\nEVALUATING - Epoch: [80][0/20]\tTime 0.989 (0.989)\tData 0.940 (0.940)\tLoss 0.5085 (0.5085)\tPrec@1 82.617 (82.617)\tPrec@5 99.609 (99.609)\t\nEVALUATING - Epoch: [80][10/20]\tTime 0.072 (0.188)\tData 0.000 (0.132)\tLoss 0.5199 (0.5522)\tPrec@1 84.570 (83.949)\tPrec@5 99.609 (99.343)\t\n\nResults - Epoch: 81\nTraining Loss 0.2038 \tTraining Prec@1 92.693 \tTraining Prec@5 99.893 \tValidation Loss 0.5474 \tValidation Prec@1 84.020 \tValidation Prec@5 99.400 \t\n\n\nStarting Epoch: 82\n\nTRAINING - Epoch: [81][0/97]\tTime 1.580 (1.580)\tData 1.311 (1.311)\tLoss 0.1676 (0.1676)\tPrec@1 93.945 (93.945)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [81][10/97]\tTime 0.161 (0.333)\tData 0.005 (0.122)\tLoss 0.1812 (0.1762)\tPrec@1 93.359 (93.839)\tPrec@5 100.000 (99.947)\t\nTRAINING - Epoch: [81][20/97]\tTime 0.176 (0.253)\tData 0.014 (0.065)\tLoss 0.1443 (0.1695)\tPrec@1 95.117 (93.964)\tPrec@5 99.805 (99.944)\t\nTRAINING - Epoch: [81][30/97]\tTime 0.166 (0.225)\tData 0.000 (0.045)\tLoss 0.1552 (0.1655)\tPrec@1 93.750 (94.109)\tPrec@5 100.000 (99.956)\t\nTRAINING - Epoch: [81][40/97]\tTime 0.153 (0.208)\tData 0.000 (0.034)\tLoss 0.1671 (0.1618)\tPrec@1 94.727 (94.293)\tPrec@5 100.000 (99.943)\t\nTRAINING - Epoch: [81][50/97]\tTime 0.209 (0.204)\tData 0.016 (0.028)\tLoss 0.1562 (0.1594)\tPrec@1 93.945 (94.413)\tPrec@5 100.000 (99.939)\t\nTRAINING - Epoch: [81][60/97]\tTime 0.165 (0.200)\tData 0.000 (0.024)\tLoss 0.1333 (0.1544)\tPrec@1 96.094 (94.682)\tPrec@5 100.000 (99.933)\t\nTRAINING - Epoch: [81][70/97]\tTime 0.155 (0.195)\tData 0.000 (0.021)\tLoss 0.1224 (0.1518)\tPrec@1 96.875 (94.773)\tPrec@5 99.805 (99.939)\t\nTRAINING - Epoch: [81][80/97]\tTime 0.171 (0.190)\tData 0.000 (0.018)\tLoss 0.1606 (0.1506)\tPrec@1 94.141 (94.825)\tPrec@5 100.000 (99.937)\t\nTRAINING - Epoch: [81][90/97]\tTime 0.150 (0.186)\tData 0.000 (0.016)\tLoss 0.1121 (0.1486)\tPrec@1 96.484 (94.892)\tPrec@5 100.000 (99.940)\t\nEVALUATING - Epoch: [81][0/20]\tTime 0.974 (0.974)\tData 0.923 (0.923)\tLoss 0.2776 (0.2776)\tPrec@1 90.430 (90.430)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [81][10/20]\tTime 0.055 (0.188)\tData 0.000 (0.133)\tLoss 0.3384 (0.3700)\tPrec@1 90.625 (89.009)\tPrec@5 99.414 (99.592)\t\n\nResults - Epoch: 82\nTraining Loss 0.1481 \tTraining Prec@1 94.934 \tTraining Prec@5 99.938 \tValidation Loss 0.3677 \tValidation Prec@1 88.980 \tValidation Prec@5 99.690 \t\n\n\nStarting Epoch: 83\n\nTRAINING - Epoch: [82][0/97]\tTime 1.392 (1.392)\tData 1.097 (1.097)\tLoss 0.1223 (0.1223)\tPrec@1 95.508 (95.508)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [82][10/97]\tTime 0.159 (0.334)\tData 0.000 (0.103)\tLoss 0.1277 (0.1329)\tPrec@1 95.508 (95.455)\tPrec@5 99.805 (99.982)\t\nTRAINING - Epoch: [82][20/97]\tTime 0.192 (0.272)\tData 0.000 (0.055)\tLoss 0.1428 (0.1314)\tPrec@1 95.898 (95.564)\tPrec@5 100.000 (99.944)\t\nTRAINING - Epoch: [82][30/97]\tTime 0.153 (0.238)\tData 0.000 (0.038)\tLoss 0.1382 (0.1317)\tPrec@1 95.117 (95.520)\tPrec@5 100.000 (99.962)\t\nTRAINING - Epoch: [82][40/97]\tTime 0.180 (0.220)\tData 0.000 (0.029)\tLoss 0.1211 (0.1310)\tPrec@1 96.289 (95.455)\tPrec@5 100.000 (99.962)\t\nTRAINING - Epoch: [82][50/97]\tTime 0.162 (0.209)\tData 0.000 (0.024)\tLoss 0.1103 (0.1289)\tPrec@1 96.680 (95.554)\tPrec@5 100.000 (99.962)\t\nTRAINING - Epoch: [82][60/97]\tTime 0.150 (0.200)\tData 0.000 (0.020)\tLoss 0.1544 (0.1316)\tPrec@1 94.922 (95.489)\tPrec@5 100.000 (99.962)\t\nTRAINING - Epoch: [82][70/97]\tTime 0.149 (0.195)\tData 0.000 (0.017)\tLoss 0.1664 (0.1299)\tPrec@1 95.117 (95.568)\tPrec@5 100.000 (99.961)\t\nTRAINING - Epoch: [82][80/97]\tTime 0.159 (0.191)\tData 0.000 (0.015)\tLoss 0.1332 (0.1306)\tPrec@1 95.703 (95.534)\tPrec@5 100.000 (99.957)\t\nTRAINING - Epoch: [82][90/97]\tTime 0.150 (0.187)\tData 0.000 (0.014)\tLoss 0.1698 (0.1316)\tPrec@1 93.945 (95.473)\tPrec@5 99.805 (99.957)\t\nEVALUATING - Epoch: [82][0/20]\tTime 1.106 (1.106)\tData 1.057 (1.057)\tLoss 0.2857 (0.2857)\tPrec@1 89.453 (89.453)\tPrec@5 100.000 (100.000)\t\nEVALUATING - Epoch: [82][10/20]\tTime 0.069 (0.195)\tData 0.024 (0.137)\tLoss 0.3349 (0.3705)\tPrec@1 89.453 (88.707)\tPrec@5 99.414 (99.609)\t\n\nResults - Epoch: 83\nTraining Loss 0.1312 \tTraining Prec@1 95.486 \tTraining Prec@5 99.958 \tValidation Loss 0.3667 \tValidation Prec@1 88.740 \tValidation Prec@5 99.660 \t\n\n\nStarting Epoch: 84\n\nTRAINING - Epoch: [83][0/97]\tTime 1.585 (1.585)\tData 1.384 (1.384)\tLoss 0.1368 (0.1368)\tPrec@1 94.531 (94.531)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [83][10/97]\tTime 0.184 (0.382)\tData 0.000 (0.151)\tLoss 0.1293 (0.1281)\tPrec@1 96.484 (95.455)\tPrec@5 99.805 (99.982)\t\nTRAINING - Epoch: [83][20/97]\tTime 0.149 (0.276)\tData 0.000 (0.079)\tLoss 0.1075 (0.1233)\tPrec@1 96.484 (95.750)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [83][30/97]\tTime 0.158 (0.240)\tData 0.000 (0.054)\tLoss 0.1348 (0.1231)\tPrec@1 95.117 (95.728)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [83][40/97]\tTime 0.157 (0.221)\tData 0.000 (0.041)\tLoss 0.1114 (0.1230)\tPrec@1 95.508 (95.698)\tPrec@5 100.000 (99.976)\t\nTRAINING - Epoch: [83][50/97]\tTime 0.166 (0.209)\tData 0.000 (0.033)\tLoss 0.0976 (0.1243)\tPrec@1 97.070 (95.669)\tPrec@5 100.000 (99.977)\t\nTRAINING - Epoch: [83][60/97]\tTime 0.165 (0.201)\tData 0.000 (0.028)\tLoss 0.1069 (0.1239)\tPrec@1 96.094 (95.719)\tPrec@5 100.000 (99.978)\t\nTRAINING - Epoch: [83][70/97]\tTime 0.163 (0.195)\tData 0.000 (0.024)\tLoss 0.1210 (0.1236)\tPrec@1 95.703 (95.772)\tPrec@5 100.000 (99.978)\t\nTRAINING - Epoch: [83][80/97]\tTime 0.152 (0.191)\tData 0.000 (0.021)\tLoss 0.1202 (0.1253)\tPrec@1 95.703 (95.730)\tPrec@5 100.000 (99.976)\t\nTRAINING - Epoch: [83][90/97]\tTime 0.149 (0.186)\tData 0.000 (0.019)\tLoss 0.1574 (0.1255)\tPrec@1 93.555 (95.714)\tPrec@5 100.000 (99.974)\t\nEVALUATING - Epoch: [83][0/20]\tTime 1.538 (1.538)\tData 1.482 (1.482)\tLoss 0.2731 (0.2731)\tPrec@1 90.625 (90.625)\tPrec@5 100.000 (100.000)\t\nEVALUATING - Epoch: [83][10/20]\tTime 0.047 (0.243)\tData 0.000 (0.186)\tLoss 0.3280 (0.3682)\tPrec@1 89.453 (89.080)\tPrec@5 99.609 (99.645)\t\n\nResults - Epoch: 84\nTraining Loss 0.1251 \tTraining Prec@1 95.743 \tTraining Prec@5 99.976 \tValidation Loss 0.3637 \tValidation Prec@1 89.200 \tValidation Prec@5 99.680 \t\n\n\nStarting Epoch: 85\n\nTRAINING - Epoch: [84][0/97]\tTime 1.370 (1.370)\tData 1.070 (1.070)\tLoss 0.1310 (0.1310)\tPrec@1 95.898 (95.898)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [84][10/97]\tTime 0.148 (0.334)\tData 0.000 (0.099)\tLoss 0.1213 (0.1132)\tPrec@1 95.898 (96.129)\tPrec@5 100.000 (99.964)\t\nTRAINING - Epoch: [84][20/97]\tTime 0.165 (0.250)\tData 0.000 (0.052)\tLoss 0.1167 (0.1138)\tPrec@1 95.508 (96.280)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [84][30/97]\tTime 0.147 (0.223)\tData 0.000 (0.036)\tLoss 0.1255 (0.1170)\tPrec@1 95.898 (96.119)\tPrec@5 100.000 (99.975)\t\nTRAINING - Epoch: [84][40/97]\tTime 0.178 (0.209)\tData 0.000 (0.028)\tLoss 0.0973 (0.1165)\tPrec@1 97.461 (96.099)\tPrec@5 99.805 (99.957)\t\nTRAINING - Epoch: [84][50/97]\tTime 0.150 (0.199)\tData 0.000 (0.022)\tLoss 0.1539 (0.1183)\tPrec@1 95.117 (96.032)\tPrec@5 99.805 (99.950)\t\nTRAINING - Epoch: [84][60/97]\tTime 0.150 (0.193)\tData 0.000 (0.019)\tLoss 0.1067 (0.1178)\tPrec@1 95.898 (96.017)\tPrec@5 100.000 (99.949)\t\nTRAINING - Epoch: [84][70/97]\tTime 0.152 (0.189)\tData 0.000 (0.017)\tLoss 0.1123 (0.1173)\tPrec@1 96.094 (96.019)\tPrec@5 100.000 (99.953)\t\nTRAINING - Epoch: [84][80/97]\tTime 0.167 (0.185)\tData 0.013 (0.015)\tLoss 0.0980 (0.1175)\tPrec@1 96.680 (96.024)\tPrec@5 100.000 (99.954)\t\nTRAINING - Epoch: [84][90/97]\tTime 0.149 (0.181)\tData 0.000 (0.013)\tLoss 0.0799 (0.1176)\tPrec@1 98.242 (96.042)\tPrec@5 100.000 (99.955)\t\nEVALUATING - Epoch: [84][0/20]\tTime 1.032 (1.032)\tData 0.947 (0.947)\tLoss 0.2774 (0.2774)\tPrec@1 91.016 (91.016)\tPrec@5 100.000 (100.000)\t\nEVALUATING - Epoch: [84][10/20]\tTime 0.052 (0.195)\tData 0.000 (0.137)\tLoss 0.3223 (0.3694)\tPrec@1 90.820 (89.080)\tPrec@5 99.219 (99.538)\t\n\nResults - Epoch: 85\nTraining Loss 0.1178 \tTraining Prec@1 96.007 \tTraining Prec@5 99.958 \tValidation Loss 0.3629 \tValidation Prec@1 89.080 \tValidation Prec@5 99.630 \t\n\n\nStarting Epoch: 86\n\nTRAINING - Epoch: [85][0/97]\tTime 1.495 (1.495)\tData 1.199 (1.199)\tLoss 0.0822 (0.0822)\tPrec@1 97.656 (97.656)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [85][10/97]\tTime 0.178 (0.334)\tData 0.000 (0.113)\tLoss 0.0989 (0.1085)\tPrec@1 97.070 (96.378)\tPrec@5 100.000 (99.964)\t\nTRAINING - Epoch: [85][20/97]\tTime 0.150 (0.254)\tData 0.000 (0.060)\tLoss 0.1203 (0.1128)\tPrec@1 95.312 (96.150)\tPrec@5 100.000 (99.972)\t\nTRAINING - Epoch: [85][30/97]\tTime 0.172 (0.226)\tData 0.000 (0.041)\tLoss 0.1189 (0.1161)\tPrec@1 96.484 (96.113)\tPrec@5 99.805 (99.943)\t\nTRAINING - Epoch: [85][40/97]\tTime 0.166 (0.209)\tData 0.000 (0.031)\tLoss 0.1214 (0.1175)\tPrec@1 94.531 (95.965)\tPrec@5 100.000 (99.948)\t\nTRAINING - Epoch: [85][50/97]\tTime 0.165 (0.200)\tData 0.000 (0.025)\tLoss 0.1506 (0.1160)\tPrec@1 94.727 (95.979)\tPrec@5 100.000 (99.954)\t\nTRAINING - Epoch: [85][60/97]\tTime 0.162 (0.193)\tData 0.000 (0.021)\tLoss 0.0906 (0.1159)\tPrec@1 97.070 (95.985)\tPrec@5 100.000 (99.958)\t\nTRAINING - Epoch: [85][70/97]\tTime 0.150 (0.188)\tData 0.000 (0.018)\tLoss 0.1307 (0.1162)\tPrec@1 93.945 (95.953)\tPrec@5 100.000 (99.961)\t\nTRAINING - Epoch: [85][80/97]\tTime 0.155 (0.184)\tData 0.001 (0.016)\tLoss 0.1247 (0.1167)\tPrec@1 96.289 (95.942)\tPrec@5 100.000 (99.964)\t\nTRAINING - Epoch: [85][90/97]\tTime 0.148 (0.180)\tData 0.000 (0.014)\tLoss 0.1231 (0.1159)\tPrec@1 95.898 (95.984)\tPrec@5 100.000 (99.959)\t\nEVALUATING - Epoch: [85][0/20]\tTime 1.324 (1.324)\tData 1.273 (1.273)\tLoss 0.2671 (0.2671)\tPrec@1 90.625 (90.625)\tPrec@5 100.000 (100.000)\t\nEVALUATING - Epoch: [85][10/20]\tTime 0.050 (0.199)\tData 0.000 (0.146)\tLoss 0.3177 (0.3656)\tPrec@1 90.430 (89.045)\tPrec@5 99.414 (99.556)\t\n\nResults - Epoch: 86\nTraining Loss 0.1158 \tTraining Prec@1 95.997 \tTraining Prec@5 99.958 \tValidation Loss 0.3596 \tValidation Prec@1 89.150 \tValidation Prec@5 99.580 \t\n\n\nStarting Epoch: 87\n\nTRAINING - Epoch: [86][0/97]\tTime 1.384 (1.384)\tData 1.100 (1.100)\tLoss 0.1019 (0.1019)\tPrec@1 97.461 (97.461)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [86][10/97]\tTime 0.162 (0.329)\tData 0.005 (0.103)\tLoss 0.1109 (0.1037)\tPrec@1 96.875 (96.715)\tPrec@5 99.805 (99.964)\t\nTRAINING - Epoch: [86][20/97]\tTime 0.157 (0.247)\tData 0.000 (0.055)\tLoss 0.0935 (0.1063)\tPrec@1 96.094 (96.512)\tPrec@5 100.000 (99.972)\t\nTRAINING - Epoch: [86][30/97]\tTime 0.149 (0.218)\tData 0.000 (0.038)\tLoss 0.1154 (0.1067)\tPrec@1 95.898 (96.447)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [86][40/97]\tTime 0.189 (0.205)\tData 0.000 (0.029)\tLoss 0.0989 (0.1076)\tPrec@1 97.070 (96.446)\tPrec@5 100.000 (99.976)\t\nTRAINING - Epoch: [86][50/97]\tTime 0.153 (0.196)\tData 0.000 (0.024)\tLoss 0.0973 (0.1085)\tPrec@1 97.461 (96.442)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [86][60/97]\tTime 0.154 (0.190)\tData 0.000 (0.020)\tLoss 0.1308 (0.1099)\tPrec@1 94.922 (96.382)\tPrec@5 100.000 (99.974)\t\nTRAINING - Epoch: [86][70/97]\tTime 0.156 (0.186)\tData 0.000 (0.017)\tLoss 0.1248 (0.1116)\tPrec@1 95.312 (96.300)\tPrec@5 100.000 (99.978)\t\nTRAINING - Epoch: [86][80/97]\tTime 0.169 (0.183)\tData 0.000 (0.015)\tLoss 0.1304 (0.1120)\tPrec@1 95.117 (96.296)\tPrec@5 100.000 (99.976)\t\nTRAINING - Epoch: [86][90/97]\tTime 0.150 (0.179)\tData 0.000 (0.014)\tLoss 0.1059 (0.1129)\tPrec@1 96.484 (96.263)\tPrec@5 100.000 (99.972)\t\nEVALUATING - Epoch: [86][0/20]\tTime 1.082 (1.082)\tData 1.028 (1.028)\tLoss 0.2804 (0.2804)\tPrec@1 90.820 (90.820)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [86][10/20]\tTime 0.048 (0.187)\tData 0.000 (0.132)\tLoss 0.3205 (0.3751)\tPrec@1 90.234 (88.920)\tPrec@5 99.414 (99.574)\t\n\nResults - Epoch: 87\nTraining Loss 0.1130 \tTraining Prec@1 96.277 \tTraining Prec@5 99.970 \tValidation Loss 0.3691 \tValidation Prec@1 88.800 \tValidation Prec@5 99.630 \t\n\n\nStarting Epoch: 88\n\nTRAINING - Epoch: [87][0/97]\tTime 1.160 (1.160)\tData 0.894 (0.894)\tLoss 0.1060 (0.1060)\tPrec@1 95.898 (95.898)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [87][10/97]\tTime 0.146 (0.332)\tData 0.000 (0.093)\tLoss 0.0998 (0.1050)\tPrec@1 97.070 (96.254)\tPrec@5 100.000 (99.982)\t\nTRAINING - Epoch: [87][20/97]\tTime 0.151 (0.251)\tData 0.000 (0.049)\tLoss 0.0913 (0.1098)\tPrec@1 97.461 (96.289)\tPrec@5 100.000 (99.963)\t\nTRAINING - Epoch: [87][30/97]\tTime 0.148 (0.222)\tData 0.000 (0.034)\tLoss 0.0969 (0.1086)\tPrec@1 97.070 (96.339)\tPrec@5 100.000 (99.962)\t\nTRAINING - Epoch: [87][40/97]\tTime 0.155 (0.208)\tData 0.000 (0.026)\tLoss 0.1113 (0.1082)\tPrec@1 96.680 (96.442)\tPrec@5 100.000 (99.967)\t\nTRAINING - Epoch: [87][50/97]\tTime 0.165 (0.198)\tData 0.000 (0.021)\tLoss 0.1356 (0.1085)\tPrec@1 95.312 (96.431)\tPrec@5 100.000 (99.966)\t\nTRAINING - Epoch: [87][60/97]\tTime 0.169 (0.193)\tData 0.000 (0.018)\tLoss 0.1411 (0.1092)\tPrec@1 94.531 (96.372)\tPrec@5 100.000 (99.971)\t\nTRAINING - Epoch: [87][70/97]\tTime 0.154 (0.188)\tData 0.005 (0.015)\tLoss 0.1177 (0.1090)\tPrec@1 95.898 (96.369)\tPrec@5 100.000 (99.975)\t\nTRAINING - Epoch: [87][80/97]\tTime 0.150 (0.185)\tData 0.000 (0.014)\tLoss 0.1092 (0.1104)\tPrec@1 96.680 (96.311)\tPrec@5 100.000 (99.976)\t\nTRAINING - Epoch: [87][90/97]\tTime 0.150 (0.181)\tData 0.000 (0.012)\tLoss 0.1305 (0.1105)\tPrec@1 95.508 (96.304)\tPrec@5 100.000 (99.976)\t\nEVALUATING - Epoch: [87][0/20]\tTime 1.105 (1.105)\tData 1.054 (1.054)\tLoss 0.2828 (0.2828)\tPrec@1 91.016 (91.016)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [87][10/20]\tTime 0.052 (0.190)\tData 0.000 (0.134)\tLoss 0.3268 (0.3764)\tPrec@1 90.234 (89.080)\tPrec@5 99.609 (99.645)\t\n\nResults - Epoch: 88\nTraining Loss 0.1096 \tTraining Prec@1 96.333 \tTraining Prec@5 99.978 \tValidation Loss 0.3682 \tValidation Prec@1 89.100 \tValidation Prec@5 99.680 \t\n\n\nStarting Epoch: 89\n\nTRAINING - Epoch: [88][0/97]\tTime 1.414 (1.414)\tData 1.123 (1.123)\tLoss 0.1027 (0.1027)\tPrec@1 96.875 (96.875)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [88][10/97]\tTime 0.159 (0.332)\tData 0.000 (0.104)\tLoss 0.0952 (0.1017)\tPrec@1 97.266 (96.680)\tPrec@5 100.000 (99.982)\t\nTRAINING - Epoch: [88][20/97]\tTime 0.154 (0.252)\tData 0.005 (0.055)\tLoss 0.1394 (0.1087)\tPrec@1 95.508 (96.522)\tPrec@5 100.000 (99.963)\t\nTRAINING - Epoch: [88][30/97]\tTime 0.178 (0.223)\tData 0.000 (0.038)\tLoss 0.1205 (0.1090)\tPrec@1 95.898 (96.421)\tPrec@5 100.000 (99.975)\t\nTRAINING - Epoch: [88][40/97]\tTime 0.164 (0.208)\tData 0.000 (0.029)\tLoss 0.0834 (0.1059)\tPrec@1 97.461 (96.542)\tPrec@5 100.000 (99.971)\t\nTRAINING - Epoch: [88][50/97]\tTime 0.168 (0.199)\tData 0.000 (0.023)\tLoss 0.1065 (0.1052)\tPrec@1 96.875 (96.569)\tPrec@5 100.000 (99.977)\t\nTRAINING - Epoch: [88][60/97]\tTime 0.154 (0.193)\tData 0.000 (0.020)\tLoss 0.1223 (0.1069)\tPrec@1 95.508 (96.507)\tPrec@5 100.000 (99.974)\t\nTRAINING - Epoch: [88][70/97]\tTime 0.150 (0.188)\tData 0.000 (0.017)\tLoss 0.1209 (0.1061)\tPrec@1 95.312 (96.517)\tPrec@5 100.000 (99.972)\t\nTRAINING - Epoch: [88][80/97]\tTime 0.167 (0.185)\tData 0.000 (0.015)\tLoss 0.1191 (0.1072)\tPrec@1 96.289 (96.484)\tPrec@5 100.000 (99.969)\t\nTRAINING - Epoch: [88][90/97]\tTime 0.150 (0.181)\tData 0.000 (0.013)\tLoss 0.1287 (0.1076)\tPrec@1 95.117 (96.437)\tPrec@5 100.000 (99.970)\t\nEVALUATING - Epoch: [88][0/20]\tTime 1.067 (1.067)\tData 1.009 (1.009)\tLoss 0.2919 (0.2919)\tPrec@1 90.039 (90.039)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [88][10/20]\tTime 0.045 (0.194)\tData 0.000 (0.141)\tLoss 0.3305 (0.3784)\tPrec@1 90.039 (89.240)\tPrec@5 99.219 (99.663)\t\n\nResults - Epoch: 89\nTraining Loss 0.1078 \tTraining Prec@1 96.450 \tTraining Prec@5 99.968 \tValidation Loss 0.3698 \tValidation Prec@1 89.270 \tValidation Prec@5 99.660 \t\n\n\nStarting Epoch: 90\n\nTRAINING - Epoch: [89][0/97]\tTime 1.378 (1.378)\tData 1.102 (1.102)\tLoss 0.0897 (0.0897)\tPrec@1 96.484 (96.484)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [89][10/97]\tTime 0.175 (0.336)\tData 0.000 (0.104)\tLoss 0.1046 (0.0975)\tPrec@1 96.680 (96.946)\tPrec@5 100.000 (99.964)\t\nTRAINING - Epoch: [89][20/97]\tTime 0.161 (0.253)\tData 0.000 (0.055)\tLoss 0.0815 (0.1002)\tPrec@1 97.852 (96.689)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [89][30/97]\tTime 0.165 (0.222)\tData 0.000 (0.037)\tLoss 0.1136 (0.1019)\tPrec@1 96.289 (96.617)\tPrec@5 99.805 (99.968)\t\nTRAINING - Epoch: [89][40/97]\tTime 0.156 (0.207)\tData 0.000 (0.029)\tLoss 0.1151 (0.1063)\tPrec@1 96.094 (96.446)\tPrec@5 100.000 (99.967)\t\nTRAINING - Epoch: [89][50/97]\tTime 0.160 (0.198)\tData 0.000 (0.023)\tLoss 0.1090 (0.1069)\tPrec@1 96.680 (96.438)\tPrec@5 100.000 (99.973)\t\nTRAINING - Epoch: [89][60/97]\tTime 0.167 (0.192)\tData 0.009 (0.020)\tLoss 0.1308 (0.1075)\tPrec@1 94.531 (96.376)\tPrec@5 100.000 (99.974)\t\nTRAINING - Epoch: [89][70/97]\tTime 0.148 (0.187)\tData 0.000 (0.017)\tLoss 0.1100 (0.1071)\tPrec@1 97.070 (96.391)\tPrec@5 99.805 (99.975)\t\nTRAINING - Epoch: [89][80/97]\tTime 0.163 (0.185)\tData 0.005 (0.015)\tLoss 0.1061 (0.1068)\tPrec@1 96.289 (96.424)\tPrec@5 100.000 (99.978)\t\nTRAINING - Epoch: [89][90/97]\tTime 0.149 (0.181)\tData 0.000 (0.013)\tLoss 0.1190 (0.1066)\tPrec@1 96.289 (96.414)\tPrec@5 100.000 (99.976)\t\nEVALUATING - Epoch: [89][0/20]\tTime 1.374 (1.374)\tData 1.315 (1.315)\tLoss 0.2832 (0.2832)\tPrec@1 90.625 (90.625)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [89][10/20]\tTime 0.044 (0.201)\tData 0.000 (0.150)\tLoss 0.3262 (0.3807)\tPrec@1 90.430 (89.364)\tPrec@5 99.414 (99.609)\t\n\nResults - Epoch: 90\nTraining Loss 0.1069 \tTraining Prec@1 96.390 \tTraining Prec@5 99.976 \tValidation Loss 0.3717 \tValidation Prec@1 89.240 \tValidation Prec@5 99.630 \t\n\n\nStarting Epoch: 91\n\nTRAINING - Epoch: [90][0/97]\tTime 1.385 (1.385)\tData 1.134 (1.134)\tLoss 0.1087 (0.1087)\tPrec@1 96.875 (96.875)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [90][10/97]\tTime 0.185 (0.330)\tData 0.000 (0.107)\tLoss 0.1046 (0.1003)\tPrec@1 96.484 (96.573)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [90][20/97]\tTime 0.169 (0.253)\tData 0.000 (0.057)\tLoss 0.1327 (0.1001)\tPrec@1 95.508 (96.689)\tPrec@5 100.000 (99.991)\t\nTRAINING - Epoch: [90][30/97]\tTime 0.153 (0.223)\tData 0.000 (0.039)\tLoss 0.0931 (0.0999)\tPrec@1 96.289 (96.648)\tPrec@5 100.000 (99.994)\t\nTRAINING - Epoch: [90][40/97]\tTime 0.150 (0.207)\tData 0.000 (0.030)\tLoss 0.1525 (0.1001)\tPrec@1 94.141 (96.646)\tPrec@5 99.609 (99.962)\t\nTRAINING - Epoch: [90][50/97]\tTime 0.156 (0.198)\tData 0.000 (0.024)\tLoss 0.1136 (0.1017)\tPrec@1 95.898 (96.592)\tPrec@5 100.000 (99.969)\t\nTRAINING - Epoch: [90][60/97]\tTime 0.161 (0.191)\tData 0.000 (0.020)\tLoss 0.0860 (0.1019)\tPrec@1 96.680 (96.577)\tPrec@5 100.000 (99.971)\t\nTRAINING - Epoch: [90][70/97]\tTime 0.148 (0.187)\tData 0.000 (0.018)\tLoss 0.1041 (0.1031)\tPrec@1 96.094 (96.515)\tPrec@5 99.805 (99.972)\t\nTRAINING - Epoch: [90][80/97]\tTime 0.162 (0.184)\tData 0.000 (0.016)\tLoss 0.1105 (0.1034)\tPrec@1 96.094 (96.499)\tPrec@5 100.000 (99.976)\t\nTRAINING - Epoch: [90][90/97]\tTime 0.151 (0.180)\tData 0.000 (0.014)\tLoss 0.0680 (0.1030)\tPrec@1 98.242 (96.504)\tPrec@5 100.000 (99.979)\t\nEVALUATING - Epoch: [90][0/20]\tTime 1.081 (1.081)\tData 1.034 (1.034)\tLoss 0.2902 (0.2902)\tPrec@1 89.648 (89.648)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [90][10/20]\tTime 0.065 (0.187)\tData 0.020 (0.134)\tLoss 0.3351 (0.3892)\tPrec@1 90.820 (88.867)\tPrec@5 99.414 (99.556)\t\n\nResults - Epoch: 91\nTraining Loss 0.1032 \tTraining Prec@1 96.507 \tTraining Prec@5 99.978 \tValidation Loss 0.3797 \tValidation Prec@1 89.050 \tValidation Prec@5 99.620 \t\n\n\nStarting Epoch: 92\n\nTRAINING - Epoch: [91][0/97]\tTime 1.545 (1.545)\tData 1.244 (1.244)\tLoss 0.0904 (0.0904)\tPrec@1 97.266 (97.266)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [91][10/97]\tTime 0.150 (0.332)\tData 0.000 (0.117)\tLoss 0.0824 (0.0999)\tPrec@1 97.461 (96.733)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [91][20/97]\tTime 0.155 (0.254)\tData 0.000 (0.062)\tLoss 0.0827 (0.0989)\tPrec@1 97.070 (96.661)\tPrec@5 100.000 (99.953)\t\nTRAINING - Epoch: [91][30/97]\tTime 0.147 (0.224)\tData 0.000 (0.042)\tLoss 0.1094 (0.1025)\tPrec@1 95.898 (96.528)\tPrec@5 100.000 (99.956)\t\nTRAINING - Epoch: [91][40/97]\tTime 0.155 (0.208)\tData 0.000 (0.032)\tLoss 0.1227 (0.1027)\tPrec@1 95.508 (96.537)\tPrec@5 100.000 (99.967)\t\nTRAINING - Epoch: [91][50/97]\tTime 0.190 (0.200)\tData 0.004 (0.026)\tLoss 0.1138 (0.1034)\tPrec@1 95.703 (96.484)\tPrec@5 100.000 (99.966)\t\nTRAINING - Epoch: [91][60/97]\tTime 0.150 (0.194)\tData 0.000 (0.022)\tLoss 0.1086 (0.1036)\tPrec@1 96.094 (96.456)\tPrec@5 100.000 (99.971)\t\nTRAINING - Epoch: [91][70/97]\tTime 0.148 (0.189)\tData 0.000 (0.019)\tLoss 0.0964 (0.1036)\tPrec@1 96.680 (96.462)\tPrec@5 100.000 (99.972)\t\nTRAINING - Epoch: [91][80/97]\tTime 0.180 (0.186)\tData 0.000 (0.017)\tLoss 0.0849 (0.1031)\tPrec@1 97.070 (96.516)\tPrec@5 100.000 (99.973)\t\nTRAINING - Epoch: [91][90/97]\tTime 0.149 (0.182)\tData 0.000 (0.015)\tLoss 0.0878 (0.1031)\tPrec@1 96.875 (96.523)\tPrec@5 100.000 (99.974)\t\nEVALUATING - Epoch: [91][0/20]\tTime 1.047 (1.047)\tData 0.997 (0.997)\tLoss 0.2880 (0.2880)\tPrec@1 90.234 (90.234)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [91][10/20]\tTime 0.055 (0.188)\tData 0.000 (0.133)\tLoss 0.3197 (0.3781)\tPrec@1 90.625 (89.116)\tPrec@5 99.414 (99.609)\t\n\nResults - Epoch: 92\nTraining Loss 0.1031 \tTraining Prec@1 96.523 \tTraining Prec@5 99.974 \tValidation Loss 0.3718 \tValidation Prec@1 89.170 \tValidation Prec@5 99.650 \t\n\n\nStarting Epoch: 93\n\nTRAINING - Epoch: [92][0/97]\tTime 0.901 (0.901)\tData 0.636 (0.636)\tLoss 0.1090 (0.1090)\tPrec@1 96.094 (96.094)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [92][10/97]\tTime 0.198 (0.334)\tData 0.000 (0.093)\tLoss 0.1029 (0.0992)\tPrec@1 97.266 (96.875)\tPrec@5 100.000 (99.964)\t\nTRAINING - Epoch: [92][20/97]\tTime 0.166 (0.252)\tData 0.000 (0.049)\tLoss 0.1198 (0.0991)\tPrec@1 95.703 (96.819)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [92][30/97]\tTime 0.149 (0.222)\tData 0.000 (0.034)\tLoss 0.0899 (0.0997)\tPrec@1 97.266 (96.743)\tPrec@5 100.000 (99.987)\t\nTRAINING - Epoch: [92][40/97]\tTime 0.149 (0.206)\tData 0.000 (0.026)\tLoss 0.1268 (0.0998)\tPrec@1 95.508 (96.670)\tPrec@5 100.000 (99.990)\t\nTRAINING - Epoch: [92][50/97]\tTime 0.161 (0.197)\tData 0.000 (0.021)\tLoss 0.0945 (0.1001)\tPrec@1 96.289 (96.630)\tPrec@5 100.000 (99.992)\t\nTRAINING - Epoch: [92][60/97]\tTime 0.167 (0.192)\tData 0.000 (0.018)\tLoss 0.1114 (0.1002)\tPrec@1 96.289 (96.644)\tPrec@5 100.000 (99.994)\t\nTRAINING - Epoch: [92][70/97]\tTime 0.155 (0.187)\tData 0.000 (0.015)\tLoss 0.1097 (0.1006)\tPrec@1 96.680 (96.614)\tPrec@5 100.000 (99.994)\t\nTRAINING - Epoch: [92][80/97]\tTime 0.171 (0.184)\tData 0.002 (0.014)\tLoss 0.0822 (0.1009)\tPrec@1 97.852 (96.605)\tPrec@5 100.000 (99.993)\t\nTRAINING - Epoch: [92][90/97]\tTime 0.150 (0.180)\tData 0.000 (0.012)\tLoss 0.0660 (0.1005)\tPrec@1 98.828 (96.611)\tPrec@5 100.000 (99.994)\t\nEVALUATING - Epoch: [92][0/20]\tTime 1.254 (1.254)\tData 1.202 (1.202)\tLoss 0.2875 (0.2875)\tPrec@1 90.039 (90.039)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [92][10/20]\tTime 0.048 (0.201)\tData 0.000 (0.148)\tLoss 0.3343 (0.3817)\tPrec@1 90.820 (89.062)\tPrec@5 99.609 (99.663)\t\n\nResults - Epoch: 93\nTraining Loss 0.0998 \tTraining Prec@1 96.637 \tTraining Prec@5 99.994 \tValidation Loss 0.3762 \tValidation Prec@1 89.140 \tValidation Prec@5 99.690 \t\n\n\nStarting Epoch: 94\n\nTRAINING - Epoch: [93][0/97]\tTime 1.503 (1.503)\tData 1.209 (1.209)\tLoss 0.1017 (0.1017)\tPrec@1 96.484 (96.484)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [93][10/97]\tTime 0.146 (0.334)\tData 0.000 (0.113)\tLoss 0.0771 (0.0912)\tPrec@1 97.461 (96.804)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [93][20/97]\tTime 0.173 (0.252)\tData 0.005 (0.060)\tLoss 0.1012 (0.0936)\tPrec@1 96.484 (96.717)\tPrec@5 100.000 (99.991)\t\nTRAINING - Epoch: [93][30/97]\tTime 0.153 (0.222)\tData 0.000 (0.041)\tLoss 0.1084 (0.0958)\tPrec@1 96.289 (96.717)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [93][40/97]\tTime 0.152 (0.207)\tData 0.000 (0.031)\tLoss 0.1051 (0.0960)\tPrec@1 95.703 (96.708)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [93][50/97]\tTime 0.169 (0.198)\tData 0.000 (0.025)\tLoss 0.0711 (0.0971)\tPrec@1 97.852 (96.706)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [93][60/97]\tTime 0.147 (0.192)\tData 0.000 (0.022)\tLoss 0.0761 (0.0958)\tPrec@1 97.266 (96.757)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [93][70/97]\tTime 0.179 (0.188)\tData 0.000 (0.019)\tLoss 0.0965 (0.0970)\tPrec@1 96.680 (96.715)\tPrec@5 100.000 (99.975)\t\nTRAINING - Epoch: [93][80/97]\tTime 0.151 (0.184)\tData 0.000 (0.016)\tLoss 0.1186 (0.0973)\tPrec@1 95.703 (96.706)\tPrec@5 100.000 (99.978)\t\nTRAINING - Epoch: [93][90/97]\tTime 0.150 (0.181)\tData 0.000 (0.015)\tLoss 0.1265 (0.0974)\tPrec@1 95.508 (96.735)\tPrec@5 100.000 (99.976)\t\nEVALUATING - Epoch: [93][0/20]\tTime 1.002 (1.002)\tData 0.954 (0.954)\tLoss 0.2925 (0.2925)\tPrec@1 89.453 (89.453)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [93][10/20]\tTime 0.050 (0.191)\tData 0.000 (0.136)\tLoss 0.3347 (0.3869)\tPrec@1 90.039 (88.849)\tPrec@5 99.609 (99.592)\t\n\nResults - Epoch: 94\nTraining Loss 0.0977 \tTraining Prec@1 96.760 \tTraining Prec@5 99.976 \tValidation Loss 0.3789 \tValidation Prec@1 88.940 \tValidation Prec@5 99.630 \t\n\n\nStarting Epoch: 95\n\nTRAINING - Epoch: [94][0/97]\tTime 1.478 (1.478)\tData 1.239 (1.239)\tLoss 0.1090 (0.1090)\tPrec@1 97.070 (97.070)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [94][10/97]\tTime 0.165 (0.334)\tData 0.000 (0.118)\tLoss 0.0972 (0.0942)\tPrec@1 97.070 (97.141)\tPrec@5 100.000 (99.947)\t\nTRAINING - Epoch: [94][20/97]\tTime 0.165 (0.253)\tData 0.006 (0.063)\tLoss 0.0812 (0.0959)\tPrec@1 97.461 (96.977)\tPrec@5 100.000 (99.953)\t\nTRAINING - Epoch: [94][30/97]\tTime 0.182 (0.225)\tData 0.000 (0.043)\tLoss 0.0993 (0.0961)\tPrec@1 97.266 (96.888)\tPrec@5 100.000 (99.962)\t\nTRAINING - Epoch: [94][40/97]\tTime 0.147 (0.208)\tData 0.000 (0.032)\tLoss 0.0761 (0.0956)\tPrec@1 97.656 (96.885)\tPrec@5 100.000 (99.971)\t\nTRAINING - Epoch: [94][50/97]\tTime 0.157 (0.198)\tData 0.000 (0.026)\tLoss 0.0839 (0.0963)\tPrec@1 97.266 (96.844)\tPrec@5 100.000 (99.969)\t\nTRAINING - Epoch: [94][60/97]\tTime 0.189 (0.193)\tData 0.000 (0.022)\tLoss 0.0886 (0.0967)\tPrec@1 96.875 (96.814)\tPrec@5 100.000 (99.971)\t\nTRAINING - Epoch: [94][70/97]\tTime 0.159 (0.188)\tData 0.000 (0.019)\tLoss 0.1102 (0.0968)\tPrec@1 95.508 (96.776)\tPrec@5 100.000 (99.972)\t\nTRAINING - Epoch: [94][80/97]\tTime 0.149 (0.185)\tData 0.000 (0.017)\tLoss 0.0993 (0.0976)\tPrec@1 96.094 (96.711)\tPrec@5 100.000 (99.969)\t\nTRAINING - Epoch: [94][90/97]\tTime 0.149 (0.181)\tData 0.000 (0.015)\tLoss 0.0924 (0.0984)\tPrec@1 97.070 (96.678)\tPrec@5 100.000 (99.970)\t\nEVALUATING - Epoch: [94][0/20]\tTime 1.299 (1.299)\tData 1.238 (1.238)\tLoss 0.2763 (0.2763)\tPrec@1 90.234 (90.234)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [94][10/20]\tTime 0.067 (0.200)\tData 0.023 (0.140)\tLoss 0.3452 (0.3869)\tPrec@1 90.625 (88.956)\tPrec@5 99.414 (99.627)\t\n\nResults - Epoch: 95\nTraining Loss 0.0989 \tTraining Prec@1 96.645 \tTraining Prec@5 99.968 \tValidation Loss 0.3780 \tValidation Prec@1 89.110 \tValidation Prec@5 99.630 \t\n\n\nStarting Epoch: 96\n\nTRAINING - Epoch: [95][0/97]\tTime 1.471 (1.471)\tData 1.186 (1.186)\tLoss 0.0856 (0.0856)\tPrec@1 97.070 (97.070)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [95][10/97]\tTime 0.210 (0.335)\tData 0.000 (0.111)\tLoss 0.1130 (0.1032)\tPrec@1 96.094 (96.626)\tPrec@5 100.000 (99.929)\t\nTRAINING - Epoch: [95][20/97]\tTime 0.183 (0.252)\tData 0.007 (0.059)\tLoss 0.1239 (0.1021)\tPrec@1 96.289 (96.680)\tPrec@5 100.000 (99.953)\t\nTRAINING - Epoch: [95][30/97]\tTime 0.151 (0.224)\tData 0.000 (0.040)\tLoss 0.1087 (0.0989)\tPrec@1 96.484 (96.667)\tPrec@5 100.000 (99.962)\t\nTRAINING - Epoch: [95][40/97]\tTime 0.173 (0.209)\tData 0.007 (0.031)\tLoss 0.0996 (0.0976)\tPrec@1 97.070 (96.708)\tPrec@5 100.000 (99.962)\t\nTRAINING - Epoch: [95][50/97]\tTime 0.156 (0.199)\tData 0.000 (0.025)\tLoss 0.0686 (0.0974)\tPrec@1 97.852 (96.703)\tPrec@5 100.000 (99.966)\t\nTRAINING - Epoch: [95][60/97]\tTime 0.156 (0.193)\tData 0.000 (0.021)\tLoss 0.0949 (0.0981)\tPrec@1 97.070 (96.654)\tPrec@5 100.000 (99.971)\t\nTRAINING - Epoch: [95][70/97]\tTime 0.149 (0.188)\tData 0.000 (0.018)\tLoss 0.1056 (0.0988)\tPrec@1 96.289 (96.641)\tPrec@5 100.000 (99.972)\t\nTRAINING - Epoch: [95][80/97]\tTime 0.153 (0.185)\tData 0.000 (0.016)\tLoss 0.0978 (0.0987)\tPrec@1 97.070 (96.658)\tPrec@5 100.000 (99.976)\t\nTRAINING - Epoch: [95][90/97]\tTime 0.150 (0.181)\tData 0.000 (0.015)\tLoss 0.0926 (0.0992)\tPrec@1 97.266 (96.652)\tPrec@5 100.000 (99.979)\t\nEVALUATING - Epoch: [95][0/20]\tTime 0.675 (0.675)\tData 0.623 (0.623)\tLoss 0.2818 (0.2818)\tPrec@1 90.430 (90.430)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [95][10/20]\tTime 0.094 (0.196)\tData 0.050 (0.147)\tLoss 0.3376 (0.3840)\tPrec@1 90.234 (89.276)\tPrec@5 99.609 (99.663)\t\n\nResults - Epoch: 96\nTraining Loss 0.0989 \tTraining Prec@1 96.656 \tTraining Prec@5 99.980 \tValidation Loss 0.3761 \tValidation Prec@1 89.220 \tValidation Prec@5 99.700 \t\n\n\nStarting Epoch: 97\n\nTRAINING - Epoch: [96][0/97]\tTime 1.572 (1.572)\tData 1.337 (1.337)\tLoss 0.1179 (0.1179)\tPrec@1 95.117 (95.117)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [96][10/97]\tTime 0.149 (0.337)\tData 0.000 (0.124)\tLoss 0.0648 (0.0927)\tPrec@1 98.633 (96.804)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [96][20/97]\tTime 0.170 (0.256)\tData 0.000 (0.065)\tLoss 0.0876 (0.0910)\tPrec@1 96.484 (96.875)\tPrec@5 100.000 (99.991)\t\nTRAINING - Epoch: [96][30/97]\tTime 0.155 (0.224)\tData 0.000 (0.045)\tLoss 0.0881 (0.0942)\tPrec@1 97.070 (96.774)\tPrec@5 100.000 (99.987)\t\nTRAINING - Epoch: [96][40/97]\tTime 0.160 (0.209)\tData 0.000 (0.034)\tLoss 0.0938 (0.0952)\tPrec@1 96.484 (96.770)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [96][50/97]\tTime 0.150 (0.199)\tData 0.000 (0.028)\tLoss 0.0983 (0.0953)\tPrec@1 96.289 (96.714)\tPrec@5 100.000 (99.985)\t\nTRAINING - Epoch: [96][60/97]\tTime 0.154 (0.193)\tData 0.000 (0.023)\tLoss 0.1059 (0.0954)\tPrec@1 95.703 (96.705)\tPrec@5 100.000 (99.984)\t\nTRAINING - Epoch: [96][70/97]\tTime 0.152 (0.188)\tData 0.000 (0.020)\tLoss 0.0888 (0.0956)\tPrec@1 96.289 (96.688)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [96][80/97]\tTime 0.153 (0.186)\tData 0.000 (0.018)\tLoss 0.0860 (0.0964)\tPrec@1 97.266 (96.665)\tPrec@5 100.000 (99.983)\t\nTRAINING - Epoch: [96][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.016)\tLoss 0.0995 (0.0967)\tPrec@1 96.875 (96.639)\tPrec@5 100.000 (99.983)\t\nEVALUATING - Epoch: [96][0/20]\tTime 1.094 (1.094)\tData 1.038 (1.038)\tLoss 0.2965 (0.2965)\tPrec@1 90.234 (90.234)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [96][10/20]\tTime 0.056 (0.191)\tData 0.000 (0.138)\tLoss 0.3361 (0.3877)\tPrec@1 90.430 (89.258)\tPrec@5 99.414 (99.609)\t\n\nResults - Epoch: 97\nTraining Loss 0.0962 \tTraining Prec@1 96.660 \tTraining Prec@5 99.984 \tValidation Loss 0.3805 \tValidation Prec@1 89.180 \tValidation Prec@5 99.670 \t\n\n\nStarting Epoch: 98\n\nTRAINING - Epoch: [97][0/97]\tTime 1.263 (1.263)\tData 0.996 (0.996)\tLoss 0.1034 (0.1034)\tPrec@1 96.484 (96.484)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [97][10/97]\tTime 0.164 (0.341)\tData 0.000 (0.093)\tLoss 0.0934 (0.0877)\tPrec@1 97.266 (97.053)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [97][20/97]\tTime 0.155 (0.254)\tData 0.000 (0.050)\tLoss 0.0976 (0.0910)\tPrec@1 95.117 (96.884)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [97][30/97]\tTime 0.156 (0.225)\tData 0.000 (0.034)\tLoss 0.0766 (0.0913)\tPrec@1 97.461 (96.869)\tPrec@5 100.000 (99.975)\t\nTRAINING - Epoch: [97][40/97]\tTime 0.153 (0.209)\tData 0.000 (0.026)\tLoss 0.0613 (0.0906)\tPrec@1 98.242 (96.894)\tPrec@5 99.805 (99.976)\t\nTRAINING - Epoch: [97][50/97]\tTime 0.154 (0.200)\tData 0.000 (0.021)\tLoss 0.0678 (0.0925)\tPrec@1 98.242 (96.798)\tPrec@5 100.000 (99.977)\t\nTRAINING - Epoch: [97][60/97]\tTime 0.170 (0.194)\tData 0.002 (0.018)\tLoss 0.1100 (0.0937)\tPrec@1 95.898 (96.817)\tPrec@5 100.000 (99.974)\t\nTRAINING - Epoch: [97][70/97]\tTime 0.150 (0.189)\tData 0.000 (0.016)\tLoss 0.0950 (0.0940)\tPrec@1 96.680 (96.765)\tPrec@5 100.000 (99.978)\t\nTRAINING - Epoch: [97][80/97]\tTime 0.152 (0.185)\tData 0.000 (0.014)\tLoss 0.0878 (0.0952)\tPrec@1 97.266 (96.740)\tPrec@5 100.000 (99.973)\t\nTRAINING - Epoch: [97][90/97]\tTime 0.150 (0.182)\tData 0.000 (0.013)\tLoss 0.0789 (0.0949)\tPrec@1 97.266 (96.746)\tPrec@5 100.000 (99.970)\t\nEVALUATING - Epoch: [97][0/20]\tTime 1.029 (1.029)\tData 0.976 (0.976)\tLoss 0.2897 (0.2897)\tPrec@1 90.234 (90.234)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [97][10/20]\tTime 0.050 (0.183)\tData 0.000 (0.130)\tLoss 0.3308 (0.3929)\tPrec@1 90.430 (89.240)\tPrec@5 99.609 (99.574)\t\n\nResults - Epoch: 98\nTraining Loss 0.0946 \tTraining Prec@1 96.754 \tTraining Prec@5 99.970 \tValidation Loss 0.3842 \tValidation Prec@1 89.160 \tValidation Prec@5 99.600 \t\n\n\nStarting Epoch: 99\n\nTRAINING - Epoch: [98][0/97]\tTime 1.459 (1.459)\tData 1.159 (1.159)\tLoss 0.0841 (0.0841)\tPrec@1 97.266 (97.266)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [98][10/97]\tTime 0.176 (0.335)\tData 0.000 (0.109)\tLoss 0.0995 (0.0959)\tPrec@1 97.070 (96.680)\tPrec@5 100.000 (99.982)\t\nTRAINING - Epoch: [98][20/97]\tTime 0.148 (0.252)\tData 0.000 (0.058)\tLoss 0.0763 (0.0918)\tPrec@1 98.047 (96.949)\tPrec@5 100.000 (99.991)\t\nTRAINING - Epoch: [98][30/97]\tTime 0.159 (0.223)\tData 0.005 (0.039)\tLoss 0.0966 (0.0933)\tPrec@1 95.898 (96.888)\tPrec@5 100.000 (99.994)\t\nTRAINING - Epoch: [98][40/97]\tTime 0.169 (0.208)\tData 0.000 (0.030)\tLoss 0.0853 (0.0929)\tPrec@1 96.484 (96.865)\tPrec@5 100.000 (99.995)\t\nTRAINING - Epoch: [98][50/97]\tTime 0.169 (0.199)\tData 0.010 (0.024)\tLoss 0.1120 (0.0936)\tPrec@1 96.094 (96.856)\tPrec@5 99.805 (99.985)\t\nTRAINING - Epoch: [98][60/97]\tTime 0.154 (0.193)\tData 0.000 (0.021)\tLoss 0.0765 (0.0939)\tPrec@1 97.656 (96.859)\tPrec@5 100.000 (99.987)\t\nTRAINING - Epoch: [98][70/97]\tTime 0.160 (0.189)\tData 0.000 (0.018)\tLoss 0.0751 (0.0941)\tPrec@1 97.461 (96.842)\tPrec@5 100.000 (99.986)\t\nTRAINING - Epoch: [98][80/97]\tTime 0.155 (0.186)\tData 0.000 (0.016)\tLoss 0.1157 (0.0952)\tPrec@1 96.289 (96.771)\tPrec@5 99.805 (99.983)\t\nTRAINING - Epoch: [98][90/97]\tTime 0.149 (0.182)\tData 0.000 (0.014)\tLoss 0.0905 (0.0956)\tPrec@1 98.047 (96.789)\tPrec@5 100.000 (99.983)\t\nEVALUATING - Epoch: [98][0/20]\tTime 1.034 (1.034)\tData 0.983 (0.983)\tLoss 0.2923 (0.2923)\tPrec@1 90.234 (90.234)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [98][10/20]\tTime 0.050 (0.194)\tData 0.000 (0.136)\tLoss 0.3418 (0.3975)\tPrec@1 89.844 (89.169)\tPrec@5 99.609 (99.609)\t\n\nResults - Epoch: 99\nTraining Loss 0.0961 \tTraining Prec@1 96.774 \tTraining Prec@5 99.984 \tValidation Loss 0.3884 \tValidation Prec@1 89.110 \tValidation Prec@5 99.630 \t\n\n\nStarting Epoch: 100\n\nTRAINING - Epoch: [99][0/97]\tTime 1.329 (1.329)\tData 1.039 (1.039)\tLoss 0.0872 (0.0872)\tPrec@1 96.875 (96.875)\tPrec@5 100.000 (100.000)\t\nTRAINING - Epoch: [99][10/97]\tTime 0.150 (0.336)\tData 0.000 (0.097)\tLoss 0.0806 (0.0941)\tPrec@1 97.266 (96.839)\tPrec@5 99.805 (99.964)\t\nTRAINING - Epoch: [99][20/97]\tTime 0.163 (0.253)\tData 0.000 (0.052)\tLoss 0.0765 (0.0904)\tPrec@1 97.070 (97.033)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [99][30/97]\tTime 0.163 (0.225)\tData 0.000 (0.035)\tLoss 0.1364 (0.0939)\tPrec@1 95.312 (96.925)\tPrec@5 100.000 (99.987)\t\nTRAINING - Epoch: [99][40/97]\tTime 0.196 (0.210)\tData 0.000 (0.027)\tLoss 0.0974 (0.0919)\tPrec@1 97.266 (96.994)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [99][50/97]\tTime 0.180 (0.200)\tData 0.000 (0.022)\tLoss 0.1257 (0.0925)\tPrec@1 95.703 (96.975)\tPrec@5 100.000 (99.981)\t\nTRAINING - Epoch: [99][60/97]\tTime 0.150 (0.194)\tData 0.000 (0.019)\tLoss 0.0905 (0.0930)\tPrec@1 97.266 (96.945)\tPrec@5 100.000 (99.984)\t\nTRAINING - Epoch: [99][70/97]\tTime 0.150 (0.189)\tData 0.000 (0.016)\tLoss 0.1126 (0.0936)\tPrec@1 95.508 (96.869)\tPrec@5 99.805 (99.983)\t\nTRAINING - Epoch: [99][80/97]\tTime 0.159 (0.185)\tData 0.000 (0.014)\tLoss 0.1079 (0.0933)\tPrec@1 95.703 (96.865)\tPrec@5 100.000 (99.978)\t\nTRAINING - Epoch: [99][90/97]\tTime 0.150 (0.181)\tData 0.000 (0.013)\tLoss 0.0903 (0.0941)\tPrec@1 97.461 (96.821)\tPrec@5 100.000 (99.979)\t\nEVALUATING - Epoch: [99][0/20]\tTime 1.281 (1.281)\tData 1.229 (1.229)\tLoss 0.2836 (0.2836)\tPrec@1 90.430 (90.430)\tPrec@5 99.805 (99.805)\t\nEVALUATING - Epoch: [99][10/20]\tTime 0.082 (0.193)\tData 0.038 (0.140)\tLoss 0.3362 (0.3935)\tPrec@1 90.430 (89.134)\tPrec@5 99.414 (99.574)\t\n\nResults - Epoch: 100\nTraining Loss 0.0936 \tTraining Prec@1 96.837 \tTraining Prec@5 99.980 \tValidation Loss 0.3848 \tValidation Prec@1 89.200 \tValidation Prec@5 99.620 \t\n\n"
]
],
[
[
"## View Results",
"_____no_output_____"
]
],
[
[
"from IPython.display import HTML\n\nHTML(filename=\"./results/resnet18_test/results.html\")",
"_____no_output_____"
],
[
"",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4ab7ffa20d79f1370cc1a9296773528595ebb6d6
| 30,285 |
ipynb
|
Jupyter Notebook
|
notebooks/afMSIT_Preprocessing.ipynb
|
szorowi1/EMOTE-afMSIT
|
16347125634f584175dcb16427db6a99740319fd
|
[
"MIT"
] | 1 |
2019-05-26T23:31:21.000Z
|
2019-05-26T23:31:21.000Z
|
notebooks/afMSIT_Preprocessing.ipynb
|
szorowi1/EMOTE-afMSIT
|
16347125634f584175dcb16427db6a99740319fd
|
[
"MIT"
] | null | null | null |
notebooks/afMSIT_Preprocessing.ipynb
|
szorowi1/EMOTE-afMSIT
|
16347125634f584175dcb16427db6a99740319fd
|
[
"MIT"
] | 1 |
2021-03-27T08:48:06.000Z
|
2021-03-27T08:48:06.000Z
| 40.006605 | 128 | 0.496781 |
[
[
[
"# Section 1: Preprocessing\n## Behavior Analysis\n### Generate trial regressors",
"_____no_output_____"
]
],
[
[
"import os\nimport numpy as np\nfrom pandas import concat, read_csv\nfrom scipy.stats import gamma\ndef normalize(arr): return (arr - arr.min()) / (arr.max() - arr.min())\n\nroot_dir = '/space/sophia/2/users/EMOTE-DBS/afMSIT/behavior'\nsubjects = ['BRTU', 'CHDR', 'CRDA', 'JADE', 'JASE', 'M5', 'MEWA', 'S2']\nthreshold = 0.005\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Load / Concatenate / Prepare Data.\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\ndf = []\nfor subject in subjects:\n \n ## Load CSV.\n csv = read_csv(os.path.join(root_dir,'%s_msit_data.txt' %subject))\n \n ## Limit columns.\n csv = csv[['SubjID','trial','iaps','DBS','interference','valence','arousal','responseTime','responseCorrect']]\n \n ## Rename columns.\n csv.columns = ['Subject', 'Trial', 'IAPS', 'DBS', 'Interference', 'Valence_Obj', 'Arousal_Obj', 'RT', 'Accuracy']\n\n ## Load IAPS ratings.\n iaps = read_csv(os.path.join(root_dir,'%s_IAPS_SAM.csv' %subject))\n iaps = iaps[['IAPS_Number','Valence','Arousal']]\n iaps.columns = ['IAPS','Valence_Subj','Arousal_Subj']\n\n ## Merge. Append.\n csv = csv.merge(iaps, on='IAPS')\n cols = ['Subject', 'Trial', 'IAPS', 'DBS', 'Interference', 'Valence_Obj', 'Arousal_Obj', \n 'Valence_Subj', 'Arousal_Subj', 'RT', 'Accuracy']\n csv = csv[cols]\n df.append(csv)\n\n## Merge data. Sort.\ndf = concat(df)\ndf['DBS'] = np.where(df['DBS']=='DBSoff',0,1)\ndf = df.sort_values(['Subject','DBS','Trial']).reset_index(drop=True)\n\n## Normalize regressors.\ndf['nsArousal'] = normalize(df.Arousal_Subj)\ndf['nsValence'] = normalize(df.Valence_Subj)\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Determine Trials for Inclusion/Exclusion.\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n\n## Set missing RTs to NaNs.\ndf['RT'] = np.where(df.Accuracy==-1, np.nan, df.RT)\ndf['Accuracy'] = np.where(df.Accuracy==-1, np.nan, df.Accuracy)\ndf['Missing'] = df.Accuracy.isnull().astype(int)\n\n## Add Error column.\ndf['Error'] = 1 - df.Accuracy\n\n## Add Post-Error Column.\ndf['PostError'] = 0\nfor subject in df.Subject.unique():\n error = df.loc[df.Subject==subject,'Error']\n posterror = np.insert(np.roll(error,1)[1:], 0, 0)\n df.loc[df.Subject==subject,'PostError'] = posterror\n\n## Iteratively detect outliers across subjects by fitting a Gamma distribution.\ndf['GammaCDF'], df['Outlier'] = 0, 0\nfor subject in df.Subject.unique():\n \n ## Fit Gamma to reaction time distribution.\n shape, loc, scale = gamma.fit(df.loc[(df.Subject==subject)&(~df.RT.isnull()),'RT'], floc=0)\n \n ## Find outliers given likelihood threshold.\n cdf = gamma.cdf(df.loc[(df.Subject==subject)&(~df.RT.isnull()),'RT'], shape, loc=loc, scale=scale)\n outliers = (cdf < threshold) | (cdf > 1 - threshold)\n \n ## Append information.\n df.loc[(df.Subject==subject)&(~df.RT.isnull()), 'GammaCDF'] += cdf\n df.loc[(df.Subject==subject)&(~df.RT.isnull()), 'Outlier'] += outliers.astype(int)\n \n## Generate exclude.\ndf['Exclude'] = np.where( df[['Missing','Error','PostError','Outlier']].sum(axis=1), 1, 0)\nprint '%s trials (%0.2f%%) excluded.' %(df.Exclude.sum(), df.Exclude.mean())\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Save.\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\ndf.to_csv('%s/afMSIT_group_data.csv' %root_dir, index=False)",
"_____no_output_____"
]
],
[
[
"## Parcellation\n### Make EMOTE Labels",
"_____no_output_____"
]
],
[
[
"import os, shutil\nimport numpy as np\nimport pylab as plt\nfrom mne import read_label, read_source_spaces, read_surface, set_log_level\nset_log_level(verbose=False)\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Define parameters.\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n\nfs_dir = '/space/sophia/2/users/EMOTE-DBS/freesurfs'\nsubject = 'BRTU'\n\nparc = 'laus250'\nlabel_dir = os.path.join(fs_dir,subject,'label',parc)\nout_dir = os.path.join(fs_dir,subject,'label','april2016')\n\nif os.path.isdir(out_dir): shutil.rmtree(out_dir)\nos.makedirs(out_dir)\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Build Left Hemisphere Labels.\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n\nhemi = 'lh'\nrr, _ = read_surface(os.path.join(fs_dir, subject, 'surf', '%s.inflated' %hemi))\nsrc = read_source_spaces(os.path.join(fs_dir, subject, 'bem', '%s-oct-6-src.fif' %subject))[0]\n\nlhdict = {'dlpfc_1-lh':['caudalmiddlefrontal_1', 'caudalmiddlefrontal_5', 'caudalmiddlefrontal_6'],\n 'dlpfc_2-lh':['caudalmiddlefrontal_2', 'caudalmiddlefrontal_3', 'caudalmiddlefrontal_4'],\n 'dlpfc_3-lh':['rostralmiddlefrontal_2', 'rostralmiddlefrontal_3'],\n 'dlpfc_4-lh':['rostralmiddlefrontal_1', 'rostralmiddlefrontal_5'],\n 'dlpfc_5-lh':['parstriangularis_2', 'parsopercularis_2'],\n 'dlpfc_6-lh':['parsopercularis_3', 'parsopercularis_4'],\n 'racc-lh':['rostralanteriorcingulate_1','rostralanteriorcingulate_2'],\n 'dacc-lh':['caudalanteriorcingulate_1','caudalanteriorcingulate_2',],\n 'pcc-lh':['posteriorcingulate_2','posteriorcingulate_3']}\n\nfor k,V in lhdict.iteritems():\n \n label = np.sum([read_label(os.path.join(label_dir,'%s-%s.label' %(v,hemi)), subject=subject) \n for v in V])\n n_vert = np.intersect1d(src['vertno'], label.vertices).shape[0]\n print '%s\\t%s' %(n_vert,k)\n label.save(os.path.join(out_dir, '%s.label' %k))\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Build Right Hemisphere Labels.\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n\nhemi = 'rh'\nrr, _ = read_surface(os.path.join(fs_dir, subject, 'surf', '%s.inflated' %hemi))\nsrc = read_source_spaces(os.path.join(fs_dir, subject, 'bem', '%s-oct-6-src.fif' %subject))[1]\n\nrhdict = {'dlpfc_1-rh':['caudalmiddlefrontal_1', 'caudalmiddlefrontal_2', 'caudalmiddlefrontal_5'],\n 'dlpfc_2-rh':['caudalmiddlefrontal_3', 'caudalmiddlefrontal_4'],\n 'dlpfc_3-rh':['rostralmiddlefrontal_2', 'rostralmiddlefrontal_3'],\n 'dlpfc_4-rh':['rostralmiddlefrontal_1', 'rostralmiddlefrontal_5'],\n 'dlpfc_5-rh':['parstriangularis_2', 'parsopercularis_1'],\n 'dlpfc_6-rh':['parsopercularis_3', 'parsopercularis_4'],\n 'racc-rh':['rostralanteriorcingulate_1','rostralanteriorcingulate_2'],\n 'dacc-rh':['caudalanteriorcingulate_1','caudalanteriorcingulate_2','caudalanteriorcingulate_3'],\n 'pcc-rh':['posteriorcingulate_2','posteriorcingulate_3']}\n\nfor k,V in rhdict.iteritems():\n label = np.sum([read_label(os.path.join(label_dir,'%s-%s.label' %(v,hemi)), subject=subject) \n for v in V])\n n_vert = np.intersect1d(src['vertno'], label.vertices).shape[0]\n print '%s\\t%s' %(n_vert,k)\n label.save(os.path.join(out_dir, '%s.label' %k))",
"_____no_output_____"
]
],
[
[
"## Preprocesing 1: Raw Data\n### Fixing MEWA: Digitization\nSomething got way messed up. Here we make MNE knows what is EEG and what is extra points.\n\nNOTE: Copied over one of the original files for MEWA and renamed it MEWA_msit_unmasked_raw.fif",
"_____no_output_____"
]
],
[
[
"import os\nimport numpy as np\nfrom mne.io import Raw\nfrom pandas import read_table\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Specify parameters.\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\nroot_dir = '/space/sophia/2/users/EMOTE-DBS/afMSIT_april2016'\nraw_file = 'MEWA_msit_unmasked_raw.fif'\nout_file = 'MEWA_msit_raw.fif'\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Load and prepare digitizations.\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n\n## Load data. Get digitization from raw.\nraw = Raw(os.path.join(root_dir,'raw',raw_file),preload=False,verbose=False)\ndigitization = raw.info['dig']\n\n## The last 101 points are extra. Set them to kind=4.\nfor d in digitization[-101:]: d['kind'] = 4\n \n## Get coordinates for EEG points (excluding ref/EOG).\nrr = np.array([d['r'] for d in dig if d['kind']==3])[:-2]\n\n## Get channels\nchs = raw.info['chs']\n\n## Update location information. This was a huge pain in the ass to figure out.\n## We ignore the first four channels (Triggers, EOG) and the last channel (STI014).\nfor ch, r in zip(chs[4:-1], rr): ch['loc'][:3] = r \n\n## Update digitization/chs.\nraw.info['dig'] = digitization\nraw.info['chs'] = chs\nraw.save(os.path.join(root_dir,'raw',out_file), overwrite=True)",
"_____no_output_____"
]
],
[
[
"### Fixing MEWA: Masking channel jumps\nTime windows were manually inspected. This step isn't strictly necessary but seemed to help with EOG projections.\n\nNOTE: Copied over one of the original files for MEWA and renamed it MEWA_msit_unmasked_raw.fif",
"_____no_output_____"
]
],
[
[
"import os\nimport numpy as np\nimport pylab as plt\nfrom mne.io import Raw, RawArray\n\n## Specify parameters.\nroot_dir = '/space/sophia/2/users/EMOTE-DBS/afMSIT_april2016'\nraw_file = 'MEWA_msit_unmasked_raw.fif'\n\n## Load data.\nraw = Raw(os.path.join(root_dir,'raw',raw_file),preload=True,verbose=False)\n\n## Get data in matrix form.\ndata = raw._data\n\n## Get list of usuable channels\nch_info = [(n,ch) for n,ch in enumerate(raw.ch_names)]\ngood_ch = [(n,ch) for n,ch in ch_info if ch not in raw.info['bads']]\ngood_ch = np.array(good_ch)[4:-1]\n\n## Make mask.\nmask = np.zeros(data.shape[1])\ntimes = [(384,394), (663,669)]\nfor t1, t2 in times:\n mask[(raw.times >= t1) & (raw.times <= t2)] += 1\nmask = mask.astype(bool)\n\n## Apply mask.\nfor ch in good_ch[:,0].astype(int):\n data[ch,mask] = 0\n\n## Make new array. Save.\nraw = RawArray(data, raw.info, first_samp=raw.first_samp)\nraw.add_eeg_average_proj()\nraw.save(os.path.join(root_dir,'raw','MEWA_msit_raw.fif'), overwrite=True, verbose=False)",
"_____no_output_____"
]
],
[
[
"### Projections: EOG",
"_____no_output_____"
]
],
[
[
"import os\nfrom mne import write_proj\nfrom mne.preprocessing import compute_proj_eog\nfrom mne.io import Raw\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Setup\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n## File params.\nroot_dir = '/space/sophia/2/users/EMOTE-DBS/afMSIT_april2016'\nsubjects = ['BRTU', 'CHDR', 'CRDA', 'JADE', 'JASE', 'M5', 'MEWA', 'S2']\nsubjects = ['MEWA']\n\n# NOTE: Not all subjects work with EOG channel = EOG. \n# Some require other frontal channels due to concatenation.\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Main Loop.\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\nfor subj in subjects: \n print 'Making EOG file for %s.' %subj\n \n ## Load files.\n raw_file = os.path.join( root_dir, 'raw', '%s_msit_raw.fif' %subj )\n raw = Raw(raw_file, preload=True, verbose=False, add_eeg_ref=False)\n raw.del_proj(0)\n ## Make EOG proj. Save.\n proj, _ = compute_proj_eog(raw, n_eeg = 4, average=True, filter_length='20s',\n reject=dict(eeg=5e-4), flat=dict(eeg=5e-8), ch_name='F2', n_jobs=3)\n write_proj(os.path.join( root_dir, 'raw', '%s_msit_eog-proj.fif' %subj ), proj)",
"_____no_output_____"
]
],
[
[
"### Projections: ECG",
"_____no_output_____"
]
],
[
[
"import os\nfrom mne import read_proj, write_proj\nfrom mne.preprocessing import compute_proj_ecg\nfrom mne.io import Raw\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Setup\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n## File params.\nroot_dir = '/space/sophia/2/users/EMOTE-DBS/afMSIT_april2016'\nsubjects = ['CHDR']\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Main Loop.\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\nfor subj in subjects: \n print 'Making ECG file for %s.' %subj\n \n ## Load files.\n raw_file = os.path.join( root_dir, 'raw', '%s_msit_raw.fif' %subj )\n eog_file = os.path.join( root_dir, 'raw', '%s_msit-proj.fif' %subj )\n raw = Raw(raw_file, preload=True, verbose=False)\n eog_proj = read_proj(eog_file)\n raw.add_proj(eog_proj, remove_existing=True)\n raw.apply_proj()\n \n ## Make ECG proj. Save.\n ecg_proj, _ = compute_proj_ecg(raw, n_eeg = 4, h_freq = 35., average=True, filter_length='20s',\n reject=dict(eeg=5e-4), flat=dict(eeg=5e-8), ch_name='P9', n_jobs=3)\n proj = eog_proj + [ecg for ecg in ecg_proj if ecg['desc'] not in [eog['desc'] for eog in eog_proj]]\n write_proj(os.path.join( root_dir, 'raw', '%s_msit-proj.fif' %subj ), proj)",
"_____no_output_____"
]
],
[
[
"## Preprocessing 2: Epoching\n### Make Forward Solutions",
"_____no_output_____"
]
],
[
[
"import os\nfrom mne import read_trans, read_bem_solution, read_source_spaces\nfrom mne import make_forward_solution, write_forward_solution\nfrom mne.io import Raw\n\n## Subject level parameters.\nsubjects = ['BRTU', 'CHDR', 'CRDA', 'JADE', 'JASE', 'M5', 'MEWA', 'S2']\ntask = 'msit'\n\n## Main loop.\nroot_dir = '/autofs/space/sophia_002/users/EMOTE-DBS/afMSIT_april2016'\nfs_dir = '/autofs/space/sophia_002/users/EMOTE-DBS/freesurfs'\n\nfor subject in subjects:\n \n print 'Making forward solution for %s.' %subject\n \n ## Load files.\n raw = Raw(os.path.join(root_dir, 'raw', '%s_msit_raw.fif' %subject), preload=False, verbose=False)\n trans = read_trans(os.path.join(fs_dir,subject,'mri','T1-neuromag','sets','COR-%s.fif' %subject))\n src = read_source_spaces(os.path.join(fs_dir,subject,'bem','%s-oct-6p-src.fif' %subject), verbose=False)\n bem = read_bem_solution(os.path.join(fs_dir,subject,'bem','%s-5120-5120-5120-bem-sol.fif' %subject), verbose=False)\n \n ## Compute and save forward solution.\n make_forward_solution(raw.info, trans, src, bem, fname=os.path.join(root_dir,'fwd','%s_msit-fwd.fif' %subject),\n meg=False, eeg=True, mindist=1.0, overwrite=True, n_jobs=3, verbose=False)\n\nprint 'Done.'",
"_____no_output_____"
]
],
[
[
"### Make Epochs",
"_____no_output_____"
]
],
[
[
"import os\nimport numpy as np\nfrom mne import compute_covariance, Epochs, EpochsArray, find_events, read_proj, pick_types, set_log_level\nfrom mne.io import Raw\nfrom pandas import read_csv\nset_log_level(verbose=False)\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Define parameters.\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n\n## Subject level parameters.\nsubjects = ['BRTU', 'CHDR', 'CRDA', 'JADE', 'JASE', 'M5', 'MEWA', 'S2']\ntask = 'msit'\n\n## Filtering parameters.\nl_freq = 0.5\nh_freq = 50\nl_trans_bandwidth = l_freq / 2.\nh_trans_bandwidth = 1.0\nfilter_length = '20s'\nn_jobs = 3\n\n## Epoching parameters.\nevent_id = dict( FN=1, FI=2, NN=3, NI=4 ) # Alik's convention, isn't he smart!?\ntmin = -1.5 # Leave some breathing room.\ntmax = 3.4 # Trial is 1900ms, leave 1500ms of room.\nresp_buffer = 1.5 # 1500ms on either side of response.\nbaseline = (-0.5,-0.1)\nreject_tmin = -0.5\nreject_tmax = 1.9\nreject = dict(eeg=150e-6)\nflat = dict(eeg=5e-7)\ndetrend = None\ndecim = 1\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Load behavior.\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\nroot_dir = '/space/sophia/2/users/EMOTE-DBS/afMSIT'\ndata_file = os.path.join( root_dir, 'behavior', 'afMSIT_group_data.csv' )\ndf = read_csv(data_file)\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Load behavior.\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n\nfor subj in subjects:\n \n print 'Loading data for %s.' %subj\n\n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n ### Load data.\n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n\n # Define paths.\n raw_file = os.path.join( root_dir, 'raw', '%s_%s_raw.fif' %(subj,task) )\n proj_file = os.path.join( root_dir, 'raw', '%s_%s-proj.fif' %(subj,task) )\n \n # Load data.\n raw = Raw(raw_file,preload=True,verbose=False)\n proj = read_proj(proj_file)\n \n ## Add projections.\n proj = [p for p in proj if 'ref' not in p['desc']]\n raw.add_proj(proj, remove_existing=True)\n raw.add_eeg_average_proj()\n raw.apply_proj()\n print raw.info['projs']\n \n ## Reduce dataframe to subject.\n data = df[df.Subject==subj]\n \n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n ### Make events.\n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n print 'Identifying events for %s.' %subj, \n events = find_events(raw, stim_channel='Trig1', output='onset', min_duration=0.25, verbose=False)\n\n # Error catching.\n if data.shape[0] != events.shape[0]: raise ValueError('Mismatching number of stimulus onsets!')\n print '%s events found.' %events.shape[0]\n \n # Update event identifiers.\n n = 1\n for dbs in [0,1]:\n for cond in [0,1]:\n ix, = np.where((data.DBS==dbs)&(data.Interference==cond))\n events[ix,-1] = n\n n+=1\n\n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n ### Filter\n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n print 'Applying bandpass filter to raw [%s, %s].' %(l_freq, h_freq)\n \n Fs = raw.info['sfreq']\n raw.filter(l_freq = l_freq, h_freq = h_freq, filter_length=filter_length, n_jobs=n_jobs,\n l_trans_bandwidth=l_trans_bandwidth, h_trans_bandwidth=h_trans_bandwidth)\n \n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n ### Make stimulus-locked epochs.\n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n \n # Build initial epochs object.\n picks = pick_types(raw.info, meg=False, eeg=True, exclude='bads')\n epochs = Epochs(raw, events, event_id=event_id, tmin=tmin, tmax=tmax, baseline=baseline, picks=picks,\n reject=reject, flat=flat, reject_tmin=reject_tmin, reject_tmax=reject_tmax, \n proj=True, detrend=detrend, decim=decim)\n \n # First round of rejections.\n epochs.drop_bad() # Remove bad epochs.\n copy = data.ix[[True if not log else False for log in epochs.drop_log]] # Update CSV based on rejections.\n \n '''NOTE: Making a new dataframe copy is just a shortcut for easy indexing between the Pandas \n DataFrame and the Epochs object. This is due to the three rounds of rejections being \n applied to the data (e.g. amplitude, behavior exclusion, equalization).'''\n \n # Drop epochs based on behavior.\n epochs.drop(copy.Exclude.astype(bool))\n \n data = data.ix[[True if not log else False for log in epochs.drop_log]]\n print '%s trials remain after rejections.' %(len(epochs))\n print epochs\n\n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n ### Make Response-locked epochs.\n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n print 'Making response-locked epochs.'\n\n # Build response-locked events.\n response_indices = raw.time_as_index(0.4 + data.RT) # Compensating for MSIT-lock.\n response_events = epochs.events.copy()\n response_events[:,0] = response_events[:,0] + response_indices\n\n # Get data.\n arr = epochs.get_data()\n times = epochs.times\n\n # Calculate lengths of response-locked epochs.\n response_times = data.RT + 0.4 # Compensating for MSIT-lock.\n response_windows = np.array([response_times-resp_buffer, response_times+resp_buffer]).T\n\n # Iteratively build epochs array.\n trials = []\n for n in xrange(len(epochs)):\n mask = (times >= response_windows[n,0]) & (times <= response_windows[n,1])\n trials.append( arr[n,:,mask] )\n trials = np.array(trials).swapaxes(1,2)\n\n # Finally, make epochs objects.\n resp_epochs = EpochsArray(trials, epochs.info, response_events, tmin=-resp_buffer, event_id=event_id,)\n \n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n ### Save data.\n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n print 'Saving epoch files.'\n epochs.save(os.path.join(root_dir,'ave','%s_%s_%s_stim-epo.fif' %(subj,task,h_freq)))\n resp_epochs.save(os.path.join(root_dir,'ave','%s_%s_%s_resp-epo.fif' %(subj,task,h_freq)))\n data.to_csv(os.path.join(root_dir,'ave','%s_%s_%s-epo.csv' %(subj,task,h_freq)), index=False)\n \n print '\\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\\n'\n \nprint 'Done.'",
"_____no_output_____"
]
],
[
[
"### Make Covariance Matrices / Inverse Solutions / Morph Maps",
"_____no_output_____"
]
],
[
[
"import os\nfrom mne import EpochsArray, read_epochs, read_forward_solution, set_log_level\nfrom mne import compute_covariance, write_cov\nfrom mne import compute_morph_matrix, read_source_spaces\nfrom mne.filter import low_pass_filter\nfrom mne.minimum_norm import make_inverse_operator, write_inverse_operator\nfrom scipy.io import savemat\nset_log_level(verbose=False)\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Define parameters.\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n\n## Subject level parameters.\nsubjects = ['BRTU', 'CHDR', 'CRDA', 'JADE', 'JASE', 'M5', 'MEWA', 'S2']\ntask = 'msit'\n\n## Analysis parameters.\nfmax = 50\n\n## Source localization parameters.\nloose = 0.2\ndepth = 0.8\n\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n### Iteratively load and prepare data.\n#~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\nroot_dir = '/autofs/space/sophia_002/users/EMOTE-DBS/afMSIT'\nfs_dir = '/autofs/space/sophia_002/users/EMOTE-DBS/freesurfs'\nsrc = read_source_spaces(os.path.join(fs_dir,'fscopy','bem','fscopy-oct-6p-src.fif'))\n\nfor subject in subjects:\n \n print 'Processing %s' %subject\n \n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n ### Load files.\n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n \n ## Load in files.\n epo_file = os.path.join(root_dir,'ave','%s_msit_%s_stim-epo.fif' %(subject,fmax))\n epochs = read_epochs(epo_file, verbose=False)\n \n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n ### Secondary objects.\n #~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~#\n \n fwd = read_forward_solution(os.path.join(root_dir, 'fwd', '%s_%s-fwd.fif' %(subject,task)), \n surf_ori=True, verbose=False)\n \n ## Compute/save noise covariance matrix & inverse operator.\n noise_cov = compute_covariance(epochs, tmin=-0.5, tmax=0.0, method='shrunk', n_jobs=1)\n write_cov(os.path.join(root_dir,'cov','%s_%s_%s-cov.fif' %(subject,task,h_freq)), noise_cov)\n inv = make_inverse_operator(epochs.info, fwd, noise_cov, loose=loose, depth=depth, verbose=False)\n write_inverse_operator(os.path.join(root_dir,'cov','%s_%s_%s-inv.fif' %(subject,task,fmax)), inv)\n\n ## Pre-compute morph matrix.\n vertices_from = [inv['src'][n]['vertno'] for n in xrange(2)]\n vertices_to = [src[n]['vertno'] for n in xrange(2)]\n morph_mat = compute_morph_matrix(subject, 'fsaverage', vertices_from=vertices_from, \n vertices_to=vertices_to,subjects_dir=fs_dir, smooth=25)\n savemat(os.path.join(root_dir, 'morph_maps', '%s-fsaverage_morph.mat' %subject),\n mdict=dict(morph_mat=morph_mat))\n \nprint 'Done.'",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ab817adbccdb00f1e307e5333f56814a42f3173
| 66,270 |
ipynb
|
Jupyter Notebook
|
site/zh-cn/tutorials/text/transformer.ipynb
|
gmb-ftcont/docs-l10n
|
8b24263ca37dbf5cb4b0c15070a3d32c7284729d
|
[
"Apache-2.0"
] | 1 |
2020-08-05T05:52:57.000Z
|
2020-08-05T05:52:57.000Z
|
site/zh-cn/tutorials/text/transformer.ipynb
|
gmb-ftcont/docs-l10n
|
8b24263ca37dbf5cb4b0c15070a3d32c7284729d
|
[
"Apache-2.0"
] | null | null | null |
site/zh-cn/tutorials/text/transformer.ipynb
|
gmb-ftcont/docs-l10n
|
8b24263ca37dbf5cb4b0c15070a3d32c7284729d
|
[
"Apache-2.0"
] | null | null | null | 32.453477 | 229 | 0.489196 |
[
[
[
"##### Copyright 2019 The TensorFlow Authors.",
"_____no_output_____"
]
],
[
[
"#@title Licensed under the Apache License, Version 2.0 (the \"License\");\n# you may not use this file except in compliance with the License.\n# You may obtain a copy of the License at\n#\n# https://www.apache.org/licenses/LICENSE-2.0\n#\n# Unless required by applicable law or agreed to in writing, software\n# distributed under the License is distributed on an \"AS IS\" BASIS,\n# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.\n# See the License for the specific language governing permissions and\n# limitations under the License.",
"_____no_output_____"
]
],
[
[
"# 理解语言的 Transformer 模型",
"_____no_output_____"
],
[
"<table class=\"tfo-notebook-buttons\" align=\"left\">\n <td>\n <a target=\"_blank\" href=\"https://tensorflow.google.cn/tutorials/text/transformer\">\n <img src=\"https://tensorflow.google.cn/images/tf_logo_32px.png\" />\n 在 tensorflow.google.cn 上查看</a>\n </td>\n <td>\n <a target=\"_blank\" href=\"https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/text/transformer.ipynb\">\n <img src=\"https://tensorflow.google.cn/images/colab_logo_32px.png\" />\n 在 Google Colab 运行</a>\n </td>\n <td>\n <a target=\"_blank\" href=\"https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/text/transformer.ipynb\">\n <img src=\"https://tensorflow.google.cn/images/GitHub-Mark-32px.png\" />\n 在 Github 上查看源代码</a>\n </td>\n <td>\n <a href=\"https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/zh-cn/tutorials/text/transformer.ipynb\"><img src=\"https://tensorflow.google.cn/images/download_logo_32px.png\" />下载此 notebook</a>\n </td>\n</table>",
"_____no_output_____"
],
[
"Note: 我们的 TensorFlow 社区翻译了这些文档。因为社区翻译是尽力而为, 所以无法保证它们是最准确的,并且反映了最新的\n[官方英文文档](https://tensorflow.google.cn/?hl=en)。如果您有改进此翻译的建议, 请提交 pull request 到\n[tensorflow/docs](https://github.com/tensorflow/docs) GitHub 仓库。要志愿地撰写或者审核译文,请加入\n[[email protected] Google Group](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-zh-cn)",
"_____no_output_____"
],
[
"本教程训练了一个 <a href=\"https://arxiv.org/abs/1706.03762\" class=\"external\">Transformer 模型</a> 用于将葡萄牙语翻译成英语。这是一个高级示例,假定您具备[文本生成(text generation)](text_generation.ipynb)和 [注意力机制(attention)](nmt_with_attention.ipynb) 的知识。\n\nTransformer 模型的核心思想是*自注意力机制(self-attention)*——能注意输入序列的不同位置以计算该序列的表示的能力。Transformer 创建了多层自注意力层(self-attetion layers)组成的堆栈,下文的*按比缩放的点积注意力(Scaled dot product attention)*和*多头注意力(Multi-head attention)*部分对此进行了说明。\n\n一个 transformer 模型用自注意力层而非 [RNNs](text_classification_rnn.ipynb) 或 [CNNs](../images/intro_to_cnns.ipynb) 来处理变长的输入。这种通用架构有一系列的优势:\n\n* 它不对数据间的时间/空间关系做任何假设。这是处理一组对象(objects)的理想选择(例如,[星际争霸单位(StarCraft units)](https://deepmind.com/blog/alphastar-mastering-real-time-strategy-game-starcraft-ii/#block-8))。\n* 层输出可以并行计算,而非像 RNN 这样的序列计算。\n* 远距离项可以影响彼此的输出,而无需经过许多 RNN 步骤或卷积层(例如,参见[场景记忆 Transformer(Scene Memory Transformer)](https://arxiv.org/pdf/1903.03878.pdf))\n* 它能学习长距离的依赖。在许多序列任务中,这是一项挑战。\n\n该架构的缺点是:\n\n* 对于时间序列,一个单位时间的输出是从*整个历史记录*计算的,而非仅从输入和当前的隐含状态计算得到。这*可能*效率较低。 \n* 如果输入*确实*有时间/空间的关系,像文本,则必须加入一些位置编码,否则模型将有效地看到一堆单词。\n\n在此 notebook 中训练完模型后,您将能输入葡萄牙语句子,得到其英文翻译。\n\n<img src=\"https://tensorflow.google.cn/images/tutorials/transformer/attention_map_portuguese.png\" width=\"800\" alt=\"Attention heatmap\">",
"_____no_output_____"
]
],
[
[
"import tensorflow_datasets as tfds\nimport tensorflow as tf\n\nimport time\nimport numpy as np\nimport matplotlib.pyplot as plt",
"_____no_output_____"
]
],
[
[
"## 设置输入流水线(input pipeline)",
"_____no_output_____"
],
[
"使用 [TFDS](https://tensorflow.google.cn/datasets) 来导入 [葡萄牙语-英语翻译数据集](https://github.com/neulab/word-embeddings-for-nmt),该数据集来自于 [TED 演讲开放翻译项目](https://www.ted.com/participate/translate).\n\n该数据集包含来约 50000 条训练样本,1100 条验证样本,以及 2000 条测试样本。",
"_____no_output_____"
]
],
[
[
"examples, metadata = tfds.load('ted_hrlr_translate/pt_to_en', with_info=True,\n as_supervised=True)\ntrain_examples, val_examples = examples['train'], examples['validation']",
"_____no_output_____"
]
],
[
[
"从训练数据集创建自定义子词分词器(subwords tokenizer)。",
"_____no_output_____"
]
],
[
[
"tokenizer_en = tfds.features.text.SubwordTextEncoder.build_from_corpus(\n (en.numpy() for pt, en in train_examples), target_vocab_size=2**13)\n\ntokenizer_pt = tfds.features.text.SubwordTextEncoder.build_from_corpus(\n (pt.numpy() for pt, en in train_examples), target_vocab_size=2**13)",
"_____no_output_____"
],
[
"sample_string = 'Transformer is awesome.'\n\ntokenized_string = tokenizer_en.encode(sample_string)\nprint ('Tokenized string is {}'.format(tokenized_string))\n\noriginal_string = tokenizer_en.decode(tokenized_string)\nprint ('The original string: {}'.format(original_string))\n\nassert original_string == sample_string",
"_____no_output_____"
]
],
[
[
"如果单词不在词典中,则分词器(tokenizer)通过将单词分解为子词来对字符串进行编码。",
"_____no_output_____"
]
],
[
[
"for ts in tokenized_string:\n print ('{} ----> {}'.format(ts, tokenizer_en.decode([ts])))",
"_____no_output_____"
],
[
"BUFFER_SIZE = 20000\nBATCH_SIZE = 64",
"_____no_output_____"
]
],
[
[
"将开始和结束标记(token)添加到输入和目标。",
"_____no_output_____"
]
],
[
[
"def encode(lang1, lang2):\n lang1 = [tokenizer_pt.vocab_size] + tokenizer_pt.encode(\n lang1.numpy()) + [tokenizer_pt.vocab_size+1]\n\n lang2 = [tokenizer_en.vocab_size] + tokenizer_en.encode(\n lang2.numpy()) + [tokenizer_en.vocab_size+1]\n \n return lang1, lang2",
"_____no_output_____"
]
],
[
[
"Note:为了使本示例较小且相对较快,删除长度大于40个标记的样本。",
"_____no_output_____"
]
],
[
[
"MAX_LENGTH = 40",
"_____no_output_____"
],
[
"def filter_max_length(x, y, max_length=MAX_LENGTH):\n return tf.logical_and(tf.size(x) <= max_length,\n tf.size(y) <= max_length)",
"_____no_output_____"
]
],
[
[
"`.map()` 内部的操作以图模式(graph mode)运行,`.map()` 接收一个不具有 numpy 属性的图张量(graph tensor)。该`分词器(tokenizer)`需要将一个字符串或 Unicode 符号,编码成整数。因此,您需要在 `tf.py_function` 内部运行编码过程,`tf.py_function` 接收一个 eager 张量,该 eager 张量有一个包含字符串值的 numpy 属性。",
"_____no_output_____"
]
],
[
[
"def tf_encode(pt, en):\n result_pt, result_en = tf.py_function(encode, [pt, en], [tf.int64, tf.int64])\n result_pt.set_shape([None])\n result_en.set_shape([None])\n\n return result_pt, result_en",
"_____no_output_____"
],
[
"train_dataset = train_examples.map(tf_encode)\ntrain_dataset = train_dataset.filter(filter_max_length)\n# 将数据集缓存到内存中以加快读取速度。\ntrain_dataset = train_dataset.cache()\ntrain_dataset = train_dataset.shuffle(BUFFER_SIZE).padded_batch(BATCH_SIZE)\ntrain_dataset = train_dataset.prefetch(tf.data.experimental.AUTOTUNE)\n\n\nval_dataset = val_examples.map(tf_encode)\nval_dataset = val_dataset.filter(filter_max_length).padded_batch(BATCH_SIZE)",
"_____no_output_____"
],
[
"pt_batch, en_batch = next(iter(val_dataset))\npt_batch, en_batch",
"_____no_output_____"
]
],
[
[
"## 位置编码(Positional encoding)\n\n因为该模型并不包括任何的循环(recurrence)或卷积,所以模型添加了位置编码,为模型提供一些关于单词在句子中相对位置的信息。\n\n位置编码向量被加到嵌入(embedding)向量中。嵌入表示一个 d 维空间的标记,在 d 维空间中有着相似含义的标记会离彼此更近。但是,嵌入并没有对在一句话中的词的相对位置进行编码。因此,当加上位置编码后,词将基于*它们含义的相似度以及它们在句子中的位置*,在 d 维空间中离彼此更近。\n\n参看 [位置编码](https://github.com/tensorflow/examples/blob/master/community/en/position_encoding.ipynb) 的 notebook 了解更多信息。计算位置编码的公式如下:\n\n$$\\Large{PE_{(pos, 2i)} = sin(pos / 10000^{2i / d_{model}})} $$\n$$\\Large{PE_{(pos, 2i+1)} = cos(pos / 10000^{2i / d_{model}})} $$",
"_____no_output_____"
]
],
[
[
"def get_angles(pos, i, d_model):\n angle_rates = 1 / np.power(10000, (2 * (i//2)) / np.float32(d_model))\n return pos * angle_rates",
"_____no_output_____"
],
[
"def positional_encoding(position, d_model):\n angle_rads = get_angles(np.arange(position)[:, np.newaxis],\n np.arange(d_model)[np.newaxis, :],\n d_model)\n \n # 将 sin 应用于数组中的偶数索引(indices);2i\n angle_rads[:, 0::2] = np.sin(angle_rads[:, 0::2])\n \n # 将 cos 应用于数组中的奇数索引;2i+1\n angle_rads[:, 1::2] = np.cos(angle_rads[:, 1::2])\n \n pos_encoding = angle_rads[np.newaxis, ...]\n \n return tf.cast(pos_encoding, dtype=tf.float32)",
"_____no_output_____"
],
[
"pos_encoding = positional_encoding(50, 512)\nprint (pos_encoding.shape)\n\nplt.pcolormesh(pos_encoding[0], cmap='RdBu')\nplt.xlabel('Depth')\nplt.xlim((0, 512))\nplt.ylabel('Position')\nplt.colorbar()\nplt.show()",
"_____no_output_____"
]
],
[
[
"## 遮挡(Masking)",
"_____no_output_____"
],
[
"遮挡一批序列中所有的填充标记(pad tokens)。这确保了模型不会将填充作为输入。该 mask 表明填充值 `0` 出现的位置:在这些位置 mask 输出 `1`,否则输出 `0`。",
"_____no_output_____"
]
],
[
[
"def create_padding_mask(seq):\n seq = tf.cast(tf.math.equal(seq, 0), tf.float32)\n \n # 添加额外的维度来将填充加到\n # 注意力对数(logits)。\n return seq[:, tf.newaxis, tf.newaxis, :] # (batch_size, 1, 1, seq_len)",
"_____no_output_____"
],
[
"x = tf.constant([[7, 6, 0, 0, 1], [1, 2, 3, 0, 0], [0, 0, 0, 4, 5]])\ncreate_padding_mask(x)",
"_____no_output_____"
]
],
[
[
"前瞻遮挡(look-ahead mask)用于遮挡一个序列中的后续标记(future tokens)。换句话说,该 mask 表明了不应该使用的条目。\n\n这意味着要预测第三个词,将仅使用第一个和第二个词。与此类似,预测第四个词,仅使用第一个,第二个和第三个词,依此类推。 ",
"_____no_output_____"
]
],
[
[
"def create_look_ahead_mask(size):\n mask = 1 - tf.linalg.band_part(tf.ones((size, size)), -1, 0)\n return mask # (seq_len, seq_len)",
"_____no_output_____"
],
[
"x = tf.random.uniform((1, 3))\ntemp = create_look_ahead_mask(x.shape[1])\ntemp",
"_____no_output_____"
]
],
[
[
"## 按比缩放的点积注意力(Scaled dot product attention)",
"_____no_output_____"
],
[
"<img src=\"https://tensorflow.google.cn/images/tutorials/transformer/scaled_attention.png\" width=\"500\" alt=\"scaled_dot_product_attention\">\n\nTransformer 使用的注意力函数有三个输入:Q(请求(query))、K(主键(key))、V(数值(value))。用于计算注意力权重的等式为:\n\n$$\\Large{Attention(Q, K, V) = softmax_k(\\frac{QK^T}{\\sqrt{d_k}}) V} $$\n\n点积注意力被缩小了深度的平方根倍。这样做是因为对于较大的深度值,点积的大小会增大,从而推动 softmax 函数往仅有很小的梯度的方向靠拢,导致了一种很硬的(hard)softmax。\n\n例如,假设 `Q` 和 `K` 的均值为0,方差为1。它们的矩阵乘积将有均值为0,方差为 `dk`。因此,*`dk` 的平方根*被用于缩放(而非其他数值),因为,`Q` 和 `K` 的矩阵乘积的均值本应该为 0,方差本应该为1,这样会获得一个更平缓的 softmax。\n\n遮挡(mask)与 -1e9(接近于负无穷)相乘。这样做是因为遮挡与缩放的 Q 和 K 的矩阵乘积相加,并在 softmax 之前立即应用。目标是将这些单元归零,因为 softmax 的较大负数输入在输出中接近于零。",
"_____no_output_____"
]
],
[
[
"def scaled_dot_product_attention(q, k, v, mask):\n \"\"\"计算注意力权重。\n q, k, v 必须具有匹配的前置维度。\n k, v 必须有匹配的倒数第二个维度,例如:seq_len_k = seq_len_v。\n 虽然 mask 根据其类型(填充或前瞻)有不同的形状,\n 但是 mask 必须能进行广播转换以便求和。\n \n 参数:\n q: 请求的形状 == (..., seq_len_q, depth)\n k: 主键的形状 == (..., seq_len_k, depth)\n v: 数值的形状 == (..., seq_len_v, depth_v)\n mask: Float 张量,其形状能转换成\n (..., seq_len_q, seq_len_k)。默认为None。\n \n 返回值:\n 输出,注意力权重\n \"\"\"\n\n matmul_qk = tf.matmul(q, k, transpose_b=True) # (..., seq_len_q, seq_len_k)\n \n # 缩放 matmul_qk\n dk = tf.cast(tf.shape(k)[-1], tf.float32)\n scaled_attention_logits = matmul_qk / tf.math.sqrt(dk)\n\n # 将 mask 加入到缩放的张量上。\n if mask is not None:\n scaled_attention_logits += (mask * -1e9) \n\n # softmax 在最后一个轴(seq_len_k)上归一化,因此分数\n # 相加等于1。\n attention_weights = tf.nn.softmax(scaled_attention_logits, axis=-1) # (..., seq_len_q, seq_len_k)\n\n output = tf.matmul(attention_weights, v) # (..., seq_len_q, depth_v)\n\n return output, attention_weights",
"_____no_output_____"
]
],
[
[
"当 softmax 在 K 上进行归一化后,它的值决定了分配到 Q 的重要程度。\n\n输出表示注意力权重和 V(数值)向量的乘积。这确保了要关注的词保持原样,而无关的词将被清除掉。",
"_____no_output_____"
]
],
[
[
"def print_out(q, k, v):\n temp_out, temp_attn = scaled_dot_product_attention(\n q, k, v, None)\n print ('Attention weights are:')\n print (temp_attn)\n print ('Output is:')\n print (temp_out)",
"_____no_output_____"
],
[
"np.set_printoptions(suppress=True)\n\ntemp_k = tf.constant([[10,0,0],\n [0,10,0],\n [0,0,10],\n [0,0,10]], dtype=tf.float32) # (4, 3)\n\ntemp_v = tf.constant([[ 1,0],\n [ 10,0],\n [ 100,5],\n [1000,6]], dtype=tf.float32) # (4, 2)\n\n# 这条 `请求(query)符合第二个`主键(key)`,\n# 因此返回了第二个`数值(value)`。\ntemp_q = tf.constant([[0, 10, 0]], dtype=tf.float32) # (1, 3)\nprint_out(temp_q, temp_k, temp_v)",
"_____no_output_____"
],
[
"# 这条请求符合重复出现的主键(第三第四个),\n# 因此,对所有的相关数值取了平均。\ntemp_q = tf.constant([[0, 0, 10]], dtype=tf.float32) # (1, 3)\nprint_out(temp_q, temp_k, temp_v)",
"_____no_output_____"
],
[
"# 这条请求符合第一和第二条主键,\n# 因此,对它们的数值去了平均。\ntemp_q = tf.constant([[10, 10, 0]], dtype=tf.float32) # (1, 3)\nprint_out(temp_q, temp_k, temp_v)",
"_____no_output_____"
]
],
[
[
"将所有请求一起*传递*。",
"_____no_output_____"
]
],
[
[
"temp_q = tf.constant([[0, 0, 10], [0, 10, 0], [10, 10, 0]], dtype=tf.float32) # (3, 3)\nprint_out(temp_q, temp_k, temp_v)",
"_____no_output_____"
]
],
[
[
"## 多头注意力(Multi-head attention)",
"_____no_output_____"
],
[
"<img src=\"https://tensorflow.google.cn/images/tutorials/transformer/multi_head_attention.png\" width=\"500\" alt=\"multi-head attention\">\n\n\n多头注意力由四部分组成:\n* 线性层并分拆成多头。\n* 按比缩放的点积注意力。\n* 多头及联。\n* 最后一层线性层。",
"_____no_output_____"
],
[
"每个多头注意力块有三个输入:Q(请求)、K(主键)、V(数值)。这些输入经过线性(Dense)层,并分拆成多头。 \n\n将上面定义的 `scaled_dot_product_attention` 函数应用于每个头(进行了广播(broadcasted)以提高效率)。注意力这步必须使用一个恰当的 mask。然后将每个头的注意力输出连接起来(用`tf.transpose` 和 `tf.reshape`),并放入最后的 `Dense` 层。\n\nQ、K、和 V 被拆分到了多个头,而非单个的注意力头,因为多头允许模型共同注意来自不同表示空间的不同位置的信息。在分拆后,每个头部的维度减少,因此总的计算成本与有着全部维度的单个注意力头相同。",
"_____no_output_____"
]
],
[
[
"class MultiHeadAttention(tf.keras.layers.Layer):\n def __init__(self, d_model, num_heads):\n super(MultiHeadAttention, self).__init__()\n self.num_heads = num_heads\n self.d_model = d_model\n \n assert d_model % self.num_heads == 0\n \n self.depth = d_model // self.num_heads\n \n self.wq = tf.keras.layers.Dense(d_model)\n self.wk = tf.keras.layers.Dense(d_model)\n self.wv = tf.keras.layers.Dense(d_model)\n \n self.dense = tf.keras.layers.Dense(d_model)\n \n def split_heads(self, x, batch_size):\n \"\"\"分拆最后一个维度到 (num_heads, depth).\n 转置结果使得形状为 (batch_size, num_heads, seq_len, depth)\n \"\"\"\n x = tf.reshape(x, (batch_size, -1, self.num_heads, self.depth))\n return tf.transpose(x, perm=[0, 2, 1, 3])\n \n def call(self, v, k, q, mask):\n batch_size = tf.shape(q)[0]\n \n q = self.wq(q) # (batch_size, seq_len, d_model)\n k = self.wk(k) # (batch_size, seq_len, d_model)\n v = self.wv(v) # (batch_size, seq_len, d_model)\n \n q = self.split_heads(q, batch_size) # (batch_size, num_heads, seq_len_q, depth)\n k = self.split_heads(k, batch_size) # (batch_size, num_heads, seq_len_k, depth)\n v = self.split_heads(v, batch_size) # (batch_size, num_heads, seq_len_v, depth)\n \n # scaled_attention.shape == (batch_size, num_heads, seq_len_q, depth)\n # attention_weights.shape == (batch_size, num_heads, seq_len_q, seq_len_k)\n scaled_attention, attention_weights = scaled_dot_product_attention(\n q, k, v, mask)\n \n scaled_attention = tf.transpose(scaled_attention, perm=[0, 2, 1, 3]) # (batch_size, seq_len_q, num_heads, depth)\n\n concat_attention = tf.reshape(scaled_attention, \n (batch_size, -1, self.d_model)) # (batch_size, seq_len_q, d_model)\n\n output = self.dense(concat_attention) # (batch_size, seq_len_q, d_model)\n \n return output, attention_weights",
"_____no_output_____"
]
],
[
[
"创建一个 `MultiHeadAttention` 层进行尝试。在序列中的每个位置 `y`,`MultiHeadAttention` 在序列中的所有其他位置运行所有8个注意力头,在每个位置y,返回一个新的同样长度的向量。",
"_____no_output_____"
]
],
[
[
"temp_mha = MultiHeadAttention(d_model=512, num_heads=8)\ny = tf.random.uniform((1, 60, 512)) # (batch_size, encoder_sequence, d_model)\nout, attn = temp_mha(y, k=y, q=y, mask=None)\nout.shape, attn.shape",
"_____no_output_____"
]
],
[
[
"## 点式前馈网络(Point wise feed forward network)",
"_____no_output_____"
],
[
"点式前馈网络由两层全联接层组成,两层之间有一个 ReLU 激活函数。",
"_____no_output_____"
]
],
[
[
"def point_wise_feed_forward_network(d_model, dff):\n return tf.keras.Sequential([\n tf.keras.layers.Dense(dff, activation='relu'), # (batch_size, seq_len, dff)\n tf.keras.layers.Dense(d_model) # (batch_size, seq_len, d_model)\n ])",
"_____no_output_____"
],
[
"sample_ffn = point_wise_feed_forward_network(512, 2048)\nsample_ffn(tf.random.uniform((64, 50, 512))).shape",
"_____no_output_____"
]
],
[
[
"## 编码与解码(Encoder and decoder)",
"_____no_output_____"
],
[
"<img src=\"https://tensorflow.google.cn/images/tutorials/transformer/transformer.png\" width=\"600\" alt=\"transformer\">",
"_____no_output_____"
],
[
"Transformer 模型与标准的[具有注意力机制的序列到序列模型(sequence to sequence with attention model)](nmt_with_attention.ipynb),遵循相同的一般模式。\n\n* 输入语句经过 `N` 个编码器层,为序列中的每个词/标记生成一个输出。\n* 解码器关注编码器的输出以及它自身的输入(自注意力)来预测下一个词。",
"_____no_output_____"
],
[
"### 编码器层(Encoder layer)\n\n每个编码器层包括以下子层:\n\n1. 多头注意力(有填充遮挡)\n2. 点式前馈网络(Point wise feed forward networks)。\n\n每个子层在其周围有一个残差连接,然后进行层归一化。残差连接有助于避免深度网络中的梯度消失问题。\n\n每个子层的输出是 `LayerNorm(x + Sublayer(x))`。归一化是在 `d_model`(最后一个)维度完成的。Transformer 中有 N 个编码器层。",
"_____no_output_____"
]
],
[
[
"class EncoderLayer(tf.keras.layers.Layer):\n def __init__(self, d_model, num_heads, dff, rate=0.1):\n super(EncoderLayer, self).__init__()\n\n self.mha = MultiHeadAttention(d_model, num_heads)\n self.ffn = point_wise_feed_forward_network(d_model, dff)\n\n self.layernorm1 = tf.keras.layers.LayerNormalization(epsilon=1e-6)\n self.layernorm2 = tf.keras.layers.LayerNormalization(epsilon=1e-6)\n \n self.dropout1 = tf.keras.layers.Dropout(rate)\n self.dropout2 = tf.keras.layers.Dropout(rate)\n \n def call(self, x, training, mask):\n\n attn_output, _ = self.mha(x, x, x, mask) # (batch_size, input_seq_len, d_model)\n attn_output = self.dropout1(attn_output, training=training)\n out1 = self.layernorm1(x + attn_output) # (batch_size, input_seq_len, d_model)\n \n ffn_output = self.ffn(out1) # (batch_size, input_seq_len, d_model)\n ffn_output = self.dropout2(ffn_output, training=training)\n out2 = self.layernorm2(out1 + ffn_output) # (batch_size, input_seq_len, d_model)\n \n return out2",
"_____no_output_____"
],
[
"sample_encoder_layer = EncoderLayer(512, 8, 2048)\n\nsample_encoder_layer_output = sample_encoder_layer(\n tf.random.uniform((64, 43, 512)), False, None)\n\nsample_encoder_layer_output.shape # (batch_size, input_seq_len, d_model)",
"_____no_output_____"
]
],
[
[
"### 解码器层(Decoder layer)\n\n每个解码器层包括以下子层:\n\n1. 遮挡的多头注意力(前瞻遮挡和填充遮挡)\n2. 多头注意力(用填充遮挡)。V(数值)和 K(主键)接收*编码器输出*作为输入。Q(请求)接收*遮挡的多头注意力子层的输出*。\n3. 点式前馈网络\n\n每个子层在其周围有一个残差连接,然后进行层归一化。每个子层的输出是 `LayerNorm(x + Sublayer(x))`。归一化是在 `d_model`(最后一个)维度完成的。\n\nTransformer 中共有 N 个解码器层。\n\n当 Q 接收到解码器的第一个注意力块的输出,并且 K 接收到编码器的输出时,注意力权重表示根据编码器的输出赋予解码器输入的重要性。换一种说法,解码器通过查看编码器输出和对其自身输出的自注意力,预测下一个词。参看按比缩放的点积注意力部分的演示。",
"_____no_output_____"
]
],
[
[
"class DecoderLayer(tf.keras.layers.Layer):\n def __init__(self, d_model, num_heads, dff, rate=0.1):\n super(DecoderLayer, self).__init__()\n\n self.mha1 = MultiHeadAttention(d_model, num_heads)\n self.mha2 = MultiHeadAttention(d_model, num_heads)\n\n self.ffn = point_wise_feed_forward_network(d_model, dff)\n \n self.layernorm1 = tf.keras.layers.LayerNormalization(epsilon=1e-6)\n self.layernorm2 = tf.keras.layers.LayerNormalization(epsilon=1e-6)\n self.layernorm3 = tf.keras.layers.LayerNormalization(epsilon=1e-6)\n \n self.dropout1 = tf.keras.layers.Dropout(rate)\n self.dropout2 = tf.keras.layers.Dropout(rate)\n self.dropout3 = tf.keras.layers.Dropout(rate)\n \n \n def call(self, x, enc_output, training, \n look_ahead_mask, padding_mask):\n # enc_output.shape == (batch_size, input_seq_len, d_model)\n\n attn1, attn_weights_block1 = self.mha1(x, x, x, look_ahead_mask) # (batch_size, target_seq_len, d_model)\n attn1 = self.dropout1(attn1, training=training)\n out1 = self.layernorm1(attn1 + x)\n \n attn2, attn_weights_block2 = self.mha2(\n enc_output, enc_output, out1, padding_mask) # (batch_size, target_seq_len, d_model)\n attn2 = self.dropout2(attn2, training=training)\n out2 = self.layernorm2(attn2 + out1) # (batch_size, target_seq_len, d_model)\n \n ffn_output = self.ffn(out2) # (batch_size, target_seq_len, d_model)\n ffn_output = self.dropout3(ffn_output, training=training)\n out3 = self.layernorm3(ffn_output + out2) # (batch_size, target_seq_len, d_model)\n \n return out3, attn_weights_block1, attn_weights_block2",
"_____no_output_____"
],
[
"sample_decoder_layer = DecoderLayer(512, 8, 2048)\n\nsample_decoder_layer_output, _, _ = sample_decoder_layer(\n tf.random.uniform((64, 50, 512)), sample_encoder_layer_output, \n False, None, None)\n\nsample_decoder_layer_output.shape # (batch_size, target_seq_len, d_model)",
"_____no_output_____"
]
],
[
[
"### 编码器(Encoder)\n\n`编码器` 包括:\n1. 输入嵌入(Input Embedding)\n2. 位置编码(Positional Encoding)\n3. N 个编码器层(encoder layers)\n\n输入经过嵌入(embedding)后,该嵌入与位置编码相加。该加法结果的输出是编码器层的输入。编码器的输出是解码器的输入。",
"_____no_output_____"
]
],
[
[
"class Encoder(tf.keras.layers.Layer):\n def __init__(self, num_layers, d_model, num_heads, dff, input_vocab_size,\n maximum_position_encoding, rate=0.1):\n super(Encoder, self).__init__()\n\n self.d_model = d_model\n self.num_layers = num_layers\n \n self.embedding = tf.keras.layers.Embedding(input_vocab_size, d_model)\n self.pos_encoding = positional_encoding(maximum_position_encoding, \n self.d_model)\n \n \n self.enc_layers = [EncoderLayer(d_model, num_heads, dff, rate) \n for _ in range(num_layers)]\n \n self.dropout = tf.keras.layers.Dropout(rate)\n \n def call(self, x, training, mask):\n\n seq_len = tf.shape(x)[1]\n \n # 将嵌入和位置编码相加。\n x = self.embedding(x) # (batch_size, input_seq_len, d_model)\n x *= tf.math.sqrt(tf.cast(self.d_model, tf.float32))\n x += self.pos_encoding[:, :seq_len, :]\n\n x = self.dropout(x, training=training)\n \n for i in range(self.num_layers):\n x = self.enc_layers[i](x, training, mask)\n \n return x # (batch_size, input_seq_len, d_model)",
"_____no_output_____"
],
[
"sample_encoder = Encoder(num_layers=2, d_model=512, num_heads=8, \n dff=2048, input_vocab_size=8500,\n maximum_position_encoding=10000)\n\nsample_encoder_output = sample_encoder(tf.random.uniform((64, 62)), \n training=False, mask=None)\n\nprint (sample_encoder_output.shape) # (batch_size, input_seq_len, d_model)",
"_____no_output_____"
]
],
[
[
"### 解码器(Decoder)",
"_____no_output_____"
],
[
"`解码器`包括:\n1. 输出嵌入(Output Embedding)\n2. 位置编码(Positional Encoding)\n3. N 个解码器层(decoder layers)\n\n目标(target)经过一个嵌入后,该嵌入和位置编码相加。该加法结果是解码器层的输入。解码器的输出是最后的线性层的输入。",
"_____no_output_____"
]
],
[
[
"class Decoder(tf.keras.layers.Layer):\n def __init__(self, num_layers, d_model, num_heads, dff, target_vocab_size,\n maximum_position_encoding, rate=0.1):\n super(Decoder, self).__init__()\n\n self.d_model = d_model\n self.num_layers = num_layers\n \n self.embedding = tf.keras.layers.Embedding(target_vocab_size, d_model)\n self.pos_encoding = positional_encoding(maximum_position_encoding, d_model)\n \n self.dec_layers = [DecoderLayer(d_model, num_heads, dff, rate) \n for _ in range(num_layers)]\n self.dropout = tf.keras.layers.Dropout(rate)\n \n def call(self, x, enc_output, training, \n look_ahead_mask, padding_mask):\n\n seq_len = tf.shape(x)[1]\n attention_weights = {}\n \n x = self.embedding(x) # (batch_size, target_seq_len, d_model)\n x *= tf.math.sqrt(tf.cast(self.d_model, tf.float32))\n x += self.pos_encoding[:, :seq_len, :]\n \n x = self.dropout(x, training=training)\n\n for i in range(self.num_layers):\n x, block1, block2 = self.dec_layers[i](x, enc_output, training,\n look_ahead_mask, padding_mask)\n \n attention_weights['decoder_layer{}_block1'.format(i+1)] = block1\n attention_weights['decoder_layer{}_block2'.format(i+1)] = block2\n \n # x.shape == (batch_size, target_seq_len, d_model)\n return x, attention_weights",
"_____no_output_____"
],
[
"sample_decoder = Decoder(num_layers=2, d_model=512, num_heads=8, \n dff=2048, target_vocab_size=8000,\n maximum_position_encoding=5000)\n\noutput, attn = sample_decoder(tf.random.uniform((64, 26)), \n enc_output=sample_encoder_output, \n training=False, look_ahead_mask=None, \n padding_mask=None)\n\noutput.shape, attn['decoder_layer2_block2'].shape",
"_____no_output_____"
]
],
[
[
"## 创建 Transformer",
"_____no_output_____"
],
[
"Transformer 包括编码器,解码器和最后的线性层。解码器的输出是线性层的输入,返回线性层的输出。",
"_____no_output_____"
]
],
[
[
"class Transformer(tf.keras.Model):\n def __init__(self, num_layers, d_model, num_heads, dff, input_vocab_size, \n target_vocab_size, pe_input, pe_target, rate=0.1):\n super(Transformer, self).__init__()\n\n self.encoder = Encoder(num_layers, d_model, num_heads, dff, \n input_vocab_size, pe_input, rate)\n\n self.decoder = Decoder(num_layers, d_model, num_heads, dff, \n target_vocab_size, pe_target, rate)\n\n self.final_layer = tf.keras.layers.Dense(target_vocab_size)\n \n def call(self, inp, tar, training, enc_padding_mask, \n look_ahead_mask, dec_padding_mask):\n\n enc_output = self.encoder(inp, training, enc_padding_mask) # (batch_size, inp_seq_len, d_model)\n \n # dec_output.shape == (batch_size, tar_seq_len, d_model)\n dec_output, attention_weights = self.decoder(\n tar, enc_output, training, look_ahead_mask, dec_padding_mask)\n \n final_output = self.final_layer(dec_output) # (batch_size, tar_seq_len, target_vocab_size)\n \n return final_output, attention_weights",
"_____no_output_____"
],
[
"sample_transformer = Transformer(\n num_layers=2, d_model=512, num_heads=8, dff=2048, \n input_vocab_size=8500, target_vocab_size=8000, \n pe_input=10000, pe_target=6000)\n\ntemp_input = tf.random.uniform((64, 62))\ntemp_target = tf.random.uniform((64, 26))\n\nfn_out, _ = sample_transformer(temp_input, temp_target, training=False, \n enc_padding_mask=None, \n look_ahead_mask=None,\n dec_padding_mask=None)\n\nfn_out.shape # (batch_size, tar_seq_len, target_vocab_size)",
"_____no_output_____"
]
],
[
[
"## 配置超参数(hyperparameters)",
"_____no_output_____"
],
[
"为了让本示例小且相对较快,已经减小了*num_layers、 d_model 和 dff* 的值。 \n\nTransformer 的基础模型使用的数值为:*num_layers=6*,*d_model = 512*,*dff = 2048*。关于所有其他版本的 Transformer,请查阅[论文](https://arxiv.org/abs/1706.03762)。\n\nNote:通过改变以下数值,您可以获得在许多任务上达到最先进水平的模型。",
"_____no_output_____"
]
],
[
[
"num_layers = 4\nd_model = 128\ndff = 512\nnum_heads = 8\n\ninput_vocab_size = tokenizer_pt.vocab_size + 2\ntarget_vocab_size = tokenizer_en.vocab_size + 2\ndropout_rate = 0.1",
"_____no_output_____"
]
],
[
[
"## 优化器(Optimizer)",
"_____no_output_____"
],
[
"根据[论文](https://arxiv.org/abs/1706.03762)中的公式,将 Adam 优化器与自定义的学习速率调度程序(scheduler)配合使用。\n\n$$\\Large{lrate = d_{model}^{-0.5} * min(step{\\_}num^{-0.5}, step{\\_}num * warmup{\\_}steps^{-1.5})}$$\n",
"_____no_output_____"
]
],
[
[
"class CustomSchedule(tf.keras.optimizers.schedules.LearningRateSchedule):\n def __init__(self, d_model, warmup_steps=4000):\n super(CustomSchedule, self).__init__()\n \n self.d_model = d_model\n self.d_model = tf.cast(self.d_model, tf.float32)\n\n self.warmup_steps = warmup_steps\n \n def __call__(self, step):\n arg1 = tf.math.rsqrt(step)\n arg2 = step * (self.warmup_steps ** -1.5)\n \n return tf.math.rsqrt(self.d_model) * tf.math.minimum(arg1, arg2)",
"_____no_output_____"
],
[
"learning_rate = CustomSchedule(d_model)\n\noptimizer = tf.keras.optimizers.Adam(learning_rate, beta_1=0.9, beta_2=0.98, \n epsilon=1e-9)",
"_____no_output_____"
],
[
"temp_learning_rate_schedule = CustomSchedule(d_model)\n\nplt.plot(temp_learning_rate_schedule(tf.range(40000, dtype=tf.float32)))\nplt.ylabel(\"Learning Rate\")\nplt.xlabel(\"Train Step\")",
"_____no_output_____"
]
],
[
[
"## 损失函数与指标(Loss and metrics)",
"_____no_output_____"
],
[
"由于目标序列是填充(padded)过的,因此在计算损失函数时,应用填充遮挡非常重要。",
"_____no_output_____"
]
],
[
[
"loss_object = tf.keras.losses.SparseCategoricalCrossentropy(\n from_logits=True, reduction='none')",
"_____no_output_____"
],
[
"def loss_function(real, pred):\n mask = tf.math.logical_not(tf.math.equal(real, 0))\n loss_ = loss_object(real, pred)\n\n mask = tf.cast(mask, dtype=loss_.dtype)\n loss_ *= mask\n \n return tf.reduce_mean(loss_)",
"_____no_output_____"
],
[
"train_loss = tf.keras.metrics.Mean(name='train_loss')\ntrain_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(\n name='train_accuracy')",
"_____no_output_____"
]
],
[
[
"## 训练与检查点(Training and checkpointing)",
"_____no_output_____"
]
],
[
[
"transformer = Transformer(num_layers, d_model, num_heads, dff,\n input_vocab_size, target_vocab_size, \n pe_input=input_vocab_size, \n pe_target=target_vocab_size,\n rate=dropout_rate)",
"_____no_output_____"
],
[
"def create_masks(inp, tar):\n # 编码器填充遮挡\n enc_padding_mask = create_padding_mask(inp)\n \n # 在解码器的第二个注意力模块使用。\n # 该填充遮挡用于遮挡编码器的输出。\n dec_padding_mask = create_padding_mask(inp)\n \n # 在解码器的第一个注意力模块使用。\n # 用于填充(pad)和遮挡(mask)解码器获取到的输入的后续标记(future tokens)。\n look_ahead_mask = create_look_ahead_mask(tf.shape(tar)[1])\n dec_target_padding_mask = create_padding_mask(tar)\n combined_mask = tf.maximum(dec_target_padding_mask, look_ahead_mask)\n \n return enc_padding_mask, combined_mask, dec_padding_mask",
"_____no_output_____"
]
],
[
[
"创建检查点的路径和检查点管理器(manager)。这将用于在每 `n` 个周期(epochs)保存检查点。",
"_____no_output_____"
]
],
[
[
"checkpoint_path = \"./checkpoints/train\"\n\nckpt = tf.train.Checkpoint(transformer=transformer,\n optimizer=optimizer)\n\nckpt_manager = tf.train.CheckpointManager(ckpt, checkpoint_path, max_to_keep=5)\n\n# 如果检查点存在,则恢复最新的检查点。\nif ckpt_manager.latest_checkpoint:\n ckpt.restore(ckpt_manager.latest_checkpoint)\n print ('Latest checkpoint restored!!')",
"_____no_output_____"
]
],
[
[
"目标(target)被分成了 tar_inp 和 tar_real。tar_inp 作为输入传递到解码器。`tar_real` 是位移了 1 的同一个输入:在 `tar_inp` 中的每个位置,`tar_real` 包含了应该被预测到的下一个标记(token)。\n\n例如,`sentence` = \"SOS A lion in the jungle is sleeping EOS\"\n\n`tar_inp` = \"SOS A lion in the jungle is sleeping\"\n\n`tar_real` = \"A lion in the jungle is sleeping EOS\"\n\nTransformer 是一个自回归(auto-regressive)模型:它一次作一个部分的预测,然后使用到目前为止的自身的输出来决定下一步要做什么。\n\n在训练过程中,本示例使用了 teacher-forcing 的方法(就像[文本生成教程](./text_generation.ipynb)中一样)。无论模型在当前时间步骤下预测出什么,teacher-forcing 方法都会将真实的输出传递到下一个时间步骤上。\n\n当 transformer 预测每个词时,*自注意力(self-attention)*功能使它能够查看输入序列中前面的单词,从而更好地预测下一个单词。\n\n为了防止模型在期望的输出上达到峰值,模型使用了前瞻遮挡(look-ahead mask)。",
"_____no_output_____"
]
],
[
[
"EPOCHS = 20",
"_____no_output_____"
],
[
"# 该 @tf.function 将追踪-编译 train_step 到 TF 图中,以便更快地\n# 执行。该函数专用于参数张量的精确形状。为了避免由于可变序列长度或可变\n# 批次大小(最后一批次较小)导致的再追踪,使用 input_signature 指定\n# 更多的通用形状。\n\ntrain_step_signature = [\n tf.TensorSpec(shape=(None, None), dtype=tf.int64),\n tf.TensorSpec(shape=(None, None), dtype=tf.int64),\n]\n\[email protected](input_signature=train_step_signature)\ndef train_step(inp, tar):\n tar_inp = tar[:, :-1]\n tar_real = tar[:, 1:]\n \n enc_padding_mask, combined_mask, dec_padding_mask = create_masks(inp, tar_inp)\n \n with tf.GradientTape() as tape:\n predictions, _ = transformer(inp, tar_inp, \n True, \n enc_padding_mask, \n combined_mask, \n dec_padding_mask)\n loss = loss_function(tar_real, predictions)\n\n gradients = tape.gradient(loss, transformer.trainable_variables) \n optimizer.apply_gradients(zip(gradients, transformer.trainable_variables))\n \n train_loss(loss)\n train_accuracy(tar_real, predictions)",
"_____no_output_____"
]
],
[
[
"葡萄牙语作为输入语言,英语为目标语言。",
"_____no_output_____"
]
],
[
[
"for epoch in range(EPOCHS):\n start = time.time()\n \n train_loss.reset_states()\n train_accuracy.reset_states()\n \n # inp -> portuguese, tar -> english\n for (batch, (inp, tar)) in enumerate(train_dataset):\n train_step(inp, tar)\n \n if batch % 50 == 0:\n print ('Epoch {} Batch {} Loss {:.4f} Accuracy {:.4f}'.format(\n epoch + 1, batch, train_loss.result(), train_accuracy.result()))\n \n if (epoch + 1) % 5 == 0:\n ckpt_save_path = ckpt_manager.save()\n print ('Saving checkpoint for epoch {} at {}'.format(epoch+1,\n ckpt_save_path))\n \n print ('Epoch {} Loss {:.4f} Accuracy {:.4f}'.format(epoch + 1, \n train_loss.result(), \n train_accuracy.result()))\n\n print ('Time taken for 1 epoch: {} secs\\n'.format(time.time() - start))",
"_____no_output_____"
]
],
[
[
"## 评估(Evaluate)",
"_____no_output_____"
],
[
"以下步骤用于评估:\n\n* 用葡萄牙语分词器(`tokenizer_pt`)编码输入语句。此外,添加开始和结束标记,这样输入就与模型训练的内容相同。这是编码器输入。\n* 解码器输入为 `start token == tokenizer_en.vocab_size`。\n* 计算填充遮挡和前瞻遮挡。\n* `解码器`通过查看`编码器输出`和它自身的输出(自注意力)给出预测。\n* 选择最后一个词并计算它的 argmax。\n* 将预测的词连接到解码器输入,然后传递给解码器。\n* 在这种方法中,解码器根据它预测的之前的词预测下一个。\n\nNote:这里使用的模型具有较小的能力以保持相对较快,因此预测可能不太正确。要复现论文中的结果,请使用全部数据集,并通过修改上述超参数来使用基础 transformer 模型或者 transformer XL。",
"_____no_output_____"
]
],
[
[
"def evaluate(inp_sentence):\n start_token = [tokenizer_pt.vocab_size]\n end_token = [tokenizer_pt.vocab_size + 1]\n \n # 输入语句是葡萄牙语,增加开始和结束标记\n inp_sentence = start_token + tokenizer_pt.encode(inp_sentence) + end_token\n encoder_input = tf.expand_dims(inp_sentence, 0)\n \n # 因为目标是英语,输入 transformer 的第一个词应该是\n # 英语的开始标记。\n decoder_input = [tokenizer_en.vocab_size]\n output = tf.expand_dims(decoder_input, 0)\n \n for i in range(MAX_LENGTH):\n enc_padding_mask, combined_mask, dec_padding_mask = create_masks(\n encoder_input, output)\n \n # predictions.shape == (batch_size, seq_len, vocab_size)\n predictions, attention_weights = transformer(encoder_input, \n output,\n False,\n enc_padding_mask,\n combined_mask,\n dec_padding_mask)\n \n # 从 seq_len 维度选择最后一个词\n predictions = predictions[: ,-1:, :] # (batch_size, 1, vocab_size)\n\n predicted_id = tf.cast(tf.argmax(predictions, axis=-1), tf.int32)\n \n # 如果 predicted_id 等于结束标记,就返回结果\n if predicted_id == tokenizer_en.vocab_size+1:\n return tf.squeeze(output, axis=0), attention_weights\n \n # 连接 predicted_id 与输出,作为解码器的输入传递到解码器。\n output = tf.concat([output, predicted_id], axis=-1)\n\n return tf.squeeze(output, axis=0), attention_weights",
"_____no_output_____"
],
[
"def plot_attention_weights(attention, sentence, result, layer):\n fig = plt.figure(figsize=(16, 8))\n \n sentence = tokenizer_pt.encode(sentence)\n \n attention = tf.squeeze(attention[layer], axis=0)\n \n for head in range(attention.shape[0]):\n ax = fig.add_subplot(2, 4, head+1)\n \n # 画出注意力权重\n ax.matshow(attention[head][:-1, :], cmap='viridis')\n\n fontdict = {'fontsize': 10}\n \n ax.set_xticks(range(len(sentence)+2))\n ax.set_yticks(range(len(result)))\n \n ax.set_ylim(len(result)-1.5, -0.5)\n \n ax.set_xticklabels(\n ['<start>']+[tokenizer_pt.decode([i]) for i in sentence]+['<end>'], \n fontdict=fontdict, rotation=90)\n \n ax.set_yticklabels([tokenizer_en.decode([i]) for i in result \n if i < tokenizer_en.vocab_size], \n fontdict=fontdict)\n \n ax.set_xlabel('Head {}'.format(head+1))\n \n plt.tight_layout()\n plt.show()",
"_____no_output_____"
],
[
"def translate(sentence, plot=''):\n result, attention_weights = evaluate(sentence)\n \n predicted_sentence = tokenizer_en.decode([i for i in result \n if i < tokenizer_en.vocab_size]) \n\n print('Input: {}'.format(sentence))\n print('Predicted translation: {}'.format(predicted_sentence))\n \n if plot:\n plot_attention_weights(attention_weights, sentence, result, plot)",
"_____no_output_____"
],
[
"translate(\"este é um problema que temos que resolver.\")\nprint (\"Real translation: this is a problem we have to solve .\")",
"_____no_output_____"
],
[
"translate(\"os meus vizinhos ouviram sobre esta ideia.\")\nprint (\"Real translation: and my neighboring homes heard about this idea .\")",
"_____no_output_____"
],
[
"translate(\"vou então muito rapidamente partilhar convosco algumas histórias de algumas coisas mágicas que aconteceram.\")\nprint (\"Real translation: so i 'll just share with you some stories very quickly of some magical things that have happened .\")",
"_____no_output_____"
]
],
[
[
"您可以为 `plot` 参数传递不同的层和解码器的注意力模块。",
"_____no_output_____"
]
],
[
[
"translate(\"este é o primeiro livro que eu fiz.\", plot='decoder_layer4_block2')\nprint (\"Real translation: this is the first book i've ever done.\")",
"_____no_output_____"
]
],
[
[
"## 总结\n\n在本教程中,您已经学习了位置编码,多头注意力,遮挡的重要性以及如何创建一个 transformer。\n\n尝试使用一个不同的数据集来训练 transformer。您可也可以通过修改上述的超参数来创建基础 transformer 或者 transformer XL。您也可以使用这里定义的层来创建 [BERT](https://arxiv.org/abs/1810.04805) 并训练最先进的模型。此外,您可以实现 beam search 得到更好的预测。",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
4ab81d369bbd9094f93fc5ac5cacdbab9261574b
| 10,728 |
ipynb
|
Jupyter Notebook
|
nlp-socher/lec09.ipynb
|
JasonWayne/course-notes
|
feff7a0636e7f8f2353c1dea24fe25296a8b33c3
|
[
"MIT"
] | null | null | null |
nlp-socher/lec09.ipynb
|
JasonWayne/course-notes
|
feff7a0636e7f8f2353c1dea24fe25296a8b33c3
|
[
"MIT"
] | null | null | null |
nlp-socher/lec09.ipynb
|
JasonWayne/course-notes
|
feff7a0636e7f8f2353c1dea24fe25296a8b33c3
|
[
"MIT"
] | null | null | null | 17.219904 | 74 | 0.477069 |
[
[
[
"##### 1",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 2",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 3",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 4",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 5",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 6",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 7",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 8",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 9",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 10",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 11",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 12",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 13",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 14",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 15",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 16",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 17",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 18",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 19",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 20",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 21",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 22",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 23",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 24",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 25",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 26",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 27",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 28",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 29",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 30",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 31",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 32",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 33",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 34",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 35",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 36",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 37",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 38",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 39",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 40",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 41",
"_____no_output_____"
],
[
"",
"_____no_output_____"
],
[
"##### 42",
"_____no_output_____"
],
[
"",
"_____no_output_____"
]
]
] |
[
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
4ab83139371d39b0ba4730bef6df78d64fb0c4fb
| 5,508 |
ipynb
|
Jupyter Notebook
|
notebooks/02-data-manipulation/2.6-subset-dataframe-by-column.ipynb
|
peymanyousefianasml/learning-apache-spark
|
675210da79cdb99b728deac0e5c1b0f9057a4357
|
[
"MIT"
] | 258 |
2017-04-28T09:58:56.000Z
|
2022-03-19T00:09:41.000Z
|
notebooks/02-data-manipulation/2.6-subset-dataframe-by-column.ipynb
|
pxu2018/learning-apache-spark
|
675210da79cdb99b728deac0e5c1b0f9057a4357
|
[
"MIT"
] | 2 |
2017-11-08T21:27:23.000Z
|
2017-11-08T21:52:13.000Z
|
notebooks/02-data-manipulation/2.6-subset-dataframe-by-column.ipynb
|
pxu2018/learning-apache-spark
|
675210da79cdb99b728deac0e5c1b0f9057a4357
|
[
"MIT"
] | 164 |
2017-05-23T17:35:11.000Z
|
2022-03-21T22:46:50.000Z
| 22.666667 | 96 | 0.433188 |
[
[
[
"# create entry points to spark\ntry:\n sc.stop()\nexcept:\n pass\nfrom pyspark import SparkContext, SparkConf\nfrom pyspark.sql import SparkSession\nsc=SparkContext()\nspark = SparkSession(sparkContext=sc)",
"_____no_output_____"
]
],
[
[
"## Example data",
"_____no_output_____"
]
],
[
[
"mtcars = spark.read.csv('../../data/mtcars.csv', inferSchema=True, header=True)\n# correct first column name\nmtcars = mtcars.withColumnRenamed('_c0', 'model')\nmtcars.show(5)",
"+-----------------+----+---+-----+---+----+-----+-----+---+---+----+----+\n| model| mpg|cyl| disp| hp|drat| wt| qsec| vs| am|gear|carb|\n+-----------------+----+---+-----+---+----+-----+-----+---+---+----+----+\n| Mazda RX4|21.0| 6|160.0|110| 3.9| 2.62|16.46| 0| 1| 4| 4|\n| Mazda RX4 Wag|21.0| 6|160.0|110| 3.9|2.875|17.02| 0| 1| 4| 4|\n| Datsun 710|22.8| 4|108.0| 93|3.85| 2.32|18.61| 1| 1| 4| 1|\n| Hornet 4 Drive|21.4| 6|258.0|110|3.08|3.215|19.44| 1| 0| 3| 1|\n|Hornet Sportabout|18.7| 8|360.0|175|3.15| 3.44|17.02| 0| 0| 3| 2|\n+-----------------+----+---+-----+---+----+-----+-----+---+---+----+----+\nonly showing top 5 rows\n\n"
]
],
[
[
"## Select columns by name\n\nWe can simply use the **select()** function to select columns by name.",
"_____no_output_____"
]
],
[
[
"mtcars.select(['hp', 'disp']).show(5)",
"+---+-----+\n| hp| disp|\n+---+-----+\n|110|160.0|\n|110|160.0|\n| 93|108.0|\n|110|258.0|\n|175|360.0|\n+---+-----+\nonly showing top 5 rows\n\n"
]
],
[
[
"## Select columns by index\n\nWe can convert indices to corresponding column names and then select columns by name.",
"_____no_output_____"
]
],
[
[
"indices = [0,3,4,7]\nselected_columns = [mtcars.columns[index] for index in indices]\nselected_columns",
"_____no_output_____"
],
[
"mtcars.select(selected_columns).show(5)",
"+-----------------+-----+---+-----+\n| model| disp| hp| qsec|\n+-----------------+-----+---+-----+\n| Mazda RX4|160.0|110|16.46|\n| Mazda RX4 Wag|160.0|110|17.02|\n| Datsun 710|108.0| 93|18.61|\n| Hornet 4 Drive|258.0|110|19.44|\n|Hornet Sportabout|360.0|175|17.02|\n+-----------------+-----+---+-----+\nonly showing top 5 rows\n\n"
]
],
[
[
"## Select columns by pattern\n\nExample: columns start with 'd'.",
"_____no_output_____"
]
],
[
[
"import re\nselected_columns = [x for x in mtcars.columns if re.compile('^d').match(x) is not None]\nselected_columns",
"_____no_output_____"
],
[
"mtcars.select(selected_columns).show(5)",
"+-----+----+\n| disp|drat|\n+-----+----+\n|160.0| 3.9|\n|160.0| 3.9|\n|108.0|3.85|\n|258.0|3.08|\n|360.0|3.15|\n+-----+----+\nonly showing top 5 rows\n\n"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4ab838057a172ded0cc2c5e9104e3b90cd505171
| 6,454 |
ipynb
|
Jupyter Notebook
|
models/call_me_model_maybe_.ipynb
|
StephenSpicer/Spotify_Music_Discovery_LS_DS_BW
|
2e61e98f45cdfeb064b58e44091ffa16d67d82b2
|
[
"MIT"
] | null | null | null |
models/call_me_model_maybe_.ipynb
|
StephenSpicer/Spotify_Music_Discovery_LS_DS_BW
|
2e61e98f45cdfeb064b58e44091ffa16d67d82b2
|
[
"MIT"
] | null | null | null |
models/call_me_model_maybe_.ipynb
|
StephenSpicer/Spotify_Music_Discovery_LS_DS_BW
|
2e61e98f45cdfeb064b58e44091ffa16d67d82b2
|
[
"MIT"
] | 5 |
2021-03-27T21:42:39.000Z
|
2021-03-31T15:12:06.000Z
| 32.432161 | 672 | 0.590951 |
[
[
[
"empty"
]
]
] |
[
"empty"
] |
[
[
"empty"
]
] |
4ab8410f3bfece94b411dc87034bcc0467a49410
| 79,885 |
ipynb
|
Jupyter Notebook
|
Check datas by Lasso, LassoCV, Ridge, RidgeCV 14.05.2017.ipynb
|
MikhailEpatko/Mashine-Learning
|
108e449deb0199a53b44a4053ae0711b49145b8d
|
[
"Apache-2.0"
] | null | null | null |
Check datas by Lasso, LassoCV, Ridge, RidgeCV 14.05.2017.ipynb
|
MikhailEpatko/Mashine-Learning
|
108e449deb0199a53b44a4053ae0711b49145b8d
|
[
"Apache-2.0"
] | null | null | null |
Check datas by Lasso, LassoCV, Ridge, RidgeCV 14.05.2017.ipynb
|
MikhailEpatko/Mashine-Learning
|
108e449deb0199a53b44a4053ae0711b49145b8d
|
[
"Apache-2.0"
] | null | null | null | 105.528402 | 38,516 | 0.853439 |
[
[
[
"import pandas as pd\nimport numpy as np\nfrom sklearn.metrics import accuracy_score\nfrom sklearn.metrics import confusion_matrix\nfrom sklearn.metrics import classification_report\nfrom sklearn.model_selection import train_test_split",
"_____no_output_____"
],
[
"dtypes = {'nt_fp': str, 'event_date': str, 'day_of_week': int, 'a_month': int, 'hour_of_day': int, 's_depcity': str, 's_adults': int, \n 's_children': int, 'ua_browser': str, 'ua_device_type': str, 'ua_platform': str, 'ip_lat': int, 'ip_long': int,\n 's_country': str}\n\nparse_dates = ['event_date']\n\ndata_df = pd.read_csv('~/Desktop/events-by-week.csv', index_col=False, dtype=dtypes, parse_dates=parse_dates)\n\ndata_df['week_of_year'] = data_df['event_date'].dt.week\n\ndata_target = data_df['s_country']\n# data_features = data_df.drop(columns=['s_country', 'event_date', 'nt_fp', 's_depcity', \n# 'ua_device_type', 'ua_browser', 'day_of_week', 'ip_lat',\n# 'ua_platform', 'hour_of_day', 'a_month'])\n\ndata_features = data_df.drop(columns=['s_country', 'event_date', 'nt_fp', 's_depcity', \n 'ua_device_type', 'ua_browser', 'day_of_week', 'week_of_year',\n 'ua_platform', 'hour_of_day', 'a_month'])\n\n\n\ndata_dummies = pd.get_dummies(data_features)\n# data_dummies = data_features\n\nindex_split = int (round(data_dummies.shape[0] * 0.75, 0))\n\ntrain_X = data_dummies[:index_split][:]\ntest_X = data_dummies[index_split:][:]\ntrain_y = data_target[:index_split][:]\ntest_y = data_target[index_split:][:]\n\nX_train = train_X.values\nX_test = test_X.values\ny_train = train_y.values\ny_test = test_y.values",
"_____no_output_____"
],
[
"from sklearn.ensemble import GradientBoostingClassifier\nfrom sklearn import preprocessing",
"_____no_output_____"
]
],
[
[
"### С нормализованными данными",
"_____no_output_____"
]
],
[
[
"data_dummies = preprocessing.normalize(data_features)\ndata_dummies.size",
"_____no_output_____"
],
[
"index_split = int (round(data_dummies.size * 0.75, 0))",
"_____no_output_____"
],
[
"X_train = data_dummies[:index_split][:]\nX_test = data_dummies[index_split:][:]\ny_train = data_target[:index_split][:]\ny_test = data_target[index_split:][:]\n\n",
"_____no_output_____"
],
[
"grb = GradientBoostingClassifier(n_estimators=50, random_state=17, learning_rate=0.01)\ngrb.fit(X_train, y_train)",
"_____no_output_____"
],
[
"# print(\"Правильность на обучающем наборе: {:.5f}\".format(grb.score(X_train, y_train)))\n# print('Правильность на тестовом наборе: {:.5f}'.format(grb.score(X_test, y_test)))",
"_____no_output_____"
],
[
"predictions = grb.predict(X_test)",
"_____no_output_____"
],
[
"for name, score in zip(data_samples.columns, grb.feature_importances_):\n print(name, score)",
"_____no_output_____"
]
],
[
[
"### Со стандартизированными данными",
"_____no_output_____"
]
],
[
[
"data_dummies = preprocessing.scale(data_features)\n\nindex_split = int (round(data_dummies.size * 0.75, 0))\n\nX_train = data_dummies[:index_split][:]\nX_test = data_dummies[index_split:][:]\ny_train = data_target[:index_split][:]\ny_test = data_target[index_split:][:]",
"_____no_output_____"
],
[
"grb = GradientBoostingClassifier(n_estimators=50, random_state=17, learning_rate=0.01)\ngrb.fit(X_train, y_train)",
"_____no_output_____"
],
[
"print(\"Правильность на обучающем наборе: {:.5f}\".format(grb.score(X_train, y_train)))\nprint('Правильность на тестовом наборе: {:.5f}'.format(grb.score(X_test, y_test)))",
"_____no_output_____"
],
[
"predictions = grb.predict(X_test)",
"_____no_output_____"
],
[
"for name, score in zip(data_samples.columns, grb.feature_importances_):\n print(name, score)",
"_____no_output_____"
],
[
"data_df.shape\n",
"_____no_output_____"
],
[
"data_features.head()",
"_____no_output_____"
]
],
[
[
"### Графики",
"_____no_output_____"
]
],
[
[
"%matplotlib inline\nfrom matplotlib import pyplot as plt\nimport seaborn as sns\n\nsns.heatmap(data_dummies.corr())",
"_____no_output_____"
],
[
"plt.rcParams['figure.figsize'] = 12, 8\nsns.violinplot(data_df['s_adults'], data_df['s_children'])",
"_____no_output_____"
],
[
"for i, col in enumerate(data_df.columns[:-1]):\n plt.subplot(5,3,i+1)\n plt.scatter(data_df[col], data_df['s_country'])\n plt.title(col)",
"_____no_output_____"
],
[
"plt.scatter(data_df['hour_of_day'], data_df['ip_long']) ",
"_____no_output_____"
]
],
[
[
"### Линейные модели",
"_____no_output_____"
]
],
[
[
"from sklearn.linear_model import LinearRegression, Lasso, Ridge\nfrom sklearn.linear_model import LassoCV, RidgeCV\nfrom sklearn.ensemble import RandomForestRegressor\nfrom sklearn.preprocessing import StandardScaler\nfrom sklearn.metrics import mean_squared_error",
"_____no_output_____"
],
[
"linreg = LinearRegression()\nlasso = Lasso(random_state=17)\nridge = Ridge(random_state=17)\nlasso_cv = LassoCV(random_state=17)\nridge_cv = RidgeCV()",
"_____no_output_____"
],
[
"scaler = StandardScaler()\n# масштабируем данные\nX_train_scaled = scaler.fit_transform(X_train)\nX_test_scaled = scaler.transform(X_test)",
"/home/mikhail/anaconda3/lib/python3.6/site-packages/sklearn/utils/validation.py:475: DataConversionWarning: Data with input dtype int64 was converted to float64 by StandardScaler.\n warnings.warn(msg, DataConversionWarning)\n"
],
[
"linreg.fit(X_train_scaled, y_train)",
"_____no_output_____"
],
[
"# среднеквадратичная ошибка между ответами на тестовой выборке и прогнозами метода наименьших квадратов\nnp.sqrt(mean_squared_error(y_test, linreg.predict(X_test_scaled)) )",
"_____no_output_____"
],
[
"pd.DataFrame(linreg.coef_, data_dummies.columns[:], columns=['coef']).sort_values(by='coef', ascending=False)",
"_____no_output_____"
],
[
"# модель, котора будет возвращать список коэффициентов:\n\ndef report(model, X_train_scaled, y_train, X_test_scaled, y_test, feature_names):\n model.fit(X_train_scaled, y_train)\n print('MSE = %f' % np.sqrt(mean_squared_error(y_test, linreg.predict(X_test_scaled))))\n print(pd.DataFrame(model.coef_, feature_names, columns=['coef']).sort_values(by='coef', ascending=False))",
"_____no_output_____"
],
[
"report(lasso, X_train_scaled, y_train, X_test_scaled, y_test, data_dummies.columns[:])",
"MSE = 119111.313968\n coef\ns_adults 5213.820585\ns_children 532.148627\nip_lat 456.908911\nip_long 345.360173\n"
],
[
"report(lasso_cv, X_train_scaled, y_train, X_test_scaled, y_test, data_dummies.columns[:])",
"MSE = 119111.313968\n coef\ns_adults 5010.467187\ns_children 336.543094\nip_lat 166.614857\nip_long 49.602080\n"
],
[
"report(ridge, X_train_scaled, y_train, X_test_scaled, y_test, data_dummies.columns[:])",
"MSE = 119111.313968\n coef\ns_adults 5214.414755\ns_children 532.865662\nip_lat 457.912122\nip_long 346.382653\n"
],
[
"report(ridge_cv, X_train_scaled, y_train, X_test_scaled, y_test, data_dummies.columns[:])",
"MSE = 119111.313968\n coef\ns_adults 5214.205696\ns_children 532.929079\nip_lat 457.897333\nip_long 346.371542\n"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab84a86bb6ad87e3d9688557022655c33514de7
| 6,684 |
ipynb
|
Jupyter Notebook
|
Airtable/Airtable_Search_data.ipynb
|
techthiyanes/awesome-notebooks
|
10ab4da1b94dfa101e908356a649609b0b17561a
|
[
"BSD-3-Clause"
] | null | null | null |
Airtable/Airtable_Search_data.ipynb
|
techthiyanes/awesome-notebooks
|
10ab4da1b94dfa101e908356a649609b0b17561a
|
[
"BSD-3-Clause"
] | null | null | null |
Airtable/Airtable_Search_data.ipynb
|
techthiyanes/awesome-notebooks
|
10ab4da1b94dfa101e908356a649609b0b17561a
|
[
"BSD-3-Clause"
] | null | null | null | 24.847584 | 283 | 0.592908 |
[
[
[
"<img width=\"10%\" alt=\"Naas\" src=\"https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160\"/>",
"_____no_output_____"
],
[
"# Airtable - Search data\n<a href=\"https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/Airtable/Airtable_Search_data.ipynb\" target=\"_parent\"><img src=\"https://naasai-public.s3.eu-west-3.amazonaws.com/open_in_naas.svg\"/></a>",
"_____no_output_____"
],
[
"**Tags:** #airtable #database #productivity #spreadsheet #naas_drivers #operations #snippet #dataframe",
"_____no_output_____"
],
[
"**Author:** [Jeremy Ravenel](https://www.linkedin.com/in/ACoAAAJHE7sB5OxuKHuzguZ9L6lfDHqw--cdnJg/)",
"_____no_output_____"
],
[
"## Input",
"_____no_output_____"
],
[
"### Import library",
"_____no_output_____"
]
],
[
[
"from naas_drivers import airtable",
"_____no_output_____"
]
],
[
[
"### Variables",
"_____no_output_____"
]
],
[
[
"API_KEY = 'API_KEY'\nBASE_KEY = 'BASE_KEY'\nTABLE_NAME = 'TABLE_NAME'",
"_____no_output_____"
]
],
[
[
"## Model",
"_____no_output_____"
],
[
"### Search data",
"_____no_output_____"
]
],
[
[
"data = airtable.connect(API_KEY,\n BASE_KEY,\n TABLE_NAME).get(view='All opportunities',\n maxRecords=20).search('Name', 'Tom')",
"_____no_output_____"
]
],
[
[
"## Output",
"_____no_output_____"
],
[
"### Display result",
"_____no_output_____"
]
],
[
[
"data",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
]
] |
4ab853f422b2c5d6b2fb32b65b6b972653248cce
| 424,830 |
ipynb
|
Jupyter Notebook
|
AI 이노베이션 스퀘어 시각지능 과정/202006/20200604/Pre-trained model predict.ipynb
|
donddog/AI_Innovation_Square_Codes
|
a04d50db011d25e00d8486146c24124c50242aa7
|
[
"MIT"
] | 1 |
2021-02-11T16:45:21.000Z
|
2021-02-11T16:45:21.000Z
|
AI 이노베이션 스퀘어 시각지능 과정/202006/20200604/Pre-trained model predict.ipynb
|
donddog/AI_Innovation_Square_Codes
|
a04d50db011d25e00d8486146c24124c50242aa7
|
[
"MIT"
] | null | null | null |
AI 이노베이션 스퀘어 시각지능 과정/202006/20200604/Pre-trained model predict.ipynb
|
donddog/AI_Innovation_Square_Codes
|
a04d50db011d25e00d8486146c24124c50242aa7
|
[
"MIT"
] | null | null | null | 392.634011 | 198,048 | 0.89896 |
[
[
[
"케라스에 저장되어있는 미리 학습된 모델을 불러오고 사용하는 방법을 알아보겠습니다. weights=None 을 넣어주면 웨이트가 랜덤으로 초기화되고 weights='imagenet' 옵션을 주면 이미지넷 데이터로 학습된 웨이트가 불러와 집니다.",
"_____no_output_____"
]
],
[
[
"from tensorflow.keras.applications.resnet50 import ResNet50 as pre_model\nfrom tensorflow.keras.applications.resnet50 import preprocess_input, decode_predictions\n\nfrom tensorflow.keras.applications.xception import Xception as pre_model\nfrom tensorflow.keras.applications.xception import preprocess_input, decode_predictions\n\nfrom tensorflow.keras.applications.vgg16 import VGG16 as pre_model\nfrom tensorflow.keras.applications.vgg16 import preprocess_input, decode_predictions\n\nfrom tensorflow.keras.applications.vgg19 import VGG19 as pre_model\nfrom tensorflow.keras.applications.vgg19 import preprocess_input, decode_predictions\n\nfrom tensorflow.keras.applications.inception_v3 import InceptionV3 as pre_model\nfrom tensorflow.keras.applications.inception_v3 import preprocess_input, decode_predictions\n\nmodel = pre_model(weights='imagenet')\nmodel.summary()",
"Model: \"inception_v3\"\n__________________________________________________________________________________________________\nLayer (type) Output Shape Param # Connected to \n==================================================================================================\ninput_1 (InputLayer) [(None, 299, 299, 3) 0 \n__________________________________________________________________________________________________\nconv2d (Conv2D) (None, 149, 149, 32) 864 input_1[0][0] \n__________________________________________________________________________________________________\nbatch_normalization (BatchNorma (None, 149, 149, 32) 96 conv2d[0][0] \n__________________________________________________________________________________________________\nactivation (Activation) (None, 149, 149, 32) 0 batch_normalization[0][0] \n__________________________________________________________________________________________________\nconv2d_1 (Conv2D) (None, 147, 147, 32) 9216 activation[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_1 (BatchNor (None, 147, 147, 32) 96 conv2d_1[0][0] \n__________________________________________________________________________________________________\nactivation_1 (Activation) (None, 147, 147, 32) 0 batch_normalization_1[0][0] \n__________________________________________________________________________________________________\nconv2d_2 (Conv2D) (None, 147, 147, 64) 18432 activation_1[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_2 (BatchNor (None, 147, 147, 64) 192 conv2d_2[0][0] \n__________________________________________________________________________________________________\nactivation_2 (Activation) (None, 147, 147, 64) 0 batch_normalization_2[0][0] \n__________________________________________________________________________________________________\nmax_pooling2d (MaxPooling2D) (None, 73, 73, 64) 0 activation_2[0][0] \n__________________________________________________________________________________________________\nconv2d_3 (Conv2D) (None, 73, 73, 80) 5120 max_pooling2d[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_3 (BatchNor (None, 73, 73, 80) 240 conv2d_3[0][0] \n__________________________________________________________________________________________________\nactivation_3 (Activation) (None, 73, 73, 80) 0 batch_normalization_3[0][0] \n__________________________________________________________________________________________________\nconv2d_4 (Conv2D) (None, 71, 71, 192) 138240 activation_3[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_4 (BatchNor (None, 71, 71, 192) 576 conv2d_4[0][0] \n__________________________________________________________________________________________________\nactivation_4 (Activation) (None, 71, 71, 192) 0 batch_normalization_4[0][0] \n__________________________________________________________________________________________________\nmax_pooling2d_1 (MaxPooling2D) (None, 35, 35, 192) 0 activation_4[0][0] \n__________________________________________________________________________________________________\nconv2d_8 (Conv2D) (None, 35, 35, 64) 12288 max_pooling2d_1[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_8 (BatchNor (None, 35, 35, 64) 192 conv2d_8[0][0] \n__________________________________________________________________________________________________\nactivation_8 (Activation) (None, 35, 35, 64) 0 batch_normalization_8[0][0] \n__________________________________________________________________________________________________\nconv2d_6 (Conv2D) (None, 35, 35, 48) 9216 max_pooling2d_1[0][0] \n__________________________________________________________________________________________________\nconv2d_9 (Conv2D) (None, 35, 35, 96) 55296 activation_8[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_6 (BatchNor (None, 35, 35, 48) 144 conv2d_6[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_9 (BatchNor (None, 35, 35, 96) 288 conv2d_9[0][0] \n__________________________________________________________________________________________________\nactivation_6 (Activation) (None, 35, 35, 48) 0 batch_normalization_6[0][0] \n__________________________________________________________________________________________________\nactivation_9 (Activation) (None, 35, 35, 96) 0 batch_normalization_9[0][0] \n__________________________________________________________________________________________________\naverage_pooling2d (AveragePooli (None, 35, 35, 192) 0 max_pooling2d_1[0][0] \n__________________________________________________________________________________________________\nconv2d_5 (Conv2D) (None, 35, 35, 64) 12288 max_pooling2d_1[0][0] \n__________________________________________________________________________________________________\nconv2d_7 (Conv2D) (None, 35, 35, 64) 76800 activation_6[0][0] \n__________________________________________________________________________________________________\nconv2d_10 (Conv2D) (None, 35, 35, 96) 82944 activation_9[0][0] \n__________________________________________________________________________________________________\nconv2d_11 (Conv2D) (None, 35, 35, 32) 6144 average_pooling2d[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_5 (BatchNor (None, 35, 35, 64) 192 conv2d_5[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_7 (BatchNor (None, 35, 35, 64) 192 conv2d_7[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_10 (BatchNo (None, 35, 35, 96) 288 conv2d_10[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_11 (BatchNo (None, 35, 35, 32) 96 conv2d_11[0][0] \n__________________________________________________________________________________________________\nactivation_5 (Activation) (None, 35, 35, 64) 0 batch_normalization_5[0][0] \n__________________________________________________________________________________________________\nactivation_7 (Activation) (None, 35, 35, 64) 0 batch_normalization_7[0][0] \n__________________________________________________________________________________________________\nactivation_10 (Activation) (None, 35, 35, 96) 0 batch_normalization_10[0][0] \n__________________________________________________________________________________________________\nactivation_11 (Activation) (None, 35, 35, 32) 0 batch_normalization_11[0][0] \n__________________________________________________________________________________________________\nmixed0 (Concatenate) (None, 35, 35, 256) 0 activation_5[0][0] \n activation_7[0][0] \n activation_10[0][0] \n activation_11[0][0] \n__________________________________________________________________________________________________\nconv2d_15 (Conv2D) (None, 35, 35, 64) 16384 mixed0[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_15 (BatchNo (None, 35, 35, 64) 192 conv2d_15[0][0] \n__________________________________________________________________________________________________\nactivation_15 (Activation) (None, 35, 35, 64) 0 batch_normalization_15[0][0] \n__________________________________________________________________________________________________\nconv2d_13 (Conv2D) (None, 35, 35, 48) 12288 mixed0[0][0] \n__________________________________________________________________________________________________\nconv2d_16 (Conv2D) (None, 35, 35, 96) 55296 activation_15[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_13 (BatchNo (None, 35, 35, 48) 144 conv2d_13[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_16 (BatchNo (None, 35, 35, 96) 288 conv2d_16[0][0] \n__________________________________________________________________________________________________\nactivation_13 (Activation) (None, 35, 35, 48) 0 batch_normalization_13[0][0] \n__________________________________________________________________________________________________\nactivation_16 (Activation) (None, 35, 35, 96) 0 batch_normalization_16[0][0] \n__________________________________________________________________________________________________\naverage_pooling2d_1 (AveragePoo (None, 35, 35, 256) 0 mixed0[0][0] \n__________________________________________________________________________________________________\nconv2d_12 (Conv2D) (None, 35, 35, 64) 16384 mixed0[0][0] \n__________________________________________________________________________________________________\nconv2d_14 (Conv2D) (None, 35, 35, 64) 76800 activation_13[0][0] \n__________________________________________________________________________________________________\nconv2d_17 (Conv2D) (None, 35, 35, 96) 82944 activation_16[0][0] \n__________________________________________________________________________________________________\nconv2d_18 (Conv2D) (None, 35, 35, 64) 16384 average_pooling2d_1[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_12 (BatchNo (None, 35, 35, 64) 192 conv2d_12[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_14 (BatchNo (None, 35, 35, 64) 192 conv2d_14[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_17 (BatchNo (None, 35, 35, 96) 288 conv2d_17[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_18 (BatchNo (None, 35, 35, 64) 192 conv2d_18[0][0] \n__________________________________________________________________________________________________\nactivation_12 (Activation) (None, 35, 35, 64) 0 batch_normalization_12[0][0] \n__________________________________________________________________________________________________\nactivation_14 (Activation) (None, 35, 35, 64) 0 batch_normalization_14[0][0] \n__________________________________________________________________________________________________\nactivation_17 (Activation) (None, 35, 35, 96) 0 batch_normalization_17[0][0] \n__________________________________________________________________________________________________\nactivation_18 (Activation) (None, 35, 35, 64) 0 batch_normalization_18[0][0] \n__________________________________________________________________________________________________\nmixed1 (Concatenate) (None, 35, 35, 288) 0 activation_12[0][0] \n activation_14[0][0] \n activation_17[0][0] \n activation_18[0][0] \n__________________________________________________________________________________________________\nconv2d_22 (Conv2D) (None, 35, 35, 64) 18432 mixed1[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_22 (BatchNo (None, 35, 35, 64) 192 conv2d_22[0][0] \n__________________________________________________________________________________________________\nactivation_22 (Activation) (None, 35, 35, 64) 0 batch_normalization_22[0][0] \n__________________________________________________________________________________________________\nconv2d_20 (Conv2D) (None, 35, 35, 48) 13824 mixed1[0][0] \n__________________________________________________________________________________________________\nconv2d_23 (Conv2D) (None, 35, 35, 96) 55296 activation_22[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_20 (BatchNo (None, 35, 35, 48) 144 conv2d_20[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_23 (BatchNo (None, 35, 35, 96) 288 conv2d_23[0][0] \n__________________________________________________________________________________________________\nactivation_20 (Activation) (None, 35, 35, 48) 0 batch_normalization_20[0][0] \n__________________________________________________________________________________________________\nactivation_23 (Activation) (None, 35, 35, 96) 0 batch_normalization_23[0][0] \n__________________________________________________________________________________________________\naverage_pooling2d_2 (AveragePoo (None, 35, 35, 288) 0 mixed1[0][0] \n__________________________________________________________________________________________________\nconv2d_19 (Conv2D) (None, 35, 35, 64) 18432 mixed1[0][0] \n__________________________________________________________________________________________________\nconv2d_21 (Conv2D) (None, 35, 35, 64) 76800 activation_20[0][0] \n__________________________________________________________________________________________________\nconv2d_24 (Conv2D) (None, 35, 35, 96) 82944 activation_23[0][0] \n__________________________________________________________________________________________________\nconv2d_25 (Conv2D) (None, 35, 35, 64) 18432 average_pooling2d_2[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_19 (BatchNo (None, 35, 35, 64) 192 conv2d_19[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_21 (BatchNo (None, 35, 35, 64) 192 conv2d_21[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_24 (BatchNo (None, 35, 35, 96) 288 conv2d_24[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_25 (BatchNo (None, 35, 35, 64) 192 conv2d_25[0][0] \n__________________________________________________________________________________________________\nactivation_19 (Activation) (None, 35, 35, 64) 0 batch_normalization_19[0][0] \n__________________________________________________________________________________________________\nactivation_21 (Activation) (None, 35, 35, 64) 0 batch_normalization_21[0][0] \n__________________________________________________________________________________________________\nactivation_24 (Activation) (None, 35, 35, 96) 0 batch_normalization_24[0][0] \n__________________________________________________________________________________________________\nactivation_25 (Activation) (None, 35, 35, 64) 0 batch_normalization_25[0][0] \n__________________________________________________________________________________________________\nmixed2 (Concatenate) (None, 35, 35, 288) 0 activation_19[0][0] \n activation_21[0][0] \n activation_24[0][0] \n activation_25[0][0] \n__________________________________________________________________________________________________\nconv2d_27 (Conv2D) (None, 35, 35, 64) 18432 mixed2[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_27 (BatchNo (None, 35, 35, 64) 192 conv2d_27[0][0] \n__________________________________________________________________________________________________\nactivation_27 (Activation) (None, 35, 35, 64) 0 batch_normalization_27[0][0] \n__________________________________________________________________________________________________\nconv2d_28 (Conv2D) (None, 35, 35, 96) 55296 activation_27[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_28 (BatchNo (None, 35, 35, 96) 288 conv2d_28[0][0] \n__________________________________________________________________________________________________\nactivation_28 (Activation) (None, 35, 35, 96) 0 batch_normalization_28[0][0] \n__________________________________________________________________________________________________\nconv2d_26 (Conv2D) (None, 17, 17, 384) 995328 mixed2[0][0] \n__________________________________________________________________________________________________\nconv2d_29 (Conv2D) (None, 17, 17, 96) 82944 activation_28[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_26 (BatchNo (None, 17, 17, 384) 1152 conv2d_26[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_29 (BatchNo (None, 17, 17, 96) 288 conv2d_29[0][0] \n__________________________________________________________________________________________________\nactivation_26 (Activation) (None, 17, 17, 384) 0 batch_normalization_26[0][0] \n__________________________________________________________________________________________________\nactivation_29 (Activation) (None, 17, 17, 96) 0 batch_normalization_29[0][0] \n__________________________________________________________________________________________________\nmax_pooling2d_2 (MaxPooling2D) (None, 17, 17, 288) 0 mixed2[0][0] \n__________________________________________________________________________________________________\nmixed3 (Concatenate) (None, 17, 17, 768) 0 activation_26[0][0] \n activation_29[0][0] \n max_pooling2d_2[0][0] \n__________________________________________________________________________________________________\nconv2d_34 (Conv2D) (None, 17, 17, 128) 98304 mixed3[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_34 (BatchNo (None, 17, 17, 128) 384 conv2d_34[0][0] \n__________________________________________________________________________________________________\nactivation_34 (Activation) (None, 17, 17, 128) 0 batch_normalization_34[0][0] \n__________________________________________________________________________________________________\nconv2d_35 (Conv2D) (None, 17, 17, 128) 114688 activation_34[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_35 (BatchNo (None, 17, 17, 128) 384 conv2d_35[0][0] \n__________________________________________________________________________________________________\nactivation_35 (Activation) (None, 17, 17, 128) 0 batch_normalization_35[0][0] \n__________________________________________________________________________________________________\nconv2d_31 (Conv2D) (None, 17, 17, 128) 98304 mixed3[0][0] \n__________________________________________________________________________________________________\nconv2d_36 (Conv2D) (None, 17, 17, 128) 114688 activation_35[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_31 (BatchNo (None, 17, 17, 128) 384 conv2d_31[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_36 (BatchNo (None, 17, 17, 128) 384 conv2d_36[0][0] \n__________________________________________________________________________________________________\nactivation_31 (Activation) (None, 17, 17, 128) 0 batch_normalization_31[0][0] \n__________________________________________________________________________________________________\nactivation_36 (Activation) (None, 17, 17, 128) 0 batch_normalization_36[0][0] \n__________________________________________________________________________________________________\nconv2d_32 (Conv2D) (None, 17, 17, 128) 114688 activation_31[0][0] \n__________________________________________________________________________________________________\nconv2d_37 (Conv2D) (None, 17, 17, 128) 114688 activation_36[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_32 (BatchNo (None, 17, 17, 128) 384 conv2d_32[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_37 (BatchNo (None, 17, 17, 128) 384 conv2d_37[0][0] \n__________________________________________________________________________________________________\nactivation_32 (Activation) (None, 17, 17, 128) 0 batch_normalization_32[0][0] \n__________________________________________________________________________________________________\nactivation_37 (Activation) (None, 17, 17, 128) 0 batch_normalization_37[0][0] \n__________________________________________________________________________________________________\naverage_pooling2d_3 (AveragePoo (None, 17, 17, 768) 0 mixed3[0][0] \n__________________________________________________________________________________________________\nconv2d_30 (Conv2D) (None, 17, 17, 192) 147456 mixed3[0][0] \n__________________________________________________________________________________________________\nconv2d_33 (Conv2D) (None, 17, 17, 192) 172032 activation_32[0][0] \n__________________________________________________________________________________________________\nconv2d_38 (Conv2D) (None, 17, 17, 192) 172032 activation_37[0][0] \n__________________________________________________________________________________________________\nconv2d_39 (Conv2D) (None, 17, 17, 192) 147456 average_pooling2d_3[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_30 (BatchNo (None, 17, 17, 192) 576 conv2d_30[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_33 (BatchNo (None, 17, 17, 192) 576 conv2d_33[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_38 (BatchNo (None, 17, 17, 192) 576 conv2d_38[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_39 (BatchNo (None, 17, 17, 192) 576 conv2d_39[0][0] \n__________________________________________________________________________________________________\nactivation_30 (Activation) (None, 17, 17, 192) 0 batch_normalization_30[0][0] \n__________________________________________________________________________________________________\nactivation_33 (Activation) (None, 17, 17, 192) 0 batch_normalization_33[0][0] \n__________________________________________________________________________________________________\nactivation_38 (Activation) (None, 17, 17, 192) 0 batch_normalization_38[0][0] \n__________________________________________________________________________________________________\nactivation_39 (Activation) (None, 17, 17, 192) 0 batch_normalization_39[0][0] \n__________________________________________________________________________________________________\nmixed4 (Concatenate) (None, 17, 17, 768) 0 activation_30[0][0] \n activation_33[0][0] \n activation_38[0][0] \n activation_39[0][0] \n__________________________________________________________________________________________________\nconv2d_44 (Conv2D) (None, 17, 17, 160) 122880 mixed4[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_44 (BatchNo (None, 17, 17, 160) 480 conv2d_44[0][0] \n__________________________________________________________________________________________________\nactivation_44 (Activation) (None, 17, 17, 160) 0 batch_normalization_44[0][0] \n__________________________________________________________________________________________________\nconv2d_45 (Conv2D) (None, 17, 17, 160) 179200 activation_44[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_45 (BatchNo (None, 17, 17, 160) 480 conv2d_45[0][0] \n__________________________________________________________________________________________________\nactivation_45 (Activation) (None, 17, 17, 160) 0 batch_normalization_45[0][0] \n__________________________________________________________________________________________________\nconv2d_41 (Conv2D) (None, 17, 17, 160) 122880 mixed4[0][0] \n__________________________________________________________________________________________________\nconv2d_46 (Conv2D) (None, 17, 17, 160) 179200 activation_45[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_41 (BatchNo (None, 17, 17, 160) 480 conv2d_41[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_46 (BatchNo (None, 17, 17, 160) 480 conv2d_46[0][0] \n__________________________________________________________________________________________________\nactivation_41 (Activation) (None, 17, 17, 160) 0 batch_normalization_41[0][0] \n__________________________________________________________________________________________________\nactivation_46 (Activation) (None, 17, 17, 160) 0 batch_normalization_46[0][0] \n__________________________________________________________________________________________________\nconv2d_42 (Conv2D) (None, 17, 17, 160) 179200 activation_41[0][0] \n__________________________________________________________________________________________________\nconv2d_47 (Conv2D) (None, 17, 17, 160) 179200 activation_46[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_42 (BatchNo (None, 17, 17, 160) 480 conv2d_42[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_47 (BatchNo (None, 17, 17, 160) 480 conv2d_47[0][0] \n__________________________________________________________________________________________________\nactivation_42 (Activation) (None, 17, 17, 160) 0 batch_normalization_42[0][0] \n__________________________________________________________________________________________________\nactivation_47 (Activation) (None, 17, 17, 160) 0 batch_normalization_47[0][0] \n__________________________________________________________________________________________________\naverage_pooling2d_4 (AveragePoo (None, 17, 17, 768) 0 mixed4[0][0] \n__________________________________________________________________________________________________\nconv2d_40 (Conv2D) (None, 17, 17, 192) 147456 mixed4[0][0] \n__________________________________________________________________________________________________\nconv2d_43 (Conv2D) (None, 17, 17, 192) 215040 activation_42[0][0] \n__________________________________________________________________________________________________\nconv2d_48 (Conv2D) (None, 17, 17, 192) 215040 activation_47[0][0] \n__________________________________________________________________________________________________\nconv2d_49 (Conv2D) (None, 17, 17, 192) 147456 average_pooling2d_4[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_40 (BatchNo (None, 17, 17, 192) 576 conv2d_40[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_43 (BatchNo (None, 17, 17, 192) 576 conv2d_43[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_48 (BatchNo (None, 17, 17, 192) 576 conv2d_48[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_49 (BatchNo (None, 17, 17, 192) 576 conv2d_49[0][0] \n__________________________________________________________________________________________________\nactivation_40 (Activation) (None, 17, 17, 192) 0 batch_normalization_40[0][0] \n__________________________________________________________________________________________________\nactivation_43 (Activation) (None, 17, 17, 192) 0 batch_normalization_43[0][0] \n__________________________________________________________________________________________________\nactivation_48 (Activation) (None, 17, 17, 192) 0 batch_normalization_48[0][0] \n__________________________________________________________________________________________________\nactivation_49 (Activation) (None, 17, 17, 192) 0 batch_normalization_49[0][0] \n__________________________________________________________________________________________________\nmixed5 (Concatenate) (None, 17, 17, 768) 0 activation_40[0][0] \n activation_43[0][0] \n activation_48[0][0] \n activation_49[0][0] \n__________________________________________________________________________________________________\nconv2d_54 (Conv2D) (None, 17, 17, 160) 122880 mixed5[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_54 (BatchNo (None, 17, 17, 160) 480 conv2d_54[0][0] \n__________________________________________________________________________________________________\nactivation_54 (Activation) (None, 17, 17, 160) 0 batch_normalization_54[0][0] \n__________________________________________________________________________________________________\nconv2d_55 (Conv2D) (None, 17, 17, 160) 179200 activation_54[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_55 (BatchNo (None, 17, 17, 160) 480 conv2d_55[0][0] \n__________________________________________________________________________________________________\nactivation_55 (Activation) (None, 17, 17, 160) 0 batch_normalization_55[0][0] \n__________________________________________________________________________________________________\nconv2d_51 (Conv2D) (None, 17, 17, 160) 122880 mixed5[0][0] \n__________________________________________________________________________________________________\nconv2d_56 (Conv2D) (None, 17, 17, 160) 179200 activation_55[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_51 (BatchNo (None, 17, 17, 160) 480 conv2d_51[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_56 (BatchNo (None, 17, 17, 160) 480 conv2d_56[0][0] \n__________________________________________________________________________________________________\nactivation_51 (Activation) (None, 17, 17, 160) 0 batch_normalization_51[0][0] \n__________________________________________________________________________________________________\nactivation_56 (Activation) (None, 17, 17, 160) 0 batch_normalization_56[0][0] \n__________________________________________________________________________________________________\nconv2d_52 (Conv2D) (None, 17, 17, 160) 179200 activation_51[0][0] \n__________________________________________________________________________________________________\nconv2d_57 (Conv2D) (None, 17, 17, 160) 179200 activation_56[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_52 (BatchNo (None, 17, 17, 160) 480 conv2d_52[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_57 (BatchNo (None, 17, 17, 160) 480 conv2d_57[0][0] \n__________________________________________________________________________________________________\nactivation_52 (Activation) (None, 17, 17, 160) 0 batch_normalization_52[0][0] \n__________________________________________________________________________________________________\nactivation_57 (Activation) (None, 17, 17, 160) 0 batch_normalization_57[0][0] \n__________________________________________________________________________________________________\naverage_pooling2d_5 (AveragePoo (None, 17, 17, 768) 0 mixed5[0][0] \n__________________________________________________________________________________________________\nconv2d_50 (Conv2D) (None, 17, 17, 192) 147456 mixed5[0][0] \n__________________________________________________________________________________________________\nconv2d_53 (Conv2D) (None, 17, 17, 192) 215040 activation_52[0][0] \n__________________________________________________________________________________________________\nconv2d_58 (Conv2D) (None, 17, 17, 192) 215040 activation_57[0][0] \n__________________________________________________________________________________________________\nconv2d_59 (Conv2D) (None, 17, 17, 192) 147456 average_pooling2d_5[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_50 (BatchNo (None, 17, 17, 192) 576 conv2d_50[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_53 (BatchNo (None, 17, 17, 192) 576 conv2d_53[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_58 (BatchNo (None, 17, 17, 192) 576 conv2d_58[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_59 (BatchNo (None, 17, 17, 192) 576 conv2d_59[0][0] \n__________________________________________________________________________________________________\nactivation_50 (Activation) (None, 17, 17, 192) 0 batch_normalization_50[0][0] \n__________________________________________________________________________________________________\nactivation_53 (Activation) (None, 17, 17, 192) 0 batch_normalization_53[0][0] \n__________________________________________________________________________________________________\nactivation_58 (Activation) (None, 17, 17, 192) 0 batch_normalization_58[0][0] \n__________________________________________________________________________________________________\nactivation_59 (Activation) (None, 17, 17, 192) 0 batch_normalization_59[0][0] \n__________________________________________________________________________________________________\nmixed6 (Concatenate) (None, 17, 17, 768) 0 activation_50[0][0] \n activation_53[0][0] \n activation_58[0][0] \n activation_59[0][0] \n__________________________________________________________________________________________________\nconv2d_64 (Conv2D) (None, 17, 17, 192) 147456 mixed6[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_64 (BatchNo (None, 17, 17, 192) 576 conv2d_64[0][0] \n__________________________________________________________________________________________________\nactivation_64 (Activation) (None, 17, 17, 192) 0 batch_normalization_64[0][0] \n__________________________________________________________________________________________________\nconv2d_65 (Conv2D) (None, 17, 17, 192) 258048 activation_64[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_65 (BatchNo (None, 17, 17, 192) 576 conv2d_65[0][0] \n__________________________________________________________________________________________________\nactivation_65 (Activation) (None, 17, 17, 192) 0 batch_normalization_65[0][0] \n__________________________________________________________________________________________________\nconv2d_61 (Conv2D) (None, 17, 17, 192) 147456 mixed6[0][0] \n__________________________________________________________________________________________________\nconv2d_66 (Conv2D) (None, 17, 17, 192) 258048 activation_65[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_61 (BatchNo (None, 17, 17, 192) 576 conv2d_61[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_66 (BatchNo (None, 17, 17, 192) 576 conv2d_66[0][0] \n__________________________________________________________________________________________________\nactivation_61 (Activation) (None, 17, 17, 192) 0 batch_normalization_61[0][0] \n__________________________________________________________________________________________________\nactivation_66 (Activation) (None, 17, 17, 192) 0 batch_normalization_66[0][0] \n__________________________________________________________________________________________________\nconv2d_62 (Conv2D) (None, 17, 17, 192) 258048 activation_61[0][0] \n__________________________________________________________________________________________________\nconv2d_67 (Conv2D) (None, 17, 17, 192) 258048 activation_66[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_62 (BatchNo (None, 17, 17, 192) 576 conv2d_62[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_67 (BatchNo (None, 17, 17, 192) 576 conv2d_67[0][0] \n__________________________________________________________________________________________________\nactivation_62 (Activation) (None, 17, 17, 192) 0 batch_normalization_62[0][0] \n__________________________________________________________________________________________________\nactivation_67 (Activation) (None, 17, 17, 192) 0 batch_normalization_67[0][0] \n__________________________________________________________________________________________________\naverage_pooling2d_6 (AveragePoo (None, 17, 17, 768) 0 mixed6[0][0] \n__________________________________________________________________________________________________\nconv2d_60 (Conv2D) (None, 17, 17, 192) 147456 mixed6[0][0] \n__________________________________________________________________________________________________\nconv2d_63 (Conv2D) (None, 17, 17, 192) 258048 activation_62[0][0] \n__________________________________________________________________________________________________\nconv2d_68 (Conv2D) (None, 17, 17, 192) 258048 activation_67[0][0] \n__________________________________________________________________________________________________\nconv2d_69 (Conv2D) (None, 17, 17, 192) 147456 average_pooling2d_6[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_60 (BatchNo (None, 17, 17, 192) 576 conv2d_60[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_63 (BatchNo (None, 17, 17, 192) 576 conv2d_63[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_68 (BatchNo (None, 17, 17, 192) 576 conv2d_68[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_69 (BatchNo (None, 17, 17, 192) 576 conv2d_69[0][0] \n__________________________________________________________________________________________________\nactivation_60 (Activation) (None, 17, 17, 192) 0 batch_normalization_60[0][0] \n__________________________________________________________________________________________________\nactivation_63 (Activation) (None, 17, 17, 192) 0 batch_normalization_63[0][0] \n__________________________________________________________________________________________________\nactivation_68 (Activation) (None, 17, 17, 192) 0 batch_normalization_68[0][0] \n__________________________________________________________________________________________________\nactivation_69 (Activation) (None, 17, 17, 192) 0 batch_normalization_69[0][0] \n__________________________________________________________________________________________________\nmixed7 (Concatenate) (None, 17, 17, 768) 0 activation_60[0][0] \n activation_63[0][0] \n activation_68[0][0] \n activation_69[0][0] \n__________________________________________________________________________________________________\nconv2d_72 (Conv2D) (None, 17, 17, 192) 147456 mixed7[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_72 (BatchNo (None, 17, 17, 192) 576 conv2d_72[0][0] \n__________________________________________________________________________________________________\nactivation_72 (Activation) (None, 17, 17, 192) 0 batch_normalization_72[0][0] \n__________________________________________________________________________________________________\nconv2d_73 (Conv2D) (None, 17, 17, 192) 258048 activation_72[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_73 (BatchNo (None, 17, 17, 192) 576 conv2d_73[0][0] \n__________________________________________________________________________________________________\nactivation_73 (Activation) (None, 17, 17, 192) 0 batch_normalization_73[0][0] \n__________________________________________________________________________________________________\nconv2d_70 (Conv2D) (None, 17, 17, 192) 147456 mixed7[0][0] \n__________________________________________________________________________________________________\nconv2d_74 (Conv2D) (None, 17, 17, 192) 258048 activation_73[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_70 (BatchNo (None, 17, 17, 192) 576 conv2d_70[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_74 (BatchNo (None, 17, 17, 192) 576 conv2d_74[0][0] \n__________________________________________________________________________________________________\nactivation_70 (Activation) (None, 17, 17, 192) 0 batch_normalization_70[0][0] \n__________________________________________________________________________________________________\nactivation_74 (Activation) (None, 17, 17, 192) 0 batch_normalization_74[0][0] \n__________________________________________________________________________________________________\nconv2d_71 (Conv2D) (None, 8, 8, 320) 552960 activation_70[0][0] \n__________________________________________________________________________________________________\nconv2d_75 (Conv2D) (None, 8, 8, 192) 331776 activation_74[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_71 (BatchNo (None, 8, 8, 320) 960 conv2d_71[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_75 (BatchNo (None, 8, 8, 192) 576 conv2d_75[0][0] \n__________________________________________________________________________________________________\nactivation_71 (Activation) (None, 8, 8, 320) 0 batch_normalization_71[0][0] \n__________________________________________________________________________________________________\nactivation_75 (Activation) (None, 8, 8, 192) 0 batch_normalization_75[0][0] \n__________________________________________________________________________________________________\nmax_pooling2d_3 (MaxPooling2D) (None, 8, 8, 768) 0 mixed7[0][0] \n__________________________________________________________________________________________________\nmixed8 (Concatenate) (None, 8, 8, 1280) 0 activation_71[0][0] \n activation_75[0][0] \n max_pooling2d_3[0][0] \n__________________________________________________________________________________________________\nconv2d_80 (Conv2D) (None, 8, 8, 448) 573440 mixed8[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_80 (BatchNo (None, 8, 8, 448) 1344 conv2d_80[0][0] \n__________________________________________________________________________________________________\nactivation_80 (Activation) (None, 8, 8, 448) 0 batch_normalization_80[0][0] \n__________________________________________________________________________________________________\nconv2d_77 (Conv2D) (None, 8, 8, 384) 491520 mixed8[0][0] \n__________________________________________________________________________________________________\nconv2d_81 (Conv2D) (None, 8, 8, 384) 1548288 activation_80[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_77 (BatchNo (None, 8, 8, 384) 1152 conv2d_77[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_81 (BatchNo (None, 8, 8, 384) 1152 conv2d_81[0][0] \n__________________________________________________________________________________________________\nactivation_77 (Activation) (None, 8, 8, 384) 0 batch_normalization_77[0][0] \n__________________________________________________________________________________________________\nactivation_81 (Activation) (None, 8, 8, 384) 0 batch_normalization_81[0][0] \n__________________________________________________________________________________________________\nconv2d_78 (Conv2D) (None, 8, 8, 384) 442368 activation_77[0][0] \n__________________________________________________________________________________________________\nconv2d_79 (Conv2D) (None, 8, 8, 384) 442368 activation_77[0][0] \n__________________________________________________________________________________________________\nconv2d_82 (Conv2D) (None, 8, 8, 384) 442368 activation_81[0][0] \n__________________________________________________________________________________________________\nconv2d_83 (Conv2D) (None, 8, 8, 384) 442368 activation_81[0][0] \n__________________________________________________________________________________________________\naverage_pooling2d_7 (AveragePoo (None, 8, 8, 1280) 0 mixed8[0][0] \n__________________________________________________________________________________________________\nconv2d_76 (Conv2D) (None, 8, 8, 320) 409600 mixed8[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_78 (BatchNo (None, 8, 8, 384) 1152 conv2d_78[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_79 (BatchNo (None, 8, 8, 384) 1152 conv2d_79[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_82 (BatchNo (None, 8, 8, 384) 1152 conv2d_82[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_83 (BatchNo (None, 8, 8, 384) 1152 conv2d_83[0][0] \n__________________________________________________________________________________________________\nconv2d_84 (Conv2D) (None, 8, 8, 192) 245760 average_pooling2d_7[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_76 (BatchNo (None, 8, 8, 320) 960 conv2d_76[0][0] \n__________________________________________________________________________________________________\nactivation_78 (Activation) (None, 8, 8, 384) 0 batch_normalization_78[0][0] \n__________________________________________________________________________________________________\nactivation_79 (Activation) (None, 8, 8, 384) 0 batch_normalization_79[0][0] \n__________________________________________________________________________________________________\nactivation_82 (Activation) (None, 8, 8, 384) 0 batch_normalization_82[0][0] \n__________________________________________________________________________________________________\nactivation_83 (Activation) (None, 8, 8, 384) 0 batch_normalization_83[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_84 (BatchNo (None, 8, 8, 192) 576 conv2d_84[0][0] \n__________________________________________________________________________________________________\nactivation_76 (Activation) (None, 8, 8, 320) 0 batch_normalization_76[0][0] \n__________________________________________________________________________________________________\nmixed9_0 (Concatenate) (None, 8, 8, 768) 0 activation_78[0][0] \n activation_79[0][0] \n__________________________________________________________________________________________________\nconcatenate (Concatenate) (None, 8, 8, 768) 0 activation_82[0][0] \n activation_83[0][0] \n__________________________________________________________________________________________________\nactivation_84 (Activation) (None, 8, 8, 192) 0 batch_normalization_84[0][0] \n__________________________________________________________________________________________________\nmixed9 (Concatenate) (None, 8, 8, 2048) 0 activation_76[0][0] \n mixed9_0[0][0] \n concatenate[0][0] \n activation_84[0][0] \n__________________________________________________________________________________________________\nconv2d_89 (Conv2D) (None, 8, 8, 448) 917504 mixed9[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_89 (BatchNo (None, 8, 8, 448) 1344 conv2d_89[0][0] \n__________________________________________________________________________________________________\nactivation_89 (Activation) (None, 8, 8, 448) 0 batch_normalization_89[0][0] \n__________________________________________________________________________________________________\nconv2d_86 (Conv2D) (None, 8, 8, 384) 786432 mixed9[0][0] \n__________________________________________________________________________________________________\nconv2d_90 (Conv2D) (None, 8, 8, 384) 1548288 activation_89[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_86 (BatchNo (None, 8, 8, 384) 1152 conv2d_86[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_90 (BatchNo (None, 8, 8, 384) 1152 conv2d_90[0][0] \n__________________________________________________________________________________________________\nactivation_86 (Activation) (None, 8, 8, 384) 0 batch_normalization_86[0][0] \n__________________________________________________________________________________________________\nactivation_90 (Activation) (None, 8, 8, 384) 0 batch_normalization_90[0][0] \n__________________________________________________________________________________________________\nconv2d_87 (Conv2D) (None, 8, 8, 384) 442368 activation_86[0][0] \n__________________________________________________________________________________________________\nconv2d_88 (Conv2D) (None, 8, 8, 384) 442368 activation_86[0][0] \n__________________________________________________________________________________________________\nconv2d_91 (Conv2D) (None, 8, 8, 384) 442368 activation_90[0][0] \n__________________________________________________________________________________________________\nconv2d_92 (Conv2D) (None, 8, 8, 384) 442368 activation_90[0][0] \n__________________________________________________________________________________________________\naverage_pooling2d_8 (AveragePoo (None, 8, 8, 2048) 0 mixed9[0][0] \n__________________________________________________________________________________________________\nconv2d_85 (Conv2D) (None, 8, 8, 320) 655360 mixed9[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_87 (BatchNo (None, 8, 8, 384) 1152 conv2d_87[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_88 (BatchNo (None, 8, 8, 384) 1152 conv2d_88[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_91 (BatchNo (None, 8, 8, 384) 1152 conv2d_91[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_92 (BatchNo (None, 8, 8, 384) 1152 conv2d_92[0][0] \n__________________________________________________________________________________________________\nconv2d_93 (Conv2D) (None, 8, 8, 192) 393216 average_pooling2d_8[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_85 (BatchNo (None, 8, 8, 320) 960 conv2d_85[0][0] \n__________________________________________________________________________________________________\nactivation_87 (Activation) (None, 8, 8, 384) 0 batch_normalization_87[0][0] \n__________________________________________________________________________________________________\nactivation_88 (Activation) (None, 8, 8, 384) 0 batch_normalization_88[0][0] \n__________________________________________________________________________________________________\nactivation_91 (Activation) (None, 8, 8, 384) 0 batch_normalization_91[0][0] \n__________________________________________________________________________________________________\nactivation_92 (Activation) (None, 8, 8, 384) 0 batch_normalization_92[0][0] \n__________________________________________________________________________________________________\nbatch_normalization_93 (BatchNo (None, 8, 8, 192) 576 conv2d_93[0][0] \n__________________________________________________________________________________________________\nactivation_85 (Activation) (None, 8, 8, 320) 0 batch_normalization_85[0][0] \n__________________________________________________________________________________________________\nmixed9_1 (Concatenate) (None, 8, 8, 768) 0 activation_87[0][0] \n activation_88[0][0] \n__________________________________________________________________________________________________\nconcatenate_1 (Concatenate) (None, 8, 8, 768) 0 activation_91[0][0] \n activation_92[0][0] \n__________________________________________________________________________________________________\nactivation_93 (Activation) (None, 8, 8, 192) 0 batch_normalization_93[0][0] \n__________________________________________________________________________________________________\nmixed10 (Concatenate) (None, 8, 8, 2048) 0 activation_85[0][0] \n mixed9_1[0][0] \n concatenate_1[0][0] \n activation_93[0][0] \n__________________________________________________________________________________________________\navg_pool (GlobalAveragePooling2 (None, 2048) 0 mixed10[0][0] \n__________________________________________________________________________________________________\npredictions (Dense) (None, 1000) 2049000 avg_pool[0][0] \n==================================================================================================\nTotal params: 23,851,784\nTrainable params: 23,817,352\nNon-trainable params: 34,432\n__________________________________________________________________________________________________\n"
]
],
[
[
"preprocess_input 을 수행하면 입력 이미지도 변경이 되는것을 알 수 있습니다.",
"_____no_output_____"
]
],
[
[
"# pip install pillow\n%matplotlib inline\n\nimport tensorflow.keras as keras\nfrom IPython.display import display\nimport matplotlib.pyplot as plt\nimport numpy as np",
"_____no_output_____"
],
[
"pil_img = keras.preprocessing.image.load_img('cat.jpg', target_size=(299, 299))\n\nprint(type(pil_img))\n\ndisplay(pil_img)",
"<class 'PIL.Image.Image'>\n"
],
[
"image = keras.preprocessing.image.img_to_array(pil_img)\nprint('image', image.shape, image.dtype)\nprint(image)",
"image (299, 299, 3) float32\n[[[12. 25. 7.]\n [11. 24. 4.]\n [12. 25. 5.]\n ...\n [18. 34. 7.]\n [23. 39. 12.]\n [20. 36. 9.]]\n\n [[12. 25. 7.]\n [11. 24. 6.]\n [ 9. 22. 4.]\n ...\n [20. 36. 7.]\n [21. 35. 12.]\n [21. 35. 12.]]\n\n [[12. 25. 8.]\n [12. 25. 7.]\n [12. 25. 7.]\n ...\n [21. 35. 9.]\n [23. 37. 12.]\n [22. 36. 11.]]\n\n ...\n\n [[81. 66. 43.]\n [82. 70. 46.]\n [84. 69. 46.]\n ...\n [30. 45. 22.]\n [29. 44. 21.]\n [33. 47. 24.]]\n\n [[82. 66. 43.]\n [85. 69. 46.]\n [84. 68. 43.]\n ...\n [33. 47. 21.]\n [32. 46. 21.]\n [33. 47. 22.]]\n\n [[83. 65. 41.]\n [86. 70. 47.]\n [82. 67. 44.]\n ...\n [31. 47. 21.]\n [29. 45. 19.]\n [30. 44. 19.]]]\n"
],
[
"images = np.expand_dims(image, axis=0)\nprint('images', images.shape, images.dtype)\nprint(images)",
"images (1, 299, 299, 3) float32\n[[[[12. 25. 7.]\n [11. 24. 4.]\n [12. 25. 5.]\n ...\n [18. 34. 7.]\n [23. 39. 12.]\n [20. 36. 9.]]\n\n [[12. 25. 7.]\n [11. 24. 6.]\n [ 9. 22. 4.]\n ...\n [20. 36. 7.]\n [21. 35. 12.]\n [21. 35. 12.]]\n\n [[12. 25. 8.]\n [12. 25. 7.]\n [12. 25. 7.]\n ...\n [21. 35. 9.]\n [23. 37. 12.]\n [22. 36. 11.]]\n\n ...\n\n [[81. 66. 43.]\n [82. 70. 46.]\n [84. 69. 46.]\n ...\n [30. 45. 22.]\n [29. 44. 21.]\n [33. 47. 24.]]\n\n [[82. 66. 43.]\n [85. 69. 46.]\n [84. 68. 43.]\n ...\n [33. 47. 21.]\n [32. 46. 21.]\n [33. 47. 22.]]\n\n [[83. 65. 41.]\n [86. 70. 47.]\n [82. 67. 44.]\n ...\n [31. 47. 21.]\n [29. 45. 19.]\n [30. 44. 19.]]]]\n"
],
[
"plt.imshow(images[0].astype('uint8'))\nplt.show()",
"_____no_output_____"
],
[
"x = preprocess_input(images)\n\nprint('x', x.shape, x.dtype)\nprint(x)",
"x (1, 299, 299, 3) float32\n[[[[-0.90588236 -0.8039216 -0.94509804]\n [-0.9137255 -0.8117647 -0.96862745]\n [-0.90588236 -0.8039216 -0.9607843 ]\n ...\n [-0.85882354 -0.73333335 -0.94509804]\n [-0.81960785 -0.69411767 -0.90588236]\n [-0.84313726 -0.7176471 -0.92941177]]\n\n [[-0.90588236 -0.8039216 -0.94509804]\n [-0.9137255 -0.8117647 -0.9529412 ]\n [-0.92941177 -0.827451 -0.96862745]\n ...\n [-0.84313726 -0.7176471 -0.94509804]\n [-0.8352941 -0.7254902 -0.90588236]\n [-0.8352941 -0.7254902 -0.90588236]]\n\n [[-0.90588236 -0.8039216 -0.9372549 ]\n [-0.90588236 -0.8039216 -0.94509804]\n [-0.90588236 -0.8039216 -0.94509804]\n ...\n [-0.8352941 -0.7254902 -0.92941177]\n [-0.81960785 -0.70980394 -0.90588236]\n [-0.827451 -0.7176471 -0.9137255 ]]\n\n ...\n\n [[-0.36470586 -0.4823529 -0.6627451 ]\n [-0.35686272 -0.45098037 -0.6392157 ]\n [-0.34117645 -0.4588235 -0.6392157 ]\n ...\n [-0.7647059 -0.64705884 -0.827451 ]\n [-0.77254903 -0.654902 -0.8352941 ]\n [-0.7411765 -0.6313726 -0.8117647 ]]\n\n [[-0.35686272 -0.4823529 -0.6627451 ]\n [-0.3333333 -0.4588235 -0.6392157 ]\n [-0.34117645 -0.46666664 -0.6627451 ]\n ...\n [-0.7411765 -0.6313726 -0.8352941 ]\n [-0.7490196 -0.6392157 -0.8352941 ]\n [-0.7411765 -0.6313726 -0.827451 ]]\n\n [[-0.3490196 -0.49019605 -0.6784314 ]\n [-0.32549018 -0.45098037 -0.6313726 ]\n [-0.35686272 -0.47450978 -0.654902 ]\n ...\n [-0.75686276 -0.6313726 -0.8352941 ]\n [-0.77254903 -0.64705884 -0.8509804 ]\n [-0.7647059 -0.654902 -0.8509804 ]]]]\n"
],
[
"plt.imshow(images[0].astype('uint8'))\nplt.show()\nplt.imshow(x[0].astype('uint8'))\nplt.show()",
"_____no_output_____"
],
[
"pred = model.predict(x)\n# decode the results into a list of tuples (class, description, probability)\nprint('prediction')\nprint(decode_predictions(pred, top=3))",
"prediction\nDownloading data from https://storage.googleapis.com/download.tensorflow.org/data/imagenet_class_index.json\n40960/35363 [==================================] - 0s 1us/step\n[[('n02123045', 'tabby', 0.30431667), ('n02123159', 'tiger_cat', 0.29704612), ('n02124075', 'Egyptian_cat', 0.21344033)]]\n"
]
]
] |
[
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ab85d534a4ee10fe12acb2602d9cb6236b9d72b
| 158,357 |
ipynb
|
Jupyter Notebook
|
examples/models/azure_aks_deep_mnist/azure_aks_deep_mnist.ipynb
|
alexlatchford/seldon-core
|
ddc446a5828901e2896a9f1aa8cb8dff4db81c52
|
[
"Apache-2.0"
] | null | null | null |
examples/models/azure_aks_deep_mnist/azure_aks_deep_mnist.ipynb
|
alexlatchford/seldon-core
|
ddc446a5828901e2896a9f1aa8cb8dff4db81c52
|
[
"Apache-2.0"
] | null | null | null |
examples/models/azure_aks_deep_mnist/azure_aks_deep_mnist.ipynb
|
alexlatchford/seldon-core
|
ddc446a5828901e2896a9f1aa8cb8dff4db81c52
|
[
"Apache-2.0"
] | null | null | null | 43.48078 | 18,784 | 0.507013 |
[
[
[
"# Azure Kubernetes Service (AKS) Deep MNIST\nIn this example we will deploy a tensorflow MNIST model in the Azure Kubernetes Service (AKS).\n\nThis tutorial will break down in the following sections:\n\n1) Train a tensorflow model to predict mnist locally\n\n2) Containerise the tensorflow model with our docker utility\n\n3) Send some data to the docker model to test it\n\n4) Install and configure Azure tools to interact with your cluster\n\n5) Use the Azure tools to create and setup AKS cluster with Seldon\n\n6) Push and run docker image through the Azure Container Registry\n\n7) Test our Elastic Kubernetes deployment by sending some data\n\n#### Let's get started! 🚀🔥\n\n## Dependencies:\n\n* Helm v2.13.1+\n* A Kubernetes cluster running v1.13 or above (minkube / docker-for-windows work well if enough RAM)\n* kubectl v1.14+\n* az CLI v2.0.66+\n* Python 3.6+\n* Python DEV requirements\n",
"_____no_output_____"
],
[
"## 1) Train a tensorflow model to predict mnist locally\nWe will load the mnist images, together with their labels, and then train a tensorflow model to predict the right labels",
"_____no_output_____"
]
],
[
[
"from tensorflow.examples.tutorials.mnist import input_data\nmnist = input_data.read_data_sets(\"MNIST_data/\", one_hot = True)\nimport tensorflow as tf\n\nif __name__ == '__main__':\n \n x = tf.placeholder(tf.float32, [None,784], name=\"x\")\n\n W = tf.Variable(tf.zeros([784,10]))\n b = tf.Variable(tf.zeros([10]))\n\n y = tf.nn.softmax(tf.matmul(x,W) + b, name=\"y\")\n\n y_ = tf.placeholder(tf.float32, [None, 10])\n\n cross_entropy = tf.reduce_mean(-tf.reduce_sum(y_ * tf.log(y), reduction_indices=[1]))\n\n train_step = tf.train.GradientDescentOptimizer(0.5).minimize(cross_entropy)\n\n init = tf.initialize_all_variables()\n\n sess = tf.Session()\n sess.run(init)\n\n for i in range(1000):\n batch_xs, batch_ys = mnist.train.next_batch(100)\n sess.run(train_step, feed_dict={x: batch_xs, y_: batch_ys})\n\n correct_prediction = tf.equal(tf.argmax(y,1), tf.argmax(y_,1))\n accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))\n print(sess.run(accuracy, feed_dict = {x: mnist.test.images, y_:mnist.test.labels}))\n\n saver = tf.train.Saver()\n\n saver.save(sess, \"model/deep_mnist_model\")",
"WARNING:tensorflow:From <ipython-input-1-559b63ab8b48>:2: read_data_sets (from tensorflow.contrib.learn.python.learn.datasets.mnist) is deprecated and will be removed in a future version.\nInstructions for updating:\nPlease use alternatives such as official/mnist/dataset.py from tensorflow/models.\nWARNING:tensorflow:From /home/alejandro/miniconda3/envs/reddit-classification/lib/python3.7/site-packages/tensorflow/contrib/learn/python/learn/datasets/mnist.py:260: maybe_download (from tensorflow.contrib.learn.python.learn.datasets.base) is deprecated and will be removed in a future version.\nInstructions for updating:\nPlease write your own downloading logic.\nWARNING:tensorflow:From /home/alejandro/miniconda3/envs/reddit-classification/lib/python3.7/site-packages/tensorflow/contrib/learn/python/learn/datasets/base.py:252: _internal_retry.<locals>.wrap.<locals>.wrapped_fn (from tensorflow.contrib.learn.python.learn.datasets.base) is deprecated and will be removed in a future version.\nInstructions for updating:\nPlease use urllib or similar directly.\nSuccessfully downloaded train-images-idx3-ubyte.gz 9912422 bytes.\nWARNING:tensorflow:From /home/alejandro/miniconda3/envs/reddit-classification/lib/python3.7/site-packages/tensorflow/contrib/learn/python/learn/datasets/mnist.py:262: extract_images (from tensorflow.contrib.learn.python.learn.datasets.mnist) is deprecated and will be removed in a future version.\nInstructions for updating:\nPlease use tf.data to implement this functionality.\nExtracting MNIST_data/train-images-idx3-ubyte.gz\nSuccessfully downloaded train-labels-idx1-ubyte.gz 28881 bytes.\nWARNING:tensorflow:From /home/alejandro/miniconda3/envs/reddit-classification/lib/python3.7/site-packages/tensorflow/contrib/learn/python/learn/datasets/mnist.py:267: extract_labels (from tensorflow.contrib.learn.python.learn.datasets.mnist) is deprecated and will be removed in a future version.\nInstructions for updating:\nPlease use tf.data to implement this functionality.\nExtracting MNIST_data/train-labels-idx1-ubyte.gz\nWARNING:tensorflow:From /home/alejandro/miniconda3/envs/reddit-classification/lib/python3.7/site-packages/tensorflow/contrib/learn/python/learn/datasets/mnist.py:110: dense_to_one_hot (from tensorflow.contrib.learn.python.learn.datasets.mnist) is deprecated and will be removed in a future version.\nInstructions for updating:\nPlease use tf.one_hot on tensors.\nSuccessfully downloaded t10k-images-idx3-ubyte.gz 1648877 bytes.\nExtracting MNIST_data/t10k-images-idx3-ubyte.gz\nSuccessfully downloaded t10k-labels-idx1-ubyte.gz 4542 bytes.\nExtracting MNIST_data/t10k-labels-idx1-ubyte.gz\nWARNING:tensorflow:From /home/alejandro/miniconda3/envs/reddit-classification/lib/python3.7/site-packages/tensorflow/contrib/learn/python/learn/datasets/mnist.py:290: DataSet.__init__ (from tensorflow.contrib.learn.python.learn.datasets.mnist) is deprecated and will be removed in a future version.\nInstructions for updating:\nPlease use alternatives such as official/mnist/dataset.py from tensorflow/models.\nWARNING:tensorflow:From /home/alejandro/miniconda3/envs/reddit-classification/lib/python3.7/site-packages/tensorflow/python/framework/op_def_library.py:263: colocate_with (from tensorflow.python.framework.ops) is deprecated and will be removed in a future version.\nInstructions for updating:\nColocations handled automatically by placer.\nWARNING:tensorflow:From /home/alejandro/miniconda3/envs/reddit-classification/lib/python3.7/site-packages/tensorflow/python/ops/math_ops.py:3066: to_int32 (from tensorflow.python.ops.math_ops) is deprecated and will be removed in a future version.\nInstructions for updating:\nUse tf.cast instead.\nWARNING:tensorflow:From /home/alejandro/miniconda3/envs/reddit-classification/lib/python3.7/site-packages/tensorflow/python/util/tf_should_use.py:193: initialize_all_variables (from tensorflow.python.ops.variables) is deprecated and will be removed after 2017-03-02.\nInstructions for updating:\nUse `tf.global_variables_initializer` instead.\n0.915\n"
]
],
[
[
"## 2) Containerise the tensorflow model with our docker utility",
"_____no_output_____"
],
[
"First you need to make sure that you have added the .s2i/environment configuration file in this folder with the following content:",
"_____no_output_____"
]
],
[
[
"!cat .s2i/environment",
"MODEL_NAME=DeepMnist\r\nAPI_TYPE=REST\r\nSERVICE_TYPE=MODEL\r\nPERSISTENCE=0\r\n"
]
],
[
[
"Now we can build a docker image named \"deep-mnist\" with the tag 0.1",
"_____no_output_____"
]
],
[
[
"!s2i build . seldonio/seldon-core-s2i-python36:0.10 deep-mnist:0.1",
"---> Installing application source...\n---> Installing dependencies ...\nLooking in links: /whl\nRequirement already satisfied: tensorflow>=1.12.0 in /usr/local/lib/python3.6/site-packages (from -r requirements.txt (line 1)) (1.13.1)\nRequirement already satisfied: keras-preprocessing>=1.0.5 in /usr/local/lib/python3.6/site-packages (from tensorflow>=1.12.0->-r requirements.txt (line 1)) (1.0.9)\nRequirement already satisfied: gast>=0.2.0 in /usr/local/lib/python3.6/site-packages (from tensorflow>=1.12.0->-r requirements.txt (line 1)) (0.2.2)\nRequirement already satisfied: absl-py>=0.1.6 in /usr/local/lib/python3.6/site-packages (from tensorflow>=1.12.0->-r requirements.txt (line 1)) (0.7.1)\nRequirement already satisfied: astor>=0.6.0 in /usr/local/lib/python3.6/site-packages (from tensorflow>=1.12.0->-r requirements.txt (line 1)) (0.7.1)\nRequirement already satisfied: keras-applications>=1.0.6 in /usr/local/lib/python3.6/site-packages (from tensorflow>=1.12.0->-r requirements.txt (line 1)) (1.0.7)\nRequirement already satisfied: six>=1.10.0 in /usr/local/lib/python3.6/site-packages (from tensorflow>=1.12.0->-r requirements.txt (line 1)) (1.12.0)\nRequirement already satisfied: termcolor>=1.1.0 in /usr/local/lib/python3.6/site-packages (from tensorflow>=1.12.0->-r requirements.txt (line 1)) (1.1.0)\nRequirement already satisfied: grpcio>=1.8.6 in /usr/local/lib/python3.6/site-packages (from tensorflow>=1.12.0->-r requirements.txt (line 1)) (1.19.0)\nRequirement already satisfied: wheel>=0.26 in /usr/local/lib/python3.6/site-packages (from tensorflow>=1.12.0->-r requirements.txt (line 1)) (0.33.1)\nRequirement already satisfied: tensorboard<1.14.0,>=1.13.0 in /usr/local/lib/python3.6/site-packages (from tensorflow>=1.12.0->-r requirements.txt (line 1)) (1.13.1)\nRequirement already satisfied: numpy>=1.13.3 in /usr/local/lib/python3.6/site-packages (from tensorflow>=1.12.0->-r requirements.txt (line 1)) (1.16.2)\nRequirement already satisfied: protobuf>=3.6.1 in /usr/local/lib/python3.6/site-packages (from tensorflow>=1.12.0->-r requirements.txt (line 1)) (3.7.0)\nRequirement already satisfied: tensorflow-estimator<1.14.0rc0,>=1.13.0 in /usr/local/lib/python3.6/site-packages (from tensorflow>=1.12.0->-r requirements.txt (line 1)) (1.13.0)\nRequirement already satisfied: h5py in /usr/local/lib/python3.6/site-packages (from keras-applications>=1.0.6->tensorflow>=1.12.0->-r requirements.txt (line 1)) (2.9.0)\nRequirement already satisfied: markdown>=2.6.8 in /usr/local/lib/python3.6/site-packages (from tensorboard<1.14.0,>=1.13.0->tensorflow>=1.12.0->-r requirements.txt (line 1)) (3.0.1)\nRequirement already satisfied: werkzeug>=0.11.15 in /usr/local/lib/python3.6/site-packages (from tensorboard<1.14.0,>=1.13.0->tensorflow>=1.12.0->-r requirements.txt (line 1)) (0.15.0)\nRequirement already satisfied: setuptools in /usr/local/lib/python3.6/site-packages (from protobuf>=3.6.1->tensorflow>=1.12.0->-r requirements.txt (line 1)) (40.8.0)\nRequirement already satisfied: mock>=2.0.0 in /usr/local/lib/python3.6/site-packages (from tensorflow-estimator<1.14.0rc0,>=1.13.0->tensorflow>=1.12.0->-r requirements.txt (line 1)) (2.0.0)\nRequirement already satisfied: pbr>=0.11 in /usr/local/lib/python3.6/site-packages (from mock>=2.0.0->tensorflow-estimator<1.14.0rc0,>=1.13.0->tensorflow>=1.12.0->-r requirements.txt (line 1)) (5.1.3)\nUrl '/whl' is ignored. It is either a non-existing path or lacks a specific scheme.\nYou are using pip version 19.0.3, however version 19.1.1 is available.\nYou should consider upgrading via the 'pip install --upgrade pip' command.\nBuild completed successfully\n"
]
],
[
[
"## 3) Send some data to the docker model to test it\nWe first run the docker image we just created as a container called \"mnist_predictor\"",
"_____no_output_____"
]
],
[
[
"!docker run --name \"mnist_predictor\" -d --rm -p 5000:5000 deep-mnist:0.1",
"9087047e368ac8f285e1f742704b4c0c7bceac7d29ee90b3b0a6ef2d61ebd15c\r\n"
]
],
[
[
"Send some random features that conform to the contract",
"_____no_output_____"
]
],
[
[
"import matplotlib.pyplot as plt\nimport numpy as np\n# This is the variable that was initialised at the beginning of the file\ni = [0]\nx = mnist.test.images[i]\ny = mnist.test.labels[i]\nplt.imshow(x.reshape((28, 28)), cmap='gray')\nplt.show()\nprint(\"Expected label: \", np.sum(range(0,10) * y), \". One hot encoding: \", y)",
"_____no_output_____"
],
[
"from seldon_core.seldon_client import SeldonClient\nimport math\nimport numpy as np\n\n# We now test the REST endpoint expecting the same result\nendpoint = \"0.0.0.0:5000\"\nbatch = x\npayload_type = \"ndarray\"\n\nsc = SeldonClient(microservice_endpoint=endpoint)\n\n# We use the microservice, instead of the \"predict\" function\nclient_prediction = sc.microservice(\n data=batch,\n method=\"predict\",\n payload_type=payload_type,\n names=[\"tfidf\"])\n\nfor proba, label in zip(client_prediction.response.data.ndarray.values[0].list_value.ListFields()[0][1], range(0,10)):\n print(f\"LABEL {label}:\\t {proba.number_value*100:6.4f} %\")",
"LABEL 0:\t 0.0064 %\nLABEL 1:\t 0.0000 %\nLABEL 2:\t 0.0155 %\nLABEL 3:\t 0.2862 %\nLABEL 4:\t 0.0003 %\nLABEL 5:\t 0.0027 %\nLABEL 6:\t 0.0000 %\nLABEL 7:\t 99.6643 %\nLABEL 8:\t 0.0020 %\nLABEL 9:\t 0.0227 %\n"
],
[
"!docker rm mnist_predictor --force",
"mnist_predictor\r\n"
]
],
[
[
"## 4) Install and configure Azure tools ",
"_____no_output_____"
],
[
"First we install the azure cli - follow specific instructions at https://docs.microsoft.com/en-us/cli/azure/install-azure-cli?view=azure-cli-latest",
"_____no_output_____"
]
],
[
[
"!curl -sL https://aka.ms/InstallAzureCLIDeb | sudo bash",
"_____no_output_____"
]
],
[
[
"#### Configure the azure CLI so it can talk to your server \n(if you are getting issues, make sure you have the permmissions to create clusters)\n\nYou must run this through a terminal and follow the instructions:\n```\naz login\n```\n\nOnce you are logged in, we can create our cluster. Run the following command, it may take a while so feel free to get a ☕.",
"_____no_output_____"
]
],
[
[
"%%bash \n# We'll create a resource group\naz group create --name SeldonResourceGroup --location westus\n# Now we create the cluster\naz aks create \\\n --resource-group SeldonResourceGroup \\\n --name SeldonCluster \\\n --node-count 1 \\\n --enable-addons monitoring \\\n --generate-ssh-keys\n --kubernetes-version 1.13.5",
"_____no_output_____"
]
],
[
[
"Once it's created we can authenticate our local `kubectl` to make sure we can talk to the azure cluster:",
"_____no_output_____"
]
],
[
[
"!az aks get-credentials --resource-group SeldonResourceGroup --name SeldonCluster",
"_____no_output_____"
]
],
[
[
"And now we can check that this has been successful by making sure that our `kubectl` context is pointing to the cluster:",
"_____no_output_____"
]
],
[
[
"!kubectl config get-contexts",
"_____no_output_____"
]
],
[
[
"## Install Seldon Core",
"_____no_output_____"
],
[
"### Before we install seldon core, we need to install HELM\nFor that, we need to create a ClusterRoleBinding for us, a ServiceAccount, and then a RoleBinding",
"_____no_output_____"
]
],
[
[
"!kubectl create clusterrolebinding kube-system-cluster-admin --clusterrole=cluster-admin --serviceaccount=kube-system:default",
"clusterrolebinding.rbac.authorization.k8s.io/kube-system-cluster-admin created\r\n"
],
[
"!kubectl create serviceaccount tiller --namespace kube-system",
"serviceaccount/tiller created\r\n"
],
[
"!kubectl apply -f tiller-role-binding.yaml",
"clusterrolebinding.rbac.authorization.k8s.io/tiller-role-binding created\r\n"
]
],
[
[
"### Once that is set-up we can install Tiller",
"_____no_output_____"
]
],
[
[
"!helm repo update",
"_____no_output_____"
],
[
"!helm init --service-account tiller",
"$HELM_HOME has been configured at /home/alejandro/.helm.\n\nTiller (the Helm server-side component) has been installed into your Kubernetes Cluster.\n\nPlease note: by default, Tiller is deployed with an insecure 'allow unauthenticated users' policy.\nTo prevent this, run `helm init` with the --tiller-tls-verify flag.\nFor more information on securing your installation see: https://docs.helm.sh/using_helm/#securing-your-helm-installation\nHappy Helming!\n"
],
[
"# Wait until Tiller finishes\n!kubectl rollout status deploy/tiller-deploy -n kube-system",
"deployment \"tiller-deploy\" successfully rolled out\r\n"
]
],
[
[
"### Now we can install SELDON. \nWe first start with the custom resource definitions (CRDs)",
"_____no_output_____"
]
],
[
[
"!helm install seldon-core-operator --name seldon-core-operator --repo https://storage.googleapis.com/seldon-charts",
"NAME: seldon-core-operator\nLAST DEPLOYED: Thu Jun 6 12:03:45 2019\nNAMESPACE: default\nSTATUS: DEPLOYED\n\nRESOURCES:\n==> v1/ClusterRole\nNAME AGE\nseldon-operator-manager-role 3s\n\n==> v1/ClusterRoleBinding\nNAME AGE\nseldon-operator-manager-rolebinding 3s\n\n==> v1/Pod(related)\nNAME READY STATUS RESTARTS AGE\nseldon-operator-controller-manager-0 1/1 Running 0 3s\n\n==> v1/Secret\nNAME TYPE DATA AGE\nseldon-operator-webhook-server-secret Opaque 4 3s\n\n==> v1/Service\nNAME TYPE CLUSTER-IP EXTERNAL-IP PORT(S) AGE\nseldon-operator-controller-manager-service ClusterIP 10.0.224.128 <none> 443/TCP 3s\n\n==> v1/StatefulSet\nNAME READY AGE\nseldon-operator-controller-manager 1/1 3s\n\n==> v1beta1/CustomResourceDefinition\nNAME AGE\nseldondeployments.machinelearning.seldon.io 3s\n\n\nNOTES:\nNOTES: TODO\n\n\n"
]
],
[
[
"And confirm they are running by getting the pods:",
"_____no_output_____"
]
],
[
[
"!kubectl rollout status statefulset.apps/seldon-operator-controller-manager -n seldon-system",
"partitioned roll out complete: 1 new pods have been updated...\r\n"
]
],
[
[
"### Now we set-up the ingress\nThis will allow you to reach the Seldon models from outside the kubernetes cluster. \n\nIn EKS it automatically creates an Elastic Load Balancer, which you can configure from the EC2 Console",
"_____no_output_____"
]
],
[
[
"!helm install stable/ambassador --name ambassador --set crds.keep=false",
"Error: release ambassador failed: serviceaccounts \"ambassador\" already exists\r\n"
]
],
[
[
"And let's wait until it's fully deployed",
"_____no_output_____"
]
],
[
[
"!kubectl rollout status deployment.apps/ambassador",
"deployment \"ambassador\" successfully rolled out\r\n"
]
],
[
[
"## Push docker image\nIn order for the EKS seldon deployment to access the image we just built, we need to push it to the Azure Container Registry (ACR) - you can check if it's been successfully created in the dashboard https://portal.azure.com/#blade/HubsExtension/BrowseResourceBlade/resourceType/Microsoft.ContainerRegistry%2Fregistries\n\nIf you have any issues please follow the official Azure documentation: https://docs.microsoft.com/en-us/azure/container-registry/container-registry-get-started-azure-cli",
"_____no_output_____"
],
[
"### First we create a registry\nMake sure you keep the `loginServer` value in the output dictionary as we'll use it below.",
"_____no_output_____"
]
],
[
[
"!az acr create --resource-group SeldonResourceGroup --name SeldonContainerRegistry --sku Basic",
"\u001b[K{- Finished ..\n \"adminUserEnabled\": false,\n \"creationDate\": \"2019-06-06T10:51:55.288108+00:00\",\n \"id\": \"/subscriptions/df7969cc-8033-4c83-b027-14c0424f039d/resourceGroups/KlawClusterResourceGroup/providers/Microsoft.ContainerRegistry/registries/SeldonContainerRegistry\",\n \"location\": \"westus\",\n \"loginServer\": \"seldoncontainerregistry.azurecr.io\",\n \"name\": \"SeldonContainerRegistry\",\n \"networkRuleSet\": null,\n \"provisioningState\": \"Succeeded\",\n \"resourceGroup\": \"KlawClusterResourceGroup\",\n \"sku\": {\n \"name\": \"Basic\",\n \"tier\": \"Basic\"\n },\n \"status\": null,\n \"storageAccount\": null,\n \"tags\": {},\n \"type\": \"Microsoft.ContainerRegistry/registries\"\n}\n\u001b[0m"
]
],
[
[
"### Make sure your local docker instance has access to the registry",
"_____no_output_____"
]
],
[
[
"!az acr login --name SeldonContainerRegistry",
"Login Succeeded\nWARNING! Your password will be stored unencrypted in /home/alejandro/.docker/config.json.\nConfigure a credential helper to remove this warning. See\nhttps://docs.docker.com/engine/reference/commandline/login/#credentials-store\n\n"
]
],
[
[
"### Now prepare docker image\nWe need to first tag the docker image before we can push it.\n\nNOTE: if you named your registry different make sure you change the value of `seldoncontainerregistry.azurecr.io`",
"_____no_output_____"
]
],
[
[
"!docker tag deep-mnist:0.1 seldoncontainerregistry.azurecr.io/deep-mnist:0.1",
"_____no_output_____"
]
],
[
[
"### And push the image\n\nNOTE: if you named your registry different make sure you change the value of `seldoncontainerregistry.azurecr.io`",
"_____no_output_____"
]
],
[
[
"!docker push seldoncontainerregistry.azurecr.io/deep-mnist:0.1",
"The push refers to repository [seldoncontainerregistry.azurecr.io/deep-mnist]\n\n\u001b[1Bb4fe3076: Preparing \n\u001b[1Be4a983d1: Preparing \n\u001b[1B74b2c556: Preparing \n\u001b[1B9472b523: Preparing \n\u001b[1Ba2a7ea60: Preparing \n\u001b[1Beddb328a: Preparing \n\u001b[1B1393f8e7: Preparing \n\u001b[1B67d6e30e: Preparing \n\u001b[1Bf19da2c9: Preparing \n\u001b[1B9ec591c4: Preparing \n\u001b[1B32b1ff99: Preparing \n\u001b[1B64f96dbc: Preparing \n\u001b[1Be6d76fd9: Preparing \n\u001b[1B11a84ad4: Preparing \n"
]
],
[
[
"## Running the Model\nWe will now run the model. As you can see we have a placeholder `\"REPLACE_FOR_IMAGE_AND_TAG\"`, which we'll replace to point to our registry.\n\nLet's first have a look at the file we'll be using to trigger the model:",
"_____no_output_____"
]
],
[
[
"!cat deep_mnist.json",
"{\r\n \"apiVersion\": \"machinelearning.seldon.io/v1alpha2\",\r\n \"kind\": \"SeldonDeployment\",\r\n \"metadata\": {\r\n \"labels\": {\r\n \"app\": \"seldon\"\r\n },\r\n \"name\": \"deep-mnist\"\r\n },\r\n \"spec\": {\r\n \"annotations\": {\r\n \"project_name\": \"Tensorflow MNIST\",\r\n \"deployment_version\": \"v1\"\r\n },\r\n \"name\": \"deep-mnist\",\r\n \"oauth_key\": \"oauth-key\",\r\n \"oauth_secret\": \"oauth-secret\",\r\n \"predictors\": [\r\n {\r\n \"componentSpecs\": [{\r\n \"spec\": {\r\n \"containers\": [\r\n {\r\n \"image\": \"REPLACE_FOR_IMAGE_AND_TAG\",\r\n \"imagePullPolicy\": \"IfNotPresent\",\r\n \"name\": \"classifier\",\r\n \"resources\": {\r\n \"requests\": {\r\n \"memory\": \"1Mi\"\r\n }\r\n }\r\n }\r\n ],\r\n \"terminationGracePeriodSeconds\": 20\r\n }\r\n }],\r\n \"graph\": {\r\n \"children\": [],\r\n \"name\": \"classifier\",\r\n \"endpoint\": {\r\n\t\t\t\"type\" : \"REST\"\r\n\t\t },\r\n \"type\": \"MODEL\"\r\n },\r\n \"name\": \"single-model\",\r\n \"replicas\": 1,\r\n\t\t\"annotations\": {\r\n\t\t \"predictor_version\" : \"v1\"\r\n\t\t}\r\n }\r\n ]\r\n }\r\n}\r\n"
]
],
[
[
"Now let's trigger seldon to run the model.\n\n### Run the deployment in your cluster\n\nNOTE: In order for this to work you need to make sure that your cluster has the permissions to pull the images. You can do this by:\n\n1) Go into the Azure Container Registry\n\n2) Select the SeldonContainerRegistry you created\n\n3) Click on \"Add a role assignment\"\n\n4) Select the AcrPull role\n\n5) Select service principle\n\n6) Find the SeldonCluster\n\n7) Wait until the role has been added\n\nWe basically have a yaml file, where we want to replace the value \"REPLACE_FOR_IMAGE_AND_TAG\" for the image you pushed",
"_____no_output_____"
]
],
[
[
"%%bash\n# Change accordingly if your registry is called differently\nsed 's|REPLACE_FOR_IMAGE_AND_TAG|seldoncontainerregistry.azurecr.io/deep-mnist:0.1|g' deep_mnist.json | kubectl apply -f -",
"seldondeployment.machinelearning.seldon.io/deep-mnist created\n"
]
],
[
[
"And let's check that it's been created.\n\nYou should see an image called \"deep-mnist-single-model...\".\n\nWe'll wait until STATUS changes from \"ContainerCreating\" to \"Running\"",
"_____no_output_____"
]
],
[
[
"!kubectl get pods",
"_____no_output_____"
]
],
[
[
"## Test the model\nNow we can test the model, let's first find out what is the URL that we'll have to use:",
"_____no_output_____"
]
],
[
[
"!kubectl get svc ambassador -o jsonpath='{.status.loadBalancer.ingress[0].ip}' ",
"52.160.64.65"
]
],
[
[
"We'll use a random example from our dataset",
"_____no_output_____"
]
],
[
[
"import matplotlib.pyplot as plt\n# This is the variable that was initialised at the beginning of the file\ni = [0]\nx = mnist.test.images[i]\ny = mnist.test.labels[i]\nplt.imshow(x.reshape((28, 28)), cmap='gray')\nplt.show()\nprint(\"Expected label: \", np.sum(range(0,10) * y), \". One hot encoding: \", y)",
"_____no_output_____"
]
],
[
[
"We can now add the URL above to send our request:",
"_____no_output_____"
]
],
[
[
"from seldon_core.seldon_client import SeldonClient\nimport math\nimport numpy as np\n\nhost = \"52.160.64.65\"\nport = \"80\" # Make sure you use the port above\nbatch = x\npayload_type = \"ndarray\"\n\nsc = SeldonClient(\n gateway=\"ambassador\", \n ambassador_endpoint=host + \":\" + port,\n namespace=\"default\",\n oauth_key=\"oauth-key\", \n oauth_secret=\"oauth-secret\")\n\nclient_prediction = sc.predict(\n data=batch, \n deployment_name=\"deep-mnist\",\n names=[\"text\"],\n payload_type=payload_type)\n\nprint(client_prediction)",
"Success:True message:\nRequest:\ndata {\n names: \"text\"\n ndarray {\n values {\n list_value {\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.3294117748737335\n }\n values {\n number_value: 0.7254902124404907\n }\n values {\n number_value: 0.6235294342041016\n }\n values {\n number_value: 0.5921568870544434\n }\n values {\n number_value: 0.2352941334247589\n }\n values {\n number_value: 0.1411764770746231\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.8705883026123047\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.9450981020927429\n }\n values {\n number_value: 0.7764706611633301\n }\n values {\n number_value: 0.7764706611633301\n }\n values {\n number_value: 0.7764706611633301\n }\n values {\n number_value: 0.7764706611633301\n }\n values {\n number_value: 0.7764706611633301\n }\n values {\n number_value: 0.7764706611633301\n }\n values {\n number_value: 0.7764706611633301\n }\n values {\n number_value: 0.7764706611633301\n }\n values {\n number_value: 0.6666666865348816\n }\n values {\n number_value: 0.2039215862751007\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.26274511218070984\n }\n values {\n number_value: 0.44705885648727417\n }\n values {\n number_value: 0.2823529541492462\n }\n values {\n number_value: 0.44705885648727417\n }\n values {\n number_value: 0.6392157077789307\n }\n values {\n number_value: 0.8901961445808411\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.8823530077934265\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.9803922176361084\n }\n values {\n number_value: 0.8980392813682556\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.5490196347236633\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.06666667014360428\n }\n values {\n number_value: 0.25882354378700256\n }\n values {\n number_value: 0.05490196496248245\n }\n values {\n number_value: 0.26274511218070984\n }\n values {\n number_value: 0.26274511218070984\n }\n values {\n number_value: 0.26274511218070984\n }\n values {\n number_value: 0.23137256503105164\n }\n values {\n number_value: 0.08235294371843338\n }\n values {\n number_value: 0.9254902601242065\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.41568630933761597\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.32549020648002625\n }\n values {\n number_value: 0.9921569228172302\n }\n values {\n number_value: 0.8196079134941101\n }\n values {\n number_value: 0.07058823853731155\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.08627451211214066\n }\n values {\n number_value: 0.9137255549430847\n }\n values {\n number_value: 1.0\n }\n values {\n number_value: 0.32549020648002625\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.5058823823928833\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.9333333969116211\n }\n values {\n number_value: 0.1725490242242813\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.23137256503105164\n }\n values {\n number_value: 0.9764706492424011\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.24313727021217346\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.5215686559677124\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.7333333492279053\n }\n values {\n number_value: 0.019607843831181526\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.03529411926865578\n }\n values {\n number_value: 0.803921639919281\n }\n values {\n number_value: 0.9725490808486938\n }\n values {\n number_value: 0.22745099663734436\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.4941176772117615\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.7137255072593689\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.29411765933036804\n }\n values {\n number_value: 0.9843137860298157\n }\n values {\n number_value: 0.9411765336990356\n }\n values {\n number_value: 0.22352942824363708\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.07450980693101883\n }\n values {\n number_value: 0.8666667342185974\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.6509804129600525\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.011764707043766975\n }\n values {\n number_value: 0.7960785031318665\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.8588235974311829\n }\n values {\n number_value: 0.13725490868091583\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.14901961386203766\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.3019607961177826\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.12156863510608673\n }\n values {\n number_value: 0.8784314393997192\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.45098042488098145\n }\n values {\n number_value: 0.003921568859368563\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.5215686559677124\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.2039215862751007\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.2392157018184662\n }\n values {\n number_value: 0.9490196704864502\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.2039215862751007\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.4745098352432251\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.8588235974311829\n }\n values {\n number_value: 0.1568627506494522\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.4745098352432251\n }\n values {\n number_value: 0.9960784912109375\n }\n values {\n number_value: 0.8117647767066956\n }\n values {\n number_value: 0.07058823853731155\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n values {\n number_value: 0.0\n }\n }\n }\n }\n}\n\nResponse:\nmeta {\n puid: \"hndttmvl5qua3beavsre2n8hrj\"\n requestPath {\n key: \"classifier\"\n value: \"seldoncontainerregistry.azurecr.io/deep-mnist:0.1\"\n }\n}\ndata {\n names: \"class:0\"\n names: \"class:1\"\n names: \"class:2\"\n names: \"class:3\"\n names: \"class:4\"\n names: \"class:5\"\n names: \"class:6\"\n names: \"class:7\"\n names: \"class:8\"\n names: \"class:9\"\n ndarray {\n values {\n list_value {\n values {\n number_value: 6.386729364749044e-05\n }\n values {\n number_value: 7.228476039955467e-09\n }\n values {\n number_value: 0.00015463074669241905\n }\n values {\n number_value: 0.00286240060813725\n }\n values {\n number_value: 2.6505524601816433e-06\n }\n values {\n number_value: 2.7261585273663513e-05\n }\n values {\n number_value: 2.7168187699544433e-08\n }\n values {\n number_value: 0.9966427087783813\n }\n values {\n number_value: 1.9810671801678836e-05\n }\n values {\n number_value: 0.00022661285765934736\n }\n }\n }\n }\n}\n\n"
]
],
[
[
"### Let's visualise the probability for each label\nIt seems that it correctly predicted the number 7",
"_____no_output_____"
]
],
[
[
"for proba, label in zip(client_prediction.response.data.ndarray.values[0].list_value.ListFields()[0][1], range(0,10)):\n print(f\"LABEL {label}:\\t {proba.number_value*100:6.4f} %\")",
"LABEL 0:\t 0.0064 %\nLABEL 1:\t 0.0000 %\nLABEL 2:\t 0.0155 %\nLABEL 3:\t 0.2862 %\nLABEL 4:\t 0.0003 %\nLABEL 5:\t 0.0027 %\nLABEL 6:\t 0.0000 %\nLABEL 7:\t 99.6643 %\nLABEL 8:\t 0.0020 %\nLABEL 9:\t 0.0227 %\n"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ab8805a30b9ae39b15d3882c0c789a1a3219008
| 108,485 |
ipynb
|
Jupyter Notebook
|
lab1/Lab 1 - Problem 1.ipynb
|
m4rquee/rl-bootcamp
|
5318a5bf5d3616a6e3914f8e5ffc1761f219df25
|
[
"MIT"
] | null | null | null |
lab1/Lab 1 - Problem 1.ipynb
|
m4rquee/rl-bootcamp
|
5318a5bf5d3616a6e3914f8e5ffc1761f219df25
|
[
"MIT"
] | null | null | null |
lab1/Lab 1 - Problem 1.ipynb
|
m4rquee/rl-bootcamp
|
5318a5bf5d3616a6e3914f8e5ffc1761f219df25
|
[
"MIT"
] | null | null | null | 185.127986 | 33,260 | 0.876573 |
[
[
[
"# Lab 1: Markov Decision Processes - Problem 1\n\n\n## Lab Instructions\nAll your answers should be written in this notebook. You shouldn't need to write or modify any other files.\n\n**You should execute every block of code to not miss any dependency.**\n\n\n*This project was developed by Peter Chen, Rocky Duan, Pieter Abbeel for the Berkeley Deep RL Bootcamp, August 2017. Bootcamp website with slides and lecture videos: https://sites.google.com/view/deep-rl-bootcamp/. It is adapted from Berkeley Deep RL Class [HW2](https://github.com/berkeleydeeprlcourse/homework/blob/c1027d83cd542e67ebed982d44666e0d22a00141/hw2/HW2.ipynb) [(license)](https://github.com/berkeleydeeprlcourse/homework/blob/master/LICENSE)*\n\n--------------------------",
"_____no_output_____"
],
[
"## Introduction\n\nThis assignment will review the two classic methods for solving Markov Decision Processes (MDPs) with finite state and action spaces.\nWe will implement value iteration (VI) and policy iteration (PI) for a finite MDP, both of which find the optimal policy in a finite number of iterations.\n\nThe experiments here will use the Frozen Lake environment, a simple gridworld MDP that is taken from `gym` and slightly modified for this assignment. In this MDP, the agent must navigate from the start state to the goal state on a 4x4 grid, with stochastic transitions.",
"_____no_output_____"
]
],
[
[
"from misc import FrozenLakeEnv, make_grader\nenv = FrozenLakeEnv()\nprint(env.__doc__)",
"\n Winter is here. You and your friends were tossing around a frisbee at the park\n when you made a wild throw that left the frisbee out in the middle of the lake.\n The water is mostly frozen, but there are a few holes where the ice has melted.\n If you step into one of those holes, you'll fall into the freezing water.\n At this time, there's an international frisbee shortage, so it's absolutely imperative that\n you navigate across the lake and retrieve the disc.\n However, the ice is slippery, so you won't always move in the direction you intend.\n The surface is described using a grid like the following\n\n SFFF\n FHFH\n FFFH\n HFFG\n\n S : starting point, safe\n F : frozen surface, safe\n H : hole, fall to your doom\n G : goal, where the frisbee is located\n\n The episode ends when you reach the goal or fall in a hole.\n You receive a reward of 1 if you reach the goal, and zero otherwise.\n\n \n"
]
],
[
[
"Let's look at what a random episode looks like.",
"_____no_output_____"
]
],
[
[
"# Some basic imports and setup\nimport numpy as np, numpy.random as nr, gym\nimport matplotlib.pyplot as plt\n%matplotlib inline\nnp.set_printoptions(precision=3)\n\n# Seed RNGs so you get the same printouts as me\nenv.seed(0); from gym.spaces import prng; prng.seed(10)\n# Generate the episode\nenv.reset()\nfor t in range(100):\n env.render()\n a = env.action_space.sample()\n ob, rew, done, _ = env.step(a)\n if done:\n break\nassert done\nenv.render();",
"\n\u001b[41mS\u001b[0mFFF\nFHFH\nFFFH\nHFFG\n (Down)\nS\u001b[41mF\u001b[0mFF\nFHFH\nFFFH\nHFFG\n (Down)\nSFFF\nF\u001b[41mH\u001b[0mFH\nFFFH\nHFFG\n"
]
],
[
[
"In the episode above, the agent falls into a hole after two timesteps. Also note the stochasticity--on the first step, the DOWN action is selected, but the agent moves to the right.\n\nWe extract the relevant information from the gym Env into the MDP class below.\nThe `env` object won't be used any further, we'll just use the `mdp` object.",
"_____no_output_____"
]
],
[
[
"class MDP(object):\n def __init__(self, P, nS, nA, desc=None):\n self.P = P # state transition and reward probabilities, explained below\n self.nS = nS # number of states\n self.nA = nA # number of actions\n self.desc = desc # 2D array specifying what each grid cell means (used for plotting)\nmdp = MDP( {s : {a : [tup[:3] for tup in tups] for (a, tups) in a2d.items()} for (s, a2d) in env.P.items()}, env.nS, env.nA, env.desc)\n\n\nprint(\"mdp.P is a two-level dict where the first key is the state and the second key is the action.\")\nprint(\"The 2D grid cells are associated with indices [0, 1, 2, ..., 15] from left to right and top to down, as in\")\nprint(np.arange(16).reshape(4,4))\nprint(\"Action indices [0, 1, 2, 3] correspond to West, South, East and North.\")\nprint(\"mdp.P[state][action] is a list of tuples (probability, nextstate, reward).\\n\")\nprint(\"For example, state 0 is the initial state, and the transition information for s=0, a=0 is \\nP[0][0] =\", mdp.P[0][0], \"\\n\")\nprint(\"As another example, state 5 corresponds to a hole in the ice, in which all actions lead to the same state with probability 1 and reward 0.\")\nfor i in range(4):\n print(\"P[5][%i] =\" % i, mdp.P[5][i])",
"mdp.P is a two-level dict where the first key is the state and the second key is the action.\nThe 2D grid cells are associated with indices [0, 1, 2, ..., 15] from left to right and top to down, as in\n[[ 0 1 2 3]\n [ 4 5 6 7]\n [ 8 9 10 11]\n [12 13 14 15]]\nAction indices [0, 1, 2, 3] correspond to West, South, East and North.\nmdp.P[state][action] is a list of tuples (probability, nextstate, reward).\n\nFor example, state 0 is the initial state, and the transition information for s=0, a=0 is \nP[0][0] = [(0.1, 0, 0.0), (0.8, 0, 0.0), (0.1, 4, 0.0)] \n\nAs another example, state 5 corresponds to a hole in the ice, in which all actions lead to the same state with probability 1 and reward 0.\nP[5][0] = [(1.0, 5, 0)]\nP[5][1] = [(1.0, 5, 0)]\nP[5][2] = [(1.0, 5, 0)]\nP[5][3] = [(1.0, 5, 0)]\n"
]
],
[
[
"## Part 1: Value Iteration",
"_____no_output_____"
],
[
"### Problem 1: implement value iteration\nIn this problem, you'll implement value iteration, which has the following pseudocode:\n\n---\nInitialize $V^{(0)}(s)=0$, for all $s$\n\nFor $i=0, 1, 2, \\dots$\n- $V^{(i+1)}(s) = \\max_a \\sum_{s'} P(s,a,s') [ R(s,a,s') + \\gamma V^{(i)}(s')]$, for all $s$\n\n---\n\nWe additionally define the sequence of greedy policies $\\pi^{(0)}, \\pi^{(1)}, \\dots, \\pi^{(n-1)}$, where\n$$\\pi^{(i)}(s) = \\arg \\max_a \\sum_{s'} P(s,a,s') [ R(s,a,s') + \\gamma V^{(i)}(s')]$$\n\nYour code will return two lists: $[V^{(0)}, V^{(1)}, \\dots, V^{(n)}]$ and $[\\pi^{(0)}, \\pi^{(1)}, \\dots, \\pi^{(n-1)}]$\n\nTo ensure that you get the same policies as the reference solution, choose the lower-index action to break ties in $\\arg \\max_a$. This is done automatically by np.argmax. This will only affect the \"# chg actions\" printout below--it won't affect the values computed.\n\n<div class=\"alert alert-warning\">\nWarning: make a copy of your value function each iteration and use that copy for the update--don't update your value function in place. \nUpdating in-place is also a valid algorithm, sometimes called Gauss-Seidel value iteration or asynchronous value iteration, but it will cause you to get different results than our reference solution (which in turn will mean that our testing code won’t be able to help in verifying your code).\n</div>",
"_____no_output_____"
]
],
[
[
"def value_iteration(mdp, gamma, nIt, grade_print=print):\n \"\"\"\n Inputs:\n mdp: MDP\n gamma: discount factor\n nIt: number of iterations, corresponding to n above\n Outputs:\n (value_functions, policies)\n \n len(value_functions) == nIt+1 and len(policies) == nIt\n \"\"\"\n grade_print(\"Iteration | max|V-Vprev| | # chg actions | V[0]\")\n grade_print(\"----------+--------------+---------------+---------\")\n Vs = [np.zeros(mdp.nS)] # list of value functions contains the initial value function V^{(0)}, which is zero\n pis = []\n \n def expected_reward(old_V, s):\n trans = mdp.P[s]\n for _, t in trans.items(): # for each action\n prod = [p * (r + gamma * old_V[s_prime]) for p, s_prime, r in t]\n yield sum(prod)\n \n def update(old_V):\n new_V = []\n new_pi = []\n exps = np.array([[*expected_reward(old_V, s)] for s in range(mdp.nS)])\n \n for exp in exps:\n b_action = np.argmax(exp)\n new_V.append(exp[b_action])\n new_pi.append(b_action)\n \n return np.array(new_V), np.array(new_pi)\n \n for it in range(nIt):\n oldpi = pis[-1] if len(pis) > 0 else None # \\pi^{(it)} = Greedy[V^{(it-1)}]. Just used for printout\n Vprev = Vs[-1] # V^{(it)}\n \n # Your code should fill in meaningful values for the following two variables\n # pi: greedy policy for Vprev (not V), \n # corresponding to the math above: \\pi^{(it)} = Greedy[V^{(it)}]\n # ** it needs to be numpy array of ints **\n # V: bellman backup on Vprev\n # corresponding to the math above: V^{(it+1)} = T[V^{(it)}]\n # ** numpy array of floats **\n \n V, pi = update(Vprev)\n \n max_diff = np.abs(V - Vprev).max()\n nChgActions=\"N/A\" if oldpi is None else (pi != oldpi).sum()\n grade_print(\"%4i | %6.5f | %4s | %5.3f\"%(it, max_diff, nChgActions, V[0]))\n Vs.append(V)\n pis.append(pi)\n return Vs, pis\n\nGAMMA = 0.95 # we'll be using this same value in subsequent problems\n\n\n# The following is the output of a correct implementation; when\n# this code block is run, your implementation's print output will be\n# compared with expected output.\n# (incorrect line in red background with correct line printed side by side to help you debug)\nexpected_output = \"\"\"Iteration | max|V-Vprev| | # chg actions | V[0]\n----------+--------------+---------------+---------\n 0 | 0.80000 | N/A | 0.000\n 1 | 0.60800 | 2 | 0.000\n 2 | 0.51984 | 2 | 0.000\n 3 | 0.39508 | 2 | 0.000\n 4 | 0.30026 | 2 | 0.000\n 5 | 0.25355 | 1 | 0.254\n 6 | 0.10478 | 0 | 0.345\n 7 | 0.09657 | 0 | 0.442\n 8 | 0.03656 | 0 | 0.478\n 9 | 0.02772 | 0 | 0.506\n 10 | 0.01111 | 0 | 0.517\n 11 | 0.00735 | 0 | 0.524\n 12 | 0.00310 | 0 | 0.527\n 13 | 0.00190 | 0 | 0.529\n 14 | 0.00083 | 0 | 0.530\n 15 | 0.00049 | 0 | 0.531\n 16 | 0.00022 | 0 | 0.531\n 17 | 0.00013 | 0 | 0.531\n 18 | 0.00006 | 0 | 0.531\n 19 | 0.00003 | 0 | 0.531\"\"\"\nVs_VI, pis_VI = value_iteration(mdp, gamma=GAMMA, nIt=20, grade_print=make_grader(expected_output))",
"Iteration | max|V-Vprev| | # chg actions | V[0]\n----------+--------------+---------------+---------\n 0 | 0.80000 | N/A | 0.000\n 1 | 0.60800 | 2 | 0.000\n 2 | 0.51984 | 2 | 0.000\n 3 | 0.39508 | 2 | 0.000\n 4 | 0.30026 | 2 | 0.000\n 5 | 0.25355 | 1 | 0.254\n 6 | 0.10478 | 0 | 0.345\n 7 | 0.09657 | 0 | 0.442\n 8 | 0.03656 | 0 | 0.478\n 9 | 0.02772 | 0 | 0.506\n 10 | 0.01111 | 0 | 0.517\n 11 | 0.00735 | 0 | 0.524\n 12 | 0.00310 | 0 | 0.527\n 13 | 0.00190 | 0 | 0.529\n 14 | 0.00083 | 0 | 0.530\n 15 | 0.00049 | 0 | 0.531\n 16 | 0.00022 | 0 | 0.531\n 17 | 0.00013 | 0 | 0.531\n 18 | 0.00006 | 0 | 0.531\n 19 | 0.00003 | 0 | 0.531\nTest succeeded\n"
]
],
[
[
"Below, we've illustrated the progress of value iteration. Your optimal actions are shown by arrows.\nAt the bottom, the value of the different states are plotted.",
"_____no_output_____"
]
],
[
[
"for (V, pi) in zip(Vs_VI[:10], pis_VI[:10]):\n plt.figure(figsize=(3,3))\n plt.imshow(V.reshape(4,4), cmap='gray', interpolation='none', clim=(0,1))\n ax = plt.gca()\n ax.set_xticks(np.arange(4)-.5)\n ax.set_yticks(np.arange(4)-.5)\n ax.set_xticklabels([])\n ax.set_yticklabels([])\n Y, X = np.mgrid[0:4, 0:4]\n a2uv = {0: (-1, 0), 1:(0, -1), 2:(1,0), 3:(-1, 0)}\n Pi = pi.reshape(4,4)\n for y in range(4):\n for x in range(4):\n a = Pi[y, x]\n u, v = a2uv[a]\n plt.arrow(x, y,u*.3, -v*.3, color='m', head_width=0.1, head_length=0.1) \n plt.text(x, y, str(env.desc[y,x].item().decode()),\n color='g', size=12, verticalalignment='center',\n horizontalalignment='center', fontweight='bold')\n plt.grid(color='b', lw=2, ls='-')\nplt.figure()\nplt.plot(Vs_VI)\nplt.title(\"Values of different states\");",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ab88a066177941a187dcde39a17c9f323e13e8a
| 321,357 |
ipynb
|
Jupyter Notebook
|
Image_Classification_NumPy.ipynb
|
leventbass/image_classification_logistic_regression
|
3e9c6db1989e429d9f6e0b29619a6d4a27041b6e
|
[
"MIT"
] | 1 |
2020-05-17T18:05:54.000Z
|
2020-05-17T18:05:54.000Z
|
Image_Classification_NumPy.ipynb
|
leventbass/image_classification_logistic_regression
|
3e9c6db1989e429d9f6e0b29619a6d4a27041b6e
|
[
"MIT"
] | null | null | null |
Image_Classification_NumPy.ipynb
|
leventbass/image_classification_logistic_regression
|
3e9c6db1989e429d9f6e0b29619a6d4a27041b6e
|
[
"MIT"
] | 2 |
2020-07-08T20:01:47.000Z
|
2021-09-27T22:55:19.000Z
| 439.012295 | 76,720 | 0.93566 |
[
[
[
"# Image Classification with Logistic Regression from Scratch with NumPy",
"_____no_output_____"
],
[
"Welcome to another jupyter notebook of implementing machine learning algorithms from scratch using only NumPy. This time we will be implementing a different version of logistic regression for a simple image classification task. I've already done a basic version of logistic regression before [here](https://github.com/leventbass/logistic_regression). This time, we will use logistic regression to classify images. I will show all necessary mathematical equations of logistic regression and how to vectorize the summations in the equations. We will be working with a subset of the famous handwritten digit dataset called MNIST. In the subset, there will only be images of digit 1 and 5. Therefore, we will be solving a binary classification problem. \n\nThis notebook includes feature extraction, model training, and evaluation steps. Let's see what we will achieve in this post in steps:\n\n* First, we wil load and visualize the dataset and extract two different set of features to build a classifier on.\n\n* We will run our logistic regression algorithm with gradient descent the representations to classify digits into 1 and 5.\n\n* We will experiment with different learning rates to find the best one.\n\n* Finally, we will evaluate the implemented models, decide which is the best performing one and visualize a decision boundary.\n\n* Once again, let's remind ourselves that we won't be using any function or library that accomplishes the task itself. For instance, we won't use scikit-learn to implement cross validation, we will use numpy for that and for all of the other tasks. ",
"_____no_output_____"
]
],
[
[
"import numpy as np\nimport matplotlib.pyplot as plt\nimport pandas as pd",
"_____no_output_____"
]
],
[
[
"\n",
"_____no_output_____"
],
[
"## Feature Extraction",
"_____no_output_____"
],
[
"Let's load the training/test data and labels as numpy arrays. All data that is used is provided in the repository in data folder. Train and test data are 1561x256 and 424x256 dimensional matrices, respectively. Each row in the aforementioned matrices corresponds to an image of a digit. The 256 pixels correspond to a 16x16 image. Label 1 is assigned to digit 1 and label -1 is assigned to digit 5.",
"_____no_output_____"
]
],
[
[
"train_x = np.load('data/train_data.npy')\ntrain_y = np.load('data/train_labels.npy')\ntest_x = np.load('data/test_data.npy')\ntest_y = np.load('data/test_labels.npy')",
"_____no_output_____"
]
],
[
[
"Now, let's display two of the digit images, one for digit 1 and one for digit 5. We will use `imshow` function of `matplotlib` library with a suitable colormap. We will first need to reshape 256 pixels to a 16x16 matrix. ",
"_____no_output_____"
]
],
[
[
"digit_1 = train_x[0].reshape((16,16))\ndigit_5 = train_x[-1].reshape((16,16))\n\nplt.subplot(121, title='Digit 1')\nplt.imshow(digit_1, cmap='gray');\nplt.subplot(122, title='Digit 5')\nplt.imshow(digit_5, cmap='gray');",
"_____no_output_____"
]
],
[
[
"**Implementing Representation 1:**\nNow, we will extract the **symmetry** and **average intensity** features to use in the model. To compute the intensity features, we compute the average pixel value of the image, and for the symmetry feature, we compute the negative of the norm of the difference between the image and its y-axis symmetrical. We will extract these two features for each image in the training and test sets. As a result, we should obtain a training data matrix of size 1561x2 and test data matrix of size 424x2.\n\nThroughout the notebook, we will refer the representation with these two features as **Representation 1** ",
"_____no_output_____"
]
],
[
[
"train_feature_1 = np.mean(train_x, axis=1)\ntest_feature_1 = np.mean(test_x, axis=1)",
"_____no_output_____"
],
[
"mirrored_image_train = np.flip(train_x.reshape((train_x.shape[0],16,16)), axis=2)\nmirrored_image_test = np.flip(test_x.reshape((test_x.shape[0],16,16)), axis=2)\n \nplt.subplot(121, title='Image')\nplt.imshow(train_x[-1].reshape((16,16)), cmap='gray');\nplt.subplot(122, title='Mirrored Image')\nplt.imshow(mirrored_image_train[-1], cmap='gray');\n",
"_____no_output_____"
],
[
"train_diff = train_x - mirrored_image_train.reshape((mirrored_image_train.shape[0],256))\ntest_diff = test_x - mirrored_image_test.reshape((mirrored_image_test.shape[0],256))\n\nnorm_train_diff = np.linalg.norm(train_diff, axis=1)\nnorm_test_diff = np.linalg.norm(test_diff, axis=1)\n\ntrain_feature_2 = -(norm_train_diff)\ntest_feature_2 = -(norm_test_diff)\n\ntrain_X_1 = np.concatenate((train_feature_1[:,np.newaxis], train_feature_2[:,np.newaxis]), axis=1)\ntest_X_1 = np.concatenate((test_feature_1[:,np.newaxis], test_feature_2[:,np.newaxis]), axis=1)",
"_____no_output_____"
]
],
[
[
"Now, let's provide two scatter plots, one for training and one for test data. The plots will contain the average intensity values in the x-axis and symmetry values in the y-axis. We will denote the data points of label 1 with blue marker shaped <font color='blue'>o</font> and the data points of label -1 with a red marker shaped <font color='red'>x</font>.",
"_____no_output_____"
]
],
[
[
"plt.figure(figsize=(6,6))\nplt.scatter(train_X_1[(train_y==1),0], train_X_1[(train_y==1),1], marker='o', color='blue', s=16)\nplt.scatter(train_X_1[(train_y==-1),0], train_X_1[(train_y==-1),1], marker='x', color='red', s=16)\nplt.title('Class Distribution of Training Data for Representation 1')\nplt.xlabel('Average Intensity')\nplt.ylabel('Symmetry')\n\nplt.figure(figsize=(6,6))\nplt.scatter(test_X_1[(test_y==1),0], test_X_1[(test_y==1),1], marker='o', color='blue', s=16)\nplt.scatter(test_X_1[(test_y==-1),0], test_X_1[(test_y==-1),1], marker='x', color='red', s=16)\nplt.title('Class Distribution of Test Data for Representation 1')\nplt.xlabel('Average Intensity')\nplt.ylabel('Symmetry');",
"_____no_output_____"
]
],
[
[
"**Implementing Representation 2:** We will come up with an alternative feature extraction approach and we will refer this representation as **Representation 2**.",
"_____no_output_____"
]
],
[
[
"train_rep2_fet1 = np.array([(i>-1).sum() for i in train_x])/(train_x.shape[0]) # feature 1 for representation 2\ntest_rep2_fet1 = np.array([(i>-1).sum() for i in test_x])/(test_x.shape[0])",
"_____no_output_____"
],
[
"train_rep2_fet2 = np.std(train_x, axis=1) # feature 2 for representation 2\ntest_rep2_fet2 = np.std(test_x, axis=1)",
"_____no_output_____"
],
[
"train_X_2 = np.concatenate((train_rep2_fet1[:,np.newaxis], train_rep2_fet2[:,np.newaxis]), axis=1)\ntest_X_2 = np.concatenate((test_rep2_fet1[:,np.newaxis], test_rep2_fet2[:,np.newaxis]), axis=1)",
"_____no_output_____"
]
],
[
[
"To create the first feature of representation 2, we sum up all of the pixel values that are higher than -1 since pixel values of -1 represent the surrounding area of the image and not itself. By summing up those values we get a number that would be clearly distinctive for image of number 5 and 1, because evidently number 5 would take up more space than number 1 when it is drawn. \n\nTo add another feature to representation 2, let's calculate the standard deviation of the images. Image of number 5 will obviously have more standard deviation than image of number 1 because of the fact that it is more dispersed throughtout the area than number 1, while pixel values of number 1 are more confined and closer to each other than the image of number 5. Hence, taking the standard deviation of pixel values would be a differentiating factor for our images.",
"_____no_output_____"
]
],
[
[
"plt.figure(figsize=(9,5))\nplt.scatter(train_X_2[(train_y==1),0], train_X_2[(train_y==1),1], marker='o', color='blue', s=16)\nplt.scatter(train_X_2[(train_y==-1),0], train_X_2[(train_y==-1),1], marker='x', color='red', s=16)\nplt.title('Class Distribution of Training Data for Representation 2')\nplt.xlabel('Average Intensity')\nplt.ylabel('Symmetry')\n\nplt.figure(figsize=(9,5))\nplt.scatter(test_X_2[(test_y==1),0], test_X_2[(test_y==1),1], marker='o', color='blue', s=16)\nplt.scatter(test_X_2[(test_y==-1),0], test_X_2[(test_y==-1),1], marker='x', color='red', s=16)\nplt.title('Class Distribution of Test Data for Representation 2')\nplt.xlabel('Length of non-white Pixels')\nplt.ylabel('Standard Deviation');\n",
"_____no_output_____"
]
],
[
[
"## Logistic Regression",
"_____no_output_____"
],
[
"Let's implement the logistic regression classifier from scratch with gradient descent and train it using Representation 1 and Representation 2 as inputs. We will concatenate 1 to our features for the intercept term, such that one data point will look like for 2-D features [1,$x_1$,$x_2$], and the model vector will be [$w_0, w_1, w_2$], where $w_0$ is the intercept parameter. \n",
"_____no_output_____"
]
],
[
[
"def data_init(X, y):\n y = y[:,np.newaxis]\n\n m = len(y)\n\n X = np.hstack((np.ones((m,1)),X))\n \n n = np.size(X,1)\n params = np.zeros((n,1))\n \n return (X, y, params)",
"_____no_output_____"
]
],
[
[
"To implement the gradient of the logistic loss with respect to $w$, first let's derive its expression:",
"_____no_output_____"
],
[
"Total cost is:\n\n$E(w) = \\frac{1}{N} \\sum_{n=1}^{N} \\ln \\left(1 + \\exp \\left(-y^{\\left(n\\right)} w^T x^{\\left(n\\right)}\\right)\\right)$\n \n\nCost for one sample is:\n\n\n$E \\left(w^{\\left(1\\right)} \\right) = \\ln \\left(1 + \\exp \\left(-y^{\\left(1\\right)} w^T x^{\\left(1\\right)} \\right) \\right)$\n \nwhere;\n\n$y = \\begin{bmatrix} y_1 \\\\ y_2 \\\\ \\vdots \\\\ y_N \\end{bmatrix}_{N\\times 1}$\n$x = \\begin{bmatrix} 1 & {x_1}^{\\left(1\\right)} & {x_2}^{\\left(1\\right)} \\\\\n 1 & {x_1}^{\\left(2\\right)} & {x_2}^{\\left(2\\right)} \\\\\n \\vdots & \\vdots & \\vdots \\\\\n 1 & {x_1}^{\\left(N\\right)} & {x_2}^{\\left(N\\right)}\\end{bmatrix}_{N\\times 3}$\n$w = \\begin{bmatrix}w_0 \\\\ w_1 \\\\ w_2 \\end{bmatrix}_{3\\times 1}$\n\n \n \nLet $z = -y^{\\left(1\\right)} w^T x^{\\left(1\\right)}$:\n\n \n\n$\\begin{aligned}\n\\frac{\\partial E}{\\partial w_0} &= \\frac{\\partial \\ln(1 + \\exp(z))}{\\partial w_0} \\\\\n &=\\frac{\\exp(z) \\frac{\\partial z}{\\partial w_0}}{1 + \\exp(z)} \n \\quad \\left( \\theta(z) = \\frac{\\exp(z)}{1 + \\exp(z)} \\right)\\\\\n &= \\theta(z) \\frac{\\partial z}{\\partial w_0} \\\\\n &= \\theta\\left(-y^{\\left(1\\right)} w^T x^{\\left(1\\right)}\\right)\n \\frac{\\partial \\left(-y^{\\left(1\\right)} w^T x^{\\left(1\\right)}\\right)}{\\partial w_0} \\\\ \n &= \\theta\\left(-y^{\\left(1\\right)} w^T x^{\\left(1\\right)}\\right)\n \\frac{\\partial \\left(-y^{\\left(1\\right)} \\left(w_0 + w_1 {x_1}^{\\left(1\\right)} + w_2 {x_2}^{\\left(1\\right)}\\right)\\right)}{\\partial w_0}\\\\\n \\frac{\\partial E}{\\partial w_0} &= \\theta\\left(-y^{\\left(1\\right)} w^T x^{\\left(1\\right)}\\right) \\left( -y^{\\left(1\\right)} \\right) \\\\\n\\frac{\\partial E}{\\partial w_1} &= \\theta\\left(-y^{\\left(1\\right)} w^T x^{\\left(1\\right)}\\right) \\left( -y^{\\left(1\\right)} {x_1}^{\\left(1\\right)} \\right)\\\\\n\\frac{\\partial E}{\\partial w_2} &= \\theta\\left(-y^{\\left(1\\right)} w^T x^{\\left(1\\right)}\\right) \\left( -y^{\\left(1\\right)} {x_2}^{\\left(1\\right)} \\right)\\\\\n\\end{aligned}$\n\n \n$\\begin{aligned}\n\\nabla E (w) &= \\frac{1}{N} \\sum_{n=1}^{N} -\\theta \\left(-y^{\\left(n\\right)} w^T x^{\\left(n\\right)}\\right) y^{\\left(n\\right)} x^{\\left(n\\right)}\\\\\n&= \\frac{1}{N} {\\left( - \\textbf{y} \\circ \\textbf{x} \\right)}^T \\cdot \\theta \\left( -\\textbf{y} \\circ \\textbf{x w} \\right)\n\\end{aligned}$\n",
"_____no_output_____"
],
[
"To prove that our implementation is converging, we will keep the loss values at each gradient descent iteration in a numpy array. To decide when to terminate the gradient descent iterations, we will check the absolute difference between the current loss value and the loss value of the previous step. If the difference is less than a small number, such as $10^{-5}$, we will exit the loop.",
"_____no_output_____"
]
],
[
[
"def sigmoid(x):\n return 1 / (1 + np.exp(-x))\n\ndef gradient_descent(X, y, params, learning_rate):\n m = len(y)\n \n cost_history = []\n i=0\n while(True):\n params = params - (learning_rate/m) * ((-y * X).T @ sigmoid(-y * (X @ params)))\n cost_history.append(compute_cost(X, y, params))\n \n if(i!=0 and abs(cost_history[i] - cost_history[i-1]) < 10**-5):\n break;\n \n i+=1\n cost_history = np.array(cost_history)\n return (cost_history, params)\n\ndef compute_cost(X, y, theta):\n N = len(y)\n cost = np.sum(np.log(1+np.exp(-y * (X @ theta)))) / N\n return cost\n ",
"_____no_output_____"
]
],
[
[
"After the training is finalized, we will plot the loss values with respect to iteration count. Obviously, we should observe a decreasing loss as the number of iterations increases. Also, we will experiment with 5 different learning rates between 0 and 1, and plot the convergence curves for each learning rate in the same plot to observe the effect of the learning rate (step size) on the convergence. ",
"_____no_output_____"
]
],
[
[
"(X, y, params) = data_init(train_X_1, train_y)\n\nlr_list = [0.001, 0.003, 0.01, 0.03, 0.1]\nc_list = ['red', 'green', 'yellow', 'blue','black']\n\nplt.figure()\nfor lr, color in zip(lr_list, c_list):\n (cost_history, params_optimal) = gradient_descent(X, y, params, lr)\n plt.plot(range(len(cost_history)),cost_history, c=color);\nplt.title(\"Convergence Graph of Cost Function\")\nplt.xlabel(\"Number of Iterations\")\nplt.ylabel(\"Cost\")\nplt.show()",
"_____no_output_____"
]
],
[
[
"## Evaluation",
"_____no_output_____"
],
[
"Now, let's train the logistic regression classifier on Representation 1 and 2 with the best learning rate we have used so far. We will report the training and test classification accuracy as:\n\n\\begin{align*}\n\\frac{\\text{number of correctly classified samples}}{\\text{total number of samples}}x100\n\\end{align*}",
"_____no_output_____"
]
],
[
[
"def predict(X, params):\n y_pred_dummy = np.round(sigmoid(X @ params))\n y_pred = np.where(y_pred_dummy==0,-1,1)\n return y_pred\n\ndef get_accuracy(y_pred, y):\n score = float(sum(y_pred == y))/ float(len(y)) * 100\n \n return score\n\ndef evaluate(train_X, train_y, test_X, test_y, learning_rate, lambda_param):\n\n (X, y, params) = data_init(train_X, train_y)\n\n (_, params_optimal_1) = gradient_descent(X, y, params, learning_rate)\n\n \n X_normalized = test_X\n\n X_test = np.hstack((np.ones((X_normalized.shape[0],1)),X_normalized))\n\n y_pred_train = predict(X, params_optimal_1)\n\n train_score = get_accuracy(y_pred_train, y)\n print('Training Score:',train_score)\n\n y_pred_test = predict(X_test, params_optimal_1)\n\n test_score = get_accuracy(y_pred_test, test_y[:,np.newaxis])\n print('Test Score:',test_score)",
"_____no_output_____"
],
[
"print('Evaluation results for Representation 1:')\nprint('-'*50)\nevaluate(train_X_1, train_y, test_X_1, test_y, 0.1, 0.0003)\n\nprint('\\nEvaluation results for Representation 2:')\nprint('-'*50)\nevaluate(train_X_2, train_y, test_X_2, test_y, 0.1, 0.0001)",
"Evaluation results for Representation 1:\n--------------------------------------------------\nTraining Score: 97.69378603459322\nTest Score: 95.04716981132076\n\nEvaluation results for Representation 2:\n--------------------------------------------------\nTraining Score: 83.27994875080077\nTest Score: 81.13207547169812\n"
]
],
[
[
"Last but not least, we will visualize the decision boundary (the line that is given by $\\mathbf{w}^{T}x=0$) obtained from the logistic regression classifier learned. For this purpose, we will only use Representation 1. Below, two scatter plots can be seen for training and test data points with the decision boundary shown on each of the plots.",
"_____no_output_____"
]
],
[
[
"(X, y, params) = data_init(train_X_1,train_y)\n\nlearning_rate = 0.1\n\n(_, params_optimal_1) = gradient_descent(X, y, params, learning_rate)\nslope = -(params_optimal_1[1] / params_optimal_1[2])\nintercept = -(params_optimal_1[0] / params_optimal_1[2])\n\ntitles = ['Training Data with Decision Boundary', 'Test Data with Decision Boundary']\n\nfor X, y, title in [(train_X_1, y, titles[0]), (test_X_1, test_y, titles[1])]:\n plt.figure(figsize=(7,7))\n plt.scatter(X[:,0],X[:,1],c=y.reshape(-1), s=14, cmap='bwr')\n ax = plt.gca()\n ax.autoscale(False)\n x_vals = np.array(ax.get_xlim())\n y_vals = intercept + (slope * x_vals)\n plt.title(title);\n plt.plot(x_vals, y_vals, c='k')\n ",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
]
] |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.