hexsha
stringlengths 40
40
| size
int64 6
14.9M
| ext
stringclasses 1
value | lang
stringclasses 1
value | max_stars_repo_path
stringlengths 6
260
| max_stars_repo_name
stringlengths 6
119
| max_stars_repo_head_hexsha
stringlengths 40
41
| max_stars_repo_licenses
list | max_stars_count
int64 1
191k
⌀ | max_stars_repo_stars_event_min_datetime
stringlengths 24
24
⌀ | max_stars_repo_stars_event_max_datetime
stringlengths 24
24
⌀ | max_issues_repo_path
stringlengths 6
260
| max_issues_repo_name
stringlengths 6
119
| max_issues_repo_head_hexsha
stringlengths 40
41
| max_issues_repo_licenses
list | max_issues_count
int64 1
67k
⌀ | max_issues_repo_issues_event_min_datetime
stringlengths 24
24
⌀ | max_issues_repo_issues_event_max_datetime
stringlengths 24
24
⌀ | max_forks_repo_path
stringlengths 6
260
| max_forks_repo_name
stringlengths 6
119
| max_forks_repo_head_hexsha
stringlengths 40
41
| max_forks_repo_licenses
list | max_forks_count
int64 1
105k
⌀ | max_forks_repo_forks_event_min_datetime
stringlengths 24
24
⌀ | max_forks_repo_forks_event_max_datetime
stringlengths 24
24
⌀ | avg_line_length
float64 2
1.04M
| max_line_length
int64 2
11.2M
| alphanum_fraction
float64 0
1
| cells
list | cell_types
list | cell_type_groups
list |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
4acb8a9e799b1dd74ad6372cfe1539e2782938a3
| 39,203 |
ipynb
|
Jupyter Notebook
|
Artificial Intelligence with PyTorch/Lesson 6_ Convolutional Neural Networks using PyTorch.ipynb
|
ZapAutomation/ZappyAIAcademyNotebooks
|
0f7ecc119c53a30188e4db557112ec2020adb827
|
[
"MIT"
] | null | null | null |
Artificial Intelligence with PyTorch/Lesson 6_ Convolutional Neural Networks using PyTorch.ipynb
|
ZapAutomation/ZappyAIAcademyNotebooks
|
0f7ecc119c53a30188e4db557112ec2020adb827
|
[
"MIT"
] | null | null | null |
Artificial Intelligence with PyTorch/Lesson 6_ Convolutional Neural Networks using PyTorch.ipynb
|
ZapAutomation/ZappyAIAcademyNotebooks
|
0f7ecc119c53a30188e4db557112ec2020adb827
|
[
"MIT"
] | null | null | null | 39,203 | 39,203 | 0.751295 |
[
[
[
"import torch\nimport torch.nn as nn\nimport torch.nn.functional as F\nimport numpy as np\nfrom torchvision import transforms, datasets",
"_____no_output_____"
],
[
"apply_transform = transforms.Compose([\ntransforms.Resize(32)\n,transforms.ToTensor()\n])\n\nBatchSize = 256\n\ntrainset = datasets.MNIST(root = './MNIST', train = True, download = True, transform = apply_transform)\ntrainLoader = torch.utils.data.DataLoader(trainset, batch_size = BatchSize, shuffle = True, num_workers = 10)\n\ntestset = datasets.MNIST(root = './MNIST', train = False, download = True, transform = apply_transform)\ntestLoader = torch.utils.data.DataLoader(testset, batch_size = BatchSize, shuffle = True, num_workers = 10)",
"Downloading http://yann.lecun.com/exdb/mnist/train-images-idx3-ubyte.gz to ./MNIST/MNIST/raw/train-images-idx3-ubyte.gz\n"
],
[
"print(len(trainLoader.dataset))\nprint(len(testLoader.dataset))",
"60000\n10000\n"
],
[
"class LeNet(nn.Module):\n def __init__(self):\n super(LeNet, self).__init__()\n self.conv1 = nn.Conv2d(1, 6, kernel_size = 5, bias = True)\n self.maxpool1 = nn.MaxPool2d(kernel_size = 2, stride = 2)\n self.conv2 = nn.Conv2d(6, 16, kernel_size = 5, bias =True)\n self.maxpool2 = nn.MaxPool2d(kernel_size = 2, stride = 2)\n self.fc1 = nn.Linear(400, 120, bias = True)\n self.fc2 = nn.Linear(120, 84, bias = True)\n self.fc3 = nn.Linear(84, 10, bias = True)\n \n def forward(self, x):\n x = F.relu(self.conv1(x))\n x = self.maxpool1(x)\n x = F.relu(self.conv2(x))\n x = self.maxpool2(x)\n x = x.view(-1, 400)\n x = F.relu(self.fc1(x))\n x = F.relu(self.fc2(x))\n x = F.relu(self.fc3(x))\n return F.log_softmax(x, dim = 1)\n \nnet = LeNet()\nprint(net)",
"\nLeNet(\n (conv1): Conv2d(1, 6, kernel_size=(5, 5), stride=(1, 1))\n (maxpool1): MaxPool2d(kernel_size=2, stride=2, padding=0, dilation=1, ceil_mode=False)\n (conv2): Conv2d(6, 16, kernel_size=(5, 5), stride=(1, 1))\n (maxpool2): MaxPool2d(kernel_size=2, stride=2, padding=0, dilation=1, ceil_mode=False)\n (fc1): Linear(in_features=400, out_features=120, bias=True)\n (fc2): Linear(in_features=120, out_features=84, bias=True)\n (fc3): Linear(in_features=84, out_features=10, bias=True)\n)\n"
],
[
"use_gpu = torch.cuda.is_available()\nif use_gpu:\n print(\"GPU is available\")\n net = net.cuda()",
"GPU is available\n"
],
[
"learning_rate = 0.01\ncriterion = nn.CrossEntropyLoss()\nnum_epochs = 50\ntrain_loss = []\ntrain_acc = []\n\nfor epoch in range(num_epochs):\n running_loss = 0.0\n running_corr = 0.0\n \n for i, data in enumerate(trainLoader):\n inputs, labels = data\n if use_gpu:\n inputs, labels = inputs.cuda(), labels.cuda()\n net.zero_grad()\n output = net(inputs)\n loss = criterion(output, labels)\n running_loss += loss\n preds = torch.argmax(output, dim = 1)\n running_corr = torch.sum(preds == labels) \n\n totalLoss = running_loss / (i + 1)\n totalLoss.backward()\n \n for f in net.parameters():\n f.data.sub_(f.grad.data * learning_rate)\n \n epoch_loss = running_loss.item()/(i+1)\n epoch_acc = running_corr.item()/60000\n\n train_loss.append(epoch_loss)\n train_acc.append(epoch_acc)\n\n print('Epoch {:.0f}/{:.0f} : Training loss: {:.4f} | Training Accuracy: {:.4f}'.format(epoch+1,num_epochs,epoch_loss,epoch_acc*100))\n",
"Epoch 1/100 : Training loss: 2.3023 | Training Accuracy: 0.0133\nEpoch 2/100 : Training loss: 2.3022 | Training Accuracy: 0.0150\nEpoch 3/100 : Training loss: 2.3022 | Training Accuracy: 0.0067\nEpoch 4/100 : Training loss: 2.3022 | Training Accuracy: 0.0117\nEpoch 5/100 : Training loss: 2.3022 | Training Accuracy: 0.0067\nEpoch 6/100 : Training loss: 2.3022 | Training Accuracy: 0.0100\nEpoch 7/100 : Training loss: 2.3022 | Training Accuracy: 0.0067\nEpoch 8/100 : Training loss: 2.3021 | Training Accuracy: 0.0133\nEpoch 9/100 : Training loss: 2.3021 | Training Accuracy: 0.0117\nEpoch 10/100 : Training loss: 2.3021 | Training Accuracy: 0.0067\nEpoch 11/100 : Training loss: 2.3021 | Training Accuracy: 0.0133\nEpoch 12/100 : Training loss: 2.3021 | Training Accuracy: 0.0050\nEpoch 13/100 : Training loss: 2.3021 | Training Accuracy: 0.0067\nEpoch 14/100 : Training loss: 2.3020 | Training Accuracy: 0.0083\nEpoch 15/100 : Training loss: 2.3020 | Training Accuracy: 0.0150\nEpoch 16/100 : Training loss: 2.3020 | Training Accuracy: 0.0183\nEpoch 17/100 : Training loss: 2.3020 | Training Accuracy: 0.0067\nEpoch 18/100 : Training loss: 2.3020 | Training Accuracy: 0.0150\nEpoch 19/100 : Training loss: 2.3020 | Training Accuracy: 0.0217\nEpoch 20/100 : Training loss: 2.3019 | Training Accuracy: 0.0083\nEpoch 21/100 : Training loss: 2.3019 | Training Accuracy: 0.0100\nEpoch 22/100 : Training loss: 2.3019 | Training Accuracy: 0.0100\nEpoch 23/100 : Training loss: 2.3019 | Training Accuracy: 0.0100\nEpoch 24/100 : Training loss: 2.3019 | Training Accuracy: 0.0200\nEpoch 25/100 : Training loss: 2.3019 | Training Accuracy: 0.0150\nEpoch 26/100 : Training loss: 2.3018 | Training Accuracy: 0.0100\nEpoch 27/100 : Training loss: 2.3018 | Training Accuracy: 0.0100\nEpoch 28/100 : Training loss: 2.3018 | Training Accuracy: 0.0083\nEpoch 29/100 : Training loss: 2.3018 | Training Accuracy: 0.0167\nEpoch 30/100 : Training loss: 2.3018 | Training Accuracy: 0.0167\nEpoch 31/100 : Training loss: 2.3018 | Training Accuracy: 0.0100\nEpoch 32/100 : Training loss: 2.3017 | Training Accuracy: 0.0150\nEpoch 33/100 : Training loss: 2.3017 | Training Accuracy: 0.0183\nEpoch 34/100 : Training loss: 2.3017 | Training Accuracy: 0.0117\nEpoch 35/100 : Training loss: 2.3017 | Training Accuracy: 0.0200\nEpoch 36/100 : Training loss: 2.3017 | Training Accuracy: 0.0167\nEpoch 37/100 : Training loss: 2.3017 | Training Accuracy: 0.0167\nEpoch 38/100 : Training loss: 2.3016 | Training Accuracy: 0.0167\nEpoch 39/100 : Training loss: 2.3016 | Training Accuracy: 0.0217\nEpoch 40/100 : Training loss: 2.3016 | Training Accuracy: 0.0183\nEpoch 41/100 : Training loss: 2.3016 | Training Accuracy: 0.0267\nEpoch 42/100 : Training loss: 2.3016 | Training Accuracy: 0.0150\nEpoch 43/100 : Training loss: 2.3016 | Training Accuracy: 0.0167\nEpoch 44/100 : Training loss: 2.3015 | Training Accuracy: 0.0233\nEpoch 45/100 : Training loss: 2.3015 | Training Accuracy: 0.0183\nEpoch 46/100 : Training loss: 2.3015 | Training Accuracy: 0.0183\nEpoch 47/100 : Training loss: 2.3015 | Training Accuracy: 0.0133\nEpoch 48/100 : Training loss: 2.3015 | Training Accuracy: 0.0233\nEpoch 49/100 : Training loss: 2.3014 | Training Accuracy: 0.0183\nEpoch 50/100 : Training loss: 2.3014 | Training Accuracy: 0.0200\nEpoch 51/100 : Training loss: 2.3014 | Training Accuracy: 0.0200\nEpoch 52/100 : Training loss: 2.3014 | Training Accuracy: 0.0167\nEpoch 53/100 : Training loss: 2.3014 | Training Accuracy: 0.0167\nEpoch 54/100 : Training loss: 2.3014 | Training Accuracy: 0.0283\nEpoch 55/100 : Training loss: 2.3013 | Training Accuracy: 0.0183\nEpoch 56/100 : Training loss: 2.3014 | Training Accuracy: 0.0050\nEpoch 57/100 : Training loss: 2.3013 | Training Accuracy: 0.0183\nEpoch 58/100 : Training loss: 2.3013 | Training Accuracy: 0.0117\nEpoch 59/100 : Training loss: 2.3013 | Training Accuracy: 0.0167\nEpoch 60/100 : Training loss: 2.3013 | Training Accuracy: 0.0200\nEpoch 61/100 : Training loss: 2.3013 | Training Accuracy: 0.0133\nEpoch 62/100 : Training loss: 2.3012 | Training Accuracy: 0.0250\nEpoch 63/100 : Training loss: 2.3012 | Training Accuracy: 0.0200\nEpoch 64/100 : Training loss: 2.3012 | Training Accuracy: 0.0250\nEpoch 65/100 : Training loss: 2.3012 | Training Accuracy: 0.0267\nEpoch 66/100 : Training loss: 2.3012 | Training Accuracy: 0.0183\nEpoch 67/100 : Training loss: 2.3011 | Training Accuracy: 0.0183\nEpoch 68/100 : Training loss: 2.3011 | Training Accuracy: 0.0200\nEpoch 69/100 : Training loss: 2.3011 | Training Accuracy: 0.0167\nEpoch 70/100 : Training loss: 2.3011 | Training Accuracy: 0.0217\nEpoch 71/100 : Training loss: 2.3011 | Training Accuracy: 0.0183\nEpoch 72/100 : Training loss: 2.3011 | Training Accuracy: 0.0167\nEpoch 73/100 : Training loss: 2.3010 | Training Accuracy: 0.0217\nEpoch 74/100 : Training loss: 2.3010 | Training Accuracy: 0.0267\nEpoch 75/100 : Training loss: 2.3010 | Training Accuracy: 0.0133\nEpoch 76/100 : Training loss: 2.3010 | Training Accuracy: 0.0250\nEpoch 77/100 : Training loss: 2.3010 | Training Accuracy: 0.0133\nEpoch 78/100 : Training loss: 2.3010 | Training Accuracy: 0.0183\nEpoch 79/100 : Training loss: 2.3009 | Training Accuracy: 0.0100\nEpoch 80/100 : Training loss: 2.3009 | Training Accuracy: 0.0133\nEpoch 81/100 : Training loss: 2.3009 | Training Accuracy: 0.0267\nEpoch 82/100 : Training loss: 2.3009 | Training Accuracy: 0.0183\nEpoch 83/100 : Training loss: 2.3009 | Training Accuracy: 0.0217\nEpoch 84/100 : Training loss: 2.3008 | Training Accuracy: 0.0200\nEpoch 85/100 : Training loss: 2.3008 | Training Accuracy: 0.0183\nEpoch 86/100 : Training loss: 2.3008 | Training Accuracy: 0.0233\nEpoch 87/100 : Training loss: 2.3008 | Training Accuracy: 0.0167\nEpoch 88/100 : Training loss: 2.3008 | Training Accuracy: 0.0183\nEpoch 89/100 : Training loss: 2.3008 | Training Accuracy: 0.0150\nEpoch 90/100 : Training loss: 2.3007 | Training Accuracy: 0.0183\nEpoch 91/100 : Training loss: 2.3007 | Training Accuracy: 0.0183\nEpoch 92/100 : Training loss: 2.3007 | Training Accuracy: 0.0183\nEpoch 93/100 : Training loss: 2.3007 | Training Accuracy: 0.0167\nEpoch 94/100 : Training loss: 2.3007 | Training Accuracy: 0.0217\nEpoch 95/100 : Training loss: 2.3007 | Training Accuracy: 0.0167\nEpoch 96/100 : Training loss: 2.3006 | Training Accuracy: 0.0200\nEpoch 97/100 : Training loss: 2.3006 | Training Accuracy: 0.0233\nEpoch 98/100 : Training loss: 2.3006 | Training Accuracy: 0.0133\nEpoch 99/100 : Training loss: 2.3006 | Training Accuracy: 0.0183\nEpoch 100/100 : Training loss: 2.3006 | Training Accuracy: 0.0333\n"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acbaebba4cb7d4f1007656790646406c153dc82
| 36,368 |
ipynb
|
Jupyter Notebook
|
Shell-exercices.ipynb
|
epfllibrary/2018-11-20-epfl
|
3d3fe8ded4dfc357d718daaaea6dc9aeb20d671a
|
[
"CC-BY-4.0"
] | 1 |
2018-07-06T07:59:00.000Z
|
2018-07-06T07:59:00.000Z
|
Shell-exercices.ipynb
|
epfllibrary/2018-11-20-epfl
|
3d3fe8ded4dfc357d718daaaea6dc9aeb20d671a
|
[
"CC-BY-4.0"
] | null | null | null |
Shell-exercices.ipynb
|
epfllibrary/2018-11-20-epfl
|
3d3fe8ded4dfc357d718daaaea6dc9aeb20d671a
|
[
"CC-BY-4.0"
] | null | null | null | 29.376414 | 489 | 0.558266 |
[
[
[
"# Software Carpentry\n\n### EPFL Library, November 2018\n",
"_____no_output_____"
],
[
"## Program\n\n| | 4 afternoons | 4 workshops |\n| :-- | :----------- | :---------- |\n| > | `Today` | `Unix Shell` |\n| | Thursday 22 | Version Control with Git |\n| | Tuesday 27 | Python I |\n| | Thursday 29 | More Python |",
"_____no_output_____"
],
[
"## Why did you decide to attend this workshop?",
"_____no_output_____"
],
[
"## Today's program\n\n| | activity |\n| :-- | :-------- |\n| 13:00 | Introducing the Unix Shell |\n| 13:15 | Navigating Files and Directories |\n| 14:00 | Working with Files and Directories |\n| 14:50 | **break** |\n| 15:20 | Loops |\n| 16:10 | Shell Scripts |\n| 16:55 | Finding Things |\n| 17:30 | Wrap-up / **END** |",
"_____no_output_____"
],
[
"## How we'll work\n\nLive coding \n\nSticky notes : use a red sticky note to say you are stuck, put the green one when all is good\n\nInstructors : Raphaël and Mathilde\n\nHelpers : Antoine, Ludovic, Raphaël and Mathilde\n\nSlides for exercices : Find the link to the slides on go.epfl.ch/swc-pad",
"_____no_output_____"
],
[
"## Introducing the Shell\n### Key points about the Shell \n\n\n- A shell is a program whose primary purpose is to read commands and run other programs.\n- The shell’s main advantages are its high action-to-keystroke ratio, its support for automating repetitive tasks, and its capacity to access networked machines.\n- The shell’s main disadvantages are its primarily textual nature and how cryptic its commands and operation can be.\n",
"_____no_output_____"
],
[
"## Navigating Files and Directories",
"_____no_output_____"
],
[
"### [Exercise] Exploring more `rm` flags\n\nWhat does the command ls do when used with the `-l` and `-h` flags?\n",
"_____no_output_____"
],
[
"### [Exercise] Listing Recursively and By Time\n\nThe command `ls -R` lists the contents of directories recursively, i.e., lists their sub-directories, sub-sub-directories, and so on at each level. The command `ls -t` lists things by time of last change, with most recently changed files or directories first. In what order does `ls -R -t` display things? ",
"_____no_output_____"
],
[
"### [Exercise] Absolute vs Relative Paths\n\nStarting from /Users/amanda/data/, which of the following commands could Amanda use to navigate to her home directory, which is /Users/amanda?\n\n 1. `cd .`\n 2. `cd /`\n 3. `cd /home/amanda`\n 4. `cd ../..`\n 5. `cd ~`\n 6. `cd home`\n 7. `cd ~/data/..`\n 8. `cd`\n 9. `cd ..`",
"_____no_output_____"
],
[
"### [Exercise] Relative Path Resolution\n\nUsing the filesystem diagram below, if pwd displays /Users/thing, what will ls -F ../backup display?\n\n 1. `../backup: No such file or directory`\n 2. `2012-12-01 2013-01-08 2013-01-27`\n 3. `2012-12-01/ 2013-01-08/ 2013-01-27/`\n 4. `original/ pnas_final/ pnas_sub/`\n \n\n\n",
"_____no_output_____"
],
[
"### [Exercise ] `ls` Reading comprehension \n\n\n\nAssuming a directory structure as in the above Figure (File System for Challenge Questions), if `pwd` displays `/Users/backup`, and `-r` tells ls to display things in reverse order, what command will display:\n\n`pnas_sub/ pnas_final/ original/`\n\n1. `ls pwd` \n2. `ls -r -F`\n3. `ls -r -F /Users/backup`\n4. Either #2 or #3 above, but not #1.",
"_____no_output_____"
],
[
"### Key Points about Navigating Files and Directories\n- The file system is responsible for managing information on the disk.\n- Information is stored in files, which are stored in directories (folders).\n- Directories can also store other directories, which forms a directory tree.\n- `cd path` changes the current working directory.\n- `ls path` prints a listing of a specific file or directory; `ls` on its own lists the current working directory.\n- `pwd` prints the user’s current working directory.\n- `/` on its own is the root directory of the whole file system.",
"_____no_output_____"
],
[
"### More key Points about Navigating Files and Directories\n- A relative path specifies a location starting from the current location.\n- An absolute path specifies a location from the root of the file system.\n- Directory names in a path are separated with `/` on Unix, but `\\`on Windows.\n- `..` means ‘the directory above the current one’; `.` on its own means ‘the current directory’.\n- Most files’ names are `something.extension`. The extension isn’t required, and doesn’t guarantee anything, but is normally used to indicate the type of data in the file.\n",
"_____no_output_____"
],
[
"## Working with Files and Directories",
"_____no_output_____"
],
[
"### [Exercise] Creating Files a Different Way\n\nWe have seen how to create text files using the `nano` editor. Now, try the following command in your home directory:\n\n```\n$ cd # go to your home directory\n$ touch my_file.txt\n```\n\n\n\n1. What did the touch command do? When you look at your home directory using the GUI file explorer, does the file show up?\n2. Use `ls -l` to inspect the files. How large is `my_file.txt`?\n3. When might you want to create a file this way?",
"_____no_output_____"
],
[
"### [Exercise] Using `rm` Safely\n\nWhat happens when we type `rm -i thesis/quotations.txt`? Why would we want this protection when using `rm`?",
"_____no_output_____"
],
[
"### [Exercise] Moving to the Current Folder\n\nAfter running the following commands, Jamie realizes that she put the files `sucrose.dat` and `maltose.dat` into the wrong folder:\n\n```\n$ ls -F\n```\n\n> analyzed/ raw/\n\n```\n$ ls -F analyzed\n```\n\n> fructose.dat glucose.dat maltose.dat sucrose.dat\n\n```\n$ cd raw/\n```\n\nFill in the blanks to move these files to the current folder (i.e., the one she is currently in):\n\n```\n$ mv ___/sucrose.dat ___/maltose.dat ___\n```",
"_____no_output_____"
],
[
"### [Exercise] Renaming Files\n\nSuppose that you created a `.txt` file in your current directory to contain a list of the statistical tests you will need to do to analyze your data, and named it: `statstics.txt`\n\nAfter creating and saving this file you realize you misspelled the filename! You want to correct the mistake, which of the following commands could you use to do so?\n\n1. `cp statstics.txt statistics.txt`\n2. `mv statstics.txt statistics.txt`\n3. `mv statstics.txt .`\n4. `cp statstics.txt .`\n",
"_____no_output_____"
],
[
"### [Exercise] Moving and Copying\n\nWhat is the output of the closing `ls` command in the sequence shown below?\n\n```\n$ pwd\n```\n\n> /Users/jamie/data\n\n```\n$ ls\n```\n> proteins.dat\n\n```\n$ mkdir recombine\n$ mv proteins.dat recombine/\n$ cp recombine/proteins.dat ../proteins-saved.dat\n$ ls\n```\n\n1. `proteins-saved.dat recombine`\n2. `recombine`\n3. `proteins.dat recombine`\n4. `proteins-saved.dat`\n",
"_____no_output_____"
],
[
"### Additional exercises\n\nIf you were quick, check out these exercises to dig a little more into details.",
"_____no_output_____"
],
[
"### [Exercise] Copy with Multiple Filenames\n\nFor this exercise, you can test the commands in the `data-shell/data` directory.\n\nIn the example below, what does `cp` do when given several filenames and a directory name?\n\n```\n$ mkdir backup\n$ cp amino-acids.txt animals.txt backup/\n```\n\nIn the example below, what does `cp` do when given three or more file names?\n\n```\n$ ls -F\n```\n\n> amino-acids.txt animals.txt backup/ elements/ morse.txt pdb/ planets.txt salmon.txt sunspot.txt\n \n```\n$ cp amino-acids.txt animals.txt morse.txt\n```",
"_____no_output_____"
],
[
"### [Exercise] Using Wildcards\n\nWhen run in the `molecules` directory, which `ls` command(s) will produce this output?\n\n`ethane.pdb methane.pdb`\n\n1. `ls *t*ane.pdb`\n2. `ls *t?ne.*`\n3. `ls *t??ne.pdb`\n4. `ls ethane.*`",
"_____no_output_____"
],
[
"### [Exercise] More on Wildcards\n\nSam has a directory containing calibration data, datasets, and descriptions of the datasets:\n\n```\n2015-10-23-calibration.txt\n2015-10-23-dataset1.txt\n2015-10-23-dataset2.txt\n2015-10-23-dataset_overview.txt\n2015-10-26-calibration.txt\n2015-10-26-dataset1.txt\n2015-10-26-dataset2.txt\n2015-10-26-dataset_overview.txt\n2015-11-23-calibration.txt\n2015-11-23-dataset1.txt\n2015-11-23-dataset2.txt\n2015-11-23-dataset_overview.txt\n```\n\nBefore heading off to another field trip, she wants to back up her data and send some datasets to her colleague Bob. Sam uses the following commands to get the job done:\n\n```\n$ cp *dataset* /backup/datasets\n$ cp ____calibration____ /backup/calibration\n$ cp 2015-____-____ ~/send_to_bob/all_november_files/\n$ cp ____ ~/send_to_bob/all_datasets_created_on_a_23rd/\n```\n\nHelp Sam by filling in the blanks.",
"_____no_output_____"
],
[
"### [Exercise] Organizing Directories and Files\n\nJamie is working on a project and she sees that her files aren’t very well organized:\n\n```\n$ ls -F\n```\n\n> analyzed/ fructose.dat raw/ sucrose.dat\n\nThe `fructose.dat` and sucrose.dat` files contain output from her data analysis. What command(s) covered in this lesson does she need to run so that the commands below will produce the output shown?\n\n```\n$ ls -F\n```\n\n> analyzed/ raw/\n\n```\n$ ls analyzed\n```\n\n> fructose.dat sucrose.dat\n\n\n",
"_____no_output_____"
],
[
"### [Exercise] Copy a folder structure but not the files\n\nYou’re starting a new experiment, and would like to duplicate the file structure from your previous experiment without the data files so you can add new data.\n\nAssume that the file structure is in a folder called ‘2016-05-18-data’, which contains a `data` folder that in turn contains folders named `raw` and `processed` that contain data files. The goal is to copy the file structure of the `2016-05-18-data` folder into a folder called `2016-05-20-data` and remove the data files from the directory you just created.\n\nWhich of the following set of commands would achieve this objective? What would the other commands do?\n\n```\n$ cp -r 2016-05-18-data/ 2016-05-20-data/\n$ rm 2016-05-20-data/raw/*\n$ rm 2016-05-20-data/processed/*\n```\n\n```\n$ rm 2016-05-20-data/raw/*\n$ rm 2016-05-20-data/processed/*\n$ cp -r 2016-05-18-data/ 2016-5-20-data/\n```\n\n```\n$ cp -r 2016-05-18-data/ 2016-05-20-data/\n$ rm -r -i 2016-05-20-data/\n```",
"_____no_output_____"
],
[
"### Key Points about Working with Files and Directories\n\n`cp old new` copies a file. \n`mkdir path` creates a new directory. \n`mv old new` moves (renames) a file or directory. \n`rm path` removes (deletes) a file. \n`*` matches zero or more characters in a filename, so `*.txt` matches all files ending in `.txt`. \n`?` matches any single character in a filename, so `?.txt` matches `a.txt` but not `any.txt`. \nUse of the Control key may be described in many ways, including `Ctrl-X`, `Control-X`, and `^X`. \nThe shell does not have a trash bin: once something is deleted, it’s really gone. \nDepending on the type of work you do, you may need a more powerful text editor than Nano. \n",
"_____no_output_____"
],
[
"## Pipes and Filters\n\n### Key Points about Pipes and Filters\n`cat` displays the contents of its inputs.\n\n`head` displays the first 10 lines of its input.\n\n`tail` displays the last 10 lines of its input.\n\n`sort` sorts its inputs.\n\n`wc` counts lines, words, and characters in its inputs.\n\n`command > file` redirects a command’s output to a file.\n\n`first | second` is a pipeline: the output of the first command is used as the input to the second.\n\n[More information on this topic](https://swcarpentry.github.io/shell-novice/04-pipefilter/index.html) on the Software Carpentry website",
"_____no_output_____"
],
[
"## Loops",
"_____no_output_____"
],
[
"### [Exercise] Variables in Loops\n\nThis exercise refers to the `data-shell/molecules` directory. `ls` gives the following output:\n\n> cubane.pdb ethane.pdb methane.pdb octane.pdb pentane.pdb propane.pdb\n\nWhat is the output of the following code?\n\n```\n$ for datafile in *.pdb\n> do\n> ls *.pdb\n> done\n```\n\nNow, what is the output of the following code?\n\n```\n$ for datafile in *.pdb\n> do\n>\tls $datafile\n> done\n```\n\nWhy do these two loops give different outputs?",
"_____no_output_____"
],
[
"### [Exercise] Limiting Sets of Files\n\nWhat would be the output of running the following loop in the `data-shell/molecules` directory?\n\n```\n$ for filename in c*\n> do\n> ls $filename \n> done\n```\n\n1. No files are listed.\n2. All files are listed.\n3. Only `cubane.pdb`, `octane.pdb` and `pentane.pdb` are listed.\n4. Only `cubane.pdb` is listed.\n",
"_____no_output_____"
],
[
"### [Exercise] Limiting Sets of Files (part 2)\n\nHow would the output differ from using this command instead?\n\n```\n$ for filename in *c*\n> do\n> ls $filename \n> done\n```\n\n1. The same files would be listed.\n2. All the files are listed this time.\n3. No files are listed this time.\n4. The files `cubane.pdb` and `octane.pdb` will be listed.\n5. Only the file `octane.pdb` will be listed.\n",
"_____no_output_____"
],
[
"### [Exercise] Saving to a File in a Loop - Part One\n\nIn the `data-shell/molecules` directory, what is the effect of this loop?\n\n```\n$ for alkanes in *.pdb\n> do\n> echo $alkanes\n> cat $alkanes > alkanes.pdb\n> done\n```\n\n1. Prints `cubane.pdb`, `ethane.pdb`, `methane.pdb`, `octane.pdb`, `pentane.pdb` and `propane.pdb`, and the text from `propane.pdb` will be saved to a file called `alkanes.pdb`.\n2. Prints `cubane.pdb`, `ethane.pdb`, and `methane.pdb`, and the text from all three files would be concatenated and saved to a file called `alkanes.pdb`.\n3. Prints `cubane.pdb`, `ethane.pdb`, `methane.pdb`, `octane.pdb`, and `pentane.pdb`, and the text from `propane.pdb` will be saved to a file called `alkanes.pdb`.\n4. None of the above.\n",
"_____no_output_____"
],
[
"### [Exercise] Saving to a File in a Loop - Part Two\n\nAlso in the `data-shell/molecules` directory, what would be the output of the following loop?\n\n```\n$ for datafile in *.pdb\n> do\n> cat $datafile >> all.pdb\n> done\n```\n\n1. All of the text from `cubane.pdb`, `ethane.pdb`, `methane.pdb`, `octane.pdb`, and `pentane.pdb` would be concatenated and saved to a file called `all.pdb`.\n2. The text from `ethane.pdb` will be saved to a file called `all.pdb`.\n3. All of the text from `cubane.pdb`, `ethane.pdb`, `methane.pdb`, `octane.pdb`, `pentane.pdb` and `propane.pdb` would be concatenated and saved to a file called `all.pdb`.\n4. All of the text from `cubane.pdb`, `ethane.pdb`, `methane.pdb`, `octane.pdb`, `pentane.pdb` and `propane.pdb` would be printed to the screen and saved to a file called `all.pdb`.\n",
"_____no_output_____"
],
[
"### Additional exercises\n\nIf you were quick, check out these exercises to dig a little more into details.",
"_____no_output_____"
],
[
"### [Exercise] Doing a Dry Run\n\nA loop is a way to do many things at once — or to make many mistakes at once if it does the wrong thing. One way to check what a loop would do is to `echo` the commands it would run instead of actually running them.\n\nSuppose we want to preview the commands the following loop will execute without actually running those commands:\n\n```\n$ for file in *.pdb\n> do\n> analyze $file > analyzed-$file\n> done\n```\n\nWhat is the difference between the two loops below, and which one would we want to run?\n\n```\n# Version 1\n$ for file in *.pdb\n> do\n> echo analyze $file > analyzed-$file\n> done\n```\n\n```\n# Version 2\n$ for file in *.pdb\n> do\n> echo \"analyze $file > analyzed-$file\"\n> done\n```\n",
"_____no_output_____"
],
[
"### [Exercise] Nested Loops\n\nSuppose we want to set up up a directory structure to organize some experiments measuring reaction rate constants with different compounds *and* different temperatures. What would be the result of the following code:\n\n```\n$ for species in cubane ethane methane\n> do\n> for temperature in 25 30 37 40\n> do\n> mkdir $species-$temperature\n> done\n> done\n```\n",
"_____no_output_____"
],
[
"### Key Points for Loops \n\n* A `for` loop repeats commands once for every thing in a list.\n* Every `for` loop needs a variable to refer to the thing it is currently operating on.\n* Use `$name` to expand a variable (i.e., get its value). `${name}` can also be used.\n* Do not use spaces, quotes, or wildcard characters such as ‘\\*’ or ‘?’ in filenames, as it complicates variable expansion.\n* Give files consistent names that are easy to match with wildcard patterns to make it easy to select them for looping.\n* Use the up-arrow key to scroll up through previous commands to edit and repeat them.\n* Use `Ctrl-R` to search through the previously entered commands.\n* Use `history` to display recent commands, and `!number` to repeat a command by number.\n",
"_____no_output_____"
],
[
"## Shell Scripts",
"_____no_output_____"
],
[
"### [Exercise] List Unique Species\n\nLeah has several hundred data files, each of which is formatted like this:\n```\n2013-11-05,deer,5\n2013-11-05,rabbit,22\n2013-11-05,raccoon,7\n2013-11-06,rabbit,19\n2013-11-06,deer,2\n2013-11-06,fox,1\n2013-11-07,rabbit,18\n2013-11-07,bear,1\n```\nAn example of this type of file is given in `data-shell/data/animal-counts/animals.txt`.\n\nWrite a shell script called `species.sh` that takes any number of filenames as command-line arguments, and uses `cut`, `sort`, and `uniq` to print a list of the unique species appearing in each of those files separately.\n\n",
"_____no_output_____"
],
[
"### [Exercise] Why Record Commands in the History Before Running Them?\n\nIf you run the command:\n\n```\n$ history | tail -n 5 > recent.sh\n```\n\nthe last command in the file is the `history` command itself, i.e., the shell has added `history` to the command log before actually running it. In fact, the shell always adds commands to the log before running them. Why do you think it does this?\n\n",
"_____no_output_____"
],
[
"### [Exercise] Variables in Shell Scripts\n\nIn the `molecules` directory, imagine you have a shell script called `script.sh` containing the following commands:\n\n```\nhead -n $2 $1\ntail -n $3 $1\n```\n\nWhile you are in the `molecules` directory, you type the following command:\n`bash script.sh '*.pdb' 1 1`\n\nWhich of the following outputs would you expect to see?\n\n1. All of the lines between the first and the last lines of each file ending in `.pdb` in the `molecules` directory\n2. The first and the last line of each file ending in `.pdb` in the `molecules directory\n3. The first and the last line of each file in the `molecules` directory\n4. An error because of the quotes around `*.pdb`\n\n",
"_____no_output_____"
],
[
"### [Exercise] Find the Longest File With a Given Extension\n\nWrite a shell script called `longest.sh` that takes the name of a directory and a filename extension as its arguments, and prints out the name of the file with the most lines in that directory with that extension. For example:\n\n`$ bash longest.sh /tmp/data pdb`\n\n\nwould print the name of the `.pdb` file in `/tmp/data` that has the most lines.\n",
"_____no_output_____"
],
[
"### Additional exercises\n\nIf you were quick, check out these exercises to dig a little more into details.",
"_____no_output_____"
],
[
"### [Exercise] Script Reading Comprehension\n\nFor this question, consider the `data-shell/molecules` directory once again. This contains a number of `.pdb` files in addition to any other files you may have created. Explain what a script called `example.sh` would do when run as `bash example.sh *.pdb` if it contained the following lines:\n```\n\n# Script 1\necho *.*\n\n# Script 2\nfor filename in $1 $2 $3\ndo\n cat $filename\ndone\n\n# Script 3\necho [email protected]\n\n",
"_____no_output_____"
],
[
"### [Exercise] Debugging Scripts\n\nSuppose you have saved the following script in a file called do-errors.sh in Nelle’s north-pacific-gyre/2012-07-03 directory:\n\n```\n# Calculate stats for data files.\nfor datafile in \"$@\"\ndo\n echo $datfile\n bash goostats $datafile stats-$datafile\ndone\n```\n\nWhen you run it:\n\n`$ bash do-errors.sh NENE*[AB].txt`\n\n\nthe output is blank. To figure out why, re-run the script using the -x option:\n\n`bash -x do-errors.sh NENE*[AB].txt`\n\nWhat is the output showing you? Which line is responsible for the error?\n",
"_____no_output_____"
],
[
"## Finding Things",
"_____no_output_____"
],
[
"### [Exercise] Using `grep`\n\nWhich command would result in the following output:\n\n```\nand the presence of absence:\n```\n\n1. `grep \"of\" haiku.txt`\n2. `grep -E \"of\" haiku.txt`\n3. `grep -w \"of\" haiku.txt`\n4. `grep -i \"of\" haiku.txt`\n",
"_____no_output_____"
],
[
"### [Exercise] Tracking a Species\n\nLeah has several hundred data files saved in one directory, each of which is formatted like this:\n\n```\n2013-11-05,deer,5\n2013-11-05,rabbit,22\n2013-11-05,raccoon,7\n2013-11-06,rabbit,19\n2013-11-06,deer,2\n```\n\nShe wants to write a shell script that takes a species as the first command-line argument and a directory as the second argument. The script should return one file called `species.txt` containing a list of dates and the number of that species seen on each date. For example using the data shown above, `rabbit.txt` would contain:\n\n```\n2013-11-05,22\n2013-11-06,19\n```\n\nPut these commands and pipes in the right order to achieve this:\n\n```\ncut -d : -f 2 \n> \n| \ngrep -w $1 -r $2 \n| \n$1.txt \ncut -d , -f 1,3 \n```\n\nHint: use `man grep` to look for how to grep text recursively in a directory and `man cut` to select more than one field in a line.\n\nAn example of such a file is provided in `data-shell/data/animal-counts/animals.txt`.",
"_____no_output_____"
],
[
"### [Exercise] Little Women\n\nYou and your friend, having just finished reading *Little Women* by Louisa May Alcott, are in an argument. Of the four sisters in the book, Jo, Meg, Beth, and Amy, your friend thinks that Jo was the most mentioned. You, however, are certain it was Amy. Luckily, you have a file `LittleWomen.txt` containing the full text of the novel (`data-shell/writing/data/LittleWomen.txt`). Using a `for` loop, how would you tabulate the number of times each of the four sisters is mentioned?\n\nHint: one solution might employ the commands `grep` and `wc` and a `|`, while another might utilize `grep` options. There is often more than one way to solve a programming task, so a particular solution is usually chosen based on a combination of yielding the correct result, elegance, readability, and speed.",
"_____no_output_____"
],
[
"### Additional exercises\n\nIf you were quick, check out these exercises to dig a little more into details.",
"_____no_output_____"
],
[
"### [Exercise] Matching and Subtracting\n\nThe `-v` flag to `grep` inverts pattern matching, so that only lines which do not match the pattern are printed. Given that, which of the following commands will find all files in `/data` whose names end in `s.txt` (e.g., `animals.txt` or `planets.txt`), but do not contain the word `net`? Once you have thought about your answer, you can test the commands in the `data-shell` directory.\n\n1. `find data -name '*s.txt' | grep -v net`\n2. `find data -name *s.txt | grep -v net`\n3. `grep -v \"temp\" $(find data -name '*s.txt')`\n4. None of the above.\n",
"_____no_output_____"
],
[
"### [Exercise] `find` Pipeline Reading Comprehension\n\nWrite a short explanatory comment for the following shell script:\n\n```\nwc -l $(find . -name '*.dat') | sort -n\n```\n",
"_____no_output_____"
],
[
"### [Exercise] Finding Files With Different Properties\n\nThe `find` command can be given several other criteria known as “tests” to locate files with specific attributes, such as creation time, size, permissions, or ownership. Use `man find` to explore these, and then write a single command to find all files in or below the current directory that were modified by the user `ahmed` in the last 24 hours.\n\nHint 1: you will need to use three tests: `-type`, `-mtime`, and `-user`.\n\nHint 2: The value for `-mtime` will need to be negative—why?",
"_____no_output_____"
],
[
"### Key Points for Finding Things\n\n* `find` finds files with specific properties that match patterns.\n* `grep` selects lines in files that match patterns.\n* `--help` is a flag supported by many bash commands, and programs that can be run from within Bash, to display more information on how to use these commands or programs.\n* `man` command displays the manual page for a given command.\n* `$(command)` inserts a command’s output in place.\n",
"_____no_output_____"
],
[
"### Additional resources\n\n| resource | description |\n| :------- | :---------- |\n| https://explainshell.com | dissects any shell command you type in |\n| https://tldr.sh | simplified and community-driven Shell manual pages |\n| https://www.shellcheck.net | checks shell scripts for common errors |\n",
"_____no_output_____"
]
]
] |
[
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
4acbafc396d26cf989357685452bf66500f1ed45
| 38,143 |
ipynb
|
Jupyter Notebook
|
data_edit.ipynb
|
Jroc561/spotify_suggester
|
5c1cec8af3969866a8bb762f1d477ccb8fac7503
|
[
"MIT"
] | null | null | null |
data_edit.ipynb
|
Jroc561/spotify_suggester
|
5c1cec8af3969866a8bb762f1d477ccb8fac7503
|
[
"MIT"
] | 2 |
2021-02-03T18:08:18.000Z
|
2021-02-04T16:38:37.000Z
|
data_edit.ipynb
|
Jroc561/spotify_suggester
|
5c1cec8af3969866a8bb762f1d477ccb8fac7503
|
[
"MIT"
] | 1 |
2021-06-17T01:11:22.000Z
|
2021-06-17T01:11:22.000Z
| 34.394049 | 219 | 0.359148 |
[
[
[
"import numpy as np\nimport pandas as pd",
"_____no_output_____"
],
[
"data = pd.read_csv(\"data/data.csv\")",
"_____no_output_____"
],
[
"data.head(5)",
"_____no_output_____"
],
[
"data['artists'] = data['artists'].str.strip('['']')",
"_____no_output_____"
],
[
"data.head()",
"_____no_output_____"
],
[
"data['artists']=data['artists'].str.strip('\"\"')",
"_____no_output_____"
],
[
"data['artists']=data['artists'].str.replace(\"'\", \"\")",
"_____no_output_____"
],
[
"data['combined'] = data['name'] + ' - ' + data['artists']",
"_____no_output_____"
],
[
"data.head(5)",
"_____no_output_____"
],
[
"data[data['name']=='SICKO MODE']",
"_____no_output_____"
],
[
"data.drop_duplicates(subset=['combined'], inplace=True)",
"_____no_output_____"
],
[
"data[data['combined']=='SICKO MODE - TRAVIS SCOTT']",
"_____no_output_____"
],
[
"data.drop('release_date', axis=1, inplace=True)",
"_____no_output_____"
],
[
"data.head()",
"_____no_output_____"
],
[
"data.to_csv(\"edited_data_v2.csv\")",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acbb2b434a17e3737ec49261150debd2178d29b
| 22,236 |
ipynb
|
Jupyter Notebook
|
Model backlog/Inference/114-tweet-inference-5fold-roberta-base-3epochs-cos.ipynb
|
dimitreOliveira/Tweet-Sentiment-Extraction
|
0a775abe9a92c4bc2db957519c523be7655df8d8
|
[
"MIT"
] | 11 |
2020-06-17T07:30:20.000Z
|
2022-03-25T16:56:01.000Z
|
Model backlog/Inference/114-tweet-inference-5fold-roberta-base-3epochs-cos.ipynb
|
dimitreOliveira/Tweet-Sentiment-Extraction
|
0a775abe9a92c4bc2db957519c523be7655df8d8
|
[
"MIT"
] | null | null | null |
Model backlog/Inference/114-tweet-inference-5fold-roberta-base-3epochs-cos.ipynb
|
dimitreOliveira/Tweet-Sentiment-Extraction
|
0a775abe9a92c4bc2db957519c523be7655df8d8
|
[
"MIT"
] | null | null | null | 31.765714 | 150 | 0.433711 |
[
[
[
"## Dependencies",
"_____no_output_____"
]
],
[
[
"import json, glob\nfrom tweet_utility_scripts import *\nfrom tweet_utility_preprocess_roberta_scripts import *\nfrom transformers import TFRobertaModel, RobertaConfig\nfrom tokenizers import ByteLevelBPETokenizer\nfrom tensorflow.keras import layers\nfrom tensorflow.keras.models import Model",
"_____no_output_____"
]
],
[
[
"# Load data",
"_____no_output_____"
]
],
[
[
"test = pd.read_csv('/kaggle/input/tweet-sentiment-extraction/test.csv')\n\nprint('Test samples: %s' % len(test))\ndisplay(test.head())",
"Test samples: 3534\n"
]
],
[
[
"# Model parameters",
"_____no_output_____"
]
],
[
[
"input_base_path = '/kaggle/input/114roberta-base/'\nwith open(input_base_path + 'config.json') as json_file:\n config = json.load(json_file)\n\nconfig",
"_____no_output_____"
],
[
"# vocab_path = input_base_path + 'vocab.json'\n# merges_path = input_base_path + 'merges.txt'\nbase_path = '/kaggle/input/qa-transformers/roberta/'\n\nvocab_path = base_path + 'roberta-base-vocab.json'\nmerges_path = base_path + 'roberta-base-merges.txt'\nconfig['base_model_path'] = base_path + 'roberta-base-tf_model.h5'\nconfig['config_path'] = base_path + 'roberta-base-config.json'\n\nmodel_path_list = glob.glob(input_base_path + '*.h5')\nmodel_path_list.sort()\nprint('Models to predict:')\nprint(*model_path_list, sep = \"\\n\")",
"Models to predict:\n/kaggle/input/114roberta-base/model_fold_1.h5\n/kaggle/input/114roberta-base/model_fold_2.h5\n/kaggle/input/114roberta-base/model_fold_3.h5\n/kaggle/input/114roberta-base/model_fold_4.h5\n/kaggle/input/114roberta-base/model_fold_5.h5\n"
]
],
[
[
"# Tokenizer",
"_____no_output_____"
]
],
[
[
"tokenizer = ByteLevelBPETokenizer(vocab_file=vocab_path, merges_file=merges_path, \n lowercase=True, add_prefix_space=True)",
"_____no_output_____"
]
],
[
[
"# Pre process",
"_____no_output_____"
]
],
[
[
"test['text'].fillna('', inplace=True)\ntest[\"text\"] = test[\"text\"].apply(lambda x: x.lower())\ntest[\"text\"] = test[\"text\"].apply(lambda x: x.strip())\n\nx_test = get_data_test(test, tokenizer, config['MAX_LEN'], preprocess_fn=preprocess_roberta_test)",
"_____no_output_____"
]
],
[
[
"# Model",
"_____no_output_____"
]
],
[
[
"module_config = RobertaConfig.from_pretrained(config['config_path'], output_hidden_states=False)\n\ndef model_fn(MAX_LEN):\n input_ids = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='input_ids')\n attention_mask = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='attention_mask')\n \n base_model = TFRobertaModel.from_pretrained(config['base_model_path'], config=module_config, name=\"base_model\")\n last_hidden_state, _ = base_model({'input_ids': input_ids, 'attention_mask': attention_mask})\n\n x = layers.Dropout(.1)(last_hidden_state)\n \n x_start = layers.Dense(1)(x)\n x_start = layers.Flatten()(x_start)\n y_start = layers.Activation('softmax', name='y_start')(x_start)\n\n x_end = layers.Dense(1)(x)\n x_end = layers.Flatten()(x_end)\n y_end = layers.Activation('softmax', name='y_end')(x_end)\n \n model = Model(inputs=[input_ids, attention_mask], outputs=[y_start, y_end])\n \n return model",
"_____no_output_____"
]
],
[
[
"# Make predictions",
"_____no_output_____"
]
],
[
[
"NUM_TEST_IMAGES = len(test)\ntest_start_preds = np.zeros((NUM_TEST_IMAGES, config['MAX_LEN']))\ntest_end_preds = np.zeros((NUM_TEST_IMAGES, config['MAX_LEN']))\n\nfor model_path in model_path_list:\n print(model_path)\n model = model_fn(config['MAX_LEN'])\n model.load_weights(model_path)\n \n test_preds = model.predict(x_test) \n test_start_preds += test_preds[0]\n test_end_preds += test_preds[1]",
"/kaggle/input/114roberta-base/model_fold_1.h5\n/kaggle/input/114roberta-base/model_fold_2.h5\n/kaggle/input/114roberta-base/model_fold_3.h5\n/kaggle/input/114roberta-base/model_fold_4.h5\n/kaggle/input/114roberta-base/model_fold_5.h5\n"
]
],
[
[
"# Post process",
"_____no_output_____"
]
],
[
[
"test['start'] = test_start_preds.argmax(axis=-1)\ntest['end'] = test_end_preds.argmax(axis=-1)\n\ntest['text_len'] = test['text'].apply(lambda x : len(x))\ntest['text_wordCnt'] = test['text'].apply(lambda x : len(x.split(' ')))\ntest[\"end\"].clip(0, test[\"text_len\"], inplace=True)\ntest[\"start\"].clip(0, test[\"end\"], inplace=True)\n\ntest['selected_text'] = test.apply(lambda x: decode(x['start'], x['end'], x['text'], config['question_size'], tokenizer), axis=1)\ntest[\"selected_text\"].fillna(test[\"text\"], inplace=True)",
"_____no_output_____"
]
],
[
[
"# Visualize predictions",
"_____no_output_____"
]
],
[
[
"display(test.head(10))",
"_____no_output_____"
]
],
[
[
"# Test set predictions",
"_____no_output_____"
]
],
[
[
"submission = pd.read_csv('/kaggle/input/tweet-sentiment-extraction/sample_submission.csv')\nsubmission['selected_text'] = test[\"selected_text\"]\nsubmission.to_csv('submission.csv', index=False)\nsubmission.head(10)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4acbc486798cbcae66d4d718fffd00ce0b527601
| 4,287 |
ipynb
|
Jupyter Notebook
|
docs/examples/data_visualization/publish_and_share/publish_visualization_gdf.ipynb
|
CartoDB/cartoframes
|
7c7392be5d15d0472ff428546c4791ed1a3842b0
|
[
"BSD-3-Clause"
] | 236 |
2017-02-28T15:17:09.000Z
|
2021-12-28T23:29:38.000Z
|
docs/examples/data_visualization/publish_and_share/publish_visualization_gdf.ipynb
|
CartoDB/cartoframes
|
7c7392be5d15d0472ff428546c4791ed1a3842b0
|
[
"BSD-3-Clause"
] | 1,332 |
2017-02-27T15:34:52.000Z
|
2022-03-07T01:31:10.000Z
|
docs/examples/data_visualization/publish_and_share/publish_visualization_gdf.ipynb
|
CartoDB/cartoframes
|
7c7392be5d15d0472ff428546c4791ed1a3842b0
|
[
"BSD-3-Clause"
] | 63 |
2017-03-14T19:02:37.000Z
|
2021-08-03T03:36:05.000Z
| 24.08427 | 272 | 0.5547 |
[
[
[
"## Publish a visualization from a GeoDataFrame\n\nThis example illustrate how to publish a visualization using a GeoDataFrame.\n\nRead more about Maps API Keys: https://carto.com/developers/auth-api/guides/types-of-API-Keys/\n\n>_Note: You'll need [CARTO Account](https://carto.com/signup) credentials to reproduce this example._",
"_____no_output_____"
]
],
[
[
"from cartoframes.auth import set_default_credentials\n\nset_default_credentials('creds.json')",
"_____no_output_____"
],
[
"from geopandas import read_file\n\ngdf = read_file('http://libs.cartocdn.com/cartoframes/files/sustainable_palm_oil_production_mills.geojson')",
"_____no_output_____"
],
[
"from cartoframes.viz import Map, Layer\n\nmap_viz = Map(Layer(gdf))",
"_____no_output_____"
],
[
"map_viz.publish(\n name='map_gdf',\n password='1234',\n if_exists='replace')",
"_____no_output_____"
]
],
[
[
"### Recommendation\n\nThe performance of visualizations can be affected by using large GeoDataFrames. There is also a size limit of 10MB per visualization published. Our recommendation in these cases is to upload the data to a table in CARTO, and use the table as the source in the Layer.",
"_____no_output_____"
]
],
[
[
"from cartoframes import to_carto\n\nto_carto(gdf, 'table_name', if_exists='replace')",
"Success! Data uploaded to table \"table_name\" correctly\n"
],
[
"map_viz = Map(Layer('table_name'))",
"_____no_output_____"
],
[
"map_viz.publish(\n name='map_table',\n password='1234',\n if_exists='replace')",
"The map has been published. The \"cartoframes_0a8a065b2cc026c5c33ee3dc269afcf1\" Maps API key with value \"YTAEiju-Utii8mgtERiEkA\" is being used for the datasets: \"table_name\". You can manage your API keys on your account.\n"
]
]
] |
[
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
]
] |
4acbc8a8e87f4620151b786a3d4af079989770e2
| 530,860 |
ipynb
|
Jupyter Notebook
|
notebooks/Redshift.ipynb
|
DoubleChimera/MachineLearningStatistics
|
3b3d5b35910e1b27a33ce580d1f3a878aa59207c
|
[
"BSD-3-Clause"
] | null | null | null |
notebooks/Redshift.ipynb
|
DoubleChimera/MachineLearningStatistics
|
3b3d5b35910e1b27a33ce580d1f3a878aa59207c
|
[
"BSD-3-Clause"
] | null | null | null |
notebooks/Redshift.ipynb
|
DoubleChimera/MachineLearningStatistics
|
3b3d5b35910e1b27a33ce580d1f3a878aa59207c
|
[
"BSD-3-Clause"
] | null | null | null | 319.21828 | 138,616 | 0.923451 |
[
[
[
"# Machine Learning and Statistics for Physicists",
"_____no_output_____"
],
[
"Material for a [UC Irvine](https://uci.edu/) course offered by the [Department of Physics and Astronomy](https://www.physics.uci.edu/).\n\nContent is maintained on [github](github.com/dkirkby/MachineLearningStatistics) and distributed under a [BSD3 license](https://opensource.org/licenses/BSD-3-Clause).\n\n##### ► [View table of contents](Contents.ipynb)",
"_____no_output_____"
]
],
[
[
"%matplotlib inline\nimport matplotlib.pyplot as plt\nimport seaborn as sns; sns.set()\nimport numpy as np\nimport pandas as pd",
"_____no_output_____"
],
[
"from mls import locate_data",
"_____no_output_____"
],
[
"from sklearn import model_selection, neighbors, tree, ensemble",
"_____no_output_____"
],
[
"import scipy.stats",
"_____no_output_____"
]
],
[
[
"## Case Study: Redshift Inference",
"_____no_output_____"
],
[
"Our goal is to predict the [cosmological redshift](https://en.wikipedia.org/wiki/Redshift) of a galaxy based on its brightness measured through 17 different filters. Redshift is a proxy for distance or, equivalently, look back time, so is a key observable for learning about past conditions in the universe.",
"_____no_output_____"
],
[
"### Load and Explore Data",
"_____no_output_____"
],
[
"Read the data to train and test on:",
"_____no_output_____"
]
],
[
[
"X = pd.read_hdf(locate_data('photoz_data.hf5'))\ny = pd.read_hdf(locate_data('photoz_targets.hf5'))",
"_____no_output_____"
],
[
"X.describe()",
"_____no_output_____"
],
[
"y.describe()",
"_____no_output_____"
],
[
"sns.pairplot(X[:500], vars=X.columns.tolist()[:6]);",
"_____no_output_____"
],
[
"plt.hist(y['Z'], bins=np.arange(0, 6, 0.2))\nplt.xlabel('Redshift $z$');\nplt.ylabel('Galaxies / ($\\Delta z=0.2$)');",
"_____no_output_____"
]
],
[
[
"### Split Data Randomly into Training and Testing Subsamples",
"_____no_output_____"
]
],
[
[
"gen = np.random.RandomState(seed=123)\nX_train, X_test, y_train, y_test = model_selection.train_test_split(X, y, test_size=0.25, random_state=gen)",
"_____no_output_____"
],
[
"print(f'{len(X)} = {len(X_train)} TRAIN + {len(X_test)} TEST')",
"17366 = 13024 TRAIN + 4342 TEST\n"
]
],
[
[
"### Nearest Neighbor Regression",
"_____no_output_____"
],
[
"Use the K-nearest neighbors (KNN) of an input sample to estimate its properties with [KNeighborsRegressor](https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.KNeighborsRegressor.html#sklearn.neighbors.KNeighborsRegressor):",
"_____no_output_____"
]
],
[
[
"knn_fit = neighbors.KNeighborsRegressor(n_jobs=8).fit(X_train, y_train)",
"_____no_output_____"
]
],
[
[
"Scores are calculated using the [coefficient of determination](https://en.wikipedia.org/wiki/Coefficient_of_determination) $R^2$, for which perfect accuracy is $R^2 = 1$:",
"_____no_output_____"
]
],
[
[
"knn_fit.score(X_train, y_train), knn_fit.score(X_test, y_test)",
"_____no_output_____"
],
[
"knn_fit.n_neighbors",
"_____no_output_____"
]
],
[
[
"#### Hyperparameter Optimization",
"_____no_output_____"
],
[
"The main hyperparameter is the value of K: the number of nearest neighbors that contribute to the final decision.",
"_____no_output_____"
]
],
[
[
"def knn_study(n=(1, 2, 4, 6, 8, 12, 16), max_score_samples=2000):\n train_score, test_score = [], []\n for n_neighbors in n:\n fit = neighbors.KNeighborsRegressor(n_neighbors=n_neighbors, n_jobs=8).fit(X_train, y_train)\n train_score.append(fit.score(X_train[:max_score_samples], y_train[:max_score_samples]))\n test_score.append(fit.score(X_test[:max_score_samples], y_test[:max_score_samples]))\n plt.plot(n, train_score, 'rx-', label='TRAIN')\n plt.plot(n, test_score, 'bo-', label='TEST')\n plt.xlabel('KNN n_neighbors')\n plt.ylabel('KNN $R^2$ score')\n plt.legend()",
"_____no_output_____"
],
[
"knn_study()",
"_____no_output_____"
]
],
[
[
"### Decision Tree Regression",
"_____no_output_____"
],
[
"Use a [binary decision tree](https://en.wikipedia.org/wiki/Decision_tree_learning) to sort each input sample into a small \"peer group\" with [DecisionTreeRegressor](https://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeRegressor.html). Note the similarity with KNN, except now we ask a set of questions to identify the \"peer group\", instead of using nearest neighbors.",
"_____no_output_____"
]
],
[
[
"gen = np.random.RandomState(seed=123)\ntree_fit = tree.DecisionTreeRegressor(random_state=gen).fit(X_train, y_train)",
"_____no_output_____"
],
[
"tree_fit.score(X_train, y_train), tree_fit.score(X_test, y_test)",
"_____no_output_____"
],
[
"tree_fit.tree_.max_depth",
"_____no_output_____"
]
],
[
[
"#### Feature Importance",
"_____no_output_____"
]
],
[
[
"importance = pd.DataFrame(\n {'feature': X.columns, 'importance': tree_fit.feature_importances_}\n).sort_values(by='importance', ascending=False)",
"_____no_output_____"
],
[
"importance.plot('feature', 'importance', 'barh', figsize=(10, 10), legend=False);",
"_____no_output_____"
]
],
[
[
"Re-train using only the 8 most important features:",
"_____no_output_____"
]
],
[
[
"importance[:8]",
"_____no_output_____"
],
[
"best_features = importance[:8]['feature']",
"_____no_output_____"
]
],
[
[
"The re-trained tree is much simpler and almost equally accurate on the test data:",
"_____no_output_____"
]
],
[
[
"tree_fit = tree.DecisionTreeRegressor(random_state=gen).fit(X_train[best_features], y_train)\ntree_fit.score(X_train[best_features], y_train), tree_fit.score(X_test[best_features], y_test)",
"_____no_output_____"
],
[
"tree_fit.tree_.max_depth",
"_____no_output_____"
]
],
[
[
"#### Hyperparameter Optimization",
"_____no_output_____"
]
],
[
[
"def tree_study(n=(3, 4, 5, 6, 8, 10, 15, 20, 25, 30, 35), seed=123):\n gen = np.random.RandomState(seed)\n train_score, test_score = [], []\n for max_depth in n:\n fit = tree.DecisionTreeRegressor(max_depth=max_depth, random_state=gen).fit(X_train[best_features], y_train)\n train_score.append(fit.score(X_train[best_features], y_train))\n test_score.append(fit.score(X_test[best_features], y_test))\n plt.plot(n, train_score, 'rx-', label='TRAIN')\n plt.plot(n, test_score, 'bo-', label='TEST')\n plt.xlabel('DecisionTree max_depth')\n plt.ylabel('DecisionTree $R^2$ score')\n plt.legend()",
"_____no_output_____"
],
[
"tree_study()",
"_____no_output_____"
]
],
[
[
"Chose a `max_depth` of 5 to minimize overfitting the training data (or choose 10 to balance overfitting with accuracy on the test data):",
"_____no_output_____"
]
],
[
[
"gen = np.random.RandomState(seed=123)\ntree_fit = tree.DecisionTreeRegressor(max_depth=5, random_state=gen).fit(X_train[best_features], y_train)\ntree_fit.score(X_train[best_features], y_train), tree_fit.score(X_test[best_features], y_test)",
"_____no_output_____"
]
],
[
[
"Note that a tree of depth $n$ sorts each sample into one of $2^n$ leaf nodes, each with a fixed prediction. This leads to a visible discretization error for small $n$, which is not necessarily a problem if the uncertainties are even larger:",
"_____no_output_____"
]
],
[
[
"y_predict = tree_fit.predict(X_test[best_features])\nplt.scatter(y_test, y_predict, lw=0)\nplt.xlabel('Target value')\nplt.ylabel('Predicted value');",
"_____no_output_____"
]
],
[
[
"#### Tree Visualization",
"_____no_output_____"
]
],
[
[
"tree.export_graphviz(tree_fit, out_file='tree.dot')",
"_____no_output_____"
],
[
"!dot -Tpng tree.dot -o tree.png",
"_____no_output_____"
],
[
"def plot_branch(path=[], fit=tree_fit, X=X_train[best_features], y=y_train.values):\n tree = fit.tree_\n n_nodes = tree.node_count\n children_left = tree.children_left\n children_right = tree.children_right\n feature = tree.feature\n threshold = tree.threshold\n # Traverse the tree using the specified path.\n node = 0\n sel = np.ones(len(X), bool)\n cut = threshold[node] \n x = X.iloc[:, feature[node]]\n print('nsel', np.count_nonzero(sel), 'cut', cut, 'value', np.mean(y[sel]))\n for below_threshold in path:\n if below_threshold:\n sel = sel & (x <= cut)\n node = children_left[node]\n else:\n sel = sel & (x > cut)\n node = children_right[node]\n cut = threshold[node] \n x = X.iloc[:, feature[node]]\n print('nsel', np.count_nonzero(sel), 'cut', cut, 'value', np.mean(y[sel]))\n fig, ax = plt.subplots(1, 2, figsize=(12, 6))\n xlim = np.percentile(x[sel], (1, 95))\n below = sel & (x <= cut)\n above = sel & (x > cut)\n ax[0].hist(x[below], range=xlim, bins=50, histtype='stepfilled', color='r', alpha=0.5)\n ax[0].hist(x[above], range=xlim, bins=50, histtype='stepfilled', color='b', alpha=0.5)\n ax[0].set_xlim(*xlim)\n ax[0].set_xlabel(X.columns[feature[node]])\n ylim = np.percentile(y, (1, 99))\n y_pred = np.empty_like(y)\n y_pred[below] = np.mean(y[below])\n y_pred[above] = np.mean(y[above]) \n mse2 = np.mean((y[sel] - y_pred[sel]) ** 2)\n n_below = np.count_nonzero(below)\n n_above = np.count_nonzero(above)\n mse = (np.var(y[below]) * n_below + np.var(y[above]) * n_above) / (n_below + n_above)\n #print('mse', mse, mse2)\n ax[1].hist(y[below], range=ylim, bins=25, histtype='stepfilled', color='r', alpha=0.5)\n ax[1].axvline(np.mean(y[below]), c='r', ls='--')\n ax[1].hist(y[above], range=ylim, bins=25, histtype='stepfilled', color='b', alpha=0.5)\n ax[1].axvline(np.mean(y[above]), c='b', ls='--')\n ax[1].set_xlabel('Redshift target')",
"_____no_output_____"
],
[
"plot_branch([])",
"nsel 13024 cut 0.007975295186042786 value 1.9733001\n"
],
[
"plot_branch([True,])",
"nsel 13024 cut 0.007975295186042786 value 1.9733001\nnsel 3044 cut 0.008564084768295288 value 3.4734519\n"
],
[
"plot_branch([False,])",
"nsel 13024 cut 0.007975295186042786 value 1.9733001\nnsel 9980 cut 0.28916649520397186 value 1.5157386\n"
]
],
[
[
"### Random Forest Regression",
"_____no_output_____"
],
[
"Use an ensemble of decision trees that are individually less accurate but collectively more accurate, with [RandomForestRegressor](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestRegressor.html). The individual trees are trained on random sub-samples of the data and the resulting \"forest\" of predictions are averaged. The random subsets for each tree are created by:\n - Using a \"bootstrap\" resampling of the rows, and\n - Finding the best split at each branch from a random subset of `max_features` features (columns).",
"_____no_output_____"
]
],
[
[
"gen = np.random.RandomState(seed=123)\nforest_fit = ensemble.RandomForestRegressor(\n n_estimators=15, max_features=0.5, random_state=gen, n_jobs=8).fit(X_train, y_train.values.reshape(-1))",
"_____no_output_____"
],
[
"forest_fit.score(X_train, y_train), forest_fit.score(X_test, y_test)",
"_____no_output_____"
]
],
[
[
"Compare the first branch for two of the trees in our forest:",
"_____no_output_____"
]
],
[
[
"plot_branch(fit=forest_fit.estimators_[0], X=X_train)",
"nsel 13024 cut 0.01966005004942417 value 1.9733001\n"
],
[
"plot_branch(fit=forest_fit.estimators_[1], X=X_train)",
"nsel 13024 cut 0.0073961850721389055 value 1.9733001\n"
]
],
[
[
"#### Hyperparameter Optimization",
"_____no_output_____"
]
],
[
[
"def forest_study(n=(1, 2, 3, 5, 10, 15, 20, 25, 30), seed=123):\n gen = np.random.RandomState(seed)\n train_score, test_score = [], []\n for n_estimators in n:\n fit = ensemble.RandomForestRegressor(\n n_estimators=n_estimators, max_features=0.5, random_state=gen, n_jobs=8).fit(\n X_train, y_train.values.reshape(-1))\n train_score.append(fit.score(X_train, y_train))\n test_score.append(fit.score(X_test, y_test))\n plt.plot(n, train_score, 'rx-', label='TRAIN')\n plt.plot(n, test_score, 'bo-', label='TEST')\n plt.xlabel('RandomForest n_estimators')\n plt.ylabel('RandomForest $R^2$ score')\n plt.legend()",
"_____no_output_____"
],
[
"forest_study()",
"_____no_output_____"
]
],
[
[
"#### Feature Importance (again)",
"_____no_output_____"
]
],
[
[
"importance = pd.DataFrame(\n {'feature': X.columns, 'importance': forest_fit.feature_importances_}\n).sort_values(by='importance', ascending=False)",
"_____no_output_____"
],
[
"importance.plot('feature', 'importance', 'barh', figsize=(10, 10), legend=False);",
"_____no_output_____"
]
],
[
[
"#### Prediction uncertainty",
"_____no_output_____"
],
[
"Since we now have multiple predictions for each sample, we can use their spread as an estimate of the uncertainty in the mean prediction:",
"_____no_output_____"
]
],
[
[
"y_pred = forest_fit.predict(X_test)",
"_____no_output_____"
],
[
"y_pred_each = np.array([tree.predict(X_test) for tree in forest_fit.estimators_])\ny_pred_each.shape",
"_____no_output_____"
],
[
"np.all(y_pred == np.mean(y_pred_each, axis=0))",
"_____no_output_____"
],
[
"y_pred_error = y_test.values.reshape(-1) - y_pred\ny_pred_spread = np.std(y_pred_each, axis=0)",
"_____no_output_____"
],
[
"plt.scatter(np.abs(y_pred_error), y_pred_spread, lw=0)\nplt.xlabel('$|y_{true} - y_{pred}|$')\nplt.ylabel('Forest prediction spread')\nplt.xlim(0, 3)\nplt.ylim(0, 3);",
"_____no_output_____"
],
[
"bins = np.linspace(-2.5, 2.5, 50)\nplt.hist(y_pred_error / y_pred_spread, bins=bins, density=True)\npull = 0.5 * (bins[1:] + bins[:-1])\nplt.plot(pull, scipy.stats.norm.pdf(pull), 'r-', label='$\\sigma=$ spread')\ncorrection = 2.0\nplt.plot(pull, correction * scipy.stats.norm.pdf(correction * pull), 'r--',\n label=('$\\sigma=%.1f \\\\times$' % correction) + 'spread')\nplt.legend()\nplt.xlabel('pull = dy / $\\sigma$')",
"_____no_output_____"
]
],
[
[
"#### \"Out-of-bag\" Testing",
"_____no_output_____"
],
[
"Combining the trees in a forest is known as \"bagging\". Since each tree leaves out some samples, we can use these omitted (aka \"out-of-bag\") samples to test our model. This means we no longer need to set aside a separate test dataset and can use all of our data for training the forest.\n\n*Technical note: since RandomForestRegressor does not support a max_samples parameter, the out-of-bag samples are only due to bootstrap sampling with replacement, which generally needs more estimators for reasonable statistics.*",
"_____no_output_____"
]
],
[
[
"gen = np.random.RandomState(seed=123)\nforest_fit = ensemble.RandomForestRegressor(\n n_estimators=100, max_features=0.5, oob_score=True, random_state=gen, n_jobs=8).fit(X, y.values.reshape(-1))",
"_____no_output_____"
],
[
"forest_fit.score(X_train, y_train), forest_fit.oob_score_",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
]
] |
4acbd7214fe69220cf5a1e8b654e3406cd37ac93
| 13,028 |
ipynb
|
Jupyter Notebook
|
jetson/.ipynb_checkpoints/simple_gotogoal_obsolete-checkpoint.ipynb
|
2788west/cerusbot
|
a667ffa951834150f9fc03f84a002b00e607a325
|
[
"Unlicense"
] | 8 |
2020-12-01T14:43:24.000Z
|
2022-03-10T04:58:27.000Z
|
jetson/.ipynb_checkpoints/simple_gotogoal_obsolete-checkpoint.ipynb
|
2788west/cerusbot
|
a667ffa951834150f9fc03f84a002b00e607a325
|
[
"Unlicense"
] | null | null | null |
jetson/.ipynb_checkpoints/simple_gotogoal_obsolete-checkpoint.ipynb
|
2788west/cerusbot
|
a667ffa951834150f9fc03f84a002b00e607a325
|
[
"Unlicense"
] | 6 |
2021-01-28T09:35:49.000Z
|
2022-03-03T04:50:00.000Z
| 33.751295 | 541 | 0.517117 |
[
[
[
"# Simple Go-To-Goal for Cerus",
"_____no_output_____"
],
[
"The following code implements a simple go-to-goal behavior for Cerus. It uses a closed feedback loop to continuously asses Cerus' state (position and heading) in the world using data from two wheel encoders. It subsequently calculates the error between a given goal location and its current pose and will attempt to minimize the error until it reaches the goal location. A P-regulator (see PID regulator) script uses the error as an input and outputs the angular velocity for the Arduino and motor controllers that drive the robot. \n\nAll models used in this program are adapted from Georgia Tech's \"Control of Mobile Robots\" by Dr. Magnus Egerstedt. ",
"_____no_output_____"
]
],
[
[
"#Import useful libraries\nimport serial\nimport time\nimport math\nimport numpy as np\nfrom traitlets import HasTraits, List",
"_____no_output_____"
],
[
"#Open a serial connection with the Arduino Mega\n#Opening a serial port on the Arduino resets it, so our encoder count is also reset to 0,0\nser = serial.Serial('COM3', 115200)",
"_____no_output_____"
],
[
"#Defining our goal location. Units are metric, real-world coordinates in an X/Y coordinate system\ngoal_x = 1\ngoal_y = 0",
"_____no_output_____"
],
[
"#Create a class for our Cerus robot\nclass Cerus():\n def __init__(self, pose_x, pose_y, pose_phi, R_wheel, N_ticks, L_track):\n self.pose_x = pose_x #X Position\n self.pose_y = pose_y #Y Position\n self.pose_phi = pose_phi #Heading\n self.R_wheel = R_wheel #wheel radius in meters\n self.N_ticks = N_ticks #encoder ticks per wheel revolution\n self.L_track = L_track #wheel track in meters\n\n#Create a Cerus instance and initialize it to a 0,0,0 world position and with some physical dimensions \ncerus = Cerus(0,0,0,0.03,900,0.23)",
"_____no_output_____"
]
],
[
[
"We'll use the Traitlets library to implement an observer pattern that will recalculate the pose of the robot every time an update to the encoder values is detected and sent to the Jetson nano by the Arduino.",
"_____no_output_____"
]
],
[
[
"#Create an encoder class with traits\nclass Encoders(HasTraits): \n encoderValues = List() #We store the left and right encoder value in a list\n \n def __init__(self, encoderValues, deltaTicks):\n self.encoderValues = encoderValues \n self.deltaTicks = deltaTicks \n\n#Create an encoder instance \nencoders = Encoders([0,0], [0,0])",
"_____no_output_____"
],
[
"#Create a function that is triggered when a change to encoders is detected\ndef monitorEncoders(change):\n if change['new']:\n \n oldVals = np.array(change['old'])\n newVals = np.array(change['new'])\n deltaTicks = newVals - oldVals\n #print(\"Old values: \", oldVals)\n #print(\"New values: \", newVals)\n #print(\"Delta values: \", deltaTicks)\n \n calculatePose(deltaTicks)\n \nencoders.observe(monitorEncoders, names = \"encoderValues\")",
"_____no_output_____"
]
],
[
[
"The functions below are helpers and will be called through our main loop.",
"_____no_output_____"
]
],
[
[
"#Create a move function that sends move commands to the Arduino\ndef move(linearVelocity, angularVelocity): \n \n command = f\"<{linearVelocity},{angularVelocity}>\"\n ser.write(str.encode(command))",
"_____no_output_____"
],
[
"#Create a function that calculates an updated pose of Cerus every time it is called\ndef calculatePose(deltaTicks):\n \n #Calculate the centerline distance moved\n distanceLeft = 2 * math.pi * cerus.R_wheel * (deltaTicks[0] / cerus.N_ticks)\n distanceRight = 2 * math.pi * cerus.R_wheel * (deltaTicks[1] / cerus.N_ticks)\n distanceCenter = (distanceLeft + distanceRight) / 2\n \n #Update the position and heading\n cerus.pose_x = round((cerus.pose_x + distanceCenter * math.cos(cerus.pose_phi)), 4)\n cerus.pose_y = round((cerus.pose_y + distanceCenter * math.sin(cerus.pose_phi)), 4)\n cerus.pose_phi = round((cerus.pose_phi + ((distanceRight - distanceLeft) / cerus.L_track)), 4)\n \n print(f\"The new position is {cerus.pose_x}, {cerus.pose_y} and the new heading is {cerus.pose_phi}.\")",
"_____no_output_____"
],
[
"#Calculate the error between Cerus' heading and the goal point\ndef calculateError():\n \n phi_desired = math.atan((goal_y - cerus.pose_y)/(goal_x - cerus.pose_x))\n \n temp = phi_desired - cerus.pose_phi\n error_heading = round((math.atan2(math.sin(temp), math.cos(temp))), 4) #ensure that error is within [-pi, pi]\n error_x = round((goal_x - cerus.pose_x), 4)\n error_y = round((goal_y - cerus.pose_y), 4)\n \n #print(\"The heading error is: \", error_heading)\n #print(\"The X error is: \", error_x)\n #print(\"The Y error is: \", error_y)\n return error_x, error_y, error_heading",
"_____no_output_____"
],
[
"atGoal = False\nconstVel = 0.2\nK = 1 #constant for our P-regulator below",
"_____no_output_____"
],
[
"#Functions to read and format encoder data received from the Serial port\ndef formatData(data):\n delimiter = \"x\"\n leftVal = \"\"\n rightVal = \"\"\n \n for i in range(len(data)): \n if data[i] == \",\":\n delimiter = \",\" \n elif delimiter != \",\" and data[i].isdigit():\n leftVal += data[i] \n elif delimiter == \",\" and data[i].isdigit():\n rightVal += data[i] \n \n leftVal, rightVal = int(leftVal), int(rightVal)\n encoders.encoderValues = [leftVal, rightVal] \n print(\"Encoders: \", encoders.encoderValues)\n \ndef handleSerial():\n #ser.readline() waits for the next line of encoder data, which is sent by Arduino every 50 ms\n if ser.inWaiting():\n \n #Get the serial data and format it\n temp = ser.readline()\n data = temp.decode()\n formatData(data)\n \n #Calculate the current pose to goal error\n error_x, error_y, error_heading = calculateError()\n print(f\"Error X: {error_x}, Error Y: {error_y}\")\n \n #If we're within 5 cm of the goal\n if error_x <= 0.05:# and error_y <= 0.05:\n print(\"Goal reached!\")\n move(0.0,0.0)\n time.sleep(0.1)\n atGoal = True\n \n #Otherwise keep driving\n else:\n omega = - (K * error_heading) \n handleSerial() \n move(constVel, 0.0)\n print(\"Moving at angular speed: \", omega)",
"_____no_output_____"
],
[
"def moveRobot():\n \n #The Arduino sends data every 50ms, we first check if data is in the buffer\n if ser.inWaiting():\n \n #Get the serial data and format it if data is in the buffer\n temp = ser.readline()\n data = temp.decode()\n formatData(data)\n \n #Calculate the current pose to goal error\n error_x, error_y, error_heading = calculateError()\n print(f\"Error X: {error_x}, Error Y: {error_y}\")\n \n #If we're within 5 cm of the goal\n if error_x <= 0.05:# and error_y <= 0.05:\n print(\"Goal reached!\")\n move(0.0,0.0)\n time.sleep(0.1)\n atGoal = True\n \n #Otherwise keep driving\n else:\n omega = - (K * error_heading) \n handleSerial() \n move(constVel, 0.0)\n print(\"Moving at angular speed: \", omega) ",
"_____no_output_____"
]
],
[
[
"This is the main part for our program that will loop over and over until Cerus has reached its goal. For our simple go-to-goal behavior, we will drive the robot at a constant speed and only adjust our heading so that we reach the goal location.\n\n__WARNING: This will move the robot!__",
"_____no_output_____"
]
],
[
[
"while not atGoal:\n try:\n moveRobot() \n \n except(KeyboardInterrupt):\n print(\"Program interrupted by user!\")\n move(0.0,0.0) #Stop motors\n break\n\n\"Loop exited...\"\nmove(0.0,0.0) #Stop motors",
"_____no_output_____"
],
[
"#Close the serial connection when done\nser.close()",
"_____no_output_____"
],
[
"atGoal = False\nconstVel = 0.2\nK = 1 #constant for our P-regulator below\n\nwhile not atGoal:\n try: \n \n #Calculate the current pose to goal error\n error_x, error_y, error_heading = calculateError()\n print(f\"Error X: {error_x}, Error Y: {error_y}\")\n \n #If we're within 5 cm of the goal\n if error_x <= 0.05 and error_y <= 0.05:\n print(\"Goal reached!\")\n move(0.0,0.0)\n time.sleep(0.1)\n atGoal = True\n\n #Otherwise keep driving\n else:\n omega = - (K * error_heading) \n handleSerial() \n move(constVel, 0.0)\n print(\"Moving at angular speed: \", omega)\n \n \n except(KeyboardInterrupt):\n print(\"Program interrupted by user!\")\n move(0.0,0.0) #Stop motors\n break\n\n\"Loop exited...\"\nmove(0.0,0.0) #Stop motors",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
]
] |
4acbd7e3547c28195ba3cd5216880c9ef077f709
| 29,907 |
ipynb
|
Jupyter Notebook
|
seq2seq/chatbot_tutorial.ipynb
|
aidiary/deep-learning-notebooks
|
95783c6ba2688e15d362a1fcfec6215861d65a9a
|
[
"MIT"
] | 1 |
2019-09-22T14:09:48.000Z
|
2019-09-22T14:09:48.000Z
|
seq2seq/chatbot_tutorial.ipynb
|
aidiary/deep-learning-notebooks
|
95783c6ba2688e15d362a1fcfec6215861d65a9a
|
[
"MIT"
] | null | null | null |
seq2seq/chatbot_tutorial.ipynb
|
aidiary/deep-learning-notebooks
|
95783c6ba2688e15d362a1fcfec6215861d65a9a
|
[
"MIT"
] | null | null | null | 36.471951 | 226 | 0.516902 |
[
[
[
"# Chatbot Tutorial\n- https://pytorch.org/tutorials/beginner/chatbot_tutorial.html",
"_____no_output_____"
]
],
[
[
"import torch\nfrom torch.jit import script, trace\nimport torch.nn as nn\nfrom torch import optim\nimport torch.nn.functional as F\nimport csv\nimport random\nimport re\nimport os\nimport unicodedata\nimport codecs\nfrom io import open\nimport itertools\nimport math\n\nUSE_CUDA = torch.cuda.is_available()\ndevice = torch.device('cuda' if USE_CUDA else 'cpu')",
"_____no_output_____"
]
],
[
[
"## データの前処理",
"_____no_output_____"
]
],
[
[
"corpus_name = 'cornell_movie_dialogs_corpus'\ncorpus = os.path.join('data', corpus_name)\n\ndef printLines(file, n=10):\n with open(file, 'rb') as datafile:\n lines = datafile.readlines()\n for line in lines[:n]:\n print(line)\n\nprintLines(os.path.join(corpus, 'movie_lines.txt'))",
"b'L1045 +++$+++ u0 +++$+++ m0 +++$+++ BIANCA +++$+++ They do not!\\n'\nb'L1044 +++$+++ u2 +++$+++ m0 +++$+++ CAMERON +++$+++ They do to!\\n'\nb'L985 +++$+++ u0 +++$+++ m0 +++$+++ BIANCA +++$+++ I hope so.\\n'\nb'L984 +++$+++ u2 +++$+++ m0 +++$+++ CAMERON +++$+++ She okay?\\n'\nb\"L925 +++$+++ u0 +++$+++ m0 +++$+++ BIANCA +++$+++ Let's go.\\n\"\nb'L924 +++$+++ u2 +++$+++ m0 +++$+++ CAMERON +++$+++ Wow\\n'\nb\"L872 +++$+++ u0 +++$+++ m0 +++$+++ BIANCA +++$+++ Okay -- you're gonna need to learn how to lie.\\n\"\nb'L871 +++$+++ u2 +++$+++ m0 +++$+++ CAMERON +++$+++ No\\n'\nb'L870 +++$+++ u0 +++$+++ m0 +++$+++ BIANCA +++$+++ I\\'m kidding. You know how sometimes you just become this \"persona\"? And you don\\'t know how to quit?\\n'\nb'L869 +++$+++ u0 +++$+++ m0 +++$+++ BIANCA +++$+++ Like my fear of wearing pastels?\\n'\n"
],
[
"# Splits each line of the file into a dictionary of fields\ndef loadLines(fileName, fields):\n lines = {}\n with open(fileName, 'r', encoding='iso-8859-1') as f:\n for line in f:\n values = line.split(' +++$+++ ')\n # Extract fields\n lineObj = {}\n for i, field in enumerate(fields):\n lineObj[field] = values[i]\n lines[lineObj['lineID']] = lineObj\n return lines",
"_____no_output_____"
],
[
"MOVIE_LINES_FIELDS = ['lineID', 'characterID', 'movieID', 'character', 'text']\nlines = loadLines(os.path.join(corpus, 'movie_lines.txt'), MOVIE_LINES_FIELDS)",
"_____no_output_____"
],
[
"lines['L1045']",
"_____no_output_____"
],
[
"# Groups fields of lines from loadLines() into conversations based on movie_conversations.txt\ndef loadConversations(fileName, lines, fields):\n conversations = []\n with open(fileName, 'r', encoding='iso-8859-1') as f:\n for line in f:\n values = line.split(' +++$+++ ')\n # Extract fields\n convObj = {}\n for i, field in enumerate(fields):\n convObj[field] = values[i]\n # Convert string to list\n utterance_id_pattern = re.compile('L[0-9]+')\n lineIds = utterance_id_pattern.findall(convObj['utteranceIDs'])\n # Reassemble lines\n convObj['lines'] = []\n for lineId in lineIds:\n convObj['lines'].append(lines[lineId])\n conversations.append(convObj)\n return conversations",
"_____no_output_____"
],
[
"MOVIE_CONVERSATIONS_FIELDS = ['character1ID', 'character2ID', 'movieID', 'utteranceIDs']\nconversations = loadConversations(os.path.join(corpus, 'movie_conversations.txt'), lines, MOVIE_CONVERSATIONS_FIELDS)",
"_____no_output_____"
],
[
"# utteranceIDsの会話系列IDがlinesに展開されていることがわかる!\nconversations[0]",
"_____no_output_____"
],
[
"# Extracts pairs of sentences from conversations\ndef extractSentencePairs(conversations):\n qa_pairs = []\n for conversation in conversations:\n # Iterate over all the lines of the conversation\n for i in range(len(conversation['lines']) - 1): # 最後の会話は回答がないので無視する\n inputLine = conversation['lines'][i]['text'].strip()\n targetLine = conversation['lines'][i + 1]['text'].strip()\n if inputLine and targetLine:\n qa_pairs.append([inputLine, targetLine])\n return qa_pairs",
"_____no_output_____"
],
[
"datafile = os.path.join(corpus, 'formatted_movie_lines.txt')\n\ndelimiter = '\\t'\ndelimiter = str(codecs.decode(delimiter, 'unicode_escape'))\n\nwith open(datafile, 'w', encoding='utf-8') as outputfile:\n writer = csv.writer(outputfile, delimiter=delimiter, lineterminator='\\n')\n for pair in extractSentencePairs(conversations):\n writer.writerow(pair)",
"_____no_output_____"
],
[
"printLines(datafile)",
"b\"Can we make this quick? Roxanne Korrine and Andrew Barrett are having an incredibly horrendous public break- up on the quad. Again.\\tWell, I thought we'd start with pronunciation, if that's okay with you.\\n\"\nb\"Well, I thought we'd start with pronunciation, if that's okay with you.\\tNot the hacking and gagging and spitting part. Please.\\n\"\nb\"Not the hacking and gagging and spitting part. Please.\\tOkay... then how 'bout we try out some French cuisine. Saturday? Night?\\n\"\nb\"You're asking me out. That's so cute. What's your name again?\\tForget it.\\n\"\nb\"No, no, it's my fault -- we didn't have a proper introduction ---\\tCameron.\\n\"\nb\"Cameron.\\tThe thing is, Cameron -- I'm at the mercy of a particularly hideous breed of loser. My sister. I can't date until she does.\\n\"\nb\"The thing is, Cameron -- I'm at the mercy of a particularly hideous breed of loser. My sister. I can't date until she does.\\tSeems like she could get a date easy enough...\\n\"\nb'Why?\\tUnsolved mystery. She used to be really popular when she started high school, then it was just like she got sick of it or something.\\n'\nb\"Unsolved mystery. She used to be really popular when she started high school, then it was just like she got sick of it or something.\\tThat's a shame.\\n\"\nb'Gosh, if only we could find Kat a boyfriend...\\tLet me see what I can do.\\n'\n"
]
],
[
[
"## Vocabularyの構築",
"_____no_output_____"
]
],
[
[
"# Default word tokens\nPAD_token = 0 # Used for padding short sentences\nSOS_token = 1 # Start-of-sentence token\nEOS_token = 2 # End-of-sentence token\n\nclass Voc:\n \n def __init__(self, name):\n self.name = name\n self.trimmed = False\n self.word2index = {}\n self.word2count = {}\n self.index2word = {PAD_token: 'PAD', SOS_token: 'SOS', EOS_token: 'EOS'}\n self.num_words = 3 # SOS, EOS, PAD\n \n def addSentence(self, sentence):\n for word in sentence.split(' '):\n self.addWord(word)\n \n def addWord(self, word):\n if word not in self.word2index:\n self.word2index[word] = self.num_words\n self.word2count[word] = 1\n self.index2word[self.num_words] = word\n self.num_words += 1\n else:\n self.word2count[word] += 1\n \n # Remove words below a certain count threshold\n def trim(self, min_count):\n if self.trimmed:\n return\n self.trimmed = True\n \n keep_words = []\n \n for k, v in self.word2count.items():\n if v >= min_count:\n keep_words.append(k)\n \n print('keep_words {} / {} = {:.4f}'.format(len(keep_words), len(self.word2index), len(keep_words) / len(self.word2index)))\n \n # Reinitialize dictionaries\n self.word2index = {}\n self.word2count = {}\n self.index2word = {PAD_token: 'PAD', SOS_token: 'SOS', EOS_token: 'EOS'}\n self.num_words = 3\n \n for word in keep_words:\n self.addWord(word)",
"_____no_output_____"
],
[
"MAX_LENGTH = 10 # Maximum sentence length to consider\n\n# Turn a Unicode string to plain ASCII\ndef unicodeToAscii(s):\n return ''.join(c for c in unicodedata.normalize('NFD', s) if unicodedata.category(c) != 'Mn')\n\n# Lowercase, trim, and remove non-letter characters\ndef normalizeString(s):\n s = unicodeToAscii(s.lower().strip())\n s = re.sub(r\"([.!?])\", r\" \\1\", s)\n s = re.sub(r\"[^a-zA-Z.!?]+\", r\" \", s)\n s = re.sub(r\"\\s+\", r\" \", s).strip()\n return s\n\n# Read query/response paiers and return a voc object\ndef readVocs(datafile, corpus_name):\n # Read the file and split into lines\n lines = open(datafile, encoding='utf-8').read().strip().split('\\n')\n # Split every line into pairs and normalize\n pairs = [[normalizeString(s) for s in l.split('\\t')] for l in lines]\n voc = Voc(corpus_name)\n return voc, pairs\n\n# Returns True iff both sentences in a pair p are under the MAX_LENGTH threshold\ndef filterPair(p):\n # Input sequences need to preserve the last word for EOS token\n return len(p[0].split(' ')) < MAX_LENGTH and len(p[1].split(' ')) < MAX_LENGTH\n\n# Filter pairs using filterPair condition\ndef filterPairs(pairs):\n return [pair for pair in pairs if filterPair(pair)]\n\n# Using the functions defined above, return a populated voc object and pairs list\n# save_dirが使われていない\ndef loadPrepareData(corpus, corpus_name, datafile, save_dir):\n print('Start preparing training data ...')\n voc, pairs = readVocs(datafile, corpus_name)\n print('Read {!s} sentence pairs'.format(len(pairs)))\n pairs = filterPairs(pairs)\n print('Trimmed to {!s} sentence pairs'.format(len(pairs)))\n print('Counting words...')\n for pair in pairs:\n voc.addSentence(pair[0])\n voc.addSentence(pair[1])\n print('Counted words:', voc.num_words)\n return voc, pairs\n\n# Load/Assemble voc and pairs\nsave_dir = os.path.join('data', 'save')\nvoc, pairs = loadPrepareData(corpus, corpus_name, datafile, save_dir)\n# Print some pairs to validate\nprint('\\npairs:')\nfor pair in pairs[:10]:\n print(pair)",
"Start preparing training data ...\nRead 221282 sentence pairs\nTrimmed to 64271 sentence pairs\nCounting words...\nCounted words: 18008\n\npairs:\n['there .', 'where ?']\n['you have my word . as a gentleman', 'you re sweet .']\n['hi .', 'looks like things worked out tonight huh ?']\n['you know chastity ?', 'i believe we share an art instructor']\n['have fun tonight ?', 'tons']\n['well no . . .', 'then that s all you had to say .']\n['then that s all you had to say .', 'but']\n['but', 'you always been this selfish ?']\n['do you listen to this crap ?', 'what crap ?']\n['what good stuff ?', 'the real you .']\n"
],
[
"MIN_COUNT = 3 # Minimum word count threshold for trimming\n\n\ndef trimRareWords(voc, pairs, MIN_COUNT):\n # Trim words used unser the MIN_COUNT from the voc\n voc.trim(MIN_COUNT)\n # Filter out pairs with trimmed words\n keep_pairs = []\n for pair in pairs:\n input_sentence = pair[0]\n output_sentence = pair[1]\n keep_input = True\n keep_output = True\n # Check input sentence\n for word in input_sentence.split(' '):\n if word not in voc.word2index:\n keep_input = False\n break\n # Check output sentence\n for word in output_sentence.split(' '):\n if word not in voc.word2index:\n keep_output = False\n break\n \n # Only keep pairs that do not contain trimmed word(s) in their input or output sentence\n if keep_input and keep_output:\n keep_pairs.append(pair)\n \n print('Trimmed from {} pairs to {}, {:.4f} of total'.format(len(pairs), len(keep_pairs), len(keep_pairs) / len(pairs)))\n return keep_pairs\n\n\n# Trim voc and pairs\npairs = trimRareWords(voc, pairs, MIN_COUNT)",
"keep_words 7823 / 18005 = 0.4345\nTrimmed from 64271 pairs to 53165, 0.8272 of total\n"
]
],
[
[
"## Minibatchの構成",
"_____no_output_____"
]
],
[
[
"def indexesFromSentence(voc, sentence):\n return [voc.word2index[word] for word in sentence.split(' ')] + [EOS_token]\n\n\ndef zeroPadding(l, fillvalue=PAD_token):\n # ここで (batch_size, max_length) => (max_length, batch_size) に転置している\n return list(itertools.zip_longest(*l, fillvalue=fillvalue))\n\n\ndef binaryMatrix(l, value=PAD_token):\n m = []\n for i, seq in enumerate(l):\n m.append([])\n for token in seq:\n if token == PAD_token:\n m[i].append(0)\n else:\n m[i].append(1)\n return m\n\n\n# Returns padded input sentence tensor and lengths\ndef inputVar(l, voc):\n indexes_batch = [indexesFromSentence(voc, sentence) for sentence in l]\n lengths = torch.tensor([len(indexes) for indexes in indexes_batch])\n padList = zeroPadding(indexes_batch)\n padVar = torch.LongTensor(padList)\n return padVar, lengths\n\n\n# Returns padded target sequence tneosr, padding mask, and max target lengths\n# maskは出力と同じサイズのテンソルでPADが入ってるところが0でそれ以外は1\ndef outputVar(l, voc):\n indexes_batch = [indexesFromSentence(voc, sentence) for sentence in l]\n max_target_len = max([len(indexes) for indexes in indexes_batch])\n padList = zeroPadding(indexes_batch)\n mask = binaryMatrix(padList)\n mask = torch.ByteTensor(mask)\n padVar = torch.LongTensor(padList)\n return padVar, mask, max_target_len\n\n\n# Returns all items for a given batch of pairs\ndef batch2TrainData(voc, pair_batch):\n # 入力文章の長い順にソートする\n pair_batch.sort(key=lambda x: len(x[0].split(' ')), reverse=True)\n input_batch, output_batch = [], []\n for pair in pair_batch:\n input_batch.append(pair[0])\n output_batch.append(pair[1])\n inp, lengths = inputVar(input_batch, voc)\n output, mask, max_target_len = outputVar(output_batch, voc)\n return inp, lengths, output, mask, max_target_len\n\n\n# Example for validation\nsmall_batch_size = 5\nbatches = batch2TrainData(voc, [random.choice(pairs) for _ in range(small_batch_size)])\ninput_variable, lengths, target_variable, mask, max_target_len = batches\n\nprint('input_variable:', input_variable)\nprint('lengths:', lengths)\nprint('target_variable:', target_variable)\nprint('mask:', mask)\nprint('max_target_len:', max_target_len)",
"input_variable: tensor([[ 25, 53, 197, 2434, 1574],\n [ 25, 301, 117, 2317, 37],\n [ 102, 140, 47, 75, 44],\n [ 112, 4449, 36, 7802, 159],\n [ 123, 403, 66, 6, 4],\n [ 40, 4, 66, 2, 2],\n [ 510, 2, 2, 0, 0],\n [ 4, 0, 0, 0, 0],\n [ 2, 0, 0, 0, 0]])\nlengths: tensor([9, 7, 7, 6, 6])\ntarget_variable: tensor([[ 389, 266, 483, 25, 5],\n [ 25, 66, 4, 105, 92],\n [1528, 2, 2, 45, 27],\n [ 21, 0, 0, 4, 95],\n [ 169, 0, 0, 2, 6],\n [ 7, 0, 0, 0, 2],\n [ 6, 0, 0, 0, 0],\n [ 2, 0, 0, 0, 0]])\nmask: tensor([[1, 1, 1, 1, 1],\n [1, 1, 1, 1, 1],\n [1, 1, 1, 1, 1],\n [1, 0, 0, 1, 1],\n [1, 0, 0, 1, 1],\n [1, 0, 0, 0, 1],\n [1, 0, 0, 0, 0],\n [1, 0, 0, 0, 0]], dtype=torch.uint8)\nmax_target_len: 8\n"
]
],
[
[
"## Seq2Seq Model",
"_____no_output_____"
]
],
[
[
"class EncoderRNN(nn.Module):\n \n def __init__(self, hidden_size, embedding, n_layers=1, dropout=0):\n super(EncoderRNN, self).__init__()\n self.n_layers = n_layers\n self.hidden_size = hidden_size\n self.embedding = embedding\n \n self.gru = nn.GRU(hidden_size, hidden_size, n_layers, dropout=(0 if n_layers == 1 else dropout), bidirectional=True)\n \n def forward(self, input_seq, input_lengths, hidden=None):\n # Convert word indexes to embedding\n embedded = self.embedding(input_seq)\n \n # Pack padded batch of sequences for RNN module\n packed = nn.utils.rnn.pack_padded_sequence(embedded, input_lengths)\n \n # Forward pass through GRU\n # output of shape (seq_len, batch, num_directions * hidden_size)\n # h_n of shape (num_layers * num_directions, batch, hidden_size)\n outputs, hidden = self.gru(packed, hidden)\n \n # Unpack padding\n outputs, _ = nn.utils.rnn.pad_packed_sequence(outputs)\n \n # Sum bidirectional GRU outputs\n # bidirectionalの場合、outputsのhidden_sizeは2倍の長さで出てくる\n outputs = outputs[:, :, :self.hidden_size] + outputs[:, :, self.hidden_size:]\n \n # Return output and final hidden state\n return outputs, hidden",
"_____no_output_____"
]
],
[
[
"## Attention",
"_____no_output_____"
]
],
[
[
"class Attn(nn.Module):\n \n def __init__(self, method, hidden_size):\n super(Attn, self).__init__()\n self.method = method\n if self.method not in ['dot', 'general', 'concat']:\n raise ValueError(self.method, 'is not an appropriate attention method.')\n self.hidden_size = hidden_size\n if self.method == 'general':\n self.attn = nn.Linear(hidden_size, hidden_size)\n elif self.method == 'concat':\n self.attn = nn.Linear(hidden_size * 2, hidden_size)\n self.v = nn.Parameter(torch.FloatTensor(hidden_size))\n \n def dot_score(self, hidden, encoder_output):\n return torch.sum(hidden * encoder_output, dim=2)\n \n def general_score(self, hidden, encoder_output):\n energy = self.attn(encoder_output)\n return torch.sum(hidden * energy, dim=2)\n\n def concat_score(self, hidden, encoder_output):\n energy = self.attn(torch.cat((hidden.expand(encoder_output.size(0), -1, -1), encoder_output), 2)).tanh()\n return torch.sum(self.v * energy, dim=2)\n \n def forward(self, hidden, encoder_outputs):\n # Calculate the attention weights (energies) based on the given method\n if self.method == 'general':\n attn_energies = self.general_score(hidden, encoder_outputs)\n elif self.method == 'concat':\n attn_energies = self.concat_score(hidden, encoder_outputs)\n elif self.method == 'dot':\n attn_energies = self.dot_score(hidden, encoder_outputs)\n \n # Transpose max_length and batch_size dimensions\n attn_energies = attn_energies.t()\n \n # Return the softmax normalized probability scores (with added dimension)\n return F.softmax(attn_energies, dim=1).unsqueeze(1)",
"_____no_output_____"
]
],
[
[
"## Decoder",
"_____no_output_____"
]
],
[
[
"class LuongAttnDecoderRNN(nn.Module):\n \n def __init__(self, attn_model, embedding, hidden_size, output_size, n_layers=1, dropout=0.1):\n super(LuongAttnDecoderRNN, self).__init__()\n \n self.attn_model = attn_model\n self.hidden_size = hidden_size\n self.output_size = output_size\n self.n_layers = n_layers\n self.dropout = dropout\n\n # Define layers\n self.embedding = embedding\n self.embedding_dropout = nn.Dropout(dropout)\n self.gru = nn.GRU(hidden_size, hidden_size, n_layers, dropout=(0 if n_layers == 1 else dropout))\n self.concat = nn.Linear(hidden_size * 2, hidden_size)\n self.out = nn.Linear(hidden_size, output_size)\n self.attn = Attn(attn_model, hidden_size)\n \n def forward(self, input_step, last_hidden, encoder_outputs):\n # Decoderは各タイムステップごとに実行される\n embedded = self.embedding(input_step)\n embedded = self.embedding_dropout(embedded)\n \n # Forward through unidirectional GRU\n rnn_output, hidden = self.gru(embedded, last_hidden)\n \n # Calculate attention weights from the current GRU output\n attn_weights = self.attn(rnn_output, encoder_outputs)\n \n # Multiply attention weights to encoder outputs to get new weighted sum context vector\n context = attn_weights.bmm(encoder_outputs.transpose(0, 1))\n \n # Concatenate weighted context vector and GRU output using Luong eq. 5\n rnn_output = rnn_output.squeeze(0)\n context = context.squeeze(1)\n concat_input = torch.cat((rnn_output, context), 1)\n concat_output = torch.tanh(self.concat(concat_input))\n \n # Predict next word using Luong eq.6\n output = self.out(concat_output)\n output = F.softmax(output, dim=1)\n \n # Return output and final hidden state\n return output, hidden",
"_____no_output_____"
]
],
[
[
"## Masked loss",
"_____no_output_____"
]
],
[
[
"def maskNLLLoss(inp, target, mask):\n nTotal = mask.sum()\n crossEntropy = -torch.log(torch.gather(inp, 1, target.view(-1, 1)).squeeze(1))\n loss = crossEntropy.masked_select(mask).mean()\n loss = loss.to(device)\n return loss, nTotal.item()",
"_____no_output_____"
]
],
[
[
"## Training",
"_____no_output_____"
]
],
[
[
"def train(input_variable, lengths, target_variable, mask, max_target_len, encoder, decoder, embedding,\n encoder_optimizer, decoder_optimizer, batch_size, clip, max_length=MAX_LENGTH):\n pass",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4acc00f633fbf119f30a07f4465a176fe2990888
| 21,774 |
ipynb
|
Jupyter Notebook
|
ForwardAutodiff.ipynb
|
Farr-PHY-604/ForwardAutodiff.jl
|
7f49ad4f7a5f885c81874d92964d4ce82f64156a
|
[
"MIT"
] | 1 |
2020-10-27T00:38:34.000Z
|
2020-10-27T00:38:34.000Z
|
ForwardAutodiff.ipynb
|
Farr-PHY-604/ForwardAutodiff.jl
|
7f49ad4f7a5f885c81874d92964d4ce82f64156a
|
[
"MIT"
] | 1 |
2019-09-13T13:22:21.000Z
|
2019-09-13T14:32:49.000Z
|
ForwardAutodiff.ipynb
|
Farr-PHY-604/ForwardAutodiff.jl
|
7f49ad4f7a5f885c81874d92964d4ce82f64156a
|
[
"MIT"
] | null | null | null | 27.251564 | 538 | 0.537935 |
[
[
[
"empty"
]
]
] |
[
"empty"
] |
[
[
"empty"
]
] |
4acc142308abcb1bf04ba531b6d95323a533efa4
| 1,652 |
ipynb
|
Jupyter Notebook
|
M05_submissions/Anaimi-rough.ipynb
|
UConn-Cooper/engineering-dynamics
|
d89a591321634905c1c0e3522a3a9f7aab5abbd3
|
[
"BSD-3-Clause"
] | 1 |
2021-02-16T23:51:30.000Z
|
2021-02-16T23:51:30.000Z
|
M05_submissions/Anaimi-rough.ipynb
|
UConn-Cooper/engineering-dynamics
|
d89a591321634905c1c0e3522a3a9f7aab5abbd3
|
[
"BSD-3-Clause"
] | 2 |
2021-02-16T01:25:06.000Z
|
2021-06-24T20:29:00.000Z
|
M05_submissions/Anaimi-rough.ipynb
|
UConn-Cooper/engineering-dynamics
|
d89a591321634905c1c0e3522a3a9f7aab5abbd3
|
[
"BSD-3-Clause"
] | 3 |
2021-02-16T01:20:32.000Z
|
2021-09-02T19:19:58.000Z
| 24.656716 | 207 | 0.535109 |
[
[
[
"from IPython.display import IFrame\nIFrame(\"https://github.com/cooperrc/engineering-dynamics/blob/cb003d9645570ea313db598ddd5c75f6b95891ed/M05_submissions/Alnaimi-rough-draft-of-Module-5-project.pdf?embed=True\", width=600, height=1500)",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code"
]
] |
4acc1e0b67b7cddf608505e9fcf39251592fd2b4
| 82,566 |
ipynb
|
Jupyter Notebook
|
Movie_recommendation_system.ipynb
|
Siddish/Movie_recommendation_system
|
9d0b1725a668a8fd99175332ca1f7db2f383ddd3
|
[
"BSD-4-Clause-UC"
] | null | null | null |
Movie_recommendation_system.ipynb
|
Siddish/Movie_recommendation_system
|
9d0b1725a668a8fd99175332ca1f7db2f383ddd3
|
[
"BSD-4-Clause-UC"
] | null | null | null |
Movie_recommendation_system.ipynb
|
Siddish/Movie_recommendation_system
|
9d0b1725a668a8fd99175332ca1f7db2f383ddd3
|
[
"BSD-4-Clause-UC"
] | null | null | null | 68.919866 | 24,584 | 0.683756 |
[
[
[
"#### This project is a code along for the article I read here: https://www.analyticsvidhya.com/blog/2020/11/create-your-own-movie-movie-recommendation-system/",
"_____no_output_____"
]
],
[
[
"# Importing required libraries and packages\nimport pandas as pd\nimport numpy as np\nfrom scipy.sparse import csr_matrix\nfrom sklearn.neighbors import NearestNeighbors\nimport matplotlib.pyplot as plt\nimport seaborn as sns",
"_____no_output_____"
],
[
"# Importing the datasets\nmovies = pd.read_csv('Movie_recommendation_datasets/movies.csv')\nratings = pd.read_csv(\"Movie_recommendation_datasets/ratings.csv\")",
"_____no_output_____"
],
[
"movies.head()\nratings.head()",
"_____no_output_____"
],
[
"final_dataset = ratings.pivot(index='movieId',columns='userId',values='rating')\nfinal_dataset.head()",
"_____no_output_____"
],
[
"final_dataset.fillna(0,inplace=True)\nfinal_dataset.head()",
"_____no_output_____"
]
],
[
[
"#### Visualizing the filters",
"_____no_output_____"
]
],
[
[
"no_user_voted = ratings.groupby('movieId')['rating'].agg('count')\nno_movies_voted = ratings.groupby('userId')['rating'].agg('count')",
"_____no_output_____"
],
[
"f,ax = plt.subplots(1,1,figsize=(16,4))\n# ratings['rating'].plot(kind='hist')\nplt.scatter(no_user_voted.index,no_user_voted,color='mediumseagreen')\nplt.axhline(y=10,color='r')\nplt.xlabel('MovieId')\nplt.ylabel('No. of users voted')\nplt.show()",
"_____no_output_____"
]
],
[
[
"#### Making the necessary modifications as per the threshold set",
"_____no_output_____"
]
],
[
[
"final_dataset = final_dataset.loc[no_user_voted[no_user_voted > 10].index,:]",
"_____no_output_____"
],
[
"f,ax = plt.subplots(1,1,figsize=(16,4))\nplt.scatter(no_movies_voted.index,no_movies_voted,color='mediumseagreen')\nplt.axhline(y=50,color='r')\nplt.xlabel('UserId')\nplt.ylabel('No. of votes by user')\nplt.show()",
"_____no_output_____"
],
[
"final_dataset=final_dataset.loc[:,no_movies_voted[no_movies_voted > 50].index]\nfinal_dataset",
"_____no_output_____"
]
],
[
[
"#### Removing sparsity",
"_____no_output_____"
]
],
[
[
"sample = np.array([[0,0,3,0,0],[4,0,0,0,2],[0,0,0,0,1]])\nsparsity = 1.0 - ( np.count_nonzero(sample) / float(sample.size) )\nprint(sparsity)",
"0.7333333333333334\n"
],
[
"csr_sample = csr_matrix(sample)\nprint(csr_sample)",
" (0, 2)\t3\n (1, 0)\t4\n (1, 4)\t2\n (2, 4)\t1\n"
],
[
"csr_data = csr_matrix(final_dataset.values)\nfinal_dataset.reset_index(inplace=True)",
"_____no_output_____"
]
],
[
[
"#### Making the movie recommendation system model",
"_____no_output_____"
]
],
[
[
"knn = NearestNeighbors(metric='cosine', algorithm='brute', n_neighbors=20, n_jobs=-1)\nknn.fit(csr_data)",
"_____no_output_____"
]
],
[
[
"#### Making the recommendation function",
"_____no_output_____"
]
],
[
[
"def get_movie_recommendation(movie_name):\n n_movies_to_reccomend = 10\n movie_list = movies[movies['title']].str.contains(movie_name) \n if len(movie_list): \n movie_idx = movie_list.iloc[0]['movieId']\n movie_idx = final_dataset[final_dataset['movieId'] == movie_idx].index[0]\n distances , indices = knn.kneighbors(csr_data[movie_idx],n_neighbors=n_movies_to_reccomend+1) \n rec_movie_indices = sorted(list(zip(indices.squeeze().tolist(),distances.squeeze().tolist())),key=lambda x: x[1])[:0:-1]\n recommend_frame = []\n for val in rec_movie_indices:\n movie_idx = final_dataset.iloc[val[0]]['movieId']\n idx = movies[movies['movieId'] == movie_idx].index\n recommend_frame.append({'Title':movies.iloc[idx]['title'].values[0],'Distance':val[1]})\n df = pd.DataFrame(recommend_frame,index=range(1,n_movies_to_reccomend+1))\n return df\n else:\n return \"No movies found. Please check your input\"",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4acc2a330f92a3749f9d62e5587ca38cddce7336
| 21,076 |
ipynb
|
Jupyter Notebook
|
Day4.ipynb
|
DamianJad/dw_matrix
|
751a5636397869cb78b8ef91bcf7bcd4dc177e47
|
[
"MIT"
] | null | null | null |
Day4.ipynb
|
DamianJad/dw_matrix
|
751a5636397869cb78b8ef91bcf7bcd4dc177e47
|
[
"MIT"
] | null | null | null |
Day4.ipynb
|
DamianJad/dw_matrix
|
751a5636397869cb78b8ef91bcf7bcd4dc177e47
|
[
"MIT"
] | null | null | null | 21,076 | 21,076 | 0.748529 |
[
[
[
"import pandas as pd\nimport numpy as np\nfrom sklearn.tree import DecisionTreeRegressor\nfrom sklearn.metrics import mean_absolute_error\nfrom sklearn.model_selection import cross_val_score",
"_____no_output_____"
],
[
"cd \"/content/drive/My Drive/Colab Notebooks/dw_matrix\"",
"/content/drive/My Drive/Colab Notebooks/dw_matrix\n"
],
[
"df = pd.read_csv('data/men_shoes.csv', low_memory=False)\ndf.shape",
"_____no_output_____"
],
[
"mean_price = np.mean(df['prices_amountmin'])\nmean_price",
"_____no_output_____"
],
[
"y_true = df.prices_amountmin\ny_pred = [mean_price] * y_true.shape[0]\n\nmean_absolute_error(y_true,y_pred)",
"_____no_output_____"
],
[
"np.log1p(df.prices_amountmin).hist(bins=100)",
"_____no_output_____"
],
[
"y_true = df.prices_amountmin\ny_pred = [np.median(y_true)] * y_true.shape[0]\n\nmean_absolute_error(y_true,y_pred)",
"_____no_output_____"
],
[
"y_true = df.prices_amountmin\nprice_log_mean = np.expm1(np.mean(np.log1p(y_true)))\ny_pred = [price_log_mean] * y_true.shape[0]\n\nmean_absolute_error(y_true,y_pred)",
"_____no_output_____"
],
[
"df['brand_cat'] = df['brand'].factorize()[0]",
"_____no_output_____"
],
[
"feats = ['brand_cat']\n\ndef runs_model(feats):\n X = df[feats].values\n y = df['prices_amountmin'].values\n\n model = DecisionTreeRegressor(max_depth=5)\n\n scores = -1 * cross_val_score(model, X, y, scoring='neg_mean_absolute_error')\n return np.mean(scores), np.std(scores)",
"_____no_output_____"
],
[
"runs_model(feats)",
"_____no_output_____"
],
[
"df.columns",
"_____no_output_____"
],
[
"df.categories.value_counts()",
"_____no_output_____"
],
[
"df['categories_cat'] = df['categories'].factorize()[0]",
"_____no_output_____"
],
[
"runs_model(['categories_cat'])",
"_____no_output_____"
],
[
"runs_model(['categories_cat','brand_cat'])",
"_____no_output_____"
],
[
"df.manufacturer.value_counts()",
"_____no_output_____"
],
[
"df['manufacturer_cat'] = df['manufacturer'].factorize()[0]",
"_____no_output_____"
],
[
"runs_model(['manufacturer_cat'])",
"_____no_output_____"
],
[
"runs_model(['manufacturer_cat','categories_cat','brand_cat'])",
"_____no_output_____"
],
[
"df['prices_merchant_cat'] = df['prices_merchant'].factorize()[0]",
"_____no_output_____"
],
[
"runs_model(['manufacturer_cat','categories_cat','brand_cat','prices_merchant_cat'])",
"_____no_output_____"
],
[
"runs_model(['categories_cat','brand_cat','prices_merchant_cat'])",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acc3273a59295d34799cff9a6194ae64bed5059
| 26,440 |
ipynb
|
Jupyter Notebook
|
06 - Work with Data.ipynb
|
LucaSavio/mslearn-dp100
|
2ab26b99f94b6fa9feeedc3c3cff8318af71a4eb
|
[
"MIT"
] | 1 |
2022-03-13T16:20:07.000Z
|
2022-03-13T16:20:07.000Z
|
06 - Work with Data.ipynb
|
vighneshutamse/mslearn-dp100
|
a427c01a9b6c50fe30c39b9da1058ea4e9fae6ea
|
[
"MIT"
] | null | null | null |
06 - Work with Data.ipynb
|
vighneshutamse/mslearn-dp100
|
a427c01a9b6c50fe30c39b9da1058ea4e9fae6ea
|
[
"MIT"
] | 1 |
2021-08-30T00:45:35.000Z
|
2021-08-30T00:45:35.000Z
| 43.774834 | 573 | 0.62969 |
[
[
[
"# Work with Data\n\nData is the foundation on which machine learning models are built. Managing data centrally in the cloud, and making it accessible to teams of data scientists who are running experiments and training models on multiple workstations and compute targets is an important part of any professional data science solution.\n\nIn this notebook, you'll explore two Azure Machine Learning objects for working with data: *datastores*, and *datasets*.",
"_____no_output_____"
],
[
"## Connect to your workspace\n\nTo get started, connect to your workspace.\n\n> **Note**: If you haven't already established an authenticated session with your Azure subscription, you'll be prompted to authenticate by clicking a link, entering an authentication code, and signing into Azure.",
"_____no_output_____"
]
],
[
[
"import azureml.core\nfrom azureml.core import Workspace\n\n# Load the workspace from the saved config file\nws = Workspace.from_config()\nprint('Ready to use Azure ML {} to work with {}'.format(azureml.core.VERSION, ws.name))",
"_____no_output_____"
]
],
[
[
"## Work with datastores\n\nIn Azure ML, *datastores* are references to storage locations, such as Azure Storage blob containers. Every workspace has a default datastore - usually the Azure storage blob container that was created with the workspace. If you need to work with data that is stored in different locations, you can add custom datastores to your workspace and set any of them to be the default.\n\n### View datastores\n\nRun the following code to determine the datastores in your workspace:",
"_____no_output_____"
]
],
[
[
"# Get the default datastore\ndefault_ds = ws.get_default_datastore()\n\n# Enumerate all datastores, indicating which is the default\nfor ds_name in ws.datastores:\n print(ds_name, \"- Default =\", ds_name == default_ds.name)",
"_____no_output_____"
]
],
[
[
"You can also view and manage datastores in your workspace on the **Datastores** page for your workspace in [Azure Machine Learning studio](https://ml.azure.com).\n\n### Upload data to a datastore\n\nNow that you have determined the available datastores, you can upload files from your local file system to a datastore so that it will be accessible to experiments running in the workspace, regardless of where the experiment script is actually being run.",
"_____no_output_____"
]
],
[
[
"default_ds.upload_files(files=['./data/diabetes.csv', './data/diabetes2.csv'], # Upload the diabetes csv files in /data\n target_path='diabetes-data/', # Put it in a folder path in the datastore\n overwrite=True, # Replace existing files of the same name\n show_progress=True)",
"_____no_output_____"
]
],
[
[
"## Work with datasets\n\nAzure Machine Learning provides an abstraction for data in the form of *datasets*. A dataset is a versioned reference to a specific set of data that you may want to use in an experiment. Datasets can be *tabular* or *file*-based.\n\n### Create a tabular dataset\n\nLet's create a dataset from the diabetes data you uploaded to the datastore, and view the first 20 records. In this case, the data is in a structured format in a CSV file, so we'll use a *tabular* dataset.",
"_____no_output_____"
]
],
[
[
"from azureml.core import Dataset\n\n# Get the default datastore\ndefault_ds = ws.get_default_datastore()\n\n#Create a tabular dataset from the path on the datastore (this may take a short while)\ntab_data_set = Dataset.Tabular.from_delimited_files(path=(default_ds, 'diabetes-data/*.csv'))\n\n# Display the first 20 rows as a Pandas dataframe\ntab_data_set.take(20).to_pandas_dataframe()",
"_____no_output_____"
]
],
[
[
"As you can see in the code above, it's easy to convert a tabular dataset to a Pandas dataframe, enabling you to work with the data using common python techniques.\n\n### Create a file Dataset\n\nThe dataset you created is a *tabular* dataset that can be read as a dataframe containing all of the data in the structured files that are included in the dataset definition. This works well for tabular data, but in some machine learning scenarios you might need to work with data that is unstructured; or you may simply want to handle reading the data from files in your own code. To accomplish this, you can use a *file* dataset, which creates a list of file paths in a virtual mount point, which you can use to read the data in the files.",
"_____no_output_____"
]
],
[
[
"#Create a file dataset from the path on the datastore (this may take a short while)\nfile_data_set = Dataset.File.from_files(path=(default_ds, 'diabetes-data/*.csv'))\n\n# Get the files in the dataset\nfor file_path in file_data_set.to_path():\n print(file_path)",
"_____no_output_____"
]
],
[
[
"### Register datasets\n\nNow that you have created datasets that reference the diabetes data, you can register them to make them easily accessible to any experiment being run in the workspace.\n\nWe'll register the tabular dataset as **diabetes dataset**, and the file dataset as **diabetes files**.",
"_____no_output_____"
]
],
[
[
"# Register the tabular dataset\ntry:\n tab_data_set = tab_data_set.register(workspace=ws, \n name='diabetes dataset',\n description='diabetes data',\n tags = {'format':'CSV'},\n create_new_version=True)\nexcept Exception as ex:\n print(ex)\n\n# Register the file dataset\ntry:\n file_data_set = file_data_set.register(workspace=ws,\n name='diabetes file dataset',\n description='diabetes files',\n tags = {'format':'CSV'},\n create_new_version=True)\nexcept Exception as ex:\n print(ex)\n\nprint('Datasets registered')",
"_____no_output_____"
]
],
[
[
"You can view and manage datasets on the **Datasets** page for your workspace in [Azure Machine Learning studio](https://ml.azure.com). You can also get a list of datasets from the workspace object:",
"_____no_output_____"
]
],
[
[
"print(\"Datasets:\")\nfor dataset_name in list(ws.datasets.keys()):\n dataset = Dataset.get_by_name(ws, dataset_name)\n print(\"\\t\", dataset.name, 'version', dataset.version)",
"_____no_output_____"
]
],
[
[
"The ability to version datasets enables you to redefine datasets without breaking existing experiments or pipelines that rely on previous definitions. By default, the latest version of a named dataset is returned, but you can retrieve a specific version of a dataset by specifying the version number, like this:\n\n```python\ndataset_v1 = Dataset.get_by_name(ws, 'diabetes dataset', version = 1)\n```\n\n\n### Train a model from a tabular dataset\n\nNow that you have datasets, you're ready to start training models from them. You can pass datasets to scripts as *inputs* in the estimator being used to run the script.\n\nRun the following two code cells to create:\n\n1. A folder named **diabetes_training_from_tab_dataset**\n2. A script that trains a classification model by using a tabular dataset that is passed to it as an argument.",
"_____no_output_____"
]
],
[
[
"import os\n\n# Create a folder for the experiment files\nexperiment_folder = 'diabetes_training_from_tab_dataset'\nos.makedirs(experiment_folder, exist_ok=True)\nprint(experiment_folder, 'folder created')",
"_____no_output_____"
],
[
"%%writefile $experiment_folder/diabetes_training.py\n# Import libraries\nimport os\nimport argparse\nfrom azureml.core import Run, Dataset\nimport pandas as pd\nimport numpy as np\nimport joblib\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.linear_model import LogisticRegression\nfrom sklearn.metrics import roc_auc_score\nfrom sklearn.metrics import roc_curve\n\n# Get the script arguments (regularization rate and training dataset ID)\nparser = argparse.ArgumentParser()\nparser.add_argument('--regularization', type=float, dest='reg_rate', default=0.01, help='regularization rate')\nparser.add_argument(\"--input-data\", type=str, dest='training_dataset_id', help='training dataset')\nargs = parser.parse_args()\n\n# Set regularization hyperparameter (passed as an argument to the script)\nreg = args.reg_rate\n\n# Get the experiment run context\nrun = Run.get_context()\n\n# Get the training dataset\nprint(\"Loading Data...\")\ndiabetes = run.input_datasets['training_data'].to_pandas_dataframe()\n\n# Separate features and labels\nX, y = diabetes[['Pregnancies','PlasmaGlucose','DiastolicBloodPressure','TricepsThickness','SerumInsulin','BMI','DiabetesPedigree','Age']].values, diabetes['Diabetic'].values\n\n# Split data into training set and test set\nX_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=0)\n\n# Train a logistic regression model\nprint('Training a logistic regression model with regularization rate of', reg)\nrun.log('Regularization Rate', np.float(reg))\nmodel = LogisticRegression(C=1/reg, solver=\"liblinear\").fit(X_train, y_train)\n\n# calculate accuracy\ny_hat = model.predict(X_test)\nacc = np.average(y_hat == y_test)\nprint('Accuracy:', acc)\nrun.log('Accuracy', np.float(acc))\n\n# calculate AUC\ny_scores = model.predict_proba(X_test)\nauc = roc_auc_score(y_test,y_scores[:,1])\nprint('AUC: ' + str(auc))\nrun.log('AUC', np.float(auc))\n\nos.makedirs('outputs', exist_ok=True)\n# note file saved in the outputs folder is automatically uploaded into experiment record\njoblib.dump(value=model, filename='outputs/diabetes_model.pkl')\n\nrun.complete()",
"_____no_output_____"
]
],
[
[
"> **Note**: In the script, the dataset is passed as a parameter (or argument). In the case of a tabular dataset, this argument will contain the ID of the registered dataset; so you could write code in the script to get the experiment's workspace from the run context, and then get the dataset using its ID; like this:\n>\n> ```\n> run = Run.get_context()\n> ws = run.experiment.workspace\n> dataset = Dataset.get_by_id(ws, id=args.training_dataset_id)\n> diabetes = dataset.to_pandas_dataframe()\n> ```\n>\n> However, Azure Machine Learning runs automatically identify arguments that reference named datasets and add them to the run's **input_datasets** collection, so you can also retrieve the dataset from this collection by specifying its \"friendly name\" (which as you'll see shortly, is specified in the argument definition in the script run configuration for the experiment). This is the approach taken in the script above.\n\nNow you can run a script as an experiment, defining an argument for the training dataset, which is read by the script.\n\n> **Note**: The **Dataset** class depends on some components in the **azureml-dataprep** package, which includes optional support for **pandas** that is used by the **to_pandas_dataframe()** method. So you need to include this package in the environment where the training experiment will be run.",
"_____no_output_____"
]
],
[
[
"from azureml.core import Experiment, ScriptRunConfig, Environment\nfrom azureml.core.conda_dependencies import CondaDependencies\nfrom azureml.widgets import RunDetails\n\n\n# Create a Python environment for the experiment\nsklearn_env = Environment(\"sklearn-env\")\n\n# Ensure the required packages are installed (we need scikit-learn, Azure ML defaults, and Azure ML dataprep)\npackages = CondaDependencies.create(conda_packages=['scikit-learn','pip'],\n pip_packages=['azureml-defaults','azureml-dataprep[pandas]'])\nsklearn_env.python.conda_dependencies = packages\n\n# Get the training dataset\ndiabetes_ds = ws.datasets.get(\"diabetes dataset\")\n\n# Create a script config\nscript_config = ScriptRunConfig(source_directory=experiment_folder,\n script='diabetes_training.py',\n arguments = ['--regularization', 0.1, # Regularizaton rate parameter\n '--input-data', diabetes_ds.as_named_input('training_data')], # Reference to dataset\n environment=sklearn_env) \n\n# submit the experiment\nexperiment_name = 'mslearn-train-diabetes'\nexperiment = Experiment(workspace=ws, name=experiment_name)\nrun = experiment.submit(config=script_config)\nRunDetails(run).show()\nrun.wait_for_completion()",
"_____no_output_____"
]
],
[
[
"> **Note:** The **--input-data** argument passes the dataset as a *named input* that includes a *friendly name* for the dataset, which is used by the script to read it from the **input_datasets** collection in the experiment run. The string value in the **--input-data** argument is actually the registered dataset's ID. As an alternative approach, you could simply pass `diabetes_ds.id`, in which case the script can access the dataset ID from the script arguments and use it to get the dataset from the workspace, but not from the **input_datasets** collection.\n\nThe first time the experiment is run, it may take some time to set up the Python environment - subsequent runs will be quicker.\n\nWhen the experiment has completed, in the widget, view the **azureml-logs/70_driver_log.txt** output log and the metrics generated by the run.\n\n### Register the trained model\n\nAs with any training experiment, you can retrieve the trained model and register it in your Azure Machine Learning workspace.",
"_____no_output_____"
]
],
[
[
"from azureml.core import Model\n\nrun.register_model(model_path='outputs/diabetes_model.pkl', model_name='diabetes_model',\n tags={'Training context':'Tabular dataset'}, properties={'AUC': run.get_metrics()['AUC'], 'Accuracy': run.get_metrics()['Accuracy']})\n\nfor model in Model.list(ws):\n print(model.name, 'version:', model.version)\n for tag_name in model.tags:\n tag = model.tags[tag_name]\n print ('\\t',tag_name, ':', tag)\n for prop_name in model.properties:\n prop = model.properties[prop_name]\n print ('\\t',prop_name, ':', prop)\n print('\\n')",
"_____no_output_____"
]
],
[
[
"### Train a model from a file dataset\n\nYou've seen how to train a model using training data in a *tabular* dataset; but what about a *file* dataset?\n\nWhen you're using a file dataset, the dataset argument passed to the script represents a mount point containing file paths. How you read the data from these files depends on the kind of data in the files and what you want to do with it. In the case of the diabetes CSV files, you can use the Python **glob** module to create a list of files in the virtual mount point defined by the dataset, and read them all into Pandas dataframes that are concatenated into a single dataframe.\n\nRun the following two code cells to create:\n\n1. A folder named **diabetes_training_from_file_dataset**\n2. A script that trains a classification model by using a file dataset that is passed to is as an *input*.",
"_____no_output_____"
]
],
[
[
"import os\n\n# Create a folder for the experiment files\nexperiment_folder = 'diabetes_training_from_file_dataset'\nos.makedirs(experiment_folder, exist_ok=True)\nprint(experiment_folder, 'folder created')",
"_____no_output_____"
],
[
"%%writefile $experiment_folder/diabetes_training.py\n# Import libraries\nimport os\nimport argparse\nfrom azureml.core import Dataset, Run\nimport pandas as pd\nimport numpy as np\nimport joblib\nfrom sklearn.model_selection import train_test_split\nfrom sklearn.linear_model import LogisticRegression\nfrom sklearn.metrics import roc_auc_score\nfrom sklearn.metrics import roc_curve\nimport glob\n\n# Get script arguments (rgularization rate and file dataset mount point)\nparser = argparse.ArgumentParser()\nparser.add_argument('--regularization', type=float, dest='reg_rate', default=0.01, help='regularization rate')\nparser.add_argument('--input-data', type=str, dest='dataset_folder', help='data mount point')\nargs = parser.parse_args()\n\n# Set regularization hyperparameter (passed as an argument to the script)\nreg = args.reg_rate\n\n# Get the experiment run context\nrun = Run.get_context()\n\n# load the diabetes dataset\nprint(\"Loading Data...\")\ndata_path = run.input_datasets['training_files'] # Get the training data path from the input\n# (You could also just use args.dataset_folder if you don't want to rely on a hard-coded friendly name)\n\n# Read the files\nall_files = glob.glob(data_path + \"/*.csv\")\ndiabetes = pd.concat((pd.read_csv(f) for f in all_files), sort=False)\n\n# Separate features and labels\nX, y = diabetes[['Pregnancies','PlasmaGlucose','DiastolicBloodPressure','TricepsThickness','SerumInsulin','BMI','DiabetesPedigree','Age']].values, diabetes['Diabetic'].values\n\n# Split data into training set and test set\nX_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=0)\n\n# Train a logistic regression model\nprint('Training a logistic regression model with regularization rate of', reg)\nrun.log('Regularization Rate', np.float(reg))\nmodel = LogisticRegression(C=1/reg, solver=\"liblinear\").fit(X_train, y_train)\n\n# calculate accuracy\ny_hat = model.predict(X_test)\nacc = np.average(y_hat == y_test)\nprint('Accuracy:', acc)\nrun.log('Accuracy', np.float(acc))\n\n# calculate AUC\ny_scores = model.predict_proba(X_test)\nauc = roc_auc_score(y_test,y_scores[:,1])\nprint('AUC: ' + str(auc))\nrun.log('AUC', np.float(auc))\n\nos.makedirs('outputs', exist_ok=True)\n# note file saved in the outputs folder is automatically uploaded into experiment record\njoblib.dump(value=model, filename='outputs/diabetes_model.pkl')\n\nrun.complete()",
"_____no_output_____"
]
],
[
[
"Just as with tabular datasets, you can retrieve a file dataset from the **input_datasets** collection by using its friendly name. You can also retrieve it from the script argument, which in the case of a file dataset contains a mount path to the files (rather than the dataset ID passed for a tabular dataset).\n\nNext we need to change the way we pass the dataset to the script - it needs to define a path from which the script can read the files. You can use either the **as_download** or **as_mount** method to do this. Using **as_download** causes the files in the file dataset to be downloaded to a temporary location on the compute where the script is being run, while **as_mount** creates a mount point from which the files can be streamed directly from the datasetore.\n\nYou can combine the access method with the **as_named_input** method to include the dataset in the **input_datasets** collection in the experiment run (if you omit this, for example by setting the argument to `diabetes_ds.as_mount()`, the script will be able to access the dataset mount point from the script arguments, but not from the **input_datasets** collection).",
"_____no_output_____"
]
],
[
[
"from azureml.core import Experiment\nfrom azureml.widgets import RunDetails\n\n\n# Get the training dataset\ndiabetes_ds = ws.datasets.get(\"diabetes file dataset\")\n\n# Create a script config\nscript_config = ScriptRunConfig(source_directory=experiment_folder,\n script='diabetes_training.py',\n arguments = ['--regularization', 0.1, # Regularizaton rate parameter\n '--input-data', diabetes_ds.as_named_input('training_files').as_download()], # Reference to dataset location\n environment=sklearn_env) # Use the environment created previously\n\n# submit the experiment\nexperiment_name = 'mslearn-train-diabetes'\nexperiment = Experiment(workspace=ws, name=experiment_name)\nrun = experiment.submit(config=script_config)\nRunDetails(run).show()\nrun.wait_for_completion()",
"_____no_output_____"
]
],
[
[
"When the experiment has completed, in the widget, view the **azureml-logs/70_driver_log.txt** output log to verify that the files in the file dataset were downloaded to a temporary folder to enable the script to read the files.\n\n### Register the trained model\n\nOnce again, you can register the model that was trained by the experiment.",
"_____no_output_____"
]
],
[
[
"from azureml.core import Model\n\nrun.register_model(model_path='outputs/diabetes_model.pkl', model_name='diabetes_model',\n tags={'Training context':'File dataset'}, properties={'AUC': run.get_metrics()['AUC'], 'Accuracy': run.get_metrics()['Accuracy']})\n\nfor model in Model.list(ws):\n print(model.name, 'version:', model.version)\n for tag_name in model.tags:\n tag = model.tags[tag_name]\n print ('\\t',tag_name, ':', tag)\n for prop_name in model.properties:\n prop = model.properties[prop_name]\n print ('\\t',prop_name, ':', prop)\n print('\\n')",
"_____no_output_____"
]
],
[
[
"> **More Information**: For more information about training with datasets, see [Training with Datasets](https://docs.microsoft.com/azure/machine-learning/how-to-train-with-datasets) in the Azure ML documentation.",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
4acc37ec60cfbbdbc7fc7195e8dd3c2cd59f84e6
| 146,927 |
ipynb
|
Jupyter Notebook
|
Notebook dump/Final_Tagging.ipynb
|
mohit-bags/BERT-Notebooks
|
71971a57d2e1eda701a93fa3bfcca75c2ec95ca8
|
[
"MIT"
] | null | null | null |
Notebook dump/Final_Tagging.ipynb
|
mohit-bags/BERT-Notebooks
|
71971a57d2e1eda701a93fa3bfcca75c2ec95ca8
|
[
"MIT"
] | null | null | null |
Notebook dump/Final_Tagging.ipynb
|
mohit-bags/BERT-Notebooks
|
71971a57d2e1eda701a93fa3bfcca75c2ec95ca8
|
[
"MIT"
] | null | null | null | 34.932715 | 331 | 0.452259 |
[
[
[
"import numpy as np\nimport pandas as pd",
"_____no_output_____"
],
[
"data=pd.read_csv(\"tech_sort1k.csv\")\ndata.head(15)",
"_____no_output_____"
],
[
"data=data.drop(columns=[\"id\",\"Note\"])",
"_____no_output_____"
],
[
"#replacing null vals with empty list\ndata['exact_matched_patt_contextual'] = [ [] if x is np.NaN else x for x in data['exact_matched_patt_contextual'] ]\n",
"_____no_output_____"
],
[
"import nltk\nimport re\nfrom bs4 import BeautifulSoup\n\nnltk.download(\"punkt\")\n\nREPLACE_BY_SPACE_RE = re.compile('[/(){}\\[\\]\\|@,;]')\nBAD_SYMBOLS_RE = re.compile('[^0-9a-z #+_]')\n\ndef clean_text(text):\n \"\"\"\n text: a string\n \n return: modified initial string\n \"\"\"\n text = BeautifulSoup(text, \"lxml\").text # HTML decoding\n text = text.lower() # lowercase text\n text = REPLACE_BY_SPACE_RE.sub(' ', text) # replace REPLACE_BY_SPACE_RE symbols by space in text\n text = BAD_SYMBOLS_RE.sub('', text) # delete symbols which are in BAD_SYMBOLS_RE from text\n return text\n",
"[nltk_data] Downloading package punkt to\n[nltk_data] /Users/mohitbagaria/nltk_data...\n[nltk_data] Package punkt is already up-to-date!\n"
],
[
"data['exact_matched_patt_contextual']=data['exact_matched_patt_contextual'].astype(str)",
"_____no_output_____"
],
[
"data.head(15)",
"_____no_output_____"
],
[
"#converting lists to strings with proper cleaning\ndata[\"to_match\"]=data[\"exact_matched_patt_contextual\"].apply(clean_text)",
"_____no_output_____"
],
[
"data.head(10)",
"_____no_output_____"
],
[
"data[\"summaries\"]=data[\"summaries\"].apply(clean_text)",
"_____no_output_____"
],
[
"def convert(lst):\n\treturn ' '.join(lst).split()",
"_____no_output_____"
],
[
"def givedata(tech_words,sentence):\n broken_list = convert([sentence])\n prev_tag=\"O\" # inital tag will be \"O\" or \"B\" or \"I\"\n tech_words=convert([tech_words])\n tag_col=[]\n for word in broken_list:\n curr_tag=\"O\" \n if word in tech_words:\n if (prev_tag=='B' or prev_tag=='I'): #if word found in dictionary\n curr_tag='I' #if prev_tag was already a tech word\n else:\n curr_tag='B' #if prev_tag was \"O\" (Not a tech word)\n prev_tag=curr_tag\n tag_col.append(curr_tag) #adding tag\n return pd.DataFrame(list(zip(broken_list, tag_col)),columns =['Word', 'Tag'])",
"_____no_output_____"
],
[
"final_data=pd.DataFrame()\n\nfor i in range(0,len(data)):\n# print(i)\n temp = givedata(data.iloc[i,2],data[\"summaries\"][i])\n length = len(temp)\n wordd=\"Sentence :\"+str(i+1) #sentence no.\n a=[wordd]*length\n temp.insert(0,\"Sentence #\",a)\n final_data = final_data.append(temp, ignore_index=True) #appending sentences in the required format",
"_____no_output_____"
],
[
"import seaborn as sns\n\nsns.countplot(final_data['Tag'])",
"/Users/mohitbagaria/anaconda3/lib/python3.7/site-packages/seaborn/_decorators.py:43: FutureWarning: Pass the following variable as a keyword arg: x. From version 0.12, the only valid positional argument will be `data`, and passing other arguments without an explicit keyword will result in an error or misinterpretation.\n FutureWarning\n"
],
[
"final_data['Tag'].value_counts()",
"_____no_output_____"
],
[
"# final_data.loc[final_data['Tag']=='I']\n# final_data.loc[final_data['Tag'].isin(['I','B'])]",
"_____no_output_____"
],
[
"\"\"\"All tagged data, without any filter\"\"\"\n# final_data.to_csv('TECH_bio_tagging.csv',index=False) ",
"_____no_output_____"
],
[
"final_data.head()",
"_____no_output_____"
],
[
"tech_dict = {}\nfor j in data['exact_matched_patt_contextual']:\n if j not in tech_dict:\n tech_dict[j] = 1\n else:\n tech_dict[j] += 1\n",
"_____no_output_____"
],
[
"tech_dict",
"_____no_output_____"
],
[
"data[\"exact_matched_patt_contextual\"] = data[\"exact_matched_patt_contextual\"].apply(eval)\n",
"_____no_output_____"
],
[
"for i, l in enumerate(data[\"exact_matched_patt_contextual\"]):\n print(\"list\",i,\"is\",type(l))\n\n\n",
"list 0 is <class 'list'>\nlist 1 is <class 'list'>\nlist 2 is <class 'list'>\nlist 3 is <class 'list'>\nlist 4 is <class 'list'>\nlist 5 is <class 'list'>\nlist 6 is <class 'list'>\nlist 7 is <class 'list'>\nlist 8 is <class 'list'>\nlist 9 is <class 'list'>\nlist 10 is <class 'list'>\nlist 11 is <class 'list'>\nlist 12 is <class 'list'>\nlist 13 is <class 'list'>\nlist 14 is <class 'list'>\nlist 15 is <class 'list'>\nlist 16 is <class 'list'>\nlist 17 is <class 'list'>\nlist 18 is <class 'list'>\nlist 19 is <class 'list'>\nlist 20 is <class 'list'>\nlist 21 is <class 'list'>\nlist 22 is <class 'list'>\nlist 23 is <class 'list'>\nlist 24 is <class 'list'>\nlist 25 is <class 'list'>\nlist 26 is <class 'list'>\nlist 27 is <class 'list'>\nlist 28 is <class 'list'>\nlist 29 is <class 'list'>\nlist 30 is <class 'list'>\nlist 31 is <class 'list'>\nlist 32 is <class 'list'>\nlist 33 is <class 'list'>\nlist 34 is <class 'list'>\nlist 35 is <class 'list'>\nlist 36 is <class 'list'>\nlist 37 is <class 'list'>\nlist 38 is <class 'list'>\nlist 39 is <class 'list'>\nlist 40 is <class 'list'>\nlist 41 is <class 'list'>\nlist 42 is <class 'list'>\nlist 43 is <class 'list'>\nlist 44 is <class 'list'>\nlist 45 is <class 'list'>\nlist 46 is <class 'list'>\nlist 47 is <class 'list'>\nlist 48 is <class 'list'>\nlist 49 is <class 'list'>\nlist 50 is <class 'list'>\nlist 51 is <class 'list'>\nlist 52 is <class 'list'>\nlist 53 is <class 'list'>\nlist 54 is <class 'list'>\nlist 55 is <class 'list'>\nlist 56 is <class 'list'>\nlist 57 is <class 'list'>\nlist 58 is <class 'list'>\nlist 59 is <class 'list'>\nlist 60 is <class 'list'>\nlist 61 is <class 'list'>\nlist 62 is <class 'list'>\nlist 63 is <class 'list'>\nlist 64 is <class 'list'>\nlist 65 is <class 'list'>\nlist 66 is <class 'list'>\nlist 67 is <class 'list'>\nlist 68 is <class 'list'>\nlist 69 is <class 'list'>\nlist 70 is <class 'list'>\nlist 71 is <class 'list'>\nlist 72 is <class 'list'>\nlist 73 is <class 'list'>\nlist 74 is <class 'list'>\nlist 75 is <class 'list'>\nlist 76 is <class 'list'>\nlist 77 is <class 'list'>\nlist 78 is <class 'list'>\nlist 79 is <class 'list'>\nlist 80 is <class 'list'>\nlist 81 is <class 'list'>\nlist 82 is <class 'list'>\nlist 83 is <class 'list'>\nlist 84 is <class 'list'>\nlist 85 is <class 'list'>\nlist 86 is <class 'list'>\nlist 87 is <class 'list'>\nlist 88 is <class 'list'>\nlist 89 is <class 'list'>\nlist 90 is <class 'list'>\nlist 91 is <class 'list'>\nlist 92 is <class 'list'>\nlist 93 is <class 'list'>\nlist 94 is <class 'list'>\nlist 95 is <class 'list'>\nlist 96 is <class 'list'>\nlist 97 is <class 'list'>\nlist 98 is <class 'list'>\nlist 99 is <class 'list'>\nlist 100 is <class 'list'>\nlist 101 is <class 'list'>\nlist 102 is <class 'list'>\nlist 103 is <class 'list'>\nlist 104 is <class 'list'>\nlist 105 is <class 'list'>\nlist 106 is <class 'list'>\nlist 107 is <class 'list'>\nlist 108 is <class 'list'>\nlist 109 is <class 'list'>\nlist 110 is <class 'list'>\nlist 111 is <class 'list'>\nlist 112 is <class 'list'>\nlist 113 is <class 'list'>\nlist 114 is <class 'list'>\nlist 115 is <class 'list'>\nlist 116 is <class 'list'>\nlist 117 is <class 'list'>\nlist 118 is <class 'list'>\nlist 119 is <class 'list'>\nlist 120 is <class 'list'>\nlist 121 is <class 'list'>\nlist 122 is <class 'list'>\nlist 123 is <class 'list'>\nlist 124 is <class 'list'>\nlist 125 is <class 'list'>\nlist 126 is <class 'list'>\nlist 127 is <class 'list'>\nlist 128 is <class 'list'>\nlist 129 is <class 'list'>\nlist 130 is <class 'list'>\nlist 131 is <class 'list'>\nlist 132 is <class 'list'>\nlist 133 is <class 'list'>\nlist 134 is <class 'list'>\nlist 135 is <class 'list'>\nlist 136 is <class 'list'>\nlist 137 is <class 'list'>\nlist 138 is <class 'list'>\nlist 139 is <class 'list'>\nlist 140 is <class 'list'>\nlist 141 is <class 'list'>\nlist 142 is <class 'list'>\nlist 143 is <class 'list'>\nlist 144 is <class 'list'>\nlist 145 is <class 'list'>\nlist 146 is <class 'list'>\nlist 147 is <class 'list'>\nlist 148 is <class 'list'>\nlist 149 is <class 'list'>\nlist 150 is <class 'list'>\nlist 151 is <class 'list'>\nlist 152 is <class 'list'>\nlist 153 is <class 'list'>\nlist 154 is <class 'list'>\nlist 155 is <class 'list'>\nlist 156 is <class 'list'>\nlist 157 is <class 'list'>\nlist 158 is <class 'list'>\nlist 159 is <class 'list'>\nlist 160 is <class 'list'>\nlist 161 is <class 'list'>\nlist 162 is <class 'list'>\nlist 163 is <class 'list'>\nlist 164 is <class 'list'>\nlist 165 is <class 'list'>\nlist 166 is <class 'list'>\nlist 167 is <class 'list'>\nlist 168 is <class 'list'>\nlist 169 is <class 'list'>\nlist 170 is <class 'list'>\nlist 171 is <class 'list'>\nlist 172 is <class 'list'>\nlist 173 is <class 'list'>\nlist 174 is <class 'list'>\nlist 175 is <class 'list'>\nlist 176 is <class 'list'>\nlist 177 is <class 'list'>\nlist 178 is <class 'list'>\nlist 179 is <class 'list'>\nlist 180 is <class 'list'>\nlist 181 is <class 'list'>\nlist 182 is <class 'list'>\nlist 183 is <class 'list'>\nlist 184 is <class 'list'>\nlist 185 is <class 'list'>\nlist 186 is <class 'list'>\nlist 187 is <class 'list'>\nlist 188 is <class 'list'>\nlist 189 is <class 'list'>\nlist 190 is <class 'list'>\nlist 191 is <class 'list'>\nlist 192 is <class 'list'>\nlist 193 is <class 'list'>\nlist 194 is <class 'list'>\nlist 195 is <class 'list'>\nlist 196 is <class 'list'>\nlist 197 is <class 'list'>\nlist 198 is <class 'list'>\nlist 199 is <class 'list'>\nlist 200 is <class 'list'>\nlist 201 is <class 'list'>\nlist 202 is <class 'list'>\nlist 203 is <class 'list'>\nlist 204 is <class 'list'>\nlist 205 is <class 'list'>\nlist 206 is <class 'list'>\nlist 207 is <class 'list'>\nlist 208 is <class 'list'>\nlist 209 is <class 'list'>\nlist 210 is <class 'list'>\nlist 211 is <class 'list'>\nlist 212 is <class 'list'>\nlist 213 is <class 'list'>\nlist 214 is <class 'list'>\nlist 215 is <class 'list'>\nlist 216 is <class 'list'>\nlist 217 is <class 'list'>\nlist 218 is <class 'list'>\nlist 219 is <class 'list'>\nlist 220 is <class 'list'>\nlist 221 is <class 'list'>\nlist 222 is <class 'list'>\nlist 223 is <class 'list'>\nlist 224 is <class 'list'>\nlist 225 is <class 'list'>\nlist 226 is <class 'list'>\nlist 227 is <class 'list'>\nlist 228 is <class 'list'>\nlist 229 is <class 'list'>\nlist 230 is <class 'list'>\nlist 231 is <class 'list'>\nlist 232 is <class 'list'>\nlist 233 is <class 'list'>\nlist 234 is <class 'list'>\nlist 235 is <class 'list'>\nlist 236 is <class 'list'>\nlist 237 is <class 'list'>\nlist 238 is <class 'list'>\nlist 239 is <class 'list'>\nlist 240 is <class 'list'>\nlist 241 is <class 'list'>\nlist 242 is <class 'list'>\nlist 243 is <class 'list'>\nlist 244 is <class 'list'>\nlist 245 is <class 'list'>\nlist 246 is <class 'list'>\nlist 247 is <class 'list'>\nlist 248 is <class 'list'>\nlist 249 is <class 'list'>\nlist 250 is <class 'list'>\nlist 251 is <class 'list'>\nlist 252 is <class 'list'>\nlist 253 is <class 'list'>\nlist 254 is <class 'list'>\nlist 255 is <class 'list'>\nlist 256 is <class 'list'>\nlist 257 is <class 'list'>\nlist 258 is <class 'list'>\nlist 259 is <class 'list'>\nlist 260 is <class 'list'>\nlist 261 is <class 'list'>\nlist 262 is <class 'list'>\nlist 263 is <class 'list'>\nlist 264 is <class 'list'>\nlist 265 is <class 'list'>\nlist 266 is <class 'list'>\nlist 267 is <class 'list'>\nlist 268 is <class 'list'>\nlist 269 is <class 'list'>\nlist 270 is <class 'list'>\nlist 271 is <class 'list'>\nlist 272 is <class 'list'>\nlist 273 is <class 'list'>\nlist 274 is <class 'list'>\nlist 275 is <class 'list'>\nlist 276 is <class 'list'>\nlist 277 is <class 'list'>\nlist 278 is <class 'list'>\nlist 279 is <class 'list'>\nlist 280 is <class 'list'>\nlist 281 is <class 'list'>\nlist 282 is <class 'list'>\nlist 283 is <class 'list'>\nlist 284 is <class 'list'>\nlist 285 is <class 'list'>\nlist 286 is <class 'list'>\nlist 287 is <class 'list'>\nlist 288 is <class 'list'>\nlist 289 is <class 'list'>\nlist 290 is <class 'list'>\nlist 291 is <class 'list'>\nlist 292 is <class 'list'>\nlist 293 is <class 'list'>\nlist 294 is <class 'list'>\nlist 295 is <class 'list'>\nlist 296 is <class 'list'>\nlist 297 is <class 'list'>\nlist 298 is <class 'list'>\nlist 299 is <class 'list'>\nlist 300 is <class 'list'>\nlist 301 is <class 'list'>\nlist 302 is <class 'list'>\nlist 303 is <class 'list'>\nlist 304 is <class 'list'>\nlist 305 is <class 'list'>\nlist 306 is <class 'list'>\nlist 307 is <class 'list'>\nlist 308 is <class 'list'>\nlist 309 is <class 'list'>\nlist 310 is <class 'list'>\nlist 311 is <class 'list'>\nlist 312 is <class 'list'>\nlist 313 is <class 'list'>\nlist 314 is <class 'list'>\nlist 315 is <class 'list'>\nlist 316 is <class 'list'>\nlist 317 is <class 'list'>\nlist 318 is <class 'list'>\nlist 319 is <class 'list'>\nlist 320 is <class 'list'>\nlist 321 is <class 'list'>\nlist 322 is <class 'list'>\nlist 323 is <class 'list'>\nlist 324 is <class 'list'>\nlist 325 is <class 'list'>\nlist 326 is <class 'list'>\nlist 327 is <class 'list'>\nlist 328 is <class 'list'>\nlist 329 is <class 'list'>\nlist 330 is <class 'list'>\nlist 331 is <class 'list'>\nlist 332 is <class 'list'>\nlist 333 is <class 'list'>\nlist 334 is <class 'list'>\nlist 335 is <class 'list'>\nlist 336 is <class 'list'>\nlist 337 is <class 'list'>\nlist 338 is <class 'list'>\nlist 339 is <class 'list'>\nlist 340 is <class 'list'>\nlist 341 is <class 'list'>\nlist 342 is <class 'list'>\nlist 343 is <class 'list'>\nlist 344 is <class 'list'>\nlist 345 is <class 'list'>\nlist 346 is <class 'list'>\nlist 347 is <class 'list'>\nlist 348 is <class 'list'>\nlist 349 is <class 'list'>\nlist 350 is <class 'list'>\nlist 351 is <class 'list'>\nlist 352 is <class 'list'>\nlist 353 is <class 'list'>\nlist 354 is <class 'list'>\nlist 355 is <class 'list'>\nlist 356 is <class 'list'>\nlist 357 is <class 'list'>\nlist 358 is <class 'list'>\nlist 359 is <class 'list'>\nlist 360 is <class 'list'>\nlist 361 is <class 'list'>\nlist 362 is <class 'list'>\nlist 363 is <class 'list'>\nlist 364 is <class 'list'>\nlist 365 is <class 'list'>\nlist 366 is <class 'list'>\nlist 367 is <class 'list'>\nlist 368 is <class 'list'>\nlist 369 is <class 'list'>\nlist 370 is <class 'list'>\nlist 371 is <class 'list'>\nlist 372 is <class 'list'>\nlist 373 is <class 'list'>\nlist 374 is <class 'list'>\nlist 375 is <class 'list'>\nlist 376 is <class 'list'>\nlist 377 is <class 'list'>\nlist 378 is <class 'list'>\nlist 379 is <class 'list'>\nlist 380 is <class 'list'>\nlist 381 is <class 'list'>\nlist 382 is <class 'list'>\nlist 383 is <class 'list'>\nlist 384 is <class 'list'>\nlist 385 is <class 'list'>\nlist 386 is <class 'list'>\nlist 387 is <class 'list'>\nlist 388 is <class 'list'>\nlist 389 is <class 'list'>\nlist 390 is <class 'list'>\nlist 391 is <class 'list'>\nlist 392 is <class 'list'>\nlist 393 is <class 'list'>\nlist 394 is <class 'list'>\nlist 395 is <class 'list'>\nlist 396 is <class 'list'>\nlist 397 is <class 'list'>\nlist 398 is <class 'list'>\nlist 399 is <class 'list'>\nlist 400 is <class 'list'>\nlist 401 is <class 'list'>\nlist 402 is <class 'list'>\nlist 403 is <class 'list'>\nlist 404 is <class 'list'>\nlist 405 is <class 'list'>\nlist 406 is <class 'list'>\nlist 407 is <class 'list'>\nlist 408 is <class 'list'>\nlist 409 is <class 'list'>\nlist 410 is <class 'list'>\nlist 411 is <class 'list'>\nlist 412 is <class 'list'>\nlist 413 is <class 'list'>\nlist 414 is <class 'list'>\nlist 415 is <class 'list'>\nlist 416 is <class 'list'>\nlist 417 is <class 'list'>\nlist 418 is <class 'list'>\nlist 419 is <class 'list'>\nlist 420 is <class 'list'>\nlist 421 is <class 'list'>\nlist 422 is <class 'list'>\nlist 423 is <class 'list'>\nlist 424 is <class 'list'>\nlist 425 is <class 'list'>\nlist 426 is <class 'list'>\nlist 427 is <class 'list'>\nlist 428 is <class 'list'>\nlist 429 is <class 'list'>\nlist 430 is <class 'list'>\nlist 431 is <class 'list'>\nlist 432 is <class 'list'>\nlist 433 is <class 'list'>\nlist 434 is <class 'list'>\nlist 435 is <class 'list'>\nlist 436 is <class 'list'>\nlist 437 is <class 'list'>\nlist 438 is <class 'list'>\nlist 439 is <class 'list'>\nlist 440 is <class 'list'>\nlist 441 is <class 'list'>\nlist 442 is <class 'list'>\nlist 443 is <class 'list'>\nlist 444 is <class 'list'>\nlist 445 is <class 'list'>\nlist 446 is <class 'list'>\nlist 447 is <class 'list'>\nlist 448 is <class 'list'>\nlist 449 is <class 'list'>\nlist 450 is <class 'list'>\nlist 451 is <class 'list'>\nlist 452 is <class 'list'>\n"
],
[
"tech_dict = {}\nfor i in data['exact_matched_patt_contextual']:\n for j in i:\n if j not in tech_dict:\n tech_dict[j] = 1\n else:\n tech_dict[j] += 1\n\n",
"_____no_output_____"
],
[
"tech_dict",
"_____no_output_____"
],
[
"len(tech_dict)",
"_____no_output_____"
],
[
"tech_list=[]\nfor i in tech_dict:\n tech_list.append(i)",
"_____no_output_____"
],
[
"tech_list",
"_____no_output_____"
],
[
"def in_techwords(tech_words,sentence):\n broken_list = convert([sentence])\n prev_tag=\"O\" # inital tag will be \"O\" or \"B\" or \"I\"\n tech_words=convert(tech_words)\n tag_col=[]\n for word in broken_list:\n curr_tag=\"O\" \n if word in tech_words:\n if (prev_tag=='B' or prev_tag=='I'): #if word found in dictionary\n curr_tag='I' #if prev_tag was already a tech word\n else:\n curr_tag='B' #if prev_tag was \"O\" (Not a tech word)\n prev_tag=curr_tag\n tag_col.append(curr_tag) #adding tag\n return pd.DataFrame(list(zip(broken_list, tag_col)),columns =['Word', 'Tag'])",
"_____no_output_____"
],
[
"final_data2=pd.DataFrame()\n\nfor i in range(0,len(data)):\n temp = in_techwords(tech_list,data[\"summaries\"][i])\n length = len(temp)\n wordd=\"Sentence :\"+str(i+1) #sentence no.\n a=[wordd]*length\n temp.insert(0,\"Sentence #\",a)\n final_data2 = final_data2.append(temp, ignore_index=True) #appending sentences in the required format",
"_____no_output_____"
],
[
"final_data2[\"Tag\"].value_counts()",
"_____no_output_____"
],
[
"def in_techwords_B(tech_words,sentence):\n broken_list = convert([sentence])\n prev_tag=\"O\" # inital tag will be \"O\" or \"B\" or \"I\"\n got_b=False\n tech_words=convert(tech_words)\n tag_col=[]\n for word in broken_list:\n curr_tag=\"O\" \n if word in tech_words:\n got_b=True\n if (prev_tag=='B' or prev_tag=='I'): #if word found in dictionary\n curr_tag='I' #if prev_tag was already a tech word\n else:\n curr_tag='B' #if prev_tag was \"O\" (Not a tech word)\n prev_tag=curr_tag\n tag_col.append(curr_tag) #adding tag\n return pd.DataFrame(list(zip(broken_list, tag_col)),columns =['Word', 'Tag']),got_b",
"_____no_output_____"
],
[
"final_data3=pd.DataFrame()\n\nfor i in range(0,len(data)):\n temp,flag = in_techwords_B(tech_list,data[\"summaries\"][i])\n length = len(temp)\n wordd=\"Sentence :\"+str(i+1) #sentence no.\n a=[wordd]*length\n temp.insert(0,\"Sentence #\",a)\n if flag==True:\n final_data3 = final_data3.append(temp, ignore_index=True) #appending sentences in the required format",
"_____no_output_____"
],
[
"final_data3[\"Tag\"].value_counts()",
"_____no_output_____"
],
[
"final_data2.to_csv(\"Combined_Match_tag.csv\")",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acc3a038f2b131165a8efda44d1c00678bea1ce
| 5,093 |
ipynb
|
Jupyter Notebook
|
SpeechText.ipynb
|
chusheng0505/Image-Captions-and-Generating-Images
|
b3af5d8327e610e5195a085d8053a835bf4d8246
|
[
"MIT"
] | null | null | null |
SpeechText.ipynb
|
chusheng0505/Image-Captions-and-Generating-Images
|
b3af5d8327e610e5195a085d8053a835bf4d8246
|
[
"MIT"
] | null | null | null |
SpeechText.ipynb
|
chusheng0505/Image-Captions-and-Generating-Images
|
b3af5d8327e610e5195a085d8053a835bf4d8246
|
[
"MIT"
] | null | null | null | 29.610465 | 96 | 0.545258 |
[
[
[
"import speech_recognition as sr\nfrom transformers import Wav2Vec2Processor, HubertForCTC,Wav2Vec2ForCTC\nimport soundfile as sf\nfrom datasets import load_dataset\nimport torch",
"_____no_output_____"
],
[
"pathSave = 'C:\\\\Users\\\\chushengtan\\\\Desktop\\\\'\nfilename = 'audio_file_test.wav'\ntimeout = 0.5\nwaiting_time = 10\n\nr = sr.Recognizer()\nwith sr.Microphone(device_index=1,sample_rate = 16000) as source:\n r.adjust_for_ambient_noise(source)\n print('請開始說話.....')\n audio = r.listen(source,\n timeout = timeout,\n phrase_time_limit = waiting_time)\n print('錄音結束.....')\nwith open(pathSave + filename,'wb') as file:\n file.write(audio.get_wav_data())",
"_____no_output_____"
]
],
[
[
"# 完成版",
"_____no_output_____"
]
],
[
[
"def speech2Wave(pathSave,filename,sample_rate = 16000,timeout = 0.5,waiting_time = 10):\n \"\"\"\n phrase_time_limit : waiting time for ending programming\n mic = sr.Microphone() # 查詢全部 microphones 的裝置名稱\n ref : https://realpython.com/python-speech-recognition/#working-with-microphones\n ref : https://github.com/Uberi/speech_recognition\n ref : https://github.com/pytorch/fairseq/tree/main/examples/hubert\n \"\"\"\n r = sr.Recognizer()\n with sr.Microphone(sample_rate = sample_rate) as source:\n r.adjust_for_ambient_noise(source)\n print('請開始說話.....')\n audio = r.listen(source,\n timeout = timeout,\n phrase_time_limit = waiting_time)\n print('錄音結束.....')\n with open(pathSave + filename,'wb') as file:\n file.write(audio.get_wav_data())\n\n \n\"\"\"\n------------------- loading model ------------------- \n\"\"\" \n\ndef Load_Model(processor_name,model_name):\n \"\"\"\n Load_Model(processor_name,model_name) : 載入使用模型\n \"\"\"\n processor = Wav2Vec2Processor.from_pretrained(processor_name)\n model = HubertForCTC.from_pretrained(model_name)\n return processor,model\n\ndef Speech2Text(audio_path,processor_name,model_name):\n \"\"\"\n Speech2Text(audio_path,processor_name,model_name) : 將語音轉換成文字\n --> audio_path : 語音檔案存放路徑位置 ; format : .wav\n \"\"\"\n processor , model = Load_Model(processor_name,model_name)\n# processor = Wav2Vec2Processor.from_pretrained(processor_name)\n# model = HubertForCTC.from_pretrained(model_name)\n speech,_ = sf.read(audio_path)\n input_values = processor(speech,return_tensors='pt',padding='longest').input_values\n logits = model(input_values).logits\n predicted_ids = torch.argmax(logits,dim=-1)\n return processor.decode(predicted_ids[0])",
"_____no_output_____"
],
[
"%%time\npathSave = 'C:\\\\Users\\\\chushengtan\\\\Desktop\\\\'\nfilename = 'audio_file_test.wav'\nspeech2Wave(pathSave=pathSave,filename=filename)",
"_____no_output_____"
],
[
"%%time\nprocessor_name = 'facebook/hubert-xlarge-ls960-ft'\nmodel_name = 'facebook/hubert-xlarge-ls960-ft'\naudio_path = 'C:\\\\Users\\\\chushengtan\\\\Desktop\\\\audio_file_test.wav'\nSpeech2Text(audio_path = audio_path,\n processor_name = processor_name,\n model_name = model_name)",
"_____no_output_____"
]
]
] |
[
"code",
"markdown",
"code"
] |
[
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
]
] |
4acc3d8abc55502c741c3f11142146239b783663
| 10,590 |
ipynb
|
Jupyter Notebook
|
temp_analysis_bonus_1_starter.ipynb
|
karahumanskiNU/sqlalchemy-challenge
|
c7d6641b71d31e47a9f61626c33442034a7fe330
|
[
"ADSL"
] | null | null | null |
temp_analysis_bonus_1_starter.ipynb
|
karahumanskiNU/sqlalchemy-challenge
|
c7d6641b71d31e47a9f61626c33442034a7fe330
|
[
"ADSL"
] | null | null | null |
temp_analysis_bonus_1_starter.ipynb
|
karahumanskiNU/sqlalchemy-challenge
|
c7d6641b71d31e47a9f61626c33442034a7fe330
|
[
"ADSL"
] | null | null | null | 25.035461 | 85 | 0.391501 |
[
[
[
"# Bonus: Temperature Analysis I",
"_____no_output_____"
]
],
[
[
"import pandas as pd\nfrom datetime import datetime as dt",
"_____no_output_____"
],
[
"# \"tobs\" is \"temperature observations\"\ntemp_df = pd.read_csv('Resources/hawaii_measurements.csv')\ntemp_df.head()",
"_____no_output_____"
],
[
"# Convert the date column format from string to datetime\ntemp_df['date']=pd.to_datetime(temp_df['date'])\ntemp_df.head()",
"_____no_output_____"
],
[
"# Set the date column as the DataFrame index\ntemp_df = temp_df.set_index('date')\ntemp_df.head()",
"_____no_output_____"
]
],
[
[
"### Compare June and December data across all years ",
"_____no_output_____"
]
],
[
[
"from scipy import stats",
"_____no_output_____"
],
[
"# Filter data for desired months\njun_df =temp_df.loc [temp_df.index.month == 6] \ndec_df = temp_df.loc [temp_df.index.month == 12] ",
"_____no_output_____"
],
[
"# Identify the average temperature for June\njune_avg = jun_df['tobs'].mean()\nprint(june_avg)",
"74.94411764705882\n"
],
[
"# Identify the average temperature for December\ndec_avg= dec_df['tobs'].mean()\nprint(dec_avg)",
"71.04152933421226\n"
],
[
"# Run paired t-test\nimport scipy.stats as st\nst.ttest_ind(jun_df['tobs'],dec_df['tobs'], equal_var=True )",
"_____no_output_____"
]
],
[
[
"### Analysis",
"_____no_output_____"
]
],
[
[
"#P value is 3.9025129038616655e-191 thus there is stat significants",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
]
] |
4acc5ce94615e68e206bce961dbcaef0f0c79b06
| 178,402 |
ipynb
|
Jupyter Notebook
|
Bike_Rentals_Forecast_Model.ipynb
|
Shrayansh19/Bike_Rentals_Forecast
|
28a053b988b04a3d0366e2b6a7e6bed7a94522c7
|
[
"MIT"
] | null | null | null |
Bike_Rentals_Forecast_Model.ipynb
|
Shrayansh19/Bike_Rentals_Forecast
|
28a053b988b04a3d0366e2b6a7e6bed7a94522c7
|
[
"MIT"
] | null | null | null |
Bike_Rentals_Forecast_Model.ipynb
|
Shrayansh19/Bike_Rentals_Forecast
|
28a053b988b04a3d0366e2b6a7e6bed7a94522c7
|
[
"MIT"
] | null | null | null | 213.911271 | 77,278 | 0.87652 |
[
[
[
"<a href=\"https://colab.research.google.com/github/Shrayansh19/Bike_Rentals_Forecast/blob/main/Bike_Rentals_Forecast_Model.ipynb\" target=\"_parent\"><img src=\"https://colab.research.google.com/assets/colab-badge.svg\" alt=\"Open In Colab\"/></a>",
"_____no_output_____"
]
],
[
[
"import numpy as np \nimport pandas as pd\nfrom sklearn import preprocessing\nfrom sklearn.preprocessing import StandardScaler \nfrom sklearn.metrics import mean_squared_error\nfrom sklearn import linear_model\nimport matplotlib.pyplot as plt\nimport os\nnp.random.seed(42)\nimport warnings\nwarnings.filterwarnings('ignore')",
"_____no_output_____"
]
],
[
[
"Loading the Dataset",
"_____no_output_____"
]
],
[
[
"from pandas import read_csv\nfilePath = '/content/bikes.csv'\nbikesData = pd.read_csv(filePath)\nprint(bikesData.info())",
"<class 'pandas.core.frame.DataFrame'>\nRangeIndex: 17379 entries, 0 to 17378\nData columns (total 17 columns):\n # Column Non-Null Count Dtype \n--- ------ -------------- ----- \n 0 instant 17379 non-null int64 \n 1 dteday 17379 non-null object \n 2 season 17379 non-null int64 \n 3 yr 17379 non-null int64 \n 4 mnth 17379 non-null int64 \n 5 hr 17379 non-null int64 \n 6 holiday 17379 non-null int64 \n 7 weekday 17379 non-null int64 \n 8 workingday 17379 non-null int64 \n 9 weathersit 17379 non-null int64 \n 10 temp 17379 non-null float64\n 11 atemp 17379 non-null float64\n 12 hum 17379 non-null float64\n 13 windspeed 17379 non-null float64\n 14 casual 17379 non-null int64 \n 15 registered 17379 non-null int64 \n 16 cnt 17379 non-null int64 \ndtypes: float64(4), int64(12), object(1)\nmemory usage: 2.3+ MB\nNone\n"
],
[
"bikesData.head(3)",
"_____no_output_____"
],
[
"columnsToDrop = ['instant','casual','registered','atemp','dteday']\n\nbikesData = bikesData.drop(columnsToDrop,1)",
"_____no_output_____"
],
[
"bikesData.head(3)",
"_____no_output_____"
],
[
"np.random.seed(42)\nfrom sklearn.model_selection import train_test_split\n\nbikesData['dayCount'] = pd.Series(range(bikesData.shape[0]))/24\n\ntrain_set, test_set = train_test_split(bikesData, test_size=0.3, random_state=42)\n\nprint(len(train_set), \"train +\", len(test_set), \"test\")\n\ntrain_set.sort_values('dayCount', axis= 0, inplace=True)\ntest_set.sort_values('dayCount', axis= 0, inplace=True)",
"12165 train + 5214 test\n"
],
[
"columnsToScale = ['temp','hum','windspeed']\n\nscaler = StandardScaler()\n\ntrain_set[columnsToScale] = scaler.fit_transform(train_set[columnsToScale])\ntest_set[columnsToScale] = scaler.transform(test_set[columnsToScale])\ntrain_set[columnsToScale].describe()\n",
"_____no_output_____"
],
[
"from sklearn.model_selection import cross_val_score\nfrom sklearn.model_selection import cross_val_predict\nfrom sklearn.tree import DecisionTreeRegressor\nfrom sklearn.linear_model import LinearRegression\nfrom sklearn.ensemble import RandomForestRegressor\nimport xgboost\nfrom xgboost import XGBRegressor\n\ntrainingCols = train_set.drop(['cnt'], axis=1)\ntrainingLabels = train_set['cnt']",
"_____no_output_____"
],
[
"#Train a Decision Tree Regressor\nfrom sklearn.tree import DecisionTreeRegressor \ndec_reg = DecisionTreeRegressor(random_state = 42)",
"_____no_output_____"
],
[
"\ndt_mae_scores = -cross_val_score(dec_reg, trainingCols, trainingLabels, cv=10, scoring=\"neg_mean_absolute_error\")\nprint(dt_mae_scores)\nprint('\\n')\ndt_mse_scores = np.sqrt(-cross_val_score(dec_reg, trainingCols, trainingLabels, cv=10, scoring=\"neg_mean_squared_error\"))\nprint(dt_mse_scores)",
"[42.94494659 50.37222679 36.95891537 44.26211997 46.99589154 71.98026316\n 58.19901316 48.87417763 50.84868421 96.46217105]\n\n\n[ 65.39786583 77.67402864 60.57274567 73.73250527 75.48574011\n 113.22922285 96.5884429 82.11639785 86.86752618 149.13680359]\n"
],
[
"lin_reg = LinearRegression()\n\nlr_mae_scores = -cross_val_score(lin_reg, trainingCols, trainingLabels, cv=10, scoring=\"neg_mean_absolute_error\")\n\nprint(lr_mae_scores)\nprint('\\n')\n\nlr_mse_scores = np.sqrt(-cross_val_score(lin_reg, trainingCols, trainingLabels, cv=10, scoring=\"neg_mean_squared_error\"))\n\nprint(lr_mse_scores)",
"[ 66.96340699 80.48809095 113.84704981 93.17230086 76.11197672\n 96.5220689 133.13798218 158.02254734 158.90195479 127.15674717]\n\n\n[ 84.63836676 111.12038541 131.88324414 119.16350622 105.17621319\n 127.72562924 174.97188817 187.31691741 205.60028279 164.30585678]\n"
],
[
"forest_reg = RandomForestRegressor(n_estimators=150, random_state=42)\nrf_mae_scores = -cross_val_score(forest_reg, trainingCols, trainingLabels, cv=10, scoring=\"neg_mean_absolute_error\")\n\nprint(rf_mae_scores)\n\nrf_mse_scores = np.sqrt(-cross_val_score(forest_reg, trainingCols, trainingLabels, cv=10, scoring=\"neg_mean_squared_error\"))\n\nprint(rf_mse_scores)",
"[33.39666393 33.54451931 28.50225692 31.78826623 36.55658724 57.81963268\n 40.96405702 40.84652961 37.57766447 84.69771382]\n[ 45.64176074 50.97205843 43.37588352 52.2640926 60.46557726\n 94.24478873 66.26045287 65.45672124 61.69916554 131.9727285 ]\n"
],
[
"from sklearn.model_selection import GridSearchCV\n\nparam_grid = [\n # combinations of hyperparameters\n {'n_estimators': [120, 150], 'max_features': [10, 12], 'max_depth': [15, 28]},\n]",
"_____no_output_____"
],
[
"grid_search = GridSearchCV(forest_reg, param_grid, cv=5, scoring='neg_mean_squared_error')",
"_____no_output_____"
],
[
"grid_search.fit(trainingCols, trainingLabels)\n\nprint(grid_search.best_estimator_)\nprint(grid_search.best_params_)",
"RandomForestRegressor(bootstrap=True, ccp_alpha=0.0, criterion='mse',\n max_depth=28, max_features=10, max_leaf_nodes=None,\n max_samples=None, min_impurity_decrease=0.0,\n min_impurity_split=None, min_samples_leaf=1,\n min_samples_split=2, min_weight_fraction_leaf=0.0,\n n_estimators=150, n_jobs=None, oob_score=False,\n random_state=42, verbose=0, warm_start=False)\n{'max_depth': 28, 'max_features': 10, 'n_estimators': 150}\n"
],
[
"feature_importances = grid_search.best_estimator_.feature_importances_\nprint(feature_importances)",
"[0.00424888 0.00145493 0.00570279 0.58348648 0.00215107 0.01790669\n 0.06993018 0.01688336 0.09373438 0.03176755 0.00907719 0.16365649]\n"
],
[
"final_model = grid_search.best_estimator_\ntest_set.sort_values('dayCount', axis= 0, inplace=True)\ntest_x_cols = (test_set.drop(['cnt'], axis=1)).columns.values\ntest_y_cols = 'cnt'\n\nX_test = test_set.loc[:,test_x_cols]\ny_test = test_set.loc[:,test_y_cols]",
"_____no_output_____"
],
[
"test_set.loc[:,'predictedCounts_test'] = final_model.predict(X_test)\n\nmse = mean_squared_error(y_test, test_set.loc[:,'predictedCounts_test'])\nfinal_mse = np.sqrt(mse)\nprint(final_mse)\ntest_set.describe()\ntimes = [9,18]\nfor time in times:\n fig = plt.figure(figsize=(8, 6))\n fig.clf()\n ax = fig.gca()\n test_set_freg_time = test_set[test_set.hr == time]\n test_set_freg_time.plot(kind = 'line', x = 'dayCount', y = 'cnt', ax = ax)\n test_set_freg_time.plot(kind = 'line', x = 'dayCount', y = 'predictedCounts_test', ax =ax)\n plt.show()",
"39.47930005837265\n"
]
]
] |
[
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acc5e36756ee666bd1da7a729e2f15f934d40e9
| 13,779 |
ipynb
|
Jupyter Notebook
|
stats/utils_et_biomarker/doc_str_example.ipynb
|
nikhil153/ET_biomarker
|
43a363f9f146c630ee9fb00539a1f8d60d8e4545
|
[
"MIT"
] | 1 |
2021-09-20T21:36:33.000Z
|
2021-09-20T21:36:33.000Z
|
stats/utils_et_biomarker/doc_str_example.ipynb
|
nikhil153/ET_biomarker
|
43a363f9f146c630ee9fb00539a1f8d60d8e4545
|
[
"MIT"
] | 4 |
2021-08-02T14:19:13.000Z
|
2022-03-22T01:27:36.000Z
|
utils_/doc_str_example.ipynb
|
Vincent-wq/ET_biomarker
|
4f7d7c499c7ddc0229955fb6432e07384b3e9711
|
[
"MIT"
] | 2 |
2021-07-21T20:38:54.000Z
|
2021-08-02T13:47:19.000Z
| 35.240409 | 90 | 0.510995 |
[
[
[
"# -*- coding: utf-8 -*-\n\"\"\"Example NumPy style docstrings.\n\nThis module demonstrates documentation as specified by the `NumPy\nDocumentation HOWTO`_. Docstrings may extend over multiple lines. Sections\nare created with a section header followed by an underline of equal length.\n\nExample\n-------\nExamples can be given using either the ``Example`` or ``Examples``\nsections. Sections support any reStructuredText formatting, including\nliteral blocks::\n\n $ python example_numpy.py\n\n\nSection breaks are created with two blank lines. Section breaks are also\nimplicitly created anytime a new section starts. Section bodies *may* be\nindented:\n\nNotes\n-----\n This is an example of an indented section. It's like any other section,\n but the body is indented to help it stand out from surrounding text.\n\nIf a section is indented, then a section break is created by\nresuming unindented text.\n\nAttributes\n----------\nmodule_level_variable1 : int\n Module level variables may be documented in either the ``Attributes``\n section of the module docstring, or in an inline docstring immediately\n following the variable.\n\n Either form is acceptable, but the two should not be mixed. Choose\n one convention to document module level variables and be consistent\n with it.\n\n\n.. _NumPy Documentation HOWTO:\n https://github.com/numpy/numpy/blob/master/doc/HOWTO_DOCUMENT.rst.txt\n\n\"\"\"\n\nmodule_level_variable1 = 12345\n\nmodule_level_variable2 = 98765\n\"\"\"int: Module level variable documented inline.\n\nThe docstring may span multiple lines. The type may optionally be specified\non the first line, separated by a colon.\n\"\"\"\n\n\ndef function_with_types_in_docstring(param1, param2):\n \"\"\"Example function with types documented in the docstring.\n\n `PEP 484`_ type annotations are supported. If attribute, parameter, and\n return types are annotated according to `PEP 484`_, they do not need to be\n included in the docstring:\n\n Parameters\n ----------\n param1 : int\n The first parameter.\n param2 : str\n The second parameter.\n\n Returns\n -------\n bool\n True if successful, False otherwise.\n\n .. _PEP 484:\n https://www.python.org/dev/peps/pep-0484/\n\n \"\"\"\n\n\ndef function_with_pep484_type_annotations(param1: int, param2: str) -> bool:\n \"\"\"Example function with PEP 484 type annotations.\n\n The return type must be duplicated in the docstring to comply\n with the NumPy docstring style.\n\n Parameters\n ----------\n param1\n The first parameter.\n param2\n The second parameter.\n\n Returns\n -------\n bool\n True if successful, False otherwise.\n\n \"\"\"\n\n\ndef module_level_function(param1, param2=None, *args, **kwargs):\n \"\"\"This is an example of a module level function.\n\n Function parameters should be documented in the ``Parameters`` section.\n The name of each parameter is required. The type and description of each\n parameter is optional, but should be included if not obvious.\n\n If \\*args or \\*\\*kwargs are accepted,\n they should be listed as ``*args`` and ``**kwargs``.\n\n The format for a parameter is::\n\n name : type\n description\n\n The description may span multiple lines. Following lines\n should be indented to match the first line of the description.\n The \": type\" is optional.\n\n Multiple paragraphs are supported in parameter\n descriptions.\n\n Parameters\n ----------\n param1 : int\n The first parameter.\n param2 : :obj:`str`, optional\n The second parameter.\n *args\n Variable length argument list.\n **kwargs\n Arbitrary keyword arguments.\n\n Returns\n -------\n bool\n True if successful, False otherwise.\n\n The return type is not optional. The ``Returns`` section may span\n multiple lines and paragraphs. Following lines should be indented to\n match the first line of the description.\n\n The ``Returns`` section supports any reStructuredText formatting,\n including literal blocks::\n\n {\n 'param1': param1,\n 'param2': param2\n }\n\n Raises\n ------\n AttributeError\n The ``Raises`` section is a list of all exceptions\n that are relevant to the interface.\n ValueError\n If `param2` is equal to `param1`.\n\n \"\"\"\n if param1 == param2:\n raise ValueError('param1 may not be equal to param2')\n return True\n\n\ndef example_generator(n):\n \"\"\"Generators have a ``Yields`` section instead of a ``Returns`` section.\n\n Parameters\n ----------\n n : int\n The upper limit of the range to generate, from 0 to `n` - 1.\n\n Yields\n ------\n int\n The next number in the range of 0 to `n` - 1.\n\n Examples\n --------\n Examples should be written in doctest format, and should illustrate how\n to use the function.\n\n >>> print([i for i in example_generator(4)])\n [0, 1, 2, 3]\n\n \"\"\"\n for i in range(n):\n yield i\n\n\nclass ExampleError(Exception):\n \"\"\"Exceptions are documented in the same way as classes.\n\n The __init__ method may be documented in either the class level\n docstring, or as a docstring on the __init__ method itself.\n\n Either form is acceptable, but the two should not be mixed. Choose one\n convention to document the __init__ method and be consistent with it.\n\n Note\n ----\n Do not include the `self` parameter in the ``Parameters`` section.\n\n Parameters\n ----------\n msg : str\n Human readable string describing the exception.\n code : :obj:`int`, optional\n Numeric error code.\n\n Attributes\n ----------\n msg : str\n Human readable string describing the exception.\n code : int\n Numeric error code.\n\n \"\"\"\n\n def __init__(self, msg, code):\n self.msg = msg\n self.code = code\n\n\nclass ExampleClass(object):\n \"\"\"The summary line for a class docstring should fit on one line.\n\n If the class has public attributes, they may be documented here\n in an ``Attributes`` section and follow the same formatting as a\n function's ``Args`` section. Alternatively, attributes may be documented\n inline with the attribute's declaration (see __init__ method below).\n\n Properties created with the ``@property`` decorator should be documented\n in the property's getter method.\n\n Attributes\n ----------\n attr1 : str\n Description of `attr1`.\n attr2 : :obj:`int`, optional\n Description of `attr2`.\n\n \"\"\"\n\n def __init__(self, param1, param2, param3):\n \"\"\"Example of docstring on the __init__ method.\n\n The __init__ method may be documented in either the class level\n docstring, or as a docstring on the __init__ method itself.\n\n Either form is acceptable, but the two should not be mixed. Choose one\n convention to document the __init__ method and be consistent with it.\n\n Note\n ----\n Do not include the `self` parameter in the ``Parameters`` section.\n\n Parameters\n ----------\n param1 : str\n Description of `param1`.\n param2 : :obj:`list` of :obj:`str`\n Description of `param2`. Multiple\n lines are supported.\n param3 : :obj:`int`, optional\n Description of `param3`.\n\n \"\"\"\n self.attr1 = param1\n self.attr2 = param2\n self.attr3 = param3 #: Doc comment *inline* with attribute\n\n #: list of str: Doc comment *before* attribute, with type specified\n self.attr4 = [\"attr4\"]\n\n self.attr5 = None\n \"\"\"str: Docstring *after* attribute, with type specified.\"\"\"\n\n @property\n def readonly_property(self):\n \"\"\"str: Properties should be documented in their getter method.\"\"\"\n return \"readonly_property\"\n\n @property\n def readwrite_property(self):\n \"\"\":obj:`list` of :obj:`str`: Properties with both a getter and setter\n should only be documented in their getter method.\n\n If the setter method contains notable behavior, it should be\n mentioned here.\n \"\"\"\n return [\"readwrite_property\"]\n\n @readwrite_property.setter\n def readwrite_property(self, value):\n value\n\n def example_method(self, param1, param2):\n \"\"\"Class methods are similar to regular functions.\n\n Note\n ----\n Do not include the `self` parameter in the ``Parameters`` section.\n\n Parameters\n ----------\n param1\n The first parameter.\n param2\n The second parameter.\n\n Returns\n -------\n bool\n True if successful, False otherwise.\n\n \"\"\"\n return True\n\n def __special__(self):\n \"\"\"By default special members with docstrings are not included.\n\n Special members are any methods or attributes that start with and\n end with a double underscore. Any special member with a docstring\n will be included in the output, if\n ``napoleon_include_special_with_doc`` is set to True.\n\n This behavior can be enabled by changing the following setting in\n Sphinx's conf.py::\n\n napoleon_include_special_with_doc = True\n\n \"\"\"\n pass\n\n def __special_without_docstring__(self):\n pass\n\n def _private(self):\n \"\"\"By default private members are not included.\n\n Private members are any methods or attributes that start with an\n underscore and are *not* special. By default they are not included\n in the output.\n\n This behavior can be changed such that private members *are* included\n by changing the following setting in Sphinx's conf.py::\n\n napoleon_include_private_with_doc = True\n\n \"\"\"\n pass\n\n def _private_without_docstring(self):\n pass",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code"
]
] |
4acc70a8a80f23de4d40e5bf986565fc6d988cce
| 56,361 |
ipynb
|
Jupyter Notebook
|
odm2api_sample_fromsqlite.ipynb
|
emiliom/stuff
|
634817b0845c22370f8adde2105d7b6c734eaa58
|
[
"CC0-1.0"
] | null | null | null |
odm2api_sample_fromsqlite.ipynb
|
emiliom/stuff
|
634817b0845c22370f8adde2105d7b6c734eaa58
|
[
"CC0-1.0"
] | 1 |
2021-10-19T18:33:08.000Z
|
2021-10-21T06:58:28.000Z
|
odm2api_sample_fromsqlite.ipynb
|
emiliom/stuff
|
634817b0845c22370f8adde2105d7b6c734eaa58
|
[
"CC0-1.0"
] | 5 |
2016-01-08T18:56:29.000Z
|
2018-05-11T18:43:18.000Z
| 68.399272 | 31,502 | 0.759727 |
[
[
[
"# odm2api demo with Little Bear SQLite sample DB\nLargely from https://github.com/ODM2/ODM2PythonAPI/blob/master/Examples/Sample.py \n- 4/25/2016. Started testing with the new `odm2` conda channel, based on the new `0.5.0-alpha` odm2api release. See my `odm2api_odm2channel` env. Ran into problems b/c the SQLite database needed to be updated to have a `SamplingFeature.FeatureGeometryWKT` field; so I added and populated it manually with `SQLite Manager`.\n- 2/7/2016. Tested successfully with `sfgeometry_em_1` branch, with my overhauls. Using `odm2api_dev` env.\n- 2/1 - 1/31. Errors with SamplingFeatures code, with latest odm2api from master (on env `odm2api_jan31test`). *The code also fails the same way with the `odm2api` env, but it does still run fine with the `odm2api_jan21` env! I'm investigating the differences between those two envs.*\n- 1/22-20,9/2016.\n\nEmilio Mayorga",
"_____no_output_____"
]
],
[
[
"%matplotlib inline\nimport matplotlib.pyplot as plt\nfrom matplotlib import dates",
"/home/mayorga/miniconda/envs/odm2api_odm2channel/lib/python2.7/site-packages/matplotlib/font_manager.py:273: UserWarning: Matplotlib is building the font cache using fc-list. This may take a moment.\n warnings.warn('Matplotlib is building the font cache using fc-list. This may take a moment.')\n"
],
[
"from odm2api.ODMconnection import dbconnection\nfrom odm2api.ODM2.services.readService import ReadODM2",
"_____no_output_____"
],
[
"# Create a connection to the ODM2 database\n# ----------------------------------------\nodm2db_fpth = '/home/mayorga/Desktop/TylerYeats/ODM2-LittleBear1.sqlite'\nsession_factory = dbconnection.createConnection('sqlite', odm2db_fpth, 2.0)\nread = ReadODM2(session_factory)",
"_____no_output_____"
],
[
"# Run some basic sample queries.\n# ------------------------------\n# Get all of the variables from the database and print their names to the console\nallVars = read.getVariables()\n\nfor x in allVars:\n print x.VariableCode + \": \" + x.VariableNameCV",
"USU36: Temperature\n"
],
[
"# Get all of the people from the database\nallPeople = read.getPeople()\n\nfor x in allPeople:\n print x.PersonFirstName + \" \" + x.PersonLastName\n\ntry:\n print \"\\n-------- Information about an Affiliation ---------\"\n allaff = read.getAffiliations()\n for x in allaff:\n print x.PersonObj.PersonFirstName + \": \" + str(x.OrganizationID)\nexcept Exception as e:\n print \"Unable to demo getAllAffiliations\", e",
"Jeff Horsburgh\n\n-------- Information about an Affiliation ---------\nUnable to demo getAllAffiliations 'NoneType' object is not iterable\n"
],
[
"allaff = read.getAffiliations()\ntype(allaff)",
"_____no_output_____"
]
],
[
[
"## SamplingFeatures tests",
"_____no_output_____"
]
],
[
[
"# from odm2api.ODM2.models import SamplingFeatures\n# read._session.query(SamplingFeatures).filter_by(SamplingFeatureTypeCV='Site').all()",
"_____no_output_____"
],
[
"# Get all of the SamplingFeatures from the database that are Sites\ntry:\n siteFeatures = read.getSamplingFeatures(type='Site')\n numSites = len(siteFeatures)\n\n for x in siteFeatures:\n print x.SamplingFeatureCode + \": \" + x.SamplingFeatureName\nexcept Exception as e:\n print \"Unable to demo getSamplingFeatures(type='Site')\", e",
"USU-LBR-Mendon: Little Bear River at Mendon Road near Mendon, Utah\n"
],
[
"read.getSamplingFeatures()",
"_____no_output_____"
],
[
"read.getSamplingFeatures(codes=['USU-LBR-Mendon'])",
"_____no_output_____"
],
[
"# Now get the SamplingFeature object for a SamplingFeature code\nsf_lst = read.getSamplingFeatures(codes=['USU-LBR-Mendon'])\nvars(sf_lst[0])",
"_____no_output_____"
],
[
"sf = sf_lst[0]",
"_____no_output_____"
],
[
"print sf, \"\\n\"\nprint type(sf)\nprint type(sf.FeatureGeometryWKT), sf.FeatureGeometryWKT\nprint type(sf.FeatureGeometry)",
"<Sites('1', '1', 'Stream', '41.718473', '-111.946402', '<SpatialReferences('1', 'None', 'Unknown', 'The spatial reference is unknown', 'None')>', 'USU-LBR-Mendon')> \n\n<class 'odm2api.ODM2.models.Sites'>\n<type 'unicode'> POINT (41.718473 -111.946402)\n<class 'geoalchemy.spatialite.SQLitePersistentSpatialElement'>\n"
],
[
"vars(sf.FeatureGeometry)",
"_____no_output_____"
],
[
"sf.FeatureGeometry.__doc__",
"_____no_output_____"
],
[
"sf.FeatureGeometry.geom_wkb, sf.FeatureGeometry.geom_wkt",
"_____no_output_____"
],
[
"# 4/25/2016: Don't know why the shape is listed 4 times ...\ntype(sf.shape()), sf.shape().wkt",
"POINT (41.718473 -111.946402)\nPOINT (41.718473 -111.946402)\nPOINT (41.718473 -111.946402)\nPOINT (41.718473 -111.946402)\n"
]
],
[
[
"## Back to the rest of the demo",
"_____no_output_____"
]
],
[
[
"read.getResults()",
"_____no_output_____"
],
[
"firstResult = read.getResults()[0]\nfirstResult.FeatureActionObj.ActionObj",
"_____no_output_____"
]
],
[
[
"### Foreign Key Example\nDrill down and get objects linked by foreign keys",
"_____no_output_____"
]
],
[
[
"try:\n # Call getResults, but return only the first result\n firstResult = read.getResults()[0]\n action_firstResult = firstResult.FeatureActionObj.ActionObj\n print \"The FeatureAction object for the Result is: \", firstResult.FeatureActionObj\n print \"The Action object for the Result is: \", action_firstResult\n print (\"\\nThe following are some of the attributes for the Action that created the Result: \\n\" +\n \"ActionTypeCV: \" + action_firstResult.ActionTypeCV + \"\\n\" +\n \"ActionDescription: \" + action_firstResult.ActionDescription + \"\\n\" +\n \"BeginDateTime: \" + str(action_firstResult.BeginDateTime) + \"\\n\" +\n \"EndDateTime: \" + str(action_firstResult.EndDateTime) + \"\\n\" +\n \"MethodName: \" + action_firstResult.MethodObj.MethodName + \"\\n\" +\n \"MethodDescription: \" + action_firstResult.MethodObj.MethodDescription)\nexcept Exception as e:\n print \"Unable to demo Foreign Key Example: \", e",
"The FeatureAction object for the Result is: <FeatureActions('1', '1', '1', )>\nThe Action object for the Result is: <Actions('1', 'Observation', '2007-08-16 16:30:00', 'An observation action that generated a time series result.')>\n\nThe following are some of the attributes for the Action that created the Result: \nActionTypeCV: Observation\nActionDescription: An observation action that generated a time series result.\nBeginDateTime: 2007-08-16 16:30:00\nEndDateTime: 2009-01-16 12:30:00\nMethodName: Quality Control Level 1 Data Series created from raw QC Level 0 data using ODM Tools.\nMethodDescription: Quality Control Level 1 Data Series created from raw QC Level 0 data using ODM Tools.\n"
]
],
[
[
"### Example of Retrieving Attributes of a Time Series Result using a ResultID",
"_____no_output_____"
]
],
[
[
"tsResult = read.getResults(ids=[1])[0]\ntype(tsResult), vars(tsResult)",
"_____no_output_____"
]
],
[
[
"**Why are `ProcessingLevelObj`, `VariableObj` and `UnitsObj` objects not shown in the above `vars()` listing!?** They **are** actually available, as demonstrated in much of the code below.",
"_____no_output_____"
]
],
[
[
"try:\n tsResult = read.getResults(ids=[1])[0]\n # Get the site information by drilling down\n sf_tsResult = tsResult.FeatureActionObj.SamplingFeatureObj\n print(\n \"Some of the attributes for the TimeSeriesResult retrieved using getResults(ids=[]): \\n\" +\n \"ResultTypeCV: \" + tsResult.ResultTypeCV + \"\\n\" +\n # Get the ProcessingLevel from the TimeSeriesResult's ProcessingLevel object\n \"ProcessingLevel: \" + tsResult.ProcessingLevelObj.Definition + \"\\n\" +\n \"SampledMedium: \" + tsResult.SampledMediumCV + \"\\n\" +\n # Get the variable information from the TimeSeriesResult's Variable object\n \"Variable: \" + tsResult.VariableObj.VariableCode + \": \" + tsResult.VariableObj.VariableNameCV + \"\\n\" +\n \"AggregationStatistic: \" + tsResult.AggregationStatisticCV + \"\\n\" +\n # Get the site information by drilling down\n \"Elevation_m: \" + str(sf_tsResult.Elevation_m) + \"\\n\" +\n \"SamplingFeature: \" + sf_tsResult.SamplingFeatureCode + \" - \" +\n sf_tsResult.SamplingFeatureName)\nexcept Exception as e:\n print \"Unable to demo Example of retrieving Attributes of a time Series Result: \", e",
"Some of the attributes for the TimeSeriesResult retrieved using getResults(ids=[]): \nResultTypeCV: Time series coverage\nProcessingLevel: Quality controlled data\nSampledMedium: Surface Water\nVariable: USU36: Temperature\nAggregationStatistic: Average\nElevation_m: 1345.0\nSamplingFeature: USU-LBR-Mendon - Little Bear River at Mendon Road near Mendon, Utah\n"
]
],
[
[
"### Example of Retrieving Time Series Result Values, then plotting them",
"_____no_output_____"
]
],
[
[
"# Get the values for a particular TimeSeriesResult\n\ntsValues = read.getResultValues(resultid=1) # Return type is a pandas dataframe\n# Print a few Time Series Values to the console\n# tsValues.set_index('ValueDateTime', inplace=True)\ntsValues.head()",
"_____no_output_____"
],
[
"# Plot the time series\ntry:\n fig = plt.figure()\n ax = fig.add_subplot(111)\n tsValues.plot(x='ValueDateTime', y='DataValue', kind='line',\n title=tsResult.VariableObj.VariableNameCV + \" at \" + \n tsResult.FeatureActionObj.SamplingFeatureObj.SamplingFeatureName,\n ax=ax)\n \n ax.set_ylabel(tsResult.VariableObj.VariableNameCV + \" (\" + \n tsResult.UnitsObj.UnitsAbbreviation + \")\")\n ax.set_xlabel(\"Date/Time\")\n ax.xaxis.set_minor_locator(dates.MonthLocator())\n ax.xaxis.set_minor_formatter(dates.DateFormatter('%b'))\n ax.xaxis.set_major_locator(dates.YearLocator())\n ax.xaxis.set_major_formatter(dates.DateFormatter('\\n%Y'))\n ax.grid(True)\nexcept Exception as e:\n print \"Unable to demo plotting of tsValues: \", e",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4acc7afb883f755c9b6da8a49be24b08d2258bf6
| 613,282 |
ipynb
|
Jupyter Notebook
|
archive/2020-04-Publishers-Usecases/5-Competing-Journals.ipynb
|
eschares/dimensions-api-lab
|
cfe6bc72631c08aabb392044abef020180dffe90
|
[
"MIT"
] | 57 |
2019-06-24T19:35:34.000Z
|
2022-02-27T14:45:10.000Z
|
archive/2020-04-Publishers-Usecases/5-Competing-Journals.ipynb
|
eschares/dimensions-api-lab
|
cfe6bc72631c08aabb392044abef020180dffe90
|
[
"MIT"
] | 6 |
2019-09-04T19:14:40.000Z
|
2021-12-09T15:54:41.000Z
|
archive/2020-04-Publishers-Usecases/5-Competing-Journals.ipynb
|
eschares/dimensions-api-lab
|
cfe6bc72631c08aabb392044abef020180dffe90
|
[
"MIT"
] | 16 |
2019-08-13T04:24:01.000Z
|
2022-03-04T07:49:11.000Z
| 105.939195 | 246,650 | 0.796942 |
[
[
[
"# Part 5: Competing Journals Analysis\n\nIn this notebook we are going to \n\n* Load the researchers impact metrics data previously extracted (see parts 1-2-3)\n* Get the full publications history for these researchers \n* Use this new publications dataset to determine which are the most frequent journals the researchers have also published in\n* Build some visualizations in order to have a quick overview of the results ",
"_____no_output_____"
],
[
"## Prerequisites: Installing the Dimensions Library and Logging in",
"_____no_output_____"
]
],
[
[
"\n# @markdown # Get the API library and login\n# @markdown Click the 'play' button on the left (or shift+enter) after entering your API credentials\n\nusername = \"\" #@param {type: \"string\"}\npassword = \"\" #@param {type: \"string\"}\nendpoint = \"https://app.dimensions.ai\" #@param {type: \"string\"}\n\n\n!pip install dimcli plotly tqdm -U --quiet\nimport dimcli\nfrom dimcli.shortcuts import *\ndimcli.login(username, password, endpoint)\ndsl = dimcli.Dsl()\n\n#\n# load common libraries\nimport time\nimport sys\nimport os\nimport json\nimport pandas as pd\nfrom pandas.io.json import json_normalize\nfrom tqdm.notebook import tqdm as progress\n\n#\n# charts libs\n# import plotly_express as px\nimport plotly.express as px\nif not 'google.colab' in sys.modules:\n # make js dependecies local / needed by html exports\n from plotly.offline import init_notebook_mode\n init_notebook_mode(connected=True)\n#\n# create output data folder\nif not(os.path.exists(\"data\")):\n os.mkdir(\"data\")",
"DimCli v0.6.7 - Succesfully connected to <https://app.dimensions.ai> (method: dsl.ini file)\n"
]
],
[
[
"## Competing Journals",
"_____no_output_____"
],
[
"From our researchers master list, we now want to extract the following:\n\n* full list of publications for a 5 year period\n* full list of journals with counts of how many publications per journal \n\nThis new dataset will let us draw up some conclusions re. which are the competing journals of the one we selected at the beginning.\n\n",
"_____no_output_____"
],
[
"### First let's reload the data obtained in previous steps",
"_____no_output_____"
]
],
[
[
"#\nresearchers = pd.read_csv(\"data/2.researchers_impact_metrics.csv\")\n#\nprint(\"Total researchers:\", len(researchers))\nresearchers.head(5)",
"Total researchers: 18341\n"
]
],
[
[
"### What the query looks like",
"_____no_output_____"
],
[
"The approach we're taking consists in pulling all publications data, so that we can count journals as a second step. \n\nThis approach may take some time (as we're potentially retrieving a lot of publications data), but it will lead to precise results. \n\nThe query template to use looks like this (for a couple of researchers only):",
"_____no_output_____"
]
],
[
[
"%%dsldf \nsearch publications where researchers.id in [\"ur.01277776417.51\", \"ur.0637651205.48\"] \n and year >= 2015 and journal is not empty \n and journal.id != \"jour.1103138\" \nreturn publications[id+journal] limit 10",
"Returned Publications: 10 (total = 149)\n"
]
],
[
[
"## Extracting all publications/journals information\n\nThis part may take some time to run (depending on how many years back one wants to go) so you may want to get a coffee while you wait.. \n",
"_____no_output_____"
]
],
[
[
"#\njournal_id = \"jour.1103138\" # Nature genetics\nstart_year = 2018\n\n# our list of researchers\nllist = list(researchers['researcher_id'])\n#\n# the query\nq2 = \"\"\"search publications \n where researchers.id in {} \n and year >= {} and journal is not empty and journal.id != \"{}\" \n return publications[id+journal+year]\"\"\"",
"_____no_output_____"
],
[
"\nVERBOSE = False\nRESEARCHER_ITERATOR_NO = 400\n\npubs = pd.DataFrame\nfor chunk in progress(list(chunks_of(llist, RESEARCHER_ITERATOR_NO))):\n # get all pubs\n query = q2.format(json.dumps(chunk), start_year, journal_id)\n res = dsl.query_iterative(query, verbose=VERBOSE)\n if pubs.empty:\n # first time, init the dataframe \n pubs = res.as_dataframe()\n else:\n pubs.append(res.as_dataframe())",
"_____no_output_____"
],
[
"# remove duplicate publications, if they have the same PUB_ID\npubs = pubs.drop_duplicates(subset=\"id\")\n# save\npubs.to_csv(\"data/5.journals-via-publications-RAW.csv\", index=False)\n# preview the data\npubs",
"_____no_output_____"
]
],
[
[
"Now we can create a journals-only dataset that includes counts per year, and grant total. ",
"_____no_output_____"
]
],
[
[
"journals = pubs.copy()\n# drop pub_id column\njournals = journals.drop(['id'], axis=1)\n# \n# add total column \njournals['total'] = journals.groupby('journal.id')['journal.id'].transform('count')\njournals['total_year'] = journals.groupby(['journal.id', 'year'])['journal.id'].transform('count')\n# \n# remove multiple counts for same journal\njournals = journals.drop_duplicates() \njournals.reset_index(drop=True)\n#\n# sort by total count\njournals = journals.sort_values('total', ascending=False)\n# #\n# # save\njournals.to_csv(\"data/5.journals-via-publications.csv\", index=False)\nprint(\"======\\nDone\")\n\n# download the data\nif COLAB_ENV:\n files.download(\"data/5.journals-via-publications.csv\")\n#preview the data \njournals.head(10)",
"======\nDone\n"
]
],
[
[
"# Visualizations",
"_____no_output_____"
]
],
[
[
"\nthreshold = 100\ntemp = journals.sort_values(\"total\", ascending=False)[:threshold]\n\npx.bar(journals[:threshold], \n x=\"journal.title\", y=\"total_year\", \n color=\"year\",\n hover_name=\"journal.title\", \n hover_data=['journal.id', 'journal.title', 'total' ], \n title=f\"Top {threshold} competitors for {journal_id} (based on publications data from {start_year})\")",
"_____no_output_____"
],
[
"threshold = 200\ntemp = journals.sort_values(\"year\", ascending=True).groupby(\"year\").head(threshold)\n\npx.bar(journals[:threshold], \n x=\"journal.title\", y=\"total_year\", \n color=\"year\",\n facet_row=\"year\",\n height=900,\n hover_name=\"journal.title\", \n hover_data=['journal.id', 'journal.title', 'total' ], \n title=f\"Top {threshold} competitors for {journal_id} - segmented by year\")",
"_____no_output_____"
]
],
[
[
"NOTE the European Neuropsychopharmacology journal has a massive jump in 2019 cause they [published a lot of conference proceedings](https://www.sciencedirect.com/journal/european-neuropsychopharmacology/issues)! See also the journal [Dimensions page](https://app.dimensions.ai/analytics/publication/overview/timeline?and_facet_source_title=jour.1101548) for comparison.. \n",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
]
] |
4acc7c07861b1b3adbb32568a9f48df1c4fb7f07
| 79,271 |
ipynb
|
Jupyter Notebook
|
Optimizer/optim.ipynb
|
KonstantinosRekoumis/CFD_CNN_Thesis
|
086c378df21ddac9e7902cac05daf514121c76a2
|
[
"MIT"
] | null | null | null |
Optimizer/optim.ipynb
|
KonstantinosRekoumis/CFD_CNN_Thesis
|
086c378df21ddac9e7902cac05daf514121c76a2
|
[
"MIT"
] | null | null | null |
Optimizer/optim.ipynb
|
KonstantinosRekoumis/CFD_CNN_Thesis
|
086c378df21ddac9e7902cac05daf514121c76a2
|
[
"MIT"
] | null | null | null | 160.467611 | 31,990 | 0.890124 |
[
[
[
"# Optimizer Notebook\n\n \n",
"_____no_output_____"
],
[
"## Julia needs to compile once 🤷 ",
"_____no_output_____"
]
],
[
[
"#Force Notebook to work on the parent Directory\nimport os \nif (\"Optimizer\" in os.getcwd()):\n os.chdir(\"..\")",
"_____no_output_____"
],
[
"from julia.api import Julia\njl = Julia(compiled_modules=False)\nfrom julia import Main\nMain.include(\"./Optimizer/eval_NN.jl\")\nNN_path = \"/home/freshstart/DiplomaThesisData/NeuralNetSaves_050/\"\n",
"_____no_output_____"
],
[
"#----- TEST ------\nRESup,RESdwn = Main.NN_eval(NN_path,\"./Optimizer/RAE_var.png\")\n",
"_____no_output_____"
]
],
[
[
"## Geometry and Image Processing\n",
"_____no_output_____"
]
],
[
[
"import Airfoil_Generation.Airfoil_Range_Creator as arg\nimport Airfoil_Generation.Images_Generator as ig\nimport numpy as np\nfrom scipy.integrate import simps\nimport matplotlib.pyplot as plt",
"_____no_output_____"
],
[
"#--- Read Geometry ---\ngeom = arg.readfile(\"./Misc/data/RAE_2822.geom\")\ngeom = (geom[0],geom[1][1:-1,:])\nplt.plot(geom[1][:,0],geom[1][:,1])\nplt.plot(geom[0][:,0],geom[0][:,1])",
"_____no_output_____"
]
],
[
[
"### Normals Calculation",
"_____no_output_____"
]
],
[
[
"def normals2D(geom,flip_n = False):\n eta = np.ndarray((len(geom)-1,2))\n\n for i in range(len(geom)-1):\n xba = geom[i+1,0]-geom[i,0]\n yba = geom[i+1,1]-geom[i,1]\n if flip_n:\n yba = - yba\n xba = - xba\n nrm2 = np.sqrt(yba**2+xba**2)\n eta[i,0] = yba/nrm2\n eta[i,1] = -xba/nrm2\n\n return eta\n \n\ndef partials2D(Cp,geom,flip_norm = True,show_norms = False):\n eta = normals2D(geom,flip_n=flip_norm)\n\n if show_norms:\n l = len(eta)\n fig, ax = plt.subplots()\n ax.plot(geom[0:-2,0],geom[0:-2,1])\n ax.quiver(geom[0:-2,0],geom[0:-2,1],eta[:,0],eta[:,1])\n\n xClCd = np.ndarray((len(Cp),3))\n\n for i in range(len(Cp)):\n for j in range(len(eta)):\n if ((Cp[i,0]<= geom[j+1,0])&(Cp[i,0]>geom[j,0])):\n xClCd[i,0] = Cp[i,0]\n xClCd[i,1] = eta[j,1]*Cp[i,1]\n xClCd[i,2] = eta[j,0]*Cp[i,1]\n break\n \n return xClCd\n\ndef calc2D(Cpup,Cpdwn,geom,show_norms = False):\n Up = partials2D(Cpup,geom[0],flip_norm = False,show_norms=show_norms)\n Dn = partials2D(Cpdwn,geom[1],flip_norm = True,show_norms=show_norms)\n Cl = -simps(Up[:,0],Up[:,1])+simps(Dn[:,0],Dn[:,1]) #invert y-axis to match the global axis\n Cd = simps(Up[:,0],Up[:,2])+simps(Dn[:,0],Dn[:,2])\n return Cl,Cd\n\nCl,Cd = calc2D(RESup,RESdwn,geom)\nprint(\"Cl = \", Cl)\nprint(\"Cd = \", Cd)\n",
"Cl = 1.017506174758391\nCd = -0.022827923892619905\n"
],
[
"phi = np.linspace(0,2*3.14159)\nx = np.array(np.cos(phi),ndmin = 2).reshape((len(phi),1))\ny = np.array(np.sin(phi),ndmin = 2).reshape((len(phi),1))\nplt.plot(x,y)\nplt.axis(\"equal\")\n\nf = np.concatenate((x,y),axis = 1)\nplt.plot(f[:,0],f[:,1])\n\na=partials2D(RESup,f,show_norms=True)",
"_____no_output_____"
]
],
[
[
"## Optimizer ",
"_____no_output_____"
]
],
[
[
"import openmdao.api as om",
"/opt/intel/intelpython3/lib/python3.7/site-packages/openmdao/utils/general_utils.py:128: OMDeprecationWarning:simple_warning is deprecated. Use openmdao.utils.om_warnings.issue_warning instead.\n/opt/intel/intelpython3/lib/python3.7/site-packages/openmdao/utils/notebook_utils.py:157: UserWarning:Tabulate is not installed. Run `pip install openmdao[notebooks]` to install required dependencies. Using ASCII for outputs.\n"
]
],
[
[
"### Class Definition and problem set-up",
"_____no_output_____"
]
],
[
[
"#---- Preparing the X coordinates for use in the optimizer -------\nX_UP = np.array(geom[0][:,0],ndmin=2)\nX_DN = np.array(geom[1][:,0],ndmin=2)\nX_UP = X_UP.reshape((X_UP.shape[1],X_UP.shape[0]))\nX_DN = X_DN.reshape((X_DN.shape[1],X_DN.shape[0]))\nY_UP = np.array(geom[0][:,1],ndmin=2)\nY_DN = np.array(geom[1][:,1],ndmin=2)\nY_UP = Y_UP.reshape((Y_UP.shape[1],Y_UP.shape[0]))\nY_DN = Y_DN.reshape((Y_DN.shape[1],Y_DN.shape[0]))\n##################################################################\nclass Airfoil(om.ExplicitComponent):\n \"\"\"\n Creates the most efficient airfoil for specific Mach and Reynolds numbers\n Changing each y-coords the deformation rate is more efficient to confine\n than flat y-coordinates\n \"\"\"\n def setup(self):\n self.add_input(\"y_up_rate\", val = 0.0)#np.zeros((len(geom[0]),1)) )\n self.add_input(\"y_dwn_rate\", val = 0.0)#np.zeros((len(geom[1]),1)) )\n\n self.add_output(\"Cl\", val = 0.0)\n self.add_output(\"Cd\", val = 0.0)\n \n\n def setup_partials(self):\n self.declare_partials(\"*\",\"*\", method = \"fd\")\n\n def compute(self, inputs, outputs):\n\n r1 = inputs[\"y_up_rate\"]\n r2 = inputs[\"y_dwn_rate\"]\n \n y1 = (1+r1)*Y_UP\n y2 = (1+r2)*Y_DN\n \n temp_geom = (np.concatenate((X_UP,y1),axis = 1),np.concatenate((X_DN,y2),axis = 1))\n ig.image_generator(np.concatenate((temp_geom[0],temp_geom[1]),axis = 0),\"./Optimizer/temp.png\",32,32)\n Cpup,Cpdwn = Main.NN_eval(NN_path,\"./Optimizer/temp.png\")\n res = calc2D(Cpup,Cpdwn,temp_geom)\n \n outputs[\"Cl\"] = res[0]\n outputs[\"Cd\"] = res[1]\n",
"_____no_output_____"
],
[
"#--------- Testing --------\nmodel = om.Group()\nmodel.add_subsystem(\"airfoil\",Airfoil(),promotes_inputs=[\"y_up_rate\",\"y_dwn_rate\"])\n\nprob = om.Problem(model)\n\nprob.setup()\nprob.run_model()\nprint(prob.get_val(\"airfoil.Cl\"))\nprob.get_val(\"airfoil.Cd\")",
"[1.01750617]\n"
]
],
[
[
"### Optimization \n",
"_____no_output_____"
]
],
[
[
"model = om.Group()\nmodel.add_subsystem(\"airfoil\",Airfoil(),promotes_inputs=[\"y_up_rate\",\"y_dwn_rate\"])\n\nprob = om.Problem(model)\n\nprob.driver = om.ScipyOptimizeDriver()\nprob.driver.options[\"optimizer\"] = \"COBYLA\"\n# prob.driver.options[\"optimizer\"] = \"SLSQP\"\n\nL_BOUND = -0.2\nU_BOUND = 0.2\n\nprob.model.add_design_var(\"y_up_rate\",lower = L_BOUND,upper= U_BOUND)\nprob.model.add_design_var(\"y_dwn_rate\",lower = L_BOUND,upper= U_BOUND)\n\nprob.model.add_objective(\"airfoil.Cl\",scaler=-1)\n# prob.model.add_objective(\"airfoil.Cd\",scaler=1)\n\nprob.setup()\nprob.run_driver();",
"Optimization Complete\n Normal return from subroutine COBYLA\n\n-----------------------------------\n\n NFVALS = 43 F =-1.527329E+00 MAXCV = 0.000000E+00\n X = 3.273863E-05 -1.046356E-04\n"
],
[
"#---------- SLSQP Optimizer ------------\nprint(\"Cl = \", prob.get_val(\"airfoil.Cl\"))\nprint(\"Cd = \", prob.get_val(\"airfoil.Cd\"))\nprint(\"Rate up = \", prob.get_val(\"y_up_rate\"))\nprint(\"Rate dwn = \", prob.get_val(\"y_dwn_rate\"))",
"Cl = [1.01752287]\nCd = [-0.02282877]\nRate up = [3.24957416e-05]\nRate dwn = [2.45606069e-08]\n"
],
[
"#----- Maximize Cl COBYLA -----\nprint(\"Cl = \", prob.get_val(\"airfoil.Cl\"))\nprint(\"Cd = \", prob.get_val(\"airfoil.Cd\"))\nprint(\"Rate up = \", prob.get_val(\"y_up_rate\"))\nprint(\"Rate dwn = \", prob.get_val(\"y_dwn_rate\"))",
"Cl = [1.52732907]\nCd = [-0.02276767]\nRate up = [3.27386321e-05]\nRate dwn = [-0.00010464]\n"
],
[
"Cpup,Cpdwn = Main.NN_eval(NN_path,\"./Optimizer/temp.png\")\nCl,Cd = calc2D(Cpup,Cpdwn,geom)\nprint(\"Cl = \", Cl)\nprint(\"Cd = \", Cd)\n",
"Cl = 1.5263322155794605\nCd = -0.022767374719605687\n"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
4acc804c2b2a0148dd55073b1805de04505f178c
| 18,279 |
ipynb
|
Jupyter Notebook
|
notebooks/03-sequential-logic.ipynb
|
ujamjar/hardcaml
|
1a834d25523417a768ebfa45cca4071a48481479
|
[
"0BSD"
] | 123 |
2015-01-01T21:34:09.000Z
|
2021-11-23T10:14:56.000Z
|
notebooks/03-sequential-logic.ipynb
|
ujamjar/hardcaml
|
1a834d25523417a768ebfa45cca4071a48481479
|
[
"0BSD"
] | 26 |
2015-03-31T13:38:18.000Z
|
2017-10-18T00:10:51.000Z
|
notebooks/03-sequential-logic.ipynb
|
ujamjar/hardcaml
|
1a834d25523417a768ebfa45cca4071a48481479
|
[
"0BSD"
] | 10 |
2015-11-08T22:07:40.000Z
|
2018-08-07T14:25:22.000Z
| 18,279 | 18,279 | 0.656655 |
[
[
[
"empty"
]
]
] |
[
"empty"
] |
[
[
"empty"
]
] |
4acc932fe9e47297ef08ca65aaaf4fc760bdcd43
| 212,663 |
ipynb
|
Jupyter Notebook
|
comp-cientifica-II-2019-2/Aulas/Heat Equation Parabollic.ipynb
|
mirandagil/university-courses
|
e70ce5262555e84cffb13e53e139e7eec21e8907
|
[
"MIT"
] | 1 |
2019-12-23T16:39:01.000Z
|
2019-12-23T16:39:01.000Z
|
comp-cientifica-II-2019-2/Aulas/Heat Equation Parabollic.ipynb
|
mirandagil/university-courses
|
e70ce5262555e84cffb13e53e139e7eec21e8907
|
[
"MIT"
] | null | null | null |
comp-cientifica-II-2019-2/Aulas/Heat Equation Parabollic.ipynb
|
mirandagil/university-courses
|
e70ce5262555e84cffb13e53e139e7eec21e8907
|
[
"MIT"
] | null | null | null | 214.378024 | 148,272 | 0.873735 |
[
[
[
"## Parabolic Equation\n\nHeat equation\n\n$$\n\\frac{\\partial T}{\\partial t} = k \\nabla^2 T\n$$",
"_____no_output_____"
]
],
[
[
"import numpy as np\nimport matplotlib.pyplot as plt\nfrom matplotlib import animation, rc\nfrom IPython.display import HTML\n\nimport imageio",
"_____no_output_____"
],
[
"##### Vectorized forward euler\n### Input: F -> Differential equation;\n### y0 -> list or scalar for initial condition;\n### ts -> list of points in time to evaluate the equation;\n### p -> list or scalar for parameters for F, default is set to 0 if F has no extra parameters;\n### Output: ys -> numpy array with all solutions for each step t, ys is a Matrix\n##### Gil Miranda - last revision 15/09/2019\ndef f_euler(F, ts, y0, p = 0):\n ys = [y0]\n h = ts[1]-ts[0]\n for tnext in ts[1:]:\n ynext = ys[-1] + F(tnext, ys[-1], p)*h\n ys.append(ynext)\n t = tnext\n return np.array(ys)",
"_____no_output_____"
],
[
"def alpha(x):\n return 0\n\ndef beta(x):\n return 0\n\ndef m_Tnew(n, i, Ts):\n if i-1 == 0:\n newT = Ts[i] + k * dt/h**2 * (Ts[i+1] - 2*Ts[i] + alpha(i))\n elif i+1 == N+1:\n newT = Ts[i] + k * dt/h**2 * (beta(i) - 2*Ts[i] + Ts[i-1])\n else:\n newT = Ts[i] + k * dt/h**2 * (Ts[i+1] - 2*Ts[i] + Ts[i-1])\n return newT\n\nN = 5\nh = 1/(N+1)\ndt = 0.1*h**2\ntf = 150\nts = np.linspace(0,tf,N+2) # grid de tempo\nxs = np.arange(0,1.1,h)\nTs = np.zeros((tf,N+2)) # Matriz de resultados\nk = 1\nfor i in range(1,N+1):\n Ts[0][i] = np.sin(np.pi*h*i) \n\nfor n in range(0,tf-1):\n for i in range(1,N+1):\n Ts[n+1][i] = m_Tnew(n, i, Ts[n])\n\n",
"_____no_output_____"
],
[
"for k in range(0,tf):\n plt.plot(ts, Ts[k])",
"_____no_output_____"
],
[
"%matplotlib notebook\nfig, ax = plt.subplots()\nax.set_xlim((0,150))\nax.set_ylim((0,1))\nline, = ax.plot([],[], lw = 2)\ndef init():\n line.set_data([], [])\n return (line,)\n\n\ndef animate(i):\n x = ts\n y = Ts[i]\n line.set_data(x, y)\n if i > 70:\n line.set_color('blue')\n if i < 70 and i > 40:\n line.set_color('orange')\n if i < 40:\n line.set_color('red')\n return (line,)\n\nanim = animation.FuncAnimation(fig, animate, init_func=init,frames=120, interval=20, blit=True)\nanim.save('heat.gif', writer = 'pillow')\n# HTML(anim.to_html5_video())\nplt.grid(alpha = 0.5)\nplt.title('Cooling Metal Bar')\nplt.show()",
"_____no_output_____"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
]
] |
4acc9d8b4b9a3004bffe3cfd8d5f2ac1e8cdf16b
| 922,689 |
ipynb
|
Jupyter Notebook
|
notes/Test code HAL potential and effective masses.ipynb
|
tkmiriya/hal_pot_code
|
3446378916df74eca887cb36c380ed0ab151192e
|
[
"MIT"
] | null | null | null |
notes/Test code HAL potential and effective masses.ipynb
|
tkmiriya/hal_pot_code
|
3446378916df74eca887cb36c380ed0ab151192e
|
[
"MIT"
] | null | null | null |
notes/Test code HAL potential and effective masses.ipynb
|
tkmiriya/hal_pot_code
|
3446378916df74eca887cb36c380ed0ab151192e
|
[
"MIT"
] | null | null | null | 1,290.474126 | 274,432 | 0.956459 |
[
[
[
"This notebook is sample of the HAL QCD potential,\nthe effective mass fitting, and the effective energy shifts of two-baryon system\nfrom compressed NBS wavefunction sample_data.\n\nIn order to decompress the wave function, hal_pot_single_ch.py requires\nbinary \"PH1.compress48\" in \"yukawa library.\"",
"_____no_output_____"
]
],
[
[
"%pylab inline",
"Populating the interactive namespace from numpy and matplotlib\n"
],
[
"import seaborn as sns\nsns.set_style('ticks', {'axes.grid': True})\nsns.set_context('poster', font_scale=2.0)\n%config InlineBackend.figure_format = 'retina'\n\nplt.rcParams['figure.figsize'] = (12.8, 9.6)\nplt.rcParams['figure.facecolor'] = 'white'\n",
"_____no_output_____"
],
[
"ls ../data/sample_data/",
"\u001b[34mBBwave.dir.S4.00\u001b[m\u001b[m/ \u001b[34mcorrelator.PS.dir\u001b[m\u001b[m/\r\nREADME.md \u001b[34mcorrelator.multi.dir\u001b[m\u001b[m/\r\n"
],
[
"# import library \nfrom hal_pot_single_ch import HAL_pot\nfrom corr_baryon import Corr_2pt_Baryon, Corr_2pt_2Baryons, Delta_Eeff",
"_____no_output_____"
]
],
[
[
"lattice spacing",
"_____no_output_____"
]
],
[
[
"ainv = 2.194e3 # MeV\nhbarc = 0.197e3 # MeV fm\nlat_unit = hbarc/ainv",
"_____no_output_____"
]
],
[
[
"analyze baryon correlator and mass\n\\begin{equation}\nm_\\mathrm{eff}(t) = \\log \\frac{C_\\mathrm{B}(t)}{C_\\mathrm{B}(t+1)}\n\\end{equation}",
"_____no_output_____"
]
],
[
[
"cb = Corr_2pt_Baryon('Xi_CG05_CG05', bin_size=1, result_dir='../data/sample_data/')",
"# Read Xi_CG05_CG05\n# total 4 conf., bin size = 1, number of samples = 4\n"
],
[
"cb.plot_meff()",
"_____no_output_____"
],
[
"fig, ax = plt.subplots()\ncb.fit_meff(fit_min=15, fit_max=20, ax=ax)\nax.set_ylim(0.65, 0.68)\nleg = ax.legend(frameon=True)\nleg.get_frame().set_edgecolor('black')\nleg.get_frame().set_linewidth(2.0)\nax.set_title('$\\Xi$-baryon effective mass');",
"_____no_output_____"
],
[
"m_red = 0.5 * 0.665 ",
"_____no_output_____"
]
],
[
[
"initialize HAL_pot object with induced mass as an input parameter",
"_____no_output_____"
]
],
[
[
"hal = HAL_pot(m_red=m_red, result_dir='../data/sample_data/',\n it0 = 10, Ns=48, channel='xixi', bin_size=2)",
"# Read Xi_CG05_CG05\n# total 4 conf., bin size = 2, number of samples = 2\n"
]
],
[
[
"calc_pot method evaluate (effective) central potential of (3S1) 1S0 channel, and central, tensor part of 3S1 channel.",
"_____no_output_____"
]
],
[
[
"# potential\nresult = hal.calc_pot(pot_type='cen', spin='1s0')",
"# calc. Rcorr results.rcorr.binned/Rcorr_1s0_xixi_t009_2bin_4conf.dat\n# binning and decompress NBS\n# binning NBS S4.00 t = 9\n--- binning NBS wavefunc. --- 000\n>> decompress results.nbs.binned/NBSwave.S4.00.t009.binned.000.dat\n--- binning NBS wavefunc. --- 001\n>> decompress results.nbs.binned/NBSwave.S4.00.t009.binned.001.dat\n>> total 2 binned NBS files\n# load results.nbs.decomp/NBSwave.S4.00.t009.binned.000.decomp.dat\n# load results.nbs.decomp/NBSwave.S4.00.t009.binned.001.decomp.dat\n# save results.rcorr.binned/Rcorr_1s0_xixi_t009_2bin_4conf.dat\n# calc. Rcorr results.rcorr.binned/Rcorr_1s0_xixi_t010_2bin_4conf.dat\n# binning and decompress NBS\n# binning NBS S4.00 t = 10\n--- binning NBS wavefunc. --- 000\n>> decompress results.nbs.binned/NBSwave.S4.00.t010.binned.000.dat\n--- binning NBS wavefunc. --- 001\n>> decompress results.nbs.binned/NBSwave.S4.00.t010.binned.001.dat\n>> total 2 binned NBS files\n# load results.nbs.decomp/NBSwave.S4.00.t010.binned.000.decomp.dat\n# load results.nbs.decomp/NBSwave.S4.00.t010.binned.001.decomp.dat\n# save results.rcorr.binned/Rcorr_1s0_xixi_t010_2bin_4conf.dat\n# calc. Rcorr results.rcorr.binned/Rcorr_1s0_xixi_t011_2bin_4conf.dat\n# binning and decompress NBS\n# binning NBS S4.00 t = 11\n--- binning NBS wavefunc. --- 000\n>> decompress results.nbs.binned/NBSwave.S4.00.t011.binned.000.dat\n--- binning NBS wavefunc. --- 001\n>> decompress results.nbs.binned/NBSwave.S4.00.t011.binned.001.dat\n>> total 2 binned NBS files\n# load results.nbs.decomp/NBSwave.S4.00.t011.binned.000.decomp.dat\n# load results.nbs.decomp/NBSwave.S4.00.t011.binned.001.decomp.dat\n# save results.rcorr.binned/Rcorr_1s0_xixi_t011_2bin_4conf.dat\n>> return jackknife samples of (effective) central\n# output potential >> results.pot/pot_1s0_cen_xixi_t010_004conf_002bin.dat\n"
]
],
[
[
"text data of potential is stored in results.pot dir.",
"_____no_output_____"
]
],
[
[
"pot = np.loadtxt('results.pot/pot_1s0_cen_xixi_t010_004conf_002bin.dat')",
"_____no_output_____"
],
[
"pot.shape",
"_____no_output_____"
]
],
[
[
"columns of potential data\n\n0, 1, 2, 3, 4, 5, 6, 7, 8\n\nr, H0-term (av, err) , d/dt-term (av, err), d2/dt2-term (av, err), total (av, err)",
"_____no_output_____"
]
],
[
[
"fig, ax = plt.subplots()\nax.errorbar(pot[:,0]*lat_unit, pot[:,1]*ainv, pot[:,2]*ainv,\n fmt='bs', mfc='none', mew=2, capsize=10, capthick=2, label=r'$H_0$-term')\n\nax.errorbar(pot[:,0]*lat_unit, pot[:,3]*ainv, pot[:,4]*ainv,\n fmt='g^', mfc='none', mew=2, capsize=10, capthick=2, label=r'$\\partial/\\partial t$-term')\n\nax.errorbar(pot[:,0]*lat_unit, pot[:,5]*ainv, pot[:,6]*ainv,\n fmt='kd', mfc='none', mew=2, capsize=10, capthick=2, label=r'$\\partial^2/\\partial t^2$-term')\n\nax.errorbar(pot[:,0]*lat_unit, pot[:,7]*ainv, pot[:,8]*ainv,\n fmt='ro', mfc='none', mew=2, capsize=10, capthick=2, label='total')\n\nax.set_ylim(-50, 50)\n\nax.axhline(0, color='black')\nax.set_xlabel(r'$r$ [fm]', size=48)\nax.set_ylabel(r'$V(r)$ [MeV]', size=48)\nleg = ax.legend(frameon=True)\nleg.get_frame().set_edgecolor('black')\nleg.get_frame().set_linewidth(2.0)",
"_____no_output_____"
]
],
[
[
"# baryon mass and energy shift",
"_____no_output_____"
],
[
"if lattice unit is given, observables are plotted in physical scale",
"_____no_output_____"
]
],
[
[
"cb = Corr_2pt_Baryon('Xi_CG05_CG05', bin_size=1, result_dir='../data/sample_data/', lat_unit=lat_unit)",
"# Read Xi_CG05_CG05\n# total 4 conf., bin size = 1, number of samples = 4\n"
],
[
"fig, ax = plt.subplots()\ncb.fit_meff(fit_min=10, fit_max=20, ax=ax)\nleg = ax.legend(frameon=True)\nleg.get_frame().set_edgecolor('black')\nleg.get_frame().set_linewidth(2.0)\nax.set_ylim(1400, 1500)",
"_____no_output_____"
]
],
[
[
"Two-baryon correlator and the effective energy\n\\begin{equation}\nE_\\mathrm{eff}(t) = \\log \\frac{C_\\mathrm{BB}(t)}{C_\\mathrm{BB}(t+1)}\n\\end{equation}",
"_____no_output_____"
]
],
[
[
"# BB two baryon correlator\ncbb = Corr_2pt_2Baryons('xixi', bin_size=1, result_dir='../data/sample_data/', lat_unit=lat_unit)",
"Read xixi\n# total 4 conf., bin size = 1, number of samples = 4\n"
],
[
"fig, ax = plt.subplots()\ncbb.fit_Eeff(ax=ax)\nax.set_ylim(2800, 3000)",
"_____no_output_____"
]
],
[
[
"effective energy shift\n\\begin{equation}\n\\Delta E_\\mathrm{eff}(t) = \\log \\frac{R(t)}{R(t+1)}\n\\end{equation}\nwith\n\\begin{equation}\nR(t) = \\frac{C_\\mathrm{BB}(t)}{\\{C_\\mathrm{B}(t)\\}^2}\n\\end{equation}",
"_____no_output_____"
]
],
[
[
"# effective energy shifts\ndEeff = Delta_Eeff('xixi', bin_size=1, result_dir='../data/sample_data', lat_unit=lat_unit)",
"Read xixi\n# total 4 conf., bin size = 1, number of samples = 4\n# Read Xi_CG05_CG05\n# total 4 conf., bin size = 1, number of samples = 4\n"
],
[
"fig, ax = plt.subplots()\ndEeff.fit_dEeff(fit_min=7, fit_max=10, spin='1s0', ax=ax)\nax.set_ylim(-10, 10)\nleg = ax.legend(frameon=True)\nleg.get_frame().set_edgecolor('black')\nleg.get_frame().set_linewidth(2.0)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4acc9dc8fd4b61473c87d23722119b07f70b5f0f
| 33,859 |
ipynb
|
Jupyter Notebook
|
functions/functions_cams.ipynb
|
esowc/adc-toolbox
|
266684a317255249170a7aa178a62d5e76938892
|
[
"MIT"
] | 6 |
2021-12-09T01:25:30.000Z
|
2021-12-19T08:42:04.000Z
|
functions/functions_cams.ipynb
|
esowc/adc-toolbox
|
266684a317255249170a7aa178a62d5e76938892
|
[
"MIT"
] | 2 |
2021-12-09T12:39:57.000Z
|
2021-12-11T23:51:07.000Z
|
functions/functions_cams.ipynb
|
esowc/adc-toolbox
|
266684a317255249170a7aa178a62d5e76938892
|
[
"MIT"
] | 10 |
2021-11-10T17:58:03.000Z
|
2022-03-25T04:37:35.000Z
| 46.129428 | 149 | 0.470746 |
[
[
[
"# CAMS functions",
"_____no_output_____"
]
],
[
[
"def get_ADS_API_key():\n \n \"\"\" Get ADS API key to download CAMS datasets\n \n Returns:\n API_key (str): ADS API key\n \"\"\"\n\n keys_path = os.path.join('/', '/'.join(\n os.getcwd().split('/')[1:3]), 'adc-toolbox', \n os.path.relpath('data/keys.txt'))\n\n try:\n keys_file = open(keys_path, 'r')\n keys = keys_file.readlines()\n environ_keys = [key.rstrip() for key in keys]\n ADS_key = environ_keys[0]\n\n except:\n \n print('ERROR: You need to create a keys.txt file in the data folder with the ADS API key.')\n print('Get your ADS API key by registering at https://ads.atmosphere.copernicus.eu/api-how-to.')\n raise KeyboardInterrupt\n\n return ADS_key",
"_____no_output_____"
],
[
"def CAMS_download(dates, start_date, end_date, component, component_nom, lat_min, lat_max, lon_min, lon_max, \n area_name, model_full_name, model_level, CAMS_UID = None, CAMS_key = None):\n\n \"\"\" Query and download the CAMS levels dataset from CDS API\n\n Args:\n dates (arr): Query dates\n start_date (str): Query start date\n end_date (str): Query end date\n component (str): Component name\n component_nom (str): Component chemical nomenclature\n lat_min (int): Minimum latitude\n lat_max (int): Maximum latitude\n lon_min (int): Minimum longitude\n lon_max (int): Maximum longitude\n area_name (str): User defined area name\n model_full_name (str): Full name of the CAMS model among:\n - 'cams-global-atmospheric-composition-forecasts' \n - 'cams-global-reanalysis-eac4-monthly'\n model_level (str): Model levels:\n - 'Simple' for total columns\n - 'Multiple' for levels\n CAMS_UID (str): ADS user ID\n CAMS_key (str): ADS key\n\n Returns:\n CAMS_product_name (str): Product name of CAMS product\n CAMS_type (str): Model type:\n - 'Forecast'\n - 'Reanalysis'\n \"\"\"\n \n # Get API key\n if CAMS_UID != None and CAMS_key != None:\n ADS_key = CAMS_UID + ':' + CAMS_key\n else:\n ADS_key = get_ADS_API_key()\n \n # Connect to the server\n c = cdsapi.Client(url = 'https://ads.atmosphere.copernicus.eu/api/v2', key = ADS_key)\n\n # Download component concentration dataset\n if model_full_name == 'cams-global-atmospheric-composition-forecasts':\n\n CAMS_type = 'Forecast'\n\n if model_level == 'Multiple':\n \n CAMS_product_name = ('CAMS_FORECAST_' + component_nom + '_137_LEVELS_' + start_date + '_' + end_date +\n '_' + area_name + '.grib')\n CAMS_product_path = os.path.join('/', '/'.join(\n os.getcwd().split('/')[1:3]), 'adc-toolbox', \n os.path.relpath('data/cams/' + component_nom + '/' + CAMS_product_name))\n \n if os.path.isfile(CAMS_product_path):\n\n print('The file exists, it will not be downloaded again.')\n \n else:\n\n print('The file does not exist, it will be downloaded.')\n c.retrieve(\n model_full_name,\n {\n 'date': start_date + '/' + end_date,\n 'type': 'forecast',\n 'format': 'grib',\n 'variable': component,\n 'model_level': [str(x + 1) for x in range(137)],\n 'time': '00:00',\n 'leadtime_hour': [str(x) for x in range(0, 24, 3)],\n 'area': [lat_max, lon_min, lat_min, lon_max],\n },\n CAMS_product_path)\n \n elif model_level == 'Single':\n\n CAMS_product_name = ('CAMS_FORECAST_' + component_nom + '_TC_' + start_date + '_' + end_date + \n '_' + area_name + '.grib')\n CAMS_product_path = os.path.join('/', '/'.join(\n os.getcwd().split('/')[1:3]), 'adc-toolbox', \n os.path.relpath('data/cams/' + component_nom + '/' + CAMS_product_name))\n \n if os.path.isfile(CAMS_product_path):\n \n print('The file exists, it will not be downloaded again.')\n \n else:\n\n print('The file does not exist, it will be downloaded.')\n c = cdsapi.Client(url = 'https://ads.atmosphere.copernicus.eu/api/v2', key = ADS_key)\n c.retrieve(\n 'cams-global-atmospheric-composition-forecasts',\n {\n 'date': start_date + '/' + end_date,\n 'type': 'forecast',\n 'format': 'grib',\n 'variable': 'total_column_' + component,\n 'time': '00:00',\n 'leadtime_hour': [str(x) for x in range(0, 24, 3)],\n 'area': [lat_max, lon_min, lat_min, lon_max],\n },\n CAMS_product_path)\n\n elif model_full_name == 'cams-global-reanalysis-eac4-monthly':\n \n CAMS_type = 'Reanalysis'\n \n if model_level == 'Single':\n\n CAMS_product_name = ('CAMS_REANALYSIS_' + component_nom + '_TC_' + start_date + '_' + end_date + \n '_' + area_name + '.grib')\n CAMS_product_path = os.path.join('/', '/'.join(\n os.getcwd().split('/')[1:3]), 'adc-toolbox', \n os.path.relpath('data/cams/' + component_nom + '/' + CAMS_product_name))\n\n if os.path.isfile(CAMS_product_path):\n\n print('The file exists, it will not be downloaded again.')\n\n else:\n\n print('The file does not exist, it will be downloaded.') \n\n months = []\n years = []\n \n for date in dates:\n\n year = date.split('-')[0]\n month = date.split('-')[1]\n\n if year not in years:\n years.append(year)\n \n if month not in months:\n months.append(month)\n \n c.retrieve(\n model_full_name,\n {\n 'format': 'grib',\n 'variable': 'total_column_' + component,\n 'year': years,\n 'month': months,\n 'product_type': 'monthly_mean',\n 'area': [lat_max, lon_min, lat_min, lon_max],\n },\n CAMS_product_path)\n\n elif model_level == 'Multiple':\n \n start_dates = pd.date_range(np.datetime64(start_date), np.datetime64(end_date), freq='MS')\n start_dates = tuple(np.unique([date.strftime('%Y-%m-%d') for date in start_dates]))\n\n end_dates = pd.date_range(np.datetime64(start_date), np.datetime64(end_date), freq='M')\n end_dates = tuple(np.unique([date.strftime('%Y-%m-%d') for date in end_dates]))\n\n # Download month by month (to avoid crashing the server)\n CAMS_product_name = []\n\n for start_date, end_date in zip(start_dates, end_dates):\n \n CAMS_product_name_month = ('CAMS_REANALYSIS_' + component_nom + '_60_LEVELS_' + start_date + '_' + end_date + \n '_' + area_name + '.grib')\n CAMS_product_path = os.path.join('/', '/'.join(\n os.getcwd().split('/')[1:3]), 'adc-toolbox', \n os.path.relpath('data/cams/' + component_nom + '/' + CAMS_product_name_month))\n\n if os.path.isfile(CAMS_product_path):\n\n print('The file exists, it will not be downloaded again.')\n\n else:\n\n print('The file does not exist, it will be downloaded.') \n c.retrieve(\n 'cams-global-reanalysis-eac4',\n {\n 'date': start_date + '/' + end_date,\n 'format': 'grib',\n 'variable': component,\n 'model_level': [str(x + 1) for x in range(60)],\n 'time': ['00:00', '03:00', '06:00', '09:00', '12:00', '15:00', '18:00', '21:00',],\n 'area': [lat_max, lon_min, lat_min, lon_max],\n },\n CAMS_product_path)\n\n CAMS_product_name.append(CAMS_product_name_month)\n\n return CAMS_product_name, CAMS_type",
"_____no_output_____"
],
[
"def CAMS_read(CAMS_product_name, component, component_nom, dates):\n\n \"\"\" Read CAMS levels dataset as xarray dataset object\n\n Args:\n CAMS_product_name (str): Product name of CAMS product\n component (str): Component name\n component_nom (str): Component chemical nomenclature\n dates (arr): Query dates\n \n Returns:\n CAMS_ds (xarray): CAMS levels dataset in xarray format\n \"\"\"\n\n # Read as xarray dataset object\n if isinstance(CAMS_product_name, list): \n CAMS_ds = xr.open_mfdataset(os.path.join('/', '/'.join(\n os.getcwd().split('/')[1:3]), 'adc-toolbox', \n os.path.relpath('data/cams/' + component_nom + '/CAMS_REANALYSIS_' + component_nom + '_60_LEVELS_*')),\n concat_dim = 'time')\n\n else:\n CAMS_ds = xr.open_dataset(os.path.join('/', '/'.join(\n os.getcwd().split('/')[1:3]), 'adc-toolbox', \n os.path.relpath('data/cams/' + component_nom + '/' + CAMS_product_name)))\n\n # Change name to component \n if 'hybrid' in CAMS_ds.keys():\n if component == 'ozone':\n CAMS_ds = CAMS_ds.rename({'go3': 'component'})\n else:\n CAMS_ds = CAMS_ds.rename({component_nom.lower(): 'component'})\n\n else: \n if component == 'ozone':\n CAMS_ds = CAMS_ds.rename({'gtco3': 'component'})\n else:\n CAMS_ds = CAMS_ds.rename({'tc' + component_nom.lower(): 'component'})\n\n if 'REANALYSIS_' + component_nom + '_TC_' in CAMS_product_name:\n \n # Remove data for dates that have been downloaded but not asked for (error of the CAMS API!)\n all_datetimes = []\n for date in dates:\n year = int(date.split('-')[0])\n month = int(date.split('-')[1])\n time_str = np.datetime64(dt.datetime(year, month, 1, 0, 0, 0, 0))\n all_datetimes.append(time_str)\n\n # Drop datetimes\n datetimes_to_delete = np.setdiff1d(CAMS_ds.time.values, np.array(all_datetimes))\n if datetimes_to_delete.size != 0:\n CAMS_ds = CAMS_ds.drop_sel(time = datetimes_to_delete) \n\n # Available dates\n dates_to_keep = np.intersect1d(CAMS_ds.time.values, np.array(all_datetimes))\n dates = tuple(dates_to_keep.astype('datetime64[M]').astype(str))\n \n # Remove step since there is only one\n CAMS_ds = CAMS_ds.drop('step')\n\n # Arrange coordinates\n CAMS_ds = CAMS_ds.assign_coords(longitude = (((CAMS_ds.longitude + 180) % 360) - 180)).sortby('longitude')\n CAMS_ds = CAMS_ds.sortby('latitude')\n\n # Assign time as dimension (when there is only one time)\n if CAMS_ds.time.values.size == 1:\n CAMS_ds = CAMS_ds.expand_dims(dim = ['time'])\n \n # Get model levels\n CAMS_levels_df = CAMS_levels(CAMS_ds, CAMS_product_name)\n\n return CAMS_ds, dates, CAMS_levels_df",
"_____no_output_____"
],
[
"def CAMS_levels(CAMS_ds, CAMS_product_name):\n\n \"\"\" Create table with information about the CAMS model levels\n\n Args:\n CAMS_ds (xarray): CAMS levels dataset in xarray format\n CAMS_product_name (str): Product name of CAMS product\n \n Returns:\n CAMS_levels_df (dataframe): Table with CAMS levels data\n \"\"\"\n \n # Read CSV table with information about the model levels\n if '60_LEVELS' in CAMS_product_name:\n CAMS_levels_df = pd.read_csv(os.path.join('/', '/'.join(\n os.getcwd().split('/')[1:3]), 'adc-toolbox', \n os.path.relpath('data/cams/60-levels-definition.csv')))\n else:\n CAMS_levels_df = pd.read_csv(os.path.join('/', '/'.join(\n os.getcwd().split('/')[1:3]), 'adc-toolbox', \n os.path.relpath('data/cams/137-levels-definition.csv')))\n\n # Drop first row and set n as index hybrid\n CAMS_levels_df = CAMS_levels_df.drop(0).reset_index(drop = True)\n CAMS_levels_df = CAMS_levels_df.set_index('n')\n CAMS_levels_df.index.names = ['hybrid']\n\n # Change important columns to numeric\n CAMS_levels_df['ph [Pa]'] = pd.to_numeric(CAMS_levels_df['ph [hPa]']) * 100\n CAMS_levels_df['Geopotential Altitude [m]'] = pd.to_numeric(CAMS_levels_df['Geopotential Altitude [m]'])\n CAMS_levels_df['Density [kg/m^3]'] = pd.to_numeric(CAMS_levels_df['Density [kg/m^3]'])\n\n # Calculate difference from geopotential altitude\n CAMS_levels_df['Depth [m]'] = CAMS_levels_df['Geopotential Altitude [m]'].diff(-1)\n CAMS_levels_df['Depth [m]'].iloc[-1] = CAMS_levels_df['Geopotential Altitude [m]'].iloc[-1]\n\n return CAMS_levels_df",
"_____no_output_____"
],
[
"def CAMS_pressure(CAMS_ds, CAMS_product_name, CAMS_levels_df, start_date, end_date, component_nom, \n lat_min, lat_max, lon_min, lon_max, area_name, CAMS_UID = None, CAMS_key = None):\n\n \"\"\" Download surface pressure and calculate levels pressure following the instructions given at:\n https://confluence.ecmwf.int/display/OIFS/4.4+OpenIFS%3A+Vertical+Resolution+and+Configurations\n\n Args:\n CAMS_ds (xarray): CAMS levels dataset in xarray format\n CAMS_product_name (str): Product name of CAMS product\n CAMS_levels_df (dataframe): Table with 137 CAMS levels data\n start_date (str): Query start date\n end_date (str): Query end date\n component_nom (str): Component chemical nomenclature\n lat_min (int): Minimum latitude\n lat_max (int): Maximum latitude\n lon_min (int): Minimum longitude\n lon_max (int): Maximum longitude\n area_name (str): User defined area name\n CAMS_UID (str): ADS user ID\n CAMS_key (str): ADS key\n \n Returns:\n CAMS_ds (xarray): CAMS levels dataset in xarray format\n \"\"\"\n \n CAMS_pressure_product_name = ('_SURFACE_PRESSURE_' + start_date + '_' + end_date +\n '_' + area_name + '.grib')\n\n # Get API key\n if CAMS_UID != None and CAMS_key != None:\n ADS_key = CAMS_UID + ':' + CAMS_key\n else:\n ADS_key = get_ADS_API_key()\n \n # Connect to the server\n c = cdsapi.Client(url = 'https://ads.atmosphere.copernicus.eu/api/v2', key = ADS_key)\n\n # Dowload surface pressure data\n if 'FORECAST' in CAMS_product_name:\n\n CAMS_surface_pressure_path = os.path.join('/', '/'.join(\n os.getcwd().split('/')[1:3]), 'adc-toolbox', \n os.path.relpath('data/cams/' + component_nom + '/CAMS_FORECAST' + CAMS_pressure_product_name))\n \n c.retrieve(\n 'cams-global-atmospheric-composition-forecasts',\n {\n 'date': start_date + '/' + end_date,\n 'type': 'forecast',\n 'format': 'grib',\n 'variable': 'surface_pressure',\n 'leadtime_hour': [str(x) for x in range(0, 24, 3)],\n 'time': '00:00',\n 'area': [lat_max, lon_min, lat_min, lon_max],\n },\n CAMS_surface_pressure_path)\n\n elif 'REANALYSIS' in CAMS_product_name:\n\n CAMS_surface_pressure_path = os.path.join('/', '/'.join(\n os.getcwd().split('/')[1:3]), 'adc-toolbox', \n os.path.relpath('data/cams/' + component_nom + '/CAMS_REANALYSIS' + CAMS_pressure_product_name))\n \n c.retrieve(\n 'cams-global-reanalysis-eac4',\n {\n 'date': start_date + '/' + end_date,\n 'format': 'grib',\n 'variable': 'surface_pressure',\n 'time': ['00:00', '03:00', '06:00',\n '09:00', '12:00', '15:00',\n '18:00', '21:00',],\n 'area': [lat_max, lon_min, lat_min, lon_max],\n },\n CAMS_surface_pressure_path)\n\n hybrid = CAMS_ds['hybrid'].data\n time = CAMS_ds['time'].data\n step = CAMS_ds['step'].data\n latitude = CAMS_ds['latitude'].data\n longitude = CAMS_ds['longitude'].data\n\n # Read surface pressure\n model_pressure_ds = xr.open_dataarray(CAMS_surface_pressure_path)\n\n # Arrange coordinates\n model_pressure_ds = model_pressure_ds.assign_coords(longitude = (((model_pressure_ds.longitude + 180) % 360) - 180)).sortby('longitude')\n model_pressure_ds = model_pressure_ds.sortby('latitude')\n \n # Assign time as dimension (when there is only one time)\n if model_pressure_ds.time.values.size == 1:\n model_pressure_ds = model_pressure_ds.expand_dims(dim = ['time'])\n \n # Transpose dimensions\n model_pressure_ds = model_pressure_ds.transpose('time', 'step', 'latitude', 'longitude')\n\n # Subset surface pressure dataset\n model_pressure_ds = subset(model_pressure_ds, bbox, sensor, component_nom, sensor_type, subset_type = 'model_subset')\n\n sp_array = xr.DataArray(\n model_pressure_ds.values,\n dims = ('time', 'step', 'latitude', 'longitude'),\n coords = {\n 'time': ('time', time),\n 'step': ('step', step),\n 'latitude': ('latitude', latitude),\n 'longitude': ('longitude', longitude),\n },\n name = 'surface_pressure'\n )\n\n a_array = xr.DataArray(\n CAMS_levels_df['a [Pa]'],\n dims = ('hybrid'),\n coords = {'hybrid': ('hybrid', hybrid),},\n name = 'a'\n )\n\n b_array = xr.DataArray(\n CAMS_levels_df['b'],\n dims = ('hybrid'),\n coords = {'hybrid': ('hybrid', hybrid),},\n name = 'b'\n )\n\n CAMS_ds['surface_pressure'] = sp_array\n CAMS_ds['a'] = a_array\n CAMS_ds['b'] = b_array\n\n CAMS_ds['pressure_1/2'] = CAMS_ds['a'] + CAMS_ds['surface_pressure'] * CAMS_ds['b']\n CAMS_ds['pressure_-1/2'] = CAMS_ds['pressure_1/2'].shift(hybrid = 1)\n CAMS_ds['pressure_-1/2'] = CAMS_ds['pressure_-1/2'].where(~np.isnan(CAMS_ds['pressure_-1/2']), 0, drop = False)\n CAMS_ds['pressure'] = 0.5 * (CAMS_ds['pressure_-1/2'] + CAMS_ds['pressure_1/2'])\n CAMS_ds = CAMS_ds.drop_vars(['a', 'b', 'surface_pressure', 'pressure_1/2', 'pressure_-1/2'])\n \n return CAMS_ds",
"_____no_output_____"
],
[
"def CAMS_get_levels_data(CAMS_ds, CAMS_product_name, CAMS_levels_df, column_type, \n lat_min, lat_max, lon_min, lon_max):\n \n \"\"\" Get the tropospheric or column model data, depending on the nature of the sensor data\n\n Args:\n CAMS_ds (xarray): CAMS levels dataset in xarray format\n CAMS_product_name (str): Product name of CAMS product\n CAMS_levels_df (dataframe): Table with 137 CAMS levels data\n column_type (str): Tropospheric or total column\n lat_min (int): Minimum latitude\n lat_max (int): Maximum latitude\n lon_min (int): Minimum longitude\n lon_max (int): Maximum longitude\n \n Returns:\n CAMS_ds (xarray): CAMS levels dataset in xarray format\n \"\"\"\n\n # Get units and calculate tropospheric columns if needed\n units = CAMS_ds.component.attrs['units'] \n\n if 'REANALYSIS' in CAMS_product_name:\n\n if column_type == 'tropospheric':\n print('The model total columns will be directly compared to the tropospheric sensor columns.')\n\n elif column_type == 'total':\n print('The model total columns will be compared to the total sensor columns.')\n\n elif 'FORECAST' in CAMS_product_name:\n\n if column_type == 'tropospheric':\n\n print('The model tropospheric columns will be compared to the tropospheric sensor columns.')\n print('The model tropospheric columns will be estimated (pressures above or equal to 300 hPa).')\n \n # Calculate levels pressure\n CAMS_ds = CAMS_pressure(CAMS_ds, CAMS_product_name, CAMS_levels_df, start_date, end_date, component_nom, \n lat_min, lat_max, lon_min, lon_max, area_name, CAMS_UID = None, CAMS_key = None)\n\n if apply_kernels == False:\n \n CAMS_ds = CAMS_ds.where(CAMS_ds.pressure >= 30000, drop = True)\n CAMS_ds = CAMS_ds.sum(dim = 'hybrid')\n CAMS_ds['component'] = CAMS_ds.component.assign_attrs({'units': units})\n\n if column_type == 'total':\n print('The model total columns will be compared to the total sensor columns.')\n\n return CAMS_ds",
"_____no_output_____"
],
[
"def CAMS_kg_kg_to_kg_m2(CAMS_ds, CAMS_levels_df, sensor, start_date, end_date, \n component_nom, apply_kernels = False, CAMS_UID = None, CAMS_key = None):\n\n \"\"\" Convert the units of the CAMS partial columns for any component from kg/kg to kg/m2. To do this,\n calculate columns above each CAMS half level assuming it is 0 at the top of the atmosphere\n\n Args:\n CAMS_ds (xarray): CAMS levels dataset in xarray format\n CAMS_levels_df (dataframe): Table with 137 CAMS levels data\n sensor (str): Name of the sensor\n start_date (str): Query start date\n end_date (str): Query end date\n component_nom (str): Component chemical nomenclature\n apply_kernels (bool): Apply (True) or not (False) the averaging kernels\n CAMS_UID (str): ADS user ID\n CAMS_key (str): ADS key\n \n Returns:\n CAMS_ds (xarray): CAMS levels dataset in xarray format\n \"\"\"\n\n # Calculate columns above each CAMS half level \n if sensor == 'tropomi' and apply_kernels == True:\n\n print('The columns above each CAMS half level will be calculated.')\n\n # Initialize new array\n CAMS_ds_all = []\n\n for time in CAMS_ds.time:\n\n # Select data for each timestep\n CAMS_ds_time_old = CAMS_ds.sel(time = time)\n\n # Initialize partial columns at the top of the atmosphere (hybrid = 1) as 0\n PC_hybrid_0 = CAMS_ds_time_old.sel(hybrid = 1)\n PC_hybrid_0['component'] = PC_hybrid_0['component'].where(PC_hybrid_0['component'] <= 0, 0, drop = False)\n PC_hybrid_0 = PC_hybrid_0.expand_dims(dim = ['hybrid'])\n\n # Create new model dataset\n PC_above_all = []\n PC_above_all.append(PC_hybrid_0)\n CAMS_ds_time_new = PC_hybrid_0\n \n for hybrid in range(1, 137):\n\n # Get current and previous partial columns and level pressures\n PC_last = CAMS_ds_time_new.component.sel(hybrid = hybrid)\n PC_current = CAMS_ds_time_old.component.sel(hybrid = hybrid + 1)\n pressure_last = CAMS_ds_time_old.pressure.sel(hybrid = hybrid)\n pressure_current = CAMS_ds_time_old.pressure.sel(hybrid = hybrid + 1)\n\n # Calculate pressure difference\n pressure_diff = pressure_current - pressure_last\n\n # Calculate partial columns above each model level\n # Units: (kg/kg * kg/m*s2) * s2/m -> kg/m2\n PC_above = CAMS_ds_time_old.sel(hybrid = hybrid + 1)\n PC_above['component'] = PC_last + PC_current * pressure_diff * (1/9.81)\n\n # Append result\n PC_above_all.append(PC_above)\n CAMS_ds_time_new = xr.concat(PC_above_all, pd.Index(range(1, hybrid + 2), name = 'hybrid'))\n\n CAMS_ds_all.append(CAMS_ds_time_new)\n\n CAMS_ds = xr.concat(CAMS_ds_all, dim = 'time')\n\n else:\n\n # Create xarray object from CAMS model levels information\n CAMS_levels_ds = CAMS_levels_df.to_xarray()\n\n # Convert units from kg/kg to kg/m3\n CAMS_ds = CAMS_ds * CAMS_levels_ds['Density [kg/m^3]']\n\n # Convert units from kg/m3 to kg/m2\n CAMS_ds = CAMS_ds * CAMS_levels_ds['Depth [m]']\n\n return CAMS_ds",
"_____no_output_____"
],
[
"def CAMS_kg_m2_to_molecules_cm2(CAMS_ds, component_mol_weight):\n\n \"\"\" Convert the units of the CAMS dataset for any component from kg/m2 to molecules/cm2\n\n Args:\n CAMS_ds (xarray): CAMS levels dataset in xarray format\n component_mol_weight (float): Component molecular weight\n\n Returns:\n CAMS_ds (xarray): CAMS levels dataset in xarray format\n \"\"\"\n\n # Convert units from kg/m2 to molecules/cm2\n NA = 6.022*10**23\n CAMS_ds['component'] = (CAMS_ds['component'] * NA * 1000) / (10000 * component_mol_weight)\n \n return CAMS_ds",
"_____no_output_____"
],
[
"def CAMS_molecules_cm2_to_DU(CAMS_ds):\n\n \"\"\" Convert the units of the CAMS dataset for any component from molecules/cm2 to DU for ozone\n\n Args:\n CAMS_ds (xarray): CAMS levels dataset in xarray format\n\n Returns:\n CAMS_ds (xarray): CAMS levels dataset in xarray format\n \"\"\"\n\n # Convert units from molecules/cm2 to DU\n CAMS_ds = CAMS_ds / (2.69*10**16)\n \n return CAMS_ds",
"_____no_output_____"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4accb3c47fb1883d6f9e4d77be42c80ce7db359e
| 40,942 |
ipynb
|
Jupyter Notebook
|
study/r/003.ipynb
|
data-block/dev
|
e7a71eda6f1c4f18b262aa07899577529b7e827c
|
[
"MIT"
] | 2 |
2019-10-29T12:19:45.000Z
|
2020-01-17T02:36:57.000Z
|
study/r/003.ipynb
|
data-block/dev
|
e7a71eda6f1c4f18b262aa07899577529b7e827c
|
[
"MIT"
] | null | null | null |
study/r/003.ipynb
|
data-block/dev
|
e7a71eda6f1c4f18b262aa07899577529b7e827c
|
[
"MIT"
] | null | null | null | 96.789598 | 476 | 0.211445 |
[
[
[
"### jitter\n- 흔들기\n\n### Barchart\n- 이산변수, 1변수\n - 2변수\n - x변수 외에도 이산 변수를 추가할 수 있음\n - 시각화\n - color\n - color = class\n - fill\n - fill = class\n - 1변수 외에 변수를 추가해도 정보를 많이 주지 않음\n - dodge\n - position = \"dodge\"\n - 3~4개의 변수가 적당함\n - fill\n - position = \"fill\"\n - %(비율)로 채워짐 ... 전체 중 비중을 따지는 것\n - 막대그래프로 그리고 따질 수는 있는데... 파이 차트로 하는 게 좋음\n \n### Histogram\n- 연속형 변수\n\n### Boxplot\n- x: 이산 변수, y: 연속 변수\n- x 변수 각각의 값에 대해서 연속 변수인 y의 분포를 볼 수 있음 (박스의 높이)\n- x변수의 값은 이산 변수면서, factor라면 변수의 값이 character라서 display가 복잡할 가능성이 높음\n\n### Pie-chart\n\n- theta = \"y\"\n - 각은 y값을 가져야 한다.",
"_____no_output_____"
],
[
"# Textmining\n## I have a dream.",
"_____no_output_____"
],
[
"# 0. Setup",
"_____no_output_____"
]
],
[
[
"library(tm) # for text mining\nlibrary(SnowballC) # for text stemming\nlibrary(wordcloud) # word-cloud generator \nlibrary(RColorBrewer) # color palettes\nlibrary(ggplot2)\nlibrary(dplyr)\n# activate(c(\"tm\", \"SnowballC\"))",
"_____no_output_____"
]
],
[
[
"# 1. Infile and prepare text (docs)",
"_____no_output_____"
]
],
[
[
"filePath <- paste0(\"http://www.sthda.com/sthda/RDoc/example-files/\",\n \"martin-luther-king-i-have-a-dream-speech.txt\")\ntext <- readLines(filePath)\ndocs <- Corpus(VectorSource(text))\n# inspect(docs)\nclass(docs)",
"_____no_output_____"
],
[
"inspect(docs)",
"<<SimpleCorpus>>\nMetadata: corpus specific: 1, document level (indexed): 0\nContent: documents: 46\n\n [1] \n [2] And so even though we face the difficulties of today and tomorrow, I still have a dream. It is a dream deeply rooted in the American dream. \n [3] \n [4] I have a dream that one day this nation will rise up and live out the true meaning of its creed: \n [5] \n [6] We hold these truths to be self-evident, that all men are created equal. \n [7] \n [8] I have a dream that one day on the red hills of Georgia, the sons of former slaves and the sons of former slave owners will be able to sit down together at the table of brotherhood. \n [9] \n[10] I have a dream that one day even the state of Mississippi, a state sweltering with the heat of injustice, sweltering with the heat of oppression, will be transformed into an oasis of freedom and justice. \n[11] \n[12] I have a dream that my four little children will one day live in a nation where they will not be judged by the color of their skin but by the content of their character. \n[13] \n[14] I have a dream today! \n[15] \n[16] I have a dream that one day, down in Alabama, with its vicious racists, with its governor having his lips dripping with the words of interposition and nullification, one day right there in Alabama little black boys and black girls will be able to join hands with little white boys and white girls as sisters and brothers. \n[17] \n[18] I have a dream today! \n[19] \n[20] I have a dream that one day every valley shall be exalted, and every hill and mountain shall be made low, the rough places will be made plain, and the crooked places will be made straight; and the glory of the Lord shall be revealed and all flesh shall see it together. \n[21] \n[22] This is our hope, and this is the faith that I go back to the South with. \n[23] \n[24] With this faith, we will be able to hew out of the mountain of despair a stone of hope. With this faith, we will be able to transform the jangling discords of our nation into a beautiful symphony of brotherhood. With this faith, we will be able to work together, to pray together, to struggle together, to go to jail together, to stand up for freedom together, knowing that we will be free one day.\n[25] \n[26] And this will be the day, this will be the day when all of God s children will be able to sing with new meaning: \n[27] \n[28] My country tis of thee, sweet land of liberty, of thee I sing. \n[29] Land where my fathers died, land of the Pilgrim s pride, \n[30] From every mountainside, let freedom ring! \n[31] And if America is to be a great nation, this must become true. \n[32] And so let freedom ring from the prodigious hilltops of New Hampshire. \n[33] Let freedom ring from the mighty mountains of New York. \n[34] Let freedom ring from the heightening Alleghenies of Pennsylvania. \n[35] Let freedom ring from the snow-capped Rockies of Colorado. \n[36] Let freedom ring from the curvaceous slopes of California. \n[37] \n[38] But not only that: \n[39] Let freedom ring from Stone Mountain of Georgia. \n[40] Let freedom ring from Lookout Mountain of Tennessee. \n[41] Let freedom ring from every hill and molehill of Mississippi. \n[42] From every mountainside, let freedom ring. \n[43] And when this happens, when we allow freedom ring, when we let it ring from every village and every hamlet, from every state and every city, we will be able to speed up that day when all of God s children, black men and white men, Jews and Gentiles, Protestants and Catholics, will be able to join hands and sing in the words of the old Negro spiritual: \n[44] Free at last! Free at last! \n[45] \n[46] Thank God Almighty, we are free at last! \n"
]
],
[
[
"# 2. Cleanup the text (docs)",
"_____no_output_____"
]
],
[
[
"# `dplyr` way\ntoSpace <- content_transformer(\n function (x , pattern) gsub(pattern, \" \", x))\ndocs <- docs %>% \n tm_map(toSpace, \"/\") %>%\n tm_map(toSpace, \"@\") %>%\n tm_map(toSpace, \"\\\\|\")",
"_____no_output_____"
],
[
"docs <- docs %>% \n tm_map(content_transformer(tolower)) %>% # Convert it to lower case\n tm_map(removeNumbers) %>% # Remove numbers\n tm_map(removeWords, stopwords(\"english\")) %>% # Remove english common stopwords\n tm_map(removeWords, c(\"blabla1\", \"blabla2\")) %>% # Remove your own stop word\n tm_map(removePunctuation) %>% # Remove punctuations \n tm_map(stripWhitespace) # Eliminate extra white spaces",
"_____no_output_____"
],
[
"inspect(docs)",
"<<SimpleCorpus>>\nMetadata: corpus specific: 1, document level (indexed): 0\nContent: documents: 46\n\n [1] \n [2] even though face difficulties today tomorrow still dream dream deeply rooted american dream \n [3] \n [4] dream one day nation will rise live true meaning creed \n [5] \n [6] hold truths selfevident men created equal \n [7] \n [8] dream one day red hills georgia sons former slaves sons former slave owners will able sit together table brotherhood \n [9] \n[10] dream one day even state mississippi state sweltering heat injustice sweltering heat oppression will transformed oasis freedom justice \n[11] \n[12] dream four little children will one day live nation will judged color skin content character \n[13] \n[14] dream today \n[15] \n[16] dream one day alabama vicious racists governor lips dripping words interposition nullification one day right alabama little black boys black girls will able join hands little white boys white girls sisters brothers \n[17] \n[18] dream today \n[19] \n[20] dream one day every valley shall exalted every hill mountain shall made low rough places will made plain crooked places will made straight glory lord shall revealed flesh shall see together \n[21] \n[22] hope faith go back south \n[23] \n[24] faith will able hew mountain despair stone hope faith will able transform jangling discords nation beautiful symphony brotherhood faith will able work together pray together struggle together go jail together stand freedom together knowing will free one day\n[25] \n[26] will day will day god s children will able sing new meaning \n[27] \n[28] country tis thee sweet land liberty thee sing \n[29] land fathers died land pilgrim s pride \n[30] every mountainside let freedom ring \n[31] america great nation must become true \n[32] let freedom ring prodigious hilltops new hampshire \n[33] let freedom ring mighty mountains new york \n[34] let freedom ring heightening alleghenies pennsylvania \n[35] let freedom ring snowcapped rockies colorado \n[36] let freedom ring curvaceous slopes california \n[37] \n[38] \n[39] let freedom ring stone mountain georgia \n[40] let freedom ring lookout mountain tennessee \n[41] let freedom ring every hill molehill mississippi \n[42] every mountainside let freedom ring \n[43] happens allow freedom ring let ring every village every hamlet every state every city will able speed day god s children black men white men jews gentiles protestants catholics will able join hands sing words old negro spiritual \n[44] free last free last \n[45] \n[46] thank god almighty free last \n"
],
[
"termMat <- TermDocumentMatrix(docs)\ntermTable <- as.matrix(termMat)\nhead(termTable, 2)\nfreqTable <- data.frame(word = rownames(termTable),\n freq = rowSums(termTable))\nfreqTable$word <- rownames(freqTable)\nfreqTable <- freqTable %>% arrange(desc(freq))\nhead(freqTable, 2)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
4accb55bfcf8c2585b071e09805f050ce9306d0c
| 60,494 |
ipynb
|
Jupyter Notebook
|
examples/noise_model.ipynb
|
tomekzaw/ewl
|
a2eeb4027e2baff7051a7ac80f886feb3b4c84cc
|
[
"MIT"
] | 6 |
2021-04-20T13:33:17.000Z
|
2021-11-26T09:28:11.000Z
|
examples/noise_model.ipynb
|
tomekzaw/ewl
|
a2eeb4027e2baff7051a7ac80f886feb3b4c84cc
|
[
"MIT"
] | 11 |
2021-04-20T22:02:20.000Z
|
2022-03-21T20:46:22.000Z
|
examples/noise_model.ipynb
|
tomekzaw/ewl
|
a2eeb4027e2baff7051a7ac80f886feb3b4c84cc
|
[
"MIT"
] | null | null | null | 169.45098 | 22,812 | 0.896469 |
[
[
[
"#!pip install ewl",
"_____no_output_____"
],
[
"%matplotlib inline",
"_____no_output_____"
],
[
"from ewl import *",
"_____no_output_____"
],
[
"from qiskit.providers.aer.noise import NoiseModel\nfrom qiskit.providers.aer.noise import QuantumError, ReadoutError\nfrom qiskit.providers.aer.noise import pauli_error",
"_____no_output_____"
],
[
"psi = (Qubit('00') + i * Qubit('11')) / sqrt2\npsi",
"_____no_output_____"
],
[
"theta1, alpha1, beta1 = sp.symbols('theta1 alpha1 beta1')\nalice = U(theta=theta1, alpha=alpha1, beta=beta1)\nalice",
"_____no_output_____"
],
[
"theta2, alpha2, beta2 = sp.symbols('theta2 alpha2 beta2')\nbob = U(theta=theta2, alpha=alpha2, beta=beta2)\nbob",
"_____no_output_____"
],
[
"payoff_matrix = Array([\n [\n [3, 5],\n [0, 1],\n ],\n [\n [3, 0],\n [5, 1],\n ],\n])\npayoff_matrix",
"_____no_output_____"
],
[
"p_error = 0.05\nbit_flip = pauli_error([('X', p_error), ('I', 1 - p_error)])\nphase_flip = pauli_error([('Z', p_error), ('I', 1 - p_error)])",
"_____no_output_____"
],
[
"noise_model = NoiseModel()\nnoise_model.add_all_qubit_quantum_error(bit_flip, ['u1', 'u2', 'u3'])\nnoise_model.add_all_qubit_quantum_error(phase_flip, ['x'], [0])\nnoise_model",
"_____no_output_____"
],
[
"ewl = EWL(psi, [alice, bob], payoff_matrix, noise_model)\newl",
"_____no_output_____"
],
[
"ewl_fixed = ewl.fix(theta1=pi / 2, alpha1=pi / 2, beta1=0,\n theta2=0, alpha2=0, beta2=0)",
"_____no_output_____"
],
[
"counts_qasm_simulator = ewl_fixed.simulate_counts()\nprint(counts_qasm_simulator)\nplot_histogram(counts_qasm_simulator)\nNone",
"{'00': 92, '10': 103, '01': 432, '11': 397}\n"
],
[
"counts_quantum_device = ewl_fixed.run()\nprint(counts_quantum_device)\nplot_histogram(counts_quantum_device)\nNone",
"Job Status: job has successfully run\n{'00': 59, '01': 549, '10': 34, '11': 382}\n"
],
[
"plot_histogram(data=[counts_qasm_simulator, counts_quantum_device],\n legend=['qasm simulator', 'quantum device'])\nNone",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acccc47a271c11da3ab14d4c6644706902d3932
| 881,936 |
ipynb
|
Jupyter Notebook
|
src/tdr_gating_example.ipynb
|
EA4FRB/sark110-python
|
ff5ace7c0d71f9de48772cdb1cedb98f5c22df8a
|
[
"MIT"
] | 2 |
2019-09-03T18:43:53.000Z
|
2019-11-06T18:26:23.000Z
|
src/tdr_gating_example.ipynb
|
EA4FRB/sark110-python
|
ff5ace7c0d71f9de48772cdb1cedb98f5c22df8a
|
[
"MIT"
] | 1 |
2020-04-03T22:59:18.000Z
|
2020-04-29T16:17:02.000Z
|
src/tdr_gating_example.ipynb
|
EA4FRB/sark110-python
|
ff5ace7c0d71f9de48772cdb1cedb98f5c22df8a
|
[
"MIT"
] | 1 |
2019-09-05T16:40:55.000Z
|
2019-09-05T16:40:55.000Z
| 993.171171 | 76,793 | 0.778239 |
[
[
[
"# SARK-110 Time Domain and Gating Example\n\nExample adapted from: https://scikit-rf.readthedocs.io/en/latest/examples/networktheory/Time%20Domain.html\n\n- Measurements with a 2.8m section of rg58 coax cable not terminated at the end \n\nThis notebooks demonstrates how to use scikit-rf for time-domain analysis and gating. A quick example is given first, followed by a more detailed explanation.\n\nS-parameters are measured in the frequency domain, but can be analyzed in time domain if you like. In many cases, measurements are not made down to DC. This implies that the time-domain transform is not complete, but it can be very useful non-theless. A major application of time-domain analysis is to use gating to isolate a single response in space. More information about the details of time domain analysis.\n\nPlease ensure that the analyzer is connected to the computer using the USB cable and in Computer Control mode.",
"_____no_output_____"
]
],
[
[
"from sark110 import *\nimport skrf as rf \nrf.stylely()\nfrom pylab import *",
"_____no_output_____"
]
],
[
[
"Enter frequency limits:",
"_____no_output_____"
]
],
[
[
"fr_start = 100000 # Frequency start in Hz\nfr_stop = 230000000 # Frequency stop in Hz\npoints = 401 # Number of points",
"_____no_output_____"
]
],
[
[
"## Utility functions",
"_____no_output_____"
]
],
[
[
"def z2vswr(rs: float, xs: float, z0=50 + 0j) -> float:\n gamma = math.sqrt((rs - z0.real) ** 2 + xs ** 2) / math.sqrt((rs + z0.real) ** 2 + xs ** 2)\n if gamma > 0.980197824:\n return 99.999\n swr = (1 + gamma) / (1 - gamma)\n return swr\n\n\ndef z2mag(r: float, x: float) -> float:\n return math.sqrt(r ** 2 + x ** 2)\n\n\ndef z2gamma(rs: float, xs: float, z0=50 + 0j) -> complex:\n z = complex(rs, xs)\n return (z - z0) / (z + z0)",
"_____no_output_____"
]
],
[
[
"## Connect to the device",
"_____no_output_____"
]
],
[
[
"sark110 = Sark110()\nsark110.open()\nsark110.connect()\nif not sark110.is_connected:\n print(\"Device not connected\")\n exit(-1)\nelse:\n print(\"Device connected\")",
"Device connected\n"
],
[
"sark110.buzzer()\nprint(sark110.fw_protocol, sark110.fw_version)",
"2566 1.5\n"
]
],
[
[
"## Acquire and plot the data",
"_____no_output_____"
]
],
[
[
"y = []\nx = []\nrs = [0]\nxs = [0]\nfor i in range(points):\n fr = int(fr_start + i * (fr_stop - fr_start) / (points - 1))\n sark110.measure(fr, rs, xs)\n x.append(fr / 1e9) # Units in GHz\n y.append(z2gamma(rs[0][0], xs[0][0]))\n\nprobe = rf.Network(frequency=x, s=y, z0=50)\nprobe.frequency.unit = 'mhz'\nprint (probe)",
"1-Port Network: '', 0.1-230.0 MHz, 401 pts, z0=[50.+0.j]\n"
]
],
[
[
"# Quick example",
"_____no_output_____"
]
],
[
[
"# we will focus on s11\ns11 = probe.s11\n\n# time-gate the first largest reflection\ns11_gated = s11.time_gate(center=0, span=50)\ns11_gated.name='gated probe'\n\n# plot frequency and time-domain s-parameters\nfigure(figsize=(8,4))\nsubplot(121)\ns11.plot_s_db()\ns11_gated.plot_s_db()\ntitle('Frequency Domain')\n\nsubplot(122)\ns11.plot_s_db_time()\ns11_gated.plot_s_db_time()\ntitle('Time Domain')\ntight_layout()",
"_____no_output_____"
]
],
[
[
"# Interpreting Time Domain\n\nNote there are two time-domain plotting functions in scikit-rf:\n\n- Network.plot_s_db_time()\n- Network.plot_s_time_db()\n\nThe difference is that the former, plot_s_db_time(), employs windowing before plotting to enhance impluse resolution. Windowing will be discussed in a bit, but for now we just use plot_s_db_time().\n\nPlotting all four s-parameters of the probe in both frequency and time-domain.",
"_____no_output_____"
]
],
[
[
"# plot frequency and time-domain s-parameters\nfigure(figsize=(8,4))\nsubplot(121)\nprobe.plot_s_db()\ntitle('Frequency Domain')\nsubplot(122)\nprobe.plot_s_db_time()\ntitle('Time Domain')\ntight_layout()",
"_____no_output_____"
]
],
[
[
"Focusing on the reflection coefficient from the waveguide port (s11), you can see there is an interference pattern present.",
"_____no_output_____"
]
],
[
[
"probe.plot_s_db(0,0)\ntitle('Reflection Coefficient From \\nWaveguide Port')",
"_____no_output_____"
]
],
[
[
"This ripple is evidence of several discrete reflections. Plotting s11 in the time-domain allows us to see where, or when, these reflections occur.",
"_____no_output_____"
]
],
[
[
"probe_s11 = probe.s11\nprobe_s11.plot_s_db_time(0,0)\ntitle('Reflection Coefficient From \\nWaveguide Port, Time Domain')\nylim(-100,0)",
"_____no_output_____"
]
],
[
[
"# Gating The Reflection of Interest\nTo isolate the reflection from the waveguide port, we can use time-gating. This can be done by using the method Network.time_gate(), and provide it an appropriate center and span (in ns). To see the effects of the gate, both the original and gated reponse are compared.",
"_____no_output_____"
]
],
[
[
"probe_s11_gated = probe_s11.time_gate(center=0, span=50)\nprobe_s11_gated.name='gated probe'\n\ns11.plot_s_db_time()\ns11_gated.plot_s_db_time()",
"_____no_output_____"
]
],
[
[
"Next, compare both responses in frequency domain to see the effect of the gate.",
"_____no_output_____"
]
],
[
[
"s11.plot_s_db()\ns11_gated.plot_s_db()",
"_____no_output_____"
]
],
[
[
"# Auto-gate\n\nThe time-gating method in skrf has an auto-gating feature which can also be used to gate the largest reflection. When no gate parameters are provided, time_gate() does the following:\n\n find the two largest peaks\n center the gate on the tallest peak\n set span to distance between two tallest peaks\n\nYou may want to plot the gated network in time-domain to see what the determined gate shape looks like.",
"_____no_output_____"
]
],
[
[
"title('Waveguide Interface of Probe') \ns11.plot_s_db(label='original')\ns11.time_gate().plot_s_db(label='autogated') #autogate on the fly",
"_____no_output_____"
]
],
[
[
"# Determining Distance\n\nTo make time-domain useful as a diagnostic tool, one would like to convert the x-axis to distance. This requires knowledge of the propagation velocity in the device. skrf provides some transmission-line models in the module skrf.media, which can be used for this.\n\nHowever...\n\nFor dispersive media, such as rectangular waveguide, the phase velocity is a function of frequency, and transforming time to distance is not straightforward. As an approximation, you can normalize the x-axis to the speed of light.\n\nAlternatively, you can simulate the a known device and compare the two time domain responses. This allows you to attribute quantatative meaning to the axes. For example, you could create an ideal delayed load as shown below. Note: the magnitude of a response behind a large impulse doesn not have meaningful units.\n",
"_____no_output_____"
]
],
[
[
"from skrf.media import DistributedCircuit\n\n# create a Media object for RG-58, based on distributed ckt values\nrg58 = DistributedCircuit(\n frequency = probe.frequency,\n C =93.5e-12,#F/m\n L =273e-9, #H/m\n R =0, #53e-3, #Ohm/m\n G =0, #S/m\n )\n\n# create an ideal delayed load, parameters are adjusted until the\n# theoretical response agrees with the measurement\ntheory = rg58.delay_load(Gamma0=rf.db_2_mag(-20),\n d=280, unit='cm')\nprobe.plot_s_db_time(0,0, label = 'Measurement')\ntheory.plot_s_db_time(label='-20dB @ 280cm from test-port')\nylim(-100,0)\nxlim(-500,500)",
"_____no_output_____"
]
],
[
[
"This plot demonstrates a few important points:\n\n the theortical delayed load is not a perfect impulse in time. This is due to the dispersion in waveguide.\n the peak of the magnitude in time domain is not identical to that specified, also due to disperison (and windowing).\n\n# What the hell is Windowing?\n\nThe 'plot_s_db_time()' function does a few things.\n\n windows the s-parameters.\n converts to time domain\n takes magnitude component, convert to dB\n calculates time-axis s\n plots\n\nA word about step 1: windowing. A FFT represents a signal with a basis of periodic signals (sinusoids). If your frequency response is not periodic, which in general it isnt, taking a FFT will introduces artifacts in the time-domain results. To minimize these effects, the frequency response is windowed. This makes the frequency response more periodic by tapering off the band-edges.\n\nWindowing is just applied to improve the plot appearance,d it does not affect the original network.\n\nIn skrf this can be done explicitly using the 'windowed()' function. By default this function uses the hamming window, but can be adjusted through arguments. The result of windowing is show below.\n",
"_____no_output_____"
]
],
[
[
"probe_w = probe.windowed()\nprobe.plot_s_db(0,0, label = 'Original')\nprobe_w.plot_s_db(0,0, label = 'Windowed')",
"_____no_output_____"
]
],
[
[
"Comparing the two time-domain plotting functions, we can see the difference between windowed and not.",
"_____no_output_____"
]
],
[
[
"probe.plot_s_time_db(0,0, label = 'Original')\nprobe_w.plot_s_time_db(0,0, label = 'Windowed')",
"_____no_output_____"
]
],
[
[
"# The end!",
"_____no_output_____"
]
],
[
[
"sark110.close()",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4accd01112eaee71292e6b2c7361b3e7908cb8cf
| 25,655 |
ipynb
|
Jupyter Notebook
|
scripts/first_figur_in_notebook.ipynb
|
JamesPMColeman/Data-Delight
|
04249be65650a1c6fa4720e74e5ace60844e3741
|
[
"MIT"
] | null | null | null |
scripts/first_figur_in_notebook.ipynb
|
JamesPMColeman/Data-Delight
|
04249be65650a1c6fa4720e74e5ace60844e3741
|
[
"MIT"
] | null | null | null |
scripts/first_figur_in_notebook.ipynb
|
JamesPMColeman/Data-Delight
|
04249be65650a1c6fa4720e74e5ace60844e3741
|
[
"MIT"
] | null | null | null | 59.249423 | 6,005 | 0.519821 |
[
[
[
"# Bokeh Libraries\nfrom bokeh.io import output_notebook\nfrom bokeh.plotting import figure, show\n\n# The figure wil be rendered into a static HTML file\noutput_notebook()\n\n# Set up a generic figure() object\nfig = figure()\n\n# view the figure\nshow(fig)",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code"
]
] |
4accd2eb6c362c2528092c8cb0d0c254b8eff7ae
| 157,155 |
ipynb
|
Jupyter Notebook
|
notebooks/data_preprocessing/immune_cells/merging/MCA_BM_merging.ipynb
|
theislab/scIB-results
|
581c3e4c1f016ba901d25f13f8c4af4d96c24fa1
|
[
"MIT"
] | 1 |
2020-10-12T10:25:38.000Z
|
2020-10-12T10:25:38.000Z
|
notebooks/data_preprocessing/immune_cells/merging/MCA_BM_merging.ipynb
|
theislab/scIB-results
|
581c3e4c1f016ba901d25f13f8c4af4d96c24fa1
|
[
"MIT"
] | 1 |
2020-10-14T11:45:54.000Z
|
2020-10-14T14:45:39.000Z
|
notebooks/data_preprocessing/immune_cells/merging/MCA_BM_merging.ipynb
|
theislab/scIB-reproducibility
|
581c3e4c1f016ba901d25f13f8c4af4d96c24fa1
|
[
"MIT"
] | null | null | null | 130.095199 | 60,720 | 0.842843 |
[
[
[
"# Mouse Bone Marrow - merging annotated samples from MCA",
"_____no_output_____"
]
],
[
[
"import scanpy as sc\nimport numpy as np\nimport scipy as sp\nimport pandas as pd\nimport matplotlib.pyplot as plt\nfrom matplotlib import rcParams\nfrom matplotlib import colors\nimport seaborn as sb\nimport glob\n\nimport rpy2.rinterface_lib.callbacks\nimport logging\n\nfrom rpy2.robjects import pandas2ri\nimport anndata2ri",
"_____no_output_____"
],
[
"# Ignore R warning messages\n#Note: this can be commented out to get more verbose R output\nrpy2.rinterface_lib.callbacks.logger.setLevel(logging.ERROR)\n\n# Automatically convert rpy2 outputs to pandas dataframes\npandas2ri.activate()\nanndata2ri.activate()\n%load_ext rpy2.ipython\n\nplt.rcParams['figure.figsize']=(8,8) #rescale figures\nsc.settings.verbosity = 3\n#sc.set_figure_params(dpi=200, dpi_save=300)\nsc.logging.print_versions()\n\nresults_file = './write/MCA_mou_BM_pp.h5ad'",
"scanpy==1.4.4.post1 anndata==0.6.22.post1 umap==0.3.10 numpy==1.17.3 scipy==1.3.0 pandas==0.24.2 scikit-learn==0.21.2 statsmodels==0.10.1 python-igraph==0.7.1 louvain==0.6.1\n"
],
[
"%%R\n# Load all the R libraries we will be using in the notebook\nlibrary(scran)",
"_____no_output_____"
]
],
[
[
"## Load \nHere we load the pre-processed datasets (which has been annotated), and the raw matrices (which won't be filtered on the gene level). ",
"_____no_output_____"
],
[
"### Raw data",
"_____no_output_____"
]
],
[
[
"file_paths = '../../Munich/datasets/mouse/MCA_boneMarrow/ckit/'\nadatas_raw = []\nfor i in glob.glob(file_paths+'*.txt.gz'):\n print(i)\n adatas_raw.append(sc.read(i, cache=True)) ",
"../../Munich/datasets/mouse/MCA_boneMarrow/ckit/BoneMarrowcKit1_dge.txt.gz\n... reading from cache file cache/..-..-Munich-datasets-mouse-MCA_boneMarrow-ckit-BoneMarrowcKit1_dge.h5ad\n../../Munich/datasets/mouse/MCA_boneMarrow/ckit/BoneMarrowcKit3_dge.txt.gz\n... reading from cache file cache/..-..-Munich-datasets-mouse-MCA_boneMarrow-ckit-BoneMarrowcKit3_dge.h5ad\n../../Munich/datasets/mouse/MCA_boneMarrow/ckit/BoneMarrowcKit2_dge.txt.gz\n... reading from cache file cache/..-..-Munich-datasets-mouse-MCA_boneMarrow-ckit-BoneMarrowcKit2_dge.h5ad\n"
],
[
"samples = ['BM_1', 'BM_3', 'BM_2']\n# Loop to annotate data\nfor i in range(len(adatas_raw)):\n adata_tmp = adatas_raw[i]\n adata_tmp = adata_tmp.transpose()\n \n #Annotate data\n adata_tmp.obs.index.rename('barcode', inplace=True)\n adata_tmp.obs['batch'] = ['MCA_'+samples[i]]*adata_tmp.n_obs\n adata_tmp.obs['study'] = ['MCA_BM']*adata_tmp.n_obs\n adata_tmp.obs['chemistry'] = ['microwell-seq']*adata_tmp.n_obs\n adata_tmp.obs['tissue'] = ['Bone_Marrow']*adata_tmp.n_obs\n adata_tmp.obs['species'] = ['Mouse']*adata_tmp.n_obs\n adata_tmp.obs['data_type'] = ['UMI']*adata_tmp.n_obs\n\n adata_tmp.var.index.names = ['gene_symbol']\n adata_tmp.var_names_make_unique()\n adatas_raw[i] = adata_tmp\n",
"_____no_output_____"
],
[
"adatas_raw[0].obs.head()",
"_____no_output_____"
],
[
"# Concatenate to unique adata object\nadata_raw = adatas_raw[0].concatenate(adatas_raw[1:], batch_key='sample_ID', index_unique=None)",
"_____no_output_____"
],
[
"adata_raw.obs.head()",
"_____no_output_____"
],
[
"adata_raw.obs.drop(columns=['sample_ID'], inplace=True)\nadata_raw.obs.head()",
"_____no_output_____"
],
[
"adata_raw.shape",
"_____no_output_____"
]
],
[
[
"### Pre-processed data",
"_____no_output_____"
]
],
[
[
"file_paths = '../../Bone_Marrow_mouse/write/'\nadatas_pp = []\nfor i in glob.glob(file_paths+'*.h5ad'):\n print(i)\n adatas_pp.append(sc.read(i, cache=True)) ",
"../../Bone_Marrow_mouse/write/MCA_BM_1_ckit.h5ad\n../../Bone_Marrow_mouse/write/MCA_BM_3_ckit.h5ad\n../../Bone_Marrow_mouse/write/MCA_BM_2_ckit.h5ad\n"
],
[
"for i in range(len(adatas_pp)):\n adata_tmp = adatas_pp[i]\n adata_obs = adata_tmp.obs.reset_index()\n adata_obs = adata_obs[['index', 'final_annotation', 'dpt_pseudotime_y', 'n_counts', 'n_genes', 'mt_frac']].rename(columns = {'index':'barcode'})\n adata_obs.set_index('barcode', inplace = True)\n adatas_pp[i].obs = adata_obs\n ",
"_____no_output_____"
],
[
"# Concatenate to unique adata object\nadata_pp = adatas_pp[0].concatenate(adatas_pp[1:], batch_key='sample_ID',\n index_unique=None)",
"_____no_output_____"
],
[
"adata_pp.obs.drop(columns=['sample_ID'], inplace = True)\nadata_pp.obs.head()",
"_____no_output_____"
],
[
"adata_raw.shape",
"_____no_output_____"
],
[
"adata_pp.shape",
"_____no_output_____"
],
[
"# Restrict to cells that passed QC and were annotated\nadata_obs_raw = adata_raw.obs.reset_index()\nadata_obs_pp = adata_pp.obs.reset_index()\nadata_merged = adata_obs_raw.merge(adata_obs_pp, on='barcode', how='left')\nadata_merged.set_index('barcode', inplace = True)\nadata_raw.obs = adata_merged\nadata_raw.obs.head()",
"_____no_output_____"
],
[
"adata_raw = adata_raw[~pd.isnull(adata_raw.obs['final_annotation'])]\nadata_raw.shape",
"_____no_output_____"
]
],
[
[
"### Normalization",
"_____no_output_____"
]
],
[
[
"# Exclude genes that are = 0 in all cells\n#Filter genes:\nprint('Total number of genes: {:d}'.format(adata_raw.n_vars))\n\n# Min 20 cells - filters out 0 count genes\nsc.pp.filter_genes(adata_raw, min_cells=1)\nprint('Number of genes after cell filter: {:d}'.format(adata_raw.n_vars))",
"Total number of genes: 15455\n"
],
[
"#Perform a clustering for scran normalization in clusters\nadata_pp = adata_raw.copy()\nsc.pp.normalize_per_cell(adata_pp, counts_per_cell_after=1e6)\nsc.pp.log1p(adata_pp)\nsc.pp.pca(adata_pp, n_comps=15, svd_solver='arpack')\nsc.pp.neighbors(adata_pp)\nsc.tl.louvain(adata_pp, key_added='groups', resolution=0.5)",
"normalizing by total count per cell\n finished (0:00:01): normalized adata.X and added 'n_counts', counts per cell before normalization (adata.obs)\ncomputing PCA with n_comps = 15\n finished (0:00:09)\ncomputing neighbors\n using 'X_pca' with n_pcs = 15\n finished: added to `.uns['neighbors']`\n 'distances', distances for each pair of neighbors\n 'connectivities', weighted adjacency matrix (0:00:08)\nrunning Louvain clustering\n using the \"louvain\" package of Traag (2017)\n finished: found 10 clusters and added\n 'groups', the cluster labels (adata.obs, categorical) (0:00:02)\n"
],
[
"# Check if the minimum number of cells per cluster is < 21:in that case, sizes will be also passed as input to the normalization\nadata_pp.obs['groups'].value_counts()",
"_____no_output_____"
],
[
"#Preprocess variables for scran normalization\ninput_groups = adata_pp.obs['groups']\ndata_mat = adata_raw.X.T",
"_____no_output_____"
],
[
"%%R -i data_mat -i input_groups -o size_factors\n\nsize_factors = computeSumFactors(data_mat, clusters=input_groups, min.mean=0.1)",
"_____no_output_____"
],
[
"#Delete adata_pp\ndel adata_pp",
"_____no_output_____"
],
[
"# Visualize the estimated size factors\nadata_raw.obs['size_factors'] = size_factors\n\nsc.pl.scatter(adata_raw, 'size_factors', 'n_counts')\nsc.pl.scatter(adata_raw, 'size_factors', 'n_genes')\n\nsb.distplot(size_factors, bins=50, kde=False)\nplt.show()",
"... storing 'batch' as categorical\n... storing 'chemistry' as categorical\n... storing 'data_type' as categorical\n... storing 'species' as categorical\n... storing 'study' as categorical\n... storing 'tissue' as categorical\n"
],
[
"#Keep the count data in a counts layer\nadata_raw.layers[\"counts\"] = adata_raw.X.copy()",
"_____no_output_____"
],
[
"#Normalize adata \nadata_raw.X /= adata_raw.obs['size_factors'].values[:,None]\nsc.pp.log1p(adata_raw)",
"_____no_output_____"
],
[
"adata_raw.write(results_file)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acce00596df324c6d13c2407edbebc992764c1b
| 375,388 |
ipynb
|
Jupyter Notebook
|
ImageProcessingScripts/Camera Based Thermal Screening/camera_based_thermal_screening.ipynb
|
rishabhjainfinal/Awesome_Python_Scripts
|
a1a65d9e1b9209da7c425a4a99d0e59d91fd1dec
|
[
"MIT"
] | 1 |
2022-01-20T10:49:30.000Z
|
2022-01-20T10:49:30.000Z
|
ImageProcessingScripts/Camera Based Thermal Screening/camera_based_thermal_screening.ipynb
|
rishabhjainfinal/Awesome_Python_Scripts
|
a1a65d9e1b9209da7c425a4a99d0e59d91fd1dec
|
[
"MIT"
] | null | null | null |
ImageProcessingScripts/Camera Based Thermal Screening/camera_based_thermal_screening.ipynb
|
rishabhjainfinal/Awesome_Python_Scripts
|
a1a65d9e1b9209da7c425a4a99d0e59d91fd1dec
|
[
"MIT"
] | 2 |
2021-10-03T16:22:08.000Z
|
2021-10-03T17:35:14.000Z
| 834.195556 | 72,904 | 0.955478 |
[
[
[
"# importing the required libraries",
"_____no_output_____"
],
[
"import os\nimport numpy as np\nimport cv2\nimport matplotlib.pyplot as plt\n\n%matplotlib inline",
"_____no_output_____"
],
[
"# function for reading the image \n# this image is taken from a video\n# and the video is taken from a thermal camera\n# converting image from BGR to RGB",
"_____no_output_____"
],
[
"def read_image(image_path):\n image = cv2.imread(image_path)\n return cv2.cvtColor(image, cv2.COLOR_BGR2RGB)",
"_____no_output_____"
],
[
"# thermal camera takes the heat/thermal energy\n# more heat means the pixel value is closer to 255\n# if it is cool then pixel value is closer to 0\n# displaying the image, where white portion means that part is having more temprature\n# and vice versa",
"_____no_output_____"
],
[
"image = read_image(\"thermal_scr_img.png\")\nplt.imshow(image)",
"_____no_output_____"
],
[
"# converting the image into grayscale\n# changing and applying the ColorMap, black and white to black and red",
"_____no_output_____"
],
[
"heatmap_gray = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)\nheatmap = cv2.applyColorMap(heatmap_gray, cv2.COLORMAP_HOT)\nheatmap = cv2.cvtColor(heatmap, cv2.COLOR_BGR2RGB)\n\nplt.imshow(heatmap)",
"_____no_output_____"
],
[
"# now taking the heatmap_gray and converting it to black and white image\n# and performing threshold operation\n# the pixels having values more than 200 will become white pixels and the one having values less than 200 will become black pixels",
"_____no_output_____"
],
[
"heatmap_gray = cv2.cvtColor(heatmap, cv2.COLOR_RGB2GRAY)\nret, binary_thresh = cv2.threshold(heatmap_gray, 200, 255, cv2.THRESH_BINARY)\nplt.imshow(binary_thresh, cmap='gray')",
"_____no_output_____"
],
[
"# then cleaning the small white pixels to calculate the temperature for bigger blocks/portions of the image\n# doing erosion operation by taking binary threshold (it makes image pixels thinner)\n# doing dilution operation by taking the image erosion (and then that's why we are removing/cleaning all small pixels)\n# kernel is some kind of filter and it changes the values of these pixels ",
"_____no_output_____"
],
[
"kernel = np.ones((5,5), np.uint8)\nimage_erosion = cv2.erode(binary_thresh, kernel, iterations=1)\nimage_opening = cv2.dilate(image_erosion, kernel, iterations=1)\n\nplt.imshow(image_opening, cmap='gray')",
"_____no_output_____"
],
[
"# now creating some masks\n# using function zeros_like() it will take all the structures like zero\n# x, y, w, h are the coordinate for rectangle\n# copying the small rectangle part from this image using mask\n# and printing the avg. value of pixels to get the temperature",
"_____no_output_____"
],
[
"contours, _ = cv2.findContours(image_opening, 1, 2)\n\ncontour = contours[11]\nmask = np.zeros_like(heatmap_gray)\nx, y, w, h = cv2.boundingRect(contour)\nmask[y:y+h, x:x+w] = image_opening[y:y+h, x:x+w]\nprint(cv2.mean(heatmap_gray, mask= mask))\nplt.imshow(mask, cmap='gray')",
"(234.61660079051381, 0.0, 0.0, 0.0)\n"
],
[
"# performing the bitwise and operator on heatmap\n# here we have created not mask",
"_____no_output_____"
],
[
"masked = cv2.bitwise_and(heatmap, heatmap, mask=~mask)\nplt.imshow(masked)",
"_____no_output_____"
],
[
"# displaying the heatmap_gray",
"_____no_output_____"
],
[
"plt.imshow(heatmap_gray)",
"_____no_output_____"
],
[
"image_with_rectangles = np.copy(heatmap)\n\nfor contour in contours:\n # rectangle over each contour\n x, y, w, h = cv2.boundingRect(contour)\n \n # mask is boolean type of matrix\n mask = np.zeros_like(heatmap_gray)\n mask[y:y+h, x:x+w] = image_opening[y:y+h, x:x+w]\n \n # temperature calculation\n temp = round(cv2.mean(heatmap_gray, mask=mask)[0] / 2.25, 2)\n \n # draw rectangles for visualisation\n image_with_rectangles = cv2.rectangle(\n image_with_rectangles, (x,y), (x+w, y+h), (0, 255, 0), 2)\n \n # write temperature for each rectangle\n cv2.putText(image_with_rectangles, f\"{temp} F\", (x,y),\n cv2.FONT_HERSHEY_SIMPLEX, 0.7, (0, 255, 0), 2, cv2.LINE_AA)\n \n plt.imshow(image_with_rectangles)",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acce9434318a396246372cd87e859d68bc527c4
| 5,598 |
ipynb
|
Jupyter Notebook
|
05_conv_layer/tf_conv_layer.ipynb
|
JiaheXu/Machine-Learning-Codebase
|
0280fbc0a048133d71fa1f8129c147d267fefc3e
|
[
"MIT"
] | null | null | null |
05_conv_layer/tf_conv_layer.ipynb
|
JiaheXu/Machine-Learning-Codebase
|
0280fbc0a048133d71fa1f8129c147d267fefc3e
|
[
"MIT"
] | null | null | null |
05_conv_layer/tf_conv_layer.ipynb
|
JiaheXu/Machine-Learning-Codebase
|
0280fbc0a048133d71fa1f8129c147d267fefc3e
|
[
"MIT"
] | null | null | null | 22.214286 | 132 | 0.479993 |
[
[
[
"import tensorflow as tf",
"_____no_output_____"
]
],
[
[
"## Conv2d(卷积核)\ntf.keras.layers.Conv2D( \n filters, kernel_size, strides=(1, 1), padding='valid', \n data_format=None, dilation_rate=(1, 1), groups=1, activation=None, \n use_bias=True, kernel_initializer='glorot_uniform', \n bias_initializer='zeros', kernel_regularizer=None, \n bias_regularizer=None, activity_regularizer=None, kernel_constraint=None, \n bias_constraint=None, **kwargs \n)",
"_____no_output_____"
]
],
[
[
"filters = 1",
"_____no_output_____"
],
[
"# 构造一个二维卷积层,它具有1个输出通道和形状为(1,2)的卷积核\nconv2d = tf.keras.layers.Conv2D(filters, (1, 2), use_bias=False)",
"_____no_output_____"
],
[
"## Padding\n## 'same': keep the size the same",
"_____no_output_____"
],
[
"# 注意!!! 这里每边都填充了1行或1列,因此总共添加了2行或2列\nconv2d = tf.keras.layers.Conv2D(filters, kernel_size=3, padding='same')",
"_____no_output_____"
],
[
"conv2d = tf.keras.layers.Conv2D(filters, kernel_size=(5, 3), padding='same')",
"_____no_output_____"
]
],
[
[
"## stride",
"_____no_output_____"
]
],
[
[
"conv2d = tf.keras.layers.Conv2D(filters, kernel_size=3, padding='same', strides=2)",
"_____no_output_____"
],
[
"conv2d = tf.keras.layers.Conv2D(filters, kernel_size=(3,5), padding='valid',strides=(3, 4))",
"_____no_output_____"
]
],
[
[
"## Pooling layer(汇聚层)\ntf.keras.layers.MaxPool2D( \n pool_size=(2, 2), \n strides=None, \n padding='valid', \n data_format=None, \n)",
"_____no_output_____"
]
],
[
[
"X = tf.reshape(tf.range(16, dtype=tf.float32), (1, 4, 4, 1))\nX",
"_____no_output_____"
]
],
[
[
"### 默认情况下,步幅与汇聚窗口的大小相同。 因此,如果我们使用形状为(3, 3)的汇聚窗口,那么默认情况下,我们得到的步幅形状为(3, 3)。",
"_____no_output_____"
]
],
[
[
"pool2d = tf.keras.layers.MaxPool2D(pool_size=[3, 3])\npool2d(X)",
"_____no_output_____"
],
[
"pool2d = tf.keras.layers.MaxPool2D(pool_size=[3, 3], padding='valid',\n strides=2)",
"_____no_output_____"
],
[
"pool2d = tf.keras.layers.MaxPool2D(pool_size=[2, 3], padding='valid',\n strides=(2, 3))",
"_____no_output_____"
]
],
[
[
"# Batch Normalization",
"_____no_output_____"
]
],
[
[
"bn = tf.keras.layers.BatchNormalization()",
"_____no_output_____"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
]
] |
4accebaa6a67a32f35f2e2742229f7dd0b6ce8c9
| 581 |
ipynb
|
Jupyter Notebook
|
notebooks/plot_avg_systemsize.ipynb
|
vwcruzeiro/ANI-Tools
|
bc44c65589148b6e6dd8065d63e73ead00d8ac9d
|
[
"MIT"
] | 19 |
2018-05-09T08:17:44.000Z
|
2021-12-26T07:02:17.000Z
|
notebooks/plot_avg_systemsize.ipynb
|
vwcruzeiro/ANI-Tools
|
bc44c65589148b6e6dd8065d63e73ead00d8ac9d
|
[
"MIT"
] | null | null | null |
notebooks/plot_avg_systemsize.ipynb
|
vwcruzeiro/ANI-Tools
|
bc44c65589148b6e6dd8065d63e73ead00d8ac9d
|
[
"MIT"
] | 11 |
2017-12-15T13:39:35.000Z
|
2021-05-15T15:06:02.000Z
| 16.6 | 34 | 0.524957 |
[] |
[] |
[] |
4accf8aded517b37dbf7abf5ceecada224e8f828
| 18,971 |
ipynb
|
Jupyter Notebook
|
Scratch/Notebooks/Compare Processing.ipynb
|
brendenpelkie/OpenAvalancheProject
|
0b35c850a5fa9376a639c6607762325cca598600
|
[
"MIT"
] | 52 |
2018-03-06T22:58:35.000Z
|
2022-02-21T15:42:02.000Z
|
Scratch/Notebooks/Compare Processing.ipynb
|
brendenpelkie/OpenAvalancheProject
|
0b35c850a5fa9376a639c6607762325cca598600
|
[
"MIT"
] | 28 |
2017-12-04T16:04:32.000Z
|
2022-02-28T20:51:28.000Z
|
Scratch/Notebooks/Compare Processing.ipynb
|
brendenpelkie/OpenAvalancheProject
|
0b35c850a5fa9376a639c6607762325cca598600
|
[
"MIT"
] | 38 |
2018-03-07T06:04:07.000Z
|
2021-12-25T20:14:08.000Z
| 32.993043 | 169 | 0.441358 |
[
[
[
"from openavalancheproject.prep_ml import PrepML",
"_____no_output_____"
],
[
"\ninterpolate = 1 #interpolation factor: whether we can to augment the data through lat/lon interpolation; 1 no interpolation, 4 is 4x interpolation\n\ndata_root = '/media/scottcha/E1/Data/OAPMLData/'\n",
"_____no_output_____"
],
[
"pml = PrepML(data_root, interpolate, date_start='2015-11-01', date_end='2017-04-30', date_train_test_cutoff='2016-11-01')",
"_____no_output_____"
],
[
"pml.regions = { \n 'Washington': ['Mt Hood', 'Olympics', 'Snoqualmie Pass', 'Stevens Pass',\n 'WA Cascades East, Central', 'WA Cascades East, North', 'WA Cascades East, South',\n 'WA Cascades West, Central', 'WA Cascades West, Mt Baker', 'WA Cascades West, South'\n ]}",
"_____no_output_____"
],
[
"%time train_labels, test_labels = pml.prep_labels()",
"Mt Hood\nOlympics\nSnoqualmie Pass\nStevens Pass\nWA Cascades East, Central\nWA Cascades East, North\nWA Cascades East, South\nWA Cascades West, Central\nWA Cascades West, Mt Baker\nWA Cascades West, South\nCPU times: user 19.2 s, sys: 703 ms, total: 19.9 s\nWall time: 20.2 s\n"
],
[
"train_labels.loc[0, 'UnifiedRegion']",
"_____no_output_____"
],
[
"train_labels.head()",
"_____no_output_____"
],
[
"d1 = pml.get_data_zarr(train_labels.loc[0,'UnifiedRegion'], train_labels.loc[0,'latitude'], train_labels.loc[0,'longitude'], 7, train_labels.loc[0,'parsed_date'])",
"_____no_output_____"
],
[
"d1",
"_____no_output_____"
],
[
"d2 = pml.get_data_zarr(test_labels.loc[0,'UnifiedRegion'], test_labels.loc[0,'latitude'], test_labels.loc[0,'longitude'], 7, test_labels.loc[0,'parsed_date'])",
"_____no_output_____"
],
[
"d2",
"_____no_output_____"
],
[
"pml2 = PrepML('/media/scottcha/E1/Data/OAPMLDataOld/', interpolate, date_start='2015-11-01', date_end='2017-04-30', date_train_test_cutoff='2016-11-01')",
"_____no_output_____"
],
[
"pml2.regions = { \n 'Washington': ['Mt Hood', 'Olympics', 'Snoqualmie Pass', 'Stevens Pass',\n 'WA Cascades East, Central', 'WA Cascades East, North', 'WA Cascades East, South',\n 'WA Cascades West, Central', 'WA Cascades West, Mt Baker', 'WA Cascades West, South'\n ]}",
"_____no_output_____"
],
[
"d3 = pml2.get_data_zarr(train_labels.loc[0,'UnifiedRegion'], train_labels.loc[0,'latitude'], train_labels.loc[0,'longitude'], 7, train_labels.loc[0,'parsed_date'])",
"_____no_output_____"
],
[
"d3",
"_____no_output_____"
],
[
"d4 = pml2.get_data_zarr(test_labels.loc[0,'UnifiedRegion'], test_labels.loc[0,'latitude'], test_labels.loc[0,'longitude'], 7, test_labels.loc[0,'parsed_date'])",
"_____no_output_____"
],
[
"d4",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acd0853e2e6d4e49588d76920c06a993639953e
| 367,418 |
ipynb
|
Jupyter Notebook
|
1_4_Feature_Vectors/3_1. HOG.ipynb
|
SDCND/CVND_Exercises
|
da2d3fce17dc5de3ec458fa71825c436f17e877f
|
[
"MIT"
] | null | null | null |
1_4_Feature_Vectors/3_1. HOG.ipynb
|
SDCND/CVND_Exercises
|
da2d3fce17dc5de3ec458fa71825c436f17e877f
|
[
"MIT"
] | null | null | null |
1_4_Feature_Vectors/3_1. HOG.ipynb
|
SDCND/CVND_Exercises
|
da2d3fce17dc5de3ec458fa71825c436f17e877f
|
[
"MIT"
] | null | null | null | 164.687584 | 238,449 | 0.840615 |
[
[
[
"## Histograms of Oriented Gradients (HOG)\n\nAs we saw with the ORB algorithm, we can use keypoints in images to do keypoint-based matching to detect objects in images. These type of algorithms work great when you want to detect objects that have a lot of consistent internal features that are not affected by the background. For example, these algorithms work well for facial detection because faces have a lot of consistent internal features that don’t get affected by the image background, such as the eyes, nose, and mouth. However, these type of algorithms don’t work so well when attempting to do more general object recognition, say for example, pedestrian detection in images. The reason is that people don’t have consistent internal features, like faces do, because the body shape and style of every person is different (see Fig. 1). This means that every person is going to have a different set of internal features, and so we need something that can more generally describe a person. \n\n<br>\n<figure>\n <img src = \"./in_cell_images/pedestrians.jpeg\" width = \"100%\" style = \"border: thin silver solid; padding: 10px\">\n <figcaption style = \"text-align:left; font-style:italic\">Fig. 1. - Pedestrians.</figcaption>\n</figure> \n<br>\n\nOne option is to try to detect pedestrians by their contours instead. Detecting objects in images by their contours (boundaries) is very challenging because we have to deal with the difficulties brought about by the contrast between the background and the foreground. For example, suppose you wanted to detect a pedestrian in an image that is walking in front of a white building and she is wearing a white coat and black pants (see Fig. 2). We can see in Fig. 2, that since the background of the image is mostly white, the black pants are going to have a very high contrast, but the coat, since it is white as well, is going to have very low contrast. In this case, detecting the edges of pants is going to be easy but detecting the edges of the coat is going to be very difficult. This is where **HOG** comes in. HOG stands for **Histograms of Oriented Gradients** and it was first introduced by Navneet Dalal and Bill Triggs in 2005.\n\n<br>\n<figure>\n <img src = \"./in_cell_images/woman.jpg\" width = \"100%\" style = \"border: thin silver solid; padding: 10px\">\n <figcaption style = \"text-align:left; font-style:italic\">Fig. 2. - High and Low Contrast.</figcaption>\n</figure> \n<br>\n\nThe HOG algorithm works by creating histograms of the distribution of gradient orientations in an image and then normalizing them in a very special way. This special normalization is what makes HOG so effective at detecting the edges of objects even in cases where the contrast is very low. These normalized histograms are put together into a feature vector, known as the HOG descriptor, that can be used to train a machine learning algorithm, such as a Support Vector Machine (SVM), to detect objects in images based on their boundaries (edges). Due to its great success and reliability, HOG has become one of the most widely used algorithms in computer vison for object detection.\n\n\nIn this notebook, you will learn:\n\n* How the HOG algorithm works\n* How to use OpenCV to create a HOG descriptor\n* How to visualize the HOG descriptor. ",
"_____no_output_____"
],
[
"# The HOG Algorithm\n\nAs its name suggests, the HOG algorithm, is based on creating histograms from the orientation of image gradients. The HOG algorithm is implemented in a series of steps:\n\n1. Given the image of particular object, set a detection window (region of interest) that covers the entire object in the image (see Fig. 3).\n\n2. Calculate the magnitude and direction of the gradient for each individual pixel in the detection window.\n\n3. Divide the detection window into connected *cells* of pixels, with all cells being of the same size (see Fig. 3). The size of the cells is a free parameter and it is usually chosen so as to match the scale of the features that want to be detected. For example, in a 64 x 128 pixel detection window, square cells 6 to 8 pixels wide are suitable for detecting human limbs.\n\n4. Create a Histogram for each cell, by first grouping the gradient directions of all pixels in each cell into a particular number of orientation (angular) bins; and then adding up the gradient magnitudes of the gradients in each angular bin (see Fig. 3). The number of bins in the histogram is a free parameter and it is usually set to 9 angular bins.\n\n5. Group adjacent cells into *blocks* (see Fig. 3). The number of cells in each block is a free parameter and all blocks must be of the same size. The distance between each block (known as the stride) is a free parameter but it is usually set to half the block size, in which case you will get overlapping blocks (*see video below*). The HOG algorithm has been shown empirically to work better with overlapping blocks.\n\n6. Use the cells contained within each block to normalize the cell histograms in that block (see Fig. 3). If you have overlapping blocks this means that most cells will be normalized with respect to different blocks (*see video below*). Therefore, the same cell may have several different normalizations.\n\n7. Collect all the normalized histograms from all the blocks into a single feature vector called the HOG descriptor.\n\n8. Use the resulting HOG descriptors from many images of the same type of object to train a machine learning algorithm, such as an SVM, to detect those type of objects in images. For example, you could use the HOG descriptors from many images of pedestrians to train an SVM to detect pedestrians in images. The training is done with both positive a negative examples of the object you want detect in the image.\n\n9. Once the SVM has been trained, a sliding window approach is used to try to detect and locate objects in images. Detecting an object in the image entails finding the part of the image that looks similar to the HOG pattern learned by the SVM.\n\n<br>\n<figure>\n <img src = \"./in_cell_images/HOG Diagram2.png\" width = \"100%\" style = \"border: thin silver solid; padding: 1px\">\n <figcaption style = \"text-align:left; font-style:italic\">Fig. 3. - HOG Diagram.</figcaption>\n</figure> \n<br>\n\n<figure>\n<video src = \"./in_cell_images/HOG Animation - Medium.mp4\" width=\"100%\" controls autoplay loop> </video>\n<figcaption style = \"text-align:left; font-style:italic\">Vid. 1. - HOG Animation.</figcaption>\n</figure> ",
"_____no_output_____"
],
[
"# Why The HOG Algorithm Works\n\nAs we learned above, HOG creates histograms by adding the magnitude of the gradients in particular orientations in localized portions of the image called *cells*. By doing this we guarantee that stronger gradients will contribute more to the magnitude of their respective angular bin, while the effects of weak and randomly oriented gradients resulting from noise are minimized. In this manner the histograms tell us the dominant gradient orientation of each cell. \n\n\n### Dealing with contrast \n\nNow, the magnitude of the dominant orientation can vary widely due to variations in local illumination and the contrast between the background and the foreground.\n\nTo account for the background-foreground contrast differences, the HOG algorithm tries to detect edges locally. In order to do this, it defines groups of cells, called **blocks**, and normalizes the histograms using this local group of cells. By normalizing locally, the HOG algorithm can detect the edges in each block very reliably; this is called **block normalization**.\n\nIn addition to using block normalization, the HOG algorithm also uses overlapping blocks to increase its performance. By using overlapping blocks, each cell contributes several independent components to the final HOG descriptor, where each component corresponds to a cell being normalized with respect to a different block. This may seem redundant but, it has been shown empirically that by normalizing each cell several times with respect to different local blocks, the performance of the HOG algorithm increases dramatically. ",
"_____no_output_____"
],
[
"### Loading Images and Importing Resources\n\nThe first step in building our HOG descriptor is to load the required packages into Python and to load our image. \n\nWe start by using OpenCV to load an image of a triangle tile. Since, the `cv2.imread()` function loads images as BGR we will convert our image to RGB so we can display it with the correct colors. As usual we will convert our BGR image to Gray Scale for analysis.",
"_____no_output_____"
]
],
[
[
"import cv2\nimport numpy as np\nimport matplotlib.pyplot as plt\n\n\n# Set the default figure size\nplt.rcParams['figure.figsize'] = [17.0, 7.0]\n\n# Load the image \nimage = cv2.imread('./images/triangle_tile.jpeg')\n\n# Convert the original image to RGB\noriginal_image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)\n\n# Convert the original image to gray scale\ngray_image = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)\n\n# Print the shape of the original and gray scale images\nprint('The original image has shape: ', original_image.shape)\nprint('The gray scale image has shape: ', gray_image.shape)\n\n# Display the images\nplt.subplot(121)\nplt.imshow(original_image)\nplt.title('Original Image')\nplt.subplot(122)\nplt.imshow(gray_image, cmap='gray')\nplt.title('Gray Scale Image')\nplt.show()",
"The original image has shape: (250, 250, 3)\nThe gray scale image has shape: (250, 250)\n"
]
],
[
[
"# Creating The HOG Descriptor\n\nWe will be using OpenCV’s `HOGDescriptor` class to create the HOG descriptor. The parameters of the HOG descriptor are setup using the `HOGDescriptor()` function. The parameters of the `HOGDescriptor()` function and their default values are given below:\n\n`cv2.HOGDescriptor(win_size = (64, 128), \n block_size = (16, 16), \n block_stride = (8, 8), \n cell_size = (8, 8), \n nbins = 9, \n win_sigma = DEFAULT_WIN_SIGMA, \n threshold_L2hys = 0.2, \n gamma_correction = true, \n nlevels = DEFAULT_NLEVELS)`\n\nParameters:\n\n* **win_size** – *Size* \nSize of detection window in pixels (*width, height*). Defines the region of interest. Must be an integer multiple of cell size.\n\n\n* **block_size** – *Size* \nBlock size in pixels (*width, height*). Defines how many cells are in each block. Must be an integer multiple of cell size and it must be smaller than the detection window. The smaller the block the finer detail you will get.\n\n\n* **block_stride** – *Size* \nBlock stride in pixels (*horizontal, vertical*). It must be an integer multiple of cell size. The `block_stride` defines the distance between adjecent blocks, for example, 8 pixels horizontally and 8 pixels vertically. Longer `block_strides` makes the algorithm run faster (because less blocks are evaluated) but the algorithm may not perform as well.\n\n\n* **cell_size** – *Size* \nCell size in pixels (*width, height*). Determines the size fo your cell. The smaller the cell the finer detail you will get.\n\n\n* **nbins** – *int* \nNumber of bins for the histograms. Determines the number of angular bins used to make the histograms. With more bins you capture more gradient directions. HOG uses unsigned gradients, so the angular bins will have values between 0 and 180 degrees.\n\n\n* **win_sigma** – *double* \nGaussian smoothing window parameter. The performance of the HOG algorithm can be improved by smoothing the pixels near the edges of the blocks by applying a Gaussian spatial window to each pixel before computing the histograms.\n\n\n* **threshold_L2hys** – *double* \nL2-Hys (Lowe-style clipped L2 norm) normalization method shrinkage. The L2-Hys method is used to normalize the blocks and it consists of an L2-norm followed by clipping and a renormalization. The clipping limits the maximum value of the descriptor vector for each block to have the value of the given threshold (0.2 by default). After the clipping the descriptor vector is renormalized as described in *IJCV*, 60(2):91-110, 2004.\n\n\n* **gamma_correction** – *bool* \nFlag to specify whether the gamma correction preprocessing is required or not. Performing gamma correction slightly increases the performance of the HOG algorithm.\n\n\n* **nlevels** – *int* \nMaximum number of detection window increases.\n\nAs we can see, the `cv2.HOGDescriptor()`function supports a wide range of parameters. The first few arguments (`block_size, block_stride, cell_size`, and `nbins`) are probably the ones you are most likely to change. The other parameters can be safely left at their default values and you will get good results. \n\nIn the code below, we will use the `cv2.HOGDescriptor()`function to set the cell size, block size, block stride, and the number of bins for the histograms of the HOG descriptor. We will then use `.compute(image)`method to compute the HOG descriptor (feature vector) for the given `image`. ",
"_____no_output_____"
]
],
[
[
"# Specify the parameters for our HOG descriptor\n\n# Cell Size in pixels (width, height). Must be smaller than the size of the detection window\n# and must be chosen so that the resulting Block Size is smaller than the detection window.\ncell_size = (6, 6)\n\n# Number of cells per block in each direction (x, y). Must be chosen so that the resulting\n# Block Size is smaller than the detection window\nnum_cells_per_block = (2, 2)\n\n# Block Size in pixels (width, height). Must be an integer multiple of Cell Size.\n# The Block Size must be smaller than the detection window\nblock_size = (num_cells_per_block[0] * cell_size[0],\n num_cells_per_block[1] * cell_size[1])\n\n# Calculate the number of cells that fit in our image in the x and y directions\nx_cells = gray_image.shape[1] // cell_size[0]\ny_cells = gray_image.shape[0] // cell_size[1]\n\n# Horizontal distance between blocks in units of Cell Size. Must be an integer and it must\n# be set such that (x_cells - num_cells_per_block[0]) / h_stride = integer.\nh_stride = 1\n\n# Vertical distance between blocks in units of Cell Size. Must be an integer and it must\n# be set such that (y_cells - num_cells_per_block[1]) / v_stride = integer.\nv_stride = 1\n\n# Block Stride in pixels (horizantal, vertical). Must be an integer multiple of Cell Size\nblock_stride = (cell_size[0] * h_stride, cell_size[1] * v_stride)\n\n# Number of gradient orientation bins\nnum_bins = 9 \n\n\n# Specify the size of the detection window (Region of Interest) in pixels (width, height).\n# It must be an integer multiple of Cell Size and it must cover the entire image. Because\n# the detection window must be an integer multiple of cell size, depending on the size of\n# your cells, the resulting detection window might be slightly smaller than the image.\n# This is perfectly ok.\nwin_size = (x_cells * cell_size[0] , y_cells * cell_size[1])\n\n# Print the shape of the gray scale image for reference\nprint('\\nThe gray scale image has shape: ', gray_image.shape)\nprint()\n\n# Print the parameters of our HOG descriptor\nprint('HOG Descriptor Parameters:\\n')\nprint('Window Size:', win_size)\nprint('Cell Size:', cell_size)\nprint('Block Size:', block_size)\nprint('Block Stride:', block_stride)\nprint('Number of Bins:', num_bins)\nprint()\n\n# Set the parameters of the HOG descriptor using the variables defined above\nhog = cv2.HOGDescriptor(win_size, block_size, block_stride, cell_size, num_bins)\n\n# Compute the HOG Descriptor for the gray scale image\nhog_descriptor = hog.compute(gray_image)",
"\nThe gray scale image has shape: (250, 250)\n\nHOG Descriptor Parameters:\n\nWindow Size: (246, 246)\nCell Size: (6, 6)\nBlock Size: (12, 12)\nBlock Stride: (6, 6)\nNumber of Bins: 9\n\n"
]
],
[
[
"# Number of Elements In The HOG Descriptor\n\nThe resulting HOG Descriptor (feature vector), contains the normalized histograms from all cells from all blocks in the detection window concatenated in one long vector. Therefore, the size of the HOG feature vector will be given by the total number of blocks in the detection window, multiplied by the number of cells per block, times the number of orientation bins:\n\n<span class=\"mathquill\">\n\\begin{equation}\n\\mbox{total_elements} = (\\mbox{total_number_of_blocks})\\mbox{ } \\times \\mbox{ } (\\mbox{number_cells_per_block})\\mbox{ } \\times \\mbox{ } (\\mbox{number_of_bins})\n\\end{equation}\n</span>\n\nIf we don’t have overlapping blocks (*i.e.* the `block_stride`equals the `block_size`), the total number of blocks can be easily calculated by dividing the size of the detection window by the block size. However, in the general case we have to take into account the fact that we have overlapping blocks. To find the total number of blocks in the general case (*i.e.* for any `block_stride` and `block_size`), we can use the formula given below:\n\n<span class=\"mathquill\">\n\\begin{equation}\n\\mbox{Total}_i = \\left( \\frac{\\mbox{block_size}_i}{\\mbox{block_stride}_i} \\right)\\left( \\frac{\\mbox{window_size}_i}{\\mbox{block_size}_i} \\right) - \\left [\\left( \\frac{\\mbox{block_size}_i}{\\mbox{block_stride}_i} \\right) - 1 \\right]; \\mbox{ for } i = x,y\n\\end{equation}\n</span>\n\nWhere <span class=\"mathquill\">Total$_x$</span>, is the total number of blocks along the width of the detection window, and <span class=\"mathquill\">Total$_y$</span>, is the total number of blocks along the height of the detection window. This formula for <span class=\"mathquill\">Total$_x$</span> and <span class=\"mathquill\">Total$_y$</span>, takes into account the extra blocks that result from overlapping. After calculating <span class=\"mathquill\">Total$_x$</span> and <span class=\"mathquill\">Total$_y$</span>, we can get the total number of blocks in the detection window by multiplying <span class=\"mathquill\">Total$_x$ $\\times$ Total$_y$</span>. The above formula can be simplified considerably because the `block_size`, `block_stride`, and `window_size`are all defined in terms of the `cell_size`. By making all the appropriate substitutions and cancelations the above formula reduces to:\n\n<span class=\"mathquill\">\n\\begin{equation}\n\\mbox{Total}_i = \\left(\\frac{\\mbox{cells}_i - \\mbox{num_cells_per_block}_i}{N_i}\\right) + 1\\mbox{ }; \\mbox{ for } i = x,y\n\\end{equation}\n</span>\n\nWhere <span class=\"mathquill\">cells$_x$</span> is the total number of cells along the width of the detection window, and <span class=\"mathquill\">cells$_y$</span>, is the total number of cells along the height of the detection window. And <span class=\"mathquill\">$N_x$</span> is the horizontal block stride in units of `cell_size` and <span class=\"mathquill\">$N_y$</span> is the vertical block stride in units of `cell_size`. \n\nLet's calculate what the number of elements for the HOG feature vector should be and check that it matches the shape of the HOG Descriptor calculated above.",
"_____no_output_____"
]
],
[
[
"# Calculate the total number of blocks along the width of the detection window\ntot_bx = np.uint32(((x_cells - num_cells_per_block[0]) / h_stride) + 1)\n\n# Calculate the total number of blocks along the height of the detection window\ntot_by = np.uint32(((y_cells - num_cells_per_block[1]) / v_stride) + 1)\n\n# Calculate the total number of elements in the feature vector\ntot_els = (tot_bx) * (tot_by) * num_cells_per_block[0] * num_cells_per_block[1] * num_bins\n\n# Print the total number of elements the HOG feature vector should have\nprint('\\nThe total number of elements in the HOG Feature Vector should be: ',\n tot_bx, 'x',\n tot_by, 'x',\n num_cells_per_block[0], 'x',\n num_cells_per_block[1], 'x',\n num_bins, '=',\n tot_els)\n\n# Print the shape of the HOG Descriptor to see that it matches the above\nprint('\\nThe HOG Descriptor has shape:', hog_descriptor.shape)\nprint()",
"\nThe total number of elements in the HOG Feature Vector should be: 40 x 40 x 2 x 2 x 9 = 57600\n\nThe HOG Descriptor has shape: (57600, 1)\n\n"
]
],
[
[
"# Visualizing The HOG Descriptor\n\nWe can visualize the HOG Descriptor by plotting the histogram associated with each cell as a collection of vectors. To do this, we will plot each bin in the histogram as a single vector whose magnitude is given by the height of the bin and its orientation is given by the angular bin that its associated with. Since any given cell might have multiple histograms associated with it, due to the overlapping blocks, we will choose to average all the histograms for each cell to produce a single histogram for each cell.\n\nOpenCV has no easy way to visualize the HOG Descriptor, so we have to do some manipulation first in order to visualize it. We will start by reshaping the HOG Descriptor in order to make our calculations easier. We will then compute the average histogram of each cell and finally we will convert the histogram bins into vectors. Once we have the vectors, we plot the corresponding vectors for each cell in an image. \n\nThe code below produces an interactive plot so that you can interact with the figure. The figure contains:\n* the grayscale image, \n* the HOG Descriptor (feature vector), \n* a zoomed-in portion of the HOG Descriptor, and \n* the histogram of the selected cell. \n\n**You can click anywhere on the gray scale image or the HOG Descriptor image to select a particular cell**. Once you click on either image a *magenta* rectangle will appear showing the cell you selected. The Zoom Window will show you a zoomed in version of the HOG descriptor around the selected cell; and the histogram plot will show you the corresponding histogram for the selected cell. The interactive window also has buttons at the bottom that allow for other functionality, such as panning, and giving you the option to save the figure if desired. The home button returns the figure to its default value.\n\n**NOTE**: If you are running this notebook in the Udacity workspace, there is around a 2 second lag in the interactive plot. This means that if you click in the image to zoom in, it will take about 2 seconds for the plot to refresh. ",
"_____no_output_____"
]
],
[
[
"%matplotlib notebook\n\nimport copy\nimport matplotlib.patches as patches\n\n# Set the default figure size\nplt.rcParams['figure.figsize'] = [9.8, 9]\n\n# Reshape the feature vector to [blocks_y, blocks_x, num_cells_per_block_x, num_cells_per_block_y, num_bins].\n# The blocks_x and blocks_y will be transposed so that the first index (blocks_y) referes to the row number\n# and the second index to the column number. This will be useful later when we plot the feature vector, so\n# that the feature vector indexing matches the image indexing.\nhog_descriptor_reshaped = hog_descriptor.reshape(tot_bx,\n tot_by,\n num_cells_per_block[0],\n num_cells_per_block[1],\n num_bins).transpose((1, 0, 2, 3, 4))\n\n# Print the shape of the feature vector for reference\nprint('The feature vector has shape:', hog_descriptor.shape)\n\n# Print the reshaped feature vector\nprint('The reshaped feature vector has shape:', hog_descriptor_reshaped.shape)\n\n# Create an array that will hold the average gradients for each cell\nave_grad = np.zeros((y_cells, x_cells, num_bins))\n\n# Print the shape of the ave_grad array for reference\nprint('The average gradient array has shape: ', ave_grad.shape) \n\n# Create an array that will count the number of histograms per cell\nhist_counter = np.zeros((y_cells, x_cells, 1))\n\n# Add up all the histograms for each cell and count the number of histograms per cell\nfor i in range (num_cells_per_block[0]):\n for j in range(num_cells_per_block[1]):\n ave_grad[i:tot_by + i,\n j:tot_bx + j] += hog_descriptor_reshaped[:, :, i, j, :]\n \n hist_counter[i:tot_by + i,\n j:tot_bx + j] += 1\n\n# Calculate the average gradient for each cell\nave_grad /= hist_counter\n \n# Calculate the total number of vectors we have in all the cells.\nlen_vecs = ave_grad.shape[0] * ave_grad.shape[1] * ave_grad.shape[2]\n\n# Create an array that has num_bins equally spaced between 0 and 180 degress in radians.\ndeg = np.linspace(0, np.pi, num_bins, endpoint = False)\n\n# Each cell will have a histogram with num_bins. For each cell, plot each bin as a vector (with its magnitude\n# equal to the height of the bin in the histogram, and its angle corresponding to the bin in the histogram). \n# To do this, create rank 1 arrays that will hold the (x,y)-coordinate of all the vectors in all the cells in the\n# image. Also, create the rank 1 arrays that will hold all the (U,V)-components of all the vectors in all the\n# cells in the image. Create the arrays that will hold all the vector positons and components.\nU = np.zeros((len_vecs))\nV = np.zeros((len_vecs))\nX = np.zeros((len_vecs))\nY = np.zeros((len_vecs))\n\n# Set the counter to zero\ncounter = 0\n\n# Use the cosine and sine functions to calculate the vector components (U,V) from their maginitudes. Remember the \n# cosine and sine functions take angles in radians. Calculate the vector positions and magnitudes from the\n# average gradient array\nfor i in range(ave_grad.shape[0]):\n for j in range(ave_grad.shape[1]):\n for k in range(ave_grad.shape[2]):\n U[counter] = ave_grad[i,j,k] * np.cos(deg[k])\n V[counter] = ave_grad[i,j,k] * np.sin(deg[k])\n \n X[counter] = (cell_size[0] / 2) + (cell_size[0] * i)\n Y[counter] = (cell_size[1] / 2) + (cell_size[1] * j)\n \n counter = counter + 1\n\n# Create the bins in degress to plot our histogram. \nangle_axis = np.linspace(0, 180, num_bins, endpoint = False)\nangle_axis += ((angle_axis[1] - angle_axis[0]) / 2)\n\n# Create a figure with 4 subplots arranged in 2 x 2\nfig, ((a,b),(c,d)) = plt.subplots(2,2)\n\n# Set the title of each subplot\na.set(title = 'Gray Scale Image\\n(Click to Zoom)')\nb.set(title = 'HOG Descriptor\\n(Click to Zoom)')\nc.set(title = 'Zoom Window', xlim = (0, 18), ylim = (0, 18), autoscale_on = False)\nd.set(title = 'Histogram of Gradients')\n\n# Plot the gray scale image\na.imshow(gray_image, cmap = 'gray')\na.set_aspect(aspect = 1)\n\n# Plot the feature vector (HOG Descriptor)\nb.quiver(Y, X, U, V, color = 'white', headwidth = 0, headlength = 0, scale_units = 'inches', scale = 5)\nb.invert_yaxis()\nb.set_aspect(aspect = 1)\n\n#b.set_facecolor('black')\nb.set_axis_bgcolor('black')\n\n# Define function for interactive zoom\ndef onpress(event):\n \n #Unless the left mouse button is pressed do nothing\n if event.button != 1:\n return\n \n # Only accept clicks for subplots a and b\n if event.inaxes in [a, b]:\n \n # Get mouse click coordinates\n x, y = event.xdata, event.ydata\n \n # Select the cell closest to the mouse click coordinates\n cell_num_x = np.uint32(x / cell_size[0])\n cell_num_y = np.uint32(y / cell_size[1])\n \n # Set the edge coordinates of the rectangle patch\n edgex = x - (x % cell_size[0])\n edgey = y - (y % cell_size[1])\n \n # Create a rectangle patch that matches the the cell selected above \n rect = patches.Rectangle((edgex, edgey),\n cell_size[0], cell_size[1],\n linewidth = 1,\n edgecolor = 'magenta',\n facecolor='none')\n \n # A single patch can only be used in a single plot. Create copies\n # of the patch to use in the other subplots\n rect2 = copy.copy(rect)\n rect3 = copy.copy(rect)\n \n # Update all subplots\n a.clear()\n a.set(title = 'Gray Scale Image\\n(Click to Zoom)')\n a.imshow(gray_image, cmap = 'gray')\n a.set_aspect(aspect = 1)\n a.add_patch(rect)\n\n b.clear()\n b.set(title = 'HOG Descriptor\\n(Click to Zoom)')\n b.quiver(Y, X, U, V, color = 'white', headwidth = 0, headlength = 0, scale_units = 'inches', scale = 5)\n b.invert_yaxis()\n b.set_aspect(aspect = 1)\n \n #b.set_facecolor('black')\n b.set_axis_bgcolor('black')\n \n b.add_patch(rect2)\n\n c.clear()\n c.set(title = 'Zoom Window')\n c.quiver(Y, X, U, V, color = 'white', headwidth = 0, headlength = 0, scale_units = 'inches', scale = 1)\n c.set_xlim(edgex - cell_size[0], edgex + (2 * cell_size[0]))\n c.set_ylim(edgey - cell_size[1], edgey + (2 * cell_size[1]))\n c.invert_yaxis()\n c.set_aspect(aspect = 1)\n \n #c.set_facecolor('black')\n c.set_axis_bgcolor('black')\n \n c.add_patch(rect3)\n\n d.clear()\n d.set(title = 'Histogram of Gradients')\n d.grid()\n d.set_xlim(0, 180)\n d.set_xticks(angle_axis)\n d.set_xlabel('Angle')\n d.bar(angle_axis,\n ave_grad[cell_num_y, cell_num_x, :],\n 180 // num_bins,\n align = 'center',\n alpha = 0.5,\n linewidth = 1.2,\n edgecolor = 'k')\n\n fig.canvas.draw()\n\n# Create a connection between the figure and the mouse click\nfig.canvas.mpl_connect('button_press_event', onpress)\nplt.show()",
"The feature vector has shape: (57600, 1)\nThe reshaped feature vector has shape: (40, 40, 2, 2, 9)\nThe average gradient array has shape: (41, 41, 9)\n"
]
],
[
[
"# Understanding The Histograms\n\nLet's take a look at a couple of snapshots of the above figure to see if the histograms for the selected cell make sense. Let's start looking at a cell that is inside a triangle and not near an edge:\n\n<br>\n<figure>\n <img src = \"./in_cell_images/snapshot1.png\" width = \"70%\" style = \"border: thin silver solid; padding: 1px\">\n <figcaption style = \"text-align:center; font-style:italic\">Fig. 4. - Histograms Inside a Triangle.</figcaption>\n</figure> \n<br>\n\nIn this case, since the triangle is nearly all of the same color there shouldn't be any dominant gradient in the selected cell. As we can clearly see in the Zoom Window and the histogram, this is indeed the case. We have many gradients but none of them clearly dominates over the other.\n\nNow let’s take a look at a cell that is near a horizontal edge:\n\n<br>\n<figure>\n <img src = \"./in_cell_images/snapshot2.png\" width = \"70%\" style = \"border: thin silver solid; padding: 1px\">\n <figcaption style = \"text-align:center; font-style:italic\">Fig. 5. - Histograms Near a Horizontal Edge.</figcaption>\n</figure> \n<br>\n\nRemember that edges are areas of an image where the intensity changes abruptly. In these cases, we will have a high intensity gradient in some particular direction. This is exactly what we see in the corresponding histogram and Zoom Window for the selected cell. In the Zoom Window, we can see that the dominant gradient is pointing up, almost at 90 degrees, since that’s the direction in which there is a sharp change in intensity. Therefore, we should expect to see the 90-degree bin in the histogram to dominate strongly over the others. This is in fact what we see. \n\nNow let’s take a look at a cell that is near a vertical edge:\n\n<br>\n<figure>\n <img src = \"./in_cell_images/snapshot3.png\" width = \"70%\" style = \"border: thin silver solid; padding: 1px\">\n <figcaption style = \"text-align:center; font-style:italic\">Fig. 6. - Histograms Near a Vertical Edge.</figcaption>\n</figure> \n<br>\n\nIn this case we expect the dominant gradient in the cell to be horizontal, close to 180 degrees, since that’s the direction in which there is a sharp change in intensity. Therefore, we should expect to see the 170-degree bin in the histogram to dominate strongly over the others. This is what we see in the histogram but we also see that there is another dominant gradient in the cell, namely the one in the 10-degree bin. The reason for this, is because the HOG algorithm is using unsigned gradients, which means 0 degrees and 180 degrees are considered the same. Therefore, when the histograms are being created, angles between 160 and 180 degrees, contribute proportionally to both the 10-degree bin and the 170-degree bin. This results in there being two dominant gradients in the cell near the vertical edge instead of just one. \n\nTo conclude let’s take a look at a cell that is near a diagonal edge.\n\n<br>\n<figure>\n <img src = \"./in_cell_images/snapshot4.png\" width = \"70%\" style = \"border: thin silver solid; padding: 1px\">\n <figcaption style = \"text-align:center; font-style:italic\">Fig. 7. - Histograms Near a Diagonal Edge.</figcaption>\n</figure> \n<br>\n\nTo understand what we are seeing, let’s first remember that gradients have an *x*-component, and a *y*-component, just like vectors. Therefore, the resulting orientation of a gradient is going to be given by the vector sum of its components. For this reason, on vertical edges the gradients are horizontal, because they only have an x-component, as we saw in Figure 4. While on horizontal edges the gradients are vertical, because they only have a y-component, as we saw in Figure 3. Consequently, on diagonal edges, the gradients are also going to be diagonal because both the *x* and *y* components are non-zero. Since the diagonal edges in the image are close to 45 degrees, we should expect to see a dominant gradient orientation in the 50-degree bin. This is in fact what we see in the histogram but, just like in Figure 4., we see there are two dominant gradients instead of just one. The reason for this is that when the histograms are being created, angles that are near the boundaries of bins, contribute proportionally to the adjacent bins. For example, a gradient with an angle of 40 degrees, is right in the middle of the 30-degree and 50-degree bin. Therefore, the magnitude of the gradient is split evenly into the 30-degree and 50-degree bin. This results in there being two dominant gradients in the cell near the diagonal edge instead of just one.\n\nNow that you know how HOG is implemented, in the workspace you will find a notebook named *Examples*. In there, you will be able set your own paramters for the HOG descriptor for various images. Have fun!",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
4acd10f8bc64567d500c666fb23d9d4a2ea8872b
| 16,996 |
ipynb
|
Jupyter Notebook
|
lecture/ds1-2020-11-12-lec12.ipynb
|
ucsb-ds/ds1-f20-content
|
25f62c7a597b98da436ca39631761c1f3feccfdd
|
[
"MIT"
] | 2 |
2020-10-14T12:43:18.000Z
|
2021-01-06T18:06:16.000Z
|
lecture/ds1-2020-11-12-lec12.ipynb
|
ucsb-ds/ds1-f20-content
|
25f62c7a597b98da436ca39631761c1f3feccfdd
|
[
"MIT"
] | 2 |
2020-10-13T05:21:15.000Z
|
2020-12-11T18:03:54.000Z
|
lecture/ds1-2020-11-12-lec12.ipynb
|
ucsb-ds/ds1-f20-content
|
25f62c7a597b98da436ca39631761c1f3feccfdd
|
[
"MIT"
] | 3 |
2021-01-05T05:30:01.000Z
|
2021-03-16T22:09:08.000Z
| 18.78011 | 82 | 0.412391 |
[
[
[
"# Running Simulations",
"_____no_output_____"
]
],
[
[
"import numpy as np\nfrom datascience import *",
"_____no_output_____"
]
],
[
[
"## `np.append` to add items to an array",
"_____no_output_____"
]
],
[
[
"fruits = make_array(\"apple\")",
"_____no_output_____"
],
[
"fruits = np.append(fruits, fruits)",
"_____no_output_____"
],
[
"fruits",
"_____no_output_____"
],
[
"fruits = np.append(fruits, fruits)",
"_____no_output_____"
],
[
"fruits",
"_____no_output_____"
],
[
"fruits = np.append(fruits, fruits)\nfruits",
"_____no_output_____"
],
[
"new_fruit = make_array(\"quince\")",
"_____no_output_____"
],
[
"new_fruit",
"_____no_output_____"
],
[
"salad = np.append(new_fruit, fruits)",
"_____no_output_____"
],
[
"len(salad)",
"_____no_output_____"
],
[
"salad",
"_____no_output_____"
]
],
[
[
"## Selecting random values from an array",
"_____no_output_____"
]
],
[
[
"np.random.choice(salad)",
"_____no_output_____"
],
[
"np.random.choice(salad)",
"_____no_output_____"
],
[
"np.random.choice(salad)",
"_____no_output_____"
],
[
"np.random.choice(salad)",
"_____no_output_____"
],
[
"np.random.choice(salad)",
"_____no_output_____"
],
[
"np.random.choice(salad)",
"_____no_output_____"
],
[
"np.random.choice(salad)",
"_____no_output_____"
],
[
"np.random.choice(salad)",
"_____no_output_____"
],
[
"np.random.choice(salad)",
"_____no_output_____"
],
[
"np.random.choice(salad)",
"_____no_output_____"
]
],
[
[
"## Simulating a coin toss",
"_____no_output_____"
]
],
[
[
"coin = make_array('H', 'T')",
"_____no_output_____"
],
[
"toss_10 = np.random.choice(coin, 10)",
"_____no_output_____"
],
[
"toss_10",
"_____no_output_____"
],
[
"sum(toss_10 == 'H')",
"_____no_output_____"
],
[
"np.count_nonzero(toss_10 == 'H')",
"_____no_output_____"
],
[
"# How often do we get 9 heads when tossing a coin 10 times\nresult = make_array()\nfor i in np.arange(10000):\n toss_10 = np.random.choice(coin, 10) # toss a coin 10 times\n num_heads = np.count_nonzero(toss_10 == 'H') # count the num of heads\n result = np.append(result, num_heads)",
"_____no_output_____"
],
[
"result",
"_____no_output_____"
],
[
"len(result)",
"_____no_output_____"
],
[
"sum(result == 9)",
"_____no_output_____"
],
[
"91/10000",
"_____no_output_____"
],
[
"for i in np.arange(0, 11):\n print(sum(result == i), \"times the coin came up with\", i, \"heads\")",
"6 times the coin came up with 0 heads\n86 times the coin came up with 1 heads\n446 times the coin came up with 2 heads\n1206 times the coin came up with 3 heads\n2075 times the coin came up with 4 heads\n2451 times the coin came up with 5 heads\n2059 times the coin came up with 6 heads\n1155 times the coin came up with 7 heads\n421 times the coin came up with 8 heads\n91 times the coin came up with 9 heads\n4 times the coin came up with 10 heads\n"
],
[
"coin_toss = Table().with_column(\"Toss\", result)",
"_____no_output_____"
],
[
"coin_toss_count = coin_toss.group(\"Toss\")",
"_____no_output_____"
],
[
"coin_toss_count",
"_____no_output_____"
],
[
"coin_toss_count.hist(\"count\")",
"_____no_output_____"
],
[
"coin_toss",
"_____no_output_____"
],
[
"coin_toss.where(\"Toss\", are.above(5)).num_rows",
"_____no_output_____"
],
[
"coin_toss.where(\"Toss\", are.below(5)).num_rows",
"_____no_output_____"
],
[
"10000 - 3730 - 3819",
"_____no_output_____"
],
[
"coin_toss.where(\"Toss\", are.equal_to(5)).num_rows",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acd235d8520f342308d8bdb11ab3651c59946b9
| 112,395 |
ipynb
|
Jupyter Notebook
|
5. Text Mining/Topic Modelling Using Gensim.ipynb
|
paragajg/DSB7
|
d37679141de3606ea571be6f675dbe8026f2a20e
|
[
"MIT"
] | 4 |
2021-05-08T03:40:33.000Z
|
2021-06-13T08:18:55.000Z
|
5. Text Mining/Topic Modelling Using Gensim.ipynb
|
paragajg/DSB7
|
d37679141de3606ea571be6f675dbe8026f2a20e
|
[
"MIT"
] | null | null | null |
5. Text Mining/Topic Modelling Using Gensim.ipynb
|
paragajg/DSB7
|
d37679141de3606ea571be6f675dbe8026f2a20e
|
[
"MIT"
] | 3 |
2021-05-01T13:54:35.000Z
|
2021-05-15T16:12:55.000Z
| 102.643836 | 69,368 | 0.63477 |
[
[
[
"### Topic Modelling Demo Code",
"_____no_output_____"
],
[
"#### Things I want to do -\n- Identify a package to build / train LDA model\n- Use visualization to explore Documents -> Topics Distribution -> Word distribution",
"_____no_output_____"
]
],
[
[
"!pip install pyLDAvis, gensim",
"_____no_output_____"
],
[
"import numpy as np\nimport pandas as pd\n\n# Visualization\nimport matplotlib.pyplot as plt\nfrom matplotlib.ticker import FuncFormatter\nimport seaborn as sns\nimport pyLDAvis.gensim\n\n# Text Preprocessing and model building\nfrom gensim.corpora import Dictionary\nimport nltk\nfrom nltk.stem import WordNetLemmatizer\nimport re\n# Iteratively read files\nimport glob\nimport os\n\n# For displaying images in ipython\nfrom IPython.display import HTML, display",
"/Users/paragpradhan/opt/anaconda3/lib/python3.8/site-packages/scipy/sparse/sparsetools.py:21: DeprecationWarning: `scipy.sparse.sparsetools` is deprecated!\nscipy.sparse.sparsetools is a private module for scipy.sparse, and should not be used.\n _deprecated()\n"
],
[
"%matplotlib inline\nplt.style.use('ggplot')\nplt.rcParams['figure.figsize'] = (14.0, 8.7)\n#warnings.filterwarnings('ignore')\npd.options.display.float_format = '{:,.2f}'.format",
"_____no_output_____"
]
],
[
[
"<h2>Latent Dirichlet Allocation</h2>\n<h3>From Documents -- DTM -- LDA Model</h3>\n\nTopic modeling aims to automatically summarize large collections of documents to facilitate organization and management, as well as search and recommendations. At the same time, it can enable the understanding of documents to the extent that humans can interpret the descriptions of topics\n\n<img src=\"images/lda2.png\" alt=\"lda\" style=\"width:60%\">\n<img src=\"images/docs_to_lda.png\" alt=\"ldaflow\" style=\"width:100%\">",
"_____no_output_____"
],
[
"### Load Data",
"_____no_output_____"
]
],
[
[
"# User defined function to read and store bbc data from multipe folders\ndef load_data(folder_names,root_path):\n fileNames = [path + '/' + 'bbc' +'/'+ folder + '/*.txt' for path,folder in zip([root_path]*len(folder_names),\n folder_names )]\n doc_list = []\n tags = folder_names\n for docs in fileNames:\n #print(docs)\n #print(type(docs))\n doc = glob.glob(docs) # glob method iterates through the all the text documents in a folder\n for text in doc:\n with open(text, encoding='latin1') as f:\n topic = docs.split('/')[-2]\n\n lines = f.readlines()\n heading = lines[0].strip()\n body = ' '.join([l.strip() for l in lines[1:]])\n doc_list.append([topic, heading, body])\n print(\"Completed loading data from folder: %s\"%topic)\n \n print(\"Completed Loading entire text\")\n \n return doc_list",
"_____no_output_____"
],
[
"folder_names = ['business','entertainment','politics','sport','tech']\ndocs = load_data(folder_names = folder_names, root_path = os.getcwd())",
"Completed loading data from folder: business\nCompleted loading data from folder: entertainment\nCompleted loading data from folder: politics\nCompleted loading data from folder: sport\nCompleted loading data from folder: tech\nCompleted Loading entire text\n"
],
[
"docs = pd.DataFrame(docs, columns=['Category', 'Heading', 'Article'])\nprint(docs.head())\nprint('\\nShape of data is {}\\n'.format(docs.shape))",
" Category Heading \\\n0 business UK economy facing 'major risks' \n1 business Aids and climate top Davos agenda \n2 business Asian quake hits European shares \n3 business India power shares jump on debut \n4 business Lacroix label bought by US firm \n\n Article \n0 The UK manufacturing sector will continue to ... \n1 Climate change and the fight against Aids are... \n2 Shares in Europe's leading reinsurers and tra... \n3 Shares in India's largest power producer, Nat... \n4 Luxury goods group LVMH has sold its loss-mak... \n\nShape of data is (2225, 3)\n\n"
]
],
[
[
"### Extract Raw Corpus",
"_____no_output_____"
]
],
[
[
"articles = docs.Article.tolist()",
"_____no_output_____"
],
[
"print(type(articles))\nprint(articles[0:2])",
"<class 'list'>\n[' The UK manufacturing sector will continue to face \"serious challenges\" over the next two years, the British Chamber of Commerce (BCC) has said. The group\\'s quarterly survey of companies found exports had picked up in the last three months of 2004 to their best levels in eight years. The rise came despite exchange rates being cited as a major concern. However, the BCC found the whole UK economy still faced \"major risks\" and warned that growth is set to slow. It recently forecast economic growth will slow from more than 3% in 2004 to a little below 2.5% in both 2005 and 2006. Manufacturers\\' domestic sales growth fell back slightly in the quarter, the survey of 5,196 firms found. Employment in manufacturing also fell and job expectations were at their lowest level for a year. \"Despite some positive news for the export sector, there are worrying signs for manufacturing,\" the BCC said. \"These results reinforce our concern over the sector\\'s persistent inability to sustain recovery.\" The outlook for the service sector was \"uncertain\" despite an increase in exports and orders over the quarter, the BCC noted. The BCC found confidence increased in the quarter across both the manufacturing and service sectors although overall it failed to reach the levels at the start of 2004. The reduced threat of interest rate increases had contributed to improved confidence, it said. The Bank of England raised interest rates five times between November 2003 and August last year. But rates have been kept on hold since then amid signs of falling consumer confidence and a slowdown in output. \"The pressure on costs and margins, the relentless increase in regulations, and the threat of higher taxes remain serious problems,\" BCC director general David Frost said. \"While consumer spending is set to decelerate significantly over the next 12-18 months, it is unlikely that investment and exports will rise sufficiently strongly to pick up the slack.\"', ' Climate change and the fight against Aids are leading the list of concerns for the first day of the World Economic Forum in the Swiss resort of Davos. Some 2,000 business and political leaders from around the globe will listen to UK Prime Minister Tony Blair\\'s opening speech on Wednesday. Mr Blair will focus on Africa\\'s development plans and global warming. Earlier in the day came an update on efforts to have 3 million people on anti-Aids drugs by the end of 2005. The World Health Organisation (WHO) said 700,000 people in poor countries were on life-extending drugs - up from 440,000 six months earlier but amounting to only 12% of the 5.8 million who needed them. A $2bn \"funding gap\" still stood in the way of hitting the 2005 target, the WHO said. The themes to be stressed by Mr Blair - whose attendance was announced at the last minute - are those he wants to dominate the UK\\'s chairmanship of the G8 group of industrialised states. Other issues to be discussed at the five-day conference range from China\\'s economic power to Iraq\\'s future after this Sunday\\'s elections. Aside from Mr Blair, more than 20 other world leaders are expected to attend including French President Jacques Chirac - due to speak by video link after bad weather delayed his helicopter - and South African President Thabo Mbeki, whose arrival has been delayed by Ivory Coast peace talks. The Ukraine\\'s new president, Viktor Yushchenko, will also be there - as will newly elected Palestinian leader Mahmoud Abbas. Showbiz figures will also put in an appearance, from U2 frontman Bono - a well-known campaigner on trade and development issues - to Angelina Jolie, a goodwill campaigner for the UN on refugees. Unlike previous years, protests against the WEF are expected to be muted. Anti-globalisation campaigners have called off a demonstration planned for the weekend. At the same time, about 100,000 people are expected to converge on the Brazilian resort of Porto Alegre for the World Social Forum - the so-called \"anti-Davos\" for campaigners against globalisation, for fair trade, and many other causes. In contrast, the Davos forum is dominated by business issues - from outsourcing to corporate leadership - with bosses of more than a fifth of the world\\'s 500 largest companies scheduled to attend. A survey published on the eve of the conference by PricewaterhouseCoopers said four in ten business leaders were \"very confident\" that their companies would see sales rise in 2005. Asian and American executives, however, were much more confident than their European counterparts. But the political discussions, focusing on Iran, Iraq and China, are likely to dominate media attention.']\n"
],
[
"wordnet_lemmatizer = WordNetLemmatizer()",
"_____no_output_____"
]
],
[
[
"### Preprocessing of Raw Text",
"_____no_output_____"
]
],
[
[
"from nltk.corpus import stopwords\nimport nltk\n# nltk.download('punkt')\n# nltk.download('wordnet')\n# nltk.download('stopwords')",
"_____no_output_____"
],
[
"# nltk.download('stopwords')",
"[nltk_data] Downloading package stopwords to\n[nltk_data] /Users/paragpradhan/nltk_data...\n[nltk_data] Unzipping corpora/stopwords.zip.\n"
],
[
"stopwords = stopwords.word('english')",
"_____no_output_____"
],
[
"# Method to preprocess my raw data\ndef preprocessText(x):\n temp = x.lower()\n temp = re.sub(r'[^\\w]', ' ', temp)\n temp = nltk.word_tokenize(temp)\n temp = [wordnet_lemmatizer.lemmatize(w) for w in temp]\n temp = [word for word in temp if word not in stopwords ]\n return temp",
"_____no_output_____"
],
[
"articles_final = [preprocessText(article) for article in articles]",
"_____no_output_____"
],
[
"articles_final[0:2]",
"_____no_output_____"
]
],
[
[
"### Transformation of Preprocessed text into Vector form using Gensim",
"_____no_output_____"
]
],
[
[
"# Create a dictionary representation of the documents.\ndictionary = Dictionary(articles_final)\n\n# Filter out words that occur less than 20 documents, or more than 50% of the documents.\ndictionary.filter_extremes(no_below=20, no_above=0.5)",
"_____no_output_____"
],
[
"print(dictionary)",
"Dictionary(3101 unique tokens: ['12', '18', '2', '2003', '2004']...)\n"
],
[
"# Bag-of-words representation of the documents.\ncorpus = [dictionary.doc2bow(doc) for doc in articles_final]",
"_____no_output_____"
],
[
"print('Number of unique tokens: %d' % len(dictionary))\nprint('Number of documents: %d' % len(corpus))",
"Number of unique tokens: 3101\nNumber of documents: 2225\n"
]
],
[
[
"### Train LDA model using Gensim",
"_____no_output_____"
]
],
[
[
"dictionary",
"_____no_output_____"
],
[
"# Train LDA model.\nfrom gensim.models import LdaModel\n\n# Set training parameters.\nnum_topics = 5\nchunksize = 2000\npasses = 10\n# iterations = 400\neval_every = None # Don't evaluate model perplexity, takes too much time.\n\n# Make a index to word dictionary.\ntemp = dictionary[0] # This is only to \"load\" the dictionary.\nid2word = dictionary.id2token\n\nmodel = LdaModel(\n corpus=corpus,\n id2word=id2word,\n chunksize=chunksize,\n alpha='auto',\n eta='auto',\n# iterations=iterations,\n num_topics=num_topics,\n passes=passes,\n eval_every=eval_every\n)",
"_____no_output_____"
]
],
[
[
"### Model exploration: Top K words in each topic",
"_____no_output_____"
]
],
[
[
"# Print the Keyword in the 10 topics\npprint.pprint(model.print_topics(num_words= 20))\ndoc_lda = model[corpus]",
"[(0,\n '0.014*\"game\" + 0.009*\"microsoft\" + 0.008*\"software\" + 0.008*\"virus\" + '\n '0.008*\"new\" + 0.008*\"firm\" + 0.007*\"file\" + 0.007*\"system\" + 0.007*\"site\" + '\n '0.007*\"program\" + 0.007*\"technology\" + 0.007*\"machine\" + 0.006*\"mr\" + '\n '0.006*\"mail\" + 0.006*\"people\" + 0.006*\"network\" + 0.006*\"user\" + '\n '0.005*\"sony\" + 0.005*\"computer\" + 0.005*\"window\"'),\n (1,\n '0.015*\"people\" + 0.012*\"service\" + 0.011*\"mobile\" + 0.009*\"phone\" + '\n '0.008*\"one\" + 0.008*\"could\" + 0.007*\"user\" + 0.007*\"technology\" + '\n '0.007*\"uk\" + 0.007*\"internet\" + 0.006*\"network\" + 0.006*\"broadband\" + '\n '0.006*\"net\" + 0.006*\"online\" + 0.006*\"system\" + 0.006*\"use\" + '\n '0.006*\"million\" + 0.006*\"new\" + 0.006*\"access\" + 0.006*\"company\"'),\n (2,\n '0.017*\"game\" + 0.009*\"film\" + 0.008*\"best\" + 0.008*\"time\" + 0.007*\"one\" + '\n '0.006*\"first\" + 0.005*\"world\" + 0.005*\"award\" + 0.005*\"player\" + '\n '0.005*\"like\" + 0.005*\"last\" + 0.005*\"two\" + 0.004*\"new\" + 0.004*\"team\" + '\n '0.004*\"back\" + 0.004*\"play\" + 0.004*\"win\" + 0.004*\"well\" + 0.004*\"star\" + '\n '0.004*\"take\"'),\n (3,\n '0.015*\"mr\" + 0.008*\"u\" + 0.007*\"â\" + 0.006*\"government\" + 0.004*\"new\" + '\n '0.004*\"company\" + 0.004*\"election\" + 0.004*\"party\" + 0.004*\"say\" + '\n '0.004*\"labour\" + 0.004*\"minister\" + 0.004*\"could\" + 0.004*\"country\" + '\n '0.004*\"last\" + 0.004*\"told\" + 0.004*\"people\" + 0.003*\"1\" + 0.003*\"one\" + '\n '0.003*\"000\" + 0.003*\"plan\"'),\n (4,\n '0.012*\"mobile\" + 0.012*\"people\" + 0.010*\"music\" + 0.010*\"technology\" + '\n '0.009*\"phone\" + 0.008*\"tv\" + 0.008*\"player\" + 0.007*\"gadget\" + '\n '0.007*\"digital\" + 0.007*\"search\" + 0.007*\"video\" + 0.006*\"one\" + 0.006*\"mr\" '\n '+ 0.006*\"could\" + 0.006*\"show\" + 0.006*\"device\" + 0.005*\"apple\" + '\n '0.005*\"service\" + 0.005*\"medium\" + 0.005*\"new\"')]\n"
]
],
[
[
"### Model Visualization using PyLDAvis",
"_____no_output_____"
]
],
[
[
"pyLDAvis.enable_notebook()\nvis = pyLDAvis.gensim.prepare(model, corpus, dictionary=dictionary)\nvis",
"_____no_output_____"
]
],
[
[
"### Assign Topic Model Numbers to original Data Frame as Column",
"_____no_output_____"
]
],
[
[
"# Assigns the topics to the documents in corpus\nlda_corpus = model[corpus]",
"_____no_output_____"
],
[
"topics = []\n\nfor doc in lda_corpus:\n temp_id = []\n temp_score = []\n for doc_tuple in doc:\n temp_id.append(doc_tuple[0])\n temp_score.append(doc_tuple[1])\n index = np.argmax(temp_score)\n topics.append(temp_id[index])",
"_____no_output_____"
],
[
"docs[\"Topic_num\"] = topics",
"_____no_output_____"
],
[
"docs.tail(n= 40)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
4acd2ba86801eb449bfb98a6d744ccd962a66c4f
| 13,215 |
ipynb
|
Jupyter Notebook
|
examples/Marks/Pyplot/Lines.ipynb
|
Chinmay4400/bqplot
|
ea1868705fc206a50de4819fe8588c8d48ecefa0
|
[
"Apache-2.0"
] | 2 |
2021-04-28T08:31:14.000Z
|
2021-08-13T19:43:55.000Z
|
examples/Marks/Pyplot/Lines.ipynb
|
Chinmay4400/bqplot
|
ea1868705fc206a50de4819fe8588c8d48ecefa0
|
[
"Apache-2.0"
] | 14 |
2020-07-13T16:36:31.000Z
|
2021-12-08T16:47:54.000Z
|
examples/Marks/Pyplot/Lines.ipynb
|
Chinmay4400/bqplot
|
ea1868705fc206a50de4819fe8588c8d48ecefa0
|
[
"Apache-2.0"
] | 1 |
2021-01-24T16:35:29.000Z
|
2021-01-24T16:35:29.000Z
| 26.91446 | 281 | 0.548241 |
[
[
[
"### Introduction\n\nThe `Lines` object provides the following features:\n\n1. Ability to plot a single set or multiple sets of y-values as a function of a set or multiple sets of x-values\n2. Ability to style the line object in different ways, by setting different attributes such as the `colors`, `line_style`, `stroke_width` etc.\n3. Ability to specify a marker at each point passed to the line. The marker can be a shape which is at the data points between which the line is interpolated and can be set through the `markers` attribute",
"_____no_output_____"
],
[
"The `Lines` object has the following attributes",
"_____no_output_____"
],
[
" | Attribute | Description | Default Value |\n|:-:|---|:-:|\n| `colors` | Sets the color of each line, takes as input a list of any RGB, HEX, or HTML color name | `CATEGORY10` |\n| `opacities` | Controls the opacity of each line, takes as input a real number between 0 and 1 | `1.0` |\n| `stroke_width` | Real number which sets the width of all paths | `2.0` |\n| `line_style` | Specifies whether a line is solid, dashed, dotted or both dashed and dotted | `'solid'` |\n| `interpolation` | Sets the type of interpolation between two points | `'linear'` |\n| `marker` | Specifies the shape of the marker inserted at each data point | `None` |\n| `marker_size` | Controls the size of the marker, takes as input a non-negative integer | `64` |\n|`close_path`| Controls whether to close the paths or not | `False` |\n|`fill`| Specifies in which way the paths are filled. Can be set to one of `{'none', 'bottom', 'top', 'inside'}`| `None` |\n|`fill_colors`| `List` that specifies the `fill` colors of each path | `[]` | \n| **Data Attribute** | **Description** | **Default Value** |\n|`x` |abscissas of the data points | `array([])` |\n|`y` |ordinates of the data points | `array([])` |\n|`color` | Data according to which the `Lines` will be colored. Setting it to `None` defaults the choice of colors to the `colors` attribute | `None` |",
"_____no_output_____"
],
[
"## pyplot's plot method can be used to plot lines with meaningful defaults",
"_____no_output_____"
]
],
[
[
"import numpy as np\nfrom pandas import date_range\nimport bqplot.pyplot as plt\nfrom bqplot import *",
"_____no_output_____"
],
[
"security_1 = np.cumsum(np.random.randn(150)) + 100.\nsecurity_2 = np.cumsum(np.random.randn(150)) + 100.",
"_____no_output_____"
]
],
[
[
"## Basic Line Chart",
"_____no_output_____"
]
],
[
[
"fig = plt.figure(title='Security 1')\naxes_options = {'x': {'label': 'Index'}, 'y': {'label': 'Price'}}\n# x values default to range of values when not specified\nline = plt.plot(security_1, axes_options=axes_options)\nfig",
"_____no_output_____"
]
],
[
[
"**We can explore the different attributes by changing each of them for the plot above:**",
"_____no_output_____"
]
],
[
[
"line.colors = ['DarkOrange']",
"_____no_output_____"
]
],
[
[
"In a similar way, we can also change any attribute after the plot has been displayed to change the plot. Run each of the cells below, and try changing the attributes to explore the different features and how they affect the plot.",
"_____no_output_____"
]
],
[
[
"# The opacity allows us to display the Line while featuring other Marks that may be on the Figure\nline.opacities = [.5]",
"_____no_output_____"
],
[
"line.stroke_width = 2.5",
"_____no_output_____"
]
],
[
[
"To switch to an area chart, set the `fill` attribute, and control the look with `fill_opacities` and `fill_colors`.",
"_____no_output_____"
]
],
[
[
"line.fill = 'bottom'\nline.fill_opacities = [0.2]",
"_____no_output_____"
],
[
"line.line_style = 'dashed'",
"_____no_output_____"
],
[
"line.interpolation = 'basis'",
"_____no_output_____"
]
],
[
[
"While a `Lines` plot allows the user to extract the general shape of the data being plotted, there may be a need to visualize discrete data points along with this shape. This is where the `markers` attribute comes in.",
"_____no_output_____"
]
],
[
[
"line.marker = 'triangle-down'",
"_____no_output_____"
]
],
[
[
"The `marker` attributes accepts the values `square`, `circle`, `cross`, `diamond`, `square`, `triangle-down`, `triangle-up`, `arrow`, `rectangle`, `ellipse`. Try changing the string above and re-running the cell to see how each `marker` type looks.",
"_____no_output_____"
],
[
"## Plotting a Time-Series",
"_____no_output_____"
],
[
"The `DateScale` allows us to plot time series as a `Lines` plot conveniently with most `date` formats.",
"_____no_output_____"
]
],
[
[
"# Here we define the dates we would like to use\ndates = date_range(start='01-01-2007', periods=150)",
"_____no_output_____"
],
[
"fig = plt.figure(title='Time Series')\naxes_options = {'x': {'label': 'Date'}, 'y': {'label': 'Security 1'}}\ntime_series = plt.plot(dates, security_1, \n axes_options=axes_options)\nfig",
"_____no_output_____"
]
],
[
[
"## Plotting multiples sets of data",
"_____no_output_____"
],
[
"The `Lines` mark allows the user to plot multiple `y`-values for a single `x`-value. This can be done by passing an `ndarray` or a list of the different `y`-values as the y-attribute of the `Lines` as shown below.",
"_____no_output_____"
]
],
[
[
"dates_new = date_range(start='06-01-2007', periods=150)",
"_____no_output_____"
]
],
[
[
"We pass each data set as an element of a `list`",
"_____no_output_____"
]
],
[
[
"fig = plt.figure()\naxes_options = {'x': {'label': 'Date'}, 'y': {'label': 'Price'}}\nline = plt.plot(dates, [security_1, security_2], \n labels=['Security 1', 'Security 2'],\n axes_options=axes_options,\n display_legend=True)\nfig",
"_____no_output_____"
]
],
[
[
"Similarly, we can also pass multiple `x`-values for multiple sets of `y`-values",
"_____no_output_____"
]
],
[
[
"line.x, line.y = [dates, dates_new], [security_1, security_2]",
"_____no_output_____"
]
],
[
[
"### Coloring Lines according to data",
"_____no_output_____"
],
[
"The `color` attribute of a `Lines` mark can also be used to encode one more dimension of data. Suppose we have a portfolio of securities and we would like to color them based on whether we have bought or sold them. We can use the `color` attribute to encode this information.",
"_____no_output_____"
]
],
[
[
"fig = plt.figure()\naxes_options = {'x': {'label': 'Date'}, \n 'y': {'label': 'Security 1'},\n 'color' : {'visible': False}}\n# add a custom color scale to color the lines\nplt.scales(scales={'color': ColorScale(colors=['Red', 'Green'])})",
"_____no_output_____"
],
[
"dates_color = date_range(start='06-01-2007', periods=150)",
"_____no_output_____"
],
[
"securities = 100. + np.cumsum(np.random.randn(150, 10), axis=0)\n# we generate 10 random price series and 10 random positions\npositions = np.random.randint(0, 2, size=10)",
"_____no_output_____"
],
[
"# We pass the color scale and the color data to the plot method\nline = plt.plot(dates_color, securities.T, color=positions, \n axes_options=axes_options)\nfig",
"_____no_output_____"
]
],
[
[
"We can also reset the colors of the Line to their defaults by setting the `color` attribute to `None`.",
"_____no_output_____"
]
],
[
[
"line.color = None",
"_____no_output_____"
]
],
[
[
"## Patches",
"_____no_output_____"
],
[
"The `fill` attribute of the `Lines` mark allows us to fill a path in different ways, while the `fill_colors` attribute lets us control the color of the `fill`",
"_____no_output_____"
]
],
[
[
"fig = plt.figure(animation_duration=1000)\npatch = plt.plot([],[],\n fill_colors=['orange', 'blue', 'red'],\n fill='inside',\n axes_options={'x': {'visible': False}, 'y': {'visible': False}},\n stroke_width=10,\n close_path=True,\n display_legend=True)\n\npatch.x = [[0, 2, 1.2, np.nan, np.nan, np.nan, np.nan], [0.5, 2.5, 1.7 , np.nan, np.nan, np.nan, np.nan], [4, 5, 6, 6, 5, 4, 3]], \npatch.y = [[0, 0, 1 , np.nan, np.nan, np.nan, np.nan], [0.5, 0.5, -0.5, np.nan, np.nan, np.nan, np.nan], [1, 1.1, 1.2, 2.3, 2.2, 2.7, 1.0]]\nfig",
"_____no_output_____"
],
[
"patch.opacities = [0.1, 0.2]",
"_____no_output_____"
],
[
"patch.x = [[2, 3, 3.2, np.nan, np.nan, np.nan, np.nan], [0.5, 2.5, 1.7, np.nan, np.nan, np.nan, np.nan], [4,5,6, 6, 5, 4, 3]]",
"_____no_output_____"
],
[
"patch.close_path = False",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
4acd39d7924dfbf6fc13b5b8e184f2088f4b1301
| 42,529 |
ipynb
|
Jupyter Notebook
|
Section 1/Implementing Activation Functions.ipynb
|
PacktPublishing/TensorFlow-for-Machine-Learning-Solutions-
|
3f258ee117bffaf18f5420fc4e6eefaab604fa02
|
[
"MIT"
] | 2 |
2019-05-19T03:32:23.000Z
|
2019-09-20T12:08:27.000Z
|
Section 1/Implementing Activation Functions.ipynb
|
PacktPublishing/TensorFlow-for-Machine-Learning-Solutions-
|
3f258ee117bffaf18f5420fc4e6eefaab604fa02
|
[
"MIT"
] | null | null | null |
Section 1/Implementing Activation Functions.ipynb
|
PacktPublishing/TensorFlow-for-Machine-Learning-Solutions-
|
3f258ee117bffaf18f5420fc4e6eefaab604fa02
|
[
"MIT"
] | 1 |
2019-05-20T17:08:36.000Z
|
2019-05-20T17:08:36.000Z
| 189.017778 | 20,582 | 0.901197 |
[
[
[
"import matplotlib.pyplot as plt\nimport numpy as np\nimport tensorflow as tf\nfrom tensorflow.python.framework import ops\nops.reset_default_graph()\nsess = tf.Session()",
"_____no_output_____"
],
[
"x_vals = np.linspace(start=-10., stop=10., num=100)",
"_____no_output_____"
],
[
"print(sess.run(tf.nn.relu([-3., 3., 10.])))\ny_relu = sess.run(tf.nn.relu(x_vals))",
"[ 0. 3. 10.]\n"
],
[
"print(sess.run(tf.nn.relu6([-3., 3., 10.])))\ny_relu6 = sess.run(tf.nn.relu6(x_vals))",
"[ 0. 3. 6.]\n"
],
[
"print(sess.run(tf.nn.sigmoid([-1., 0., 1.])))\ny_sigmoid = sess.run(tf.nn.sigmoid(x_vals))",
"[ 0.26894143 0.5 0.7310586 ]\n"
],
[
"print(sess.run(tf.nn.tanh([-1., 0., 1.])))\ny_tanh = sess.run(tf.nn.tanh(x_vals))",
"[-0.76159418 0. 0.76159418]\n"
],
[
"print(sess.run(tf.nn.softsign([-1., 0., 1.])))\ny_softsign = sess.run(tf.nn.softsign(x_vals))",
"[-0.5 0. 0.5]\n"
],
[
"print(sess.run(tf.nn.softplus([-1., 0., 1.])))\ny_softplus = sess.run(tf.nn.softplus(x_vals))",
"[ 0.31326166 0.69314718 1.31326163]\n"
],
[
"print(sess.run(tf.nn.elu([-1., 0., 1.])))\ny_elu = sess.run(tf.nn.elu(x_vals))",
"[-0.63212055 0. 1. ]\n"
],
[
"plt.plot(x_vals, y_softplus, 'r--', label='Softplus', linewidth=2)\nplt.plot(x_vals, y_relu, 'b:', label='ReLU', linewidth=2)\nplt.plot(x_vals, y_relu6, 'g-.', label='ReLU6', linewidth=2)\nplt.plot(x_vals, y_elu, 'k-', label='ExpLU', linewidth=0.5)\nplt.ylim([-1.5,7])\nplt.legend(loc='upper left')\nplt.show()\n\nplt.plot(x_vals, y_sigmoid, 'r--', label='Sigmoid', linewidth=2)\nplt.plot(x_vals, y_tanh, 'b:', label='Tanh', linewidth=2)\nplt.plot(x_vals, y_softsign, 'g-.', label='Softsign', linewidth=2)\nplt.ylim([-2,2])\nplt.legend(loc='upper left')\nplt.show()",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acd40c5f9ed7fae572b1e8673bef411e02136b3
| 155,011 |
ipynb
|
Jupyter Notebook
|
notebooks/poisoning_attack_feature_collision.ipynb
|
tagomaru/adversarial-robustness-toolbox
|
b0d86c4489f92275d828e7cde7e9d0a34181f2fe
|
[
"MIT"
] | 1 |
2022-01-31T15:17:20.000Z
|
2022-01-31T15:17:20.000Z
|
notebooks/poisoning_attack_feature_collision.ipynb
|
proteanblank/adversarial-robustness-toolbox
|
8415d693b7bddd1d24d58d34e8b228d9c1487627
|
[
"MIT"
] | null | null | null |
notebooks/poisoning_attack_feature_collision.ipynb
|
proteanblank/adversarial-robustness-toolbox
|
8415d693b7bddd1d24d58d34e8b228d9c1487627
|
[
"MIT"
] | 1 |
2022-03-22T05:30:31.000Z
|
2022-03-22T05:30:31.000Z
| 379 | 63,944 | 0.935005 |
[
[
[
"# Clean-Label Feature Collision Attacks on a Keras Classifier",
"_____no_output_____"
],
[
"In this notebook, we will learn how to use ART to run a clean-label feature collision poisoning attack on a neural network trained with Keras. We will be training our data on a subset of the CIFAR-10 dataset. The methods described are derived from [this paper](https://arxiv.org/abs/1804.00792) by Shafahi, Huang, et. al. 2018.",
"_____no_output_____"
]
],
[
[
"import os, sys\nfrom os.path import abspath\n\nmodule_path = os.path.abspath(os.path.join('..'))\nif module_path not in sys.path:\n sys.path.append(module_path)\n\nimport warnings\nwarnings.filterwarnings('ignore')\n\nfrom keras.models import load_model\n\nfrom art import config\nfrom art.utils import load_dataset, get_file\nfrom art.estimators.classification import KerasClassifier\nfrom art.attacks.poisoning import FeatureCollisionAttack\n\nimport numpy as np\n\n%matplotlib inline\nimport matplotlib.pyplot as plt\n\nnp.random.seed(301)",
"Using TensorFlow backend.\n"
],
[
"(x_train, y_train), (x_test, y_test), min_, max_ = load_dataset('cifar10')\n\nnum_samples_train = 1000\nnum_samples_test = 1000\nx_train = x_train[0:num_samples_train]\ny_train = y_train[0:num_samples_train]\nx_test = x_test[0:num_samples_test]\ny_test = y_test[0:num_samples_test]\n\nclass_descr = ['airplane', 'automobile', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']",
"_____no_output_____"
]
],
[
[
"## Load Model to be Attacked\n\nIn this example, we using a RESNET50 model pretrained on the CIFAR dataset.",
"_____no_output_____"
]
],
[
[
"path = get_file('cifar_alexnet.h5',extract=False, path=config.ART_DATA_PATH,\n url='https://www.dropbox.com/s/ta75pl4krya5djj/cifar_alexnet.h5?dl=1')\nclassifier_model = load_model(path)\nclassifier = KerasClassifier(clip_values=(min_, max_), model=classifier_model, use_logits=False, \n preprocessing=(0.5, 1))",
"_____no_output_____"
]
],
[
[
"## Choose Target Image from Test Set",
"_____no_output_____"
]
],
[
[
"target_class = \"bird\" # one of ['airplane', 'automobile', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']\ntarget_label = np.zeros(len(class_descr))\ntarget_label[class_descr.index(target_class)] = 1\ntarget_instance = np.expand_dims(x_test[np.argmax(y_test, axis=1) == class_descr.index(target_class)][3], axis=0)\n\nfig = plt.imshow(target_instance[0])\n\nprint('true_class: ' + target_class)\nprint('predicted_class: ' + class_descr[np.argmax(classifier.predict(target_instance), axis=1)[0]])\n\nfeature_layer = classifier.layer_names[-2]",
"true_class: bird\npredicted_class: bird\n"
]
],
[
[
"## Poison Training Images to Misclassify Test\n\nThe attacker wants to make it such that whenever a prediction is made on this particular cat the output will be a horse.",
"_____no_output_____"
]
],
[
[
"base_class = \"frog\" # one of ['airplane', 'automobile', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']\nbase_idxs = np.argmax(y_test, axis=1) == class_descr.index(base_class)\nbase_instances = np.copy(x_test[base_idxs][:10])\nbase_labels = y_test[base_idxs][:10]\n\nx_test_pred = np.argmax(classifier.predict(base_instances), axis=1)\nnb_correct_pred = np.sum(x_test_pred == np.argmax(base_labels, axis=1))\n\nprint(\"New test data to be poisoned (10 images):\")\nprint(\"Correctly classified: {}\".format(nb_correct_pred))\nprint(\"Incorrectly classified: {}\".format(10-nb_correct_pred))",
"New test data to be poisoned (10 images):\nCorrectly classified: 9\nIncorrectly classified: 1\n"
],
[
"plt.figure(figsize=(10,10))\nfor i in range(0, 9):\n pred_label, true_label = class_descr[x_test_pred[i]], class_descr[np.argmax(base_labels[i])]\n plt.subplot(330 + 1 + i)\n fig=plt.imshow(base_instances[i])\n fig.axes.get_xaxis().set_visible(False)\n fig.axes.get_yaxis().set_visible(False)\n fig.axes.text(0.5, -0.1, pred_label + \" (\" + true_label + \")\", fontsize=12, transform=fig.axes.transAxes, \n horizontalalignment='center')",
"_____no_output_____"
]
],
[
[
"The captions on the images can be read: `predicted label (true label)`",
"_____no_output_____"
],
[
"## Creating Poison Frogs",
"_____no_output_____"
]
],
[
[
"attack = FeatureCollisionAttack(classifier, target_instance, feature_layer, max_iter=10, similarity_coeff=256, watermark=0.3)\npoison, poison_labels = attack.poison(base_instances)",
"100%|██████████| 10/10 [00:01<00:00, 9.76it/s]\n100%|██████████| 10/10 [00:00<00:00, 16.56it/s]\n100%|██████████| 10/10 [00:00<00:00, 15.97it/s]\n100%|██████████| 10/10 [00:00<00:00, 17.06it/s]\n100%|██████████| 10/10 [00:00<00:00, 16.40it/s]\n100%|██████████| 10/10 [00:00<00:00, 15.85it/s]\n100%|██████████| 10/10 [00:00<00:00, 17.06it/s]\n100%|██████████| 10/10 [00:00<00:00, 15.25it/s]\n100%|██████████| 10/10 [00:00<00:00, 15.91it/s]\n100%|██████████| 10/10 [00:00<00:00, 16.80it/s]\n"
],
[
"poison_pred = np.argmax(classifier.predict(poison), axis=1)\nplt.figure(figsize=(10,10))\nfor i in range(0, 9):\n pred_label, true_label = class_descr[poison_pred[i]], class_descr[np.argmax(poison_labels[i])]\n plt.subplot(330 + 1 + i)\n fig=plt.imshow(poison[i])\n fig.axes.get_xaxis().set_visible(False)\n fig.axes.get_yaxis().set_visible(False)\n fig.axes.text(0.5, -0.1, pred_label + \" (\" + true_label + \")\", fontsize=12, transform=fig.axes.transAxes, \n horizontalalignment='center')",
"_____no_output_____"
]
],
[
[
"Notice how the network classifies most of theses poison examples as frogs, and it's not incorrect to do so. The examples look mostly froggy. A slight watermark of the target instance is also added to push the poisons closer to the target class in feature space.",
"_____no_output_____"
],
[
"## Training with Poison Images",
"_____no_output_____"
]
],
[
[
"classifier.set_learning_phase(True)\nprint(x_train.shape)\nprint(base_instances.shape)\nadv_train = np.vstack([x_train, poison])\nadv_labels = np.vstack([y_train, poison_labels])\nclassifier.fit(adv_train, adv_labels, nb_epochs=5, batch_size=4)",
"(1000, 32, 32, 3)\n(10, 32, 32, 3)\nEpoch 1/5\n252/252 [==============================] - 34s 136ms/step - loss: 0.4635 - acc: 0.92061s - loss: 0.4711 - a\nEpoch 2/5\n252/252 [==============================] - 26s 101ms/step - loss: 0.3405 - acc: 0.9325\nEpoch 3/5\n252/252 [==============================] - 26s 102ms/step - loss: 0.2121 - acc: 0.9534\nEpoch 4/5\n252/252 [==============================] - 26s 102ms/step - loss: 0.1950 - acc: 0.9742\nEpoch 5/5\n252/252 [==============================] - 30s 118ms/step - loss: 0.1888 - acc: 0.9712\n"
]
],
[
[
"## Fooled Network Misclassifies Bird",
"_____no_output_____"
]
],
[
[
"fig = plt.imshow(target_instance[0])\n\nprint('true_class: ' + target_class)\nprint('predicted_class: ' + class_descr[np.argmax(classifier.predict(target_instance), axis=1)[0]])",
"true_class: bird\npredicted_class: frog\n"
]
],
[
[
"These attacks allow adversaries who can poison your dataset the ability to mislabel any particular target instance of their choosing without manipulating labels.",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
4acd5b1acd7922d7aa0d4e5e8604f88898c84fe7
| 6,499 |
ipynb
|
Jupyter Notebook
|
example_nets/batch_normalization_example.ipynb
|
ZhiangChen/soft_arm
|
7df9585695452985887b09013317015ea4b73aea
|
[
"MIT"
] | null | null | null |
example_nets/batch_normalization_example.ipynb
|
ZhiangChen/soft_arm
|
7df9585695452985887b09013317015ea4b73aea
|
[
"MIT"
] | null | null | null |
example_nets/batch_normalization_example.ipynb
|
ZhiangChen/soft_arm
|
7df9585695452985887b09013317015ea4b73aea
|
[
"MIT"
] | null | null | null | 33.328205 | 120 | 0.530235 |
[
[
[
"import tensorflow as tf\nimport numpy as np\nfrom tensorflow.examples.tutorials.mnist import input_data\nmnist = input_data.read_data_sets('MNIST_data', one_hot=True)\ntest_data = mnist.test\ntrain_data = mnist.train\nvalid_data = mnist.validation",
"Successfully downloaded train-images-idx3-ubyte.gz 9912422 bytes.\nExtracting MNIST_data/train-images-idx3-ubyte.gz\nSuccessfully downloaded train-labels-idx1-ubyte.gz 28881 bytes.\nExtracting MNIST_data/train-labels-idx1-ubyte.gz\nSuccessfully downloaded t10k-images-idx3-ubyte.gz 1648877 bytes.\nExtracting MNIST_data/t10k-images-idx3-ubyte.gz\nSuccessfully downloaded t10k-labels-idx1-ubyte.gz 4542 bytes.\nExtracting MNIST_data/t10k-labels-idx1-ubyte.gz\n"
],
[
"epsilon = 1e-3\n\nclass FC(object):\n def __init__(self, learning_rate=0.01):\n self.lr = learning_rate\n self.sess = tf.Session()\n self.x = tf.placeholder(tf.float32,[None, 784], 'x')\n self.y_ = tf.placeholder(tf.float32, [None, 10], 'y_')\n self.training = tf.placeholder(tf.bool, name='training')\n self._build_net(self.x,'FC')\n\n with tf.variable_scope('Accuracy'):\n self.correct_prediction = tf.equal(tf.argmax(self.y,1), tf.argmax(self.y_,1))\n self.accuracy = tf.reduce_mean(tf.cast(self.correct_prediction, tf.float32))\n with tf.variable_scope('Train'):\n self.loss = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(labels=self.y_, logits=self.y)) \n extra_update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)\n with tf.control_dependencies(extra_update_ops):\n self.train_opt = tf.train.GradientDescentOptimizer(self.lr).minimize(self.loss)\n self.sess.run(tf.global_variables_initializer())\n \n def batch_norm_wrapper(self, inputs, is_training, decay = 0.999):\n\n scale = tf.Variable(tf.ones([inputs.get_shape()[-1]]))\n beta = tf.Variable(tf.zeros([inputs.get_shape()[-1]]))\n pop_mean = tf.Variable(tf.zeros([inputs.get_shape()[-1]]), trainable=False)\n pop_var = tf.Variable(tf.ones([inputs.get_shape()[-1]]), trainable=False)\n\n if is_training==tf.constant(True):\n batch_mean, batch_var = tf.nn.moments(inputs,[0])\n train_mean = tf.assign(pop_mean,\n pop_mean * decay + batch_mean * (1 - decay))\n train_var = tf.assign(pop_var,\n pop_var * decay + batch_var * (1 - decay))\n with tf.control_dependencies([train_mean, train_var]):\n return tf.nn.batch_normalization(inputs,\n batch_mean, batch_var, beta, scale, epsilon)\n else:\n return tf.nn.batch_normalization(inputs,\n pop_mean, pop_var, beta, scale, epsilon)\n \n def _build_net(self, x, scope):\n with tf.variable_scope(scope):\n bn = tf.layers.batch_normalization(x, axis=1, training=self.training, name = 'bn')\n #bn = self.batch_norm_wrapper(x, self.training)\n hidden = tf.layers.dense(bn, 50, activation=tf.nn.relu, name='l1')\n self.y = tf.layers.dense(hidden, 10, name='o')\n \n def learn(self, x, y):\n loss,_ = self.sess.run([self.loss,self.train_opt],{self.x:x, self.y_:y, self.training:True})\n return loss\n \n def inference(self, x, y=None):\n y = self.sess.run(self.y,{self.x:x, self.training:False})\n #loss,_ = self.sess.run(self.loss,{self.x:x, self.y_:y, self.training:False})\n return y\n \nfc = FC()",
"_____no_output_____"
],
[
"OUTPUT_GRAPH = True\nif OUTPUT_GRAPH:\n tf.summary.FileWriter(\"logs/\", fc.sess.graph)\n",
"_____no_output_____"
],
[
"for i in range(1000):\n batch = train_data.next_batch(100)\n loss = fc.learn(batch[0],batch[1])\n if i%200 == 0:\n print(loss)\n \nbatch = valid_data.next_batch(5000)\nprint(\"validation accuracy: %f\" % fc.sess.run(fc.accuracy,{fc.x:batch[0], fc.y_:batch[1], fc.training:True}))",
"2.70122\n0.862728\n0.519033\n0.401137\n0.317988\nvalidation accuracy: 0.917200\n"
],
[
"nm = 1000\ncount = 0\nfor _ in range(nm):\n t = test_data.next_batch(1)\n x = t[0]\n y = fc.inference(x)\n a = np.argmax(y,axis=1)\n b = np.argmax(t[1],axis=1)\n if a==b:\n count += 1\nprint count/float(nm)",
"0.914\n"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code"
]
] |
4acd64f921c9c074e789375cf81033dfd5c37399
| 17,433 |
ipynb
|
Jupyter Notebook
|
notebooks/05-strings-and-files.ipynb
|
tjsuki/EPETutorials
|
f3e54ebaad0bf7a251625d936f9e34f47d26bc83
|
[
"MIT"
] | null | null | null |
notebooks/05-strings-and-files.ipynb
|
tjsuki/EPETutorials
|
f3e54ebaad0bf7a251625d936f9e34f47d26bc83
|
[
"MIT"
] | null | null | null |
notebooks/05-strings-and-files.ipynb
|
tjsuki/EPETutorials
|
f3e54ebaad0bf7a251625d936f9e34f47d26bc83
|
[
"MIT"
] | null | null | null | 26.215038 | 373 | 0.565651 |
[
[
[
"# Strings\n\nLesson goals:\n\n1. Examine the string class in greater detail.\n2. Use `open()` to open, read, and write to files.\n\n\nTo start understanding the string type, let's use the built in helpsystem.",
"_____no_output_____"
]
],
[
[
"help(str)",
"_____no_output_____"
]
],
[
[
"The help page for string is very long, and it may be easier to keep it open\nin a browser window by going to the [online Python\ndocumentation](http://docs.python.org/library/stdtypes.html#sequence-types-str-unicode-list-tuple-bytearray-buffer-xrange)\nwhile we talk about its properties.\n\nAt its heart, a string is just a sequence of characters. Basic strings are\ndefined using single or double quotes.",
"_____no_output_____"
]
],
[
[
"s = \"This is a string.\"\ns2 = 'This is another string that uses single quotes'",
"_____no_output_____"
]
],
[
[
"The reason for having two types of quotes to define a string is\nemphasized in these examples:",
"_____no_output_____"
]
],
[
[
"s = \"Bob's mom called to say hello.\"\ns = 'Bob's mom called to say hello.'",
"_____no_output_____"
]
],
[
[
"The second one should be an error: Python interprets it as `s = 'Bob'` then the\nrest of the line breaks the language standard.\n\nCharacters in literal strings must come from the ASCII character set,\nwhich is a set of 127 character codes that is used by all modern\nprogramming languages and computers. Unfortunately, ASCII does not have\nroom for non-Roman characters like accents or Eastern scripts. Unicode\nstrings in Python are specified with a leading u:",
"_____no_output_____"
]
],
[
[
"u = u'abcdé'",
"_____no_output_____"
]
],
[
[
"For the rest of this lecture, we will deal with ASCII strings, because\nmost scientific data that is stored as text is stored with ASCII.",
"_____no_output_____"
],
[
"## Escape Characters \nHow can you have multiline line strings in python? We can represent an \"enter\" using the escape character '\\n'. An [escape character](https://en.wikipedia.org/wiki/Escape_character) starts with a '\\\\' and is followed by another character. This invokes an alternative interpretation in the string. Try running the example below to see how \\n changes the string: ",
"_____no_output_____"
]
],
[
[
"s = \"Hello\\n World\"\nprint(s)",
"_____no_output_____"
]
],
[
[
"Notice how it didn't print \\n, but replaced it with a newline. There are more characters like this, such as \\t which is replaced with a tab or \\b that is equal to a backspace. Use [this guide](https://linuxconfig.org/list-of-python-escape-sequence-characters-with-examples) to print the following output as one string:",
"_____no_output_____"
],
[
"\"If you think you can do a thing or think you can't do a thing, you're right.\" \n - Henry Ford\n /\\\n/ \\",
"_____no_output_____"
]
],
[
[
"s = \"Your string here\"\nprint(s) # do not modify the print statement",
"_____no_output_____"
]
],
[
[
"## Working with Strings\n\nStrings are iterables, which means many of the ideas from lists can also\nbe applied directly to string manipulation. For instance, characters can\nbe accessed individually or in sequences:",
"_____no_output_____"
]
],
[
[
"s = 'abcdefghijklmnopqrstuvwxyz'\ns[0]",
"_____no_output_____"
],
[
"s[-1]",
"_____no_output_____"
],
[
"s[1:4] #this include char at index 1, but excludes char at index 4",
"_____no_output_____"
]
],
[
[
"They can also be compared using sort and equals.",
"_____no_output_____"
]
],
[
[
"'str1' == 'str2'",
"_____no_output_____"
],
[
"'str1' == 'str1'",
"_____no_output_____"
],
[
"'str1' < 'str2'",
"_____no_output_____"
]
],
[
[
"In the help screen, which we looked at above, there are lots of\nfunctions that look like this:\n\n | __add__(...)\n | x.__add__(y) <==> x+y\n\n | __le__(...)\n | x.__le__(y) <==> x<y\n\nThese are special Python functions that interpret operations like < and \\+.\nWe'll talk more about these in the next lecture on Classes.\n\nSome special functions introduce handy text functions.",
"_____no_output_____"
],
[
"**Hands on example**\n\nTry each of the following functions on a few strings. What do these\nfunctions do?",
"_____no_output_____"
]
],
[
[
"s = \"This is a string \"",
"_____no_output_____"
],
[
"s.startswith(\"This\")",
"_____no_output_____"
],
[
"s.split(\" \")",
"_____no_output_____"
],
[
"s.strip() # This won't change every string!",
"_____no_output_____"
],
[
"s.capitalize()",
"_____no_output_____"
],
[
"s.lower()",
"_____no_output_____"
],
[
"s.upper()",
"_____no_output_____"
]
],
[
[
"## Formatting\nTry printing \"Dave was traveling at 50 mph for 4.5 hours\" using these given variables:",
"_____no_output_____"
]
],
[
[
"name = \"Dave\"\nv = 50\nt = 4.5",
"_____no_output_____"
]
],
[
[
"Here is an easy way to do this is using string formatting:",
"_____no_output_____"
]
],
[
[
"print(\"%s was traveling at %d mph for %f hours\" % (name, v, t))",
"_____no_output_____"
]
],
[
[
"**%s** is used to represent a string (name), **%d** is used to represented an integer (v), and **%f** is replaced with a float (t). Now, try printing this data as \"Dave drove 10 miles faster than Sally for 4.5 hours.\"",
"_____no_output_____"
]
],
[
[
"name = \"Sally\"\nv1 = 40\nprint(\"Your print here\" % ())",
"_____no_output_____"
]
],
[
[
"## File I/O\n\nPython has a built-in function called \"open()\" that can be used to\nmanipulate files. The help information for open is below:",
"_____no_output_____"
]
],
[
[
"help(open)",
"_____no_output_____"
]
],
[
[
"The main two parameters we'll need to worry about are 'file', the name of the\nfile, and 'mode', which determines whether we can read from or write to the file. `open(...)` returns a file object, which acts like a pointer into the file, similarily to how an assigned variable can 'point' to a list/array. \nAn example will make this clear. In the code below, I've opened a file\nthat contains one line:\n\n $ cat ./OtherFiles/testfile.txt\n abcde\n fghij\n\nNow let's open this file in Python:",
"_____no_output_____"
]
],
[
[
"f = open('./OtherFiles/testfile.txt','r')",
"_____no_output_____"
]
],
[
[
"The second input, 'r' means I want to open the file for reading only. I\ncannot write to this handle. The read() command will read a specified\nnumber of bytes:",
"_____no_output_____"
]
],
[
[
"s = f.read(3)\nprint(s)",
"_____no_output_____"
]
],
[
[
"We read the first three characters, where each character is a byte long.\nWe can see that the file handle points to the 4th byte (index number 3)\nin the file:",
"_____no_output_____"
]
],
[
[
"f.tell() # which index we are pointing at",
"_____no_output_____"
],
[
"f.read(1) # read the 1st byte, starting from where the file handle is pointing",
"_____no_output_____"
],
[
"f.close() # close the old handle",
"_____no_output_____"
],
[
"f.read() # can't read anymore because the file is closed.",
"_____no_output_____"
]
],
[
[
"The file we are using is a long series of characters, but two of the\ncharacters are new line characters. If we looked at the file in\nsequence, it would look like \"abcdenfghijn\". Separating a file into\nlines is popular enough that there are two ways to read whole lines in a\nfile. The first is to use the `readlines()` method:",
"_____no_output_____"
]
],
[
[
"f = open('OtherFiles/testfile.txt','r')\nlines = f.readlines()\nprint(lines)\nf.close() # Always close the file when you are done with it",
"_____no_output_____"
]
],
[
[
"A very important point about the readline method is that it *keeps* the\nnewline character at the end of each line. You can use the `strip()`\nmethod to get rid of the escape characters at the beggining and end of the string.\n\nFile handles are also iterable, which means we can use them in for loops\nor list extensions. You will learn more about this iteration later:",
"_____no_output_____"
]
],
[
[
"f = open('OtherFiles/testfile.txt','r')\nlines = [line.strip() for line in f]\nf.close()\nprint(lines)",
"_____no_output_____"
],
[
"lines = []\nf = open('OtherFiles/testfile.txt','r')\nfor line in f:\n lines.append(line.strip())\nf.close()\nprint(lines)",
"_____no_output_____"
]
],
[
[
"These are equivalent operations. It's often best to handle a file one\nline at a time, particularly when the file is so large it might not fit\nin memory.\n\nThe other half of the story is writing output to files. We'll talk about\ntwo techniques: writing to the shell and writing to files directly.\n\nIf your program only creates one stream of output, it's often a good\nidea to write to the shell using the print function. There are several\nadvantages to this strategy, including the fact that it allows the user\nto select where they want to store the output without worrying about any\ncommand line flags. You can use \"\\>\" to direct the output of your\nprogram to a file or use \"|\" to pipe it to another program (this was covered in the 01-shell notebook).\n\nSometimes, you need to direct your output directly to a file handle. For\ninstance, if your program produces two output streams, you may want to\nassign two open file handles. Opening a file for reading simply requires\nchanging the second option from 'r' to 'w' or 'a'.\n\n*Caution!* Opening a file with the 'w' option means start writing *at\nthe beginning*, which may overwrite old material. If you want to append\nto the file without losing what is already there, open it with 'a'.\n\nWriting to a file uses the `write()` command, which accepts a string. Check outfile.txt before and after running the following code.",
"_____no_output_____"
]
],
[
[
"outfile = open('OtherFiles/outfile.txt','w')\noutfile.write('This is the first line!')\noutfile.close()",
"_____no_output_____"
]
],
[
[
"Another way to write to a file is to use `writelines()`, which accepts a\nlist of strings and writes them in order. \n\n*Caution!* `writelines()` does not\nappend newlines. If you really want to write a newline at the end of\neach string in the list, add it yourself.",
"_____no_output_____"
],
[
"### Aside About File Editing\n\nHow is it possible that you can edit a file in place. You can use `f.seek()`\nand `f.tell()` to verify that even if your file handle is pointing to the\nmiddle of a file, write commands go to the end of the file in append\nmode. The best way to change a file is to open a temporary file in\n/tmp/, fill it, and then move it to overwrite the original. On large\nclusters, /tmp/ is often local to each node, which means it reduces I/O\nbottlenecks associated with writing large amounts of data.",
"_____no_output_____"
],
[
"## Exercise\nFind the index of the string 'needle' in the file OtherFiles/haystack.txt using the `.find()` method of a String. Recall that `file.read()` will return the file as a string.",
"_____no_output_____"
]
],
[
[
"f = None; # open the file here\nn = 0 # find the index here\nprint(\"Needle at Index %d\" % n) # n should be 185\nf.close()",
"_____no_output_____"
]
],
[
[
"Create a new file OtherFiles/haystack1.txt by opening in \"w+\" mode. \"a+\" and \"r+\" also create a file if it does not exist. Write the contents of haystack.txt into this new file and add an extra 'needle' at the end.",
"_____no_output_____"
]
],
[
[
"# Your code here\nf = None;\nf.close()\n\n# Prints the file to check your answer; don't write anything here\nf = open(\"OtherFiles/haystack1.txt\",\"r\")\nprint(f.read())\nf.close()",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4acd713ee2bd7d2a1d2bbdb5efb40741cce1566e
| 196,956 |
ipynb
|
Jupyter Notebook
|
web/12/Canonical_economic_models.ipynb
|
NumEconCopenhagen/lectures-2022
|
0bbd4a7af3f26009ec92e874ace095e998ea1fab
|
[
"MIT"
] | 5 |
2022-02-13T06:41:31.000Z
|
2022-03-29T10:24:48.000Z
|
web/12/Canonical_economic_models.ipynb
|
NumEconCopenhagen/lectures-2022
|
0bbd4a7af3f26009ec92e874ace095e998ea1fab
|
[
"MIT"
] | null | null | null |
web/12/Canonical_economic_models.ipynb
|
NumEconCopenhagen/lectures-2022
|
0bbd4a7af3f26009ec92e874ace095e998ea1fab
|
[
"MIT"
] | 10 |
2022-02-07T11:51:15.000Z
|
2022-03-09T07:20:02.000Z
| 119.22276 | 31,232 | 0.869377 |
[
[
[
"# Lecture 12: Canonical Economic Models",
"_____no_output_____"
],
[
"[Download on GitHub](https://github.com/NumEconCopenhagen/lectures-2022)\n\n[<img src=\"https://mybinder.org/badge_logo.svg\">](https://mybinder.org/v2/gh/NumEconCopenhagen/lectures-2022/master?urlpath=lab/tree/12/Canonical_economic_models.ipynb)",
"_____no_output_____"
],
[
"1. [OverLapping Generations (OLG) model](#OverLapping-Generations-(OLG)-model)\n2. [Ramsey model](#Ramsey-model)\n3. [Further perspectives](#Further-perspectives)\n",
"_____no_output_____"
],
[
"You will learn how to solve **two canonical economic models**:\n\n1. The **overlapping generations (OLG) model**\n2. The **Ramsey model**\n\n**Main take-away:** Hopefully inspiration to analyze such models on your own.",
"_____no_output_____"
]
],
[
[
"%load_ext autoreload\n%autoreload 2\n\nimport numpy as np\nfrom scipy import optimize\n\n# plotting\nimport matplotlib.pyplot as plt\nplt.style.use('seaborn-whitegrid')\nplt.rcParams.update({'font.size': 12})\n\n# models\nfrom OLGModel import OLGModelClass\nfrom RamseyModel import RamseyModelClass",
"_____no_output_____"
]
],
[
[
"<a id=\"OverLapping-Generations-(OLG)-model\"></a>\n\n# 1. OverLapping Generations (OLG) model",
"_____no_output_____"
],
[
"## 1.1 Model description",
"_____no_output_____"
],
[
"**Time:** Discrete and indexed by $t\\in\\{0,1,\\dots\\}$.",
"_____no_output_____"
],
[
"**Demographics:** Population is constant. A life consists of\ntwo periods, *young* and *old*.",
"_____no_output_____"
],
[
"**Households:** As young a household supplies labor exogenously, $L_{t}=1$, and earns a after tax wage $(1-\\tau_w)w_{t}$. Consumption as young and old\nare denoted by $C_{1t}$ and $C_{2t+1}$. The after-tax return on saving is $(1-\\tau_{r})r_{t+1}$. Utility is\n \n$$\n\\begin{aligned}\nU & =\\max_{s_{t}\\in[0,1]}\\frac{C_{1t}^{1-\\sigma}}{1-\\sigma}+\\beta\\frac{C_{1t+1}^{1-\\sigma}}{1-\\sigma},\\,\\,\\,\\beta > -1, \\sigma > 0\\\\\n & \\text{s.t.}\\\\\n & S_{t}=s_{t}(1-\\tau_{w})w_{t}\\\\\n & C_{1t}=(1-s_{t})(1-\\tau_{w})w_{t}\\\\\n & C_{2t+1}=(1+(1-\\tau_{r})r_{t+1})S_{t}\n\\end{aligned}\n$$\n \nThe problem is formulated in terms of the saving rate $s_t\\in[0,1]$.",
"_____no_output_____"
],
[
"**Firms:** Firms rent capital $K_{t-1}$ at the rental rate $r_{t}^{K}$,\nand hires labor $E_{t}$ at the wage rate $w_{t}$. Firms have access\nto the production function\n \n$$\n\\begin{aligned}\nY_{t}=F(K_{t-1},E_{t})=(\\alpha K_{t-1}^{-\\theta}+(1-\\alpha)E_{t}^{-\\theta})^{\\frac{1}{-\\theta}},\\,\\,\\,\\theta>-1,\\alpha\\in(0,1)\n\\end{aligned}\n$$\n \nProfits are\n \n$$\n\\begin{aligned}\n\\Pi_{t}=Y_{t}-w_{t}E_{t}-r_{t}^{K}K_{t-1}\n\\end{aligned}\n$$",
"_____no_output_____"
],
[
"**Government:** Choose public consumption, $G_{t}$, and tax rates $\\tau_w \\in [0,1]$ and $\\tau_r \\in [0,1]$. Total tax revenue is\n \n$$\n\\begin{aligned}\nT_{t} &=\\tau_r r_{t} (K_{t-1}+B_{t-1})+\\tau_w w_{t} \n\\end{aligned}\n$$\n \nGovernment debt accumulates according to\n \n$$\n\\begin{aligned}\nB_{t} &=(1+r^b_{t})B_{t-1}-T_{t}+G_{t} \n\\end{aligned}\n$$\n \nA *balanced budget* implies $G_{t}=T_{t}-r_{t}B_{t-1}$.",
"_____no_output_____"
],
[
"**Capital:** Depreciates with a rate of $\\delta \\in [0,1]$.",
"_____no_output_____"
],
[
"**Equilibrium:**\n\n1. Households maximize utility\n2. Firms maximize profits\n3. No-arbitrage between bonds and capital\n\n $$\n r_{t}=r_{t}^{K}-\\delta=r_{t}^{b}\n $$\n\n4. Labor market clears: $E_{t}=L_{t}=1$\n5. Goods market clears: $Y_{t}=C_{1t}+C_{2t}+G_{t}+I_{t}$\n6. Asset market clears: $S_{t}=K_{t}+B_{t}$\n7. Capital follows its law of motion: $K_{t}=(1-\\delta)K_{t-1}+I_{t}$",
"_____no_output_____"
],
[
"**For more details on the OLG model:** See chapter 3-4 [here](https://web.econ.ku.dk/okocg/VM/VM-general/Material/Chapters-VM.htm).",
"_____no_output_____"
],
[
"## 1.2 Solution and simulation",
"_____no_output_____"
],
[
"**Implication of profit maximization:** From FOCs\n\n$$\n\\begin{aligned}\nr_{t}^{k} & =F_{K}(K_{t-1},E_{t})=\\alpha K_{t-1}^{-\\theta-1}Y_{t}^{1+\\theta}\\\\\nw_{t} & =F_{E}(K_{t-1},E_{t})=(1-\\alpha)E_{t}^{-\\theta-1}Y_{t}^{1+\\theta}\n\\end{aligned}\n$$\n\n**Implication of utility maximization:** From FOC\n\n$$\n\\begin{aligned}\nC_{1t}^{-\\sigma}=\\beta (1+(1-\\tau_r)r_{t+1})C_{2t+1}^{-\\sigma}\n\\end{aligned}\n$$",
"_____no_output_____"
],
[
"**Simulation algorithm:** At the beginning of period $t$, the\neconomy can be summarized in the state variables $K_{t-1}$ and $B_{t-1}$. *Before* $s_t$ is known, we can calculate:\n\n$$\n\\begin{aligned}\nY_{t} & =F(K_{t-1},1)\\\\\nr_{t}^{k} & =F_{K}(K_{t-1},1)\\\\\nw_{t} & =F_{E}(K_{t-1},1)\\\\\nr_{t} & =r^k_{t}-\\delta\\\\\nr_{t}^{b} & =r_{t}\\\\\n\\tilde{r}_{t} & =(1-\\tau_{r})r_{t}\\\\\nC_{2t} & =(1+\\tilde{r}_{t})(K_{t-1}+B_{t-1})\\\\\nT_{t} & =\\tau_{r}r_{t}(K_{t-1}+B_{t-1})+\\tau_{w}w_{t}\\\\\nB_{t} & =(1+r^b_{t})B_{t-1}+T_{t}-G_{t}\\\\\n\\end{aligned}\n$$\n\n*After* $s_t$ is known we can calculate:\n\n$$\n\\begin{aligned}\nC_{1t} & = (1-s_{t})(1-\\tau_{w})w_{t}\\\\\nI_{t} & =Y_{t}-C_{1t}-C_{2t}-G_{t}\\\\\nK_{t} & =(1-\\delta)K_{t-1} + I_t\n\\end{aligned}\n$$",
"_____no_output_____"
],
[
"**Solution algorithm:** Simulate forward choosing $s_{t}$ so\nthat we always have\n\n$$\n\\begin{aligned}\nC_{1t}^{-\\sigma}=\\beta(1+\\tilde{r}_{t+1})C_{2t+1}^{-\\sigma}\n\\end{aligned}\n$$",
"_____no_output_____"
],
[
"**Implementation:**\n\n1. Use a bisection root-finder to determine $s_t$\n2. Low $s_t$: A lot of consumption today. Low marginal utility. LHS < RHS.\n3. High $s_t$: Little consumption today. High marginal utility. LHS > RHS.\n4. Problem: Too low $s_t$ might not be feasible if $B_t > 0$.\n\n**Note:** Never errors in the Euler-equation due to *perfect foresight*.",
"_____no_output_____"
],
[
"**Question:** Are all the requirements for the equilibrium satisfied?",
"_____no_output_____"
],
[
"## 1.3 Test case",
"_____no_output_____"
],
[
"1. Production is Cobb-Douglas ($\\theta = 0$)\n2. Utility is logarithmic ($\\sigma = 1$)\n3. The government is not doing anything ($\\tau_w=\\tau_r=0$, $T_t = G_t = 0$ and $B_t = 0$) \n\n**Analytical steady state:** It can be proven\n\n$$ \\lim_{t\\rightarrow\\infty} K_t = \\left(\\frac{1-\\alpha}{1+1/\\beta}\\right)^{\\frac{1}{1-\\alpha}} $$",
"_____no_output_____"
],
[
"**Setup:**",
"_____no_output_____"
]
],
[
[
"model = OLGModelClass()\npar = model.par # SimpeNamespace\nsim = model.sim # SimpeNamespace\n\n# a. production\npar.production_function = 'cobb-douglas'\npar.theta = 0.0\n\n# b. households\npar.sigma = 1.0\n\n# c. government\npar.tau_w = 0.0\npar.tau_r = 0.0\nsim.balanced_budget[:] = True # G changes to achieve this\n\n# d. initial values\nK_ss = ((1-par.alpha)/((1+1.0/par.beta)))**(1/(1-par.alpha))\npar.K_lag_ini = 0.1*K_ss",
"initializing the model:\ncalling .setup()\ncalling .allocate()\n"
]
],
[
[
"### Simulate first period manually",
"_____no_output_____"
]
],
[
[
"from OLGModel import simulate_before_s, simulate_after_s, find_s_bracket, calc_euler_error",
"_____no_output_____"
]
],
[
[
"**Make a guess:**",
"_____no_output_____"
]
],
[
[
"s_guess = 0.41",
"_____no_output_____"
]
],
[
[
"**Evaluate first period:**",
"_____no_output_____"
]
],
[
[
"# a. initialize\nsim.K_lag[0] = par.K_lag_ini\nsim.B_lag[0] = par.B_lag_ini\n\nsimulate_before_s(par,sim,t=0)\nprint(f'{sim.C2[0] = : .4f}')\n\nsimulate_after_s(par,sim,s=s_guess,t=0)\nprint(f'{sim.C1[0] = : .4f}')\n\nsimulate_before_s(par,sim,t=1)\nprint(f'{sim.C2[1] = : .4f}')\nprint(f'{sim.rt[1] = : .4f}')\n\nLHS_Euler = sim.C1[0]**(-par.sigma)\nRHS_Euler = (1+sim.rt[1])*par.beta * sim.C2[1]**(-par.sigma)\nprint(f'euler-error = {LHS_Euler-RHS_Euler:.8f}')",
"sim.C2[0] = 0.0973\nsim.C1[0] = 0.1221\nsim.C2[1] = 0.1855\nsim.rt[1] = 1.1871\neuler-error = -0.22834540\n"
]
],
[
[
"**Implemented as function:**",
"_____no_output_____"
]
],
[
[
"euler_error = calc_euler_error(s_guess,par,sim,t=0)\nprint(f'euler-error = {euler_error:.8f}')",
"euler-error = -0.22834540\n"
]
],
[
[
"**Find bracket to search in:**",
"_____no_output_____"
]
],
[
[
"s_min,s_max = find_s_bracket(par,sim,t=0,do_print=True);",
"euler-error for s = 0.99999999 = 483321577.17005599\neuler-error for s = 0.50000000 = 2.76183762\neuler-error for s = 0.25000001 = -7.36489999\nbracket to search in with opposite signed errors:\n[ 0.25000001- 0.50000000]\n"
]
],
[
[
"**Call root-finder:**",
"_____no_output_____"
]
],
[
[
"obj = lambda s: calc_euler_error(s,par,sim,t=0)\nresult = optimize.root_scalar(obj,bracket=(s_min,s_max),method='bisect')\nprint(result)",
" converged: True\n flag: 'converged'\n function_calls: 39\n iterations: 37\n root: 0.41666666666653274\n"
]
],
[
[
"**Check result:**",
"_____no_output_____"
]
],
[
[
"euler_error = calc_euler_error(result.root,par,sim,t=0)\nprint(f'euler-error = {euler_error:.8f}')",
"euler-error = -0.00000000\n"
]
],
[
[
"### Full simulation",
"_____no_output_____"
]
],
[
[
"model.simulate()",
"simulation done in 0.06 secs\n"
]
],
[
[
"**Check euler-errors:**",
"_____no_output_____"
]
],
[
[
"for t in range(5):\n LHS_Euler = sim.C1[t]**(-par.sigma)\n RHS_Euler = (1+sim.rt[t+1])*par.beta * sim.C2[t+1]**(-par.sigma)\n print(f't = {t:2d}: euler-error = {LHS_Euler-RHS_Euler:.8f}')",
"t = 0: euler-error = -0.00000000\nt = 1: euler-error = -0.00000000\nt = 2: euler-error = -0.00000000\nt = 3: euler-error = -0.00000000\nt = 4: euler-error = -0.00000000\n"
]
],
[
[
"**Plot and check with analytical solution:**",
"_____no_output_____"
]
],
[
[
"fig = plt.figure(figsize=(6,6/1.5))\nax = fig.add_subplot(1,1,1)\nax.plot(model.sim.K_lag,label=r'$K_{t-1}$')\nax.axhline(K_ss,ls='--',color='black',label='analytical steady state')\nax.legend(frameon=True)\nfig.tight_layout()\nK_lag_old = model.sim.K_lag.copy()",
"_____no_output_____"
]
],
[
[
"**Task:** Test if the starting point matters?",
"_____no_output_____"
],
[
"**Additional check:** Not much should change with only small parameter changes. ",
"_____no_output_____"
]
],
[
[
"# a. production (close to cobb-douglas)\npar.production_function = 'ces'\npar.theta = 0.001\n\n# b. household (close to logarithmic)\npar.sigma = 1.1\n\n# c. goverment (weakly active)\npar.tau_w = 0.001\npar.tau_r = 0.001\n\n# d. simulate\nmodel.simulate()",
"simulation done in 0.06 secs\n"
],
[
"fig = plt.figure(figsize=(6,6/1.5))\nax = fig.add_subplot(1,1,1)\nax.plot(model.sim.K_lag,label=r'$K_{t-1}$')\nax.plot(K_lag_old,label=r'$K_{t-1}$ ($\\theta = 0.0, \\sigma = 1.0$, inactive government)')\nax.axhline(K_ss,ls='--',color='black',label='analytical steady state (wrong)')\nax.legend(frameon=True)\nfig.tight_layout()",
"_____no_output_____"
]
],
[
[
"## 1.4 Active government",
"_____no_output_____"
]
],
[
[
"model = OLGModelClass()\npar = model.par\nsim = model.sim",
"initializing the model:\ncalling .setup()\ncalling .allocate()\n"
]
],
[
[
"**Baseline:**",
"_____no_output_____"
]
],
[
[
"model.simulate()",
"simulation done in 0.02 secs\n"
],
[
"fig = plt.figure(figsize=(6,6/1.5))\nax = fig.add_subplot(1,1,1)\nax.plot(sim.K_lag/(sim.Y),label=r'$\\frac{K_{t-1}}{Y_t}$')\nax.plot(sim.B_lag/(sim.Y),label=r'$\\frac{B_{t-1}}{Y_t}$')\nax.legend(frameon=True)\nfig.tight_layout()",
"_____no_output_____"
]
],
[
[
"**Remember steady state:**",
"_____no_output_____"
]
],
[
[
"K_ss = sim.K_lag[-1]\nB_ss = sim.B_lag[-1]\nG_ss = sim.G[-1]",
"_____no_output_____"
]
],
[
[
"**Spending spree of 5% in $T=3$ periods:**",
"_____no_output_____"
]
],
[
[
"# a. start from steady state\npar.K_lag_ini = K_ss\npar.B_lag_ini = B_ss\n\n# b. spending spree\nT0 = 0\ndT = 3\nsim.G[T0:T0+dT] = 1.05*G_ss\nsim.balanced_budget[:T0] = True #G adjusts\nsim.balanced_budget[T0:T0+dT] = False # B adjusts\nsim.balanced_budget[T0+dT:] = True # G adjusts",
"_____no_output_____"
]
],
[
[
"**Simulate:**",
"_____no_output_____"
]
],
[
[
"model.simulate()",
"simulation done in 0.07 secs\n"
]
],
[
[
"**Crowding-out of capital:**",
"_____no_output_____"
]
],
[
[
"fig = plt.figure(figsize=(6,6/1.5))\nax = fig.add_subplot(1,1,1)\nax.plot(sim.K/(sim.Y),label=r'$\\frac{K_{t-1}}{Y_t}$')\nax.plot(sim.B/(sim.Y),label=r'$\\frac{B_{t-1}}{Y_t}$')\nax.legend(frameon=True)\nfig.tight_layout()",
"_____no_output_____"
]
],
[
[
"**Question:** Would the households react today if the spending spree is say 10 periods in the future? ",
"_____no_output_____"
],
[
"## 1.5 Getting an overview",
"_____no_output_____"
],
[
"1. Spend 3 minutes looking at `OLGModel.py`\n2. Write one question at [https://b.socrative.com/login/student/](https://b.socrative.com/login/student/) with `ROOM=NUMECON`",
"_____no_output_____"
],
[
"## 1.6 Potential analysis and extension",
"_____no_output_____"
],
[
"**Potential analysis:**\n\n1. Over-accumulation of capital relative to golden rule?\n2. Calibration to actual data\n3. Generational inequality\n4. Multiple equilibria",
"_____no_output_____"
],
[
"**Extensions:**\n\n1. Add population and technology growth\n2. More detailed tax and transfer system\n3. Utility and productive effect of government consumption/investment\n4. Endogenous labor supply\n5. Bequest motive\n6. Uncertain returns on capital\n7. Additional assets (e.g. housing)\n8. More than two periods in the life-cycle (life-cycle)\n9. More than one dynasty (cross-sectional inequality dynamics)",
"_____no_output_____"
],
[
"<a id=\"Ramsey-model\"></a>\n\n# 2. Ramsey model",
"_____no_output_____"
],
[
"... also called the Ramsey-Cass-Koopman model.",
"_____no_output_____"
],
[
"## 2.1 Model descripton",
"_____no_output_____"
],
[
"**Time:** Discrete and indexed by $t\\in\\{0,1,\\dots\\}$.",
"_____no_output_____"
],
[
"**Demographics::** Population is constant. Everybody lives forever.",
"_____no_output_____"
],
[
"**Household:** Households supply labor exogenously, $L_{t}=1$, and earns a wage $w_{t}$. The return on saving is $r_{t+1}$. Utility is \n\n$$\n\\begin{aligned}\nU & =\\max_{\\{C_{t}\\}_{t=0}^{\\infty}}\\sum_{t=0}^{\\infty}\\beta^{t}\\frac{C_{t}^{1-\\sigma}}{1-\\sigma},\\beta\\in(0,1),\\sigma>0\\\\\n & \\text{s.t.}\\\\\n & M_{t}=(1+r_{t})N_{t-1}+w_{t}\\\\\n & N_{t}=M_{t}-C_{t}\n\\end{aligned}\n$$\n\nwhere $M_{t}$ is cash-on-hand and $N_{t}$ is end-of-period assets.",
"_____no_output_____"
],
[
"**Firms:** Firms rent capital $K_{t-1}$ at the rental rate $r_{t}^{K}$\nand hires labor $E_{t}$ at the wage rate $w_{t}$. Firms have access\nto the production function\n \n$$\n\\begin{aligned}\nY_{t}= F(K_{t-1},E_{t})=A_t(\\alpha K_{t-1}^{-\\theta}+(1-\\alpha)E_{t}^{-\\theta})^{\\frac{1}{-\\theta}},\\,\\,\\,\\theta>-1,\\alpha\\in(0,1),A_t>0\n\\end{aligned}\n$$\n\nProfits are\n\n$$\n\\begin{aligned}\n\\Pi_{t}=Y_{t}-w_{t}E_{t}-r_{t}^{K}K_{t-1}\n\\end{aligned}\n$$",
"_____no_output_____"
],
[
"**Equilibrium:**\n\n1. Households maximize utility\n2. Firms maximize profits\n3. Labor market clear: $E_{t}=L_{t}=1$\n4. Goods market clear: $Y_{t}=C_{t}+I_{t}$\n5. Asset market clear: $N_{t}=K_{t}$ and $r_{t}=r_{t}^{k}-\\delta$\n6. Capital follows its law of motion: $K_{t}=(1-\\delta)K_{t-1}+I_{t}$\n\n**Implication of profit maximization:** From FOCs\n\n$$\n\\begin{aligned}\nr_{t}^{k} & = F_{K}(K_{t-1},E_{t})=A_t \\alpha K_{t-1}^{-\\theta-1}Y_{t}^{-1}\\\\\nw_{t} & = F_{E}(K_{t-1},E_{t})=A_t (1-\\alpha)E_{t}^{-\\theta-1}Y_{t}^{-1}\n\\end{aligned}\n$$\n\n**Implication of utility maximization:** From FOCs\n\n$$\n\\begin{aligned}\nC_{t}^{-\\sigma}=\\beta(1+r_{t+1})C_{t+1}^{-\\sigma}\n\\end{aligned}\n$$",
"_____no_output_____"
],
[
"**Solution algorithm:** \n\nWe can summarize the model in the **non-linear equation system**\n\n$$\n\\begin{aligned}\n\\boldsymbol{H}(\\boldsymbol{K},\\boldsymbol{C},K_{-1})=\\left[\\begin{array}{c}\nH_{0}\\\\\nH_{1}\\\\\n\\begin{array}{c}\n\\vdots\\end{array}\n\\end{array}\\right]=\\left[\\begin{array}{c}\n0\\\\\n0\\\\\n\\begin{array}{c}\n\\vdots\\end{array}\n\\end{array}\\right]\n\\end{aligned}\n$$\n\nwhere $\\boldsymbol{K} = [K_0,K_1\\dots]$, $\\boldsymbol{C} = [C_0,C_1\\dots]$, and\n\n$$\n\\begin{aligned}\nH_{t}\n=\\left[\\begin{array}{c}\nC_{t}^{-\\sigma}-\\beta(1+r_{t+1})C_{t+1}^{-\\sigma}\\\\\nK_{t}-[(1-\\delta)K_{t-1}+Y_t-C_{t}]\n\\end{array}\\right]\n=\\left[\\begin{array}{c}\nC_{t}^{-\\sigma}-\\beta(1+F_{K}(K_{t},1))C_{t+1}^{-\\sigma}\\\\\nK_{t}-[(1-\\delta)K_{t-1} + F(K_{t-1},1)-C_{t}])\n\\end{array}\\right]\n\\end{aligned}\n$$\n\n**Path:** We refer to $\\boldsymbol{K}$ and $\\boldsymbol{C}$ as *transition paths*.",
"_____no_output_____"
],
[
"**Implementation:** We solve this equation system in **two steps**:\n\n1. Assume all variables are in steady state after some **truncation horizon**.\n1. Calculate the numerical **jacobian** of $\\boldsymbol{H}$ wrt. $\\boldsymbol{K}$\nand $\\boldsymbol{C}$ around the steady state\n2. Solve the equation system using a **hand-written Broyden-solver**\n\n**Note:** The equation system can also be solved directly using `scipy.optimize.root`.\n\n**Remember:** The jacobian is just a gradient. I.e. the matrix of what the implied errors are in $\\boldsymbol{H}$ when a *single* $K_t$ or $C_t$ change.",
"_____no_output_____"
],
[
"## 2.2 Solution",
"_____no_output_____"
]
],
[
[
"model = RamseyModelClass()\npar = model.par\nss = model.ss\npath = model.path",
"initializing the model:\ncalling .setup()\ncalling .allocate()\n"
]
],
[
[
"**Find steady state:** \n\n1. Target steady-state capital-output ratio, $K_{ss}/Y_{ss}$ of 4.0.\n2. Force steady-state output $Y_{ss} = 1$.\n3. Adjust $\\beta$ and $A_{ss}$ to achieve this.",
"_____no_output_____"
]
],
[
[
"model.find_steady_state(KY_ss=4.0)",
"Y_ss = 1.0000\nK_ss/Y_ss = 4.0000\nrk_ss = 0.0750\nr_ss = 0.0250\nw_ss = 0.7000\nA = 0.6598\nbeta = 0.9756\n"
]
],
[
[
"**Test that errors and the path are 0:**",
"_____no_output_____"
]
],
[
[
"# a. set initial value\npar.K_lag_ini = ss.K\n\n# b. set path\npath.A[:] = ss.A\npath.C[:] = ss.C\npath.K[:] = ss.K\n\n# c. check errors\nerrors_ss = model.evaluate_path_errors()\nassert np.allclose(errors_ss,0.0)",
"_____no_output_____"
],
[
"model.calculate_jacobian()",
"_____no_output_____"
]
],
[
[
"**Solve:**",
"_____no_output_____"
]
],
[
[
"par.K_lag_ini = 0.50*ss.K # start away from steady state\nmodel.solve() # find transition path",
" it = 0 -> max. abs. error = 2.08774760\n it = 1 -> max. abs. error = 0.03407048\n it = 2 -> max. abs. error = 0.04084472\n it = 3 -> max. abs. error = 0.00495803\n it = 4 -> max. abs. error = 0.01354190\n it = 5 -> max. abs. error = 0.01209108\n it = 6 -> max. abs. error = 0.00397825\n it = 7 -> max. abs. error = 0.00192043\n it = 8 -> max. abs. error = 0.00097483\n it = 9 -> max. abs. error = 0.00009018\n it = 10 -> max. abs. error = 0.00010485\n it = 11 -> max. abs. error = 0.00000476\n it = 12 -> max. abs. error = 0.00000737\n it = 13 -> max. abs. error = 0.00000045\n it = 14 -> max. abs. error = 0.00000038\n it = 15 -> max. abs. error = 0.00000002\n it = 16 -> max. abs. error = 0.00000002\n it = 17 -> max. abs. error = 0.00000000\n"
],
[
"fig = plt.figure(figsize=(6,6/1.5))\nax = fig.add_subplot(1,1,1)\nax.plot(path.K_lag,label=r'$K_{t-1}$')\nax.legend(frameon=True)\nfig.tight_layout()",
"_____no_output_____"
]
],
[
[
"## 2.3 Comparison with scipy solution",
"_____no_output_____"
],
[
"**Note:** scipy computes the jacobian internally",
"_____no_output_____"
]
],
[
[
"model_scipy = RamseyModelClass()\nmodel_scipy.par.solver = 'scipy'\nmodel_scipy.find_steady_state(KY_ss=4.0)\nmodel_scipy.par.K_lag_ini = 0.50*model_scipy.ss.K\nmodel_scipy.path.A[:] = model_scipy.ss.A\nmodel_scipy.solve()",
"initializing the model:\ncalling .setup()\ncalling .allocate()\nY_ss = 1.0000\nK_ss/Y_ss = 4.0000\nrk_ss = 0.0750\nr_ss = 0.0250\nw_ss = 0.7000\nA = 0.6598\nbeta = 0.9756\n"
],
[
"fig = plt.figure(figsize=(6,6/1.5))\nax = fig.add_subplot(1,1,1)\nax.plot(path.K_lag,label=r'$K_{t-1}$, broyden')\nax.plot(model_scipy.path.K_lag,ls='--',label=r'$K_{t-1}$, scipy')\nax.legend(frameon=True)\nfig.tight_layout()",
"_____no_output_____"
]
],
[
[
"## 2.4 Persistent technology shock",
"_____no_output_____"
],
[
"**Shock:**",
"_____no_output_____"
]
],
[
[
"par.K_lag_ini = ss.K # start from steady state\npath.A[:] = 0.95**np.arange(par.Tpath)*0.1*ss.A + ss.A # shock path",
"_____no_output_____"
]
],
[
[
"**Terminology:** This is called an MIT-shock. Households do not expect shocks. Know the full path of the shock when it arrives. Continue to believe no future shocks will happen.",
"_____no_output_____"
],
[
"**Solve:**",
"_____no_output_____"
]
],
[
[
"model.solve()",
" it = 0 -> max. abs. error = 0.10000000\n it = 1 -> max. abs. error = 0.00096551\n it = 2 -> max. abs. error = 0.00004937\n it = 3 -> max. abs. error = 0.00000248\n it = 4 -> max. abs. error = 0.00000040\n it = 5 -> max. abs. error = 0.00000006\n it = 6 -> max. abs. error = 0.00000000\n"
],
[
"fig = plt.figure(figsize=(2*6,6/1.5))\n\nax = fig.add_subplot(1,2,1)\nax.set_title('Capital, $K_{t-1}$')\nax.plot(path.K_lag)\n\nax = fig.add_subplot(1,2,2)\nax.plot(path.A)\nax.set_title('Technology, $A_t$')\n\nfig.tight_layout()",
"_____no_output_____"
]
],
[
[
"**Question:** Could a much more persistent shock be problematic?",
"_____no_output_____"
],
[
"## 2.5 Future persistent technology shock",
"_____no_output_____"
],
[
"**Shock happing after period $H$:**",
"_____no_output_____"
]
],
[
[
"par.K_lag_ini = ss.K # start from steady state\n\n# shock\nH = 50\npath.A[:] = ss.A\npath.A[H:] = 0.95**np.arange(par.Tpath-H)*0.1*ss.A + ss.A",
"_____no_output_____"
]
],
[
[
"**Solve:**",
"_____no_output_____"
]
],
[
[
"model.solve()",
" it = 0 -> max. abs. error = 0.10000000\n it = 1 -> max. abs. error = 0.00267237\n it = 2 -> max. abs. error = 0.00015130\n it = 3 -> max. abs. error = 0.00000241\n it = 4 -> max. abs. error = 0.00000025\n it = 5 -> max. abs. error = 0.00000002\n it = 6 -> max. abs. error = 0.00000000\n"
],
[
"fig = plt.figure(figsize=(2*6,6/1.5))\n\nax = fig.add_subplot(1,2,1)\nax.set_title('Capital, $K_{t-1}$')\nax.plot(path.K_lag)\n\nax = fig.add_subplot(1,2,2)\nax.plot(path.A)\nax.set_title('Technology, $A_t$')\n\nfig.tight_layout()",
"_____no_output_____"
],
[
"par.K_lag_ini = path.K[30]\npath.A[:] = ss.A\nmodel.solve()",
" it = 0 -> max. abs. error = 0.05301739\n it = 1 -> max. abs. error = 0.00001569\n it = 2 -> max. abs. error = 0.00000010\n it = 3 -> max. abs. error = 0.00000000\n"
]
],
[
[
"**Take-away:** Households are forward looking and responds before the shock hits.",
"_____no_output_____"
],
[
"## 2.6 Getting an overview",
"_____no_output_____"
],
[
"1. Spend 3 minutes looking at `RamseyModel.py`\n2. Write one question at [https://b.socrative.com/login/student/](https://b.socrative.com/login/student/) with `ROOM=NUMECON`",
"_____no_output_____"
],
[
"## 2.7 Potential analysis and extension",
"_____no_output_____"
],
[
"**Potential analysis:**\n\n1. Different shocks (e.g. discount factor)\n2. Multiple shocks\n3. Permanent shocks ($\\rightarrow$ convergence to new steady state)\n4. Transition speed",
"_____no_output_____"
],
[
"**Extensions:**\n\n1. Add a government and taxation\n2. Endogenous labor supply\n3. Additional assets (e.g. housing)\n4. Add nominal rigidities (New Keynesian)",
"_____no_output_____"
],
[
"<a id=\"Further-perspectives\"></a>\n\n# 3. Further perspectives",
"_____no_output_____"
],
[
"**The next steps beyond this course:** \n\n1. The **Bewley-Huggett-Aiyagari** model. A multi-period OLG model or Ramsey model with households making decisions *under uncertainty and borrowing constraints* as in lecture 11 under \"dynamic optimization\". Such heterogenous agent models are used in state-of-the-art research, see [Quantitative Macroeconomics with Heterogeneous Households](https://www.annualreviews.org/doi/abs/10.1146/annurev.economics.050708.142922). \n\n2. Further adding nominal rigidities this is called a **Heterogenous Agent New Keynesian (HANK)** model. See [Macroeconomics with HANK models](https://drive.google.com/file/d/16Qq7NJ_AZh5NmjPFSrLI42mfT7EsCUeH/view).\n\n3. This extends the **Representative Agent New Keynesian (RANK)** model, which itself is a Ramsey model extended with nominal rigidities. \n\n4. The final frontier is including **aggregate risk**, which either requires linearization or using a **Krussell-Smith method**. Solving the model in *sequence-space* as we did with the Ramsey model is a frontier method (see [here](https://github.com/shade-econ/sequence-jacobian/#sequence-space-jacobian)).\n\n\n\n",
"_____no_output_____"
],
[
"**Next lecture:** Agent Based Models",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
4acd7343fe8217a3ca205d2d79062a31fe53d4e3
| 5,703 |
ipynb
|
Jupyter Notebook
|
examples/kubeflow_metadata_example.ipynb
|
google/nitroml
|
5eabdbe6de85ff7fdae4fefda7547c0c031f9431
|
[
"Apache-2.0"
] | 43 |
2020-09-13T18:07:15.000Z
|
2022-01-05T19:05:28.000Z
|
examples/kubeflow_metadata_example.ipynb
|
google/nitroml
|
5eabdbe6de85ff7fdae4fefda7547c0c031f9431
|
[
"Apache-2.0"
] | 4 |
2020-09-14T13:15:09.000Z
|
2021-11-21T11:21:13.000Z
|
examples/kubeflow_metadata_example.ipynb
|
google/nitroml
|
5eabdbe6de85ff7fdae4fefda7547c0c031f9431
|
[
"Apache-2.0"
] | 5 |
2020-09-14T13:03:04.000Z
|
2021-10-21T01:55:48.000Z
| 27.418269 | 137 | 0.590566 |
[
[
[
"##### Copyright 2020 Google LLC.",
"_____no_output_____"
],
[
"#@title Licensed under the Apache License, Version 2.0 (the \"License\");\n# you may not use this file except in compliance with the License.\n# You may obtain a copy of the License at\n#\n# https://www.apache.org/licenses/LICENSE-2.0\n#\n# Unless required by applicable law or agreed to in writing, software\n# distributed under the License is distributed on an \"AS IS\" BASIS,\n# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.\n# See the License for the specific language governing permissions and\n# limitations under the License.",
"_____no_output_____"
]
],
[
[
"# We are using NitroML on Kubeflow: \n\nThis notebook allows users to analyze NitroML benchmark results.",
"_____no_output_____"
]
],
[
[
"# This notebook assumes you have followed the following steps to setup port-forwarding:\n\n# Step 1: Configure your cluster with gcloud\n# `gcloud container clusters get-credentials <cluster_name> --zone <cluster-zone> --project <project-id>\n\n# Step 2: Get the port where the gRPC service is running on the cluster\n# `kubectl get configmap metadata-grpc-configmap -o jsonpath={.data}`\n# Use `METADATA_GRPC_SERVICE_PORT` in the next step. The default port used is 8080.\n\n# Step 3: Port forwarding\n# `kubectl port-forward deployment/metadata-grpc-deployment 9898:<METADATA_GRPC_SERVICE_PORT>`\n\n# Troubleshooting\n# If getting error related to Metadata (For examples, Transaction already open). Try restarting the metadata-grpc-service using:\n# `kubectl rollout restart deployment metadata-grpc-deployment` ",
"_____no_output_____"
],
[
"import sys, os\nPROJECT_DIR=os.path.join(sys.path[0], '..')\n%cd {PROJECT_DIR}",
"_____no_output_____"
],
[
"from ml_metadata.proto import metadata_store_pb2\nfrom ml_metadata.metadata_store import metadata_store\nfrom nitroml.benchmark import results",
"_____no_output_____"
]
],
[
[
"## Connect to the ML Metadata (MLMD) database\n\nFirst we need to connect to our MLMD database which stores the results of our\nbenchmark runs.",
"_____no_output_____"
]
],
[
[
"connection_config = metadata_store_pb2.MetadataStoreClientConfig()\n\nconnection_config.host = 'localhost'\nconnection_config.port = 9898\n\nstore = metadata_store.MetadataStore(connection_config)",
"_____no_output_____"
]
],
[
[
"## Display benchmark results\n\nNext we load and visualize `pd.DataFrame` containing our benchmark results.\nThese results contain contextual features such as the pipeline ID, and \nbenchmark metrics as computed by the downstream Evaluators. If your\nbenchmark included an `EstimatorTrainer` component, its hyperparameters may also\ndisplay in the table below.",
"_____no_output_____"
]
],
[
[
"#@markdown ### Choose how to aggregate metrics:\nmean = False #@param { type: \"boolean\" }\nstdev = False #@param { type: \"boolean\" }\nmin_and_max = False #@param { type: \"boolean\" }\n\nagg = []\nif mean:\n agg.append(\"mean\")\nif stdev:\n agg.append(\"std\")\nif min_and_max:\n agg += [\"min\", \"max\"]\n\ndf = results.overview(store, metric_aggregators=agg)",
"_____no_output_____"
],
[
"df.head()",
"_____no_output_____"
]
],
[
[
"### We can display an interactive table using qgrid\n\nPlease follow the latest instructions on downloading qqgrid package from here: https://github.com/quantopian/qgrid",
"_____no_output_____"
]
],
[
[
"import qgrid\nqgid_wdget = qgrid.show_grid(df, show_toolbar=True)\nqgid_wdget",
"_____no_output_____"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
]
] |
4acd9e646d2c0a2201725cd0a566264c90b3fa98
| 2,603 |
ipynb
|
Jupyter Notebook
|
murphy-book/chapter04/q02.ipynb
|
yusueliu/murphy-book
|
71d62cc083a683fb861be1e5acb8eeb948b00c54
|
[
"Apache-2.0"
] | 2 |
2019-03-25T22:22:23.000Z
|
2019-09-29T20:46:58.000Z
|
murphy-book/chapter04/q02.ipynb
|
yusueliu/murphy-book
|
71d62cc083a683fb861be1e5acb8eeb948b00c54
|
[
"Apache-2.0"
] | null | null | null |
murphy-book/chapter04/q02.ipynb
|
yusueliu/murphy-book
|
71d62cc083a683fb861be1e5acb8eeb948b00c54
|
[
"Apache-2.0"
] | 1 |
2021-12-24T01:14:12.000Z
|
2021-12-24T01:14:12.000Z
| 26.292929 | 138 | 0.472916 |
[
[
[
"## Exercise 4.2 Uncorrelated and Gaussian does not imply independent unless jointly Gaussian\nLet $X ∼ \\mathcal{N}(0,1)$ and $Y = WX$, where $p(W = −1) = p(W = 1) = 0.5$. It is clear that $X$ and $Y$ are\nnot independent, since $Y$ is a function of $X$. \n\na. Show $Y \\sim \\mathcal{N}(0,1)$.",
"_____no_output_____"
],
[
"$$\nP(Y\\le x) = \\mathbb{E}[P(Y \\le x |W)] = P(X\\le x)P(W = 1) + P(-X \\le x)P(W=-1) = \\Phi(x)\n$$\n\n$\\Phi(x)$ is the cumulative density function of the standard normal distribution.",
"_____no_output_____"
],
[
"**Second solution**\n\n$$\np(y) = p(wx) = p(\\mathrm{sign}(w) x)\n$$\n\nwhere $\\mathrm{sign}(w)$ is the sign function. The normal is always symmetrical around its mean, so we have $p(x) = p(-x)$, and \n\n$$\np(y) = p(\\mathrm{sign}(w)x) = p(x)\n$$",
"_____no_output_____"
],
[
"b. Show $\\mathrm{cov}[X, Y ] = 0$. Thus $X$ and $Y$ are uncorrelated but dependent, even though they are Gaussian. \n\nHint: use the definition of covariance \n\n$$\\mathrm{cov} [X, Y ] = \\mathbb{E} [XY ] − \\mathbb{E} [X] \\mathbb{E} [Y ]\n$$ \n\nand the **rule of iterated expectation**\n\n$$\n\\mathbb{E} [X Y ] = \\mathbb{E} [\\mathbb{E} [X Y |W ]]\n$$",
"_____no_output_____"
],
[
"**Solution**:\n\n\\begin{aligned}\n\\mathbb{E}[XY] & = \\mathbb{E}[\\mathbb{E}[XY|W]] = 0.5\\mathbb{E}[XY|W=-1] + 0.5\\mathbb{E}[XY|W=1] \\\\\n& = 0.5(\\mathbb{E}[-X^2] + \\mathbb{E}[X^2]) \\\\\n& = 0.5\\mathbb{E}[X^2 - X^2] = 0\n\\end{aligned}\n\n\nHence $\\mathrm{cov}[X, Y] = \\mathbb{E}[XY]-\\mathbb{E}[X]\\mathbb{E}[Y] = 0 \\implies \\rho(X, Y) = 0$",
"_____no_output_____"
]
]
] |
[
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
4acdabc441786b974fcde7074d47777b73ffbacb
| 6,865 |
ipynb
|
Jupyter Notebook
|
notebooks/Group Metrics.ipynb
|
janhavi13/fairlearn
|
042bc3bcc1c1fb0d59845b931b2be4c109a3f6ae
|
[
"MIT"
] | 1 |
2020-10-25T17:18:45.000Z
|
2020-10-25T17:18:45.000Z
|
notebooks/Group Metrics.ipynb
|
janhavi13/fairlearn
|
042bc3bcc1c1fb0d59845b931b2be4c109a3f6ae
|
[
"MIT"
] | 6 |
2021-03-11T00:38:07.000Z
|
2022-02-27T07:50:00.000Z
|
notebooks/Group Metrics.ipynb
|
janhavi13/fairlearn
|
042bc3bcc1c1fb0d59845b931b2be4c109a3f6ae
|
[
"MIT"
] | 1 |
2021-05-30T06:38:27.000Z
|
2021-05-30T06:38:27.000Z
| 34.154229 | 364 | 0.621996 |
[
[
[
"# Group Metrics\n\nThe `fairlearn` package contains algorithms which enable machine learning models to minimise disparity between groups. The `metrics` portion of the package provides the means required to verify that the mitigation algorithms are succeeding.",
"_____no_output_____"
]
],
[
[
"import numpy as np\nimport pandas as pd\nimport sklearn.metrics as skm",
"_____no_output_____"
]
],
[
[
"## Ungrouped Metrics\n\nAt their simplest, metrics take a set of 'true' values $Y_{true}$ (from the input data) and predicted values $Y_{pred}$ (by applying the model to the input data), and use these to compute a measure. For example, the _recall_ or _true positive rate_ is given by\n\\begin{equation}\nP( Y_{pred}=1 | Y_{true}=1 )\n\\end{equation}\nThat is, a measure of whether the model finds all the positive cases in the input data. The `scikit-learn` package implements this in [sklearn.metrics.recall_score](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.recall_score.html).\n\nSuppose we have the following data:",
"_____no_output_____"
]
],
[
[
"Y_true = [0, 1, 1, 1, 1, 0, 1, 0, 1, 0, 0, 0, 1, 1, 1, 1]\nY_pred = [0, 0, 1, 0, 1, 1, 1, 0, 0, 1, 1, 1, 1, 0, 0, 1]",
"_____no_output_____"
]
],
[
[
"we can see that the prediction is 1 in five of the ten cases where the true value is 1, so we expect the recall to be 0.0.5:",
"_____no_output_____"
]
],
[
[
"skm.recall_score(Y_true, Y_pred)",
"_____no_output_____"
]
],
[
[
"## Metrics with Grouping\n\nWhen considering fairness, each row of input data will have an associated group label $g \\in G$, and we will want to know how the metric behaves for each $g$. To help with this, Fairlearn provides wrappers, which take an existing (ungrouped) metric function, and apply it to each group within a set of data.\n\nSuppose in addition to the $Y_{true}$ and $Y_{pred}$ above, we had the following set of labels:",
"_____no_output_____"
]
],
[
[
"group_membership_data = ['d', 'a', 'c', 'b', 'b', 'c', 'c', 'c', 'b', 'd', 'c', 'a', 'b', 'd', 'c', 'c']\n\ndf = pd.DataFrame({ 'Y_true': Y_true, 'Y_pred': Y_pred, 'group_membership_data': group_membership_data})\ndf",
"_____no_output_____"
]
],
[
[
"We can see that for the groups 'a' and 'd' the recall is 0 (none of the true positives were identified), for 'b' the recall is 0.5 and for 'c' the recall is 0.75.\n\nThe `fairlearn.metrics.group_summary` routine can calculate all of these for us. This takes as arguments an _ungrouped_ metric (such as `sklearn.metrics.recall_score`), the arrays $Y_{true}$ and $Y_{pred}$ and an array of group labels (and optionally, if the ungrouped metric supports it, an array of sample weights), and produces `GroupMetricResult` object:",
"_____no_output_____"
]
],
[
[
"import fairlearn.metrics as flm\n\ngroup_metrics = flm.group_summary(skm.recall_score, Y_true, Y_pred, sensitive_features=group_membership_data, sample_weight=None)\n\nprint(\"Overall recall = \", group_metrics.overall)\nprint(\"recall by groups = \", group_metrics.by_group)",
"_____no_output_____"
]
],
[
[
"Note that the overall recall is the same as that calculated above in the Ungrouped Metric section, while the `by_group` dictionary matches the values we calculated by inspection from the table above.\n\nIn addition to these basic scores, `fairlearn.metrics` also provides convenience functions to recover the maximum and minimum values of the metric across groups and also the difference and ratio between the maximum and minimum:",
"_____no_output_____"
]
],
[
[
"print(\"min recall over groups = \", flm.group_min_from_summary(group_metrics))\nprint(\"max recall over groups = \", flm.group_max_from_summary(group_metrics))\nprint(\"difference in recall = \", flm.difference_from_summary(group_metrics))\nprint(\"ratio in recall = \", flm.ratio_from_summary(group_metrics))",
"_____no_output_____"
]
],
[
[
"## Supported Ungrouped Metrics\n\nTo be used by `group_summary`, the supplied Python function must take arguments of `y_true` and `y_pred`:\n```python\nmy_metric_func(y_true, y_pred)\n```\nAn additional argument of `sample_weight` is also supported:\n```python\nmy_metric_with_weight(y_true, y_pred, sample_weight=None)\n```\nThe `sample_weight` argument is always invoked by name, and _only_ if the user supplies a `sample_weight` argument.",
"_____no_output_____"
],
[
"## Convenience Wrapper\n\nRather than require a call to `group_summary` each time, we also provide a function which turns an ungrouped metric into a grouped one. This is called `make_metric_group_summary`:",
"_____no_output_____"
]
],
[
[
"recall_score_group_summary = flm.make_metric_group_summary(skm.recall_score)\n\nresults = recall_score_group_summary(Y_true, Y_pred, sensitive_features=group_membership_data)\n\nprint(\"Overall recall = \", results.overall)\nprint(\"recall by groups = \", results.by_group)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"raw",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"raw"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
]
] |
4acdb967d0588010dc3d4cac31c338db1f544eb2
| 79,376 |
ipynb
|
Jupyter Notebook
|
main.ipynb
|
thomle295/CartPole_RL
|
7bc4426473eabfb5f961b22778d1089a56b6aa67
|
[
"MIT"
] | null | null | null |
main.ipynb
|
thomle295/CartPole_RL
|
7bc4426473eabfb5f961b22778d1089a56b6aa67
|
[
"MIT"
] | null | null | null |
main.ipynb
|
thomle295/CartPole_RL
|
7bc4426473eabfb5f961b22778d1089a56b6aa67
|
[
"MIT"
] | null | null | null | 79.694779 | 19,227 | 0.69972 |
[
[
[
"<a href=\"https://colab.research.google.com/github/thomle295/CartPole_RL/blob/main/main.ipynb\" target=\"_parent\"><img src=\"https://colab.research.google.com/assets/colab-badge.svg\" alt=\"Open In Colab\"/></a>",
"_____no_output_____"
]
],
[
[
"from google.colab import drive\ndrive.mount('/content/gdrive', force_remount=True)",
"Mounted at /content/gdrive\n"
],
[
"%cd '/content/gdrive/My Drive/Research/Reinforcement_Learning/CartPole/'",
"/content/gdrive/My Drive/Research/Reinforcement_Learning/CartPole\n"
],
[
"import urllib.request\nurllib.request.urlretrieve('http://www.atarimania.com/roms/Roms.rar','Roms.rar')\n!pip install tensorflow==2.3.0\n!pip install gym\n!pip install keras\n!pip install keras-rl2\n!pip install unrar\n!unrar x Roms.rar\n!mkdir rars\n!mv HC\\ ROMS.zip rars\n!mv ROMS.zip rars\n!python -m atari_py.import_roms rars",
"Requirement already satisfied: tensorflow==2.3.0 in /usr/local/lib/python3.7/dist-packages (2.3.0)\nRequirement already satisfied: tensorflow-estimator<2.4.0,>=2.3.0 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (2.3.0)\nRequirement already satisfied: gast==0.3.3 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (0.3.3)\nRequirement already satisfied: opt-einsum>=2.3.2 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (3.3.0)\nRequirement already satisfied: termcolor>=1.1.0 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (1.1.0)\nRequirement already satisfied: google-pasta>=0.1.8 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (0.2.0)\nRequirement already satisfied: tensorboard<3,>=2.3.0 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (2.7.0)\nRequirement already satisfied: grpcio>=1.8.6 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (1.43.0)\nRequirement already satisfied: six>=1.12.0 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (1.15.0)\nRequirement already satisfied: absl-py>=0.7.0 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (0.12.0)\nRequirement already satisfied: wheel>=0.26 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (0.37.1)\nRequirement already satisfied: numpy<1.19.0,>=1.16.0 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (1.18.5)\nRequirement already satisfied: wrapt>=1.11.1 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (1.13.3)\nRequirement already satisfied: h5py<2.11.0,>=2.10.0 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (2.10.0)\nRequirement already satisfied: scipy==1.4.1 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (1.4.1)\nRequirement already satisfied: astunparse==1.6.3 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (1.6.3)\nRequirement already satisfied: keras-preprocessing<1.2,>=1.1.1 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (1.1.2)\nRequirement already satisfied: protobuf>=3.9.2 in /usr/local/lib/python3.7/dist-packages (from tensorflow==2.3.0) (3.17.3)\nRequirement already satisfied: werkzeug>=0.11.15 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow==2.3.0) (1.0.1)\nRequirement already satisfied: tensorboard-data-server<0.7.0,>=0.6.0 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow==2.3.0) (0.6.1)\nRequirement already satisfied: requests<3,>=2.21.0 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow==2.3.0) (2.23.0)\nRequirement already satisfied: google-auth-oauthlib<0.5,>=0.4.1 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow==2.3.0) (0.4.6)\nRequirement already satisfied: google-auth<3,>=1.6.3 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow==2.3.0) (1.35.0)\nRequirement already satisfied: tensorboard-plugin-wit>=1.6.0 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow==2.3.0) (1.8.1)\nRequirement already satisfied: markdown>=2.6.8 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow==2.3.0) (3.3.6)\nRequirement already satisfied: setuptools>=41.0.0 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow==2.3.0) (60.5.0)\nRequirement already satisfied: pyasn1-modules>=0.2.1 in /usr/local/lib/python3.7/dist-packages (from google-auth<3,>=1.6.3->tensorboard<3,>=2.3.0->tensorflow==2.3.0) (0.2.8)\nRequirement already satisfied: cachetools<5.0,>=2.0.0 in /usr/local/lib/python3.7/dist-packages (from google-auth<3,>=1.6.3->tensorboard<3,>=2.3.0->tensorflow==2.3.0) (4.2.4)\nRequirement already satisfied: rsa<5,>=3.1.4 in /usr/local/lib/python3.7/dist-packages (from google-auth<3,>=1.6.3->tensorboard<3,>=2.3.0->tensorflow==2.3.0) (4.8)\nRequirement already satisfied: requests-oauthlib>=0.7.0 in /usr/local/lib/python3.7/dist-packages (from google-auth-oauthlib<0.5,>=0.4.1->tensorboard<3,>=2.3.0->tensorflow==2.3.0) (1.3.0)\nRequirement already satisfied: importlib-metadata>=4.4 in /usr/local/lib/python3.7/dist-packages (from markdown>=2.6.8->tensorboard<3,>=2.3.0->tensorflow==2.3.0) (4.10.0)\nRequirement already satisfied: typing-extensions>=3.6.4 in /usr/local/lib/python3.7/dist-packages (from importlib-metadata>=4.4->markdown>=2.6.8->tensorboard<3,>=2.3.0->tensorflow==2.3.0) (3.10.0.2)\nRequirement already satisfied: zipp>=0.5 in /usr/local/lib/python3.7/dist-packages (from importlib-metadata>=4.4->markdown>=2.6.8->tensorboard<3,>=2.3.0->tensorflow==2.3.0) (3.7.0)\nRequirement already satisfied: pyasn1<0.5.0,>=0.4.6 in /usr/local/lib/python3.7/dist-packages (from pyasn1-modules>=0.2.1->google-auth<3,>=1.6.3->tensorboard<3,>=2.3.0->tensorflow==2.3.0) (0.4.8)\nRequirement already satisfied: idna<3,>=2.5 in /usr/local/lib/python3.7/dist-packages (from requests<3,>=2.21.0->tensorboard<3,>=2.3.0->tensorflow==2.3.0) (2.10)\nRequirement already satisfied: certifi>=2017.4.17 in /usr/local/lib/python3.7/dist-packages (from requests<3,>=2.21.0->tensorboard<3,>=2.3.0->tensorflow==2.3.0) (2021.10.8)\nRequirement already satisfied: chardet<4,>=3.0.2 in /usr/local/lib/python3.7/dist-packages (from requests<3,>=2.21.0->tensorboard<3,>=2.3.0->tensorflow==2.3.0) (3.0.4)\nRequirement already satisfied: urllib3!=1.25.0,!=1.25.1,<1.26,>=1.21.1 in /usr/local/lib/python3.7/dist-packages (from requests<3,>=2.21.0->tensorboard<3,>=2.3.0->tensorflow==2.3.0) (1.24.3)\nRequirement already satisfied: oauthlib>=3.0.0 in /usr/local/lib/python3.7/dist-packages (from requests-oauthlib>=0.7.0->google-auth-oauthlib<0.5,>=0.4.1->tensorboard<3,>=2.3.0->tensorflow==2.3.0) (3.1.1)\nRequirement already satisfied: gym in /usr/local/lib/python3.7/dist-packages (0.17.3)\nRequirement already satisfied: numpy>=1.10.4 in /usr/local/lib/python3.7/dist-packages (from gym) (1.18.5)\nRequirement already satisfied: pyglet<=1.5.0,>=1.4.0 in /usr/local/lib/python3.7/dist-packages (from gym) (1.5.0)\nRequirement already satisfied: cloudpickle<1.7.0,>=1.2.0 in /usr/local/lib/python3.7/dist-packages (from gym) (1.3.0)\nRequirement already satisfied: scipy in /usr/local/lib/python3.7/dist-packages (from gym) (1.4.1)\nRequirement already satisfied: future in /usr/local/lib/python3.7/dist-packages (from pyglet<=1.5.0,>=1.4.0->gym) (0.16.0)\nRequirement already satisfied: keras in /usr/local/lib/python3.7/dist-packages (2.7.0)\nRequirement already satisfied: keras-rl2 in /usr/local/lib/python3.7/dist-packages (1.0.5)\nRequirement already satisfied: tensorflow in /usr/local/lib/python3.7/dist-packages (from keras-rl2) (2.3.0)\nRequirement already satisfied: gast==0.3.3 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (0.3.3)\nRequirement already satisfied: wrapt>=1.11.1 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (1.13.3)\nRequirement already satisfied: grpcio>=1.8.6 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (1.43.0)\nRequirement already satisfied: scipy==1.4.1 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (1.4.1)\nRequirement already satisfied: termcolor>=1.1.0 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (1.1.0)\nRequirement already satisfied: astunparse==1.6.3 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (1.6.3)\nRequirement already satisfied: h5py<2.11.0,>=2.10.0 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (2.10.0)\nRequirement already satisfied: tensorflow-estimator<2.4.0,>=2.3.0 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (2.3.0)\nRequirement already satisfied: tensorboard<3,>=2.3.0 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (2.7.0)\nRequirement already satisfied: opt-einsum>=2.3.2 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (3.3.0)\nRequirement already satisfied: protobuf>=3.9.2 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (3.17.3)\nRequirement already satisfied: google-pasta>=0.1.8 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (0.2.0)\nRequirement already satisfied: six>=1.12.0 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (1.15.0)\nRequirement already satisfied: wheel>=0.26 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (0.37.1)\nRequirement already satisfied: absl-py>=0.7.0 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (0.12.0)\nRequirement already satisfied: numpy<1.19.0,>=1.16.0 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (1.18.5)\nRequirement already satisfied: keras-preprocessing<1.2,>=1.1.1 in /usr/local/lib/python3.7/dist-packages (from tensorflow->keras-rl2) (1.1.2)\nRequirement already satisfied: google-auth<3,>=1.6.3 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (1.35.0)\nRequirement already satisfied: setuptools>=41.0.0 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (60.5.0)\nRequirement already satisfied: tensorboard-data-server<0.7.0,>=0.6.0 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (0.6.1)\nRequirement already satisfied: requests<3,>=2.21.0 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (2.23.0)\nRequirement already satisfied: tensorboard-plugin-wit>=1.6.0 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (1.8.1)\nRequirement already satisfied: markdown>=2.6.8 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (3.3.6)\nRequirement already satisfied: google-auth-oauthlib<0.5,>=0.4.1 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (0.4.6)\nRequirement already satisfied: werkzeug>=0.11.15 in /usr/local/lib/python3.7/dist-packages (from tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (1.0.1)\nRequirement already satisfied: pyasn1-modules>=0.2.1 in /usr/local/lib/python3.7/dist-packages (from google-auth<3,>=1.6.3->tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (0.2.8)\nRequirement already satisfied: rsa<5,>=3.1.4 in /usr/local/lib/python3.7/dist-packages (from google-auth<3,>=1.6.3->tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (4.8)\nRequirement already satisfied: cachetools<5.0,>=2.0.0 in /usr/local/lib/python3.7/dist-packages (from google-auth<3,>=1.6.3->tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (4.2.4)\nRequirement already satisfied: requests-oauthlib>=0.7.0 in /usr/local/lib/python3.7/dist-packages (from google-auth-oauthlib<0.5,>=0.4.1->tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (1.3.0)\nRequirement already satisfied: importlib-metadata>=4.4 in /usr/local/lib/python3.7/dist-packages (from markdown>=2.6.8->tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (4.10.0)\nRequirement already satisfied: typing-extensions>=3.6.4 in /usr/local/lib/python3.7/dist-packages (from importlib-metadata>=4.4->markdown>=2.6.8->tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (3.10.0.2)\nRequirement already satisfied: zipp>=0.5 in /usr/local/lib/python3.7/dist-packages (from importlib-metadata>=4.4->markdown>=2.6.8->tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (3.7.0)\nRequirement already satisfied: pyasn1<0.5.0,>=0.4.6 in /usr/local/lib/python3.7/dist-packages (from pyasn1-modules>=0.2.1->google-auth<3,>=1.6.3->tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (0.4.8)\nRequirement already satisfied: certifi>=2017.4.17 in /usr/local/lib/python3.7/dist-packages (from requests<3,>=2.21.0->tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (2021.10.8)\nRequirement already satisfied: urllib3!=1.25.0,!=1.25.1,<1.26,>=1.21.1 in /usr/local/lib/python3.7/dist-packages (from requests<3,>=2.21.0->tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (1.24.3)\nRequirement already satisfied: idna<3,>=2.5 in /usr/local/lib/python3.7/dist-packages (from requests<3,>=2.21.0->tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (2.10)\nRequirement already satisfied: chardet<4,>=3.0.2 in /usr/local/lib/python3.7/dist-packages (from requests<3,>=2.21.0->tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (3.0.4)\nRequirement already satisfied: oauthlib>=3.0.0 in /usr/local/lib/python3.7/dist-packages (from requests-oauthlib>=0.7.0->google-auth-oauthlib<0.5,>=0.4.1->tensorboard<3,>=2.3.0->tensorflow->keras-rl2) (3.1.1)\nRequirement already satisfied: unrar in /usr/local/lib/python3.7/dist-packages (0.4)\n\nUNRAR 5.50 freeware Copyright (c) 1993-2017 Alexander Roshal\n\n\nExtracting from Roms.rar\n\nExtracting HC ROMS.zip \b\b\b\b 36%\b\b\b\b\b OK \nExtracting ROMS.zip \b\b\b\b 74%\b\b\b\b 99%\b\b\b\b\b OK \nAll OK\nmkdir: cannot create directory ‘rars’: File exists\ncopying adventure.bin from HC ROMS/BY ALPHABET (PAL)/A-G/Adventure (PAL).bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/adventure.bin\ncopying air_raid.bin from HC ROMS/BY ALPHABET (PAL)/A-G/Air Raid (PAL).bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/air_raid.bin\ncopying alien.bin from HC ROMS/BY ALPHABET (PAL)/A-G/REMAINING NTSC ORIGINALS/Alien.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/alien.bin\ncopying crazy_climber.bin from HC ROMS/BY ALPHABET (PAL)/A-G/REMAINING NTSC ORIGINALS/Crazy Climber.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/crazy_climber.bin\ncopying elevator_action.bin from HC ROMS/BY ALPHABET (PAL)/A-G/REMAINING NTSC ORIGINALS/Elevator Action (Prototype).bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/elevator_action.bin\ncopying gravitar.bin from HC ROMS/BY ALPHABET (PAL)/A-G/REMAINING NTSC ORIGINALS/Gravitar.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/gravitar.bin\ncopying keystone_kapers.bin from HC ROMS/BY ALPHABET (PAL)/H-R/Keystone Kapers (PAL).bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/keystone_kapers.bin\ncopying king_kong.bin from HC ROMS/BY ALPHABET (PAL)/H-R/King Kong (PAL).bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/king_kong.bin\ncopying laser_gates.bin from HC ROMS/BY ALPHABET (PAL)/H-R/Laser Gates (PAL).bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/laser_gates.bin\ncopying mr_do.bin from HC ROMS/BY ALPHABET (PAL)/H-R/Mr. Do! (PAL).bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/mr_do.bin\ncopying pacman.bin from HC ROMS/BY ALPHABET (PAL)/H-R/Pac-Man (PAL).bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/pacman.bin\ncopying jamesbond.bin from HC ROMS/BY ALPHABET (PAL)/H-R/REMAINING NTSC ORIGINALS/James Bond 007.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/jamesbond.bin\ncopying koolaid.bin from HC ROMS/BY ALPHABET (PAL)/H-R/REMAINING NTSC ORIGINALS/Kool-Aid Man.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/koolaid.bin\ncopying krull.bin from HC ROMS/BY ALPHABET (PAL)/H-R/REMAINING NTSC ORIGINALS/Krull.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/krull.bin\ncopying montezuma_revenge.bin from HC ROMS/BY ALPHABET (PAL)/H-R/REMAINING NTSC ORIGINALS/Montezuma's Revenge - Featuring Panama Joe.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/montezuma_revenge.bin\ncopying star_gunner.bin from HC ROMS/BY ALPHABET (PAL)/S-Z/REMAINING NTSC ORIGINALS/Stargunner.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/star_gunner.bin\ncopying time_pilot.bin from HC ROMS/BY ALPHABET (PAL)/S-Z/REMAINING NTSC ORIGINALS/Time Pilot.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/time_pilot.bin\ncopying up_n_down.bin from HC ROMS/BY ALPHABET (PAL)/S-Z/REMAINING NTSC ORIGINALS/Up 'n Down.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/up_n_down.bin\ncopying sir_lancelot.bin from HC ROMS/BY ALPHABET (PAL)/S-Z/Sir Lancelot (PAL).bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/sir_lancelot.bin\ncopying amidar.bin from HC ROMS/BY ALPHABET/A-G/Amidar.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/amidar.bin\ncopying asteroids.bin from HC ROMS/BY ALPHABET/A-G/Asteroids [no copyright].bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/asteroids.bin\ncopying atlantis.bin from HC ROMS/BY ALPHABET/A-G/Atlantis.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/atlantis.bin\ncopying bank_heist.bin from HC ROMS/BY ALPHABET/A-G/Bank Heist.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/bank_heist.bin\ncopying battle_zone.bin from HC ROMS/BY ALPHABET/A-G/Battlezone.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/battle_zone.bin\ncopying beam_rider.bin from HC ROMS/BY ALPHABET/A-G/Beamrider.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/beam_rider.bin\ncopying berzerk.bin from HC ROMS/BY ALPHABET/A-G/Berzerk.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/berzerk.bin\ncopying bowling.bin from HC ROMS/BY ALPHABET/A-G/Bowling.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/bowling.bin\ncopying boxing.bin from HC ROMS/BY ALPHABET/A-G/Boxing.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/boxing.bin\ncopying breakout.bin from HC ROMS/BY ALPHABET/A-G/Breakout - Breakaway IV.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/breakout.bin\ncopying carnival.bin from HC ROMS/BY ALPHABET/A-G/Carnival.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/carnival.bin\ncopying centipede.bin from HC ROMS/BY ALPHABET/A-G/Centipede.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/centipede.bin\ncopying chopper_command.bin from HC ROMS/BY ALPHABET/A-G/Chopper Command.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/chopper_command.bin\ncopying defender.bin from HC ROMS/BY ALPHABET/A-G/Defender.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/defender.bin\ncopying demon_attack.bin from HC ROMS/BY ALPHABET/A-G/Demon Attack.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/demon_attack.bin\ncopying donkey_kong.bin from HC ROMS/BY ALPHABET/A-G/Donkey Kong.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/donkey_kong.bin\ncopying double_dunk.bin from HC ROMS/BY ALPHABET/A-G/Double Dunk.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/double_dunk.bin\ncopying enduro.bin from HC ROMS/BY ALPHABET/A-G/Enduro.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/enduro.bin\ncopying fishing_derby.bin from HC ROMS/BY ALPHABET/A-G/Fishing Derby.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/fishing_derby.bin\ncopying freeway.bin from HC ROMS/BY ALPHABET/A-G/Freeway.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/freeway.bin\ncopying frogger.bin from HC ROMS/BY ALPHABET/A-G/Frogger.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/frogger.bin\ncopying frostbite.bin from HC ROMS/BY ALPHABET/A-G/Frostbite.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/frostbite.bin\ncopying galaxian.bin from HC ROMS/BY ALPHABET/A-G/Galaxian.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/galaxian.bin\ncopying gopher.bin from HC ROMS/BY ALPHABET/A-G/Gopher.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/gopher.bin\ncopying hero.bin from HC ROMS/BY ALPHABET/H-R/H.E.R.O..bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/hero.bin\ncopying ice_hockey.bin from HC ROMS/BY ALPHABET/H-R/Ice Hockey.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/ice_hockey.bin\ncopying journey_escape.bin from HC ROMS/BY ALPHABET/H-R/Journey Escape.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/journey_escape.bin\ncopying kaboom.bin from HC ROMS/BY ALPHABET/H-R/Kaboom!.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/kaboom.bin\ncopying kangaroo.bin from HC ROMS/BY ALPHABET/H-R/Kangaroo.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/kangaroo.bin\ncopying kung_fu_master.bin from HC ROMS/BY ALPHABET/H-R/Kung-Fu Master.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/kung_fu_master.bin\ncopying lost_luggage.bin from HC ROMS/BY ALPHABET/H-R/Lost Luggage [no opening scene].bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/lost_luggage.bin\ncopying ms_pacman.bin from HC ROMS/BY ALPHABET/H-R/Ms. Pac-Man.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/ms_pacman.bin\ncopying name_this_game.bin from HC ROMS/BY ALPHABET/H-R/Name This Game.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/name_this_game.bin\ncopying phoenix.bin from HC ROMS/BY ALPHABET/H-R/Phoenix.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/phoenix.bin\ncopying pitfall.bin from HC ROMS/BY ALPHABET/H-R/Pitfall! - Pitfall Harry's Jungle Adventure.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/pitfall.bin\ncopying pooyan.bin from HC ROMS/BY ALPHABET/H-R/Pooyan.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/pooyan.bin\ncopying private_eye.bin from HC ROMS/BY ALPHABET/H-R/Private Eye.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/private_eye.bin\ncopying qbert.bin from HC ROMS/BY ALPHABET/H-R/Q-bert.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/qbert.bin\ncopying riverraid.bin from HC ROMS/BY ALPHABET/H-R/River Raid.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/riverraid.bin\ncopying road_runner.bin from patched version of HC ROMS/BY ALPHABET/H-R/Road Runner.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/road_runner.bin\ncopying robotank.bin from HC ROMS/BY ALPHABET/H-R/Robot Tank.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/robotank.bin\ncopying seaquest.bin from HC ROMS/BY ALPHABET/S-Z/Seaquest.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/seaquest.bin\ncopying skiing.bin from HC ROMS/BY ALPHABET/S-Z/Skiing.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/skiing.bin\ncopying solaris.bin from HC ROMS/BY ALPHABET/S-Z/Solaris.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/solaris.bin\ncopying space_invaders.bin from HC ROMS/BY ALPHABET/S-Z/Space Invaders.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/space_invaders.bin\ncopying surround.bin from HC ROMS/BY ALPHABET/S-Z/Surround - Chase.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/surround.bin\ncopying tennis.bin from HC ROMS/BY ALPHABET/S-Z/Tennis.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/tennis.bin\ncopying trondead.bin from HC ROMS/BY ALPHABET/S-Z/TRON - Deadly Discs.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/trondead.bin\ncopying tutankham.bin from HC ROMS/BY ALPHABET/S-Z/Tutankham.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/tutankham.bin\ncopying venture.bin from HC ROMS/BY ALPHABET/S-Z/Venture.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/venture.bin\ncopying pong.bin from HC ROMS/BY ALPHABET/S-Z/Video Olympics - Pong Sports.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/pong.bin\ncopying video_pinball.bin from HC ROMS/BY ALPHABET/S-Z/Video Pinball - Arcade Pinball.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/video_pinball.bin\ncopying wizard_of_wor.bin from HC ROMS/BY ALPHABET/S-Z/Wizard of Wor.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/wizard_of_wor.bin\ncopying yars_revenge.bin from HC ROMS/BY ALPHABET/S-Z/Yars' Revenge.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/yars_revenge.bin\ncopying zaxxon.bin from HC ROMS/BY ALPHABET/S-Z/Zaxxon.bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/zaxxon.bin\ncopying assault.bin from HC ROMS/NTSC VERSIONS OF PAL ORIGINALS/Assault (AKA Sky Alien) (1983) (Bomb - Onbase) (CA281).bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/assault.bin\ncopying asterix.bin from ROMS/Asterix (AKA Taz) (07-27-1983) (Atari, Jerome Domurat, Steve Woita) (CX2696) (Prototype).bin to /usr/local/lib/python3.7/dist-packages/atari_py/atari_roms/asterix.bin\n"
],
[
"!pip install gym pyvirtualdisplay > /dev/null 2>&1\n!apt-get install -y xvfb python-opengl ffmpeg > /dev/null 2>&1\n!apt-get update > /dev/null 2>&1\n!apt-get install cmake > /dev/null 2>&1\n!pip install --upgrade setuptools 2>&1\n!pip install ez_setup > /dev/null 2>&1\n!pip install gym[atari] > /dev/null 2>&1",
"Requirement already satisfied: setuptools in /usr/local/lib/python3.7/dist-packages (60.5.0)\n"
]
],
[
[
"#Helper Functions",
"_____no_output_____"
]
],
[
[
"import gym\n\ndef query_environment(name):\n env = gym.make(name)\n spec = gym.spec(name)\n print(f\"Action Space: {env.action_space}\")\n print(f\"Observation Space: {env.observation_space}\")\n print(f\"Max Episode Steps: {spec.max_episode_steps}\")\n print(f\"Nondeterministic: {spec.nondeterministic}\")\n print(f\"Reward Range: {env.reward_range}\")\n print(f\"Reward Threshold: {spec.reward_threshold}\")",
"_____no_output_____"
],
[
"import gym\nfrom gym.wrappers import Monitor\nimport glob\nimport io\nimport base64\nfrom IPython.display import HTML\nfrom pyvirtualdisplay import Display\nfrom IPython import display as ipythondisplay\n\ndisplay = Display(visible=0, size=(1400, 900))\ndisplay.start()\n\n\"\"\"\nUtility functions to enable video recording of gym environment \nand displaying it.\nTo enable video, just do \"env = wrap_env(env)\"\"\n\"\"\"\n\ndef show_video():\n mp4list = glob.glob('video/*.mp4')\n if len(mp4list) > 0:\n mp4 = mp4list[0]\n video = io.open(mp4, 'r+b').read()\n encoded = base64.b64encode(video)\n ipythondisplay.display(HTML(data='''<video alt=\"test\" autoplay \n loop controls style=\"height: 400px;\">\n <source src=\"data:video/mp4;base64,{0}\" type=\"video/mp4\" />\n </video>'''.format(encoded.decode('ascii'))))\n else: \n print(\"Could not find video\")\n \n\ndef wrap_env(env):\n env = Monitor(env, './video', force=True)\n return env",
"_____no_output_____"
]
],
[
[
"## Check Env OpenAI Gym",
"_____no_output_____"
]
],
[
[
"query_environment('CartPole-v0')",
"Action Space: Discrete(2)\nObservation Space: Box(-3.4028234663852886e+38, 3.4028234663852886e+38, (4,), float32)\nMax Episode Steps: 200\nNondeterministic: False\nReward Range: (-inf, inf)\nReward Threshold: 195.0\n"
],
[
"env = wrap_env(gym.make('CartPole-v0'))\n\nobservation = env.reset()\n\nwhile True:\n \n env.render()\n \n #your agent goes here\n action = env.action_space.sample() \n \n observation, reward, done, info = env.step(action) \n \n \n if done: \n break;\n \nenv.close()\nshow_video()",
"_____no_output_____"
],
[
"import random\nenv = wrap_env(gym.make(\"CartPole-v0\"))\nepisodes = 10\nfor episode in range(1, episodes+1):\n state = env.reset()\n done = False\n score = 0 \n \n while not done:\n env.render()\n action = random.choice([0,1])\n n_state, reward, done, info = env.step(action)\n score+=reward\n print('Episode:{} Score:{}'.format(episode, score))",
"Episode:1 Score:19.0\nEpisode:2 Score:25.0\nEpisode:3 Score:15.0\nEpisode:4 Score:30.0\nEpisode:5 Score:29.0\nEpisode:6 Score:30.0\nEpisode:7 Score:26.0\nEpisode:8 Score:17.0\nEpisode:9 Score:15.0\nEpisode:10 Score:10.0\n"
]
],
[
[
"#Create a Deep Learning Model",
"_____no_output_____"
]
],
[
[
"import numpy as np\nfrom tensorflow.keras.models import Sequential\nfrom tensorflow.keras.layers import Dense, Flatten\nfrom tensorflow.keras.optimizers import Adam",
"_____no_output_____"
],
[
"def build_model(states, actions):\n model = Sequential()\n model.add(Flatten(input_shape=(1,states)))\n model.add(Dense(24, activation='relu'))\n model.add(Dense(24, activation='relu'))\n model.add(Dense(actions, activation='linear'))\n return model",
"_____no_output_____"
],
[
"env = gym.make('CartPole-v0')\nstates = env.observation_space.shape[0]\nactions = env.action_space.n\nmodel = build_model(states, actions)\nmodel.summary()",
"Model: \"sequential\"\n_________________________________________________________________\nLayer (type) Output Shape Param # \n=================================================================\nflatten (Flatten) (None, 4) 0 \n_________________________________________________________________\ndense (Dense) (None, 24) 120 \n_________________________________________________________________\ndense_1 (Dense) (None, 24) 600 \n_________________________________________________________________\ndense_2 (Dense) (None, 2) 50 \n=================================================================\nTotal params: 770\nTrainable params: 770\nNon-trainable params: 0\n_________________________________________________________________\n"
]
],
[
[
"# Build Agent",
"_____no_output_____"
]
],
[
[
"from rl.agents import DQNAgent\nfrom rl.policy import BoltzmannQPolicy\nfrom rl.memory import SequentialMemory",
"_____no_output_____"
],
[
"def build_agent(model, actions):\n policy = BoltzmannQPolicy()\n memory = SequentialMemory(limit=50000, window_length=1)\n dqn = DQNAgent(model=model, memory=memory, policy=policy, \n nb_actions=actions, nb_steps_warmup=10, target_model_update=1e-2)\n return dqn",
"_____no_output_____"
],
[
"env = gym.make('CartPole-v0')\nactions = env.action_space.n\nstates = env.observation_space.shape[0]\nmodel = build_model(states, actions)\ndqn = build_agent(model, actions)\ndqn.compile(Adam(lr=1e-3), metrics=['mae'])\ndqn.fit(env, nb_steps=50000, visualize=False, verbose=1)",
"Training for 50000 steps ...\nInterval 1 (0 steps performed)\n\r 1/10000 [..............................] - ETA: 21:32 - reward: 1.0000"
],
[
"scores = dqn.test(env, nb_episodes=100, visualize=False)\nprint(np.mean(scores.history['episode_reward']))",
"Testing for 100 episodes ...\nEpisode 1: reward: 200.000, steps: 200\nEpisode 2: reward: 200.000, steps: 200\nEpisode 3: reward: 200.000, steps: 200\nEpisode 4: reward: 200.000, steps: 200\nEpisode 5: reward: 200.000, steps: 200\nEpisode 6: reward: 200.000, steps: 200\nEpisode 7: reward: 200.000, steps: 200\nEpisode 8: reward: 200.000, steps: 200\nEpisode 9: reward: 200.000, steps: 200\nEpisode 10: reward: 200.000, steps: 200\nEpisode 11: reward: 200.000, steps: 200\nEpisode 12: reward: 200.000, steps: 200\nEpisode 13: reward: 200.000, steps: 200\nEpisode 14: reward: 200.000, steps: 200\nEpisode 15: reward: 200.000, steps: 200\nEpisode 16: reward: 200.000, steps: 200\nEpisode 17: reward: 200.000, steps: 200\nEpisode 18: reward: 200.000, steps: 200\nEpisode 19: reward: 200.000, steps: 200\nEpisode 20: reward: 200.000, steps: 200\nEpisode 21: reward: 200.000, steps: 200\nEpisode 22: reward: 200.000, steps: 200\nEpisode 23: reward: 200.000, steps: 200\nEpisode 24: reward: 200.000, steps: 200\nEpisode 25: reward: 200.000, steps: 200\nEpisode 26: reward: 200.000, steps: 200\nEpisode 27: reward: 200.000, steps: 200\nEpisode 28: reward: 200.000, steps: 200\nEpisode 29: reward: 200.000, steps: 200\nEpisode 30: reward: 200.000, steps: 200\nEpisode 31: reward: 200.000, steps: 200\nEpisode 32: reward: 200.000, steps: 200\nEpisode 33: reward: 200.000, steps: 200\nEpisode 34: reward: 200.000, steps: 200\nEpisode 35: reward: 200.000, steps: 200\nEpisode 36: reward: 200.000, steps: 200\nEpisode 37: reward: 200.000, steps: 200\nEpisode 38: reward: 200.000, steps: 200\nEpisode 39: reward: 200.000, steps: 200\nEpisode 40: reward: 200.000, steps: 200\nEpisode 41: reward: 200.000, steps: 200\nEpisode 42: reward: 200.000, steps: 200\nEpisode 43: reward: 200.000, steps: 200\nEpisode 44: reward: 200.000, steps: 200\nEpisode 45: reward: 200.000, steps: 200\nEpisode 46: reward: 200.000, steps: 200\nEpisode 47: reward: 200.000, steps: 200\nEpisode 48: reward: 200.000, steps: 200\nEpisode 49: reward: 200.000, steps: 200\nEpisode 50: reward: 200.000, steps: 200\nEpisode 51: reward: 200.000, steps: 200\nEpisode 52: reward: 200.000, steps: 200\nEpisode 53: reward: 200.000, steps: 200\nEpisode 54: reward: 200.000, steps: 200\nEpisode 55: reward: 200.000, steps: 200\nEpisode 56: reward: 200.000, steps: 200\nEpisode 57: reward: 200.000, steps: 200\nEpisode 58: reward: 200.000, steps: 200\nEpisode 59: reward: 200.000, steps: 200\nEpisode 60: reward: 200.000, steps: 200\nEpisode 61: reward: 200.000, steps: 200\nEpisode 62: reward: 200.000, steps: 200\nEpisode 63: reward: 200.000, steps: 200\nEpisode 64: reward: 200.000, steps: 200\nEpisode 65: reward: 200.000, steps: 200\nEpisode 66: reward: 200.000, steps: 200\nEpisode 67: reward: 200.000, steps: 200\nEpisode 68: reward: 200.000, steps: 200\nEpisode 69: reward: 200.000, steps: 200\nEpisode 70: reward: 200.000, steps: 200\nEpisode 71: reward: 200.000, steps: 200\nEpisode 72: reward: 200.000, steps: 200\nEpisode 73: reward: 200.000, steps: 200\nEpisode 74: reward: 200.000, steps: 200\nEpisode 75: reward: 200.000, steps: 200\nEpisode 76: reward: 200.000, steps: 200\nEpisode 77: reward: 200.000, steps: 200\nEpisode 78: reward: 200.000, steps: 200\nEpisode 79: reward: 200.000, steps: 200\nEpisode 80: reward: 200.000, steps: 200\nEpisode 81: reward: 200.000, steps: 200\nEpisode 82: reward: 200.000, steps: 200\nEpisode 83: reward: 200.000, steps: 200\nEpisode 84: reward: 200.000, steps: 200\nEpisode 85: reward: 200.000, steps: 200\nEpisode 86: reward: 200.000, steps: 200\nEpisode 87: reward: 200.000, steps: 200\nEpisode 88: reward: 200.000, steps: 200\nEpisode 89: reward: 200.000, steps: 200\nEpisode 90: reward: 200.000, steps: 200\nEpisode 91: reward: 200.000, steps: 200\nEpisode 92: reward: 200.000, steps: 200\nEpisode 93: reward: 200.000, steps: 200\nEpisode 94: reward: 200.000, steps: 200\nEpisode 95: reward: 200.000, steps: 200\nEpisode 96: reward: 200.000, steps: 200\nEpisode 97: reward: 200.000, steps: 200\nEpisode 98: reward: 200.000, steps: 200\nEpisode 99: reward: 200.000, steps: 200\nEpisode 100: reward: 200.000, steps: 200\n200.0\n"
],
[
"_ = dqn.test(env, nb_episodes=15, visualize=True)",
"Testing for 15 episodes ...\nEpisode 1: reward: 200.000, steps: 200\nEpisode 2: reward: 200.000, steps: 200\nEpisode 3: reward: 200.000, steps: 200\nEpisode 4: reward: 200.000, steps: 200\nEpisode 5: reward: 200.000, steps: 200\nEpisode 6: reward: 200.000, steps: 200\nEpisode 7: reward: 200.000, steps: 200\nEpisode 8: reward: 200.000, steps: 200\nEpisode 9: reward: 200.000, steps: 200\nEpisode 10: reward: 200.000, steps: 200\nEpisode 11: reward: 200.000, steps: 200\nEpisode 12: reward: 200.000, steps: 200\nEpisode 13: reward: 200.000, steps: 200\nEpisode 14: reward: 200.000, steps: 200\nEpisode 15: reward: 200.000, steps: 200\n"
],
[
"dqn.save_weights('dqn_weights.h5f', overwrite=True)",
"_____no_output_____"
]
],
[
[
"#Reloading Agent",
"_____no_output_____"
]
],
[
[
"env = wrap_env(gym.make('CartPole-v0'))\nactions = env.action_space.n\nstates = env.observation_space.shape[0]\nmodel = build_model(states, actions)\ndqn = build_agent(model, actions)\ndqn.compile(Adam(lr=1e-3), metrics=['mae'])",
"_____no_output_____"
],
[
"dqn.load_weights('dqn_weights.h5f')",
"_____no_output_____"
],
[
"_ = dqn.test(env, nb_episodes=5, visualize=True)",
"Testing for 5 episodes ...\nEpisode 1: reward: 200.000, steps: 200\nEpisode 2: reward: 200.000, steps: 200\nEpisode 3: reward: 200.000, steps: 200\nEpisode 4: reward: 200.000, steps: 200\nEpisode 5: reward: 200.000, steps: 200\n"
],
[
"show_video()",
"_____no_output_____"
],
[
"",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
]
] |
4acdbaad28bf4f657518d906b85c636da18253ad
| 24,304 |
ipynb
|
Jupyter Notebook
|
src/week_3/Assignment 2 cooperation in space.ipynb
|
lyoussifou/EPA1324_open
|
1b9fed7555406c6d2f17483e1ddd174f249a965c
|
[
"BSD-3-Clause"
] | 6 |
2020-11-18T08:08:25.000Z
|
2021-12-06T18:38:17.000Z
|
src/week_3/Assignment 2 cooperation in space.ipynb
|
lyoussifou/EPA1324_open
|
1b9fed7555406c6d2f17483e1ddd174f249a965c
|
[
"BSD-3-Clause"
] | null | null | null |
src/week_3/Assignment 2 cooperation in space.ipynb
|
lyoussifou/EPA1324_open
|
1b9fed7555406c6d2f17483e1ddd174f249a965c
|
[
"BSD-3-Clause"
] | 29 |
2020-11-12T10:14:57.000Z
|
2022-01-20T12:31:03.000Z
| 52.606061 | 1,267 | 0.604798 |
[
[
[
"\n# The importance of space\nAgent based models are useful when the aggregate system behavior emerges out of local interactions amongst the agents. In the model of the evolution of cooperation, we created a set of agents and let all agents play against all other agents. Basically, we pretended as if all our agents were perfectly mixed. In practice, however, it is much more common that agents only interact with some, but not all, other agents. For example, in models of epidemiology, social interactions are a key factors. Thus, interactions are dependend on your social network. In other situations, our behavior might be based on what we see around us. Phenomena like fashion are at least partly driven by seeing what others are doing and mimicking this behavior. The same is true for many animals. Flocking dynamics as exhibited by starling, or shoaling behavior in fish, can be explained by the animal looking at its neirest neighbors and staying within a given distance of them. In agent based models, anything that structures the interaction amongst agents is typically called a space. This space can be a 2d or 3d space with euclidian distances (as in models of flocking and shoaling), it can also be a grid structure (as we will show below), or it can be a network structure. \n\nMESA comes with several spaces that we can readily use. These are\n\n* **SingleGrid;** an 'excel-like' space with each agent occopying a single grid cell\n* **MultiGrid;** like grid, but with more than one agent per grid cell\n* **HexGrid;** like grid, but on a hexagonal grid (*e.g.*, the board game Catan) thus changing who your neighbours are\n* **ConinuousSpace;** a 2d continous space were agents can occupy any coordinate\n* **NetworkGrid;** a network structure were one or more agents occupy a given node.\n\nA key concern when using a none-networked space, is to think carefull about what happens at the edges of the space. In a basic implementation, agents in for example the top left corner has only 2 neighbors, while an agent in the middle has four neighbors. This can give rise to artifacts in the results. Basically, the dynamics at the edges are different from the behavior further away from the edges. It is therefore quite common to use a torus, or donut, shape for the space. In this way, there is no longer any edge and artifacts are thus removed.\n\n\n# The emergence of cooperation in space\nThe documentation of MESA on the different spaces is quite limited. Therefore, this assignment is largely a tutorial continuing on the evolution of cooperation. \n\nWe make the following changes to the model\n\n* Each agent gets a position, which is an x,y coordinate indicating the grid cell the agent occupies.\n* The model has a grid, with an agent of random class. We initialize the model with equal probabilities for each type of class.\n* All agents play against their neighbors. On a grid, neighborhood can be defined in various ways. For example, a Von Neumann neighborhood contains the four cells that share a border with the central cell. A Moore neighborhood with distance one contains 8 cells by also considering the diagonal. Below, we use a neighborhood distance of 1, and we do include diagonal neighbors. So we set Moore to True. Feel free to experiment with this model by setting it to False, \n* The evolutionary dynamic, after all agents having played, is that each agent compares its scores to its neighbors. It will adopt whichever strategy within its neighborhood performed best.\n* Next to using a SingleGrid from MESA, we also use a DataCollector to handle collecting statistics.\n\nBelow, I discuss in more detail the code containing the most important modifications.\n",
"_____no_output_____"
]
],
[
[
"from collections import deque, Counter, defaultdict\nfrom enum import Enum\nfrom itertools import combinations\nfrom math import floor\n\nimport numpy as np\nimport matplotlib.pyplot as plt\nimport pandas as pd\n\nfrom mesa import Model, Agent\nfrom mesa.space import SingleGrid\nfrom mesa.datacollection import DataCollector\n\n\nclass Move(Enum):\n COOPERATE = 1\n DEFECT = 2\n\n\nclass AxelrodAgent(Agent):\n \"\"\"An Abstract class from which all strategies have to be derived\n \n \n Attributes\n ----------\n points : int\n pos : tuple\n \n \"\"\"\n def __init__(self, unique_id, pos, model):\n super().__init__(unique_id, model)\n self.points = 0\n self.pos = pos\n\n def step(self):\n '''\n This function defines the move and any logic for deciding\n on the move goes here.\n \n Returns\n -------\n Move.COOPERATE or Move.DEFECT\n \n '''\n raise NotImplemetedError\n\n def receive_payoff(self, payoff, my_move, opponent_move):\n '''receive payoff and the two moves resulting in this payoff\n \n Parameters\n ----------\n payoff : int\n my_move : {Move.COOPERATE, Move.DEFECT}\n opponements_move : {Move.COOPERATE, Move.DEFECT}\n \n '''\n self.points += payoff\n \n def reset(self):\n '''\n This function is called after playing N iterations against\n another player.\n '''\n raise NotImplementedError\n\n\nclass TitForTat(AxelrodAgent):\n \"\"\"This class defines the following strategy: play nice, unless, \n in the previous move, the other player betrayed you.\"\"\"\n \n def __init__(self, unique_id, pos, model):\n super().__init__(unique_id, pos, model)\n self.opponent_last_move = Move.COOPERATE\n \n def step(self):\n return self.opponent_last_move\n \n def receive_payoff(self, payoff, my_move, opponent_move):\n super().receive_payoff(payoff, my_move, opponent_move)\n self.opponent_last_move = opponent_move\n \n def reset(self):\n self.opponent_last_move = Move.COOPERATE\n\n\nclass ContriteTitForTat(AxelrodAgent):\n \"\"\"This class defines the following strategy: play nice, unless, \n in the previous two moves, the other player betrayed you.\"\"\" \n \n def __init__(self, unique_id, pos, model):\n super().__init__(unique_id, pos, model)\n self.opponent_last_two_moves = deque([Move.COOPERATE, Move.COOPERATE], maxlen=2)\n \n def step(self):\n if (self.opponent_last_two_moves[0] == Move.DEFECT) and\\\n (self.opponent_last_two_moves[1] == Move.DEFECT):\n return Move.DEFECT\n else:\n return Move.COOPERATE\n \n def receive_payoff(self, payoff, my_move, opponent_move):\n super().receive_payoff(payoff, my_move, opponent_move)\n self.opponent_last_two_moves.append(opponent_move)\n \n def reset(self):\n self.opponent_last_two_moves = deque([Move.COOPERATE, Move.COOPERATE], maxlen=2) \n\n\nclass NoisySpatialEvolutionaryAxelrodModel(Model):\n \n def __init__(self, N, noise_level=0.01, seed=None,\n height=20, width=20,):\n super().__init__(seed=seed)\n self.noise_level = noise_level\n self.num_iterations = N\n self.agents = set()\n\n self.payoff_matrix = {}\n self.payoff_matrix[(Move.COOPERATE, Move.COOPERATE)] = (2, 2)\n self.payoff_matrix[(Move.COOPERATE, Move.DEFECT)] = (0, 3)\n self.payoff_matrix[(Move.DEFECT, Move.COOPERATE)] = (3, 0)\n self.payoff_matrix[(Move.DEFECT, Move.DEFECT)] = (1, 1) \n \n self.grid = SingleGrid(width, height, torus=True)\n \n strategies = AxelrodAgent.__subclasses__()\n num_strategies = len(strategies)\n self.agent_id = 0\n \n for cell in self.grid.coord_iter():\n _, x, y = cell\n pos = (x, y)\n \n self.agent_id += 1\n\n strategy_index = int(floor(self.random.random()*num_strategies))\n \n agent = strategies[strategy_index](self.agent_id, pos, self)\n self.grid.position_agent(agent, (x, y))\n self.agents.add(agent)\n \n self.datacollector = DataCollector(model_reporters={klass.__name__:klass.__name__\n for klass in strategies})\n\n def count_agent_types(self):\n counter = Counter()\n for agent in self.agents:\n counter[agent.__class__.__name__] += 1\n \n for k,v in counter.items():\n setattr(self, k, v)\n\n def step(self):\n '''Advance the model by one step.'''\n self.count_agent_types()\n self.datacollector.collect(self)\n\n for (agent_a, x, y) in self.grid.coord_iter():\n for agent_b in self.grid.neighbor_iter((x,y), moore=True):\n for _ in range(self.num_iterations):\n move_a = agent_a.step()\n move_b = agent_b.step()\n\n #insert noise in movement\n if self.random.random() < self.noise_level:\n if move_a == Move.COOPERATE:\n move_a = Move.DEFECT\n else:\n move_a = Move.COOPERATE\n if self.random.random() < self.noise_level:\n if move_b == Move.COOPERATE:\n move_b = Move.DEFECT\n else:\n move_b = Move.COOPERATE\n\n payoff_a, payoff_b = self.payoff_matrix[(move_a, move_b)]\n\n agent_a.receive_payoff(payoff_a, move_a, move_b)\n agent_b.receive_payoff(payoff_b, move_b, move_a)\n agent_a.reset()\n agent_b.reset()\n\n # evolution\n # tricky, we need to determine for each grid cell\n # is a change needed, if so, log position, agent, and type to change to\n agents_to_change = []\n for agent_a in self.agents:\n neighborhood = self.grid.iter_neighbors(agent_a.pos, moore=True,\n include_center=True)\n neighborhood = ([n for n in neighborhood])\n neighborhood.sort(key=lambda x:x.points, reverse=True)\n best_strategy = neighborhood[0].__class__\n # if best type of strategy in neighborhood is\n # different from strategy type of agent, we need\n # to change our strategy\n if not isinstance(agent_a, best_strategy):\n agents_to_change.append((agent_a, best_strategy))\n\n for entry in agents_to_change:\n agent, klass = entry\n self.agents.remove(agent)\n self.grid.remove_agent(agent)\n \n pos = agent.pos\n self.agent_id += 1\n \n new_agent = klass(self.agent_id, pos, self)\n self.grid.position_agent(new_agent, pos)\n self.agents.add(new_agent)\n",
"_____no_output_____"
]
],
[
[
"\nIn the `__init__`, we now instantiate a SingleGrid, with a specified width and height. We set the kwarg torus to True indicating we are using a donut shape grid to avoid edge effects. Next, we fill this grid with random agents of the different types. This can be implemented in various ways. What I do here is using a list with the different classes (*i.e.*, types of strategies). By drawing a random number from a unit interval, multiplying it with the lenght of the list of classes and flooring the resulting number to an integer, I now have a random index into this list with the different classes. Next, I can get the class from the list and instantiate the agent object.\n\nSome minor points with instantiating the agents. First, we give the agent a position, called pos, this is a default attribute assumed by MESA. We also still need a unique ID for the agent, we do this with a simple counter (`self.agent_id`). `self.grid.coord_iter` is a method on the grid. It returns an iterator over the cells in the grid. This iterator returns the agent occupying the cell and the x and y coordinate. Since the first item is `null` because we are filling the grid, we can ignore this. We do this by using the underscore variable name (`_`). This is a python convention. \n\nOnce we have instantiated the agent, we place the agent in the grid and add it to our collection of agents. If you look in more detail at the model class, you will see that I use a set for agents, rather than a list. The reason for this is that we are going to remove agents in the evolutionary phase. Removing agents from a list is memory and compute intensive, while it is computationally easy and cheap when we have a set. \n\n```python\nself.grid = SingleGrid(width, height, torus=True)\n\nstrategies = AxelrodAgent.__subclasses__()\nnum_strategies = len(strategies)\nself.agent_id = 0\n\nfor cell in self.grid.coord_iter():\n _, x, y = cell\n pos = (x, y)\n\n self.agent_id += 1\n\n strategy_index = int(floor(self.random.random()*num_strategies))\n\n agent = strategies[strategy_index](self.agent_id, pos, self)\n self.grid.position_agent(agent, (x, y))\n self.agents.add(agent)\n```\n\nWe also use a DataCollector. This is a default class provided by MESA that can be used for keeping track of relevant statistics. It can store both model level variables as well as agent level variables. Here we are only using model level variables (i.e. attributes of the model). Specifically, we are going to have an attribute on the model for each type of agent strategy (i.e. classes). This attribute is the current count of agents in the grid of the specific type. To implement this, we need to do several things.\n\n1. initialize a data collector instance\n2. at every step update the current count of agents of each strategy\n3. collect the current counts with the data collector.\n\nFor step 1, we set a DataCollector as an attribute. This datacollector needs to know the names of the attributes on the model it needs to collect. So we pass a dict as kwarg to model_reporters. This dict has as key the name by which the variable will be known in the DataCollector. As value, I pass the name of the attribute on the model, but it can also be a function or method which returns a number. Note th at the ``klass`` misspelling is deliberate. The word ``class`` is protected in Python, so you cannot use it as a variable name. It is common practice to use ``klass`` instead in the rare cases were you are having variable refering to a specific class.\n\n```python\nself.datacollector = DataCollector(model_reporters={klass.__name__:klass.__name__\n for klass in strategies})\n```\n\nFor step 2, we need to count at every step the number of agents per strategy type. To help keep track of this, we define a new method, `count_agent_types`. The main magic is the use of `setattr` which is a standard python function for setting attributes to a particular value on a given object. This reason for writing our code this way is that we automatically adapt our attributes to the classes of agents we have, rather than hardcoding the agent classes as attributes on our model. If we now add new classes of agents, we don't need to change the model code itself. There is also a ``getattr`` function, which is used by for example the DataCollector to get the values for the specified attribute names. \n\n```python\ndef count_agent_types(self):\n counter = Counter()\n for agent in self.agents:\n counter[agent.__class__.__name__] += 1\n\n for k,v in counter.items():\n setattr(self, k, v) \n\n```\n\nFor step 3, we modify the first part of the ``step`` method. We first count the types of agents and next collect this data with the datacollector.\n```python\nself.count_agent_types()\nself.datacollector.collect(self)\n```\n\nThe remainder of the ``step`` method has also been changed quite substantially. First, We have to change against whom each agent is playing. We do this by iterating over all agents in the model. Next, we use the grid to give us the neighbors of a given agent. By setting the kwarg ``moore`` to ``True``, we indicate that we include also our diagonal neighbors. Next, we play as we did before in the noisy version of the Axelrod model.\n\n```python\nfor agent_a in self.agents:\n for agent_b in self.grid.neighbor_iter(agent_a.pos, moore=True):\n for _ in range(self.num_iterations):\n```\n\nSecond, we have to add the evolutionary dynamic. This is a bit tricky. First, we loop again over all agents in the model. We check its neighbors and see which strategy performed best. If this is of a different type (``not isinstance(agent_a, best_strategy)``, we add it to a list of agents that needs to be changed and the type of agent to which it needs to be changed. Once we know all agents that need to be changed, we can make this change. \n\nMaking the change is quite straighforward. We remove the agent from the set of agents (`self.agents`) and from the grid. Next we get the position of the agent, we increment our unique ID counter, and create a new agent. This new agent is than added to the grid and to the set of agents. \n\n```python\n# evolution\nagents_to_change = []\nfor agent_a in self.agents:\n neighborhood = self.grid.iter_neighbors(agent_a.pos, moore=True,\n include_center=True)\n neighborhood = ([n for n in neighborhood])\n neighborhood.sort(key=lambda x:x.points, reverse=True)\n best_strategy = neighborhood[0].__class__\n # if best type of strategy in neighborhood is\n # different from strategy type of agent, we need\n # to change our strategy\n if not isinstance(agent_a, best_strategy):\n agents_to_change.append((agent_a, best_strategy))\n\nfor entry in agents_to_change:\n agent, klass = entry\n self.agents.remove(agent)\n self.grid.remove_agent(agent)\n\n pos = agent.pos\n self.agent_id += 1\n\n new_agent = klass(self.agent_id, pos, self)\n self.grid.position_agent(new_agent, pos)\n self.agents.add(new_agent)\n\n```\n\n## Assignment 1\nCan you explain why we need to first loop over all agents before we are changing a given agent to a different strategy?",
"_____no_output_____"
],
[
"## Assignment 2\nAdd all agents classes (i.e., strategie) from the previous assignment to this model. Note that you might have to update the ``__init__`` method to reflect the new pos keyword argument and attribute. ",
"_____no_output_____"
],
[
"\n## Assignment 3\nRun the model for 50 steps, and with 200 rounds of the iterated game. Use the defaults for all other keyword arguments.\n\nPlot the results. \n\n*hint: the DataCollector can return the statistics it has collected as a dataframe, which in turn you can plot directly.*",
"_____no_output_____"
],
[
"This new model is quite a bit noisier than previously. We have a random initialization of the grid and depending on the initial neighborhood, different evolutionary dynamics can happen. On top, we have the noise in game play, and the random agent. \n\n\n## Assignment 4\nLet's explore the model for 10 replications. Run the model 10 times, with 200 rounds of the iterated prisoners dilemma. Run each model for fifty steps. Plot the results for each run. \n\n1. Can you say anything generalizable about the behavioral dynamics of the model?\n2. What do you find striking in the results and why?\n3. If you compare the results for this spatially explicit version of the Emergence of Cooperation with the none spatially explicit version, what are the most important differences in dynamics. Can you explain why adding local interactions results in these changes? ",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
4acdc08b429eeb1b780de97b82eeb7cc4c98b8bc
| 815,759 |
ipynb
|
Jupyter Notebook
|
Supervised_Classification_of_IMG_DATA_1-Abordagem diferente para o alinhamentos dos dados.ipynb
|
SimaoBarbosa/Land_cover_classification
|
20f5d957cbcd664648d04b9f37aaa81b1000b283
|
[
"MIT"
] | null | null | null |
Supervised_Classification_of_IMG_DATA_1-Abordagem diferente para o alinhamentos dos dados.ipynb
|
SimaoBarbosa/Land_cover_classification
|
20f5d957cbcd664648d04b9f37aaa81b1000b283
|
[
"MIT"
] | null | null | null |
Supervised_Classification_of_IMG_DATA_1-Abordagem diferente para o alinhamentos dos dados.ipynb
|
SimaoBarbosa/Land_cover_classification
|
20f5d957cbcd664648d04b9f37aaa81b1000b283
|
[
"MIT"
] | null | null | null | 375.925806 | 415,312 | 0.927323 |
[
[
[
"Code adapted from https://github.com/patrickcgray/open-geo-tutorial",
"_____no_output_____"
]
],
[
[
"from IPython.display import Audio, display\nfrom timeit import default_timer as timer\nstart = timer()\ndef color_stretch(image, index):\n colors = image[:, :, index].astype(np.float64)\n for b in range(colors.shape[2]):\n colors[:, :, b] = rasterio.plot.adjust_band(colors[:, :, b])\n return colors\ndef alert():\n global start\n end= timer()\n print(\"\\n\\nDuration in minutes : \" + str((end - start)/60) + \" minutes.\" )\n display(Audio(url='https://sound.peal.io/ps/audios/000/000/537/original/woo_vu_luvub_dub_dub.wav', autoplay=True))",
"_____no_output_____"
]
],
[
[
"# Preparing Dataset",
"_____no_output_____"
],
[
"#### Cutting images for faster process",
"_____no_output_____"
]
],
[
[
"# 0 to 10980 for this img_data\n#training\n#xmin = 6700\n#xmax = 6900\n#ymin = 6700\n#ymax = 6900\nxmin = 6700\nxmax = 7000\nymin = 6600\nymax = 6900\nxmin_test = 6500\nxmax_test = 6700\nymin_test = 6750\nymax_test = 6950\n\nshapemin = min(xmin,ymin,xmin_test,ymin_test)\nshapemax = max(xmax,ymax,xmax_test,ymax_test)\nprint(shapemin)\nshapemax",
"6500\n"
],
[
"import rasterio\nfrom rasterio.mask import mask\nimport geopandas as gp\nimport numpy as np\nimport shapely\nfrom shapely import geometry\nfrom shapely.geometry import shape, Point, LineString, Polygon , mapping\nimport matplotlib.pyplot as plt\nfrom rasterio.plot import show\nimport pyproj\nfrom pyproj import CRS\nimport fiona\nimport os # we need os to do some basic file operations\nimport re\nfrom rasterio.plot import adjust_band\nfrom rasterio.plot import reshape_as_raster, reshape_as_image\nimport matplotlib.pyplot as plt\nfrom rasterio.plot import show\nfrom pyproj import transform\nfrom pyproj import Proj\nimport ast",
"_____no_output_____"
],
[
"# create a products directory within the data dir which won't be uploaded to Github\nimg_dir = './data/'\n\n# check to see if the dir it exists, if not, create it\nif not os.path.exists(img_dir):\n os.makedirs(img_dir)\n\n# filepath for image we're writing out\nimg_fp = img_dir + 'sentinel_bands_img_data_1.tif'",
"_____no_output_____"
],
[
"full_dataset = rasterio.open(img_fp)\nimg_rows, img_cols = full_dataset.shape\nimg_bands = full_dataset.count\nprint(full_dataset.shape) # dimensions\nprint(full_dataset.count) # bands",
"(10980, 10980)\n4\n"
],
[
"with rasterio.open(img_fp) as src:\n # may need to reduce this image size if your kernel crashes, takes a lot of memory\n img = src.read()[:, : , : ]\n\n# Take our full image and reshape into long 2d array (nrow * ncol, nband) for classification\n",
"_____no_output_____"
],
[
"img_train = img[:, xmin : xmax , ymin : ymax ]\nimg_test = img[:, xmin_test : xmax_test , ymin_test : ymax_test ]\nreshaped_img_train = reshape_as_image(img_train)\nreshaped_img_test = reshape_as_image(img_test)\n\nfig, axs = plt.subplots(2,1,figsize=(15,15))\nimg_stretched_train = color_stretch(reshaped_img_train, [2, 1, 0])\naxs[0].imshow(img_stretched_train)\n\nimg_stretched_test = color_stretch(reshaped_img_test, [2, 1, 0])\naxs[1].imshow(img_stretched_test)",
"_____no_output_____"
]
],
[
[
"# Intercepting label data with cutting image",
"_____no_output_____"
]
],
[
[
"# Open the dataset from the file\nshapefile = gp.read_file('./data/shapefiles/shapefile_compressed/shp_2012_IMG_DATA_1.shp')\nshapefile = shapefile.filter(['Legenda','geometry','CLC2012'])\nshapefile.crs = {'init': 'epsg:4326'}",
"_____no_output_____"
],
[
"shapefile",
"_____no_output_____"
],
[
"shapefile.bounds",
"_____no_output_____"
],
[
"from pyproj import transform\nfrom pyproj import Proj\ndef cut_bounds( xmin , xmax , ymin , ymax ):\n # this will get our four corner points\n xmin = xmin \n ymin = ymin\n xmax = xmax\n ymax = ymax\n raster_gps_points = full_dataset.transform * (xmin, ymin),full_dataset.transform * (xmax, ymin),full_dataset.transform * (xmax, ymax), full_dataset.transform * (xmin, ymax),\n # Project all longitudes, latitudes using the pyproj package\n p1 = Proj(full_dataset.crs) # our current crs\n print(p1)\n print('raster bounds in current crs :\\n', raster_gps_points)\n return raster_gps_points\nraster_gps_points = cut_bounds( shapemin , shapemax , shapemin , shapemax )",
"Proj('+proj=utm +zone=29 +datum=WGS84 +units=m +no_defs', preserve_units=True)\nraster bounds in current crs :\n ((464960.0, 4335040.0), (469960.0, 4335040.0), (469960.0, 4330040.0), (464960.0, 4330040.0))\n"
],
[
"def shapefile_from_rasterbounds(raster_gps_points,shapefile) :\n polygon = Polygon( list(raster_gps_points))\n print(\"Polygon raster gps points\",polygon)\n imageBounds = gp.GeoDataFrame(crs=CRS.from_epsg(32629))\n imageBounds.geometry = [polygon]\n imageBounds = imageBounds.to_crs(fiona.crs.from_epsg(4326))\n print(\"Imaged cutted bounds\",imageBounds.bounds)\n shapefile = shapefile.to_crs(fiona.crs.from_epsg(4326))\n print(\"Shapefile bounds\",shapefile.bounds)\n intersected = gp.sjoin( shapefile, imageBounds, how='inner', op='intersects', lsuffix='left', rsuffix='right')\n intersected = intersected.reset_index()\n print(\"Intersected df\",intersected)\n print(\"Intersected crs\",intersected.crs)\n print(\"Intersected bounds\",intersected.bounds)\n return intersected\nshapefile = shapefile_from_rasterbounds(raster_gps_points,shapefile)",
"Polygon raster gps points POLYGON ((464960 4335040, 469960 4335040, 469960 4330040, 464960 4330040, 464960 4335040))\n"
],
[
"import re\ndef class_from_CLCcode(clc) :\n mega_classes = {\n '^1.*' : 'Territórios artificializados',\n '^2.*' : 'Agricultura',\n '^31.*' : 'Floresta',\n '^3.4' : 'Floresta',\n '^32[123]' : 'Vegetação natural',\n '^33[123]' : 'Espaços descobertos ou com vegetação esparsa',\n '^41.*' : 'Zonas húmidas',\n '^42[13]' : 'Zonas húmidas',\n '^5.*' : 'Corpos de água',\n '^422' : 'Corpos de água'\n }\n for mega_class_key in mega_classes :\n searched = re.search(mega_class_key, str(clc))\n if searched is not None:\n return mega_classes[mega_class_key]\n return 'Not Defined'\n\nx = shapefile['CLC2012'][0] \nprint (x)\nclass_from_CLCcode (x) ",
"111\n"
],
[
"for i in shapefile.index:\n clc = shapefile.at[i, 'CLC2012']\n new_class = class_from_CLCcode(clc)\n shapefile.at[i, 'Mega_Legenda'] = new_class\nshapefile.head()",
"_____no_output_____"
]
],
[
[
"# Setting up label data with bands",
"_____no_output_____"
]
],
[
[
"def setup_shafile_to_data(shapefile,full_dataset) :\n shapefile = shapefile.filter(['Legenda','Mega_Legenda', 'geometry','CLC2012'])\n print(\"Data crs: \" + str(full_dataset.crs))\n print(\"Before crs: \" + str(shapefile.crs))\n if shapefile.crs != full_dataset.crs :\n shapefile = shapefile.to_crs(full_dataset.crs)\n print(\"After crs: \" + str(shapefile.crs))\n return shapefile\nshapefile = setup_shafile_to_data(shapefile,full_dataset)\nprint(\"SIZE:\",shapefile.size)\nshapefile.head()",
"Data crs: EPSG:32629\nBefore crs: {'init': 'epsg:4326'}\n"
]
],
[
[
"Choose target column = Legenda or Mega_Legenda",
"_____no_output_____"
]
],
[
[
"Class = 'Mega_Legenda'\n#Class = 'Legenda' ",
"_____no_output_____"
],
[
"unique= np.unique(shapefile[Class])\nprint('List of Land Cover Classes:')\nunique",
"List of Land Cover Classes:\n"
],
[
"import matplotlib.pyplot as plt\nunique, counts = np.unique(shapefile[Class], return_counts=True)\nplt.bar(unique, counts, 1)\nplt.title('Class Frequency')\nplt.xlabel('Class')\nplt.ylabel('Frequency')\nplt.show()\n\nprint('Number of different classes :' + str(len(unique)))\nprint(unique)",
"_____no_output_____"
],
[
"band_count = full_dataset.count\ntransf = full_dataset.transform\nfull_dataset_array = img\ncutted =full_dataset_array[:, xmin : xmax , ymin : ymax ]\ncutted_test = full_dataset_array [:, xmin_test : xmax_test , ymin_test : ymax_test ]\nprint(cutted.shape)\ncutted_test.shape",
"(4, 300, 300)\n"
],
[
"import numba\nfrom numba import jit,prange\n\n#@jit(parallel=True) \ndef setup_cell_noglobals(i,j,xmin,xmax, ymin, ymax , cutted ,transf , band_count,shapefile):\n coordinates = transf * (xmin + i,ymin + j)\n point = Point(coordinates)\n geometry = shapefile['geometry']\n geometry = geometry[geometry.contains(point)]\n if geometry.size>0 :\n return geometry.index[0]\n else : \n print(\"Not found in shapefile\")\n return -1\n\n#@jit(parallel=True) \ndef setup_X_and_y_noglobals(xmin,xmax, ymin, ymax , cutted ,transf , band_count,shapefile):\n rX = np.array([], dtype=np.int8).reshape(0,band_count) # pixels for training\n ry = np.array([], dtype=np.string_) # labels for training\n for i in range(xmax-xmin):\n print(i)\n for j in range(ymax-ymin):\n result = setup_cell_noglobals(i,j,xmin,xmax, ymin, ymax , cutted ,transf , band_count,shapefile)\n if (result!=-1) : \n target = shapefile.at[ result ,Class]\n list_x = []\n for band in range(band_count):\n list_x.append( cutted[band][i][j] )\n x_entry = np.array( list_x )\n rX = np.vstack((rX, x_entry ))\n ry = np.append(ry,target)\n return rX,ry",
"_____no_output_____"
],
[
"X,y = setup_X_and_y_noglobals(xmin,xmax, ymin, ymax, cutted ,transf , band_count,shapefile)\nX_test,y_test = setup_X_and_y_noglobals(xmin_test,xmax_test, ymin_test, ymax_test, cutted_test ,transf , band_count,shapefile)\nprint(y.shape)\nprint(y_test.shape)",
"0\n1\n2\n3\n4\n5\n6\n7\n8\n9\n10\n11\n12\n13\n14\n15\n16\n17\n18\n19\n20\n21\n22\n23\n24\n25\n26\n27\n28\n29\n30\n31\n32\n33\n34\n35\n36\n37\n38\n39\n40\n41\n42\n43\n44\n45\n46\n47\n48\n49\n50\n51\n52\n53\n54\n55\n56\n57\n58\n59\n60\n61\n62\n63\n64\n65\n66\n67\n68\n69\n70\n71\n72\n73\n74\n75\n76\n77\n78\n79\n80\n81\n82\n83\n84\n85\n86\n87\n88\n89\n90\n91\n92\n93\n94\n95\n96\n97\n98\n99\n100\n101\n102\n103\n104\n105\n106\n107\n108\n109\n110\n111\n112\n113\n114\n115\n116\n117\n118\n119\n120\n121\n122\n123\n124\n125\n126\n127\n128\n129\n130\n131\n132\n133\n134\n135\n136\n137\n138\n139\n140\n141\n142\n143\n144\n145\n146\n147\n148\n149\n150\n151\n152\n153\n154\n155\n156\n157\n158\n159\n160\n161\n162\n163\n164\n165\n166\n167\n168\n169\n170\n171\n172\n173\n174\n175\n176\n177\n178\n179\n180\n181\n182\n183\n184\n185\n186\n187\n188\n189\n190\n191\n192\n193\n194\n195\n196\n197\n198\n199\n200\n201\n202\n203\n204\n205\n206\n207\n208\n209\n210\n211\n212\n213\n214\n215\n216\n217\n218\n219\n220\n221\n222\n223\n224\n225\n226\n227\n228\n229\n230\n231\n232\n233\n234\n235\n236\n237\n238\n239\n240\n241\n242\n243\n244\n245\n246\n247\n248\n249\n250\n251\n252\n253\n254\n255\n256\n257\n258\n259\n260\n261\n262\n263\n264\n265\n266\n267\n268\n269\n270\n271\n272\n273\n274\n275\n276\n277\n278\n279\n280\n281\n282\n283\n284\n285\n286\n287\n288\n289\n290\n291\n292\n293\n294\n295\n296\n297\n298\n299\n0\n1\n2\n3\n4\n5\n6\n7\n8\n9\n10\n11\n12\n13\n14\n15\n16\n17\n18\n19\n20\n21\n22\n23\n24\n25\n26\n27\n28\n29\n30\n31\n32\n33\n34\n35\n36\n37\n38\n39\n40\n41\n42\n43\n44\n45\n46\n47\n48\n49\n50\n51\n52\n53\n54\n55\n56\n57\n58\n59\n60\n61\n62\n63\n64\n65\n66\n67\n68\n69\n70\n71\n72\n73\n74\n75\n76\n77\n78\n79\n80\n81\n82\n83\n84\n85\n86\n87\n88\n89\n90\n91\n92\n93\n94\n95\n96\n97\n98\n99\n100\n101\n102\n103\n104\n105\n106\n107\n108\n109\n110\n111\n112\n113\n114\n115\n116\n117\n118\n119\n120\n121\n122\n123\n124\n125\n126\n127\n128\n129\n130\n131\n132\n133\n134\n135\n136\n137\n138\n139\n140\n141\n142\n143\n144\n145\n146\n147\n148\n149\n150\n151\n152\n153\n154\n155\n156\n157\n158\n159\n160\n161\n162\n163\n164\n165\n166\n167\n168\n169\n170\n171\n172\n173\n174\n175\n176\n177\n178\n179\n180\n181\n182\n183\n184\n185\n186\n187\n188\n189\n190\n191\n192\n193\n194\n195\n196\n197\n198\n199\n(90000,)\n(40000,)\n"
],
[
"# What are our classification labels?\n\nlabels = np.unique( y )\nprint('The training data include {n} classes: {classes}\\n'.format(n=labels.size, classes=labels))\n\nlabels_test = np.unique( y_test )\nprint('The testing data include {n} classes: {classes}\\n'.format(n=labels_test.size, classes=labels_test))\n\n# We will need a \"X\" matrix containing our features, and a \"y\" array containing our labels\nprint('Our X matrix is sized: {sz}'.format(sz=X.shape))\nprint('Our y array is sized: {sz}'.format(sz=y.shape))\n\nprint('Our X_test matrix is sized: {sz}'.format(sz=X_test.shape))\nprint('Our y_test array is sized: {sz}'.format(sz=y_test.shape))\n",
"The training data include 5 classes: ['Agricultura' 'Corpos de água' 'Floresta' 'Territórios artificializados'\n 'Vegetação natural']\n\nThe testing data include 3 classes: ['Agricultura' 'Corpos de água' 'Territórios artificializados']\n\nOur X matrix is sized: (90000, 4)\nOur y array is sized: (90000,)\nOur X_test matrix is sized: (40000, 4)\nOur y_test array is sized: (40000,)\n"
],
[
"fig, ax = plt.subplots(1,3, figsize=[20,8])\n\n# numbers 1-4\nband_count = np.arange(1,5)\n\nclasses = np.unique(y)\nfor class_type in classes:\n band_intensity = np.mean(X[y==class_type, :], axis=0)\n ax[0].plot(band_count, band_intensity, label=class_type)\n ax[1].plot(band_count, band_intensity, label=class_type)\n ax[2].plot(band_count, band_intensity, label=class_type)\n# plot them as lines\n\n# Add some axis labels\nax[0].set_xlabel('Band #')\nax[0].set_ylabel('Reflectance Value')\nax[1].set_ylabel('Reflectance Value')\nax[1].set_xlabel('Band #')\nax[2].set_ylabel('Reflectance Value')\nax[2].set_xlabel('Band #')\n#ax[0].set_ylim(32,38)\nax[1].set_ylim(32,38)\nax[2].set_ylim(70,140)\n#ax.set\nax[1].legend(loc=\"upper right\")\n# Add a title\nax[0].set_title('Band Intensities Full Overview')\nax[1].set_title('Band Intensities Lower Ref Subset')\nax[2].set_title('Band Intensities Higher Ref Subset')",
"_____no_output_____"
],
[
"def str_class_to_int(class_array):\n class_array[class_array == 'Territórios artificializados'] = 0\n class_array[class_array == 'Agricultura'] = 1\n class_array[class_array == 'Floresta'] = 2\n class_array[class_array == 'Vegetação natural'] = 3\n class_array[class_array == 'Espaços descobertos ou com vegetação esparsa'] = 4\n class_array[class_array == 'Zonas húmidas'] = 5\n class_array[class_array == 'Corpos de água'] = 6\n return(class_array.astype(int))\ndef str_class_to_int_v2(class_array,class_test):\n uniques = np.unique(class_array)\n i=0\n for class_name in uniques :\n class_array[class_array == class_name ] = i\n class_test[class_test == class_name ] = i\n i+=1\n return(class_array.astype(int) , class_test.astype(int) ) ",
"_____no_output_____"
],
[
"print(np.unique(y))\nprint(np.unique(y_test))\ny = str_class_to_int(y)\ny_test = str_class_to_int(y_test)\nprint(np.unique(y))\nprint(np.unique(y_test))\n",
"['Agricultura' 'Corpos de água' 'Floresta' 'Territórios artificializados'\n 'Vegetação natural']\n['Agricultura' 'Corpos de água' 'Territórios artificializados']\n[0 1 2 3 6]\n[0 1 6]\n"
]
],
[
[
"# Supervised Classification Algorithm",
"_____no_output_____"
]
],
[
[
"from sklearn.svm import LinearSVC\nfrom sklearn.gaussian_process import GaussianProcessClassifier\nfrom sklearn.naive_bayes import GaussianNB\nfrom sklearn.neighbors import KNeighborsClassifier\n#model = LinearSVC(C=1,verbose=1)\nmodel = KNeighborsClassifier()\nmodel.fit(X, y)\nscore = model.score(X_test, y_test)\nscore",
"_____no_output_____"
],
[
"class_prediction = model.predict(reshaped_img_test.reshape(-1, 4))\nbefore = class_prediction.shape\n# Reshape our classification map back into a 2D matrix so we can visualize it\nclass_prediction = class_prediction.reshape(reshaped_img_test[:, :, 0].shape)\nclass_prediction.shape",
"_____no_output_____"
],
[
"real_img = y_test\nprint(before)\nprint(real_img.shape)\nprint(y_test.shape)\n# Reshape our classification map back into a 2D matrix so we can visualize it\nreal_class = real_img.reshape(reshaped_img_test[:, :, 0].shape) #fixme\nreal_class.shape",
"(40000,)\n(40000,)\n(40000,)\n"
],
[
"real_class = real_class.astype(int)\nprint(real_class)\nclass_prediction = class_prediction.astype(int)\nclass_prediction",
"[[6 6 6 ... 6 6 6]\n [6 6 6 ... 6 6 6]\n [6 6 6 ... 6 6 6]\n ...\n [6 6 6 ... 0 0 0]\n [6 6 6 ... 0 0 0]\n [6 6 6 ... 0 0 0]]\n"
]
],
[
[
"# Visualizing results ",
"_____no_output_____"
]
],
[
[
" \n# find the highest pixel value in the prediction image\nn = int(np.max(class_prediction))\n\n# next setup a colormap for our map\ncolors = dict((\n (0, (60, 60, 60, 255)), # Grey - Territórios artificializados\n (1, (20,230,20, 255)), # Light Green - Agricultura\n (2, (5, 80, 5, 255)), # Dark Green - Floresta\n (3, (170, 200, 20, 255)), # Yellow - Vegetação natural\n (4, (50, 20, 0, 255)), # Brown - Espaços descobertos ou com vegetação esparsa\n (5, (0, 250, 250, 255)), # Light Blue - Zonas húmidas\n (6, (0, 0, 200, 255)), # Dark Blue - Corpos de água\n))\n\n# Put 0 - 255 as float 0 - 1\nfor k in colors:\n v = colors[k]\n _v = [_v / 255.0 for _v in v]\n colors[k] = _v\n \nindex_colors = [colors[key] if key in colors else \n (255, 255, 255, 0) for key in range(0, n+1)]\n\ncmap = plt.matplotlib.colors.ListedColormap(index_colors, 'Classification', n+1)",
"_____no_output_____"
],
[
"def discrete_matshow(data):\n #get discrete colormap\n cmap = plt.get_cmap('RdBu', np.max(data)-np.min(data)+1)\n # set limits .5 outside true range\n mat = plt.matshow(data,cmap=cmap,vmin = np.min(data)-.5, vmax = np.max(data)+.5)\n #tell the colorbar to tick at integers\n cax = plt.colorbar(mat, ticks=np.arange(np.min(data),np.max(data)+1))",
"_____no_output_____"
],
[
"\nfig, axs = plt.subplots(1,2,figsize=(15,15))\n\n\naxs[0].imshow(img_stretched_test)\n\nprediction_plot= axs[1].imshow(class_prediction, cmap=cmap, interpolation='none')\nfig.colorbar(prediction_plot, ax=axs[1],ticks=range(0,7),fraction=0.046, pad=0.04)\n\n#real_plot = axs[2].imshow(real_class, cmap=cmap, interpolation='none')\n#fig.colorbar(real_plot, ax=axs[2],ticks=range(0,6),fraction=0.046, pad=0.04)\n\nfig.show()",
"C:\\ProgramData\\Anaconda3\\envs\\geo_py37\\lib\\site-packages\\ipykernel_launcher.py:12: UserWarning: Matplotlib is currently using module://ipykernel.pylab.backend_inline, which is a non-GUI backend, so cannot show the figure.\n if sys.path[0] == '':\n"
],
[
"fig, axs = plt.subplots(1,2,figsize=(15,15))\n\naxs[0].imshow(img_stretched_test[50:150, 50:150])\nprediction_plot = axs[1].imshow(class_prediction[50:150, 50:150], cmap=cmap, interpolation='none')\nfig.colorbar(prediction_plot, ax=axs[1],ticks=range(0,7),fraction=0.046, pad=0.04)\nfig.show()\n",
"C:\\ProgramData\\Anaconda3\\envs\\geo_py37\\lib\\site-packages\\ipykernel_launcher.py:6: UserWarning: Matplotlib is currently using module://ipykernel.pylab.backend_inline, which is a non-GUI backend, so cannot show the figure.\n \n"
],
[
"alert() # audio alert",
"\n\nDuration in minutes : 20.00997302 minutes.\n"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
]
] |
4acdc757fe978d9df543242c6731e297e9a0cf07
| 479,821 |
ipynb
|
Jupyter Notebook
|
ADS-Fall2018/WineRecommender.ipynb
|
peperaj/XBUS-507-01.Applied_Data_Science
|
0928b9aa000bfaabef13a3a59c0c80c68307013b
|
[
"MIT"
] | 9 |
2019-06-21T21:38:46.000Z
|
2022-01-11T18:16:22.000Z
|
ADS-Fall2018/WineRecommender.ipynb
|
peperaj/XBUS-507-01.Applied_Data_Science
|
0928b9aa000bfaabef13a3a59c0c80c68307013b
|
[
"MIT"
] | 3 |
2019-08-24T18:56:27.000Z
|
2020-01-04T19:26:32.000Z
|
ADS-Fall2018/WineRecommender.ipynb
|
peperaj/XBUS-507-01.Applied_Data_Science
|
0928b9aa000bfaabef13a3a59c0c80c68307013b
|
[
"MIT"
] | 13 |
2018-09-28T21:39:41.000Z
|
2022-01-11T18:16:25.000Z
| 142.50698 | 150,116 | 0.819064 |
[
[
[
"## Getting Data",
"_____no_output_____"
]
],
[
[
"#import os \n#import requests \n\n#DATASET = (\n# \"https://archive.ics.uci.edu/ml/machine-learning-databases/abalone/abalone.data\",\n# \"https://archive.ics.uci.edu/ml/machine-learning-databases/abalone/abalone.names\"\n#)\n\n#def download_data(path='data', urls=DATASET):\n# if not os.path.exists(path):\n# os.mkdir(path) \n# \n# for url in urls:\n# response = requests.get(url)\n# name = os.path.basename(url) \n# with open(os.path.join(path, name), 'wb') as f: \n# f.write(response.content)\n\n#download_data()\n\n\n#DOWNLOAD AND LOAD IN DATA FROM URL!!!!!!!!!!!!!!!!!!!\n#import requests\n#import io\n#data = io.BytesIO(requests.get('URL HERE'))\n#whitedata = pd.read_csv(data.content)\n#whitedata.head()",
"_____no_output_____"
]
],
[
[
"## Load Data",
"_____no_output_____"
]
],
[
[
"import pandas as pd\nimport yellowbrick\nimport matplotlib.pyplot as plt\nimport seaborn as sns\n\nimport numpy as np\nimport io\nimport sklearn\n\ncolumns = [\n \"fixed acidity\",\n \"volatile acidity\",\n \"citric acid\",\n \"residual sugar\",\n \"chlorides\",\n \"free sulfur dioxide\",\n \"total sulfur dioxide\",\n \"density\",\n \"pH\",\n \"sulphates\",\n \"alcohol\",\n \"quality\"\n]\n\nreddata = pd.read_csv('data/winequality-red.csv', sep=\";\")\nwhitedata = pd.read_csv('data/winequality-white.csv', sep=\";\")",
"_____no_output_____"
]
],
[
[
"## Check it out",
"_____no_output_____"
]
],
[
[
"whitedata.head(10)",
"_____no_output_____"
],
[
"whitedata.describe()",
"_____no_output_____"
],
[
"whitedata.info()",
"<class 'pandas.core.frame.DataFrame'>\nRangeIndex: 4898 entries, 0 to 4897\nData columns (total 12 columns):\nfixed acidity 4898 non-null float64\nvolatile acidity 4898 non-null float64\ncitric acid 4898 non-null float64\nresidual sugar 4898 non-null float64\nchlorides 4898 non-null float64\nfree sulfur dioxide 4898 non-null float64\ntotal sulfur dioxide 4898 non-null float64\ndensity 4898 non-null float64\npH 4898 non-null float64\nsulphates 4898 non-null float64\nalcohol 4898 non-null float64\nquality 4898 non-null int64\ndtypes: float64(11), int64(1)\nmemory usage: 459.3 KB\n"
],
[
"whitedata.pH.describe()",
"_____no_output_____"
],
[
"whitedata['poor'] = np.where(whitedata['quality'] < 5, 1, 0)",
"_____no_output_____"
],
[
"whitedata['poor'].value_counts()",
"_____no_output_____"
],
[
"whitedata['expected'] = np.where(whitedata['quality'] > 4, 1, 0)\nwhitedata['expected'].value_counts()",
"_____no_output_____"
],
[
"whitedata['expected'].describe()",
"_____no_output_____"
],
[
"whitedata.head()",
"_____no_output_____"
],
[
"#set up the figure size\n%matplotlib inline\nplt.rcParams['figure.figsize'] = (30, 30)\n\n#make the subplot\nfig, axes = plt.subplots(nrows = 6, ncols = 2)\n\n#specify the features of intersest\nnum_features = ['pH', 'alcohol', 'citric acid', 'chlorides', 'residual sugar', 'free sulfur dioxide', 'total sulfur dioxide', 'density', 'sulphates']\n\nxaxes = num_features\nyaxes = ['Counts', 'Counts', 'Counts', 'Counts', 'Counts', 'Counts', 'Counts', 'Counts', 'Counts']\n\n#draw the histogram\naxes = axes.ravel()\nfor idx, ax in enumerate(axes):\n ax.hist(whitedata[num_features[idx]].dropna(), bins = 30)\n ax.set_xlabel(xaxes[idx], fontsize = 20)\n ax.set_ylabel(yaxes[idx], fontsize = 20)\n ax.tick_params(axis = 'both', labelsize = 20)",
"_____no_output_____"
],
[
"\nfeatures = ['pH', 'alcohol', 'citric acid', 'residual sugar', 'chlorides', 'free sulfur dioxide', 'total sulfur dioxide', 'density', 'sulphates']\nclasses = ['unexpected', 'expected']\n\nX = whitedata[features].as_matrix()\ny = whitedata['quality'].as_matrix()",
"C:\\Users\\Trypt\\AppData\\Local\\Continuum\\anaconda3\\lib\\site-packages\\ipykernel_launcher.py:5: FutureWarning: Method .as_matrix will be removed in a future version. Use .values instead.\n \"\"\"\nC:\\Users\\Trypt\\AppData\\Local\\Continuum\\anaconda3\\lib\\site-packages\\ipykernel_launcher.py:6: FutureWarning: Method .as_matrix will be removed in a future version. Use .values instead.\n \n"
],
[
"from sklearn.naive_bayes import GaussianNB",
"_____no_output_____"
],
[
"viz = GaussianNB()\nviz.fit(X, y)\nviz.score(X, y)",
"_____no_output_____"
],
[
"from sklearn.naive_bayes import MultinomialNB",
"_____no_output_____"
],
[
"viz = MultinomialNB()\nviz.fit(X, y)\nviz.score(X, y)",
"_____no_output_____"
],
[
"from sklearn.naive_bayes import BernoulliNB",
"_____no_output_____"
],
[
"viz = BernoulliNB()\nviz.fit(X, y)\nviz.score(X, y)",
"_____no_output_____"
]
],
[
[
"## Features and Targets",
"_____no_output_____"
]
],
[
[
"y = whitedata[\"quality\"]",
"_____no_output_____"
],
[
"y",
"_____no_output_____"
],
[
"X = whitedata.iloc[:,1:-1]\nX",
"_____no_output_____"
],
[
"from sklearn.ensemble import RandomForestClassifier as rfc\n\nestimator = rfc(n_estimators=7)\nestimator.fit(X,y)",
"_____no_output_____"
],
[
"y_hat = estimator.predict(X)\nprint(y_hat)",
"_____no_output_____"
],
[
"%matplotlib notebook\n\nimport numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\n\nfrom sklearn.ensemble import RandomForestClassifier\nfrom sklearn.model_selection import cross_val_score",
"_____no_output_____"
],
[
"data = pd.read_csv(\".../data/occupency.csv\")\ndata.head()",
"_____no_output_____"
],
[
"X = data(['temperature', 'relative humidity', 'light', 'CO2', 'humidity'])\ny = data('occupency')",
"_____no_output_____"
],
[
"def n_estimators_tuning(X, y, min_estimator=1, max_estimator=50, ax=None, save=None):\n \n if ax is None:\n _, ax = plt.subplots()\n \n mean = []\n stds = []\n n_estimators = np.arrange(min_estimators, mx_estimators+1)\n \n for n in n_estimators:\n model = RandomForestClassifier(n_estimators=n)\n scores = cross_val_score(model, X, y, cv=cv)\n means.append(scores.mean())\n stds.append(scores.std())\n \n means = np.array(means)\n stds = np.array(stds)\n \n ax.plot(n_estimatrors, scores, label='CV+{} scores'.forest(cv))\n ax.fill_between(n_estimators, means-stds, means+stds, alpha=0.3)\n \n max_score = means.max()\n max_score_idx = np.where(means==max_score)[0]\n ax.hline#???????????????\n \n \n \n ax.set_xlim(min_estimators, max_estimators)\n ax.set_xlabel(\"n_estimators\")\n ax.set_ylabel(\"F1 Score\")\n ax.set_title(\"Random Forest Hyperparameter Tuning\")\n ax.legend(loc='best')\n \n if save:\n plt.savefig(save)\n \n return ax\n #print(scores)\n\nn_estimators_tuning(X, y)",
"_____no_output_____"
],
[
"whitedata[winecolor] = 0\nreddata[winecolor] = 1",
"_____no_output_____"
],
[
"df3 = [whitedata, reddata]\ndf = pd.concat(df3)\ndf.reset_index(drop = True, inplace = True)\ndf.isnull().sum()",
"C:\\Users\\Trypt\\AppData\\Local\\Continuum\\anaconda3\\lib\\site-packages\\ipykernel_launcher.py:2: FutureWarning: Sorting because non-concatenation axis is not aligned. A future version\nof pandas will change to not sort by default.\n\nTo accept the future behavior, pass 'sort=True'.\n\nTo retain the current behavior and silence the warning, pass sort=False\n\n \n"
],
[
"whitedata.head()",
"_____no_output_____"
],
[
"reddata.head()",
"_____no_output_____"
],
[
"df = df.drop(columns=['poor', 'expected'])\ndf.isnull().sum()",
"_____no_output_____"
],
[
"df.head()",
"_____no_output_____"
],
[
"df.describe()",
"_____no_output_____"
],
[
"df['recommended'] = np.where(df['quality'] < 6, 0, 1)",
"_____no_output_____"
],
[
"df.head(50)",
"_____no_output_____"
],
[
"df[\"quality\"].value_counts().sort_values(ascending = False)",
"_____no_output_____"
],
[
"df[\"recommended\"].value_counts().sort_values(ascending = False)",
"_____no_output_____"
],
[
"df['recommended'] = np.where(df['quality'] < 6, 0, 1)\ndf[\"recommended\"].value_counts().sort_values(ascending = False)",
"_____no_output_____"
],
[
"from pandas.plotting import radviz\nplt.figure(figsize=(8,8))\nradviz(df, 'recommended', color=['blue', 'red'])\nplt.show()",
"_____no_output_____"
],
[
"features = ['pH', 'alcohol', 'citric acid', 'residual sugar', 'chlorides', 'free sulfur dioxide', 'total sulfur dioxide', 'density', 'sulphates']\n#classes = ['unexpected', 'expected']\n\nX = df[features].as_matrix()\ny = df['recommended'].as_matrix()",
"C:\\Users\\Trypt\\AppData\\Local\\Continuum\\anaconda3\\lib\\site-packages\\ipykernel_launcher.py:4: FutureWarning: Method .as_matrix will be removed in a future version. Use .values instead.\n after removing the cwd from sys.path.\nC:\\Users\\Trypt\\AppData\\Local\\Continuum\\anaconda3\\lib\\site-packages\\ipykernel_launcher.py:5: FutureWarning: Method .as_matrix will be removed in a future version. Use .values instead.\n \"\"\"\n"
],
[
"viz = GaussianNB()\nviz.fit(X, y)\nviz.score(X, y)",
"_____no_output_____"
],
[
"viz = MultinomialNB()\nviz.fit(X, y)\nviz.score(X, y)",
"_____no_output_____"
],
[
"viz = BernoulliNB()\nviz.fit(X, y)\nviz.score(X, y)",
"_____no_output_____"
],
[
"from sklearn.ensemble import RandomForestClassifier\nviz = RandomForestClassifier()\nviz.fit(X, y)\nviz.score(X, y)",
"_____no_output_____"
],
[
"from sklearn.metrics import (auc, roc_curve, recall_score, accuracy_score, confusion_matrix, classification_report, f1_score, precision_score)\nfrom sklearn.model_selection import train_test_split\nX_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=11)\npredicted = viz.predict(X_test)\ncm = confusion_matrix(y_test, predicted)\nfig = plt.figure(figsize=(7, 5))\nax = plt.subplot()\ncm1 = (cm.astype(np.float64) / cm.sum(axis=1, keepdims=1))\ncmap = sns.cubehelix_palette(light=1, as_cmap=True)\nsns.heatmap(cm1, annot=True, ax = ax, cmap=cmap); #annot=True to annotate cells\n# labels, title and ticks\nax.set_xlabel('Features');\nax.set_ylabel('Recommended'); \nax.set_title('Normalized confusion matrix'); \nax.xaxis.set_ticklabels(['Good', 'Bad']); \nax.yaxis.set_ticklabels(['Good', 'Bad']); \nprint(cm)",
"[[457 2]\n [ 5 836]]\n"
],
[
"# Recursive Feature Elimination (RFE) \nfrom sklearn.feature_selection import (chi2, RFE)\n\nmodel = RandomForestClassifier() \nrfe = RFE(model, 38) \nfit = rfe.fit(X, y) \nprint(\"Num Features: \", fit.n_features_) \nprint(\"Selected Features: \", fit.support_) \nprint(\"Feature Ranking: \", fit.ranking_)",
"Num Features: 9\nSelected Features: [ True True True True True True True True True]\nFeature Ranking: [1 1 1 1 1 1 1 1 1]\n"
],
[
"from sklearn.model_selection import StratifiedKFold\nfrom sklearn.datasets import make_classification\n\nfrom yellowbrick.features import RFECV\n\nsns.set(font_scale=3)\n\ncv = StratifiedKFold(5)\noz = RFECV(RandomForestClassifier(), cv=cv, scoring='f1')\n\noz.fit(X, y)\noz.poof()",
"_____no_output_____"
],
[
"# Ridge \n# Create a new figure \n#mpl.rcParams['axes.prop_cycle'] = cycler('color', ['red']) \nfrom yellowbrick.features.importances import FeatureImportances\nfrom sklearn.linear_model import (LogisticRegression, LogisticRegressionCV, RidgeClassifier, Ridge, Lasso, ElasticNet)\nfig = plt.gcf() \nfig.set_size_inches(10,10) \nax = plt.subplot(311) \nlabels = features \nviz = FeatureImportances(Ridge(alpha=0.5), ax=ax, labels=labels, relative=False) \nax.spines['right'].set_visible(False) \nax.spines['top'].set_visible(False) \nax.grid(False) # Fit and display \nviz.fit(X, y) \nviz.poof()",
"_____no_output_____"
],
[
"estimator = RandomForestClassifier(class_weight='balanced')\ny_pred_proba = RandomForestClassifier(X_test) \n#y_pred_proba[:5] \ndef plot_roc_curve(y_test, y_pred_proba): \n fpr, tpr, thresholds = roc_curve(y_test, y_pred_proba[:, 1]) \n roc_auc = auc(fpr, tpr) \n plt.plot(fpr, tpr, label='ROC curve (area = %0.3f)' % roc_auc, color='darkblue') \n plt.plot([0, 1], [0, 1], 'k--') # random predictions curve \n plt.xlim([0.0, 1.0]) \n plt.ylim([0.0, 1.0]) \n plt.xlabel('False Positive Rate or (1 - Specifity)') \n plt.ylabel('True Positive Rate or (Sensitivity)') \n plt.title('Receiver Operating Characteristic') \n plt.legend(loc=\"lower right\") \n plt.grid(False) \n\nplot_roc_curve(y_test, y_pred_proba)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acdcc1765b7374cd36535e0992a6d0816bc1f44
| 48,866 |
ipynb
|
Jupyter Notebook
|
Project 1 - Part B Submission Files/Project 1 - Part B - L18-1827.ipynb
|
taimurzahid/FAST-Deep-Learning
|
cfb2417f0bed43838e5c2401105f7f03552c6734
|
[
"MIT"
] | null | null | null |
Project 1 - Part B Submission Files/Project 1 - Part B - L18-1827.ipynb
|
taimurzahid/FAST-Deep-Learning
|
cfb2417f0bed43838e5c2401105f7f03552c6734
|
[
"MIT"
] | null | null | null |
Project 1 - Part B Submission Files/Project 1 - Part B - L18-1827.ipynb
|
taimurzahid/FAST-Deep-Learning
|
cfb2417f0bed43838e5c2401105f7f03552c6734
|
[
"MIT"
] | null | null | null | 89.009107 | 23,888 | 0.73149 |
[
[
[
"from __future__ import print_function\nimport keras\nfrom keras.datasets import mnist\nfrom keras.models import Sequential\nfrom keras.layers import Dense, Dropout, Flatten\nfrom keras.layers import Conv2D, MaxPooling2D\nfrom keras import backend as K\nimport matplotlib.pyplot as plt\nfrom keras.callbacks import TensorBoard\nimport matplotlib\nimport matplotlib.pyplot as plt\nimport numpy as np\nfrom time import time\n",
"_____no_output_____"
],
[
"batch_size = 128\nnum_classes = 10\nepochs = 10\n# input image dimensions\nimg_rows, img_cols = 28, 28",
"_____no_output_____"
],
[
"# the data, split between train and test sets\n(x_train, y_train), (x_test, y_test) = mnist.load_data()",
"Downloading data from https://s3.amazonaws.com/img-datasets/mnist.npz\n11493376/11490434 [==============================] - 1s 0us/step\n"
],
[
"# the data, split between train and test sets\n(x_train, y_train), (x_test, y_test) = mnist.load_data()\n\nif K.image_data_format() == 'channels_first':\n x_train = x_train.reshape(x_train.shape[0], 1, img_rows, img_cols)\n x_test = x_test.reshape(x_test.shape[0], 1, img_rows, img_cols)\n input_shape = (1, img_rows, img_cols)\nelse:\n x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)\n x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)\n input_shape = (img_rows, img_cols, 1)\n\nx_train = x_train.astype('float32')\nx_test = x_test.astype('float32')\nx_train /= 255\nx_test /= 255\nprint('x_train shape:', x_train.shape)\nprint(x_train.shape[0], 'train samples')\nprint(x_test.shape[0], 'test samples')",
"x_train shape: (60000, 28, 28, 1)\n60000 train samples\n10000 test samples\n"
],
[
"# convert class vectors to binary class matrices\ny_train = keras.utils.to_categorical(y_train, num_classes)\ny_test = keras.utils.to_categorical(y_test, num_classes)",
"_____no_output_____"
],
[
"#tbCallBack = keras.callbacks.TensorBoard(log_dir='./Graph', histogram_freq=2, write_graph=True, write_images=True)\ncallback=[keras.callbacks.TensorBoard(log_dir=\"/tmp/mnist/2\".format(time()), histogram_freq=1, write_graph=True, write_images=True)]\n\nmodel = Sequential()\nmodel.add(Conv2D(64, kernel_size=(3, 3),\n activation='relu',\n input_shape=input_shape))\nmodel.add(Conv2D(64, (3, 3), activation='relu'))\nmodel.add(Conv2D(128, (3, 3), activation='relu'))\nmodel.add(MaxPooling2D(pool_size=(2, 2)))\nmodel.add(MaxPooling2D(pool_size=(2, 2)))\nmodel.add(MaxPooling2D(pool_size=(2, 2)))\nmodel.add(Flatten())\nmodel.add(Dense(128, activation='relu'))\nmodel.add(Dropout(0.25))\nmodel.add(Dense(num_classes, activation='softmax'))\n\nmodel.compile(loss=keras.losses.categorical_crossentropy,\n optimizer=keras.optimizers.Adadelta(),\n metrics=['accuracy'])\n\nX = model.fit(x_train, y_train,\n batch_size=batch_size,\n epochs=epochs,\n verbose=1,\n validation_data=(x_test, y_test),\n callbacks=callback)\n\nplt.plot(X.history['loss'])\nplt.plot(X.history['val_loss'])\nplt.title('model loss')\nplt.ylabel('loss')\nplt.xlabel('epoch')\nplt.legend(['train', 'validation'], loc='upper left')\nplt.show()\n\n\nweights, biases = model.layers[0].get_weights()\n\nprint(\"Weights\\n\")\nprint(weights)\n\nscore = model.evaluate(x_test, y_test, verbose=0)\nprint('Test loss:', score[0])\nprint('Test accuracy:', score[1])",
"Train on 60000 samples, validate on 10000 samples\nEpoch 1/10\n60000/60000 [==============================] - 21s 351us/step - loss: 0.3508 - acc: 0.8880 - val_loss: 0.1011 - val_acc: 0.9665\nEpoch 2/10\n60000/60000 [==============================] - 18s 300us/step - loss: 0.0878 - acc: 0.9720 - val_loss: 0.0541 - val_acc: 0.9840\nEpoch 3/10\n60000/60000 [==============================] - 18s 299us/step - loss: 0.0623 - acc: 0.9811 - val_loss: 0.0555 - val_acc: 0.9826\nEpoch 4/10\n60000/60000 [==============================] - 18s 299us/step - loss: 0.0516 - acc: 0.9847 - val_loss: 0.0398 - val_acc: 0.9882\nEpoch 5/10\n60000/60000 [==============================] - 18s 297us/step - loss: 0.0434 - acc: 0.9871 - val_loss: 0.0452 - val_acc: 0.9862\nEpoch 6/10\n60000/60000 [==============================] - 18s 299us/step - loss: 0.0370 - acc: 0.9883 - val_loss: 0.0362 - val_acc: 0.9893\nEpoch 7/10\n60000/60000 [==============================] - 18s 300us/step - loss: 0.0329 - acc: 0.9901 - val_loss: 0.0383 - val_acc: 0.9885\nEpoch 8/10\n60000/60000 [==============================] - 18s 300us/step - loss: 0.0296 - acc: 0.9907 - val_loss: 0.0339 - val_acc: 0.9886\nEpoch 9/10\n60000/60000 [==============================] - 18s 297us/step - loss: 0.0260 - acc: 0.9918 - val_loss: 0.0324 - val_acc: 0.9900\nEpoch 10/10\n60000/60000 [==============================] - 18s 299us/step - loss: 0.0245 - acc: 0.9925 - val_loss: 0.0408 - val_acc: 0.9888\n"
],
[
"LOG_DIR = '/tmp/mnist/2'\nget_ipython().system_raw(\n 'tensorboard --logdir {} --host 0.0.0.0 --port 6006 &'\n .format(LOG_DIR)\n)\n# Install\n! npm install -g localtunnel\n\n# Tunnel port 6006 (TensorBoard assumed running)\nget_ipython().system_raw('lt --port 6006 >> url.txt 2>&1 &')\n\n# Get url\n! cat url.txt",
"\u001b[K\u001b[?25h/tools/node/bin/lt -> /tools/node/lib/node_modules/localtunnel/bin/client\n\u001b[K\u001b[?25h+ [email protected]\nadded 54 packages from 31 contributors in 4.984s\nyour url is: https://young-dog-46.localtunnel.me\n"
],
[
"weights, biases = model.layers[0].get_weights()\nprint(biases)",
"[ 5.2305091e-02 4.2329561e-02 2.2895745e-04 5.2398529e-02\n -1.6769962e-03 3.7942608e-03 5.2037787e-02 3.5299581e-02\n 7.9607598e-02 -8.6277332e-03 -4.2480580e-03 -2.0624446e-03\n 4.5489490e-02 7.3770866e-02 -1.6724458e-03 4.3317840e-02\n 9.3861036e-02 1.4329603e-01 9.3229800e-02 -1.9087589e-03\n 2.5267934e-03 7.8527488e-02 2.6613273e-02 6.3304435e-03\n 9.8840922e-02 5.2497875e-02 -1.0301483e-02 3.7837710e-04\n 2.7502151e-02 3.6927410e-03 5.5295322e-03 -2.1472098e-03\n 1.2749669e-01 -5.4282760e-03 -5.0160423e-05 2.5811609e-02\n 6.6094242e-02 1.5407995e-02 -6.3875001e-03 -5.9412839e-03\n -5.2355146e-03 9.5629036e-02 1.6479639e-03 6.5808021e-03\n 3.2558650e-02 5.9396092e-02 2.5926530e-03 4.3830305e-02\n -6.6156511e-04 4.5381743e-02 7.1374851e-04 3.6877085e-02\n -2.3744851e-03 -3.5023768e-03 -4.4187387e-03 1.6534025e-03\n 2.8542482e-04 2.2166800e-03 5.5086050e-02 2.9888425e-02\n 7.4517909e-03 3.2116991e-02 -1.0055948e-02 -2.2171314e-03]\n"
],
[
"def plot_weights(layers,x,y):\n weights, biases = model.layers[0].get_weights()\n \n fig = plt.figure(figsize=(106,108), dpi=80)\n \n for j in range(len(weights)):\n ax = fig.add_subplot(y,x,j+1)\n ax.matshow(weights[j][0], cmap = matplotlib.cm.binary)\n \n plt.xticks(np.array([]))\n plt.yticks(np.array([]))\n plt.rcParams['figure.figsize'] = 4.5, 4.\n plt.tight_layout(pad=0.4, w_pad=0.5, h_pad=1.0)\n return plt\n \n",
"_____no_output_____"
],
[
"plot_weights(model.layers[0],8,4)",
"_____no_output_____"
],
[
"",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acdd6a6b33c800acda55d4bb11e42e6debd02e2
| 12,179 |
ipynb
|
Jupyter Notebook
|
examples/cli.ipynb
|
astrogewgaw/priwo
|
cc3ef488b6e9a36fb5d633d6c2c6135274b2c7b7
|
[
"MIT"
] | 7 |
2021-04-24T12:16:11.000Z
|
2021-12-31T14:46:35.000Z
|
examples/cli.ipynb
|
astrogewgaw/priwo
|
cc3ef488b6e9a36fb5d633d6c2c6135274b2c7b7
|
[
"MIT"
] | null | null | null |
examples/cli.ipynb
|
astrogewgaw/priwo
|
cc3ef488b6e9a36fb5d633d6c2c6135274b2c7b7
|
[
"MIT"
] | null | null | null | 60.895 | 402 | 0.445849 |
[
[
[
"# priwo on the command line\n\nYou can also use priwo on the command line. When you installed [**priwo**](https://github.com/astrogewgaw/priwo), it added a `priwo` command to your `PATH`. To check it out, just do (note that the exclamation mark just tells the notebook to run a shell command; you should try it out without it):",
"_____no_output_____"
]
],
[
[
"!priwo",
"Usage: priwo [OPTIONS] COMMAND [ARGS]...\n\n The priwo command line tool.\n\nOptions:\n --version Show the version and exit.\n -h, --help Show this message and exit.\n\nCommands:\n available Show the available formats in priwo.\n max Show the maximum size set in priwo.\n peek Take a peek into any pulsar data file.\n"
]
],
[
[
"This just displays some a help message. We have already met two of the three commands currently supported by [**priwo**](https://github.com/astrogewgaw/priwo). But we still have to check out `peek`. As the name suggests, it lets you take a peek into the data and metadata in any pulsar data file, provided the format is supported by [**priwo**](https://github.com/astrogewgaw/priwo). For example:",
"_____no_output_____"
]
],
[
[
"!priwo peek data/Lband_DM62.00.dat",
"╭────────────────── Lband_DM62.00.dat ───────────────────╮\n│ \u001b[1mbsname\u001b[0m \u001b[3mtempdir/Lband_DM62.00\u001b[0m │\n│ \u001b[1mtelescope\u001b[0m \u001b[3mGBT\u001b[0m │\n│ \u001b[1minstrument\u001b[0m \u001b[3munset\u001b[0m │\n│ \u001b[1mobject\u001b[0m \u001b[3mMystery_PSR\u001b[0m │\n│ \u001b[1mrastr\u001b[0m \u001b[3m16:43:38.1000\u001b[0m │\n│ \u001b[1mdecstr\u001b[0m \u001b[3m-12:24:58.7000\u001b[0m │\n│ \u001b[1mobserver\u001b[0m \u001b[3munset\u001b[0m │\n│ \u001b[1mmjd\u001b[0m \u001b[1;91m53010.48095599331\u001b[0m │\n│ \u001b[1mbary\u001b[0m \u001b[1;92mTrue\u001b[0m │\n│ \u001b[1mnsamp\u001b[0m \u001b[1;91m134400\u001b[0m │\n│ \u001b[1mtsamp\u001b[0m \u001b[1;91m0.000288\u001b[0m │\n│ \u001b[1mbreaks\u001b[0m \u001b[1;92mTrue\u001b[0m │\n│ \u001b[1memband\u001b[0m \u001b[3mRadio\u001b[0m │\n│ \u001b[1mbdiam\u001b[0m \u001b[1;91m530.0\u001b[0m │\n│ \u001b[1mdm\u001b[0m \u001b[1;91m62.0\u001b[0m │\n│ \u001b[1mcfreq\u001b[0m \u001b[1;91m1352.5\u001b[0m │\n│ \u001b[1mbw\u001b[0m \u001b[1;91m96.0\u001b[0m │\n│ \u001b[1mnchan\u001b[0m \u001b[1;91m96\u001b[0m │\n│ \u001b[1mchanwid\u001b[0m \u001b[1;91m1.0\u001b[0m │\n│ \u001b[1manalyst\u001b[0m \u001b[3mmaximus\u001b[0m │\n│ \u001b[1mnotes\u001b[0m \u001b[3mProject ID unset, Date: 2004-01-06T11:38:09.\u001b[0m │\n│ \u001b[3m2 polns were summed. Samples have 4 bits.\u001b[0m │\n│ \u001b[1monoffs\u001b[0m \u001b[3m(0, 131406)\u001b[0m │\n│ \u001b[3m(134399, 134399)\u001b[0m │\n│ \u001b[1mdata\u001b[0m Array of size \u001b[1m531000\u001b[0m and dimensions \u001b[1m531000\u001b[0m │\n╰────────────────────────────────────────────────────────╯\n"
]
],
[
[
"Note that `peek` doesn't show you the arrays of data, but only tells you the size and the dimensions of the arrays. It does this because it can often not display the entire array to you on the terminal. For example, take a `*.pfd` file:",
"_____no_output_____"
]
],
[
[
"!priwo peek data/Lband_DM62.00_ACCEL_Cand_1.pfd",
"╭──────────── Lband_DM62.00_ACCEL_Cand_1.pfd ─────────────╮\n│ \u001b[1mnumdms\u001b[0m \u001b[1;91m0\u001b[0m │\n│ \u001b[1mnumperiods\u001b[0m \u001b[1;91m257\u001b[0m │\n│ \u001b[1mnumpdots\u001b[0m \u001b[1;91m257\u001b[0m │\n│ \u001b[1mnsub\u001b[0m \u001b[1;91m1\u001b[0m │\n│ \u001b[1mnpart\u001b[0m \u001b[1;91m64\u001b[0m │\n│ \u001b[1mproflen\u001b[0m \u001b[1;91m64\u001b[0m │\n│ \u001b[1mnumchan\u001b[0m \u001b[1;91m1\u001b[0m │\n│ \u001b[1mpstep\u001b[0m \u001b[1;91m2\u001b[0m │\n│ \u001b[1mpdstep\u001b[0m \u001b[1;91m4\u001b[0m │\n│ \u001b[1mdmstep\u001b[0m \u001b[1;91m2\u001b[0m │\n│ \u001b[1mndmfact\u001b[0m \u001b[1;91m3\u001b[0m │\n│ \u001b[1mnpfact\u001b[0m \u001b[1;91m2\u001b[0m │\n│ \u001b[1mfilename\u001b[0m \u001b[3m../Lband_DM62.00.dat\u001b[0m │\n│ \u001b[1mcandname\u001b[0m \u001b[3mACCEL_Cand_1\u001b[0m │\n│ \u001b[1mtelescope\u001b[0m \u001b[3mGBT\u001b[0m │\n│ \u001b[1mpgdev\u001b[0m \u001b[3m../Lband_DM62.00_ACCEL_Cand_1.pfd.ps/CPS\u001b[0m │\n│ \u001b[1mrastr\u001b[0m \u001b[3m16:43:38.1000\u001b[0m │\n│ \u001b[1mdecstr\u001b[0m \u001b[3m-12:24:58.7000\u001b[0m │\n│ \u001b[1mtsamp\u001b[0m \u001b[1;91m0.000288\u001b[0m │\n│ \u001b[1mstartT\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1mendT\u001b[0m \u001b[1;91m1.0\u001b[0m │\n│ \u001b[1mtepoch\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1mbepoch\u001b[0m \u001b[1;91m53010.48095599331\u001b[0m │\n│ \u001b[1mavgoverc\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1mlofreq\u001b[0m \u001b[1;91m1352.5\u001b[0m │\n│ \u001b[1mchanwidth\u001b[0m \u001b[1;91m1.0\u001b[0m │\n│ \u001b[1mbestdm\u001b[0m \u001b[1;91m62.0\u001b[0m │\n│ \u001b[1mtopopow\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1m_t\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1mtopop1\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1mtopop2\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1mtopop3\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1mbarypow\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1m_b\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1mbaryp1\u001b[0m \u001b[1;91m9.275145008336503\u001b[0m │\n│ \u001b[1mbaryp2\u001b[0m \u001b[1;91m-0.0003640671516059178\u001b[0m │\n│ \u001b[1mbaryp3\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1mfoldpow\u001b[0m \u001b[1;91m1.0\u001b[0m │\n│ \u001b[1m_f\u001b[0m \u001b[1;91m4.55674234629144e-41\u001b[0m │\n│ \u001b[1mfoldp1\u001b[0m \u001b[1;91m0.12271618716931217\u001b[0m │\n│ \u001b[1mfoldp2\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1mfoldp3\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1morbp\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1morbe\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1morbx\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1morbw\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1morbt\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1morbpd\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1morbwd\u001b[0m \u001b[1;91m0.0\u001b[0m │\n│ \u001b[1mdms\u001b[0m Array of size \u001b[1m0\u001b[0m and dimensions \u001b[1m0\u001b[0m │\n│ \u001b[1mperiods\u001b[0m Array of size \u001b[1m257\u001b[0m and dimensions \u001b[1m257\u001b[0m │\n│ \u001b[1mpdots\u001b[0m Array of size \u001b[1m257\u001b[0m and dimensions \u001b[1m257\u001b[0m │\n│ \u001b[1mprofs\u001b[0m Array of size \u001b[1m4096\u001b[0m and dimensions \u001b[1m64 x 1 x 64\u001b[0m │\n│ \u001b[1mstats\u001b[0m Array of size \u001b[1m448\u001b[0m and dimensions \u001b[1m64 x 1 x 7\u001b[0m │\n│ \u001b[1mnumprofs\u001b[0m \u001b[1;91m64\u001b[0m │\n╰─────────────────────────────────────────────────────────╯\n"
]
],
[
[
"As you can see, there are a lot of arrays, and they are huge (atleast for the terminal). `peek` helps you get a first look at pulsar data of all kinds, before you actually start analysing stuff.",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
4acdd6c8d4e0b5b6926a01785e2bc477bbad3fd8
| 104,232 |
ipynb
|
Jupyter Notebook
|
test/forms/DKRZ_CDP/DKRZ_CDP_ki_123.ipynb
|
IS-ENES-Data/submission_forms
|
d8fbe7437f604cd0153ef18b9e848d572b188155
|
[
"Apache-2.0"
] | null | null | null |
test/forms/DKRZ_CDP/DKRZ_CDP_ki_123.ipynb
|
IS-ENES-Data/submission_forms
|
d8fbe7437f604cd0153ef18b9e848d572b188155
|
[
"Apache-2.0"
] | 6 |
2016-07-25T11:16:29.000Z
|
2018-05-31T13:58:17.000Z
|
test/forms/DKRZ_CDP/DKRZ_CDP_ki_123.ipynb
|
IS-ENES-Data/submission_forms
|
d8fbe7437f604cd0153ef18b9e848d572b188155
|
[
"Apache-2.0"
] | null | null | null | 207.633466 | 68,014 | 0.795936 |
[
[
[
"# Generic DKRZ national archive ingest form \n\n\nThis form is intended to request data to be made locally available in the DKRZ nationl data archive besides the Data which is ingested as part of the CMIP6 replication. For replication requests a separate form is available. \n\nPlease provide information on the following aspects of your data ingest request:\n\n* scientific context of data\n* specific data access rights\n* technical details, like\n * amount of data\n * source of data",
"_____no_output_____"
]
],
[
[
"from dkrz_forms import form_widgets\nform_widgets.show_status('form-submission')",
"_____no_output_____"
]
],
[
[
"## Please provide information to unlock your form\n\n- last name\n- password",
"_____no_output_____"
]
],
[
[
"from dkrz_forms import form_handler, form_widgets\n\n#please provide your last name - replacing ... below\nMY_LAST_NAME = \"ki\" \n\n\nform_info = form_widgets.check_pwd(MY_LAST_NAME)\n\nsf = form_handler.init_form(form_info)\nform = sf.sub.entity_out.form_info",
"Enter your form key: ········\n---- Your Name: ki ki\n---- Your email: ki\n---- Name of this submission form: DKRZ_CDP_ki_123\nForm Handler: Initialized form for project: DKRZ_CDP\nDKRZ_CDP\n/home/stephan/tmp/Repos/form_repo/DKRZ_CDP\nDKRZ_CDP_ki_123.ipynb\n/home/stephan/Repos/ENES-EUDAT/submission_forms/test/forms/DKRZ_CDP/DKRZ_CDP_ki_123.ipynb\n/home/stephan/tmp/Repos/form_repo/DKRZ_CDP/DKRZ_CDP_ki_123.ipynb\nsubmission form intitialized: sf\n(For the curious: the sf object is used in the following to store and manage all your information)\n"
],
[
"import pprint\nfrom dkrz_forms import form_handler\npprint.pprint(form_handler.form_to_dict(sf))\n\n",
"{'__doc__': '\\n Form object for data replication related information\\n \\n Workflow steps related sub-forms (filled by data manager):\\n - sub: data submission form\\n - rev: data review_form\\n - ing: data ingest form\\n - qua: data quality assurance form\\n - pub: data publication form\\n \\n Each workfow step form is structured according to\\n - entity_in : input information for this step\\n - entity_out: output information for this step\\n - agent: information related to responsible party for this step\\n - activity: information related the workflow step execution\\n \\n End user provided form information is stored in:\\n \\n _this_form_object.sub.entity_out.form \\n \\n The the documentation of the sub.entity_out subform for \\n the end-user filled information entities \\n ',\n 'form_dir': '/home/stephan/Repos/ENES-EUDAT/submission_forms/test/forms/DKRZ_CDP',\n 'ing': {'__doc__': '\\n Attributes characterizing the data ingest workflow step:\\n - entity_in : data review report\\n - entity_out : data ingest report\\n - agent : person or tool related ingest step information \\n - activity : information on the ingest process\\n ',\n 'activity': {'__doc__': '\\n Attributes characterizing the data ingest activity:\\n - status: status information\\n - timestamp_started\\n - timestamp_finished : data ingest timing information\\n - comment : free text comment\\n - ticket_id: related RT ticket number\\n - ingest_report: dictionary with ingest related information (tbd.)\\n ',\n 'comment': '',\n 'ingest_report': {},\n 'status': '',\n 'ticket_id': 0,\n 'timestamp_finished': '',\n 'timestamp_started': ''},\n 'agent': {'__doc__': ' \\n Attributes characterizing the person or tool managing the data ingest:\\n - responsible_person: \\n ',\n 'responsible_person': ''},\n 'entity_in': {'__doc__': \"\\n Attributes characterizing the review results:\\n - data: time stamp of last results\\n - tag: git tag for repo conatining report information\\n - repo: (gitlab) repo containing report information,\\n - comment: free text comment for this review,\\n - status : review status information: ok, undef, uncomplete, error,\\n - dict: report details in dictionary,\\n - dict_type': predefined value: review_report \\n \",\n 'comment': '',\n 'date': '',\n 'dict': '',\n 'dict_type': 'review_report',\n 'repo': '',\n 'status': '',\n 'tag': ''},\n 'entity_out': {'__doc__': \"\\n Attributes characterizing the ingest report summary:\\n - date: 'last modification date,\\n - tag : '(git) tag for repo containing report information,\\n - repo: '(gitlab) repo containing report information,\\n - comment : 'free text comment for this review,\\n - status : 'data ingest status information: ok, undef, uncomplete, error,\\n - dict: 'report details in dictionary,\\n - dict_type: 'predefined value: ingest_report \\n \",\n 'comment': '',\n 'date': '',\n 'dict': '',\n 'dict_type': 'ingest_report',\n 'repo': '',\n 'status': '',\n 'tag': ''}},\n 'project': 'DKRZ_CDP',\n 'pub': {'__doc__': '\\n Attributes characterizing the data publication workflow step:\\n - entity_in: \\n - entity_out:\\n - agent:\\n - activity:\\n ',\n 'activity': {'__doc__': ' Attributes characterizing the data publication activity:\\n - status: status information\\n - timestamp_started\\n - timestamp_finished : data ingest timing information\\n - comment : free text comment\\n - ticket_id: related RT ticket number\\n - follow_up_ticket: in case new data has to be provided\\n ',\n 'comment': '',\n 'follow_up_ticket': '',\n 'status': '',\n 'ticket_id': '',\n 'timestamp_finished': '',\n 'timestamp_started': ''},\n 'agent': {'__doc__': '\\n Attributes characterizing the persons performing the data publication:\\n - responsible_person: person name\\n - publication_tool: string characterizing the publication tool\\n ',\n 'publication_tool': '',\n 'responsible_person': ''},\n 'entity_in': {'__doc__': \"\\n Attributes characterizing the quality report summary:\\n - date: 'last modification date,\\n - tag : '(git) tag for repo containing report information,\\n - repo: '(gitlab) repo containing report information,\\n - comment : 'free text comment for this review,\\n - status : 'data ingest status information: ok, undef, uncomplete, error,\\n - dict: 'report details in dictionary,\\n - dict_type: 'predefined value: ingest_report \\n \",\n 'comment': '',\n 'date': '',\n 'dict': '',\n 'dict_type': 'qua_report',\n 'repo': '',\n 'status': '',\n 'tag': ''},\n 'entity_out': {'__doc__': \" \\n Attributes characterizing the data publication report\\n - date: last modification date,\\n - tag : (git) tag for repo containing report information,\\n - repo: (gitlab) repo containing report information,\\n - comment : free text comment for this review,\\n - status : data ingest status information: ok, undef, uncomplete, error,\\n - dict: 'report details in dictionary,\\n - dict_type: predefined value: publication_report, \\n \",\n 'comment': '',\n 'date': '',\n 'dict': '',\n 'dict_type': 'publication_report',\n 'facet_string': '# e.g. project=A&model=B& ....',\n 'repo': '',\n 'status': '',\n 'tag': ''}},\n 'qua': {'__doc__': '\\n Attributes characterizing the data quality assurance step:\\n - entity_in: data ingest report\\n - entity_out: data quality assurance report\\n - agent: person and tool responsible for qua checking\\n - activity: info on qua checking process\\n ',\n 'activity': {'__doc__': '\\n Attributes characterizing the data quality assurance activity:\\n - status: status information\\n - timestamp_started\\n - timestamp_finished : data ingest timing information\\n - comment : free text comment\\n - ticket_id: related RT ticket number\\n - follow_up_ticket: in case new data has to be provided\\n - quality_report: dictionary with quality related information (tbd.)\\n ',\n 'comment': '',\n 'follow_up_ticket': '',\n 'status': '',\n 'ticket_id': '',\n 'timestamp_finished': '',\n 'timestamp_started': ''},\n 'agent': {'__doc__': ' \\n Attributes characterizing the person or tool managing the data ingest:\\n - responsible_person: \\n ',\n 'responsible_person': ''},\n 'entity_in': {'__doc__': \"\\n Attributes characterizing the ingest report summary:\\n - date: 'last modification date,\\n - tag : '(git) tag for repo containing report information,\\n - repo: '(gitlab) repo containing report information,\\n - comment : 'free text comment for this review,\\n - status : 'data ingest status information: ok, undef, uncomplete, error,\\n - dict: 'report details in dictionary,\\n - dict_type: 'predefined value: ingest_report \\n \",\n 'comment': '',\n 'date': '',\n 'dict': '',\n 'dict_type': 'ingest_report',\n 'repo': '',\n 'status': '',\n 'tag': ''},\n 'entity_out': {'__doc__': \"\\n Attributes characterizing the quality report summary:\\n - date: 'last modification date,\\n - tag : '(git) tag for repo containing report information,\\n - repo: '(gitlab) repo containing report information,\\n - comment : 'free text comment for this review,\\n - status : 'data ingest status information: ok, undef, uncomplete, error,\\n - dict: 'report details in dictionary,\\n - dict_type: 'predefined value: ingest_report \\n \",\n 'comment': '',\n 'date': '',\n 'dict': '',\n 'dict_type': 'qua_report',\n 'repo': '',\n 'status': '',\n 'tag': ''}},\n 'rev': {'__doc__': \" \\n Attributes characterizing the data submission review step:\\n - entity_in : submission form\\n - entity_out: adapted submission form\\n - agent: person or tool related information\\n - activity': submission activity related information\\n \",\n 'activity': {'__doc__': '\\n Attributes characterizing the form review activity:\\n - ticket_url: assigned RT Ticket\\n - ticket_id: RT Ticket id \\n - review_comment: free text comment \\n - review_status: progress status of review\\n - review_report: dictionary with review results and issues\\n ',\n 'review_comment': '',\n 'review_status': '',\n 'ticket_id': 0,\n 'ticket_url': ''},\n 'agent': {'__doc__': ' \\n Attributes characterizing the person or tool checking the form:\\n - responsible_person: \\n ',\n 'responsible_person': ''},\n 'entity_in': {'__doc__': \" \\n Attributes characterizing the data submission workflow step:\\n - entity_in : Input (form template and init info)\\n - entity_out: Output (submission form information)\\n - agent: person or tool related information\\n - activity': submission activity related information\\n \",\n 'activity': {'__doc__': '\\n Attributes characterizing the form submission activity:\\n \\n - submission_comment : free text comment\\n - submission_method : How the submission was generated and submitted to DKRZ: email or DKRZ form server based \\n ',\n 'pwd': ' password to access form ',\n 'submission_comment': '',\n 'submission_method': ''},\n 'agent': {'__doc__': 'Attributes characterizing the person responsible for form completion and submission:\\n \\n - last_name: Last name of the person responsible for the submission form content\\n - first_name: Corresponding first name\\n - email: Valid user email address: all follow up activities will use this email to contact end user\\n - key_word : user provided key word to remember and separate submission\\n ',\n 'email': '',\n 'first_name': '',\n 'key_word': '',\n 'last_name': ''},\n 'entity_in': {'__doc__': '\\n Attributes defining the form template used:\\n \\n - source_path: path to the form template used (jupyter notebook) \\n - form_template_version: version string for the form template used\\n - tag: git tag of repo containing templates (=source code repo in most cases)\\n ',\n 'form_template_version': '',\n 'source_path': '',\n 'tag': ''},\n 'entity_out': {'__doc__': \"\\n Attributes characterizing the form submission process and context:\\n - form: Form object for all end user provided information\\n - form_name: consistent prefix for the form name (postfix=.ipynb and .json)\\n - form_repo: git repo where forms are stored (before submission)\\n - form_json, form_repo_path: full paths to json and ipynb representations\\n - form_dir: directory where form are served to the notebook interface\\n - status: status information\\n - checks_done: form consistency checks done\\n - tag: 'git tag of submission form in submission form repo',\\n - repo: '(gitlab) repo where the tag relates to','\\n \\n \",\n 'checks_done': '',\n 'form': '',\n 'form_dir': '',\n 'form_info': {},\n 'form_json': '',\n 'form_name': '',\n 'form_path': '',\n 'form_repo': '',\n 'form_repo_path': '',\n 'repo': '',\n 'status': '',\n 'tag': ''}},\n 'entity_out': {'__doc__': \"\\n Attributes characterizing the review results:\\n - data: time stamp of last results\\n - tag: git tag for repo conatining report information\\n - repo: (gitlab) repo containing report information,\\n - comment: free text comment for this review,\\n - status : review status information: ok, undef, uncomplete, error,\\n - dict: report details in dictionary,\\n - dict_type': predefined value: review_report \\n \",\n 'comment': '',\n 'date': '',\n 'dict': '',\n 'dict_type': 'review_report',\n 'repo': '',\n 'status': '',\n 'tag': ''}},\n 'sub': {'__doc__': \" \\n Attributes characterizing the data submission workflow step:\\n - entity_in : Input (form template and init info)\\n - entity_out: Output (submission form information)\\n - agent: person or tool related information\\n - activity': submission activity related information\\n \",\n 'activity': {'__doc__': '\\n Attributes characterizing the form submission activity:\\n \\n - submission_comment : free text comment\\n - submission_method : How the submission was generated and submitted to DKRZ: email or DKRZ form server based \\n ',\n 'keyword': u'123',\n 'pwd': '4VY08A',\n 'submission_comment': '',\n 'submission_method': ''},\n 'agent': {'__doc__': 'Attributes characterizing the person responsible for form completion and submission:\\n \\n - last_name: Last name of the person responsible for the submission form content\\n - first_name: Corresponding first name\\n - email: Valid user email address: all follow up activities will use this email to contact end user\\n - key_word : user provided key word to remember and separate submission\\n ',\n 'email': u'kim',\n 'first_name': u'ki',\n 'key_word': '',\n 'last_name': u'ki'},\n 'entity_in': {'__doc__': '\\n Attributes defining the form template used:\\n \\n - source_path: path to the form template used (jupyter notebook) \\n - form_template_version: version string for the form template used\\n - tag: git tag of repo containing templates (=source code repo in most cases)\\n ',\n 'form_json': u'/home/stephan/Repos/ENES-EUDAT/submission_forms/test/forms/DKRZ_CDP/DKRZ_CDP_ki_123.json',\n 'form_path': u'/home/stephan/Repos/ENES-EUDAT/submission_forms/test/forms/DKRZ_CDP/DKRZ_CDP_ki_123.ipynb',\n 'form_template_version': '',\n 'source_path': '',\n 'tag': ''},\n 'entity_out': {'__doc__': \"\\n Attributes characterizing the form submission process and context:\\n - form: Form object for all end user provided information\\n - form_name: consistent prefix for the form name (postfix=.ipynb and .json)\\n - form_repo: git repo where forms are stored (before submission)\\n - form_json, form_repo_path: full paths to json and ipynb representations\\n - form_dir: directory where form are served to the notebook interface\\n - status: status information\\n - checks_done: form consistency checks done\\n - tag: 'git tag of submission form in submission form repo',\\n - repo: '(gitlab) repo where the tag relates to','\\n \\n \",\n 'checks_done': '',\n 'form': {'__doc__': '\\n Form object for data replication related information\\n \\n Workflow steps related sub-forms (filled by data manager):\\n - sub: data submission form\\n - rev: data review_form\\n - ing: data ingest form\\n - qua: data quality assurance form\\n - pub: data publication form\\n \\n Each workfow step form is structured according to\\n - entity_in : input information for this step\\n - entity_out: output information for this step\\n - agent: information related to responsible party for this step\\n - activity: information related the workflow step execution\\n \\n End user provided form information is stored in:\\n \\n _this_form_object.sub.entity_out.form \\n \\n The the documentation of the sub.entity_out subform for \\n the end-user filled information entities \\n ',\n 'project': 'CMIP6_CDP',\n 'workflow': [('sub', 'data_submission'),\n ('rev',\n 'data_submission_review'),\n ('ing', 'data_ingest'),\n ('qua',\n 'data_quality_assurance'),\n ('pub', 'data_publication')]},\n 'form_dir': '',\n 'form_info': {'access_group': '....',\n 'access_rights': '....',\n 'best_ingest_before': '....',\n 'context_comment': '....',\n 'data_path': '....',\n 'data_type': '....',\n 'directory_structure': '...',\n 'directory_structure_comment': '...',\n 'directory_structure_convention': '...',\n 'metadata_comment': '...',\n 'metadata_convention_name': '...',\n 'scientific_context': '...',\n 'terms_of_use': '....',\n 'usage': '....'},\n 'form_json': u'/home/stephan/tmp/Repos/form_repo/DKRZ_CDP/DKRZ_CDP_ki_123.json',\n 'form_name': u'DKRZ_CDP_ki_123',\n 'form_path': '',\n 'form_repo': '/home/stephan/tmp/Repos/form_repo/DKRZ_CDP',\n 'form_repo_path': u'/home/stephan/tmp/Repos/form_repo/DKRZ_CDP/DKRZ_CDP_ki_123.ipynb',\n 'pwd': '4VY08A',\n 'repo': '',\n 'status': '',\n 'tag': ''},\n 'id': '79ed688a-3280-11e7-b816-080027f178b4',\n 'submission_repo': '/home/stephan/tmp/Repos/submission_repo/DKRZ_CDP',\n 'timestamp': '2017-05-06 19:21:44.081274'},\n 'workflow': [('sub', 'data_submission'),\n ('rev', 'data_submission_review'),\n ('ing', 'data_ingest'),\n ('qua', 'data_quality_assurance'),\n ('pub', 'data_publication')]}\n"
]
],
[
[
"## Please provide the following information\n\nPlease provide some generic context information about the data, which should be availabe as part of the DKRZ CMIP Data Pool (CDP)",
"_____no_output_____"
]
],
[
[
"# (informal) type of data\nform.data_type = \"....\" # e.g. model data, observational data, .. \n# # free text describing scientific context of data\nform.scientific_context =\"...\" \n# free text describing the expected usage as part of the DKRZ CMIP Data pool\nform.usage = \"....\" \n# free text describing access rights (who is allowed to read the data)\nform.access_rights = \"....\"\n# generic terms of policy information\nform.terms_of_use = \"....\" # e.g. unrestricted, restricted\n# any additional comment on context\nform.access_group = \"....\"\nform.context_comment = \"....\"",
"_____no_output_____"
]
],
[
[
"## technical information concerning your request",
"_____no_output_____"
]
],
[
[
"# information on where the data is stored and can be accessed\n# e.g. file system path if on DKRZ storage, url etc. if on web accessible resources (cloud,thredds server,..)\nform.data_path = \"....\"\n\n# timing constraints, when the data ingest should be completed \n# (e.g. because the data source is only accessible in specific time frame) \nform.best_ingest_before = \"....\"\n\n# directory structure information, especially \nform.directory_structure = \"...\" # e.g. institute/experiment/file.nc\nform.directory_structure_convention = \"...\" # e.g. CMIP5, CMIP6, CORDEX, your_convention_name\nform.directory_structure_comment = \"...\" # free text, e.g. with link describing the directory structure convention you used\n\n# metadata information\nform.metadata_convention_name = \"...\" # e.g. CF1.6 etc. None if not applicable\nform.metadata_comment = \"...\" # information about metadata, e.g. links to metadata info etc. ",
"_____no_output_____"
]
],
[
[
"## Check your submission form\n\nPlease evaluate the following cell to check your submission form.\n\nIn case of errors, please go up to the corresponden information cells and update your information accordingly...",
"_____no_output_____"
]
],
[
[
"# to be completed .. ",
"_____no_output_____"
]
],
[
[
"# Save your form\n\nyour form will be stored (the form name consists of your last name plut your keyword)",
"_____no_output_____"
]
],
[
[
"form_handler.save_form(sf,\"..my comment..\") # edit my comment info ",
"\n\nForm Handler - save form status message:\n/home/stephan/Repos/ENES-EUDAT/submission_forms/test/forms/DKRZ_CDP/DKRZ_CDP_ki_123.ipynb\n/home/stephan/tmp/Repos/form_repo/DKRZ_CDP/DKRZ_CDP_ki_123.ipynb\n --- form stored in transfer format in: /home/stephan/tmp/Repos/form_repo/DKRZ_CDP/DKRZ_CDP_ki_123.json\n \n --- commit message:[master d29a38a] Form Handler: submission form for user ki saved using prefix DKRZ_CDP_ki_123 ## ..my comment..\n 2 files changed, 53 insertions(+), 7 deletions(-)\n"
]
],
[
[
"# officially submit your form\nthe form will be submitted to the DKRZ team to process\nyou also receive a confirmation email with a reference to your online form for future modifications ",
"_____no_output_____"
]
],
[
[
"form_handler.email_form_info(sf)\nform_handler.form_submission(sf)",
"This form is not hosted at DKRZ! Thus form information is stored locally on your computer \n\nHere is a summary of the generated and stored information:\n-- form for project: DKRZ_CDP\n-- form name: DKRZ_CDP_ki_123\n"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4acde16fcc439831004565b824a88da045d4d7ca
| 109,699 |
ipynb
|
Jupyter Notebook
|
japan_company_list/scrape_wikipedia.ipynb
|
ierika/public_notebooks
|
e096c0f077a29a48de8dc7cd234ac75eec5964ac
|
[
"MIT"
] | null | null | null |
japan_company_list/scrape_wikipedia.ipynb
|
ierika/public_notebooks
|
e096c0f077a29a48de8dc7cd234ac75eec5964ac
|
[
"MIT"
] | null | null | null |
japan_company_list/scrape_wikipedia.ipynb
|
ierika/public_notebooks
|
e096c0f077a29a48de8dc7cd234ac75eec5964ac
|
[
"MIT"
] | null | null | null | 260.567696 | 97,110 | 0.640325 |
[
[
[
"import pandas as pd\nimport requests\nfrom bs4 import BeautifulSoup\nfrom pathlib import Path",
"_____no_output_____"
],
[
"cwd = Path.cwd()\nhtml_directory = cwd / 'html'\n\nif not html_directory.exists():\n html_directory.mkdir()",
"_____no_output_____"
]
],
[
[
"### Get response and save to a file",
"_____no_output_____"
]
],
[
[
"# Get the HTML and cache it in local disk\nurl = 'https://en.wikipedia.org/wiki/list_of_companies_of_Japan'\noutput_file = html_directory / 'wikipedia_page.html'\n\nif not output_file.exists():\n html_content = requests.get(url).content\n html_content = html_content.decode()\n output_file.write_text(html_content, encoding='UTF-8')",
"_____no_output_____"
]
],
[
[
"### Parse HTML page",
"_____no_output_____"
]
],
[
[
"soup = BeautifulSoup(output_file.read_text(), 'html.parser')\ntype(soup)",
"_____no_output_____"
],
[
"tables = soup.find_all(class_='wikitable')\nprint(len(tables))\ntables[:10]",
"26\n"
],
[
"filtered_table_list = []\ntarget_column_names = ['English', 'Japanese', 'Rōmaji', 'TSE']\n\nprint(f'Unfiltered table list: {len(tables)}')\n\nfor table in tables:\n column_names = table.find_all('th')\n column_names = [c.text.strip() for c in column_names]\n column_names = set(column_names)\n\n if column_names == set(target_column_names):\n filtered_table_list.append(table)\n\nprint(f'Filtered table list: {len(filtered_table_list)}')",
"Unfiltered table list: 26\nFiltered table list: 24\n"
],
[
"# Get all the data points we need from the rows\ncompany_list = []\n\nfor table in filtered_table_list:\n row_list = table.find_all('tr')[1:]\n for row in row_list:\n cells = row.find_all('td')\n cells = [c.get_text().strip() for c in cells]\n\n def get_company_dict(cells):\n def get_value(idx):\n try:\n return cells[idx]\n except IndexError:\n return None\n\n try:\n company = {\n 'English': get_value(0),\n 'Japanese': get_value(1),\n 'Romaji': get_value(2),\n 'TSE': get_value(3),\n }\n return company\n except IndexError as e:\n print(cells)\n print(str(e))\n\n company_list.append(get_company_dict(cells))\n\nprint(len(company_list))\ncompany_list[:5]",
"601\n"
]
],
[
[
"### Convert company list to a pandas dataframe",
"_____no_output_____"
]
],
[
[
"df = pd.DataFrame(company_list, dtype='object')\ndf.info()\ndf.head()",
"<class 'pandas.core.frame.DataFrame'>\nRangeIndex: 601 entries, 0 to 600\nData columns (total 4 columns):\n # Column Non-Null Count Dtype \n--- ------ -------------- ----- \n 0 English 601 non-null object\n 1 Japanese 601 non-null object\n 2 Romaji 601 non-null object\n 3 TSE 598 non-null object\ndtypes: object(4)\nmemory usage: 18.9+ KB\n"
]
],
[
[
"Clean data",
"_____no_output_____"
]
],
[
[
"df['TSE'] = df['TSE'].str.replace('TYO: ', '').str.strip()\ndf.head()",
"_____no_output_____"
],
[
"df.to_csv('japanese_companies_wiki.csv', index=False, encoding='UTF-8')\nprint('Finished')",
"Finished\n"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4acde986eeaedd513325ac07ec04a6ad7fb5bb66
| 50,778 |
ipynb
|
Jupyter Notebook
|
what_is_torch_nn.ipynb
|
hopezh/pytorch_official_tutorials
|
b2fc0207b6cdcae5c590a441cbfd8b6161c2bc86
|
[
"MIT"
] | null | null | null |
what_is_torch_nn.ipynb
|
hopezh/pytorch_official_tutorials
|
b2fc0207b6cdcae5c590a441cbfd8b6161c2bc86
|
[
"MIT"
] | null | null | null |
what_is_torch_nn.ipynb
|
hopezh/pytorch_official_tutorials
|
b2fc0207b6cdcae5c590a441cbfd8b6161c2bc86
|
[
"MIT"
] | null | null | null | 33.037085 | 4,984 | 0.595041 |
[
[
[
"# What is torch.nn ?",
"_____no_output_____"
],
[
"## MNIST data setup",
"_____no_output_____"
],
[
"We will use the classic MNIST dataset, which consists of black-and-white images of hand-drawn digits (between 0 and 9).\n\nWe will use pathlib for dealing with paths (part of the Python 3 standard library), and will download the dataset using requests. We will only import modules when we use them, so you can see exactly what’s being used at each point.",
"_____no_output_____"
]
],
[
[
"from pathlib import Path\nimport requests\n\nDATA_PATH = Path('data')\nPATH = DATA_PATH / 'mnist'\n\nPATH.mkdir(parents=True, exist_ok=True)\n\nURL = 'https://github.com/pytorch/tutorials/raw/master/_static/'\nFILENAME = 'mnist.pkl.gz'\n\nif not (PATH / FILENAME).exists():\n content = requests.get(URL + FILENAME).content\n (PATH / FILENAME).open('wb').write(content)",
"_____no_output_____"
],
[
"import pickle\nimport gzip \n\nwith gzip.open((PATH / FILENAME).as_posix(), 'rb') as f:\n ((x_train, y_train), (x_valid, y_valid), _) = pickle.load(f, encoding='latin-1')",
"_____no_output_____"
]
],
[
[
"Each image is 28 x 28, and is being stored as a flattened row of length 784 (=28x28). Let’s take a look at one; we need to reshape it to 2d first.\n",
"_____no_output_____"
]
],
[
[
"from matplotlib import pyplot\nimport numpy as np\n\nprint(x_train.shape)\npyplot.imshow(x_train[0].reshape((28, 28)), cmap='gray')",
"(50000, 784)\n"
]
],
[
[
"PyTorch uses torch.tensor, rather than numpy arrays, so we need to convert our data.",
"_____no_output_____"
]
],
[
[
"import torch\n\nx_train, y_train, x_valid, y_valid = map(\n torch.tensor, (x_train, y_train, x_valid, y_valid)\n)\n\nn, c = x_train.shape\n\nx_train, x_train.shape, y_train.min(), y_train.max()\n\nprint(x_train, y_train)\nprint(x_train.shape)\nprint(y_train.min(), y_train.max())",
"tensor([[0., 0., 0., ..., 0., 0., 0.],\n [0., 0., 0., ..., 0., 0., 0.],\n [0., 0., 0., ..., 0., 0., 0.],\n ...,\n [0., 0., 0., ..., 0., 0., 0.],\n [0., 0., 0., ..., 0., 0., 0.],\n [0., 0., 0., ..., 0., 0., 0.]]) tensor([5, 0, 4, ..., 8, 4, 8])\ntorch.Size([50000, 784])\ntensor(0) tensor(9)\n"
]
],
[
[
"## neural net from scratch (no torch.nn)\n\nLet’s first create a model using nothing but PyTorch tensor operations. We’re assuming you’re already familiar with the basics of neural networks. (If you’re not, you can learn them at [course.fast.ai](https://course.fast.ai/)).\n\nPyTorch provides methods to create random or zero-filled tensors, which we will use to create our weights and bias for a simple linear model. These are just regular tensors, with one very special addition: we tell PyTorch that they require a gradient. This causes PyTorch to record all of the operations done on the tensor, so that it can calculate the gradient during back-propagation _automatically_!\n\nFor the weights, we set `requires_grad` **after** the initialization, since we don’t want that step included in the gradient. (Note that a trailing `_` in PyTorch signifies that the operation is performed in-place.)",
"_____no_output_____"
],
[
"We are initializing the weights here with [Xavier initialisation](http://proceedings.mlr.press/v9/glorot10a/glorot10a.pdf) (by multiplying with 1/sqrt(n)).",
"_____no_output_____"
]
],
[
[
"import math\n\n# 10 neurons in the output layer\nweights = torch.randn(784, 10) / math.sqrt(784)\nweights.requires_grad_() # set requires_grad in-place \nbias = torch.zeros(10, requires_grad=True) ",
"_____no_output_____"
],
[
"print(weights.shape, bias.shape)",
"torch.Size([784, 10]) torch.Size([10])\n"
]
],
[
[
"Thanks to PyTorch’s ability to calculate gradients automatically, we can use any standard Python function (or callable object) as a model! \n\nSo let’s just write a plain **matrix multiplication** and **broadcasted addition** to create a simple **linear model**. \n\nWe also need an **activation function**, so we’ll write _log_softmax_ and use it. \n\nRemember: although PyTorch provides lots of pre-written loss functions, activation functions, and so forth, you can easily write your own using plain python. PyTorch will even create fast GPU or vectorized CPU code for your function automatically.",
"_____no_output_____"
]
],
[
[
"def log_softmax(x):\n return x - x.exp().sum(-1).log().unsqueeze(-1)\n\ndef model(xb):\n return log_softmax( xb @ weights + bias )",
"_____no_output_____"
]
],
[
[
"In the above, the `@` stands for the **dot product** operation. \n\nWe will call our function on **one batch** of data (in this case, 64 images). \n\nThis is one $\\color{yellow}{\\textbf{foward}}$ pass. \n\nNote that our predictions won’t be any better than random at this stage, since we start with random weights.",
"_____no_output_____"
]
],
[
[
"bs = 64 # batch size\n\nxb = x_train[0 : bs] # one mini-batch from x\npreds = model(xb)\nprint(preds[0], preds.shape)",
"tensor([-2.3681, -2.4970, -2.5899, -1.9262, -2.0757, -2.3267, -2.2474, -1.9873,\n -2.5858, -2.7785], grad_fn=<SelectBackward>) torch.Size([64, 10])\n"
]
],
[
[
"As you see, the preds tensor contains not only the tensor values, but also a gradient function. We’ll use this later to do backprop.\n\nLet’s implement **negative log-likelihood** to use as the **loss function** (again, we can just use standard Python):",
"_____no_output_____"
]
],
[
[
"def nll(input, target):\n return -input[range(target.shape[0]), target].mean()\n\nloss_func = nll",
"_____no_output_____"
]
],
[
[
"Let’s check our loss with our random model, so we can see if we improve after a backprop pass later.",
"_____no_output_____"
]
],
[
[
"yb = y_train[0:bs]\nprint(loss_func(preds, yb))",
"tensor(2.3301, grad_fn=<NegBackward>)\n"
]
],
[
[
"Let’s also implement a function to calculate the **accuracy** of our model. \n\nFor each prediction, if the index with the largest value matches the target value, then the prediction was correct.",
"_____no_output_____"
]
],
[
[
"def accuracy(out, yb):\n preds = torch.argmax(out, dim=1)\n return (preds == yb).float().mean()",
"_____no_output_____"
],
[
"print(accuracy(preds, yb))",
"tensor(0.1562)\n"
]
],
[
[
"We can now run a training loop. For each iteration, we will:\n\n- select a mini-batch of data (of size bs)\n- use the model to make predictions\n- calculate the loss\n- loss.backward() updates the gradients of the model, in this case, weights and bias.\n\nWe now use these gradients to update the weights and bias. We do this within the `torch.no_grad()` context manager, because we do not want these actions to be recorded for our next calculation of the gradient. You can read more about how PyTorch’s Autograd records operations [here](https://pytorch.org/docs/stable/notes/autograd.html).\n\nWe then set the gradients to zero, so that we are ready for the next loop. Otherwise, our gradients would record a running tally of all the operations that had happened (i.e. `loss.backward()` **_adds_** the gradients to whatever is already stored, rather than replacing them).",
"_____no_output_____"
],
[
"You can use the standard python debugger to step through PyTorch code, allowing you to check the various variable values at each step. Uncomment `set_trace()` below to try it out.",
"_____no_output_____"
]
],
[
[
"from IPython.core.debugger import set_trace\n\nlr = 0.5\nepochs = 2\n\nfor epoch in range(epochs):\n for i in range( (n-1) // bs + 1 ):\n # set_trace()\n start_i = i * bs\n end_i = start_i + bs\n xb = x_train[start_i : end_i]\n yb = y_train[start_i : end_i]\n pred = model(xb)\n loss = loss_func(pred, yb)\n \n loss.backward()\n \n with torch.no_grad():\n weights -= weights.grad * lr\n bias -= bias.grad * lr\n weights.grad.zero_()\n bias.grad.zero_()\n",
"_____no_output_____"
]
],
[
[
"That’s it: we’ve created and trained a minimal neural network (in this case, a logistic regression, since we have no hidden layers) entirely from scratch!\n\nLet’s check the loss and accuracy and compare those to what we got earlier. We expect that the loss will have decreased and accuracy to have increased, and they have.",
"_____no_output_____"
]
],
[
[
"print( \n loss_func(model(xb), yb), \n accuracy(model(xb), yb) \n)",
"tensor(0.0827, grad_fn=<NegBackward>) tensor(1.)\n"
]
],
[
[
"## Using torch.nn.functional\n\nWe will now refactor our code, so that it does the same thing as before, only we’ll start taking advantage of PyTorch’s `nn` classes to make it more concise and flexible. At each step from here, we should be making our code one or more of: **shorter, more understandable, and/or more flexible**.\n\nThe first and easiest step is to make our code shorter by replacing our hand-written activation and loss functions with those from `torch.nn.functional` (which is generally imported into the namespace `F` by convention). This module contains all the **functions** in the `torch.nn` library (whereas other parts of the library contain **classes**). As well as a wide range of loss and activation functions, you’ll also find here some convenient functions for creating neural nets, such as pooling functions. (There are also functions for doing convolutions, linear layers, etc, but as we’ll see, these are usually better handled using other parts of the library.)\n\nIf you’re using negative log likelihood loss and log softmax activation, then Pytorch provides a single function `F.cross_entropy` that combines the two. So we can even remove the activation function from our model.",
"_____no_output_____"
]
],
[
[
"import torch.nn.functional as F \n\nloss_func = F.cross_entropy\n\ndef model(xb):\n return xb @ weights + bias",
"_____no_output_____"
]
],
[
[
"Note that we no longer call `log_softmax` in the model function. \n\nLet’s confirm that our loss and accuracy are the same as before:",
"_____no_output_____"
]
],
[
[
"print( \n loss_func(model(xb), yb), \n accuracy(model(xb), yb) \n)",
"tensor(0.0827, grad_fn=<NllLossBackward>) tensor(1.)\n"
]
],
[
[
"## Refactor using nn.Module\n\nNext up, we’ll use `nn.Module` and `nn.Parameter`, for a clearer and more concise training loop. \n\nWe subclass `nn.Module` (which itself is a class and able to keep track of state). \n\nIn this case, we want to create a class that holds our weights, bias, and method for the forward step. \n\n`nn.Module` has a number of **attributes** and **methods** (such as `.parameters()` and `.zero_grad()`) which we will be using.",
"_____no_output_____"
]
],
[
[
"from torch import nn \n\nclass Mnist_Logistic(nn.Module):\n def __init__(self):\n super().__init__()\n self.weights = nn.Parameter( torch.randn(784, 10) / math.sqrt(784) )\n self.bias = nn.Parameter( torch.zeros(10) )\n \n def forward(self, xb):\n return xb @ self.weights + self.bias",
"_____no_output_____"
]
],
[
[
"Since we’re now using an **object** instead of just using a function, we first have to **instantiate** our model:",
"_____no_output_____"
]
],
[
[
"model = Mnist_Logistic()",
"_____no_output_____"
]
],
[
[
"Now we can calculate the loss in the same way as before. \n\nNote that `nn.Module` objects are used as if they are **functions** (i.e they are callable), but behind the scenes Pytorch will call our forward method automatically.",
"_____no_output_____"
]
],
[
[
"print(loss_func(model(xb), yb))",
"tensor(2.2570, grad_fn=<NllLossBackward>)\n"
]
],
[
[
"Previously for our training loop we had to update the values for each parameter by name, and manually zero out the grads for each parameter separately, like this:\n\n```\nwith torch.no_grad():\n weights -= weights.grad * lr\n bias -= bias.grad * lr\n weights.grad.zero_()\n bias.grad.zero_()\n```\n\nNow we can take advantage of `model.parameters()` and `model.zero_grad()` (which are both defined by PyTorch for nn.Module) to make those steps more concise and less prone to the error of forgetting some of our parameters, particularly if we had a more complicated model:",
"_____no_output_____"
]
],
[
[
"with torch.no_grad():\n for p in model.parameters(): p -= p.grad * lr\n model.zero_grad()",
"_____no_output_____"
]
],
[
[
"We’ll wrap our little training loop in a fit function so we can run it again later.",
"_____no_output_____"
]
],
[
[
"def fit():\n for epoch in range(epochs):\n for i in range( (n-1) // bs + 1 ):\n # set_trace()\n start_i = i * bs\n end_i = start_i + bs\n xb = x_train[start_i : end_i]\n yb = y_train[start_i : end_i]\n pred = model(xb)\n loss = loss_func(pred, yb)\n \n loss.backward()\n \n with torch.no_grad():\n for p in model.parameters(): \n p -= p.grad * lr\n model.zero_grad()",
"_____no_output_____"
],
[
"fit()",
"_____no_output_____"
]
],
[
[
"Let’s double-check that our loss has gone down:",
"_____no_output_____"
]
],
[
[
"print(loss_func(model(xb), yb))",
"tensor(0.0810, grad_fn=<NllLossBackward>)\n"
]
],
[
[
"## Refactor using nn.Linear\n\nWe continue to refactor our code. \n\nInstead of manually defining and initializing `self.weights` and `self.bias`, and calculating `xb @ self.weights + self.bias`, we will instead use the Pytorch class `nn.Linear` for a **linear** layer, which does all that for us. \n\nPytorch has many types of predefined layers that can greatly simplify our code, and often makes it faster too.",
"_____no_output_____"
]
],
[
[
"class Mnist_Logistic(nn.Module):\n def __init__(self):\n super().__init__()\n self.lin = nn.Linear(784, 10)\n \n def forward(self, xb):\n return self.lin(xb)",
"_____no_output_____"
]
],
[
[
"We instantiate our model and calculate the loss in the same way as before:",
"_____no_output_____"
]
],
[
[
"model = Mnist_Logistic()\nprint(loss_func(model(xb), yb))",
"tensor(2.3793, grad_fn=<NllLossBackward>)\n"
]
],
[
[
"We are still able to use our same fit method as before.",
"_____no_output_____"
]
],
[
[
"fit()\nprint(loss_func(model(xb), yb))",
"tensor(0.0820, grad_fn=<NllLossBackward>)\n"
]
],
[
[
"## Refactor using optim\n\nPytorch also has a package with various optimization algorithms, `torch.optim`. \n\nWe can use the `step` method from our optimizer to take a forward step, instead of manually updating each parameter.\n\nThis will let us replace our previous manually coded optimization step:\n\n```\nwith torch.no_grad():\n for p in model.parameters(): p -= p.grad * lr\n model.zero_grad()\n```\n\nand instead use just:\n\n```\nopt.step()\nopt.zero_grad()\n```\n\n`optim.zero_grad()` resets the gradient to 0 and we need to call it before computing the gradient for the next minibatch. \n\nWe’ll define a little function to create our model and optimizer so we can reuse it in the future.",
"_____no_output_____"
]
],
[
[
"from torch import optim \n\ndef get_model():\n model = Mnist_Logistic()\n return model, optim.SGD(model.parameters(), lr=lr)\n\nmodel, opt = get_model()\nprint('before training:', loss_func(model(xb), yb))\n\nfor epoch in range(epochs):\n for i in range( (n-1) // bs + 1 ):\n # set_trace()\n start_i = i * bs\n end_i = start_i + bs\n xb = x_train[start_i : end_i]\n yb = y_train[start_i : end_i]\n pred = model(xb)\n loss = loss_func(pred, yb)\n \n loss.backward()\n \n opt.step()\n opt.zero_grad()\n\nprint('after training:', loss_func(model(xb), yb)) ",
"before training: tensor(2.2926, grad_fn=<NllLossBackward>)\nafter training: tensor(0.0807, grad_fn=<NllLossBackward>)\n"
]
],
[
[
"## Refactor using Dataset\n\nPyTorch has an abstract `Dataset` class. A Dataset can be anything that has a `__len__` function (called by Python’s standard len function) and a `__getitem__` function as a way of indexing into it. \n\n[This tutorial](https://pytorch.org/tutorials/beginner/data_loading_tutorial.html) walks through a nice example of creating a custom `FacialLandmarkDataset` class as a subclass of `Dataset`.\n\nPyTorch’s `TensorDataset` is a Dataset wrapping tensors. By defining a length and way of indexing, this also gives us a way to iterate, index, and slice along the first dimension of a tensor. This will make it easier to access both the **independent** and **dependent** variables in the same line as we train.",
"_____no_output_____"
]
],
[
[
"from torch.utils.data import TensorDataset",
"_____no_output_____"
]
],
[
[
"Both `x_train` and `y_train` can be combined in a single `TensorDataset`, which will be easier to iterate over and slice.",
"_____no_output_____"
]
],
[
[
"train_ds = TensorDataset(x_train, y_train)",
"_____no_output_____"
]
],
[
[
"Previously, we had to iterate through minibatches of x and y values separately:\n\n```\nxb = x_train[start_i:end_i]\nyb = y_train[start_i:end_i]\n```\n\nNow, we can do these two steps together:\n\n```\nxb,yb = train_ds[i*bs : i*bs+bs]\n\n```",
"_____no_output_____"
]
],
[
[
"model, opt = get_model()\n\nfor epoch in range(epochs):\n for i in range( (n-1) // bs + 1 ):\n xb, yb = train_ds[i * bs : i * bs + bs]\n pred = model(xb)\n loss = loss_func(pred, yb)\n \n loss.backward()\n \n opt.step()\n opt.zero_grad()\n\nprint(loss_func(model(xb), yb))",
"tensor(0.0829, grad_fn=<NllLossBackward>)\n"
]
],
[
[
"## Refactor using DataLoader\n\nPytorch’s `DataLoader` is responsible for managing **batches**. \n\nYou can create a `DataLoader` from any Dataset. `DataLoader` makes it easier to **iterate** over batches. \n\nRather than having to use `train_ds[i*bs : i*bs+bs]`, the DataLoader gives us each **minibatch** automatically.",
"_____no_output_____"
]
],
[
[
"from torch.utils.data import DataLoader\n\ntrain_ds = TensorDataset(x_train, y_train)\ntrain_dl = DataLoader(train_ds, batch_size=bs)",
"_____no_output_____"
]
],
[
[
"Previously, our loop iterated over batches (xb, yb) like this:\n\n```\nfor i in range((n-1)//bs + 1):\n xb,yb = train_ds[i*bs : i*bs+bs]\n pred = model(xb)\n```\n\nNow, our loop is much cleaner, as (xb, yb) are loaded automatically from the data loader:\n\n```\nfor xb,yb in train_dl:\n pred = model(xb)\n```",
"_____no_output_____"
]
],
[
[
"model, opt = get_model()\n\nfor epoch in range(epochs):\n for xb, yb in train_dl:\n pred = model(xb)\n loss = loss_func(pred, yb)\n \n loss.backward()\n \n opt.step()\n opt.zero_grad()\n\nprint(loss_func(model(xb), yb))",
"tensor(0.0826, grad_fn=<NllLossBackward>)\n"
]
],
[
[
"Thanks to Pytorch’s `nn.Module`, `nn.Parameter`, `Dataset`, and `DataLoader`, our training loop is now dramatically smaller and easier to understand. \n\nLet’s now try to add the basic features necessary to create effective models in practice.",
"_____no_output_____"
],
[
"## Add validation\n\nIn section 1, we were just trying to get a reasonable training loop set up for use on our training data.\n\nIn reality, you always should also have a **validation set**, in order to identify if you are overfitting.\n\n**Shuffling** the training data is important to prevent correlation between batches and overfitting. \n\nOn the other hand, the validation loss will be identical whether we shuffle the validation set or not. Since shuffling takes extra time, it makes no sense to shuffle the validation data.\n\nWe’ll use a batch size for the validation set that is twice as large as that for the training set. This is because the **validation set does not need _backpropagation_** and thus takes less memory (it doesn’t need to store the gradients). We take advantage of this to use a larger batch size and compute the loss more quickly.",
"_____no_output_____"
]
],
[
[
"train_ds = TensorDataset(x_train, y_train)\ntrain_dl = DataLoader(train_ds, batch_size=bs)\n\nvalid_ds = TensorDataset(x_valid, y_valid)\nvalid_dl = DataLoader(valid_ds, batch_size=bs*2)",
"_____no_output_____"
]
],
[
[
"We will calculate and print the **validation loss** at the end of each epoch.\n\n(Note that we always call **`model.train()`** before training, and **`model.eval()`** before inference, because these are used by layers such as `nn.BatchNorm2d` and `nn.Dropout` to ensure appropriate behaviour for these different phases.)",
"_____no_output_____"
]
],
[
[
"model, opt = get_model()\n\nfor epoch in range(epochs):\n model.train()\n \n for xb, yb in train_dl:\n pred = model(xb)\n loss = loss_func(pred, yb)\n \n loss.backward()\n \n opt.step()\n opt.zero_grad()\n\n model.eval()\n \n with torch.no_grad():\n valid_loss = sum(loss_func(model(xb), yb) for xb, yb in valid_dl) \n \n print(epoch, valid_loss / len(valid_dl))",
"0 tensor(0.3089)\n1 tensor(0.2899)\n"
]
],
[
[
"## Create fit() and get_data()\n\nWe’ll now do a little refactoring of our own. \n\nSince we go through a similar process twice of calculating the loss for both the training set and the validation set, let’s make that into its own function, `loss_batch`, which computes the loss for one batch.\n\nWe pass an optimizer in for the training set, and use it to perform backprop. \n\nFor the validation set, we don’t pass an optimizer, so the method doesn’t perform backprop.",
"_____no_output_____"
]
],
[
[
"def loss_batch(model, loss_func, xb, yb, opt=None):\n loss = loss_func(model(xb), yb)\n \n if opt is not None:\n loss.backward()\n opt.step()\n opt.zero_grad()\n \n return loss.item(), len(xb)",
"_____no_output_____"
]
],
[
[
"`fit` runs the necessary operations to train our model and compute the training and validation losses for each epoch.",
"_____no_output_____"
]
],
[
[
"import numpy as np \n\ndef fit(epoch, model, loss_func, opt, train_dl, valid_dl):\n for epoch in range(epochs):\n model.train()\n \n for xb, yb in train_dl:\n loss_batch(model, loss_func, xb, yb, opt)\n\n model.eval()\n \n with torch.no_grad():\n losses, nums = zip(\n *[ loss_batch(model, loss_func, xb, yb) for xb, yb in valid_dl ]\n )\n \n val_loss = np.sum(np.multiply(losses, nums)) / np.sum(nums)\n \n print(epoch, ':\\t', val_loss)",
"_____no_output_____"
]
],
[
[
"`get_data` returns dataloaders for the training and validation sets.",
"_____no_output_____"
]
],
[
[
"def get_data(train_dl, valid_dl, bs):\n return (\n DataLoader(train_dl, batch_size=bs, shuffle=True), \n DataLoader(valid_dl, batch_size=bs*2), \n )",
"_____no_output_____"
]
],
[
[
"Now, our whole process of obtaining the data loaders and fitting the model can be run in 3 lines of code:",
"_____no_output_____"
]
],
[
[
"train_dl, valid_dl = get_data(train_ds, valid_ds, bs)\nmodel, opt = get_model()\nfit(epochs, model, loss_func, opt, train_dl, valid_dl)",
"0 :\t 0.35734008679389956\n1 :\t 0.28959022200107576\n"
]
],
[
[
"You can use these basic 3 lines of code to train a wide variety of models. \n\nLet’s see if we can use them to train a convolutional neural network (CNN)!",
"_____no_output_____"
],
[
"## Switch to CNN\n\nWe are now going to build our neural network with three convolutional layers. \n\nBecause none of the functions in the previous section assume anything about the model form, we’ll be able to use them to train a CNN without any modification.\n\nWe will use Pytorch’s predefined `Conv2d` class as our convolutional layer. \n\nWe define a CNN with 3 convolutional layers. Each convolution is followed by a ReLU. At the end, we perform an average pooling. (Note that **`view`** is PyTorch’s version of numpy’s reshape)",
"_____no_output_____"
]
],
[
[
"import torch.nn.functional as F\n\nclass Mnist_CNN(nn.Module):\n def __init__(self):\n super().__init__()\n \n self.conv1 = nn.Conv2d(1, 16, kernel_size=3, stride=2, padding=1)\n self.conv2 = nn.Conv2d(16, 16, kernel_size=3, stride=2, padding=1)\n self.conv3 = nn.Conv2d(16, 10, kernel_size=3, stride=2, padding=1)\n \n def forward(self, xb):\n xb = xb.view(-1, 1, 28, 28)\n xb = F.relu(self.conv1(xb))\n xb = F.relu(self.conv2(xb))\n xb = F.relu(self.conv3(xb))\n xb = F.avg_pool2d(xb, 4)\n return xb.view(-1, xb.size(1))",
"_____no_output_____"
]
],
[
[
"`Momentum` is a variation on stochastic gradient descent that takes previous updates into account as well and generally leads to faster training.",
"_____no_output_____"
]
],
[
[
"lr = 0.1 \nmodel = Mnist_CNN()\nopt = optim.SGD(model.parameters(), lr=lr, momentum=0.9)\n\nfit(epochs, model, loss_func, opt, train_dl, valid_dl)",
"0 :\t 0.3153703056097031\n1 :\t 0.257355453813076\n"
]
],
[
[
"## nn.Sequential\n\ntorch.nn has another handy class we can use to simplify our code: `Sequential`. \n\nA `Sequential` object runs each of the modules contained within it, in a sequential manner. This is a simpler way of writing our neural network.\n\nTo take advantage of this, we need to be able to easily define a custom layer from a given function. \n\nFor instance, PyTorch doesn’t have a **view** layer, and we need to create one for our network. \n\nThe following class `Lambda` will create a layer that we can then use when defining a network with Sequential.",
"_____no_output_____"
]
],
[
[
"class Lambda(nn.Module):\n def __init__(self, func):\n super().__init__()\n self.func = func\n \n def forward(self, x):\n return self.func(x)\n\ndef preprocess(x):\n return x.view(-1, 1, 28, 28)",
"_____no_output_____"
]
],
[
[
"The model created with `Sequential` is simply:",
"_____no_output_____"
]
],
[
[
"model = nn.Sequential(\n Lambda(preprocess), \n nn.Conv2d(1, 16, kernel_size=3, stride=2, padding=1), \n nn.ReLU(), \n nn.Conv2d(16, 16, kernel_size=3, stride=2, padding=1),\n nn.ReLU(), \n nn.Conv2d(16, 10, kernel_size=3, stride=2, padding=1),\n nn.ReLU(), \n nn.AvgPool2d(4), \n Lambda(lambda x: x.view(x.size(0), -1)), \n)\n\nopt = optim.SGD(model.parameters(), lr=lr, momentum=0.9)\n\nfit(epochs, model, loss_func, opt, train_dl, valid_dl)",
"0 :\t 0.31578762679100036\n1 :\t 0.2491037842988968\n"
]
],
[
[
"## Wrapping DataLoader\n\nOur CNN is fairly concise, but it only works with MNIST, because:\n- It assumes the input is a 28*28 long vector\n- It assumes that the final CNN grid size is 4*4 (since that’s the average pooling kernel size we used)\n\nLet’s get rid of these two assumptions, so our model works with any 2d single channel image. First, we can remove the initial Lambda layer by moving the data preprocessing into a generator:",
"_____no_output_____"
]
],
[
[
"def preprocess(x, y):\n return x.view(-1, 1, 28, 28), y\n\nclass WrappedDataLoader:\n def __init__(self, dl, func):\n self.dl = dl\n self.func = func \n \n def __len__(self):\n return len(self.dl)\n\n def __iter__(self):\n batches = iter(self.dl)\n for b in batches:\n yield (self.func(*b))\n \ntrain_dl, valid_dl = get_data(train_ds, valid_ds, bs)\ntrain_dl = WrappedDataLoader(train_dl, preprocess)\nvalid_dl = WrappedDataLoader(valid_dl, preprocess)",
"_____no_output_____"
]
],
[
[
"Next, we can replace `nn.AvgPool2d` with `nn.AdaptiveAvgPool2d`, which allows us to define the size of the output tensor we want, rather than the input tensor we have. As a result, our model will work with any size input.",
"_____no_output_____"
]
],
[
[
"model = nn.Sequential(\n nn.Conv2d(1, 16, kernel_size=3, stride=2, padding=1), \n nn.ReLU(), \n nn.Conv2d(16, 16, kernel_size=3, stride=2, padding=1),\n nn.ReLU(), \n nn.Conv2d(16, 10, kernel_size=3, stride=2, padding=1),\n nn.ReLU(), \n nn.AdaptiveAvgPool2d(1), \n Lambda(lambda x: x.view(x.size(0), -1)), \n)\n\nopt = optim.SGD(model.parameters(), lr=lr, momentum=0.9)\n\nfit(epochs, model, loss_func, opt, train_dl, valid_dl)",
"0 :\t 0.3755627481341362\n1 :\t 0.22480471715927125\n"
]
],
[
[
"## Using your GPU\n\nIf you’re lucky enough to have access to a CUDA-capable GPU (you can rent one for about $0.50/hour from most cloud providers) you can use it to speed up your code. First check that your GPU is working in Pytorch:",
"_____no_output_____"
]
],
[
[
"print(torch.cuda.is_available())",
"False\n"
]
],
[
[
"And then create a device object for it:",
"_____no_output_____"
]
],
[
[
"dev = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')",
"_____no_output_____"
]
],
[
[
"Let’s update preprocess to move batches to the GPU:",
"_____no_output_____"
]
],
[
[
"def preprocess(x, y):\n return x.view(-1, 1, 28, 28).to(dev), y.to(dev)\n\ntrain_dl, valid_dl = get_data(train_ds, valid_ds, bs)\ntrain_dl = WrappedDataLoader(train_dl, preprocess)\nvalid_dl = WrappedDataLoader(valid_dl, preprocess)",
"_____no_output_____"
]
],
[
[
"Finally, we can move our model to the GPU.",
"_____no_output_____"
]
],
[
[
"model.to(dev)\n\nopt = optim.SGD(model.parameters(), lr=lr, momentum=0.9)\n\nfit(epochs, model, loss_func, opt, train_dl, valid_dl)",
"0 :\t 0.18121731426119805\n1 :\t 0.16951641612648963\n"
]
],
[
[
"## Closing thoughts\n\n\nWe now have a general data pipeline and training loop which you can use for training many types of models using Pytorch. To see how simple training a model can now be, take a look at the mnist_sample sample notebook.\n\nOf course, there are many things you’ll want to add, such as data augmentation, hyperparameter tuning, monitoring training, transfer learning, and so forth. These features are available in the fastai library, which has been developed using the same design approach shown in this tutorial, providing a natural next step for practitioners looking to take their models further.\n\nWe promised at the start of this tutorial we’d explain through example each of torch.nn, torch.optim, Dataset, and DataLoader. So let’s summarize what we’ve seen:\n\n- torch.nn \n - Module: creates a callable which behaves like a function, but can also contain state(such as neural net layer weights). It knows what Parameter (s) it contains and can zero all their gradients, loop through them for weight updates, etc.\n \n - Parameter: a wrapper for a tensor that tells a Module that it has weights that need updating during backprop. Only tensors with the requires_grad attribute set are updated\n \n - functional: a module(usually imported into the F namespace by convention) which contains activation functions, loss functions, etc, as well as non-stateful versions of layers such as convolutional and linear layers.\n\n- torch.optim: Contains optimizers such as SGD, which update the weights of Parameter during the backward step\n\n- Dataset: An abstract interface of objects with a __len__ and a __getitem__, including classes provided with Pytorch such as TensorDataset\n\n- DataLoader: Takes any Dataset and creates an iterator which returns batches of data.",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
4ace1006c83e63715b49e355d2d6c647127eaf43
| 774,207 |
ipynb
|
Jupyter Notebook
|
notebooks/13-isolines-isochrones.ipynb
|
zh-yao/osmnx-examples
|
58f44376c03851e4e5c72bffaa8eb87f94238731
|
[
"MIT"
] | 1 |
2019-09-11T08:07:16.000Z
|
2019-09-11T08:07:16.000Z
|
notebooks/13-isolines-isochrones.ipynb
|
TheSuguser/osmnx-examples
|
58f44376c03851e4e5c72bffaa8eb87f94238731
|
[
"MIT"
] | null | null | null |
notebooks/13-isolines-isochrones.ipynb
|
TheSuguser/osmnx-examples
|
58f44376c03851e4e5c72bffaa8eb87f94238731
|
[
"MIT"
] | null | null | null | 2,707.017483 | 265,204 | 0.962482 |
[
[
[
"# Draw an isochrone map with OSMnx\n\nHow far can you travel on foot in 15 minutes?\n\n - [Overview of OSMnx](http://geoffboeing.com/2016/11/osmnx-python-street-networks/)\n - [GitHub repo](https://github.com/gboeing/osmnx)\n - [Examples, demos, tutorials](https://github.com/gboeing/osmnx-examples)\n - [Documentation](https://osmnx.readthedocs.io/en/stable/)\n - [Journal article/citation](http://geoffboeing.com/publications/osmnx-complex-street-networks/)",
"_____no_output_____"
]
],
[
[
"import geopandas as gpd\nimport matplotlib.pyplot as plt\nimport networkx as nx\nimport osmnx as ox\nfrom descartes import PolygonPatch\nfrom shapely.geometry import Point, LineString, Polygon\nox.config(log_console=True, use_cache=True)\nox.__version__",
"_____no_output_____"
],
[
"# configure the place, network type, trip times, and travel speed\nplace = 'Berkeley, CA, USA'\nnetwork_type = 'walk'\ntrip_times = [5, 10, 15, 20, 25] #in minutes\ntravel_speed = 4.5 #walking speed in km/hour",
"_____no_output_____"
]
],
[
[
"## Download and prep the street network",
"_____no_output_____"
]
],
[
[
"# download the street network\nG = ox.graph_from_place(place, network_type=network_type)",
"_____no_output_____"
],
[
"# find the centermost node and then project the graph to UTM\ngdf_nodes = ox.graph_to_gdfs(G, edges=False)\nx, y = gdf_nodes['geometry'].unary_union.centroid.xy\ncenter_node = ox.get_nearest_node(G, (y[0], x[0]))\nG = ox.project_graph(G)",
"_____no_output_____"
],
[
"# add an edge attribute for time in minutes required to traverse each edge\nmeters_per_minute = travel_speed * 1000 / 60 #km per hour to m per minute\nfor u, v, k, data in G.edges(data=True, keys=True):\n data['time'] = data['length'] / meters_per_minute",
"_____no_output_____"
]
],
[
[
"## Plots nodes you can reach on foot within each time\n\nHow far can you walk in 5, 10, 15, 20, and 25 minutes from the origin node? We'll use NetworkX to induce a subgraph of G within each distance, based on trip time and travel speed.",
"_____no_output_____"
]
],
[
[
"# get one color for each isochrone\niso_colors = ox.get_colors(n=len(trip_times), cmap='Reds', start=0.3, return_hex=True)",
"_____no_output_____"
],
[
"# color the nodes according to isochrone then plot the street network\nnode_colors = {}\nfor trip_time, color in zip(sorted(trip_times, reverse=True), iso_colors):\n subgraph = nx.ego_graph(G, center_node, radius=trip_time, distance='time')\n for node in subgraph.nodes():\n node_colors[node] = color\nnc = [node_colors[node] if node in node_colors else 'none' for node in G.nodes()]\nns = [20 if node in node_colors else 0 for node in G.nodes()]\nfig, ax = ox.plot_graph(G, fig_height=8, node_color=nc, node_size=ns, node_alpha=0.8, node_zorder=2)",
"_____no_output_____"
]
],
[
[
"## Plot the time-distances as isochrones\n\nHow far can you walk in 5, 10, 15, 20, and 25 minutes from the origin node? We'll use a convex hull, which isn't perfectly accurate. A concave hull would be better, but shapely doesn't offer that.",
"_____no_output_____"
]
],
[
[
"# make the isochrone polygons\nisochrone_polys = []\nfor trip_time in sorted(trip_times, reverse=True):\n subgraph = nx.ego_graph(G, center_node, radius=trip_time, distance='time')\n node_points = [Point((data['x'], data['y'])) for node, data in subgraph.nodes(data=True)]\n bounding_poly = gpd.GeoSeries(node_points).unary_union.convex_hull\n isochrone_polys.append(bounding_poly)",
"_____no_output_____"
],
[
"# plot the network then add isochrones as colored descartes polygon patches\nfig, ax = ox.plot_graph(G, fig_height=8, show=False, close=False, edge_color='k', edge_alpha=0.2, node_color='none')\nfor polygon, fc in zip(isochrone_polys, iso_colors):\n patch = PolygonPatch(polygon, fc=fc, ec='none', alpha=0.6, zorder=-1)\n ax.add_patch(patch)\nplt.show()",
"_____no_output_____"
]
],
[
[
"## Or, plot isochrones as buffers to get more faithful isochrones than convex hulls can offer\n\nin the style of http://kuanbutts.com/2017/12/16/osmnx-isochrones/",
"_____no_output_____"
]
],
[
[
"def make_iso_polys(G, edge_buff=25, node_buff=50, infill=False):\n isochrone_polys = []\n for trip_time in sorted(trip_times, reverse=True):\n subgraph = nx.ego_graph(G, center_node, radius=trip_time, distance='time')\n\n node_points = [Point((data['x'], data['y'])) for node, data in subgraph.nodes(data=True)]\n nodes_gdf = gpd.GeoDataFrame({'id': subgraph.nodes()}, geometry=node_points)\n nodes_gdf = nodes_gdf.set_index('id')\n\n edge_lines = []\n for n_fr, n_to in subgraph.edges():\n f = nodes_gdf.loc[n_fr].geometry\n t = nodes_gdf.loc[n_to].geometry\n edge_lines.append(LineString([f,t]))\n\n n = nodes_gdf.buffer(node_buff).geometry\n e = gpd.GeoSeries(edge_lines).buffer(edge_buff).geometry\n all_gs = list(n) + list(e)\n new_iso = gpd.GeoSeries(all_gs).unary_union\n \n # try to fill in surrounded areas so shapes will appear solid and blocks without white space inside them\n if infill:\n new_iso = Polygon(new_iso.exterior)\n isochrone_polys.append(new_iso)\n return isochrone_polys\n\nisochrone_polys = make_iso_polys(G, edge_buff=25, node_buff=0, infill=True)\nfig, ax = ox.plot_graph(G, fig_height=8, show=False, close=False, edge_color='k', edge_alpha=0.2, node_color='none')\nfor polygon, fc in zip(isochrone_polys, iso_colors):\n patch = PolygonPatch(polygon, fc=fc, ec='none', alpha=0.6, zorder=-1)\n ax.add_patch(patch)\nplt.show()",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ace1187fe2ee0822cd6114129b4778294314ae3
| 28,434 |
ipynb
|
Jupyter Notebook
|
python3.6/.ipynb_checkpoints/python15-checkpoint.ipynb
|
gjbr5/python-link-e-learning
|
ed18f9537af29d7edd75686c9203d31b0a46ae5e
|
[
"MIT"
] | null | null | null |
python3.6/.ipynb_checkpoints/python15-checkpoint.ipynb
|
gjbr5/python-link-e-learning
|
ed18f9537af29d7edd75686c9203d31b0a46ae5e
|
[
"MIT"
] | null | null | null |
python3.6/.ipynb_checkpoints/python15-checkpoint.ipynb
|
gjbr5/python-link-e-learning
|
ed18f9537af29d7edd75686c9203d31b0a46ae5e
|
[
"MIT"
] | 6 |
2020-09-04T10:16:59.000Z
|
2020-12-03T01:47:03.000Z
| 19.329708 | 335 | 0.441795 |
[
[
[
"***\n***\n# 15. 파이썬 함수\n***\n***",
"_____no_output_____"
],
[
"***\n## 1 함수의 정의와 호출\n***\n- 함수: 여러 개의 Statement들을 하나로 묶은 단위\n- 함수 사용의 장점\n - 반복적인 수행이 가능하다\n - 코드를 논리적으로 이해하는 데 도움을 준다\n - 코드의 일정 부분을 별도의 논리적 개념으로 독립화할 수 있음\n - 수학에서 복잡한 개념을 하나의 단순한 기호로 대치하는 것과 비슷",
"_____no_output_____"
],
[
"### 1-1 간단한 함수의 정의\n- 함수 정의시 사용하는 키워드: def",
"_____no_output_____"
]
],
[
[
"def add(a, b):\n return a + b\n\nprint(add(1, 2))\nprint()\n\ndef myabs(x):\n if x < 0 : \n x = -x\n return x\n\nprint(abs(-4))\nprint(myabs(-4))",
"3\n\n4\n4\n"
]
],
[
[
"### 1-2 함수 객체와 함수 호출\n- 함수의 이름 자체는 함수 객체의 레퍼런스(Reference)를 지니고 있다.",
"_____no_output_____"
]
],
[
[
"def add(a, b):\n return a + b\n\nprint(add)",
"<function add at 0x104e88bf8>\n"
],
[
"c = add(10, 30)\nprint(c)",
"40\n"
]
],
[
[
"- 함수 이름에 저장된 레퍼런스를 다른 변수에 할당하여 그 변수를 이용한 함수 호출 가능 ",
"_____no_output_____"
]
],
[
[
"f = add\nprint(f(4, 5))",
"9\n"
],
[
"print(f)\n\nprint(f is add)",
"<function add at 0x104e88bf8>\nTrue\n"
]
],
[
[
"- 함수의 몸체에는 최소한 한개 이상의 statement가 존재해야 함\n - 아무런 내용이 없는 몸체를 지닌 함수를 만들 때에는 pass 라는 키워드를 몸체에 적어주어야 함",
"_____no_output_____"
]
],
[
[
"def simpleFunction():\n pass\n\nsimpleFunction()",
"_____no_output_____"
]
],
[
[
"- 함수 사용 예",
"_____no_output_____"
]
],
[
[
"def addmember(members, newmember):\n if newmember not in members: # 기존 멤버가 아니면\n members.append(newmember) # 추가\n\nmembers = ['hwang', 'lee', 'park', 'youn'] # 리스트에 초기 멤버 설정\n\naddmember(members, 'jo') # 새로운 멤버 추가\n\naddmember(members, 'hwang') # (이미 존재하는) 새로운 멤버 추가\n\nprint(members)",
"['hwang', 'lee', 'park', 'youn', 'jo']\n"
]
],
[
[
"### 1-3 함수 인수값 전달방법\n- 파이썬에서의 인수값 전달 방법\n - 기본적으로 값에 의한 호출(Call-by-Value)\n - 하지만 변수에 저장된 값이 참조값(Reference)이므로 실제로는 참조에 의한 호출(Call-by-Reference)로 실행됨",
"_____no_output_____"
],
[
"- 함수 인자에 변경불가능(Immutable) 객체인 숫자값을 전달\n - 함수 내에서 다른 숫자값으로 치환 --> 의미 없는 인자 전달",
"_____no_output_____"
]
],
[
[
"def f1(b):\n b = 100\n\na = 200\nf1(a)\nprint(a)",
"200\n"
]
],
[
[
"",
"_____no_output_____"
],
[
"- 함수 인자에 변경불가능(Immutable) 객체인 문자열을 전달\n - 함수 내에서 다른 문자열로 치환 --> 의미 없는 인자 전달",
"_____no_output_____"
]
],
[
[
"def f2(b):\n b = \"abc\"\n\na = \"def\"\nf2(a)\nprint(a)",
"def\n"
]
],
[
[
"",
"_____no_output_____"
],
[
"- 함수 인자에 변경불가능(Immutable) 객체인 튜플을 전달\n - 함수 내에서 다른 튜플로 치환 --> 의미 없는 인자 전달",
"_____no_output_____"
]
],
[
[
"def f3(b):\n b = (1,2,3)\n\na = (4,5,6)\nf3(a)\nprint(a)",
"(4, 5, 6)\n"
]
],
[
[
"",
"_____no_output_____"
],
[
"- 함수 인자에 변경가능한(Mutable)한 객체인 리스트 전달 및 내용 수정\n - 전형적인 함수 인자 전달법 및 활용법",
"_____no_output_____"
]
],
[
[
"def f4(b):\n b[1] = 10\n\na = [4,5,6]\nf4(a)\nprint(a)",
"[4, 10, 6]\n"
]
],
[
[
"",
"_____no_output_____"
],
[
"- 함수 인자에 변경가능한(Mutable)한 객체인 사전 전달 및 내용 수정\n - 전형적인 함수 인자 전달법 및 활용법",
"_____no_output_____"
]
],
[
[
"def f5(b):\n b['a'] = 10\n\na = {\"a\":1, \"b\":2}\nf5(a)\nprint(a)",
"{'a': 10, 'b': 2}\n"
]
],
[
[
"",
"_____no_output_____"
],
[
"### 1-4 반환(return)문",
"_____no_output_____"
],
[
"- 인수 없이 return 문을 사용하면 실제로는 None 객체가 전달된다.\n - None 객체: 파이썬 내장 객체로서 아무 값도 없음을 나타내기 위한 객체",
"_____no_output_____"
]
],
[
[
"def nothing():\n return\n\nprint(nothing())",
"None\n"
]
],
[
[
"- return문 없이 리턴하기",
"_____no_output_____"
]
],
[
[
"def print_menu():\n print('1. Snack')\n print('2. Snake')\n print('3. Snick')\n \nprint_menu()",
"1. Snack\n2. Snake\n3. Snick\n"
]
],
[
[
"- return문이 없는 함수라 해도, 실제로는 None 객체가 리턴됨",
"_____no_output_____"
]
],
[
[
"a = print_menu()\n\nprint(a)",
"1. Snack\n2. Snake\n3. Snick\nNone\n"
]
],
[
[
"- 한 개의 값을 리턴할 때",
"_____no_output_____"
]
],
[
[
"def abs_function(x):\n if x < 0 : return -x\n return x\n\nprint(abs_function(-10))",
"10\n"
]
],
[
[
"- 두 개 이상의 값을 리턴할 때",
"_____no_output_____"
]
],
[
[
"def swap(x, y):\n return y, x # 튜플로 리턴된다.\n\na = 10\nb = 20\nprint(a, b)\nprint()\n\na, b = swap(a, b) # 결과적으로 a, b = b, a와 동일\nprint(a, b)\nprint()\n\na = 10\nb = 20\nx = swap(a, b)\nprint(x[0], x[1]) # 하나의 이름으로 튜플을 받아서 처리할 수 도있다.",
"10 20\n\n20 10\n\n20 10\n"
]
],
[
[
"- 새로운 리스트를 리턴하는 함수의 예\n - 문자열 리스트를 받아서 각 문자열의 길이 정보를 지닌 리스트를 리턴 ",
"_____no_output_____"
]
],
[
[
"def length_list(l):\n res = []\n for el in l:\n res.append(len(el))\n return res\n\nl = ['python', 'pyson', 'pythong', 'pydon']\nprint(length_list(l))",
"[6, 5, 7, 5]\n"
],
[
"l = ['python', 'pyson', 'pythong', 'pydon']\nprint([len(s) for s in l])",
"[6, 5, 7, 5]\n"
]
],
[
[
"### 1-5 함수 인자에 대한 동적인 자료형 결정\n- 파이썬에서는 모든 객체는 동적으로 (실행 시간에) 그 타입이 결정된다.\n- 그러므로, 함수 인자는 함수가 호출되는 순간 해당 인자에 전달되는 객체에 따라 그 타입이 결정된다.\n - 함수 몸체 내에서 사용되는 여러가지 연산자들은 함수 호출시에 결정된 객체 타입에 맞게 실행된다.",
"_____no_output_____"
]
],
[
[
"def add(a, b):\n return a + b\n\nc = add(1, 3.4)\nd = add('dynamic', 'typing')\ne = add(['list'], ['and', 'list'])\nprint(c)\nprint(d)\nprint(e)",
"4.4\ndynamictyping\n['list', 'and', 'list']\n"
]
],
[
[
"***\n## 2 함수 인수 처리\n***",
"_____no_output_____"
],
[
"### 2-1 기본 인수 값\n- 기본 인수 값\n - 함수를 호출할 때 인수를 넘겨주지 않아도 인수가 기본적으로 가지는 값",
"_____no_output_____"
]
],
[
[
"def incr(a, step=1):\n return a + step\n\nb = 1\nb = incr(b) # 1 증가\nprint(b)\n\nb = incr(b, 10) # 10 증가\nprint(b)",
"2\n12\n"
]
],
[
[
"- [주의] 함수 정의를 할 때 기본 값을 지닌 인수 뒤에 일반적인 인수가 올 수 없음",
"_____no_output_____"
]
],
[
[
"def incr(step=1, a):\n return a + step",
"_____no_output_____"
]
],
[
[
"- 함수 정의 시에 여러 개의 기본 인수 값 정의 가능",
"_____no_output_____"
]
],
[
[
"def incr(a, step=1, step2=10):\n return a + step + step2\n\nprint(incr(10))",
"21\n"
]
],
[
[
"### 2-2 키워드 인수\n- 키워드 인수\n - 인수 값 전달 시에 인수 이름과 함께 값을 전달하는 방식을 일컫는다.",
"_____no_output_____"
]
],
[
[
"def area(height, width):\n return height * width\n\n#순서가 아닌 이름으로 값이 전달\na = area(height='height string ', width=3)\nprint(a)\n\nb = area(width=20, height=10)\nprint(b)",
"height string height string height string \n200\n"
]
],
[
[
"- 함수를 호출 할 때에 키워드 인수는 마지막에 놓여져야 한다.",
"_____no_output_____"
]
],
[
[
"print(area(20, width=5))",
"100\n"
]
],
[
[
"- [주의] 함수 호출시에 키워드 인수 뒤에 일반 인수 값이 올 수 없다. ",
"_____no_output_____"
]
],
[
[
"area(width=5, 20)\n# 기존: SyntaxError: non-keyword arg after keyword arg",
"_____no_output_____"
]
],
[
[
"- 기본 인수값 및 키워드 인수의 혼용",
"_____no_output_____"
]
],
[
[
"def incr(a, step=1, step2=10, step3=100):\n return a + step + step2 + step3\n\nprint(incr(10, 2, step2=100))",
"212\n"
]
],
[
[
"- 함수 호출 시에 키워드 인수 뒤에 일반 인수 값이 오면 에러 ",
"_____no_output_____"
]
],
[
[
"def incr(a, step=1, step2=10, step3=100):\n return a + step + step2 + step3\n\nprint(incr(10, 2, step2=100, 200))\n# 기존: SyntaxError: non-keyword arg after keyword arg",
"_____no_output_____"
],
[
"def incr(a, step=1, step2=10, step3=100):\n return a + step + step2 + step3\n\nprint(incr(10, 2, step2=100, step3=200))",
"312\n"
]
],
[
[
"### 2-3 가변 인수 리스트",
"_____no_output_____"
],
[
"- 함수 정의시에 일반적인 인수 선언 뒤에 <code>*args</code> 형식의 인수로 가변 인수를 선언할 수 있음\n- The special syntax <code>*args</code> in function definitions in python is used to pass a variable number of arguments to a function.",
"_____no_output_____"
]
],
[
[
"def myFun(*args):\n print(type(args), args)\n print()\n for arg in args:\n print(arg) \n \nmyFun('Hello', 'Welcome', 'to', 'GeeksforGeeks')",
"<class 'tuple'> ('Hello', 'Welcome', 'to', 'GeeksforGeeks')\n\nHello\nWelcome\nto\nGeeksforGeeks\n"
]
],
[
[
"- 함수 호출시 넣어주는 인수 값들 중 일반 인수에 할당되는 값을 제외한 나머지 값들만을 지닌 튜플 객체가 할당된다.",
"_____no_output_____"
]
],
[
[
"def varg(a, *args):\n print(a, args)\n\nvarg(1)\nvarg(2, 3)\nvarg(2, 3, 4, 5, 6)",
"1 ()\n2 (3,)\n2 (3, 4, 5, 6)\n"
]
],
[
[
"- C언어의 printf문과 유사한 형태의 printf 정의 방법",
"_____no_output_____"
]
],
[
[
"def printf(format, *args):\n print(format % args)\n\nprintf(\"I've spent %d days and %d night to do this\", 6, 5)",
"I've spent 6 days and 5 night to do this\n"
]
],
[
[
"- The special syntax ****kwargs** in function definitions in python is used to pass a keyworded, variable-length argument list. ",
"_____no_output_____"
]
],
[
[
"def myFun(**kwargs):\n print(type(kwargs), kwargs)\n print()\n for key, value in kwargs.items(): \n print(\"{0}: {1}\".format(key, value)) \n \nmyFun(first ='Geeks', mid ='for', last='Geeks') ",
"<class 'dict'> {'first': 'Geeks', 'mid': 'for', 'last': 'Geeks'}\n\nfirst: Geeks\nmid: for\nlast: Geeks\n"
],
[
"def myFun(arg1, **kwargs): \n for key, value in kwargs.items(): \n print (\"{0} - {1}: {2}\".format(arg1, key, value)) \n \nmyFun(\"Hi\", first ='Geeks', mid ='for', last='Geeks') ",
"Hi - first: Geeks\nHi - mid: for\nHi - last: Geeks\n"
]
],
[
[
"### 2-5 튜플 인수와 사전 인수로 함수 호출하기\n- 함수 호출에 사용될 인수값들이 튜플에 있다면 \"<code>*튜플변수</code>\"를 이용하여 함수 호출이 가능",
"_____no_output_____"
]
],
[
[
"def h(a, b, c):\n print(a, b, c)\n \nargs = (1, 2, 3)\nh(*args)",
"1 2 3\n"
]
],
[
[
"- 함수 호출에 사용될 인수값들이 사전에 있다면 \"<code>**사전변수</code>\"를 이용하여 함수 호출이 가능",
"_____no_output_____"
]
],
[
[
"def h(a, b, c):\n print(a, b, c)\n \ndargs = {'aa':1, 'bb':2, 'cc':3}\nh(*dargs)\n\ndargs = {'a':1, 'b':2, 'c':3}\nh(**dargs)",
"aa bb cc\n1 2 3\n"
],
[
"def myFun(arg1, arg2, arg3): \n print(\"arg1:\", arg1) \n print(\"arg2:\", arg2) \n print(\"arg3:\", arg3) \n \nargs = (\"Geeks\", \"for\", \"Geeks\") \nmyFun(*args) \n \nkwargs = {\"arg1\" : \"Geeks\", \"arg2\" : \"for\", \"arg3\" : \"Geeks\"} \nmyFun(**kwargs)",
"arg1: Geeks\narg2: for\narg3: Geeks\narg1: Geeks\narg2: for\narg3: Geeks\n"
],
[
"def function(**arg):\n for i in arg:\n print (i, arg[i])\n\nfunction(a=1, b=2, c=3, d=4)",
"a 1\nb 2\nc 3\nd 4\n"
]
],
[
[
"- Asterisks in Python: what they are and how to use them\n - https://treyhunner.com/2018/10/asterisks-in-python-what-they-are-and-how-to-use-them/",
"_____no_output_____"
]
],
[
[
"a, *b = [1, 2, 3, 4]\nprint(a)\nprint(b)",
"1\n[2, 3, 4]\n"
],
[
"def func(a, *b):\n print(a, b)\n \nfunc(1, 2, 3, 4)",
"1 (2, 3, 4)\n"
],
[
"numbers = [2, 1, 3, 4, 7]\nmore_numbers = [*numbers, 11, 18]\nprint(more_numbers)",
"[2, 1, 3, 4, 7, 11, 18]\n"
],
[
"print(*more_numbers)",
"2 1 3 4 7 11 18\n"
],
[
"fruits = ['lemon', 'pear', 'watermelon', 'tomato']\nprint(fruits[0], fruits[1], fruits[2], fruits[3])\n\nprint(*fruits)",
"lemon pear watermelon tomato\nlemon pear watermelon tomato\n"
],
[
"def transpose_list(list_of_lists):\n return [list(row) for row in zip(*list_of_lists)]\n\nt_l = transpose_list([[1, 4, 7], [2, 5, 8], [3, 6, 9]])\nprint(t_l)",
"[[1, 2, 3], [4, 5, 6], [7, 8, 9]]\n"
],
[
"date_info = {'year': \"2020\", 'month': \"01\", 'day': \"01\"}\n\nfilename = \"{year}-{month}-{day}.txt\".format(**date_info)\n\nprint(filename)",
"2020-01-01.txt\n"
],
[
"fruits = ['lemon', 'pear', 'watermelon', 'tomato']\nnumbers = [2, 1, 3, 4, 7]\nprint(*numbers, *fruits)",
"2 1 3 4 7 lemon pear watermelon tomato\n"
],
[
"date_info = {'year': \"2020\", 'month': \"01\", 'day': \"01\"}\ntrack_info = {'artist': \"Beethoven\", 'title': 'Symphony No 5'}\n\nfilename = \"{year}-{month}-{day}-{artist}-{title}.txt\".format(\n **date_info,\n **track_info,\n)\n\nprint(filename)",
"2020-01-01-Beethoven-Symphony No 5.txt\n"
],
[
"from random import randint\n\ndef roll(*dice):\n print(type(dice), dice)\n return sum([randint(1, die) for die in dice])\n\nprint(roll(20))\n\nprint(roll(6, 6))\n\nprint(roll(6, 6, 6))",
"<class 'tuple'> (20,)\n17\n<class 'tuple'> (6, 6)\n10\n<class 'tuple'> (6, 6, 6)\n12\n"
],
[
"def tag(tag_name, **attributes):\n attribute_list = [\"{0}='{1}'\".format(name, value) for name, value in attributes.items()]\n return \"<{0} \".format(tag_name) + ' '.join(attribute_list) + \">\"\n\nprint(tag('a', href=\"http://treyhunner.com\"))\n\nprint(tag('img', height=20, width=40, src=\"face.jpg\"))",
"<a href='http://treyhunner.com'>\n<img height='20' width='40' src='face.jpg'>\n"
]
],
[
[
"<p style='text-align: right;'>참고 문헌: 파이썬(열혈강의)(개정판 VER.2), 이강성, FreeLec, 2005년 8월 29일</p>",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
]
] |
4ace3348ea4b1f75cab868edc5b450d814e18add
| 212,861 |
ipynb
|
Jupyter Notebook
|
titanic/Titanic + feature engineering + turning xgboost.ipynb
|
wutienyang/kaggle_competitions
|
d06a074b9e691328d6111797d391df07ed4ee9ee
|
[
"MIT"
] | null | null | null |
titanic/Titanic + feature engineering + turning xgboost.ipynb
|
wutienyang/kaggle_competitions
|
d06a074b9e691328d6111797d391df07ed4ee9ee
|
[
"MIT"
] | null | null | null |
titanic/Titanic + feature engineering + turning xgboost.ipynb
|
wutienyang/kaggle_competitions
|
d06a074b9e691328d6111797d391df07ed4ee9ee
|
[
"MIT"
] | null | null | null | 91.789996 | 31,682 | 0.775304 |
[
[
[
"# Imports\nimport pandas as pd\nimport numpy as np\n\n# machine learning\nfrom sklearn import svm\nfrom sklearn.ensemble import VotingClassifier\nfrom sklearn.ensemble import GradientBoostingClassifier\nfrom sklearn.ensemble import AdaBoostClassifier\nfrom sklearn.model_selection import cross_val_score\nfrom sklearn.ensemble import RandomForestClassifier\nfrom sklearn import preprocessing\n\n# xgboost\nimport xgboost as xgb\n\n# matplotlib\nimport matplotlib.pyplot as plt\n%matplotlib inline",
"_____no_output_____"
],
[
"# 自定義的function\n\n# 算 accuracy, precision, recall\ndef performance(clf, X_train, Y_train, cv_num = 4):\n scores = cross_val_score(clf, X_train, Y_train, cv=cv_num , scoring='precision')\n print \"precision is {}\".format(scores.mean())\n \n scores = cross_val_score(clf, X_train, Y_train, cv=cv_num , scoring='recall')\n print \"recall is {}\".format(scores.mean())\n\n scores = cross_val_score(clf, X_train, Y_train, cv=cv_num , scoring='accuracy')\n print \"accuracy is {}\".format(scores.mean())",
"_____no_output_____"
],
[
"# get titanic & test csv files as a DataFrame\ntrain = pd.read_csv(\"/Users/wy/notebook/kaggle_competitions/titanic/train.csv\")\ntest = pd.read_csv(\"/Users/wy/notebook/kaggle_competitions/titanic/test.csv\")\ntest_passengerId = test['PassengerId']",
"_____no_output_____"
],
[
"train.info()\nprint \"--------------\"\ntest.info()",
"<class 'pandas.core.frame.DataFrame'>\nRangeIndex: 891 entries, 0 to 890\nData columns (total 12 columns):\nPassengerId 891 non-null int64\nSurvived 891 non-null int64\nPclass 891 non-null int64\nName 891 non-null object\nSex 891 non-null object\nAge 714 non-null float64\nSibSp 891 non-null int64\nParch 891 non-null int64\nTicket 891 non-null object\nFare 891 non-null float64\nCabin 204 non-null object\nEmbarked 889 non-null object\ndtypes: float64(2), int64(5), object(5)\nmemory usage: 83.6+ KB\n--------------\n<class 'pandas.core.frame.DataFrame'>\nRangeIndex: 418 entries, 0 to 417\nData columns (total 11 columns):\nPassengerId 418 non-null int64\nPclass 418 non-null int64\nName 418 non-null object\nSex 418 non-null object\nAge 332 non-null float64\nSibSp 418 non-null int64\nParch 418 non-null int64\nTicket 418 non-null object\nFare 417 non-null float64\nCabin 91 non-null object\nEmbarked 418 non-null object\ndtypes: float64(2), int64(4), object(5)\nmemory usage: 36.0+ KB\n"
],
[
"#Combine into data:\ntrain['source']= 'train'\ntest['source'] = 'test'\ndata=pd.concat([train, test],ignore_index=True)\ndata.shape",
"_____no_output_____"
],
[
"# 稍微看一下 data長怎樣\ndata.head()",
"_____no_output_____"
]
],
[
[
"##Check missing:",
"_____no_output_____"
]
],
[
[
"data.apply(lambda x: sum(x.isnull()))",
"_____no_output_____"
]
],
[
[
"##Look at categories of all object variables:",
"_____no_output_____"
]
],
[
[
"var = ['Sex','Ticket','Cabin','Embarked']\nfor v in var:\n print '\\nFrequency count for variable %s'%v\n print data[v].value_counts()",
"\nFrequency count for variable Sex\nmale 843\nfemale 466\nName: Sex, dtype: int64\n\nFrequency count for variable Ticket\nCA. 2343 11\nCA 2144 8\n1601 8\nS.O.C. 14879 7\nPC 17608 7\n347077 7\n347082 7\n3101295 7\n347088 6\n113781 6\n382652 6\n19950 6\n16966 5\nPC 17757 5\nW./C. 6608 5\n349909 5\n113503 5\n220845 5\n4133 5\nPC 17760 4\nPC 17755 4\n24160 4\nLINE 4\n17421 4\n230136 4\n12749 4\nSC/Paris 2123 4\n113760 4\n36928 4\nC.A. 34651 4\n ..\nC.A. 24579 1\n347062 1\n323951 1\n233478 1\n315088 1\n248723 1\n347079 1\n248726 1\n347074 1\n347075 1\n347076 1\nC.A. 18723 1\n250652 1\n347073 1\n330935 1\n248659 1\n330932 1\n330931 1\n233866 1\n345498 1\n362316 1\n382653 1\n382651 1\n382650 1\n373450 1\n364859 1\n240261 1\n31028 1\n3101276 1\nSCO/W 1585 1\nName: Ticket, dtype: int64\n\nFrequency count for variable Cabin\nC23 C25 C27 6\nB57 B59 B63 B66 5\nG6 5\nD 4\nB96 B98 4\nF2 4\nF4 4\nC78 4\nF33 4\nC22 C26 4\nB58 B60 3\nE101 3\nB51 B53 B55 3\nC101 3\nA34 3\nE34 3\nC68 2\nC7 2\nC62 C64 2\nE25 2\nC92 2\nC93 2\nB77 2\nB78 2\nE24 2\nE50 2\nB35 2\nD10 D12 2\nC2 2\nE121 2\n ..\nA23 1\nA26 1\nA29 1\nB24 1\nC105 1\nB4 1\nC128 1\nD45 1\nD6 1\nB3 1\nC53 1\nE58 1\nD34 1\nB102 1\nA32 1\nE17 1\nA16 1\nF E69 1\nD38 1\nE39 E41 1\nA14 1\nE52 1\nC91 1\nB73 1\nB39 1\nC95 1\nC99 1\nB37 1\nB30 1\nA10 1\nName: Cabin, dtype: int64\n\nFrequency count for variable Embarked\nS 914\nC 270\nQ 123\nName: Embarked, dtype: int64\n"
]
],
[
[
"##Missing values on Embarked",
"_____no_output_____"
]
],
[
[
"fig = plt.figure(figsize=(8, 5))\nax = fig.add_subplot(111)\nax = data.boxplot(column='Fare', by=['Embarked','Pclass'], ax=ax)\nplt.axhline(y=80, color='green')\nax.set_title('', y=1.1)\n\ndata[data.Embarked.isnull()][['Fare', 'Pclass', 'Embarked']]",
"_____no_output_____"
]
],
[
[
"####From the above boxplot, we should replace NA with C because most people who had Pclass 1 and Fare 80 would be Embarked C",
"_____no_output_____"
]
],
[
[
"data['Embarked'].fillna('C', inplace=True)",
"_____no_output_____"
]
],
[
[
"##Missing values on Fare",
"_____no_output_____"
]
],
[
[
"fig = plt.figure(figsize=(8, 5))\nax = fig.add_subplot(111)\ndata[(data.Pclass==3)&(data.Embarked=='S')].Fare.hist(bins=100, ax=ax)\ndata[data.Fare.isnull()][['Pclass', 'Fare', 'Embarked']]\nplt.xlabel('Fare')\nplt.ylabel('Frequency')\nplt.title('Histogram of Fare, Plcass 3 and Embarked S')\n\ndata[data.Fare.isnull()][['Pclass', 'Fare', 'Embarked']]",
"_____no_output_____"
],
[
"print (\"The top 5 most common value of Fare\")\ndata[(data.Pclass==3)&(data.Embarked=='S')].Fare.value_counts().head()",
"The top 5 most common value of Fare\n"
],
[
"data['Fare'].fillna(8.05, inplace=True)",
"_____no_output_____"
]
],
[
[
"###Replace the missing value of Cabin with U0",
"_____no_output_____"
]
],
[
[
"data['Cabin_Missing'] = data['Cabin'].apply(lambda x: 1 if pd.isnull(x) else 0)\ndata['Cabin'].fillna('U0', inplace=True)",
"_____no_output_____"
]
],
[
[
"##Feature Engineering",
"_____no_output_____"
],
[
"###Create a feature, Names, to store the length of words in name.",
"_____no_output_____"
]
],
[
[
"import re\ndata['Names'] = data['Name'].map(lambda x: len(re.split(' ', x)))",
"_____no_output_____"
]
],
[
[
"###Create a feature, Title.",
"_____no_output_____"
]
],
[
[
"title = data['Name'].map(lambda x: re.compile(', (.*?)\\.').findall(x)[0])\ntitle[title=='Mme'] = 'Mrs'\ntitle[title.isin(['Ms','Mlle'])] = 'Miss'\ntitle[title.isin(['Don', 'Jonkheer'])] = 'Sir'\ntitle[title.isin(['Dona', 'Lady', 'the Countess'])] = 'Lady'\ntitle[title.isin(['Capt', 'Col', 'Major', 'Dr', 'Officer', 'Rev'])] = 'Officer'\ndata['Title'] = title\ndel title",
"_____no_output_____"
]
],
[
[
"###Create a feature, Deck. It may represents the socioeconomic status.",
"_____no_output_____"
]
],
[
[
"deck = data['Cabin'].map( lambda x : re.compile(\"([a-zA-Z]+)\").search(x).group())\ndata['Deck'] = deck\ndel deck",
"_____no_output_____"
],
[
"data.head()",
"_____no_output_____"
]
],
[
[
"###Create a feature, Room. It may represents the geo lacation.",
"_____no_output_____"
]
],
[
[
"checker = re.compile(\"([0-9]+)\")\ndef roomNum(x):\n nums = checker.search(x)\n if nums:\n return int(nums.group())+1\n else:\n return 1\nrooms = data['Cabin'].map(lambda x: roomNum(x))",
"_____no_output_____"
],
[
"data['Cabin_Room'] = rooms / rooms.sum()\ndel checker, roomNum",
"_____no_output_____"
],
[
"data.head()",
"_____no_output_____"
]
],
[
[
"###Create a feature, Group_num. It may represents the size of family.",
"_____no_output_____"
]
],
[
[
"data['Group_num'] = data['Parch'] + data['SibSp'] + 1",
"_____no_output_____"
]
],
[
[
"###Create a feature, Group_size. When the size is between 2 and 4, more people are survived.",
"_____no_output_____"
]
],
[
[
"def groupSize(x):\n if x > 4 :\n return 'L'\n elif x == 1 :\n return 'S'\n else:\n return 'M'\ngroup_size = data['Group_num'].map(lambda x: groupSize(x))\ndata['Group_size'] = group_size",
"_____no_output_____"
],
[
"data.head()",
"_____no_output_____"
],
[
"data.dtypes",
"_____no_output_____"
]
],
[
[
"###Normalized the fare.",
"_____no_output_____"
]
],
[
[
"from sklearn.preprocessing import StandardScaler\nscaler = StandardScaler()\ndata['Nor_Fare'] = pd.Series(scaler.fit_transform(data['Fare'].values.reshape(-1,1)).reshape(-1), index=data.index)",
"_____no_output_____"
]
],
[
[
"###Numerical Coding:",
"_____no_output_____"
]
],
[
[
"from sklearn.preprocessing import LabelEncoder\nle = LabelEncoder()\nvar_to_encode = ['Embarked','Sex','Deck','Group_size','Title']\nfor col in var_to_encode:\n data[col] = le.fit_transform(data[col])",
"_____no_output_____"
]
],
[
[
"###One-Hot Coding",
"_____no_output_____"
]
],
[
[
"data = pd.get_dummies(data, columns=var_to_encode)\ndata.columns",
"_____no_output_____"
]
],
[
[
"###Predict Age",
"_____no_output_____"
]
],
[
[
"label_y = data[data['source'] == 'train']['Survived']",
"_____no_output_____"
],
[
"from sklearn.model_selection import train_test_split\ndata.drop(labels=['PassengerId', 'Name', 'Cabin', 'Survived', 'Ticket', 'Fare'], axis=1, inplace=True)\n\nX = data[data['Age'].notnull()].drop(['Age','source'], axis=1)\ny = data[data['Age'].notnull()].Age\nX_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)\n\nfrom sklearn.metrics import mean_absolute_error\nfrom sklearn.metrics import make_scorer\nfrom sklearn.model_selection import GridSearchCV\n\ndef get_model(estimator, parameters, X_train, y_train, scoring): \n model = GridSearchCV(estimator, param_grid=parameters, scoring=scoring)\n model.fit(X_train, y_train)\n return model.best_estimator_\n\nimport xgboost as xgb\n\nXGB = xgb.XGBRegressor(max_depth=4, seed= 42)\nscoring = make_scorer(mean_absolute_error, greater_is_better=False)\nparameters = {'reg_alpha':np.linspace(0.1,1.0,5), 'reg_lambda': np.linspace(1.0,3.0,5)}\nreg_xgb = get_model(XGB, parameters, X_train, y_train, scoring)\nprint (reg_xgb)\n\nprint (\"Mean absolute error of test data: {}\".format(mean_absolute_error(y_test, reg_xgb.predict(X_test))))",
"XGBRegressor(base_score=0.5, colsample_bylevel=1, colsample_bytree=1, gamma=0,\n learning_rate=0.1, max_delta_step=0, max_depth=4,\n min_child_weight=1, missing=None, n_estimators=100, nthread=-1,\n objective='reg:linear', reg_alpha=0.77500000000000002,\n reg_lambda=3.0, scale_pos_weight=1, seed=42, silent=True,\n subsample=1)\nMean absolute error of test data: 7.74417822386\n"
],
[
"fig = plt.figure(figsize=(15, 6))\nalpha = 0.5\ndata['Age'].value_counts().plot(kind='density', color='#FA2379', label='Before', alpha=alpha)\n\npred = reg_xgb.predict(data[data['Age'].isnull()].drop(['Age','source'], axis=1))\ndata.set_value(data['Age'].isnull(), 'Age', pred)\n\ndata['Age'].value_counts().plot(kind='density', label='After', alpha=alpha)\nplt.xlabel('Age')\nplt.title(\"What's the distribution of Age after predicting?\" )\nplt.legend(loc='best')\nplt.grid()",
"_____no_output_____"
]
],
[
[
"###Separate train & test:",
"_____no_output_____"
]
],
[
[
"# label_y\ntrain = data.loc[data['source']=='train']\ntest = data.loc[data['source']=='test']\n\ntrain.drop('source',axis=1,inplace=True)\ntest.drop('source',axis=1,inplace=True)",
"/Users/wy/anaconda/envs/condapy2.7/lib/python2.7/site-packages/IPython/kernel/__main__.py:5: SettingWithCopyWarning: \nA value is trying to be set on a copy of a slice from a DataFrame\n\nSee the caveats in the documentation: http://pandas.pydata.org/pandas-docs/stable/indexing.html#indexing-view-versus-copy\n/Users/wy/anaconda/envs/condapy2.7/lib/python2.7/site-packages/IPython/kernel/__main__.py:6: SettingWithCopyWarning: \nA value is trying to be set on a copy of a slice from a DataFrame\n\nSee the caveats in the documentation: http://pandas.pydata.org/pandas-docs/stable/indexing.html#indexing-view-versus-copy\n"
]
],
[
[
"##Build Model",
"_____no_output_____"
]
],
[
[
"def modelfit(alg, train, label_y,useTrainCV=True, cv_folds=5, early_stopping_rounds=50):\n \n if useTrainCV:\n xgb_param = alg.get_xgb_params()\n xgtrain = xgb.DMatrix(train, label=label_y)\n \n cvresult = xgb.cv(xgb_param, xgtrain, num_boost_round=alg.get_params()['n_estimators'], nfold=cv_folds,\n metrics='auc', early_stopping_rounds=early_stopping_rounds)\n alg.set_params(n_estimators=cvresult.shape[0])\n \n #Fit the algorithm on the data\n alg.fit(train, label_y,eval_metric='auc')\n\n #Predict training set:\n dtrain_predictions = alg.predict(train)\n dtrain_predprob = alg.predict_proba(train)[:,1]\n \n #Print model report:\n print \"\\nModel Report\"\n print \"Accuracy : %.4g\" % metrics.accuracy_score(label_y, dtrain_predictions)\n print \"AUC Score (Train): %f\" % metrics.roc_auc_score(label_y, dtrain_predprob)\n \n feat_imp = pd.Series(alg.booster().get_fscore()).sort_values(ascending=False)\n feat_imp.plot(kind='bar', title='Feature Importances')\n plt.ylabel('Feature Importance Score')",
"_____no_output_____"
],
[
"from xgboost.sklearn import XGBClassifier\nfrom sklearn import cross_validation, metrics\n\nxgb1 = XGBClassifier(\n learning_rate =0.1,\n n_estimators=1000,\n max_depth=5,\n min_child_weight=1,\n gamma=0,\n subsample=0.8,\n colsample_bytree=0.8,\n objective= 'binary:logistic',\n nthread=4,\n# scale_pos_weight=1,\n seed=27)\nmodelfit(xgb1, train, label_y)",
"\nModel Report\nAccuracy : 0.9046\nAUC Score (Train): 0.965328\n"
],
[
"#Grid seach on subsample and max_features\n#Choose all predictors except target & IDcols\nparam_test1 = {\n 'max_depth':range(3,10,2),\n 'min_child_weight':range(1,6,2)\n}\ngsearch1 = GridSearchCV(estimator = XGBClassifier( learning_rate =0.1, n_estimators=140, max_depth=5,\n min_child_weight=1, gamma=0, subsample=0.8, colsample_bytree=0.8,\n objective= 'binary:logistic', nthread=4, seed=27), \n param_grid = param_test1, scoring='roc_auc',n_jobs=4,iid=False, cv=5)\ngsearch1.fit(train,label_y)\ngsearch1.grid_scores_, gsearch1.best_params_, gsearch1.best_score_",
"/Users/wy/anaconda/envs/condapy2.7/lib/python2.7/site-packages/sklearn/model_selection/_search.py:662: DeprecationWarning: The grid_scores_ attribute was deprecated in version 0.18 in favor of the more elaborate cv_results_ attribute. The grid_scores_ attribute will not be available from 0.20\n DeprecationWarning)\n"
],
[
"#Grid seach on subsample and max_features\n#Choose all predictors except target & IDcols\nparam_test2 = {\n 'max_depth':[8,9,10,11,12],\n 'min_child_weight':[4,5,6]\n}\ngsearch2 = GridSearchCV(estimator = XGBClassifier( learning_rate=0.1, n_estimators=140, max_depth=5,\n min_child_weight=2, gamma=0, subsample=0.8, colsample_bytree=0.8,\n objective= 'binary:logistic', nthread=4,seed=27), \n param_grid = param_test2, scoring='roc_auc',n_jobs=4,iid=False, cv=5)\ngsearch2.fit(train,label_y)\ngsearch2.grid_scores_, gsearch2.best_params_, gsearch2.best_score_",
"/Users/wy/anaconda/envs/condapy2.7/lib/python2.7/site-packages/sklearn/model_selection/_search.py:662: DeprecationWarning: The grid_scores_ attribute was deprecated in version 0.18 in favor of the more elaborate cv_results_ attribute. The grid_scores_ attribute will not be available from 0.20\n DeprecationWarning)\n"
],
[
"#Grid seach on subsample and max_features\n#Choose all predictors except target & IDcols\nparam_test3 = {\n 'gamma':[i/10.0 for i in range(0,15)]\n}\ngsearch3 = GridSearchCV(estimator = XGBClassifier( learning_rate =0.1, n_estimators=140, max_depth=10,\n min_child_weight=5, gamma=0, subsample=0.8, colsample_bytree=0.8,\n objective= 'binary:logistic', nthread=4,seed=27), \n param_grid = param_test3, scoring='roc_auc',n_jobs=4,iid=False, cv=5)\ngsearch3.fit(train,label_y)\ngsearch3.grid_scores_, gsearch3.best_params_, gsearch3.best_score_",
"/Users/wy/anaconda/envs/condapy2.7/lib/python2.7/site-packages/sklearn/model_selection/_search.py:662: DeprecationWarning: The grid_scores_ attribute was deprecated in version 0.18 in favor of the more elaborate cv_results_ attribute. The grid_scores_ attribute will not be available from 0.20\n DeprecationWarning)\n"
],
[
"xgb2 = XGBClassifier(\n learning_rate =0.1,\n n_estimators=1000,\n max_depth=10,\n min_child_weight=5,\n gamma=0.9,\n subsample=0.8,\n colsample_bytree=0.8,\n objective= 'binary:logistic',\n nthread=4,\n seed=27)\nmodelfit(xgb2, train, label_y)",
"\nModel Report\nAccuracy : 0.8866\nAUC Score (Train): 0.943739\n"
],
[
"#Grid seach on subsample and max_features\n#Choose all predictors except target & IDcols\nparam_test4 = {\n 'subsample':[i/10.0 for i in range(6,10)],\n 'colsample_bytree':[i/10.0 for i in range(6,10)]\n}\ngsearch4 = GridSearchCV(estimator = XGBClassifier( learning_rate =0.1, n_estimators=177, max_depth=4,\n min_child_weight=6, gamma=0, subsample=0.8, colsample_bytree=0.8,\n objective= 'binary:logistic', nthread=4, scale_pos_weight=1,seed=27), \n param_grid = param_test4, scoring='roc_auc',n_jobs=4,iid=False, cv=5)\ngsearch4.fit(train,label_y)\ngsearch4.grid_scores_, gsearch4.best_params_, gsearch4.best_score_",
"/Users/wy/anaconda/envs/condapy2.7/lib/python2.7/site-packages/sklearn/model_selection/_search.py:662: DeprecationWarning: The grid_scores_ attribute was deprecated in version 0.18 in favor of the more elaborate cv_results_ attribute. The grid_scores_ attribute will not be available from 0.20\n DeprecationWarning)\n"
],
[
"#Grid seach on subsample and max_features\n#Choose all predictors except target & IDcols\nparam_test5 = {\n 'reg_alpha':[1e-5, 1e-2, 0.1, 1, 100]\n}\ngsearch5 = GridSearchCV(estimator = XGBClassifier( learning_rate =0.1, n_estimators=177, max_depth=4,\n min_child_weight=6, gamma=0.1, subsample=0.8, colsample_bytree=0.8,\n objective= 'binary:logistic', nthread=4, scale_pos_weight=1,seed=27), \n param_grid = param_test5, scoring='roc_auc',n_jobs=4,iid=False, cv=5)\ngsearch5.fit(train,label_y)\ngsearch5.grid_scores_, gsearch5.best_params_, gsearch5.best_score_",
"/Users/wy/anaconda/envs/condapy2.7/lib/python2.7/site-packages/sklearn/model_selection/_search.py:662: DeprecationWarning: The grid_scores_ attribute was deprecated in version 0.18 in favor of the more elaborate cv_results_ attribute. The grid_scores_ attribute will not be available from 0.20\n DeprecationWarning)\n"
],
[
"#Grid seach on subsample and max_features\n#Choose all predictors except target & IDcols\nparam_test6 = {\n 'reg_alpha':[0, 0.01, 0.05, 0.1, 0.16, 0.19]\n}\ngsearch6 = GridSearchCV(estimator = XGBClassifier( learning_rate =0.1, n_estimators=177, max_depth=4,\n min_child_weight=6, gamma=0.1, subsample=0.8, colsample_bytree=0.8,\n objective= 'binary:logistic', nthread=4, scale_pos_weight=1,seed=27), \n param_grid = param_test6, scoring='roc_auc',n_jobs=4,iid=False, cv=5)\ngsearch6.fit(train,label_y)\ngsearch6.grid_scores_, gsearch6.best_params_, gsearch6.best_score_",
"/Users/wy/anaconda/envs/condapy2.7/lib/python2.7/site-packages/sklearn/model_selection/_search.py:662: DeprecationWarning: The grid_scores_ attribute was deprecated in version 0.18 in favor of the more elaborate cv_results_ attribute. The grid_scores_ attribute will not be available from 0.20\n DeprecationWarning)\n"
],
[
"xgb3 = XGBClassifier(\n learning_rate =0.1,\n n_estimators=1000,\n max_depth=10,\n min_child_weight=5,\n gamma=0.9,\n subsample=0.7,\n colsample_bytree=0.7,\n reg_alpha=0.1,\n objective= 'binary:logistic',\n nthread=4,\n seed=27)\nmodelfit(xgb3, train, label_y)",
"\nModel Report\nAccuracy : 0.8866\nAUC Score (Train): 0.948585\n"
],
[
"xgb4 = XGBClassifier(\n learning_rate =0.01,\n n_estimators=5000,\n max_depth=10,\n min_child_weight=5,\n gamma=0.9,\n subsample=0.7,\n colsample_bytree=0.7,\n reg_alpha=0.1,\n objective= 'binary:logistic',\n nthread=4,\n scale_pos_weight=1,\n seed=27)\nmodelfit(xgb4, train, label_y)",
"\nModel Report\nAccuracy : 0.8462\nAUC Score (Train): 0.887035\n"
]
],
[
[
"##Make submission",
"_____no_output_____"
]
],
[
[
"test_predict = xgb4.predict(test)",
"_____no_output_____"
],
[
"submission = pd.DataFrame({\n \"PassengerId\": test_passengerId,\n \"Survived\": test_predict\n })",
"_____no_output_____"
],
[
"submission['Survived'] = submission['Survived'].astype('int64')",
"_____no_output_____"
],
[
"submission.to_csv('/Users/wy/Desktop/titanic_xgboost2.csv', index=False)",
"_____no_output_____"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
4ace33cdc7c252352bffb13bd399936520cde478
| 4,260 |
ipynb
|
Jupyter Notebook
|
tutorials/.ipynb_checkpoints/UsingDonutsHooks-checkpoint.ipynb
|
jmccormac01/Donuts
|
ba1dbb8f6f721e52008611a87bdc1b8900b5bd66
|
[
"Unlicense"
] | 14 |
2016-03-26T10:20:51.000Z
|
2020-03-29T21:07:06.000Z
|
tutorials/.ipynb_checkpoints/UsingDonutsHooks-checkpoint.ipynb
|
jmccormac01/DONUTS
|
ba1dbb8f6f721e52008611a87bdc1b8900b5bd66
|
[
"Unlicense"
] | 56 |
2016-03-26T10:52:06.000Z
|
2021-12-26T09:29:54.000Z
|
tutorials/.ipynb_checkpoints/UsingDonutsHooks-checkpoint.ipynb
|
jmccormac01/DONUTS
|
ba1dbb8f6f721e52008611a87bdc1b8900b5bd66
|
[
"Unlicense"
] | 8 |
2016-03-26T10:21:14.000Z
|
2019-12-10T15:56:37.000Z
| 27.843137 | 323 | 0.529812 |
[
[
[
"# Using pre/post construction hooks in Donuts\n\nThe Donuts ```Image``` class has 2 special functions. One that works on an image before the standard procedure and one that works on it afterwards. You can sub-class the ```Image``` class and assign your own functions to the hooks to add extra functionality to Donuts. Below is an example of a pre-construction hook. ",
"_____no_output_____"
]
],
[
[
"import os\nfrom astropy.io import fits\nimport donuts",
"_____no_output_____"
],
[
"os.chdir('/Users/jmcc/Desktop/NG2325-2844_803')\ntest_file = \"NG2325-2844_803_IMAGE80320190523073419.fits\"\n\n# open the test file so we can see the values\nwith fits.open(test_file) as ff:\n data = ff[0].data\n\nprint(data)",
"[[942 929 941 ... 945 946 941]\n [936 929 944 ... 934 937 934]\n [944 940 949 ... 937 940 929]\n ...\n [941 936 940 ... 942 946 936]\n [943 937 935 ... 937 937 943]\n [936 936 943 ... 937 944 931]]\n"
],
[
"# sub class the Image class and redefine the preconstruct_hook function\nclass NewImage(donuts.image.Image):\n def preconstruct_hook(self):\n \"\"\"\n Here we have access to the full raw_image, untrimmed or \n uncropped in any way. In this example we simply remove \n 100 counts from each pixel as a demonstration of the hook\n \"\"\"\n self.raw_image = self.raw_image - 100.\n\n# Instantiate the new Image class and pass it to donuts\n# All the other class methods ae inherited as normal\ncustom_image_class = NewImage\n\n# now we initalise donuts using the modified Image class with our own special feature\nd = donuts.Donuts(test_file, image_class=custom_image_class)\n\n# now we check the raw_image data and it should show 100 counts less than above\nd.reference_image.raw_image",
"WARNING: Exposure time keyword \"EXPTIME\" not found, assuming 1.0 [donuts.image]\n"
],
[
"# finally we measure the shift of the image wrt itself, which should be 0 pixels\nshift = d.measure_shift(test_file)\nprint(shift.x, shift.y)",
"WARNING: Exposure time keyword \"EXPTIME\" not found, assuming 1.0 [donuts.image]\n"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
4ace344eb82a120dec6bc38837f063561ab40945
| 52,504 |
ipynb
|
Jupyter Notebook
|
data/KNN.ipynb
|
kumailZaidi12/Face-Recognition
|
263f60d27fae48e25af47b674da82f59364f0046
|
[
"MIT"
] | 2 |
2018-10-12T15:23:52.000Z
|
2018-10-22T22:01:49.000Z
|
data/KNN.ipynb
|
kumailZaidi12/Face-Recognition
|
263f60d27fae48e25af47b674da82f59364f0046
|
[
"MIT"
] | null | null | null |
data/KNN.ipynb
|
kumailZaidi12/Face-Recognition
|
263f60d27fae48e25af47b674da82f59364f0046
|
[
"MIT"
] | 2 |
2018-10-22T20:34:05.000Z
|
2018-10-23T18:00:57.000Z
| 218.766667 | 41,988 | 0.923739 |
[
[
[
"import numpy as np\nfrom matplotlib import pyplot as plt",
"_____no_output_____"
],
[
"dist = np.random.normal(2,.5,1000)\ndist2 = np.random.normal(0,1,1000)\n# print(dist.shape)",
"_____no_output_____"
],
[
"plt.hist(dist,100)\nplt.hist(dist2,100,alpha=0.5)\nplt.show()",
"_____no_output_____"
]
],
[
[
"# Multivarate case",
"_____no_output_____"
]
],
[
[
"monkey_data= np.random.multivariate_normal([1,2],[[1,0.5],[0.5,1]],1000)\nchimp_data= np.random.multivariate_normal([4,4],[[1,0.5],[0.5,1]],1000)\n\nprint(monkey_data.shape)",
"(1000, 2)\n"
],
[
"plt.scatter(monkey_data[:,0],monkey_data[:,1],label='Monkey')\nplt.scatter(chimp_data[:,0],chimp_data[:,1],alpha=0.5,label='Chimpanzee')\n\n\nquery_x = np.array([3,2])\nplt.scatter(query_x[0],query_x[1],c='k',label='Query Point')\nplt.legend()\nplt.show()",
"_____no_output_____"
]
],
[
[
"#K-Nearest Nieghbours \n ",
"_____no_output_____"
]
],
[
[
"data = np.zeros((2000,3))\n\ndata[:1000,:-1] = monkey_data\ndata[1000:,:-1] = chimp_data\ndata[:1000,-1] = 0\ndata[1000:,-1] = 1\n\n\nprint(data.shape)",
"(2000, 3)\n"
],
[
"x_train = data[:,:-1]\ny_train = data[:,-1]\nprint(x_train.shape)\nprint(y_train.shape)\n",
"(2000, 2)\n(2000,)\n"
],
[
"def dist(x1,x2):\n return np.sqrt(((x1-x2)**2).sum())\n\n\ndef knn(x_train,y_train,query_x,k=5):\n dist_vals=[]\n m=x_train.shape[0]\n for ix in range(m):\n d=dist(query_x,x_train[ix])\n dist_vals.append((d,y_train[ix]))\n dist_vals = sorted(dist_vals)\n dist_vals=dist_vals[:k]\n \n y=np.array(dist_vals)\n t=np.unique(y[:,1],return_counts=True)\n index=t[1].argmax()\n prediction=t[0][index]\n \n print (prediction)\n ",
"_____no_output_____"
],
[
"knn(x_train,y_train,query_x)",
"1.0\n"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
4ace4b3dc5d20bde7b4464a3815e4ab224c60171
| 640,439 |
ipynb
|
Jupyter Notebook
|
notebooks/kmer_frequency/kmer_frequency_Bacillus.ipynb
|
susiegriggo/ProtvecHierachy
|
05508eb9bf7eb51dec1309694ad1c789b32f02e2
|
[
"MIT"
] | null | null | null |
notebooks/kmer_frequency/kmer_frequency_Bacillus.ipynb
|
susiegriggo/ProtvecHierachy
|
05508eb9bf7eb51dec1309694ad1c789b32f02e2
|
[
"MIT"
] | null | null | null |
notebooks/kmer_frequency/kmer_frequency_Bacillus.ipynb
|
susiegriggo/ProtvecHierachy
|
05508eb9bf7eb51dec1309694ad1c789b32f02e2
|
[
"MIT"
] | null | null | null | 1,404.471491 | 220,264 | 0.955285 |
[
[
[
"# Kmer frequency Bacillus\n\nGenerate code to embed Bacillus sequences by calculating kmer frequency\n\nimport to note that this requires biopython version 1.77. Alphabet was deprecated in 1.78 (September 2020). Alternatively we could not reduce the alphabet though the kmer frequency table is sparse so could be a computational nightmare. \n\nUsing the murphy10 reduced alphabet. There are other amino acid reduced alphabets could be tried as well https://biopython.org/docs/1.75/api/Bio.Alphabet.Reduced.html\n\nSequences containing 'X' have been excluded. The murphy10 alphabet has been used with options to try alphabets with a different number of amino acids. Sequeces longer than 1024 amino acids are also excluded. ",
"_____no_output_____"
]
],
[
[
"#imports \nimport numpy as np \nimport pandas as pd\nfrom Bio.Seq import Seq\nfrom Bio import Alphabet\nfrom Bio.Alphabet import Reduced\nimport itertools\nfrom Bio import SeqIO\nfrom sklearn.decomposition import PCA\nfrom sklearn.manifold import TSNE\nfrom sklearn.preprocessing import StandardScaler\nfrom matplotlib import pyplot as plt\nfrom collections import Counter\nimport seaborn as sns\nfrom sklearn.utils.extmath import randomized_svd \nfrom scipy.spatial import distance \nimport random",
"_____no_output_____"
],
[
"def seq_3mers(sequence): \n \"\"\"Takes a sequence to overlapping 3-mers\"\"\"\n \n seq_size = len(sequence) \n seq_3mers = list() #intialise list \n \n #iterate through sequence to obtain 3mers \n for i in range (1,seq_size-1):\n seq_3mers.append(sequence[i-1]+sequence[i]+sequence[i+1])\n \n return seq_3mers\n\ndef murphy10(seq_str): \n \"\"\"Takes an amino acid sequence using the standard 20 amino acid code to reduced 10 letter alphabet. \n This funcseqs_swiss_keystion requires biopython version 1.77 or lower. Input is a a string of amino acids\"\"\"\n \n #turn starting sequence into a sequence object \n intial_seq = Seq(seq_str, Alphabet.ProteinAlphabet())\n \n #intialise sequence object \n new_seq = Seq('', Alphabet.Reduced.Murphy10())\n \n #iterate through the letters in the sequence and convert to murphy10 \n for aa in intial_seq: \n new_seq += Alphabet.Reduced.murphy_10_tab[aa]\n \n return str(new_seq)\n\ndef seq_vector(seq, embedding): \n \"\"\"Embeds a sequence as a kmer frequency embedding\"\"\"\n \n #break the seq into kmers \n seq_kmers = seq_3mers(seq)\n \n #intialise a vector for the sequence to be embedded \n seq_vec = np.zeros(len(embedding))\n \n #iterate through the kmers in the sequence \n for kmer in seq_kmers: \n \n #add the kmer vector to make the sequence vector \n seq_vec += embedding[kmer]\n \n #divide the sequence by the number of kmers (number of kmer counts) (NOT SURE IF THIS IS CORRECT - PLAY AROUND WITH this)\n seq_vec = seq_vec/len(seq_kmers)\n \n return seq_vec \n\ndef embedkmers_seqs(seqs, embedding): \n \"\"\"Embed a list of sequences with a dataframe of kmer frequency\"\"\"\n \n #intialise an array to hold the embeddings\n embed_kmerfreq = np.zeros((len(seqs), len(embedding)))\n \n #iterate through the sequences \n for i in range(len(seqs)): \n #get the sequence \n seq = seqs[i]\n \n #get the vector\n seq_vec= seq_vector(seq, embedding)\n \n #add the sequnce vector to the embeddings matrix \n embed_kmerfreq[i] = seq_vec \n \n return embed_kmerfreq \n ",
"_____no_output_____"
],
[
"#import the embedding sequences \nembed_seqs_dict = SeqIO.index(\"../../sequences/bacillus_embeddingset.fa\", 'fasta')\nembed_seqs_keys = list(embed_seqs_dict.keys()) #gives md5 hashes of the sequences \nembed_seqs = [str(embed_seqs_dict.get(key).seq) for key in embed_seqs_keys]\n\n#get a random subset of 16763 sequences to embed (this was the number of sequences embedded for bacteroides)\nrandint = random.sample(range(len(embed_seqs)), 16763) \nembed_seqs_keys = embed_seqs_keys\n\n#determine which sequences contain the invalid character 'X' and remove them from the set of sequences to embed \nembed_seqs_containsX = ['X' not in seqs for seqs in embed_seqs]\nkeys_containsX = [embed_seqs_keys[i] for i in range(len(embed_seqs_keys)) if embed_seqs_containsX[i] == True]\nembed_seqs = [str(embed_seqs_dict.get(key).seq) for key in keys_containsX]\nembed_seqs_keys = keys_containsX \n\n#remove sequences which contain more than 1024 amino acids \nembed_seqs_1024 = [len(seqs)<= 1024 for seqs in embed_seqs]\nkeys_1024 = [embed_seqs_keys[i] for i in range(len(embed_seqs)) if embed_seqs_1024[i] == True]\nembed_seqs = [str(embed_seqs_dict.get(key).seq) for key in keys_1024]\nembed_seqs_keys = keys_1024\n\n#generate a list of all possible kmeres for the murphy10 alphabet \nmurphy10_sub = Alphabet.Reduced.murphy_10_tab \nmurphy10_l = set([d[1] for d in list(murphy10_sub.items())]) #list of letters in the murphy10 alphabet \nk = 3 #intialise the length of the kmer \nkmers = [''.join(kmer) for kmer in list(itertools.product(murphy10_l, repeat = k))]\n\n#intialise idnetity matrix size of kmers to represent the kmer embedding (each 1 is denotes a different kmer)\nkmerfreq = np.identity(len(kmers))\n#represent as a dataframe \nkmerfreq_df = pd.DataFrame(kmerfreq) \nkmerfreq_df.columns = kmers\nkmerfreq_df.index = kmers\n\n#convert the embedded sequences to murphy 10 \nembed_seqs_murphy10 = [murphy10(seq) for seq in embed_seqs]\n#embed the sequences \nembed_kmerfreq = embedkmers_seqs(embed_seqs_murphy10, kmerfreq_df)",
"_____no_output_____"
],
[
"#read in the ontology info and filter out the sequences we choose to ignore \n#need to read in some file which will map the sequences to the known hierachical classification (KEGG, Subsystems)\nsub_sys = pd.read_csv('../../subsystems_labels/bacillus.ids.tsv', sep = '\\t', header = None)\nsub_sys.columns = ['fig_ID', 'species', 'superclass', 'class', 'subclass', 'subsystem', 'product role_name']\n\n#get the file which takes the md5 hashes to the fig IDs \nmd5_fig = pd.read_csv('../../subsystems_labels/bacillus.md5.ids', sep = '\\t', header = None)\nmd5_fig.columns = ['md5', 'fig_ID']\n\n#assemble as a dictionary which takes a seqence key to the ontological represenation \nseqs_keys_figID = pd.concat([md5_fig[md5_fig['md5'] == key] for key in embed_seqs_keys])\n\n#convert the embedded sequences to murphy 10 \nembed_seqs_murphy10 = [murphy10(seq) for seq in embed_seqs]\nembed_kmerfreq = embedkmers_seqs(embed_seqs_murphy10, kmerfreq_df)",
"_____no_output_____"
],
[
"#make dictionaries for subclass/superclass but this time include the entry 'dual'\nfig2subclass = sub_sys[['fig_ID', 'subclass']].drop_duplicates()\nfig2subsystem = sub_sys[['fig_ID', 'subsystem']].drop_duplicates()\n\n#change fig_IDs which have more than one subclass to 'dual'\nduplicate_subclasses = pd.DataFrame(fig2subclass['fig_ID'].value_counts())\nduplicate_subclasses = duplicate_subclasses[duplicate_subclasses['fig_ID'] > 1].index.values\nduplicate_removed_subclasses = fig2subclass[~fig2subclass['fig_ID'].isin(duplicate_subclasses)]\n\ndual_vec = ['dual' for i in range(0,len(duplicate_subclasses))]\ndual_subclasses = pd.DataFrame({'fig_ID': duplicate_subclasses, 'subclass': dual_vec})\nfig2subclass = pd.concat([duplicate_removed_subclasses, dual_subclasses], axis = 0)\n\n#change fig_IDs which have more than one subsystem to 'dual'\nduplicate_subsystems = pd.DataFrame(fig2subsystem['fig_ID'].value_counts())\nduplicate_subsystems = duplicate_subsystems[duplicate_subsystems['fig_ID'] > 1].index.values\nduplicate_removed_subsystems = fig2subsystem[~fig2subsystem['fig_ID'].isin(duplicate_subsystems)]\n\ndual_vec = ['dual' for i in range(0,len(duplicate_subsystems))]\ndual_subsystems = pd.DataFrame({'fig_ID': duplicate_subsystems, 'subsystem': dual_vec})\nfig2subsystem = pd.concat([duplicate_removed_subsystems, dual_subsystems], axis = 0)\n\n#make these dataframes into dictionaries \nsubclass_dict = dict(zip(fig2subclass['fig_ID'].values, fig2subclass['subclass'].values))\nsubsystem_dict = dict(zip(fig2subsystem['fig_ID'].values, fig2subsystem['subsystem'].values))\n\n#add columns to dataframes for the subsystem and subclasses \nseqs_keys_figID['Subsystem'] = [subsystem_dict.get(fig_id) for fig_id in seqs_keys_figID['fig_ID']]\nseqs_keys_figID['Subclass'] = [subclass_dict.get(fig_id) for fig_id in seqs_keys_figID['fig_ID']]\n\n#collapse by subclass and subsystem \nseqs_subclass = seqs_keys_figID[['md5', 'Subclass']].drop_duplicates()\nseqs_subsystem = seqs_keys_figID[['md5', 'Subsystem']].drop_duplicates()\n\nseqs_subsystem = seqs_keys_figID[['md5', 'Subsystem']].drop_duplicates()\nseqs_subsystem_count = Counter(seqs_subsystem['md5'].values)\ncount_df = pd.DataFrame.from_dict(seqs_subsystem_count, orient='index').reset_index()\nduplicates = count_df[count_df[0]>1] #this gives the 2 sequences with duplicates\nduplicates_md5 = duplicates['index'].values\nduplicates_df = seqs_subsystem[seqs_subsystem['md5'].isin(duplicates_md5)]\nduplicates_idx = duplicates_df[duplicates_df['Subsystem'] != 'dual'].index.values\n\nseqs_subsystem = seqs_subsystem.drop(duplicates_idx)\n\nseqs_subclass = seqs_subclass.replace('dual', 'CO2 fixation and C-1 compound metabolism')\ncmap = ('#a6cee3', '#1f78b4', '#b2df8a', '#33a02c', '#fb9a99', '#e31a1c', '#fdbf6f', '#ff7f00', '#cab2d6', '#6a3d9a' ) #colour map for the plots ",
"_____no_output_____"
],
[
"#save the embedding - useful to save as can take a long time to run \nembed_kmerfreqDf = pd.DataFrame(embed_kmerfreq, index = embed_seqs_keys)\nembed_kmerfreqDf.columns = kmers\nembed_kmerfreqDf.to_csv('kmer_frequency_bacillus.csv')",
"_____no_output_____"
],
[
"#do the PCA \nembedding_scaled = StandardScaler().fit_transform(embed_kmerfreq)\n\npca = PCA()\nembedding_pca = pca.fit_transform(embedding_scaled)\n\n#do the scree plot - see how the PCA went \nper_var = np.round(pca.explained_variance_ratio_* 100, decimals=1)\nlabels = ['PC' + str(x) for x in range(1, len(per_var)+1)]\nplt.bar(x=range(1,len(per_var)+1), height=per_var, tick_label=labels)\nplt.ylabel('Percentage of Explained Variance')\nplt.xlabel('Principal Component')\nplt.title('Scree Plot')\nplt.show()\n\n#plot the PCA \nlabels = ['PC' + str(x) for x in range(1, len(per_var)+1)]\npca_df = pd.DataFrame(embedding_pca, columns=labels)\npca_df.index = embed_seqs_keys\npca_df['Subclass'] = seqs_subclass['Subclass'].values\n\n#sort the pca df by the subclass labels - allows to keep colours consistent between models \npca_df = pca_df.sort_values('Subclass')\n\n#get the labels for the plot \nx_label = 'PC1 ('+str(np.round(per_var[0],2))+\"%)\"\ny_label = 'PC2 ('+str(np.round(per_var[1],2))+\"%)\"\n\nsns.set(font_scale = 1.5)\nfig, ax = plt.subplots(1, 1, figsize = (12,8))\nax.set_facecolor('white')\nplt.xlabel(x_label, fontsize = 18)\nplt.ylabel(y_label, fontsize = 18)\nsns.scatterplot(x = 'PC1', y = 'PC2', hue = 'Subclass',data = pca_df, legend = 'full', s = 8,linewidth=0, alpha = 0.7, palette = cmap)\nax.spines['left'].set_color('black')\nax.spines['bottom'].set_color('black')\nplt.legend([],[], frameon=False)",
"_____no_output_____"
],
[
"#do the tSNE\ntsne = TSNE(perplexity = 50, learning_rate = 100)\nembedding_tsne = tsne.fit_transform(embedding_scaled)\n\n#plot it scatter plot \ntsne_df = pd.DataFrame(embedding_tsne, columns = ['Dimension 1', 'Dimension 2'])\ntsne_df.index = embed_seqs_keys\n\n#colour by subclass \ntsne_df['Subclass'] = seqs_subclass['Subclass'].values\n\n#sort so that the colouring is consistent \ntsne_df = tsne_df.sort_values('Subclass')\n\nsns.set(font_scale = 1.5)\nfig, ax = plt.subplots(1, 1, figsize = (12,8))\nax.set_facecolor('white')\nplt.xlabel('Dimension 1', fontsize = 18)\nplt.ylabel('Dimension 2', fontsize = 18)\nsns.scatterplot(x = 'Dimension 1', y = 'Dimension 2', hue = 'Subclass',data = tsne_df, s = 8,linewidth=0, alpha = 0.7, palette = cmap)\nax.spines['left'].set_color('black')\nax.spines['bottom'].set_color('black')\nplt.legend([],[], frameon=False)",
"_____no_output_____"
],
[
"tsne_df = pd.DataFrame(embedding_tsne, columns = ['Dimension 1', 'Dimension 2'])\n\n#colour by subclass \ntsne_df['Subclass'] = seqs_subclass['Subclass'].values\ntsne_df.index = embed_seqs_keys\n\n#sort so that the colouring is consistent \ntsne_df = tsne_df.sort_values('Subclass')\n\n\nsns.set(font_scale = 1.5)\nfig, ax = plt.subplots(1, 1, figsize = (12,8))\nax.set_facecolor('white')\nplt.xlabel('Dimension 1', fontsize = 18)\nplt.ylabel('Dimension 2', fontsize = 18)\nsns.scatterplot(x = 'Dimension 1', y = 'Dimension 2', hue = 'Subclass',data = tsne_df, s = 8,linewidth=0, alpha = 0.7, palette = cmap)\nax.spines['left'].set_color('black')\nax.spines['bottom'].set_color('black')\nplt.legend([],[], frameon=False)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ace4e4ae000a380f6e7eb4a1afeb885252a123d
| 5,908 |
ipynb
|
Jupyter Notebook
|
notebook/pandas_datareader_morningstar.ipynb
|
puyopop/python-snippets
|
9d70aa3b2a867dd22f5a5e6178a5c0c5081add73
|
[
"MIT"
] | 174 |
2018-05-30T21:14:50.000Z
|
2022-03-25T07:59:37.000Z
|
notebook/pandas_datareader_morningstar.ipynb
|
puyopop/python-snippets
|
9d70aa3b2a867dd22f5a5e6178a5c0c5081add73
|
[
"MIT"
] | 5 |
2019-08-10T03:22:02.000Z
|
2021-07-12T20:31:17.000Z
|
notebook/pandas_datareader_morningstar.ipynb
|
puyopop/python-snippets
|
9d70aa3b2a867dd22f5a5e6178a5c0c5081add73
|
[
"MIT"
] | 53 |
2018-04-27T05:26:35.000Z
|
2022-03-25T07:59:37.000Z
| 27.607477 | 89 | 0.457854 |
[
[
[
"import pandas_datareader.data as web\nimport datetime\nimport matplotlib.pyplot as plt",
"_____no_output_____"
],
[
"start = datetime.datetime(2012, 1, 1)\nend = datetime.datetime(2017, 12, 31)",
"_____no_output_____"
],
[
"f = web.DataReader('SNE', 'morningstar', start, end)",
"_____no_output_____"
],
[
"print(f.head())",
" Close High Low Open Volume\nSymbol Date \nSNE 2012-01-02 18.04 18.04 18.04 18.04 0\n 2012-01-03 18.38 18.50 18.28 18.28 1414748\n 2012-01-04 18.22 18.27 18.14 18.24 1146367\n 2012-01-05 17.70 17.85 17.60 17.83 1464843\n 2012-01-06 17.44 17.57 17.37 17.57 594057\n"
],
[
"f2 = web.DataReader(['SNE', 'AAPL'], 'morningstar', start, end)",
"_____no_output_____"
],
[
"print(type(f2.index))\nprint(f2.head())\nprint(f2.tail())",
"<class 'pandas.core.indexes.multi.MultiIndex'>\n Close High Low Open Volume\nSymbol Date \nSNE 2012-01-02 18.04 18.04 18.04 18.04 0\n 2012-01-03 18.38 18.50 18.28 18.28 1414748\n 2012-01-04 18.22 18.27 18.14 18.24 1146367\n 2012-01-05 17.70 17.85 17.60 17.83 1464843\n 2012-01-06 17.44 17.57 17.37 17.57 594057\n Close High Low Open Volume\nSymbol Date \nAAPL 2017-12-25 175.01 175.01 175.010 175.01 0\n 2017-12-26 170.57 171.47 169.679 170.80 33185536\n 2017-12-27 170.60 170.78 169.710 170.10 21498213\n 2017-12-28 171.08 171.85 170.480 171.00 16480187\n 2017-12-29 169.23 170.59 169.220 170.52 25999922\n"
],
[
"f2_u = f2.unstack(0)\nprint(f2_u.head())",
" Close High Low Open \\\nSymbol AAPL SNE AAPL SNE AAPL SNE AAPL SNE \nDate \n2012-01-02 57.8571 18.04 57.8571 18.04 57.8571 18.04 57.8571 18.04 \n2012-01-03 58.7471 18.38 58.9286 18.50 58.4286 18.28 58.5000 18.28 \n2012-01-04 59.0629 18.22 59.2400 18.27 58.4686 18.14 58.6000 18.24 \n2012-01-05 59.7186 17.70 59.7929 17.85 58.9529 17.60 59.2786 17.83 \n2012-01-06 60.3429 17.44 60.3929 17.57 59.8886 17.37 59.9671 17.57 \n\n Volume \nSymbol AAPL SNE \nDate \n2012-01-02 0 0 \n2012-01-03 75564699 1414748 \n2012-01-04 65061108 1146367 \n2012-01-05 67816805 1464843 \n2012-01-06 79596412 594057 \n"
],
[
"print(f2_u['Close'].head())",
"Symbol AAPL SNE\nDate \n2012-01-02 57.8571 18.04\n2012-01-03 58.7471 18.38\n2012-01-04 59.0629 18.22\n2012-01-05 59.7186 17.70\n2012-01-06 60.3429 17.44\n"
],
[
"f2_u['Close'].plot(title='SNE vs AAPL', grid=True)\n# plt.show()\nplt.savefig('data/dst/pandas_datareader_morningstar.png')",
"_____no_output_____"
],
[
"f2_u['Close', 'AAPL'] /= f2_u['Close'].loc[f2_u.index[0], 'AAPL']\nf2_u['Close', 'SNE'] /= f2_u['Close'].loc[f2_u.index[0], 'SNE']",
"_____no_output_____"
],
[
"f2_u['Close'].plot(title='SNE vs AAPL', grid=True)\n# plt.show()\nplt.savefig('data/dst/pandas_datareader_morningstar_normalize.png')",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ace50e744aeb321e67700f9a3f2f0c552cd6989
| 37,320 |
ipynb
|
Jupyter Notebook
|
docs/tutorials/raven.ipynb
|
louisfh/opensoundscape
|
3a038e68297743d61338bc946e5577fc4d78d420
|
[
"MIT"
] | 30 |
2019-02-21T16:51:51.000Z
|
2022-03-17T22:32:10.000Z
|
docs/tutorials/raven.ipynb
|
louisfh/opensoundscape
|
3a038e68297743d61338bc946e5577fc4d78d420
|
[
"MIT"
] | 253 |
2019-03-04T17:58:18.000Z
|
2022-03-30T23:04:49.000Z
|
docs/tutorials/raven.ipynb
|
louisfh/opensoundscape
|
3a038e68297743d61338bc946e5577fc4d78d420
|
[
"MIT"
] | 9 |
2019-08-20T17:14:45.000Z
|
2022-03-28T19:33:11.000Z
| 39.492063 | 288 | 0.532771 |
[
[
[
"# Raven annotations\nRaven Sound Analysis Software enables users to inspect spectrograms, draw time and frequency boxes around sounds of interest, and label these boxes with species identities. OpenSoundscape contains functionality to prepare and use these annotations for machine learning.",
"_____no_output_____"
],
[
"## Download annotated data\nWe published an example Raven-annotated dataset here: https://doi.org/10.1002/ecy.3329",
"_____no_output_____"
]
],
[
[
"from opensoundscape.commands import run_command\nfrom pathlib import Path",
"_____no_output_____"
]
],
[
[
"Download the zipped data here:",
"_____no_output_____"
]
],
[
[
"link = \"https://esajournals.onlinelibrary.wiley.com/action/downloadSupplement?doi=10.1002%2Fecy.3329&file=ecy3329-sup-0001-DataS1.zip\"\nname = 'powdermill_data.zip'\nout = run_command(f\"wget -O powdermill_data.zip {link}\")",
"_____no_output_____"
]
],
[
[
"Unzip the files to a new directory, `powdermill_data/`",
"_____no_output_____"
]
],
[
[
"out = run_command(\"unzip powdermill_data.zip -d powdermill_data\")",
"_____no_output_____"
]
],
[
[
"Keep track of the files we have now so we can delete them later.",
"_____no_output_____"
]
],
[
[
"files_to_delete = [Path(\"powdermill_data\"), Path(\"powdermill_data.zip\")]",
"_____no_output_____"
]
],
[
[
"## Preprocess Raven data\n\nThe `opensoundscape.raven` module contains preprocessing functions for Raven data, including:\n* `annotation_check` - for all the selections files, make sure they all contain labels\n* `lowercase_annotations` - lowercase all of the annotations\n* `generate_class_corrections` - create a CSV to see whether there are any weird names\n * Modify the CSV as needed. If you need to look up files you can use `query_annotations`\n * Can be used in `SplitterDataset`\n* `apply_class_corrections` - replace incorrect labels with correct labels\n* `query_annotations` - look for files that contain a particular species or a typo",
"_____no_output_____"
]
],
[
[
"import pandas as pd\nimport opensoundscape.raven as raven\nimport opensoundscape.audio as audio",
"_____no_output_____"
],
[
"raven_files_raw = Path(\"./powdermill_data/Annotation_Files/\")",
"_____no_output_____"
]
],
[
[
"### Check Raven files have labels\n\nCheck that all selections files contain labels under one column name. In this dataset the labels column is named `\"species\"`.",
"_____no_output_____"
]
],
[
[
"raven.annotation_check(directory=raven_files_raw, col='species')",
"All rows in powdermill_data/Annotation_Files contain labels in column `species`\n"
]
],
[
[
"### Create lowercase files\n\nConvert all the text in the files to lowercase to standardize them. Save these to a new directory. They will be saved with the same filename but with \".lower\" appended.",
"_____no_output_____"
]
],
[
[
"raven_directory = Path('./powdermill_data/Annotation_Files_Standardized')\nif not raven_directory.exists(): raven_directory.mkdir()\nraven.lowercase_annotations(directory=raven_files_raw, out_dir=raven_directory)",
"_____no_output_____"
]
],
[
[
"Check that the outputs are saved as expected.",
"_____no_output_____"
]
],
[
[
"list(raven_directory.glob(\"*.lower\"))[:5]",
"_____no_output_____"
]
],
[
[
"### Generate class corrections\n\nThis function generates a table that can be modified by hand to correct labels with typos in them. It identifies the unique labels in the provided column (here `\"species\"`) in all of the lowercase files in the directory `raven_directory`.\n\nFor instance, the generated table could be something like the following:\n```\nraw,corrected\nsparrow,sparrow\nsparow,sparow\ngoose,goose\n```",
"_____no_output_____"
]
],
[
[
"print(raven.generate_class_corrections(directory=raven_directory, col='species'))",
"raw,corrected\namcr,amcr\namgo,amgo\namre,amre\namro,amro\nbaor,baor\nbaww,baww\nbbwa,bbwa\nbcch,bcch\nbggn,bggn\nbhco,bhco\nbhvi,bhvi\nblja,blja\nbrcr,brcr\nbtnw,btnw\nbwwa,bwwa\ncang,cang\ncarw,carw\ncedw,cedw\ncora,cora\ncoye,coye\ncswa,cswa\ndowo,dowo\neato,eato\neawp,eawp\nhawo,hawo\nheth,heth\nhowa,howa\nkewa,kewa\nlowa,lowa\nnawa,nawa\nnoca,noca\nnofl,nofl\noven,oven\npiwo,piwo\nrbgr,rbgr\nrbwo,rbwo\nrcki,rcki\nrevi,revi\nrsha,rsha\nrwbl,rwbl\nscta,scta\nswth,swth\ntuti,tuti\nveer,veer\nwbnu,wbnu\nwitu,witu\nwoth,woth\nybcu,ybcu\n\n"
]
],
[
[
"The released dataset has no need for class corrections, but if it did, we could save the return text to a CSV and use the CSV to apply corrections to future dataframes.",
"_____no_output_____"
],
[
"### Query annotations\nThis function can be used to print all annotations of a particular class, e.g. \"amro\" (American Robin)",
"_____no_output_____"
]
],
[
[
"output = raven.query_annotations(directory=raven_directory, cls='amro', col='species', print_out=True)",
"=================================================================================================\npowdermill_data/Annotation_Files_Standardized/Recording_4_Segment_16.Table.1.selections.txt.lower\n=================================================================================================\n\n selection view channel begin time (s) end time (s) \\\n85 86 spectrogram 1 1 77.634876 82.129659 \n93 94 spectrogram 1 1 84.226733 86.313096 \n98 99 spectrogram 1 1 88.825438 91.272182 \n107 108 spectrogram 1 1 96.028977 97.552840 \n111 112 spectrogram 1 1 99.990354 100.914517 \n116 117 spectrogram 1 1 104.327755 108.656087 \n122 123 spectrogram 1 1 109.525937 112.021391 \n129 130 spectrogram 1 1 113.765766 117.386474 \n137 138 spectrogram 1 1 121.053454 121.383161 \n141 142 spectrogram 1 1 124.864220 129.139630 \n154 155 spectrogram 1 1 132.583749 135.017840 \n162 163 spectrogram 1 1 139.602300 142.087527 \n168 169 spectrogram 1 1 143.969913 146.785822 \n176 177 spectrogram 1 1 149.282840 151.873748 \n210 211 spectrogram 1 1 170.636021 174.123521 \n225 226 spectrogram 1 1 178.252401 181.670619 \n238 239 spectrogram 1 1 184.176135 188.110226 \n250 251 spectrogram 1 1 190.244089 192.858862 \n267 268 spectrogram 1 1 203.737856 204.958310 \n277 278 spectrogram 1 1 211.662233 216.270763 \n\n low freq (hz) high freq (hz) species \n85 1539.7 3668.7 amro \n93 1349.6 3630.6 amro \n98 1539.7 4029.8 amro \n107 1159.5 3573.6 amro \n111 1539.7 3440.4 amro \n116 1368.6 3041.4 amro \n122 1577.7 3041.4 amro \n129 1602.9 3831.4 amro \n137 1993.9 2813.1 amro \n141 1558.7 4200.9 amro \n154 2186.0 3782.7 amro \n162 1634.7 4200.9 amro \n168 1748.8 3687.7 amro \n176 1634.7 3744.7 amro \n210 1444.7 4162.9 amro \n225 1798.4 3831.4 amro \n238 1653.7 3592.6 amro \n250 1615.7 3687.7 amro \n267 1563.1 4230.8 amro \n277 1646.5 4189.1 amro \n\n=================================================================================================\npowdermill_data/Annotation_Files_Standardized/Recording_4_Segment_01.Table.1.selections.txt.lower\n=================================================================================================\n\n selection view channel begin time (s) end time (s) \\\n188 189 spectrogram 1 1 247.263069 249.107387 \n201 202 spectrogram 1 1 263.512160 264.851933 \n\n low freq (hz) high freq (hz) species \n188 1249.2 2419.2 amro \n201 1229.4 2558.0 amro \n\n"
]
],
[
[
"## Split Raven annotations and audio files\n\nThe Raven module's `raven_audio_split_and_save` function enables splitting of both audio data and associated annotations. It requires that the annotation and audio filenames are unique, and that corresponding annotation and audiofilenames are named the same filenames as each other.",
"_____no_output_____"
]
],
[
[
"audio_directory = Path('./powdermill_data/Recordings/')\ndestination = Path('./powdermill_data/Split_Recordings')\nout = raven.raven_audio_split_and_save(\n \n # Where to look for Raven files\n raven_directory = raven_directory,\n \n # Where to look for audio files\n audio_directory = audio_directory,\n \n # The destination to save clips and the labels CSV to \n destination = destination,\n \n # The column name of the labels\n col = 'species',\n \n # Desired audio sample rate\n sample_rate = 22050,\n \n # Desired duration of clips\n clip_duration = 5,\n \n # Verbose (uncomment the next line to see progress--this cell takes a while to run)\n #verbose=True,\n)",
"Found 77 sets of matching audio files and selection tables out of 77 audio files and 77 selection tables\n"
]
],
[
[
"The results of the splitting are saved in the destination folder under the name `labels.csv`.",
"_____no_output_____"
]
],
[
[
"labels = pd.read_csv(destination.joinpath(\"labels.csv\"), index_col='filename')\nlabels.head()",
"_____no_output_____"
]
],
[
[
"The `raven_audio_split_and_save` function contains several options. Notable options are:\n* `clip_duration`: the length of the clips\n* `clip_overlap`: the overlap, in seconds, between clips\n* `final_clip`: what to do with the final clip if it is not exactly `clip_duration` in length (see API docs for more details)\n* `labeled_clips_only`: whether to only save labeled clips\n* `min_label_length`: minimum length, in seconds, of an annotation for a clip to be considered labeled. For instance, if an annotation only overlaps 0.1s with a 5s clip, you might want to exclude it with `min_label_length=0.2`.\n* `species`: a subset of species to search for labels of (by default, finds all species labels in dataset)\n* `dry_run`: if `True`, produces print statements and returns dataframe of labels, but does not save files.\n* `verbose`: if `True`, prints more information, e.g. clip-by-clip progress.",
"_____no_output_____"
],
[
"For instance, let's extract labels for one species, American Redstart (AMRE) only saving clips that contain at least 0.5s of label for that species. The \"verbose\" flag causes the function to print progress splitting each clip.",
"_____no_output_____"
]
],
[
[
"btnw_split_dir = Path('./powdermill_data/btnw_recordings')\nout = raven.raven_audio_split_and_save(\n raven_directory = raven_directory,\n audio_directory = audio_directory,\n destination = btnw_split_dir,\n col = 'species',\n sample_rate = 22050,\n clip_duration = 5,\n clip_overlap = 0,\n verbose=True,\n species='amre',\n labeled_clips_only=True,\n min_label_len=1\n)",
"Found 77 sets of matching audio files and selection tables out of 77 audio files and 77 selection tables\nMaking directory powdermill_data/btnw_recordings\n1. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_13.mp3\n2. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_33.mp3\n3. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_26.mp3\n4. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_19.mp3\n5. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_11.mp3\n6. Finished powdermill_data/Recordings/Recording_2/Recording_2_Segment_13.mp3\n7. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_29.mp3\n8. Finished powdermill_data/Recordings/Recording_2/Recording_2_Segment_01.mp3\n9. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_15.mp3\n10. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_20.mp3\n11. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_12.mp3\n12. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_36.mp3\n13. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_25.mp3\n14. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_26.mp3\n15. Finished powdermill_data/Recordings/Recording_2/Recording_2_Segment_14.mp3\n16. Finished powdermill_data/Recordings/Recording_2/Recording_2_Segment_10.mp3\n17. Finished powdermill_data/Recordings/Recording_2/Recording_2_Segment_11.mp3\n18. Finished powdermill_data/Recordings/Recording_3/Recording_3_Segment_01.mp3\n19. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_32.mp3\n20. Finished powdermill_data/Recordings/Recording_2/Recording_2_Segment_03.mp3\n21. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_07.mp3\n22. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_04.mp3\n23. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_16.mp3\n24. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_30.mp3\n25. Finished powdermill_data/Recordings/Recording_2/Recording_2_Segment_02.mp3\n26. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_19.mp3\n27. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_12.mp3\n28. Finished powdermill_data/Recordings/Recording_2/Recording_2_Segment_08.mp3\n29. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_10.mp3\n30. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_20.mp3\n31. Finished powdermill_data/Recordings/Recording_2/Recording_2_Segment_12.mp3\n32. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_14.mp3\n33. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_16.mp3\n34. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_25.mp3\n35. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_09.mp3\n36. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_17.mp3\n37. Finished powdermill_data/Recordings/Recording_2/Recording_2_Segment_07.mp3\n38. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_02.mp3\n39. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_02.mp3\n40. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_08.mp3\n41. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_09.mp3\n42. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_05.mp3\n43. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_08.mp3\n44. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_05.mp3\n45. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_18.mp3\n46. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_14.mp3\n47. Finished powdermill_data/Recordings/Recording_2/Recording_2_Segment_09.mp3\n48. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_23.mp3\n49. Finished powdermill_data/Recordings/Recording_2/Recording_2_Segment_06.mp3\n50. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_34.mp3\n51. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_10.mp3\n52. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_27.mp3\n53. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_06.mp3\n54. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_31.mp3\n55. Finished powdermill_data/Recordings/Recording_2/Recording_2_Segment_04.mp3\n56. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_24.mp3\n57. Finished powdermill_data/Recordings/Recording_2/Recording_2_Segment_05.mp3\n58. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_22.mp3\n59. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_18.mp3\n60. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_01.mp3\n61. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_21.mp3\n62. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_24.mp3\n63. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_03.mp3\n64. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_01.mp3\n65. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_04.mp3\n66. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_15.mp3\n67. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_13.mp3\n68. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_07.mp3\n69. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_11.mp3\n70. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_21.mp3\n71. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_28.mp3\n72. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_17.mp3\n73. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_03.mp3\n74. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_35.mp3\n75. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_06.mp3\n76. Finished powdermill_data/Recordings/Recording_4/Recording_4_Segment_23.mp3\n77. Finished powdermill_data/Recordings/Recording_1/Recording_1_Segment_22.mp3\n"
]
],
[
[
"The labels CSV only has a column for the species of interest:",
"_____no_output_____"
]
],
[
[
"btnw_labels = pd.read_csv(btnw_split_dir.joinpath(\"labels.csv\"), index_col='filename')\nbtnw_labels.head()",
"_____no_output_____"
]
],
[
[
"The split files and associated labels csv can now be used to train machine learning models (see additional tutorials).\n\nThe command below cleans up after the tutorial is done -- only run it if you want to delete all of the files.",
"_____no_output_____"
]
],
[
[
"from shutil import rmtree\nfor file in files_to_delete:\n if file.is_dir():\n rmtree(file)\n else:\n file.unlink()",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4ace65eb90c85961daf0e182356b4df83b021143
| 13,385 |
ipynb
|
Jupyter Notebook
|
Coursera/Data Visualization with Python-IBM/Week-1/Excercise/Basic-Plot-in-Matplotlib.ipynb
|
manipiradi/Online-Courses-Learning
|
2a4ce7590d1f6d1dfa5cfde632660b562fcff596
|
[
"MIT"
] | 331 |
2019-10-22T09:06:28.000Z
|
2022-03-27T13:36:03.000Z
|
Coursera/Data Visualization with Python-IBM/Week-1/Excercise/Basic-Plot-in-Matplotlib.ipynb
|
manipiradi/Online-Courses-Learning
|
2a4ce7590d1f6d1dfa5cfde632660b562fcff596
|
[
"MIT"
] | 8 |
2020-04-10T07:59:06.000Z
|
2022-02-06T11:36:47.000Z
|
Coursera/Data Visualization with Python-IBM/Week-1/Excercise/Basic-Plot-in-Matplotlib.ipynb
|
manipiradi/Online-Courses-Learning
|
2a4ce7590d1f6d1dfa5cfde632660b562fcff596
|
[
"MIT"
] | 572 |
2019-07-28T23:43:35.000Z
|
2022-03-27T22:40:08.000Z
| 159.345238 | 7,264 | 0.917968 |
[
[
[
"%matplotlib inline\nfrom matplotlib import pyplot as plt",
"_____no_output_____"
],
[
"plt.plot(5, 5, 'o')\nplt.show()",
"_____no_output_____"
],
[
"plt.plot(5, 5, 'o')\nplt.ylabel('Y-Label')\nplt.xlabel('X-Label')\nplt.title('Ploting Example')\n\nplt.show()",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code"
]
] |
4ace7617a92e8920c0c0083b682c523a051e88f5
| 45,549 |
ipynb
|
Jupyter Notebook
|
Big-Data-Clusters/CU9/public/content/cert-management/cer043-install-master-certs.ipynb
|
WilliamAntonRohm/tigertoolbox
|
f6aa9ccbbed06d41fd7161f02a4e65871a3feaa9
|
[
"MIT"
] | 541 |
2019-05-07T11:41:25.000Z
|
2022-03-29T17:33:19.000Z
|
Big-Data-Clusters/CU9/public/content/cert-management/cer043-install-master-certs.ipynb
|
sqlworldwide/tigertoolbox
|
2abcb62a09daf0116ab1ab9c9dd9317319b23297
|
[
"MIT"
] | 89 |
2019-05-09T14:23:52.000Z
|
2022-01-13T20:21:04.000Z
|
Big-Data-Clusters/CU9/public/content/cert-management/cer043-install-master-certs.ipynb
|
sqlworldwide/tigertoolbox
|
2abcb62a09daf0116ab1ab9c9dd9317319b23297
|
[
"MIT"
] | 338 |
2019-05-08T05:45:16.000Z
|
2022-03-28T15:35:03.000Z
| 45,549 | 45,549 | 0.630596 |
[
[
[
"# CER043 - Install signed Master certificates\n\nThis notebook installs into the Big Data Cluster the certificates signed\nusing:\n\n- [CER033 - Sign Master certificates with generated\n CA](../cert-management/cer033-sign-master-generated-certs.ipynb)\n\n## Steps\n\n### Parameters",
"_____no_output_____"
]
],
[
[
"app_name = \"master\"\nscaledset_name = \"master\"\ncontainer_name = \"mssql-server\"\ncommon_name = \"master-svc\"\nuser = \"mssql\"\ngroup = \"mssql\"\nmode = \"550\"\n\nprefix_keyfile_name = \"sql\"\ncertificate_names = {\"master-0\" : \"master-0-certificate.pem\", \"master-1\" : \"master-1-certificate.pem\", \"master-2\" : \"master-2-certificate.pem\"}\nkey_names = {\"master-0\" : \"master-0-privatekey.pem\", \"master-1\" : \"master-1-privatekey.pem\", \"master-2\" : \"master-2-privatekey.pem\"}\n\ntest_cert_store_root = \"/var/opt/secrets/test-certificates\"\n\ntimeout = 600 # amount of time to wait before cluster is healthy: default to 10 minutes\ncheck_interval = 10 # amount of time between health checks - default 10 seconds\nmin_pod_count = 10 # minimum number of healthy pods required to assert health",
"_____no_output_____"
]
],
[
[
"### Common functions\n\nDefine helper functions used in this notebook.",
"_____no_output_____"
]
],
[
[
"# Define `run` function for transient fault handling, suggestions on error, and scrolling updates on Windows\nimport sys\nimport os\nimport re\nimport platform\nimport shlex\nimport shutil\nimport datetime\n\nfrom subprocess import Popen, PIPE\nfrom IPython.display import Markdown\n\nretry_hints = {} # Output in stderr known to be transient, therefore automatically retry\nerror_hints = {} # Output in stderr where a known SOP/TSG exists which will be HINTed for further help\ninstall_hint = {} # The SOP to help install the executable if it cannot be found\n\ndef run(cmd, return_output=False, no_output=False, retry_count=0, base64_decode=False, return_as_json=False):\n \"\"\"Run shell command, stream stdout, print stderr and optionally return output\n\n NOTES:\n\n 1. Commands that need this kind of ' quoting on Windows e.g.:\n\n kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='data-pool')].metadata.name}\n\n Need to actually pass in as '\"':\n\n kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='\"'data-pool'\"')].metadata.name}\n\n The ' quote approach, although correct when pasting into Windows cmd, will hang at the line:\n \n `iter(p.stdout.readline, b'')`\n\n The shlex.split call does the right thing for each platform, just use the '\"' pattern for a '\n \"\"\"\n MAX_RETRIES = 5\n output = \"\"\n retry = False\n\n # When running `azdata sql query` on Windows, replace any \\n in \"\"\" strings, with \" \", otherwise we see:\n #\n # ('HY090', '[HY090] [Microsoft][ODBC Driver Manager] Invalid string or buffer length (0) (SQLExecDirectW)')\n #\n if platform.system() == \"Windows\" and cmd.startswith(\"azdata sql query\"):\n cmd = cmd.replace(\"\\n\", \" \")\n\n # shlex.split is required on bash and for Windows paths with spaces\n #\n cmd_actual = shlex.split(cmd)\n\n # Store this (i.e. kubectl, python etc.) to support binary context aware error_hints and retries\n #\n user_provided_exe_name = cmd_actual[0].lower()\n\n # When running python, use the python in the ADS sandbox ({sys.executable})\n #\n if cmd.startswith(\"python \"):\n cmd_actual[0] = cmd_actual[0].replace(\"python\", sys.executable)\n\n # On Mac, when ADS is not launched from terminal, LC_ALL may not be set, which causes pip installs to fail\n # with:\n #\n # UnicodeDecodeError: 'ascii' codec can't decode byte 0xc5 in position 4969: ordinal not in range(128)\n #\n # Setting it to a default value of \"en_US.UTF-8\" enables pip install to complete\n #\n if platform.system() == \"Darwin\" and \"LC_ALL\" not in os.environ:\n os.environ[\"LC_ALL\"] = \"en_US.UTF-8\"\n\n # When running `kubectl`, if AZDATA_OPENSHIFT is set, use `oc`\n #\n if cmd.startswith(\"kubectl \") and \"AZDATA_OPENSHIFT\" in os.environ:\n cmd_actual[0] = cmd_actual[0].replace(\"kubectl\", \"oc\")\n\n # To aid supportability, determine which binary file will actually be executed on the machine\n #\n which_binary = None\n\n # Special case for CURL on Windows. The version of CURL in Windows System32 does not work to\n # get JWT tokens, it returns \"(56) Failure when receiving data from the peer\". If another instance\n # of CURL exists on the machine use that one. (Unfortunately the curl.exe in System32 is almost\n # always the first curl.exe in the path, and it can't be uninstalled from System32, so here we\n # look for the 2nd installation of CURL in the path)\n if platform.system() == \"Windows\" and cmd.startswith(\"curl \"):\n path = os.getenv('PATH')\n for p in path.split(os.path.pathsep):\n p = os.path.join(p, \"curl.exe\")\n if os.path.exists(p) and os.access(p, os.X_OK):\n if p.lower().find(\"system32\") == -1:\n cmd_actual[0] = p\n which_binary = p\n break\n\n # Find the path based location (shutil.which) of the executable that will be run (and display it to aid supportability), this\n # seems to be required for .msi installs of azdata.cmd/az.cmd. (otherwise Popen returns FileNotFound) \n #\n # NOTE: Bash needs cmd to be the list of the space separated values hence shlex.split.\n #\n if which_binary == None:\n which_binary = shutil.which(cmd_actual[0])\n\n # Display an install HINT, so the user can click on a SOP to install the missing binary\n #\n if which_binary == None:\n print(f\"The path used to search for '{cmd_actual[0]}' was:\")\n print(sys.path)\n\n if user_provided_exe_name in install_hint and install_hint[user_provided_exe_name] is not None:\n display(Markdown(f'HINT: Use [{install_hint[user_provided_exe_name][0]}]({install_hint[user_provided_exe_name][1]}) to resolve this issue.'))\n\n raise FileNotFoundError(f\"Executable '{cmd_actual[0]}' not found in path (where/which)\")\n else: \n cmd_actual[0] = which_binary\n\n start_time = datetime.datetime.now().replace(microsecond=0)\n\n print(f\"START: {cmd} @ {start_time} ({datetime.datetime.utcnow().replace(microsecond=0)} UTC)\")\n print(f\" using: {which_binary} ({platform.system()} {platform.release()} on {platform.machine()})\")\n print(f\" cwd: {os.getcwd()}\")\n\n # Command-line tools such as CURL and AZDATA HDFS commands output\n # scrolling progress bars, which causes Jupyter to hang forever, to\n # workaround this, use no_output=True\n #\n\n # Work around a infinite hang when a notebook generates a non-zero return code, break out, and do not wait\n #\n wait = True \n\n try:\n if no_output:\n p = Popen(cmd_actual)\n else:\n p = Popen(cmd_actual, stdout=PIPE, stderr=PIPE, bufsize=1)\n with p.stdout:\n for line in iter(p.stdout.readline, b''):\n line = line.decode()\n if return_output:\n output = output + line\n else:\n if cmd.startswith(\"azdata notebook run\"): # Hyperlink the .ipynb file\n regex = re.compile(' \"(.*)\"\\: \"(.*)\"') \n match = regex.match(line)\n if match:\n if match.group(1).find(\"HTML\") != -1:\n display(Markdown(f' - \"{match.group(1)}\": \"{match.group(2)}\"'))\n else:\n display(Markdown(f' - \"{match.group(1)}\": \"[{match.group(2)}]({match.group(2)})\"'))\n\n wait = False\n break # otherwise infinite hang, have not worked out why yet.\n else:\n print(line, end='')\n\n if wait:\n p.wait()\n except FileNotFoundError as e:\n if install_hint is not None:\n display(Markdown(f'HINT: Use {install_hint} to resolve this issue.'))\n\n raise FileNotFoundError(f\"Executable '{cmd_actual[0]}' not found in path (where/which)\") from e\n\n exit_code_workaround = 0 # WORKAROUND: azdata hangs on exception from notebook on p.wait()\n\n if not no_output:\n for line in iter(p.stderr.readline, b''):\n try:\n line_decoded = line.decode()\n except UnicodeDecodeError:\n # NOTE: Sometimes we get characters back that cannot be decoded(), e.g.\n #\n # \\xa0\n #\n # For example see this in the response from `az group create`:\n #\n # ERROR: Get Token request returned http error: 400 and server \n # response: {\"error\":\"invalid_grant\",# \"error_description\":\"AADSTS700082: \n # The refresh token has expired due to inactivity.\\xa0The token was \n # issued on 2018-10-25T23:35:11.9832872Z\n #\n # which generates the exception:\n #\n # UnicodeDecodeError: 'utf-8' codec can't decode byte 0xa0 in position 179: invalid start byte\n #\n print(\"WARNING: Unable to decode stderr line, printing raw bytes:\")\n print(line)\n line_decoded = \"\"\n pass\n else:\n\n # azdata emits a single empty line to stderr when doing an hdfs cp, don't\n # print this empty \"ERR:\" as it confuses.\n #\n if line_decoded == \"\":\n continue\n \n print(f\"STDERR: {line_decoded}\", end='')\n\n if line_decoded.startswith(\"An exception has occurred\") or line_decoded.startswith(\"ERROR: An error occurred while executing the following cell\"):\n exit_code_workaround = 1\n\n # inject HINTs to next TSG/SOP based on output in stderr\n #\n if user_provided_exe_name in error_hints:\n for error_hint in error_hints[user_provided_exe_name]:\n if line_decoded.find(error_hint[0]) != -1:\n display(Markdown(f'HINT: Use [{error_hint[1]}]({error_hint[2]}) to resolve this issue.'))\n\n # Verify if a transient error, if so automatically retry (recursive)\n #\n if user_provided_exe_name in retry_hints:\n for retry_hint in retry_hints[user_provided_exe_name]:\n if line_decoded.find(retry_hint) != -1:\n if retry_count < MAX_RETRIES:\n print(f\"RETRY: {retry_count} (due to: {retry_hint})\")\n retry_count = retry_count + 1\n output = run(cmd, return_output=return_output, retry_count=retry_count)\n\n if return_output:\n if base64_decode:\n import base64\n return base64.b64decode(output).decode('utf-8')\n else:\n return output\n\n elapsed = datetime.datetime.now().replace(microsecond=0) - start_time\n\n # WORKAROUND: We avoid infinite hang above in the `azdata notebook run` failure case, by inferring success (from stdout output), so\n # don't wait here, if success known above\n #\n if wait: \n if p.returncode != 0:\n raise SystemExit(f'Shell command:\\n\\n\\t{cmd} ({elapsed}s elapsed)\\n\\nreturned non-zero exit code: {str(p.returncode)}.\\n')\n else:\n if exit_code_workaround !=0 :\n raise SystemExit(f'Shell command:\\n\\n\\t{cmd} ({elapsed}s elapsed)\\n\\nreturned non-zero exit code: {str(exit_code_workaround)}.\\n')\n\n print(f'\\nSUCCESS: {elapsed}s elapsed.\\n')\n\n if return_output:\n if base64_decode:\n import base64\n return base64.b64decode(output).decode('utf-8')\n else:\n return output\n\n\n\n# Hints for tool retry (on transient fault), known errors and install guide\n#\nretry_hints = {'azdata': ['Endpoint sql-server-master does not exist', 'Endpoint livy does not exist', 'Failed to get state for cluster', 'Endpoint webhdfs does not exist', 'Adaptive Server is unavailable or does not exist', 'Error: Address already in use', 'Login timeout expired (0) (SQLDriverConnect)', 'SSPI Provider: No Kerberos credentials available', ], 'kubectl': ['A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond', ], 'python': [ ], }\nerror_hints = {'azdata': [['Please run \\'azdata login\\' to first authenticate', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['The token is expired', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Reason: Unauthorized', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Max retries exceeded with url: /api/v1/bdc/endpoints', 'SOP028 - azdata login', '../common/sop028-azdata-login.ipynb'], ['Look at the controller logs for more details', 'TSG027 - Observe cluster deployment', '../diagnose/tsg027-observe-bdc-create.ipynb'], ['provided port is already allocated', 'TSG062 - Get tail of all previous container logs for pods in BDC namespace', '../log-files/tsg062-tail-bdc-previous-container-logs.ipynb'], ['Create cluster failed since the existing namespace', 'SOP061 - Delete a big data cluster', '../install/sop061-delete-bdc.ipynb'], ['Failed to complete kube config setup', 'TSG067 - Failed to complete kube config setup', '../repair/tsg067-failed-to-complete-kube-config-setup.ipynb'], ['Data source name not found and no default driver specified', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Can\\'t open lib \\'ODBC Driver 17 for SQL Server', 'SOP069 - Install ODBC for SQL Server', '../install/sop069-install-odbc-driver-for-sql-server.ipynb'], ['Control plane upgrade failed. Failed to upgrade controller.', 'TSG108 - View the controller upgrade config map', '../diagnose/tsg108-controller-failed-to-upgrade.ipynb'], ['NameError: name \\'azdata_login_secret_name\\' is not defined', 'SOP013 - Create secret for azdata login (inside cluster)', '../common/sop013-create-secret-for-azdata-login.ipynb'], ['ERROR: No credentials were supplied, or the credentials were unavailable or inaccessible.', 'TSG124 - \\'No credentials were supplied\\' error from azdata login', '../repair/tsg124-no-credentials-were-supplied.ipynb'], ['Please accept the license terms to use this product through', 'TSG126 - azdata fails with \\'accept the license terms to use this product\\'', '../repair/tsg126-accept-license-terms.ipynb'], ], 'kubectl': [['no such host', 'TSG010 - Get configuration contexts', '../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb'], ['No connection could be made because the target machine actively refused it', 'TSG056 - Kubectl fails with No connection could be made because the target machine actively refused it', '../repair/tsg056-kubectl-no-connection-could-be-made.ipynb'], ], 'python': [['Library not loaded: /usr/local/opt/unixodbc', 'SOP012 - Install unixodbc for Mac', '../install/sop012-brew-install-odbc-for-sql-server.ipynb'], ['WARNING: You are using pip version', 'SOP040 - Upgrade pip in ADS Python sandbox', '../install/sop040-upgrade-pip.ipynb'], ], }\ninstall_hint = {'azdata': [ 'SOP063 - Install azdata CLI (using package manager)', '../install/sop063-packman-install-azdata.ipynb' ], 'kubectl': [ 'SOP036 - Install kubectl command line interface', '../install/sop036-install-kubectl.ipynb' ], }\n\n\nprint('Common functions defined successfully.')",
"_____no_output_____"
]
],
[
[
"### Get the Kubernetes namespace for the big data cluster\n\nGet the namespace of the Big Data Cluster use the kubectl command line\ninterface .\n\n**NOTE:**\n\nIf there is more than one Big Data Cluster in the target Kubernetes\ncluster, then either:\n\n- set \\[0\\] to the correct value for the big data cluster.\n- set the environment variable AZDATA_NAMESPACE, before starting Azure\n Data Studio.",
"_____no_output_____"
]
],
[
[
"# Place Kubernetes namespace name for BDC into 'namespace' variable\n\nif \"AZDATA_NAMESPACE\" in os.environ:\n namespace = os.environ[\"AZDATA_NAMESPACE\"]\nelse:\n try:\n namespace = run(f'kubectl get namespace --selector=MSSQL_CLUSTER -o jsonpath={{.items[0].metadata.name}}', return_output=True)\n except:\n from IPython.display import Markdown\n print(f\"ERROR: Unable to find a Kubernetes namespace with label 'MSSQL_CLUSTER'. SQL Server Big Data Cluster Kubernetes namespaces contain the label 'MSSQL_CLUSTER'.\")\n display(Markdown(f'HINT: Use [TSG081 - Get namespaces (Kubernetes)](../monitor-k8s/tsg081-get-kubernetes-namespaces.ipynb) to resolve this issue.'))\n display(Markdown(f'HINT: Use [TSG010 - Get configuration contexts](../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb) to resolve this issue.'))\n display(Markdown(f'HINT: Use [SOP011 - Set kubernetes configuration context](../common/sop011-set-kubernetes-context.ipynb) to resolve this issue.'))\n raise\n\nprint(f'The SQL Server Big Data Cluster Kubernetes namespace is: {namespace}')",
"_____no_output_____"
]
],
[
[
"### Create a temporary directory to stage files",
"_____no_output_____"
]
],
[
[
"# Create a temporary directory to hold configuration files\n\nimport tempfile\n\ntemp_dir = tempfile.mkdtemp()\n\nprint(f\"Temporary directory created: {temp_dir}\")",
"_____no_output_____"
]
],
[
[
"### Helper function to save configuration files to disk",
"_____no_output_____"
]
],
[
[
"# Define helper function 'save_file' to save configuration files to the temporary directory created above\nimport os\nimport io\n\ndef save_file(filename, contents):\n with io.open(os.path.join(temp_dir, filename), \"w\", encoding='utf8', newline='\\n') as text_file:\n text_file.write(contents)\n\n print(\"File saved: \" + os.path.join(temp_dir, filename))\n\nprint(\"Function `save_file` defined successfully.\")",
"_____no_output_____"
]
],
[
[
"### Instantiate Kubernetes client",
"_____no_output_____"
]
],
[
[
"# Instantiate the Python Kubernetes client into 'api' variable\n\nimport os\nfrom IPython.display import Markdown\n\ntry:\n from kubernetes import client, config\n from kubernetes.stream import stream\nexcept ImportError: \n\n # Install the Kubernetes module\n import sys\n !{sys.executable} -m pip install kubernetes \n \n try:\n from kubernetes import client, config\n from kubernetes.stream import stream\n except ImportError:\n display(Markdown(f'HINT: Use [SOP059 - Install Kubernetes Python module](../install/sop059-install-kubernetes-module.ipynb) to resolve this issue.'))\n raise\n\nif \"KUBERNETES_SERVICE_PORT\" in os.environ and \"KUBERNETES_SERVICE_HOST\" in os.environ:\n config.load_incluster_config()\nelse:\n try:\n config.load_kube_config()\n except:\n display(Markdown(f'HINT: Use [TSG118 - Configure Kubernetes config](../repair/tsg118-configure-kube-config.ipynb) to resolve this issue.'))\n raise\n\napi = client.CoreV1Api()\n\nprint('Kubernetes client instantiated')",
"_____no_output_____"
]
],
[
[
"### Helper functions for waiting for the cluster to become healthy",
"_____no_output_____"
]
],
[
[
"import threading\nimport time\nimport sys\nimport os\nfrom IPython.display import Markdown\n\nisRunning = True\n\ndef all_containers_ready(pod):\n \"\"\"helper method returns true if all the containers within the given pod are ready\n\n Arguments:\n pod {v1Pod} -- Metadata retrieved from the api call to.\n \"\"\"\n \n return all(map(lambda c: c.ready is True, pod.status.container_statuses))\n\n\ndef pod_is_ready(pod):\n \"\"\"tests that the pod, and all containers are ready\n\n Arguments:\n pod {v1Pod} -- Metadata retrieved from api call.\n \"\"\"\n\n return \"job-name\" in pod.metadata.labels or (pod.status.phase == \"Running\" and all_containers_ready(pod))\n\n\ndef waitReady():\n \"\"\"Waits for all pods, and containers to become ready.\n \"\"\"\n while isRunning:\n try:\n time.sleep(check_interval)\n pods = get_pods()\n allReady = len(pods.items) >= min_pod_count and all(map(pod_is_ready, pods.items))\n\n if allReady:\n return True\n else:\n display(Markdown(get_pod_failures(pods)))\n display(Markdown(f\"cluster not healthy, rechecking in {check_interval} seconds.\"))\n except Exception as ex:\n last_error_message = str(ex)\n display(Markdown(last_error_message))\n time.sleep(check_interval)\n\ndef get_pod_failures(pods=None):\n \"\"\"Returns a status message for any pods that are not ready.\n \"\"\"\n results = \"\"\n if not pods:\n pods = get_pods()\n\n for pod in pods.items:\n if \"job-name\" not in pod.metadata.labels:\n if pod.status and pod.status.container_statuses:\n for container in filter(lambda c: c.ready is False, pod.status.container_statuses):\n results = results + \"Container {0} in Pod {1} is not ready. Reported status: {2} <br/>\".format(container.name, pod.metadata.name, container.state) \n else:\n results = results + \"Pod {0} is not ready. <br/>\".format(pod.metadata.name)\n return results\n\n\ndef get_pods():\n \"\"\"Returns a list of pods by namespace, or all namespaces if no namespace is specified\n \"\"\"\n pods = None\n if namespace is not None:\n display(Markdown(f'Checking namespace {namespace}'))\n pods = api.list_namespaced_pod(namespace, _request_timeout=30) \n else:\n display(Markdown('Checking all namespaces'))\n pods = api.list_pod_for_all_namespaces(_request_timeout=30)\n return pods\n\ndef wait_for_cluster_healthy():\n isRunning = True\n mt = threading.Thread(target=waitReady)\n mt.start()\n mt.join(timeout=timeout)\n\n if mt.is_alive():\n raise SystemExit(\"Timeout waiting for all cluster to be healthy.\")\n \n isRunning = False",
"_____no_output_____"
]
],
[
[
"### Get name of the ‘Running’ `controller` `pod`",
"_____no_output_____"
]
],
[
[
"# Place the name of the 'Running' controller pod in variable `controller`\n\ncontroller = run(f'kubectl get pod --selector=app=controller -n {namespace} -o jsonpath={{.items[0].metadata.name}} --field-selector=status.phase=Running', return_output=True)\n\nprint(f\"Controller pod name: {controller}\")",
"_____no_output_____"
]
],
[
[
"### Get the name of the `master` `pods`",
"_____no_output_____"
]
],
[
[
"# Place the name of the master pods in variable `pods`\n\npodNames = run(f'kubectl get pod --selector=app=master -n {namespace} -o jsonpath={{.items[*].metadata.name}}', return_output=True)\npods = podNames.split(\" \")\n\nprint(f\"Master pod names: {pods}\")",
"_____no_output_____"
]
],
[
[
"### Validate certificate common name and alt names",
"_____no_output_____"
]
],
[
[
"import json\nfrom urllib.parse import urlparse\n\nkubernetes_default_record_name = 'kubernetes.default'\nkubernetes_default_svc_prefix = 'kubernetes.default.svc'\ndefault_dns_suffix = 'svc.cluster.local'\ndns_suffix = ''\n\nnslookup_output=run(f'kubectl exec {controller} -c controller -n {namespace} -- bash -c \"nslookup {kubernetes_default_record_name} > /tmp/nslookup.out; cat /tmp/nslookup.out; rm /tmp/nslookup.out\" ', return_output=True)\n\nname = re.findall('Name:\\s+(.[^,|^\\s|^\\n]+)', nslookup_output)\n\nif not name or kubernetes_default_svc_prefix not in name[0]:\n dns_suffix = default_dns_suffix\nelse:\n dns_suffix = 'svc' + name[0].replace(kubernetes_default_svc_prefix, '')\n\npods.sort()\n \nfor pod_name in pods:\n\n alt_names = \"\"\n bdc_fqdn = \"\"\n\n alt_names += f\"DNS.1 = {common_name}\\n\"\n alt_names += f\"DNS.2 = {common_name}.{namespace}.{dns_suffix} \\n\"\n\n hdfs_vault_svc = \"hdfsvault-svc\"\n bdc_config = run(\"azdata bdc config show\", return_output=True)\n bdc_config = json.loads(bdc_config)\n\n dns_counter = 3 # DNS.1 and DNS.2 are already in the certificate template\n\n # Stateful set related DNS names\n #\n if app_name == \"gateway\" or app_name == \"master\":\n alt_names += f'DNS.{str(dns_counter)} = {pod_name}.{common_name}\\n'\n dns_counter = dns_counter + 1\n alt_names += f'DNS.{str(dns_counter)} = {pod_name}.{common_name}.{namespace}.{dns_suffix}\\n'\n dns_counter = dns_counter + 1\n\n # AD related DNS names\n #\n if \"security\" in bdc_config[\"spec\"] and \"activeDirectory\" in bdc_config[\"spec\"][\"security\"]:\n domain_dns_name = bdc_config[\"spec\"][\"security\"][\"activeDirectory\"][\"domainDnsName\"]\n subdomain_name = bdc_config[\"spec\"][\"security\"][\"activeDirectory\"][\"subdomain\"]\n\n if subdomain_name:\n bdc_fqdn = f\"{subdomain_name}.{domain_dns_name}\"\n else:\n bdc_fqdn = f\"{namespace}.{domain_dns_name}\"\n \n alt_names += f\"DNS.{str(dns_counter)} = {common_name}.{bdc_fqdn}\\n\"\n dns_counter = dns_counter + 1\n\n if app_name == \"gateway\" or app_name == \"master\":\n alt_names += f'DNS.{str(dns_counter)} = {pod_name}.{bdc_fqdn}\\n'\n dns_counter = dns_counter + 1\n\n # Endpoint DNS names for bdc certificates\n #\n if app_name in bdc_config[\"spec\"][\"resources\"]:\n app_name_endpoints = bdc_config[\"spec\"][\"resources\"][app_name][\"spec\"][\"endpoints\"]\n for endpoint in app_name_endpoints:\n if \"dnsName\" in endpoint:\n alt_names += f'DNS.{str(dns_counter)} = {endpoint[\"dnsName\"]}\\n'\n dns_counter = dns_counter + 1\n \n # Endpoint DNS names for control plane certificates\n #\n if app_name == \"controller\" or app_name == \"mgmtproxy\":\n bdc_endpoint_list = run(\"azdata bdc endpoint list\", return_output=True)\n bdc_endpoint_list = json.loads(bdc_endpoint_list)\n\n # Parse the DNS host name from:\n #\n # \"endpoint\": \"https://monitor.aris.local:30777\"\n # \n for endpoint in bdc_endpoint_list:\n if endpoint[\"name\"] == app_name:\n url = urlparse(endpoint[\"endpoint\"])\n alt_names += f\"DNS.{str(dns_counter)} = {url.hostname}\\n\"\n dns_counter = dns_counter + 1\n\n # Special case for the controller certificate\n #\n if app_name == \"controller\":\n alt_names += f\"DNS.{str(dns_counter)} = localhost\\n\"\n dns_counter = dns_counter + 1\n\n # Add hdfsvault-svc host for key management calls.\n #\n alt_names += f\"DNS.{str(dns_counter)} = {hdfs_vault_svc}\\n\"\n dns_counter = dns_counter + 1\n\n # Add hdfsvault-svc FQDN for key management calls.\n #\n if bdc_fqdn:\n alt_names += f\"DNS.{str(dns_counter)} = {hdfs_vault_svc}.{bdc_fqdn}\\n\"\n dns_counter = dns_counter + 1\n\n required_dns_names = re.findall('DNS\\.[0-9] = ([^,|^\\s|^\\n]+)', alt_names)\n\n # Get certificate common name and DNS names\n # use nameopt compat, to generate CN= format on all versions of openssl\n # \n cert = run(f'kubectl exec {controller} -c controller -n {namespace} -- openssl x509 -nameopt compat -in {test_cert_store_root}/{app_name}/{certificate_names[pod_name]} -text -noout', return_output=True)\n subject = re.findall('Subject:(.+)', cert)[0]\n certficate_common_name = re.findall('CN=(.[^,|^\\s|^\\n]+)', subject)[0]\n certficate_dns_names = re.findall('DNS:(.[^,|^\\s|^\\n]+)', cert)\n\n # Validate the common name\n #\n if (common_name != certficate_common_name):\n run(f'kubectl exec {controller} -c controller -n {namespace} -- bash -c \"rm -rf {test_cert_store_root}/{app_name}\"')\n raise SystemExit(f'Certficate common name does not match the expected one: {common_name}')\n\n # Validate the DNS names\n #\n if not all(dns_name in certficate_dns_names for dns_name in required_dns_names):\n run(f'kubectl exec {controller} -c controller -n {namespace} -- bash -c \"rm -rf {test_cert_store_root}/{app_name}\"')\n raise SystemExit(f'Certficate does not have all required DNS names: {required_dns_names}')",
"_____no_output_____"
]
],
[
[
"### Copy certifcate files from `controller` to local machine",
"_____no_output_____"
]
],
[
[
"import os\n\ncwd = os.getcwd()\nos.chdir(temp_dir) # Use chdir to workaround kubectl bug on Windows, which incorrectly processes 'c:\\' on kubectl cp cmd line \n\nfor pod_name in pods:\n\n run(f'kubectl cp {controller}:{test_cert_store_root}/{app_name}/{certificate_names[pod_name]} {certificate_names[pod_name]} -c controller -n {namespace}')\n run(f'kubectl cp {controller}:{test_cert_store_root}/{app_name}/{key_names[pod_name]} {key_names[pod_name]} -c controller -n {namespace}')\n\nos.chdir(cwd)",
"_____no_output_____"
]
],
[
[
"### Copy certifcate files from local machine to `controldb`",
"_____no_output_____"
]
],
[
[
"import os\n\ncwd = os.getcwd()\nos.chdir(temp_dir) # Workaround kubectl bug on Windows, can't put c:\\ on kubectl cp cmd line \n \nfor pod_name in pods:\n run(f'kubectl cp {certificate_names[pod_name]} controldb-0:/var/opt/mssql/{certificate_names[pod_name]} -c mssql-server -n {namespace}')\n run(f'kubectl cp {key_names[pod_name]} controldb-0:/var/opt/mssql/{key_names[pod_name]} -c mssql-server -n {namespace}')\n\nos.chdir(cwd)",
"_____no_output_____"
]
],
[
[
"### Get the `controller-db-rw-secret` secret\n\nGet the controller SQL symmetric key password for decryption.",
"_____no_output_____"
]
],
[
[
"import base64\n\ncontroller_db_rw_secret = run(f'kubectl get secret/controller-db-rw-secret -n {namespace} -o jsonpath={{.data.encryptionPassword}}', return_output=True)\ncontroller_db_rw_secret = base64.b64decode(controller_db_rw_secret).decode('utf-8')\n\nprint(\"controller_db_rw_secret retrieved\")",
"_____no_output_____"
]
],
[
[
"### Update the files table with the certificates through opened SQL connection",
"_____no_output_____"
]
],
[
[
"import os\n\nsql = f\"\"\"\nOPEN SYMMETRIC KEY ControllerDbSymmetricKey DECRYPTION BY PASSWORD = '{controller_db_rw_secret}'\n\nDECLARE @FileData VARBINARY(MAX), @Key uniqueidentifier;\nSELECT @Key = KEY_GUID('ControllerDbSymmetricKey');\n\n\"\"\"\n \nfor pod_name in pods:\n\n insert = f\"\"\"\n SELECT TOP 1 @FileData = doc.BulkColumn FROM OPENROWSET(BULK N'/var/opt/mssql/{certificate_names[pod_name]}', SINGLE_BLOB) AS doc;\n EXEC [dbo].[sp_set_file_data_encrypted] @FilePath = '/config/scaledsets/{scaledset_name}/pods/{pod_name}/containers/{container_name}/files/{prefix_keyfile_name}-certificate.pem',\n @Data = @FileData,\n @KeyGuid = @Key,\n @Version = '0',\n @User = '{user}',\n @Group = '{group}',\n @Mode = '{mode}';\n\n SELECT TOP 1 @FileData = doc.BulkColumn FROM OPENROWSET(BULK N'/var/opt/mssql/{key_names[pod_name]}', SINGLE_BLOB) AS doc;\n EXEC [dbo].[sp_set_file_data_encrypted] @FilePath = '/config/scaledsets/{scaledset_name}/pods/{pod_name}/containers/{container_name}/files/{prefix_keyfile_name}-privatekey.pem',\n @Data = @FileData,\n @KeyGuid = @Key,\n @Version = '0',\n @User = '{user}',\n @Group = '{group}',\n @Mode = '{mode}';\n\n \"\"\"\n\n sql += insert\n\nsave_file(\"insert_certificates.sql\", sql)\n\ncwd = os.getcwd()\nos.chdir(temp_dir) # Workaround kubectl bug on Windows, can't put c:\\ on kubectl cp cmd line \n\nrun(f'kubectl cp insert_certificates.sql controldb-0:/var/opt/mssql/insert_certificates.sql -c mssql-server -n {namespace}')\n\nrun(f\"\"\"kubectl exec controldb-0 -c mssql-server -n {namespace} -- bash -c \"SQLCMDPASSWORD=`cat /var/run/secrets/credentials/mssql-sa-password/password` /opt/mssql-tools/bin/sqlcmd -b -U sa -d controller -i /var/opt/mssql/insert_certificates.sql\" \"\"\")\n\n# Clean up\nrun(f\"\"\"kubectl exec controldb-0 -c mssql-server -n {namespace} -- bash -c \"rm /var/opt/mssql/insert_certificates.sql\" \"\"\")\n\nfor pod_name in pods:\n\n run(f\"\"\"kubectl exec controldb-0 -c mssql-server -n {namespace} -- bash -c \"rm /var/opt/mssql/{certificate_names[pod_name]}\" \"\"\")\n run(f\"\"\"kubectl exec controldb-0 -c mssql-server -n {namespace} -- bash -c \"rm /var/opt/mssql/{key_names[pod_name]}\" \"\"\")\n\nos.chdir(cwd)",
"_____no_output_____"
]
],
[
[
"### Clear out the controller_db_rw_secret variable",
"_____no_output_____"
]
],
[
[
"controller_db_rw_secret= \"\"",
"_____no_output_____"
]
],
[
[
"### Get the name of the `master` `pods`",
"_____no_output_____"
]
],
[
[
"# Place the name of the master pods in variable `pods`\n\npodNames = run(f'kubectl get pod --selector=app=master -n {namespace} -o jsonpath={{.items[*].metadata.name}}', return_output=True)\npods = podNames.split(\" \")\n\nprint(f\"Master pod names: {pods}\")",
"_____no_output_____"
]
],
[
[
"### Restart Pods",
"_____no_output_____"
]
],
[
[
"import threading\nimport time\n\nif len(pods) == 1:\n # One master pod indicates non-HA environment, just delete it\n run(f'kubectl delete pod {pods[0]} -n {namespace}')\n wait_for_cluster_healthy()\nelse:\n # HA setup, delete secondaries before primary\n timeout_s = 300\n check_interval_s = 20\n\n master_primary_svc_ip = run(f'kubectl get service master-p-svc -n {namespace} -o jsonpath={{.spec.clusterIP}}', return_output=True) \n master_password = run(f'kubectl exec master-0 -c mssql-server -n {namespace} -- cat /var/run/secrets/credentials/pool/mssql-system-password', return_output=True) \n\n def get_number_of_unsynchronized_replicas(result):\n cmd = 'select count(*) from sys.dm_hadr_database_replica_states where synchronization_state <> 2'\n res = run(f\"kubectl exec controldb-0 -c mssql-server -n {namespace} -- /opt/mssql-tools/bin/sqlcmd -S {master_primary_svc_ip} -U system -P {master_password} -h -1 -q \\\"SET NOCOUNT ON; {cmd}\\\" \", return_output=True)\n rows = res.strip().split(\"\\n\")\n\n result[0] = int(rows[0])\n return True\n\n def get_primary_replica():\n cmd = 'select distinct replica_server_name from sys.dm_hadr_database_replica_states s join sys.availability_replicas r on s.replica_id = r.replica_id where is_primary_replica = 1'\n res = run(f\"kubectl exec controldb-0 -c mssql-server -n {namespace} -- /opt/mssql-tools/bin/sqlcmd -S {master_primary_svc_ip} -U system -P {master_password} -h -1 -q \\\"SET NOCOUNT ON; {cmd}\\\" \", return_output=True)\n\n rows = res.strip().split(\"\\n\")\n return rows[0]\n\n def get_secondary_replicas():\n cmd = 'select distinct replica_server_name from sys.dm_hadr_database_replica_states s join sys.availability_replicas r on s.replica_id = r.replica_id where is_primary_replica = 0'\n res = run(f\"kubectl exec controldb-0 -c mssql-server -n {namespace} -- /opt/mssql-tools/bin/sqlcmd -S {master_primary_svc_ip} -U system -P {master_password} -h -1 -q \\\"SET NOCOUNT ON; {cmd}\\\" \", return_output=True)\n\n rows = res.strip().split(\"\\n\")\n res = []\n for row in rows:\n if (row != \"\" and \"Sqlcmd: Warning\" not in row):\n res.append(row.strip())\n return res\n\n def all_replicas_syncrhonized():\n while True:\n unsynchronized_replicas_cnt = len(pods)\n rows = [None]\n time.sleep(check_interval_s)\n\n getNumberOfReplicasThread = threading.Thread(target=get_number_of_unsynchronized_replicas, args=(rows,) )\n getNumberOfReplicasThread.start()\n getNumberOfReplicasThread.join(timeout=timeout_s)\n\n if getNumberOfReplicasThread.is_alive():\n raise SystemExit(\"Timeout getting the number of unsynchronized replicas.\")\n\n unsynchronized_replicas_cnt = rows[0]\n if (unsynchronized_replicas_cnt == 0):\n return True\n\n def wait_for_replicas_to_synchronize():\n waitForReplicasToSynchronizeThread = threading.Thread(target=all_replicas_syncrhonized)\n waitForReplicasToSynchronizeThread.start()\n waitForReplicasToSynchronizeThread.join(timeout=timeout_s)\n\n if waitForReplicasToSynchronizeThread.is_alive():\n raise SystemExit(\"Timeout waiting for all replicas to be synchronized.\")\n \n secondary_replicas = get_secondary_replicas()\n\n for replica in secondary_replicas:\n wait_for_replicas_to_synchronize()\n run(f'kubectl delete pod {replica} -n {namespace}')\n\n primary_replica = get_primary_replica() \n wait_for_replicas_to_synchronize()\n\n key = \"/var/run/secrets/certificates/sqlha/mssql-ha-operator-controller-client/mssql-ha-operator-controller-client-privatekey.pem\"\n cert = \"/var/run/secrets/certificates/sqlha/mssql-ha-operator-controller-client/mssql-ha-operator-controller-client-certificate.pem\"\n content_type_header = \"Content-Type: application/json\"\n authorization_header = \"Authorization: Certificate\"\n data = f'{{\"TargetReplicaName\":\"{secondary_replicas[0]}\",\"ForceFailover\":\"false\"}}'\n request_url = f'https://controller-svc:443/internal/api/v1/bdc/services/sql/resources/master/availabilitygroups/containedag/failover'\n\n manual_failover_api_command = f\"curl -sS --key {key} --cert {cert} -X POST --header '{content_type_header}' --header '{authorization_header}' --data '{data}' {request_url}\"\n\n operator_pod = run(f'kubectl get pod --selector=app=mssql-operator -n {namespace} -o jsonpath={{.items[0].metadata.name}}', return_output=True)\n\n run(f'kubectl exec {operator_pod} -c mssql-ha-operator -n {namespace} -- {manual_failover_api_command}')\n\n wait_for_replicas_to_synchronize()\n\n run(f'kubectl delete pod {primary_replica} -n {namespace}')\n wait_for_replicas_to_synchronize()",
"_____no_output_____"
]
],
[
[
"### Clean up certificate staging area\n\nRemove the certificate files generated on disk (they have now been\nplaced in the controller database).",
"_____no_output_____"
]
],
[
[
"cmd = f\"rm -r {test_cert_store_root}/{app_name}\"\n\nrun(f'kubectl exec {controller} -c controller -n {namespace} -- bash -c \"{cmd}\"')",
"_____no_output_____"
]
],
[
[
"### Clean up temporary directory for staging configuration files",
"_____no_output_____"
]
],
[
[
"# Delete the temporary directory used to hold configuration files\n\nimport shutil\n\nshutil.rmtree(temp_dir)\n\nprint(f'Temporary directory deleted: {temp_dir}')",
"_____no_output_____"
],
[
"print(\"Notebook execution is complete.\")",
"_____no_output_____"
]
],
[
[
"Related\n-------\n\n- [CER023 - Create Master certificates](../cert-management/cer023-create-master-certs.ipynb)\n- [CER033 - Sign Master certificates with generated CA](../cert-management/cer033-sign-master-generated-certs.ipynb)\n- [CER044 - Install signed Controller certificate](../cert-management/cer044-install-controller-cert.ipynb)\n",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
]
] |
4acead9b758c100586bbefcd16344b3d6d1fe377
| 77,226 |
ipynb
|
Jupyter Notebook
|
src/Test_quera_waveforms.ipynb
|
maolinml/amazon-braket-sdk-python
|
48431815c75be592ec1ee1c5952d90e777df0a13
|
[
"Apache-2.0"
] | null | null | null |
src/Test_quera_waveforms.ipynb
|
maolinml/amazon-braket-sdk-python
|
48431815c75be592ec1ee1c5952d90e777df0a13
|
[
"Apache-2.0"
] | null | null | null |
src/Test_quera_waveforms.ipynb
|
maolinml/amazon-braket-sdk-python
|
48431815c75be592ec1ee1c5952d90e777df0a13
|
[
"Apache-2.0"
] | null | null | null | 388.070352 | 14,404 | 0.944345 |
[
[
[
"from braket.analog.quera.waveforms import Waveform, LinearWaveform, ConstantWaveform, \\\nCompositeWaveform, CustomWaveform, InterpolatedWaveform\n\nimport numpy as np",
"_____no_output_____"
],
[
"wf1 = LinearWaveform(2, 10, 20)\nprint(f\"duration = {wf1.duration}\")\nprint(f\"first_value = {wf1.first_value}\")\nprint(f\"last_value = {wf1.last_value}\")\nprint(f\"integral = {wf1.integral}\")\nprint(f\"slope = {wf1.slope}\")\nprint(wf1.samples)\nwf1.draw()\nwf1 = wf1.change_duration(4)\nwf1.draw()",
"duration = 2\nfirst_value = 10.0\nlast_value = 20.0\nintegral = 0.03\nslope = 5.0\n[10. 20.]\n"
],
[
"wf2 = ConstantWaveform(2, 12)\nwf2.draw()",
"_____no_output_____"
],
[
"wf3 = CompositeWaveform(wf1, wf2)\nwf3.draw()",
"_____no_output_____"
],
[
"wf4 = CustomWaveform(np.linspace(0, 10, 20))\nwf4.draw()",
"_____no_output_____"
],
[
"wf5 = InterpolatedWaveform(4, [0.1, 0.2] , [0.5, 0.9])\nwf5.draw()",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acec6cc10cbffd9ebca38b5dea0ba0b7634f07b
| 190,634 |
ipynb
|
Jupyter Notebook
|
dtw.ipynb
|
tahoemph/ipython_exploration
|
a9fb33ccb383801192eb8ac7fd9727f0ea0305d8
|
[
"BSD-2-Clause"
] | null | null | null |
dtw.ipynb
|
tahoemph/ipython_exploration
|
a9fb33ccb383801192eb8ac7fd9727f0ea0305d8
|
[
"BSD-2-Clause"
] | null | null | null |
dtw.ipynb
|
tahoemph/ipython_exploration
|
a9fb33ccb383801192eb8ac7fd9727f0ea0305d8
|
[
"BSD-2-Clause"
] | null | null | null | 240.699495 | 61,019 | 0.899462 |
[
[
[
"empty"
]
]
] |
[
"empty"
] |
[
[
"empty"
]
] |
4aced8cf2be71dece0150e77d98852ff766b1b5a
| 104,874 |
ipynb
|
Jupyter Notebook
|
es12/12.2.ipynb
|
lorycontixd/PNS
|
23f61833ae75a2425d05ca9757ef471aec930654
|
[
"MIT"
] | null | null | null |
es12/12.2.ipynb
|
lorycontixd/PNS
|
23f61833ae75a2425d05ca9757ef471aec930654
|
[
"MIT"
] | null | null | null |
es12/12.2.ipynb
|
lorycontixd/PNS
|
23f61833ae75a2425d05ca9757ef471aec930654
|
[
"MIT"
] | null | null | null | 215.347023 | 29,776 | 0.89926 |
[
[
[
"# <span style=\"color:orange\"> Exercise 12.2 </span>\n\n## <span style=\"color:green\"> Task </span>\nChange the architecture of your DNN using convolutional layers. Use `Conv2D`, `MaxPooling2D`, `Dropout`, but also do not forget `Flatten`, a standard `Dense` layer and `soft-max` in the end. I have merged step 2 and 3 in the following definition of `create_CNN()` that **<span style=\"color:red\">you should complete</span>**<br><br>\nThe cell below shows some information about the dataset, such as the size and a set of 10 images to reveal the appearance of the data.",
"_____no_output_____"
]
],
[
[
"import tensorflow as tf\ntf.get_logger().setLevel('INFO')\nfrom keras.datasets import mnist\nfrom matplotlib import pyplot as plt\nfrom numba import jit, njit\n# load dataset\n(trainX, trainy), (testX, testy) = mnist.load_data()\n# summarize loaded dataset\nprint('Train: X=%s, y=%s' % (trainX.shape, trainy.shape))\nprint('Test: X=%s, y=%s' % (testX.shape, testy.shape))\n# plot first few images\nfor i in range(9):\n # define subplot\n plt.subplot(330 + 1 + i)\n # plot raw pixel data\n plt.imshow(trainX[i], cmap=plt.get_cmap('gray'))\n# show the figure\nplt.show()",
"Train: X=(60000, 28, 28), y=(60000,)\nTest: X=(10000, 28, 28), y=(10000,)\n"
]
],
[
[
"### Preparing Data\n\nThe following functions respectively load the MNIST dataset and scale the pixels of the images. In fact, the images are made up of pixels whose values are integers describing a color between black and white, or an integer from 0 to 255. Therefore, the second function normalizes the values of the pixels to the range [0,1] after having converted them to floats.",
"_____no_output_____"
]
],
[
[
"from numpy import mean\nfrom numpy import std\nfrom matplotlib import pyplot\nfrom sklearn.model_selection import KFold\nfrom tensorflow.keras.datasets import mnist\nfrom tensorflow.keras.utils import to_categorical\nfrom tensorflow.keras.models import Sequential\nfrom tensorflow.keras.layers import Conv2D\nfrom tensorflow.keras.layers import MaxPooling2D\nfrom tensorflow.keras.layers import Dense\nfrom tensorflow.keras.layers import Flatten\nfrom tensorflow.keras.optimizers import SGD\n \n# load train and test dataset\ndef load_dataset():\n # load dataset\n (trainX, trainY), (testX, testY) = mnist.load_data()\n # reshape dataset to have a single channel\n trainX = trainX.reshape((trainX.shape[0], 28, 28, 1))\n testX = testX.reshape((testX.shape[0], 28, 28, 1))\n # one hot encode target values\n trainY = to_categorical(trainY)\n testY = to_categorical(testY)\n print(len(trainX),\"-\",len(trainY),\"-\",len(testX),\"-\",len(testY))\n return trainX, trainY, testX, testY\n\n# scale pixels\ndef prep_pixels(train, test):\n # convert from integers to floats\n train_norm = train.astype('float32')\n test_norm = test.astype('float32')\n # normalize to range 0-1\n train_norm = train_norm / 255.0\n test_norm = test_norm / 255.0\n # return normalized images\n return train_norm, test_norm",
"_____no_output_____"
]
],
[
[
"### Defining model\nThe model that I used to define the convolutional neural network starts with a Convolutional Layer with a moderate amount of filters (32) and a filter size of (3,3). The layer is followed by a Max Pooling Layer, while the filter map is then flattened using a Flatten layer.\nThe output layer should consist of 10 nodes because the images must be classified across 10 different classes, while the same layer should follow the softmax activation function. The softmax activation function $\\sigma : {\\rm I\\!R}^K \\longrightarrow [0,1]^K$ is defined by: $$ \\sigma(\\vec{z})_i = \\frac{e^{z_i}}{\\sum_{j=1}^{K}e^{z_j}}, $$\nused to normalize the output of a neural network to a probability distribution. Between the convolutional layers and the output layer, I introduced a dense layer with 100 neurons and a ReLU activation function. <br>\nThe model uses a Stochastic Descent Gradient (SDG) optimizer with a learning rate of 0.01 and a momentum of 0.9, a categorical cross-entropy loss function and will monitor the classification accuracy metric.",
"_____no_output_____"
]
],
[
[
"# define CNN model\ndef define_model(summarize:bool=False,save:bool=False):\n model = Sequential()\n model.add(Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_uniform', input_shape=(28, 28, 1)))\n model.add(MaxPooling2D((2, 2)))\n model.add(Flatten())\n model.add(Dense(100, activation='relu', kernel_initializer='he_uniform'))\n model.add(Dense(10, activation='softmax'))\n # compile model\n opt = SGD(learning_rate=0.01, momentum=0.9)\n model.compile(optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])\n if summarize:\n model.summary()\n if save:\n from pathlib import Path\n dir = \"./models/\"\n Path(dir).mkdir(parents=True, exist_ok=True)\n print(\"[MODEL] Saved model to directory: \"+dir)\n model.save(dir)\n return model",
"_____no_output_____"
]
],
[
[
"#### Using a 5-fold validation\n<img src=\"../media/images/kfold-validation.png\" alt=\"Five-fold cross validation\" width=\"800\" height=\"600\">\n<br>The model will be evaluated using a five-fold cross validation. In general, a k-fold cross validation is used to effectively split a small part of the full dataset for testing purposes, while the rest is used for training purposes. The K-fold cross validation splits the dataset into K partitions of equal size, so that K-1 can be used for training while the last one can be used for testing. Finally, the procedure is repeated K times, with a different partition for each iteration. In the case of a five-fold cross validation, the dataset is split into 80/20, each time with a different portion used for testing. <br>\nThe model is trained with a standard batch size of 32 examples and with 10 epochs. The latter is lower than usual, primarily because the five-fold validation process requires five times for computational power than manual validation, while the number is still valid for accurate results. ",
"_____no_output_____"
]
],
[
[
"# evaluate a model using k-fold cross-validation\n\ndef evaluate_model(dataX, dataY, n_folds=5, model=None):\n scores, histories = list(), list()\n # prepare cross validation\n kfold = KFold(n_folds, shuffle=True, random_state=1)\n # enumerate splits\n iter = 0\n \n for train_ix, test_ix in kfold.split(dataX):\n print(\"[EVAL] Running iteration {}\".format(iter))\n # define model\n if model is None:\n if iter<n_folds-1:\n model = define_model(save=False)\n else:\n model = define_model(save=True)\n else:\n model = model\n # select rows for train and test\n trainX, trainY, testX, testY = dataX[train_ix], dataY[train_ix], dataX[test_ix], dataY[test_ix]\n # fit model\n history = model.fit(trainX, trainY, epochs=10, batch_size=32, validation_data=(testX, testY), verbose=0)\n # evaluate model\n _, acc = model.evaluate(testX, testY, verbose=0)\n print('> %.3f' % (acc * 100.0))\n # stores scores\n scores.append(acc)\n histories.append(history)\n iter += 1\n return scores, histories",
"_____no_output_____"
]
],
[
[
"### Validation\nThe evaluation method defined in the previous cell returns a list of accuracy scores and training histories used to summarize and validate the model. The diagnostics function aims to plot the model performance for training and testing during each fold of the five-fold cross-validation, for both the loss and the accuracy of the model. The performance is then summarized by calculated the mean and standard deviation of the accuracy scores.",
"_____no_output_____"
]
],
[
[
"# plot diagnostic learning curves ---> training\ndef summarize_diagnostics(histories):\n fig, ax = plt.subplots(2,1, gridspec_kw={'hspace': 0.6})\n for i in range(len(histories)):\n ax[0].set_title('Cross Entropy Loss')\n ax[0].plot(histories[i].history['loss'], color='blue', label='train')\n ax[0].plot(histories[i].history['val_loss'], color='orange', label='test')\n ax[0].set_xlabel(\"Epoch\")\n ax[0].set_xlabel(\"Loss\")\n # plot accuracy\n ax[1].set_title('Classification Accuracy')\n ax[1].plot(histories[i].history['accuracy'], color='blue', label='train')\n ax[1].plot(histories[i].history['val_accuracy'], color='orange', label='test')\n ax[1].set_xlabel(\"Epoch\")\n ax[1].set_ylabel(\"Accuracy\")\n pyplot.show()\n \n# summarize model performance ---> testing\ndef summarize_performance(scores):\n # print summary\n print('Accuracy: mean=%.3f std=%.3f, n=%d' % (mean(scores)*100, std(scores)*100, len(scores)))\n # box and whisker plots of results\n plt.boxplot(scores) \n plt.ylabel(\"Accuracy\")\n plt.show()\n \n# run the test harness for evaluating a model\ndef run(model=None):\n # load dataset\n print(\"\\r[RUN] Loading dataset..\")\n trainX, trainY, testX, testY = load_dataset()\n # prepare pixel data\n print(\"\\r[RUN] Preparing pixels\")\n trainX, testX = prep_pixels(trainX, testX)\n # evaluate model\n print(\"\\r[RUN] Creating and evaluating the model\")\n scores, histories = evaluate_model(trainX, trainY, model=model)\n # learning curves\n print(\"\\r[RUN] Learning curves\")\n summarize_diagnostics(histories)\n # summarize estimated performance\n print(\"\\r[RUN] Evaluating performance\")\n summarize_performance(scores)",
"_____no_output_____"
],
[
"run()",
"[RUN] Loading dataset..\n60000 - 60000 - 10000 - 10000\n[RUN] Preparing pixels\n[RUN] Creating and evaluating the model\n[EVAL] Running iteration 0\n> 98.600\n[EVAL] Running iteration 1\n> 99.842\n[EVAL] Running iteration 2\n> 99.992\n[EVAL] Running iteration 3\n> 100.000\n[EVAL] Running iteration 4\n> 100.000\n[RUN] Learning curves\n"
]
],
[
[
"## <span style=\"color:green\"> Results </span>\nWhen running the CNN, each fold of the cross-validation produces an excellent accuracy score, which means that the network is classifying the images very well. The plots show how the model is learning during each fold, where both <span style=\"color:blue\"> training </span> and <span style=\"color:orange\"> testing </span> curves converge, meaning that the data is not being over-fitted or under-fitted.<br>\nTo summarize the model performance, the model has an estimated accuracy of 98.7\\%, which is a good result. This is also shown in a box and whisker plot, which exposes quantities like the maximum value, the minimum value, the median, the first and third quartile of the data.<br>\n\n## <span style=\"color:green\"> How to improve the model </span>\nThe following section explores how the CNN model could be improved to correctly predict and classify images.\n\n### Learning rate\nProbably, the most impactful variable is the learning rate of the Stochastic Descent Gradient, which is the hyperparameter that defines the rate at which the weights of the model are updated during training. It is a difficult task to find a good balance for its value, since a value that is too large can cause the model to converge too quickly to an inaccurate solution, while a value that is too low can cause the model to get stuck or take very long times to converge.\n\n### Batch Normalization\nBatch normaization is a process to standardize the inputs to a layer in a Neural Network. This can accelerate the learning pace of a model and sometimes improve its accuracy. The Batch Normalization layer is typically introduced after convolutional and fully-connected layers, to standardize inputs so that the mean is zero and the standard deviation is one. Finally, the standardized outputs can be eventually scaled to extract the new mean and standard deviation for the final output.<br>\nFor this specific CNN model, I've decided to introduce the Batch Normalization layer after the activation function, between the convolutional and the max pooling layers:<br>",
"_____no_output_____"
]
],
[
[
"from keras.layers import BatchNormalization\n# define CNN model\ndef define_model(summarize:bool=False,save:bool=False):\n model = Sequential()\n model.add(Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_uniform', input_shape=(28, 28, 1)))\n model.add(BatchNormalization())\n model.add(MaxPooling2D((2, 2)))\n model.add(Flatten())\n model.add(Dense(100, activation='relu', kernel_initializer='he_uniform'))\n model.add(Dense(10, activation='softmax'))\n # compile model\n opt = SGD(learning_rate =0.01, momentum=0.9)\n model.compile(optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])\n if summarize:\n model.summary()\n if save:\n from pathlib import Path\n dir = \"./models/\"\n Path(dir).mkdir(parents=True, exist_ok=True)\n print(\"[MODEL] Saved model to directory: \"+dir)\n model.save(dir)\n return model",
"_____no_output_____"
],
[
"run()",
"[RUN] Loading dataset..\n60000 - 60000 - 10000 - 10000\n[RUN] Preparing pixels\n[RUN] Creating and evaluating the model\n[EVAL] Running iteration 0\n> 98.575\n[EVAL] Running iteration 1\n> 99.783\n[EVAL] Running iteration 2\n> 100.000\n[EVAL] Running iteration 3\n> 100.000\n[EVAL] Running iteration 4\n> 100.000\n[RUN] Learning curves\n"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4acee7a45d6b77d8ee266d7e669e0acfe5b1ebbb
| 414,262 |
ipynb
|
Jupyter Notebook
|
Data Mining/Untitled1.ipynb
|
vkgpt11/GreatLakesAssignments
|
14daf30c617f79ab43df51ac8d82ef0afb9c9bf8
|
[
"Apache-2.0"
] | null | null | null |
Data Mining/Untitled1.ipynb
|
vkgpt11/GreatLakesAssignments
|
14daf30c617f79ab43df51ac8d82ef0afb9c9bf8
|
[
"Apache-2.0"
] | null | null | null |
Data Mining/Untitled1.ipynb
|
vkgpt11/GreatLakesAssignments
|
14daf30c617f79ab43df51ac8d82ef0afb9c9bf8
|
[
"Apache-2.0"
] | null | null | null | 72.423427 | 48,142 | 0.686829 |
[
[
[
"d = data.frame(age=rnorm(1000,45,7.6), inc=rnorm(1000,1000,1200),height=runif(1000,140,180))",
"_____no_output_____"
],
[
"head(d)",
"_____no_output_____"
],
[
"as.data.frame(scale(d)) -> d1",
"_____no_output_____"
],
[
"kmeans(d1,3) -> c1",
"_____no_output_____"
],
[
"c1",
"_____no_output_____"
],
[
"c1$withinss",
"_____no_output_____"
],
[
"c1$betweenss",
"_____no_output_____"
],
[
"c1$size",
"_____no_output_____"
],
[
"c1$tot.withinss",
"_____no_output_____"
],
[
"c1$totss",
"_____no_output_____"
],
[
"d$cluster <- c1$cluster",
"_____no_output_____"
],
[
"head(d)",
"_____no_output_____"
],
[
"boxplot(height~cluster,data=d)",
"_____no_output_____"
],
[
"boxplot(age~cluster,data=d)",
"_____no_output_____"
],
[
"kmeans(d1,5) -> c1\nc1",
"_____no_output_____"
]
],
[
[
"# Hierarchical Clustering ",
"_____no_output_____"
]
],
[
[
"dist(d1) -> dt",
"_____no_output_____"
],
[
"hclust(dt,method = \"single\") -> hsin",
"_____no_output_____"
],
[
"plot(hsin)",
"_____no_output_____"
],
[
"plot(hsin, hang=1)",
"_____no_output_____"
],
[
"hclust(dt,method = \"complete\") -> hcomp\nplot(hcomp)\n",
"_____no_output_____"
],
[
"hclust(dt,method = \"complete\") -> havg\nplot(havg)",
"_____no_output_____"
],
[
"cutree(hcomp,4) -> hcom4",
"_____no_output_____"
],
[
"hcom4",
"_____no_output_____"
],
[
"d$hcluster <- hcom4",
"_____no_output_____"
],
[
"d",
"_____no_output_____"
],
[
"boxplot(height~hcluster, data=d)",
"_____no_output_____"
],
[
"boxplot(age~hcluster, data=d)",
"_____no_output_____"
],
[
"boxplot(inc~hcluster, data=d) ",
"_____no_output_____"
],
[
"aov(inc ~ as.factor(hcluster),data=d) ->ft\nsummary(ft)",
"_____no_output_____"
],
[
"aov(age ~ as.factor(hcluster),data=d) ->ft\nsummary(ft)",
"_____no_output_____"
],
[
"aov(height ~ as.factor(hcluster),data=d) ->ft\nsummary(ft)",
"_____no_output_____"
]
],
[
[
"## they are statistically significant but we will check for significance using silhouette test",
"_____no_output_____"
]
],
[
[
"install.packages('cluster',repos = \"http://cran.us.r-project.org\")",
"Installing package into 'C:/Users/vikas/Documents/R/win-library/3.3'\n(as 'lib' is unspecified)\n"
],
[
"library(cluster)",
"Warning message:\n\"package 'cluster' was built under R version 3.3.3\""
],
[
"silhouette(d$hcluster,dt)",
"_____no_output_____"
],
[
" plot(silhouette(d$hcluster,dt),col=1:4, border=NA)",
"_____no_output_____"
]
],
[
[
"## Mixed Data set",
"_____no_output_____"
]
],
[
[
"d2 <- data.frame(age= rnorm(10,34,2),gender=sample(0:1,10,repl=T),cal=sample(1:4,10,repl=T),satis=sample(1:5,10,repl=T))",
"_____no_output_____"
],
[
"str(d2)",
"'data.frame':\t10 obs. of 4 variables:\n $ age : num 33.8 33.5 37.5 30.8 32.6 ...\n $ gender: int 0 0 0 1 1 1 1 0 1 0\n $ cal : int 4 3 1 2 1 3 2 4 2 2\n $ satis : int 5 5 4 1 3 3 1 5 2 2\n"
],
[
"d2$gender <- as.factor(d2$gender)\nd2$cal <- as.factor(d2$cal)\nd2$satis <- as.ordered(d2$satis) ",
"_____no_output_____"
],
[
"str(d2)",
"'data.frame':\t10 obs. of 4 variables:\n $ age : num 33.8 33.5 37.5 30.8 32.6 ...\n $ gender: Factor w/ 2 levels \"0\",\"1\": 1 1 1 2 2 2 2 1 2 1\n $ cal : Factor w/ 4 levels \"1\",\"2\",\"3\",\"4\": 4 3 1 2 1 3 2 4 2 2\n $ satis : Ord.factor w/ 5 levels \"1\"<\"2\"<\"3\"<\"4\"<..: 5 5 4 1 3 3 1 5 2 2\n"
],
[
"daisy(d2,metho,metric=\"gower\") -> dt1",
"_____no_output_____"
],
[
"dt1",
"_____no_output_____"
],
[
"hclust(dt1,method=\"single\") -> hclustmixed\nplot(hclustmixed)",
"_____no_output_____"
],
[
"d2",
"_____no_output_____"
],
[
"print(max(d2$age))\nprint(min(d2$age))\nprint(max(d2$age)-min(d2$age))",
"[1] 37.48073\n[1] 30.764\n[1] 6.716723\n"
],
[
"cutree(hclustmixed,3) -> clustervector",
"_____no_output_____"
],
[
"clustervector",
"_____no_output_____"
],
[
"hclustmixed.",
"_____no_output_____"
],
[
"> df2 <- df1[sample(nrow(df1)),]",
"_____no_output_____"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4aceec92d6747e13da49c05b2851e7c6158fe13a
| 57,880 |
ipynb
|
Jupyter Notebook
|
Loan Cancellation Problem-tf3.ipynb
|
jnsxlx/Deep_learning
|
90f62f6063fb2d20dfd6f355c70423f51547c127
|
[
"MIT"
] | null | null | null |
Loan Cancellation Problem-tf3.ipynb
|
jnsxlx/Deep_learning
|
90f62f6063fb2d20dfd6f355c70423f51547c127
|
[
"MIT"
] | null | null | null |
Loan Cancellation Problem-tf3.ipynb
|
jnsxlx/Deep_learning
|
90f62f6063fb2d20dfd6f355c70423f51547c127
|
[
"MIT"
] | null | null | null | 39.400953 | 251 | 0.439409 |
[
[
[
"# To support both python 2 and python 3\nfrom __future__ import division, print_function, unicode_literals\n\n# Common imports\nimport os\nimport pickle\nimport timeit\n\n# numpy settings\nimport numpy as np\nnp.random.seed(42) # to make this notebook's output stable across runs\n\n# pandas settings\nimport pandas as pd\npd.set_option('display.max_columns', None)\n\n# matplotlib settings\n%matplotlib inline\nimport matplotlib\nimport matplotlib.pyplot as plt\nplt.rcParams['axes.labelsize'] = 14\nplt.rcParams['xtick.labelsize'] = 12\nplt.rcParams['ytick.labelsize'] = 12\n\n# Datasets\nDATASET_PATH = os.path.join(\".\", \"datasets\")\n\n# Figures\nIMAGE_PATH = os.path.join(\".\", \"images\")\nif not os.path.isdir(IMAGE_PATH):\n os.makedirs(IMAGE_PATH)\n\ndef save_fig(fig_id, tight_layout=True):\n path = os.path.join(IMAGE_PATH, fig_id + \".png\")\n print(\"Saving figure\", fig_id)\n if tight_layout:\n plt.tight_layout()\n plt.savefig(path, format='png', dpi=300)\n\n# Models\nMODEL_PATH = os.path.join(\".\", \"models\")\nif not os.path.isdir(MODEL_PATH):\n os.makedirs(MODEL_PATH)\n \n# Features\nFEATURE_PATH = os.path.join(\".\", \"features\")\nif not os.path.isdir(FEATURE_PATH):\n os.makedirs(FEATURE_PATH)\n\ndef save_features(feature_score, model_name):\n path = os.path.join(FEATURE_PATH, model_name + \".csv\")\n with open(path, 'w') as f:\n for idx in range(len(feature_score)):\n f.write(\"{0},{1}\\n\".format(feature_score[idx][0], feature_score[idx][1]))\n\n# Color for print\nclass color:\n PURPLE = '\\033[95m'\n CYAN = '\\033[96m'\n DARKCYAN = '\\033[36m'\n BLUE = '\\033[94m'\n GREEN = '\\033[92m'\n YELLOW = '\\033[93m'\n RED = '\\033[91m'\n BOLD = '\\033[1m'\n UNDERLINE = '\\033[4m'\n END = '\\033[0m'\n\n \n# Print a line\ndef print_lines(num, length):\n for i in range(num):\n print('-'*length)",
"_____no_output_____"
]
],
[
[
"***\n## Data Preprocessing\n\nThis section preprocesses the raw data.",
"_____no_output_____"
]
],
[
[
"# Import packages\nfrom sklearn.base import BaseEstimator, TransformerMixin\nfrom sklearn.feature_selection import SelectKBest",
"_____no_output_____"
],
[
"# Load data\nwith open(os.path.join(DATASET_PATH, 'new_theorem_data.p'), 'rb') as f:\n u = pickle._Unpickler(f)\n u.encoding = 'latin1'\n loan_data = u.load()",
"_____no_output_____"
]
],
[
[
"Labels are extracted and removed from the dataset.",
"_____no_output_____"
]
],
[
[
"# Lables processing\nloan_labels = loan_data['EnumListingStatus']\nloan_labels.replace(to_replace=7, value=0, inplace=True) # cancelled loans\nloan_labels.replace(to_replace=6, value=1, inplace=True) # originated loans\n# loan_labels.shape",
"_____no_output_____"
],
[
"# Correct data type\nbool_list = ['BoolIsLender']\nloan_data[bool_list] = loan_data[bool_list].astype(bool)\n\ncat_list = ['EnumListingCategory', 'EnumLoanFractionalType']\nfor cat in cat_list:\n loan_data[cat] = loan_data[cat].astype('object')",
"_____no_output_____"
]
],
[
[
"The feature selection needs careful treatment. \nSome features are identified as unnecessary either manually or by through model-based approach and are removed from the features. \nSpeficically, features with low scores (given by the classfier) are removed. ",
"_____no_output_____"
]
],
[
[
"# Drop unnecessary features\nfeature_drop_list = ['EnumListingStatus',\n 'ListingID',\n 'DateWholeLoanEnd',\n 'NumPublicRecords12',\n 'NumOpenTradesDelinqOrPastDue6',\n 'BoolInGroup',\n 'BoolOwnsHome',\n 'BoolIsLender',\n 'BoolPartialFundingApproved',\n 'BoolEverWholeLoan',\n 'BoolIsFractionalLoan',\n 'NumTradesCurr30DPDOrDerog6',\n 'NumTradesDelinqOrPastDue6',\n 'EnumLoanFractionalType',\n ]\nloan_data.drop(feature_drop_list, axis=1, inplace=True)\n \nprosper_list = ['NumPriorProsperLoans61dpd',\n 'NumPriorProsperLoans31dpd',\n 'NumPriorProsperLoansEarliestPayOff',\n 'NumPriorProsperLoansOnTimePayments',\n 'DolPriorProsperLoansPrincipalBorrowed',\n 'DolPriorProsperLoansPrincipalOutstanding',\n 'DolPriorProsperLoansBalanceOutstanding',\n 'NumPriorProsperLoansLatePayments',\n 'NumPriorProsperLoansCyclesBilled',\n 'NumPriorProsperLoansLateCycles',\n 'DolMaxPriorProsperLoan',\n 'DolMinPriorProsperLoan']",
"_____no_output_____"
]
],
[
[
"The following code defines a class that returns a sub-dataframe that consists of specified data types from a dataframe.",
"_____no_output_____"
]
],
[
[
"# Data selection\n\nclass DataFrameSelector(BaseEstimator, TransformerMixin):\n def __init__(self, dtypes):\n self.dtypes = dtypes\n def fit(self, X, y=None):\n return self\n def transform(self, X):\n return X.select_dtypes(self.dtypes).copy()\n\n\n# Test\n# DataFrameSelector([\"int64\"]).transform(loan_data).head()",
"_____no_output_____"
]
],
[
[
"The following code handles numerical features. `NAN` values are filled with 0 expcet for `FracDebtToIncomeRatio` and `ProsperScore`, which are filled with `mean` values. \n\nI tried to add some new features such as `IncomePaymentRatio` and `IncomeAmountRatio`. However, it turns out these new features do not contribute to the classification accuracy and thus are not adopted.\n\nIn fact, when time permits, it is recommended to generate various combinations of new features using both heuristics and pure techniques such as `PolynomialFeatures()` from `sklearn`. A good understanding of the loan market will definitely help.",
"_____no_output_____"
]
],
[
[
"# Numerical Feature Handling\n\nclass NumericalTransformer(BaseEstimator, TransformerMixin):\n def __init__(self):\n pass\n def fit(self, X, y=None):\n return self\n def transform(self, X, y=None):\n # Replace NA values\n for col in ['FracDebtToIncomeRatio', 'ProsperScore']:\n X[col].fillna(X[col].mean(), inplace=True)\n X.fillna(0, inplace=True)\n \n # Drop some features\n# if y is not None:\n# # Select K best features, X is 2-d array with each column representing a feature\n# kbest_selector = SelectKBest(lambda X, y: np.array(map(lambda x:mic(x, y), X.T)), k=2) \n# kbest_selector.fit_transform(X.values, y)\n# selected_cols = kbest_selector.get_support(indices=True)\n# X = X[selected_cols]\n\n # Generate new features\n# X['IncomePaymentRatio'] = X['DolMonthlyIncome'] / X['DolMonthlyLoanPayment']\n# X['IncomeAmountRatio'] = X['DolMonthlyIncome'] / X['DolLoanAmountRequested']\n \n return X\n\n\n# Test\n# loan_data_num = NumericalTransformer().transform(DataFrameSelector(['float64', 'int64']).transform(loan_data))\n# loan_data_num.head()",
"_____no_output_____"
]
],
[
[
"The following code handles categorical features. `NAN` values are filled with `-`. \nCategorical features are encoded as integers. Another popular encoding, `One-hot Enconding`, is also tested. \nYet, the benefit of using `One-hot Enconding` is not obvious here and thus is not used.",
"_____no_output_____"
]
],
[
[
"# Categorical features handling\n\nfrom sklearn.preprocessing import LabelEncoder\nfrom sklearn.preprocessing import OneHotEncoder\n\nclass CategoricalTransformer(BaseEstimator, TransformerMixin):\n def __init__(self, encoding='integer'):\n self.encoding = encoding\n def fit(self, X, y=None):\n return self\n def transform(self, X):\n # Replace NAN values\n X.fillna('-', inplace=True)\n \n # Encode\n le = LabelEncoder()\n X = X.apply(le.fit_transform) \n if self.encoding == 'onehot':\n ohe = OneHotEncoder()\n X = ohe.fit_transform(X.values)\n \n return X\n\n\n# Test\n# loan_data_cat = CategoricalTransformer().transform(DataFrameSelector(['object']).transform(loan_data))\n# loan_data_cat",
"_____no_output_____"
]
],
[
[
"The following code handles Datetime features. NAN values are filled with `2017-12-31 23:59:59`.\nAll Datetime values are converted to days from `1900-01-01 00:00:00`.\n\nTwo new features are added, i.e., `CreditLength` and `ListingTime`. ",
"_____no_output_____"
]
],
[
[
"# Datetime features handling\n\nclass DatetimeTransformer(BaseEstimator, TransformerMixin):\n def __init__(self):\n pass\n def fit(self, X, y=None):\n return self\n def transform(self, X):\n # Replace NA values\n X.fillna(pd.Timestamp('2017-12-31 23:59:59'), inplace=True)\n \n # Convert to days\n X['TimeBase'] = pd.Timestamp('1900-01-01 00:00:00')\n for col in X.columns:\n if col == 'TimeBase':\n pass\n else:\n X[col] = (X[col] - X['TimeBase']).dt.days\n X.drop(['TimeBase'], axis=1, inplace=True)\n \n # Generate new features\n X['CreditLength'] = X[\"DateCreditPulled\"] - X[\"DateFirstCredit\"]\n X['ListingTime'] = X[\"DateListingStart\"] - X[\"DateListingCreation\"]\n \n return X\n\n\n# Test\n# loan_data_dt = DatetimeTransformer().transform(DataFrameSelector(['datetime64']).transform(loan_data))\n# loan_data_dt.head()",
"_____no_output_____"
]
],
[
[
"The following code handles bool features. All values are converted to 0 or 1.",
"_____no_output_____"
]
],
[
[
"# Bool feature handling\n\nclass BoolTransformer(BaseEstimator, TransformerMixin):\n def __init__(self):\n pass\n def fit(self, X, y=None):\n return self\n def transform(self, X):\n X = X.astype(int)\n return X\n\n\n# Test\n# loan_data_bool = BoolTransformer().transform(DataFrameSelector(['bool']).transform(loan_data))\n# loan_data_bool.head()",
"_____no_output_____"
],
[
"# Scaler\nfrom sklearn.preprocessing import StandardScaler\n\nclass Scaler(BaseEstimator, TransformerMixin):\n def __init__(self):\n pass\n def fit(self, X, y=None):\n return self\n def transform(self, X):\n X = pd.DataFrame(StandardScaler().fit_transform(X.values),\n index=X.index, columns=X.columns)\n return X\n\n \n# Test\n# loan_data_scaled = Scaler().fit_transform(DataFrameSelector(['int64']).transform(loan_data))\n# loan_data_scaled.head()",
"_____no_output_____"
],
[
"# Transformation pipelines\n\nfrom sklearn.pipeline import Pipeline \n\nnum_pipeline = Pipeline([\n ('selector', DataFrameSelector(['float64', 'int64'])),\n ('transformer', NumericalTransformer()),\n ('scaler', Scaler()),\n ])\n\ncat_pipeline = Pipeline([\n ('selector', DataFrameSelector(['object'])),\n ('transformer', CategoricalTransformer()),\n ])\n \ndt_pipeline = Pipeline([\n ('selector', DataFrameSelector(['datetime64'])),\n ('transformer', DatetimeTransformer()),\n ('scaler', Scaler()),\n ])\n\nbool_pipeline = Pipeline([\n ('selector', DataFrameSelector(['bool'])),\n ('transformer', BoolTransformer()),\n ])\n\n# Pandas dataframe returned\nloan_data_clean = num_pipeline.fit_transform(loan_data)\nloan_data_clean = loan_data_clean.join(cat_pipeline.fit_transform(loan_data))\nloan_data_clean = loan_data_clean.join(dt_pipeline.fit_transform(loan_data))\nloan_data_clean = loan_data_clean.join(bool_pipeline.fit_transform(loan_data))\n# loan_data_clean.head()",
"/usr/local/lib/python3.6/site-packages/sklearn/utils/validation.py:475: DataConversionWarning: Data with input dtype int64 was converted to float64 by StandardScaler.\n warnings.warn(msg, DataConversionWarning)\n"
],
[
"# # Prepare data using FeatureUnion\n\n# from sklearn.pipeline import FeatureUnion\n\n# data_pipeline = FeatureUnion(transformer_list=[\n# (\"num_pipeline\", num_pipeline),\n# (\"cat_pipeline\", cat_pipeline),\n# (\"dt_pipeline\", dt_pipeline),\n# (\"bool_pipeline\", bool_pipeline)\n# ])\n\n# # Numpy array returned\n# loan_data_clean = data_pipeline.fit_transform(loan_data)",
"_____no_output_____"
],
[
"# Drop unnecessary features\nnull_cols = [\n 'DolPriorProsperLoansPrincipalBorrowed',\n 'DolPriorProsperLoansPrincipalOutstanding',\n 'DolPriorProsperLoansBalanceOutstanding',\n 'NumPriorProsperLoansCyclesBilled',\n 'NumPriorProsperLoansOnTimePayments',\n 'NumPriorProsperLoansLateCycles',\n 'NumPriorProsperLoansLatePayments',\n 'DolMaxPriorProsperLoan',\n 'DolMinPriorProsperLoan',\n 'NumPriorProsperLoansEarliestPayOff',\n 'NumPriorProsperLoans31dpd',\n 'NumPriorProsperLoans61dpd',\n 'DolMonthlyIncome'\n ]\nloan_data_clean.drop(null_cols, axis=1, inplace=True)",
"_____no_output_____"
],
[
"def drop_date_columns(data, col):\n data.drop(col, axis=1, inplace=True)\n return None",
"_____no_output_____"
],
[
"drop_date_columns(loan_data_clean, 'DateCreditPulled')\ndrop_date_columns(loan_data_clean, 'DateListingStart')\ndrop_date_columns(loan_data_clean, 'DateListingCreation')\ndrop_date_columns(loan_data_clean, 'DateFirstCredit')\ndrop_date_columns(loan_data_clean, 'DateWholeLoanStart')",
"_____no_output_____"
],
[
"# Split the data into a training set and a test set\n\n# Regular sampling\nfrom sklearn.model_selection import train_test_split\n\n# Full set for mode tunning\nX_train, X_test, y_train, y_test = train_test_split(loan_data_clean, loan_labels, test_size=0.2, random_state=42)\nprint([X_train.shape, y_train.shape, X_test.shape, y_test.shape])",
"[(201975, 56), (201975,), (50494, 56), (50494,)]\n"
],
[
"loan_data_clean.head()",
"_____no_output_____"
]
],
[
[
"## `Deep Learning`",
"_____no_output_____"
],
[
"#### Split data into 80/2 training/test",
"_____no_output_____"
]
],
[
[
"train_size = 80000\ntest_size = 2000\nX_train_s, y_train_s = X_train[:train_size], y_train[:train_size]\nX_test_s, y_test_s = X_test[:test_size], y_test[:test_size]\n\nprint([X_train_s.shape, y_train_s.shape, X_test_s.shape, y_test_s.shape])\nDL_X_train, DL_X_test = X_train_s.values, X_test_s.values\nDL_Y_train, DL_Y_test = y_train_s.values, y_test_s.values \nDL_Y_train = np.array([DL_Y_train, -(DL_Y_train-1)]).T\nDL_Y_test = np.array([DL_Y_test, -(DL_Y_test-1)]).T",
"[(80000, 56), (80000,), (2000, 56), (2000,)]\n"
],
[
"import tensorflow as tf",
"_____no_output_____"
],
[
"# Parameters\nlr_0 = 0.001\ndecay_rate = 9/1500\ntraining_epochs = 1000\nbatch_size = 256\ndisplay_step = 100\ndrop_rate_1 = 0.25\ndrop_rate_2 = 0.5\n\n\n# Network Parameters\nn_classes = 2 # Number of classes to predict\nn_inputs = X_train_s.shape[1] # Number of feature\nn_hidden_1 = 1024 # 1st layer number of features\nn_hidden_2 = 10 # 2st layer number of features\nn_hidden_3 = 128 # 3nd layer number of features\nn_hidden_4 = 128 # 4nd layer number of features\nn_hidden_5 = 64 # 5nd layer number of features\nn_hidden_6 = 64 # 6nd layer number of features\nn_hidden_7 = 64 # 7nd layer number of features",
"_____no_output_____"
]
],
[
[
"#### Construct 7-layer CNN with regularization, drop-out, batch normalization and relu as activation function",
"_____no_output_____"
]
],
[
[
"def dense(inputs, num_outputs, activation_fn):\n regularizer = tf.contrib.layers.l2_regularizer(scale=0.2)\n return tf.contrib.layers.fully_connected(inputs, num_outputs, activation_fn=None, weights_regularizer=regularizer)\n\ndef convnn(inputs, filt, knsize, activation_fn):\n return tf.layers.conv1d(inputs=inputs, filters=filt, kernel_size=knsize, padding=\"same\", activation=None)\n\ndef pooling(inputs):\n return tf.layers.max_pooling1d(inputs=inputs, pool_size=2, strides=2)\n \ndef dense_relu(inputs, num_outputs, is_training, scope):\n with tf.variable_scope(scope):\n h1 = dense(inputs, num_outputs, scope)\n return tf.nn.relu(h1, 'relu')\n\ndef dense_batch_relu(inputs, num_outputs, is_training, scope):\n with tf.variable_scope(scope):\n h1 = dense(inputs, num_outputs, scope)\n h2 = tf.contrib.layers.batch_norm(h1, center=True, scale=True, is_training=is_training, scope='bn')\n return tf.nn.relu(h2, 'relu')\n\ndef cnn_batch_relu(inputs, filt, knsize, is_training, scope):\n with tf.variable_scope(scope):\n h1 = convnn(inputs, filt, knsize, scope)\n# h2 = tf.contrib.layers.batch_norm(h1, center=True, scale=True, is_training=is_training, scope='bn')\n return tf.nn.relu(h1, 'relu')\n \ndef dense_dropout_relu(inputs, num_outputs, dropout_rate, is_training, scope):\n with tf.variable_scope(scope): \n h1 = dense(inputs, num_outputs, scope)\n h2 = tf.layers.dropout(inputs=h1, rate=dropout_rate, training=is_training)\n return tf.nn.relu(h2, 'relu')\n",
"_____no_output_____"
]
],
[
[
"#### Implement CNN and output accuracy and loss",
"_____no_output_____"
]
],
[
[
"tf.reset_default_graph()\nX = tf.placeholder('float32', [None, n_inputs, 1], name='X')\nY = tf.placeholder('float32', (None, n_classes), name='Y')\nis_training = tf.placeholder(tf.bool, name='is_training')\nlearning_rate = tf.placeholder('float32', name='lr')\n\ninput_layer = tf.reshape(X, [-1, n_inputs, 1])\nh1 = cnn_batch_relu(X, 32, 5, is_training=is_training, scope='cnn1')\nh2 = pooling(h1)\nh2_dr = tf.layers.dropout(inputs=h2, rate=drop_rate_1, training=is_training)\nh3 = cnn_batch_relu(h2_dr, 64, 5, is_training=is_training, scope='cnn2')\nh4 = pooling(h3)\nh4_dr = tf.layers.dropout(inputs=h4, rate=drop_rate_1, training=is_training)\nh4_flatten = tf.contrib.layers.flatten(h4_dr)\nh5 = dense_dropout_relu(inputs=h4_flatten, dropout_rate=drop_rate_2, num_outputs=n_hidden_1, is_training=is_training, scope='layer5')\nlogits = dense(inputs=h5, num_outputs=2, activation_fn='logits') \n\nwith tf.name_scope('accuracy'):\n accuracy = tf.reduce_mean(tf.cast(tf.equal(tf.argmax(Y, 1), tf.argmax(logits, 1)), 'float32'))\n\nwith tf.name_scope('loss'):\n loss = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits_v2(labels=Y, logits=logits))\n\nupdate_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)\nwith tf.control_dependencies(update_ops):\n # Ensures that we execute the update_ops before performing the train_step\n optimizer = tf.train.AdamOptimizer(learning_rate).minimize(loss)\n# optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)\n# gvs = optimizer.compute_gradients(loss)\n# capped_gvs = [(tf.clip_by_value(grad, -5., 5.), var) for grad, var in gvs]\n# train_op = optimizer.apply_gradients(capped_gvs)",
"_____no_output_____"
],
[
"init = tf.global_variables_initializer()\nwith tf.Session() as sess:\n sess.run(init)\n # Training cycle\n for epoch in range(training_epochs):\n avg_cost = 0.\n n_batches = int(len(DL_X_train)/batch_size)\n X_batches = np.array_split(DL_X_train, n_batches)\n Y_batches = np.array_split(DL_Y_train, n_batches)\n lr_value = 1/(1 + decay_rate * epoch) * lr_0\n # Loop over all batches\n for i in range(n_batches):\n batch_X, batch_Y = X_batches[i], Y_batches[i]\n batch_X = np.reshape(batch_X, (np.shape(batch_X)[0], n_inputs, 1))\n # Run optimization op (backprop) and cost op (to get loss value)\n# _, c = sess.run([optimizer, loss], feed_dict={X: batch_X, Y: batch_Y, is_training:1, learning_rate:lr_value})\n _, c = sess.run([optimizer, loss], feed_dict={X: batch_X, Y: batch_Y, is_training:1, learning_rate:lr_value})\n # Compute average loss\n avg_cost += c / n_batches\n # Display logs per epoch step\n if epoch % display_step == 0:\n print(\"Epoch:\", '%04d' % (epoch+1), \"cost=\", \"{:.9f}\".format(avg_cost))\n print(\"Optimization Finished!\")\n\n # Test model\n correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(Y, 1))\n # Calculate accuracy\n accuracy = tf.reduce_mean(tf.cast(correct_prediction, \"float\"))\n DL_X_train_tr = np.reshape(DL_X_train, (np.shape(DL_X_train)[0], n_inputs, 1))\n DL_X_test_tr = np.reshape(DL_X_test, (np.shape(DL_X_test)[0], n_inputs, 1))\n print(\"Training Accuracy:\", accuracy.eval({X: DL_X_train_tr, Y: DL_Y_train, is_training:1}))\n print(\"Test Accuracy:\", accuracy.eval({X: DL_X_test_tr, Y: DL_Y_test, is_training:0}))\n# global result \n# result = tf.argmax(pred, 1).eval({X: DL_X_test, Y: DL_Y_test})\n\n# # plot the cost\n# plt.plot(np.squeeze(loss))\n# plt.ylabel('cost')\n# plt.xlabel('iterations (per tens)')\n# plt.title(\"Learning rate =\" + str(learning_rate))\n# plt.show()",
"Epoch: 0001 cost= 31.617279328\nEpoch: 0101 cost= 0.561617220\nEpoch: 0201 cost= 0.543271641\nEpoch: 0301 cost= 0.521064070\nEpoch: 0401 cost= 0.506781861\nEpoch: 0501 cost= 0.497414144\nEpoch: 0601 cost= 0.488500934\nEpoch: 0701 cost= 0.480860585\nEpoch: 0801 cost= 0.474410257\nEpoch: 0901 cost= 0.471704989\nOptimization Finished!\nTraining Accuracy: 0.771\nTest Accuracy: 0.6925\n"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4acef280ef1fb7d47e354b8ff36845f601873254
| 522,349 |
ipynb
|
Jupyter Notebook
|
notebooks/Exploration and Processing Data.ipynb
|
Kolawole-Ikeoluwa-Joshua/Titanic-Data-science
|
02307448e34a9fcf7ae11f0225bad5fde4b5f7e4
|
[
"MIT"
] | null | null | null |
notebooks/Exploration and Processing Data.ipynb
|
Kolawole-Ikeoluwa-Joshua/Titanic-Data-science
|
02307448e34a9fcf7ae11f0225bad5fde4b5f7e4
|
[
"MIT"
] | null | null | null |
notebooks/Exploration and Processing Data.ipynb
|
Kolawole-Ikeoluwa-Joshua/Titanic-Data-science
|
02307448e34a9fcf7ae11f0225bad5fde4b5f7e4
|
[
"MIT"
] | null | null | null | 74.953221 | 49,508 | 0.720415 |
[
[
[
"# Exploring and Processing Data - Part 1",
"_____no_output_____"
]
],
[
[
"# imports\nimport pandas as pd\nimport numpy as np\nimport os",
"_____no_output_____"
]
],
[
[
"# Import Data",
"_____no_output_____"
]
],
[
[
"# set the path of the raw data\nraw_data_path = os.path.join(os.path.pardir, 'data', 'raw')\ntrain_file_path = os.path.join(raw_data_path, 'train.csv')\ntest_file_path = os.path.join(raw_data_path, 'test.csv')",
"_____no_output_____"
],
[
"# read the data with all default parameters\ntrain_df = pd.read_csv(train_file_path, index_col='PassengerId')\ntest_df = pd.read_csv(test_file_path, index_col='PassengerId')",
"_____no_output_____"
],
[
"# get the type\ntype(train_df)",
"_____no_output_____"
]
],
[
[
"# Basic Structure",
"_____no_output_____"
]
],
[
[
"# information about dataframe\ntrain_df.info()",
"<class 'pandas.core.frame.DataFrame'>\nInt64Index: 891 entries, 1 to 891\nData columns (total 11 columns):\nSurvived 891 non-null int64\nPclass 891 non-null int64\nName 891 non-null object\nSex 891 non-null object\nAge 714 non-null float64\nSibSp 891 non-null int64\nParch 891 non-null int64\nTicket 891 non-null object\nFare 891 non-null float64\nCabin 204 non-null object\nEmbarked 889 non-null object\ndtypes: float64(2), int64(4), object(5)\nmemory usage: 83.5+ KB\n"
],
[
"test_df.info()",
"<class 'pandas.core.frame.DataFrame'>\nInt64Index: 418 entries, 892 to 1309\nData columns (total 10 columns):\nPclass 418 non-null int64\nName 418 non-null object\nSex 418 non-null object\nAge 332 non-null float64\nSibSp 418 non-null int64\nParch 418 non-null int64\nTicket 418 non-null object\nFare 417 non-null float64\nCabin 91 non-null object\nEmbarked 418 non-null object\ndtypes: float64(2), int64(3), object(5)\nmemory usage: 35.9+ KB\n"
],
[
"# add survived column in test dataframe\ntest_df['Survived'] = -888",
"_____no_output_____"
],
[
"df = pd.concat((train_df, test_df), axis=0)",
"C:\\Users\\Scarllee\\Anaconda3\\lib\\site-packages\\ipykernel_launcher.py:1: FutureWarning: Sorting because non-concatenation axis is not aligned. A future version\nof pandas will change to not sort by default.\n\nTo accept the future behavior, pass 'sort=True'.\n\nTo retain the current behavior and silence the warning, pass sort=False\n\n \"\"\"Entry point for launching an IPython kernel.\n"
],
[
"df.info()",
"<class 'pandas.core.frame.DataFrame'>\nInt64Index: 1309 entries, 1 to 1309\nData columns (total 11 columns):\nAge 1046 non-null float64\nCabin 295 non-null object\nEmbarked 1307 non-null object\nFare 1308 non-null float64\nName 1309 non-null object\nParch 1309 non-null int64\nPclass 1309 non-null int64\nSex 1309 non-null object\nSibSp 1309 non-null int64\nSurvived 1309 non-null int64\nTicket 1309 non-null object\ndtypes: float64(2), int64(4), object(5)\nmemory usage: 122.7+ KB\n"
],
[
"# to get top 5 rows\ndf.head()",
"_____no_output_____"
],
[
"# use to get n-rows\ndf.head(10)",
"_____no_output_____"
],
[
"# use last 5 rows\ndf.tail()",
"_____no_output_____"
],
[
"# column selection using dot\ndf.Name",
"_____no_output_____"
],
[
"# selecting column using square brackets\ndf['Name']",
"_____no_output_____"
],
[
"# multiple selection\ndf[['Name', 'Age']]",
"_____no_output_____"
],
[
"# Indexing : use loc for label based indexing\n# all columns\ndf.loc[5:10,]",
"_____no_output_____"
],
[
"# selecting column range\ndf.loc[5:10, 'Age' : 'Pclass']",
"_____no_output_____"
],
[
"# discrete columns\ndf.loc[5:10, ['Age', 'Fare', 'Embarked']]",
"_____no_output_____"
],
[
"# indexing using iloc for postion based indexing\ndf.iloc[5:10, 3:8]",
"_____no_output_____"
],
[
"# filter rows based on the condition\nmale_passengers = df.loc[df.Sex == 'male',:]\nprint('Number of male passengers in first class: {0}'.format(len(male_passengers)))",
"Number of male passengers in first class: 843\n"
],
[
"# use & or | operators to build complex logic\nmale_passengers_first_class = df.loc[((df.Sex == 'male') & (df.Pclass == 1)),:]\nprint('Number of male passengers in first class: {0}'.format(len(male_passengers_first_class)))",
"Number of male passengers in first class: 179\n"
]
],
[
[
"# Summary Statistics",
"_____no_output_____"
]
],
[
[
"# to get summary statistics for all numeric columns\ndf.describe()",
"_____no_output_____"
],
[
"# numerical feature\n# centrality measures\n\nprint('Mean Fare : {0}'.format(df.Fare.mean()))\nprint('Median Fare : {0}'.format(df.Fare.median()))",
"Mean Fare : 33.2954792813456\nMedian Fare : 14.4542\n"
],
[
"# dispersion measures\nprint('Min fare : {0}'.format(df.Fare.min()))\nprint('Max fare : {0}'.format(df.Fare.max()))\nprint('Fare Range : {0}'.format(df.Fare.max() - df.Fare.min()))\nprint('25 Percentile : {0}'.format(df.Fare.quantile(.25)))\nprint('50 Percentile : {0}'.format(df.Fare.quantile(.5)))\nprint('75 Percentile : {0}'.format(df.Fare.quantile(.75)))\nprint('Variance fare : {0}'.format(df.Fare.var()))\nprint('Standard Deviation fare : {0}'.format(df.Fare.std()))\n\n\n",
"Min fare : 0.0\nMax fare : 512.3292\nFare Range : 512.3292\n25 Percentile : 7.8958\n50 Percentile : 14.4542\n75 Percentile : 31.275\nVariance fare : 2678.959737892894\nStandard Deviation fare : 51.75866823917414\n"
],
[
"# magic function to view plot\n%matplotlib inline",
"_____no_output_____"
],
[
"# box-whisker plot\ndf.Fare.plot(kind='box')",
"_____no_output_____"
],
[
"# get statistics for all columns including non-numeric ones\ndf.describe(include='all')",
"_____no_output_____"
],
[
"# categorical column: counts\ndf.Sex.value_counts()",
"_____no_output_____"
],
[
"# categorical column: proportions\ndf.Sex.value_counts(normalize=True)",
"_____no_output_____"
],
[
"# apply on other columns\ndf[df.Survived != -888].Survived.value_counts()",
"_____no_output_____"
],
[
"# count : passenger class\ndf.Pclass.value_counts()",
"_____no_output_____"
],
[
"# visualize counts\ndf.Pclass.value_counts().plot(kind='bar')",
"_____no_output_____"
],
[
"# to set title, color, and rotate labels\ndf.Pclass.value_counts().plot(kind='bar', rot = 0, title='Class wise passenger count', color='c');",
"_____no_output_____"
]
],
[
[
"# Distributions",
"_____no_output_____"
],
[
"## Univariate distributions",
"_____no_output_____"
]
],
[
[
"# use hist to create histogram\ndf.Age.plot(kind='hist', title='Histogram for Age', color='c');",
"_____no_output_____"
],
[
"# adding or removing bins\ndf.Age.plot(kind='hist', title='Histogram dor Age', color='c', bins=20);",
"_____no_output_____"
],
[
"# kde for density plot\ndf.Age.plot(kind='kde', title='Density plot for Age', color='c');",
"_____no_output_____"
],
[
"# histogram for fare\ndf.Fare.plot(kind='hist', title='histogram for Fare', color='c', bins=20);",
"_____no_output_____"
],
[
"print('skewness for age : {0:.2f}'.format(df.Age.skew()))\nprint('skewness for fare : {0:.2f}'.format(df.Fare.skew()))",
"skewness for age : 0.41\nskewness for fare : 4.37\n"
]
],
[
[
"### Bi-variate distribution",
"_____no_output_____"
]
],
[
[
"# scatter plot for bi-variate distribution\ndf.plot.scatter(x='Age', y='Fare', color='c', title='scatter plot : Age vs Fare');",
"_____no_output_____"
],
[
"# use to set the opacity of the dots\ndf.plot.scatter(x='Age', y='Fare', color='c', title='scatter plot : Age vs Fare', alpha=0.1);",
"_____no_output_____"
],
[
"df.plot.scatter(x='Pclass', y='Fare', color='c', title='scatter plot : Passenger class vs Fare', alpha=0.15);",
"_____no_output_____"
]
],
[
[
"# Grouping and Aggregations",
"_____no_output_____"
]
],
[
[
"# group by\ndf.groupby('Sex').Age.median()",
"_____no_output_____"
],
[
"# creating groups based on pclass\ndf.groupby(['Pclass']).Fare.median()",
"_____no_output_____"
],
[
"df.groupby(['Pclass'])['Fare', 'Age'].median()",
"_____no_output_____"
],
[
"# extracting more than one summary statistic\ndf.groupby(['Pclass']).agg({'Fare' : 'mean', 'Age' : 'median'})",
"_____no_output_____"
],
[
"# more complicated aggregations\naggregations = {\n 'Fare': { # work on the \"Fare\" column\n 'mean_Fare': 'mean',\n 'median_Fare': 'median',\n 'max_Fare': max,\n 'min_Fare': np.min\n },\n 'Age': { # work on the \"Age\" column\n 'median_Age': 'median',\n 'min_Age': min,\n 'max-Age': max,\n 'range_Age': lambda x: max(x) - min(x) #calculate the age range per group\n }\n}",
"_____no_output_____"
],
[
"df.groupby(['Pclass']).agg(aggregations)",
"C:\\Users\\Scarllee\\Anaconda3\\lib\\site-packages\\pandas\\core\\groupby\\groupby.py:4658: FutureWarning: using a dict with renaming is deprecated and will be removed in a future version\n return super(DataFrameGroupBy, self).aggregate(arg, *args, **kwargs)\n"
],
[
"df.groupby(['Pclass', 'Embarked']).Fare.median()",
"_____no_output_____"
]
],
[
[
"# Crosstabs",
"_____no_output_____"
]
],
[
[
"# crosstabs on Sex and Pclass\npd.crosstab(df.Sex, df.Pclass)",
"_____no_output_____"
],
[
"pd.crosstab(df.Sex, df.Pclass).plot(kind='bar');",
"_____no_output_____"
]
],
[
[
"# Pivot table",
"_____no_output_____"
]
],
[
[
"# Pivot table\n\ndf.pivot_table(index='Sex', columns='Pclass', values='Age', aggfunc='mean')",
"_____no_output_____"
],
[
"df.groupby(['Sex', 'Pclass']).Age.mean()",
"_____no_output_____"
],
[
"df.groupby(['Sex', 'Pclass']).Age.mean().unstack()",
"_____no_output_____"
]
],
[
[
"# Data Munging: Working with missing values",
"_____no_output_____"
]
],
[
[
"# detect missing values\ndf.info()",
"<class 'pandas.core.frame.DataFrame'>\nInt64Index: 1309 entries, 1 to 1309\nData columns (total 11 columns):\nAge 1046 non-null float64\nCabin 295 non-null object\nEmbarked 1307 non-null object\nFare 1308 non-null float64\nName 1309 non-null object\nParch 1309 non-null int64\nPclass 1309 non-null int64\nSex 1309 non-null object\nSibSp 1309 non-null int64\nSurvived 1309 non-null int64\nTicket 1309 non-null object\ndtypes: float64(2), int64(4), object(5)\nmemory usage: 162.7+ KB\n"
]
],
[
[
"### Feature: Embarked",
"_____no_output_____"
]
],
[
[
"# extract rows with Embarked as Null\ndf[df.Embarked.isnull()]",
"_____no_output_____"
],
[
"# find most common embarkment point\ndf.Embarked.value_counts()",
"_____no_output_____"
],
[
"# which embarkment point has higher survival count\npd.crosstab(df[df.Survived != -888].Survived, df[df.Survived != -888].Embarked)",
"_____no_output_____"
],
[
"# input the missing value with 'S'\n# df.loc[df.Embarked.isnull(), 'Embarked'] = 'S'\n# df.Embarked.fillna('S', inplace=True)",
"_____no_output_____"
],
[
"# Option 2 : explore the fare of each class for each embarkment point\ndf.groupby(['Pclass', 'Embarked']).Fare.median()",
"_____no_output_____"
],
[
"# replace the missing value with 'C'\ndf.Embarked.fillna('C', inplace=True)",
"_____no_output_____"
],
[
"# check if any null value is remaining\ndf[df.Embarked.isnull()]",
"_____no_output_____"
],
[
"# check info again\ndf.info()",
"<class 'pandas.core.frame.DataFrame'>\nInt64Index: 1309 entries, 1 to 1309\nData columns (total 11 columns):\nAge 1046 non-null float64\nCabin 295 non-null object\nEmbarked 1309 non-null object\nFare 1308 non-null float64\nName 1309 non-null object\nParch 1309 non-null int64\nPclass 1309 non-null int64\nSex 1309 non-null object\nSibSp 1309 non-null int64\nSurvived 1309 non-null int64\nTicket 1309 non-null object\ndtypes: float64(2), int64(4), object(5)\nmemory usage: 162.7+ KB\n"
]
],
[
[
"### Feature: Fare",
"_____no_output_____"
]
],
[
[
"df[df.Fare.isnull()]",
"_____no_output_____"
],
[
"median_fare = df.loc[(df.Pclass == 3) & (df.Embarked == 'S'), 'Fare'].median()\nprint(median_fare)",
"8.05\n"
],
[
"df.Fare.fillna(median_fare, inplace=True)",
"_____no_output_____"
],
[
"df.info()",
"<class 'pandas.core.frame.DataFrame'>\nInt64Index: 1309 entries, 1 to 1309\nData columns (total 11 columns):\nAge 1046 non-null float64\nCabin 295 non-null object\nEmbarked 1309 non-null object\nFare 1309 non-null float64\nName 1309 non-null object\nParch 1309 non-null int64\nPclass 1309 non-null int64\nSex 1309 non-null object\nSibSp 1309 non-null int64\nSurvived 1309 non-null int64\nTicket 1309 non-null object\ndtypes: float64(2), int64(4), object(5)\nmemory usage: 162.7+ KB\n"
]
],
[
[
"### Feature : Age",
"_____no_output_____"
]
],
[
[
"# set maximum number of rows to be displayed\npd.options.display.max_rows = 15",
"_____no_output_____"
],
[
"# return null rows\ndf[df.Age.isnull()]",
"_____no_output_____"
]
],
[
[
"### option 1: Replace all missing age with mean value",
"_____no_output_____"
]
],
[
[
"df.Age.plot(kind='hist', bins=20, color='c');",
"_____no_output_____"
],
[
"# get mean\ndf.Age.mean()",
"_____no_output_____"
]
],
[
[
"#### issue: due to few high values of 70's and 80's pushing the overall mean",
"_____no_output_____"
]
],
[
[
"# replace the missing values\n# df.Age.fillna(df.Age.mean(), inplace=True)",
"_____no_output_____"
]
],
[
[
"### option 2: replace with median age of gender",
"_____no_output_____"
]
],
[
[
"# median values\ndf.groupby('Sex').Age.median()",
"_____no_output_____"
],
[
"# boxplot to visualize\ndf[df.Age.notnull()].boxplot('Age', 'Sex');",
"_____no_output_____"
],
[
"# replace:\n# age_sex_median = df.groupby('Sex').Age.transform('median')\n# df.Age.fillna(age_sex_mediaan, inplace=True)",
"_____no_output_____"
]
],
[
[
"### option 3: replace with median age of Pclass\n",
"_____no_output_____"
]
],
[
[
"df[df.Age.notnull()].boxplot('Age', 'Pclass');",
"_____no_output_____"
],
[
"# replace:\n# pclass_age_median = df.groupby('Pclass').Age.transform('median')\n# df.Age.fillna(age_sex_mediaan, inplace=True)",
"_____no_output_____"
]
],
[
[
"### option 4: replace with median age of title",
"_____no_output_____"
]
],
[
[
"df.Name",
"_____no_output_____"
],
[
"# Function to extract the title from the name\ndef getTitle(name):\n first_name_with_title = name.split(',')[1]\n title = first_name_with_title.split('.')[0]\n title = title.strip().lower()\n return title",
"_____no_output_____"
],
[
"# use map function to apply the function on each Name value row i\ndf.Name.map(lambda x : getTitle(x)) # alternatively you can use : df.Name.map(getTitle)",
"_____no_output_____"
],
[
"df.Name.map(lambda x : getTitle(x)).unique()",
"_____no_output_____"
],
[
"# function to extract the title from the name\ndef GetTitle(name):\n title_group = {'mr' : 'Mr',\n 'mrs' : 'Mrs',\n 'miss' : 'Miss',\n 'master' : 'Master',\n 'don' : 'Sir',\n 'rev' : 'Sir',\n 'dr' : 'Officer',\n 'mme' : 'Mrs',\n 'ms' : 'Mrs',\n 'major' : 'Officer',\n 'lady' : 'Lady',\n 'sir' : 'Sir',\n 'mlle' : 'Miss',\n 'col' : 'Officer',\n 'capt' : 'Officer',\n 'the countess' : 'Lady',\n 'jonkheer' : 'Sir',\n 'dona' : 'Lady'\n }\n \n first_name_with_title = name.split(',')[1]\n title = first_name_with_title.split('.')[0]\n title = title.strip().lower()\n return title_group[title]",
"_____no_output_____"
],
[
"# create Title feature\ndf['Title'] = df.Name.map(lambda x : GetTitle(x))",
"_____no_output_____"
],
[
"# head\ndf.head()",
"_____no_output_____"
],
[
"# Box plot of Age with title\ndf[df.Age.notnull()].boxplot('Age', 'Title');",
"_____no_output_____"
],
[
"# replace missing values\ntitle_age_median = df.groupby('Title').Age.transform('median')\ndf.Age.fillna(title_age_median, inplace=True)",
"_____no_output_____"
],
[
"# check info\ndf.info()",
"<class 'pandas.core.frame.DataFrame'>\nInt64Index: 1309 entries, 1 to 1309\nData columns (total 12 columns):\nAge 1309 non-null float64\nCabin 295 non-null object\nEmbarked 1309 non-null object\nFare 1309 non-null float64\nName 1309 non-null object\nParch 1309 non-null int64\nPclass 1309 non-null int64\nSex 1309 non-null object\nSibSp 1309 non-null int64\nSurvived 1309 non-null int64\nTicket 1309 non-null object\nTitle 1309 non-null object\ndtypes: float64(2), int64(4), object(6)\nmemory usage: 172.9+ KB\n"
]
],
[
[
"# Working with outliers",
"_____no_output_____"
],
[
"# Age",
"_____no_output_____"
]
],
[
[
"# use histogram to understand distribution\ndf.Age.plot(kind='hist', bins=20, color='c');",
"_____no_output_____"
],
[
"df.loc[df.Age > 70]",
"_____no_output_____"
]
],
[
[
"# Fare",
"_____no_output_____"
]
],
[
[
"# histogram for fare\ndf.Fare.plot(kind='hist', title='histogram for Fare', bins=20, color='c');",
"_____no_output_____"
],
[
"# box plot to indentiify outliers\ndf.Fare.plot(kind='box');",
"_____no_output_____"
],
[
"# look into the outliers\ndf.loc[df.Fare == df.Fare.max()]",
"_____no_output_____"
],
[
"# Try some transformation to reduce the skewness\nLogFare = np.log(df.Fare + 1.0) # Adding 1 to accomodate zero fares : log(0) is not defined",
"_____no_output_____"
],
[
"# Histogram of Logfare\nLogFare.plot(kind='hist', color='c', bins=20);",
"_____no_output_____"
],
[
"# binning\npd.qcut(df.Fare, 4)",
"_____no_output_____"
],
[
"pd.qcut(df.Fare, 4, labels=['very_low', 'low', 'high', 'very_high']) #discretization",
"_____no_output_____"
],
[
"pd.qcut(df.Fare, 4, labels=['very_low', 'low', 'high', 'very_high']).value_counts().plot(kind='bar', color='c', rot=0);",
"_____no_output_____"
],
[
"# create fare bin feature\ndf['Fare_Bin'] = pd.qcut(df.Fare, 4, labels=['very_low', 'low', 'high', 'very_high'])",
"_____no_output_____"
]
],
[
[
"# Feature Engineering",
"_____no_output_____"
],
[
"### Feature: Age State (Adult or Child)",
"_____no_output_____"
]
],
[
[
"# AgeState based on Age\ndf['AgeState'] = np.where(df['Age'] >= 18, 'Adult', 'Child')",
"_____no_output_____"
],
[
"# AgeState counts\ndf['AgeState'].value_counts()",
"_____no_output_____"
],
[
"pd.crosstab(df[df.Survived != -888].Survived, df[df.Survived != -888].AgeState)",
"_____no_output_____"
]
],
[
[
"### Feature : FamilySize",
"_____no_output_____"
]
],
[
[
"# Family : Adding Parents with Siblings\ndf['FamilySize'] = df.Parch + df.SibSp + 1 # 1 for self",
"_____no_output_____"
],
[
"# explore the family feature\ndf['FamilySize'].plot(kind='hist', color='c');",
"_____no_output_____"
],
[
"# Explore families with max family members\ndf.loc[df.FamilySize == df.FamilySize.max(), ['Name', 'Survived', 'FamilySize', 'Ticket']]",
"_____no_output_____"
],
[
"pd.crosstab(df[df.Survived != -888].Survived, df[df.Survived != -888].FamilySize)",
"_____no_output_____"
]
],
[
[
"### Feature : IsMother",
"_____no_output_____"
]
],
[
[
"# a Lady who is aged more than 18 who has at least a child and is married\ndf['IsMother'] = np.where(((df.Sex == 'female') & (df.Parch > 0) & (df.Age > 18) & (df.Title != 'Miss')), 1, 0)",
"_____no_output_____"
],
[
"# crosstab with IsMother\npd.crosstab(df[df.Survived != -888].Survived, df[df.Survived != -888].IsMother)",
"_____no_output_____"
]
],
[
[
"### Feature : Deck",
"_____no_output_____"
]
],
[
[
"# explore Cabin values\ndf.Cabin",
"_____no_output_____"
],
[
"# use unique to get unqiue values for cabin feature\ndf.Cabin.unique()",
"_____no_output_____"
],
[
"# look at the cabin = T\ndf.loc[df.Cabin == 'T']",
"_____no_output_____"
],
[
"# set the value to NaN\ndf.loc[df.Cabin == 'T', 'Cabin'] = np.NaN",
"_____no_output_____"
],
[
"df.Cabin.unique()",
"_____no_output_____"
],
[
"# extract the first character of cabin string o the deck\ndef get_deck(cabin):\n return np.where(pd.notnull(cabin),str(cabin)[0].upper(), 'Z')\ndf['Deck'] = df['Cabin'].map(lambda x : get_deck(x))",
"_____no_output_____"
],
[
"# check counts\ndf.Deck.value_counts()",
"_____no_output_____"
],
[
"# use crosstab to look into survived feature cabin wise\npd.crosstab(df[df.Survived != -888].Survived, df[df.Survived != -888].Deck)",
"_____no_output_____"
],
[
"# info command\ndf.info()",
"<class 'pandas.core.frame.DataFrame'>\nInt64Index: 1309 entries, 1 to 1309\nData columns (total 17 columns):\nAge 1309 non-null float64\nCabin 294 non-null object\nEmbarked 1309 non-null object\nFare 1309 non-null float64\nName 1309 non-null object\nParch 1309 non-null int64\nPclass 1309 non-null int64\nSex 1309 non-null object\nSibSp 1309 non-null int64\nSurvived 1309 non-null int64\nTicket 1309 non-null object\nTitle 1309 non-null object\nFare_Bin 1309 non-null category\nAgeState 1309 non-null object\nFamilySize 1309 non-null int64\nIsMother 1309 non-null int32\nDeck 1309 non-null object\ndtypes: category(1), float64(2), int32(1), int64(5), object(8)\nmemory usage: 210.2+ KB\n"
]
],
[
[
"# Categorical Feature Encoding",
"_____no_output_____"
]
],
[
[
"# Sex or Gender\ndf['IsMale'] = np.where(df.Sex == 'male', 1, 0)",
"_____no_output_____"
],
[
"# columns Deck, Pclass, Title, AgeState\ndf = pd.get_dummies(df, columns=['Deck', 'Pclass', 'Title', 'Fare_Bin', 'Embarked', 'AgeState'])",
"_____no_output_____"
],
[
"print(df.info())",
"<class 'pandas.core.frame.DataFrame'>\nInt64Index: 1309 entries, 1 to 1309\nData columns (total 39 columns):\nAge 1309 non-null float64\nCabin 294 non-null object\nFare 1309 non-null float64\nName 1309 non-null object\nParch 1309 non-null int64\nSex 1309 non-null object\nSibSp 1309 non-null int64\nSurvived 1309 non-null int64\nTicket 1309 non-null object\nFamilySize 1309 non-null int64\nIsMother 1309 non-null int32\nIsMale 1309 non-null int32\nDeck_A 1309 non-null uint8\nDeck_B 1309 non-null uint8\nDeck_C 1309 non-null uint8\nDeck_D 1309 non-null uint8\nDeck_E 1309 non-null uint8\nDeck_F 1309 non-null uint8\nDeck_G 1309 non-null uint8\nDeck_Z 1309 non-null uint8\nPclass_1 1309 non-null uint8\nPclass_2 1309 non-null uint8\nPclass_3 1309 non-null uint8\nTitle_Lady 1309 non-null uint8\nTitle_Master 1309 non-null uint8\nTitle_Miss 1309 non-null uint8\nTitle_Mr 1309 non-null uint8\nTitle_Mrs 1309 non-null uint8\nTitle_Officer 1309 non-null uint8\nTitle_Sir 1309 non-null uint8\nFare_Bin_very_low 1309 non-null uint8\nFare_Bin_low 1309 non-null uint8\nFare_Bin_high 1309 non-null uint8\nFare_Bin_very_high 1309 non-null uint8\nEmbarked_C 1309 non-null uint8\nEmbarked_Q 1309 non-null uint8\nEmbarked_S 1309 non-null uint8\nAgeState_Adult 1309 non-null uint8\nAgeState_Child 1309 non-null uint8\ndtypes: float64(2), int32(2), int64(4), object(4), uint8(27)\nmemory usage: 197.2+ KB\nNone\n"
]
],
[
[
"# Drop and Reorder Columns",
"_____no_output_____"
]
],
[
[
"# drop columns\ndf.drop(['Cabin', 'Name', 'Ticket', 'Parch', 'SibSp', 'Sex'], axis=1, inplace=True)",
"_____no_output_____"
],
[
"# reorder columns\ncolumns = [column for column in df.columns if column != 'Survived']\ncolumns = ['Survived'] + columns\ndf = df[columns]",
"_____no_output_____"
],
[
"# check data frame status\ndf.info()",
"<class 'pandas.core.frame.DataFrame'>\nInt64Index: 1309 entries, 1 to 1309\nData columns (total 33 columns):\nSurvived 1309 non-null int64\nAge 1309 non-null float64\nFare 1309 non-null float64\nFamilySize 1309 non-null int64\nIsMother 1309 non-null int32\nIsMale 1309 non-null int32\nDeck_A 1309 non-null uint8\nDeck_B 1309 non-null uint8\nDeck_C 1309 non-null uint8\nDeck_D 1309 non-null uint8\nDeck_E 1309 non-null uint8\nDeck_F 1309 non-null uint8\nDeck_G 1309 non-null uint8\nDeck_Z 1309 non-null uint8\nPclass_1 1309 non-null uint8\nPclass_2 1309 non-null uint8\nPclass_3 1309 non-null uint8\nTitle_Lady 1309 non-null uint8\nTitle_Master 1309 non-null uint8\nTitle_Miss 1309 non-null uint8\nTitle_Mr 1309 non-null uint8\nTitle_Mrs 1309 non-null uint8\nTitle_Officer 1309 non-null uint8\nTitle_Sir 1309 non-null uint8\nFare_Bin_very_low 1309 non-null uint8\nFare_Bin_low 1309 non-null uint8\nFare_Bin_high 1309 non-null uint8\nFare_Bin_very_high 1309 non-null uint8\nEmbarked_C 1309 non-null uint8\nEmbarked_Q 1309 non-null uint8\nEmbarked_S 1309 non-null uint8\nAgeState_Adult 1309 non-null uint8\nAgeState_Child 1309 non-null uint8\ndtypes: float64(2), int32(2), int64(2), uint8(27)\nmemory usage: 135.9 KB\n"
]
],
[
[
"# Save Processed Dataset",
"_____no_output_____"
]
],
[
[
"processed_data_path = os.path.join(os.path.pardir, 'data', 'processed')\nwrite_train_path = os.path.join(processed_data_path, 'train.csv')\nwrite_test_path = os.path.join(processed_data_path, 'test.csv')",
"_____no_output_____"
],
[
"# train data\ndf.loc[df.Survived != -888].to_csv(write_train_path)\n# test data\ncolumns = [column for column in df.columns if column != 'Survived']\ndf.loc[df.Survived == -888, columns].to_csv(write_test_path)",
"_____no_output_____"
]
],
[
[
"# Building the data processing script",
"_____no_output_____"
]
],
[
[
"get_processed_data_script_file = os.path.join(os.path.pardir, 'src','data','get_processed_data.py')",
"_____no_output_____"
],
[
"%%writefile $get_processed_data_script_file\nimport numpy as np\nimport pandas as pd\nimport os\n\ndef read_data():\n # set the path of the raw data\n raw_data_path = os.path.join(os.path.pardir, 'data', 'raw')\n train_file_path = os.path.join(raw_data_path, 'train.csv')\n test_file_path = os.path.join(raw_data_path, 'test.csv')\n \n # read the data with all default parameters\n train_df = pd.read_csv(train_file_path, index_col='PassengerId')\n test_df = pd.read_csv(test_file_path, index_col='PassengerId')\n test_df['Survived'] = -888\n df = pd.concat((train_df, test_df), axis=0)\n return df\n\n\ndef process_data(df):\n # using the method chaining concept\n return (df\n # create title attribute - then add this\n .assign(Title = lambda x: x.Name.map(get_title))\n # working missing values - start with this\n .pipe(fill_missing_values)\n # create fare bin feature\n .assign(Fare_Bin = lambda x: pd.qcut(x.Fare, 4, labels=['very_low', 'low', 'high', 'very_high']))\n # create age state\n .assign(AgeState = lambda x: np.where(x.Age >= 18, 'Adult', 'Child'))\n .assign(FamilySize = lambda x: x.Parch + x.SibSp + 1)\n .assign(IsMother = lambda x : np.where(((x.Sex == 'female') & (x.Parch > 0) & (x.Age > 18) & (x.Title != 'Miss')), 1, 0))\n # create deck feature\n .assign(Cabin = lambda x: np.where(x.Cabin == 'T', np.nan, x.Cabin))\n .assign(Deck = lambda x: x.Cabin.map(get_deck))\n # feature encoding\n .assign(IsMale = lambda x : np.where(x.Sex == 'male', 1, 0))\n .pipe(pd.get_dummies, columns=['Deck', 'Pclass', 'Title', 'Fare_Bin', 'Embarked', 'AgeState'])\n # add code to drop unnecessary columns\n .drop(['Cabin', 'Name', 'Ticket', 'Parch', 'SibSp', 'Sex'], axis=1)\n # reorder columns\n .pipe(reorder_columns)\n )\n \n \ndef get_title(name):\n title_group = {'mr' : 'Mr',\n 'mrs' : 'Mrs',\n 'miss' : 'Miss',\n 'master' : 'Master',\n 'don' : 'Sir',\n 'rev' : 'Sir',\n 'dr' : 'Officer',\n 'mme' : 'Mrs',\n 'ms' : 'Mrs',\n 'major' : 'Officer',\n 'lady' : 'Lady',\n 'sir' : 'Sir',\n 'mlle' : 'Miss',\n 'col' : 'Officer',\n 'capt' : 'Officer',\n 'the countess' : 'Lady',\n 'jonkheer' : 'Sir',\n 'dona' : 'Lady'\n }\n \n first_name_with_title = name.split(',')[1]\n title = first_name_with_title.split('.')[0]\n title = title.strip().lower()\n return title_group[title]\n\n\ndef get_deck(cabin):\n return np.where(pd.notnull(cabin), str(cabin)[0].upper(), 'Z')\n\n\n\ndef fill_missing_values(df):\n # embarked\n df.Embarked.fillna('C', inplace=True)\n # fare\n median_fare = df[(df.Pclass == 3) & (df.Embarked == 'S')]['Fare'].median()\n df.Fare.fillna(median_fare, inplace=True)\n # age \n title_age_median = df.groupby('Title').Age.transform('median')\n df.Age.fillna(title_age_median, inplace=True)\n return df\n\n\ndef reorder_columns(df):\n columns = [column for column in df.columns if column != 'Survived']\n columns = ['Survived'] + columns\n df = df[columns]\n return df\n\n\ndef write_data(df):\n processed_data_path = os.path.join(os.path.pardir, 'data', 'processed')\n write_train_path = os.path.join(processed_data_path, 'train.csv')\n write_test_path = os.path.join(processed_data_path, 'test.csv')\n # train data\n df[df.Survived != -888].to_csv(write_train_path)\n # test data\n columns = [column for column in df.columns if column != 'Survived']\n df[df.Survived == -888][columns].to_csv(write_test_path)\n \nif __name__ == '__main__':\n df = read_data()\n df = process_data(df)\n write_data(df)",
"Overwriting ..\\src\\data\\get_processed_data.py\n"
],
[
"!python $get_processed_data_script_file",
"..\\src\\data\\get_processed_data.py:15: FutureWarning: Sorting because non-concatenation axis is not aligned. A future version\nof pandas will change to not sort by default.\n\nTo accept the future behavior, pass 'sort=True'.\n\nTo retain the current behavior and silence the warning, pass sort=False\n\n df = pd.concat((train_df, test_df), axis=0)\n"
]
],
[
[
"# Advanced visualization using MatPlotlib",
"_____no_output_____"
]
],
[
[
"import matplotlib.pyplot as plt\n%matplotlib inline",
"_____no_output_____"
],
[
"plt.hist(df.Age)",
"_____no_output_____"
],
[
"plt.hist(df.Age, bins=20, color='c')\nplt.show()",
"_____no_output_____"
],
[
"plt.hist(df.Age, bins=20, color='c')\nplt.title('Histogram : Age')\nplt.xlabel('Bins')\nplt.ylabel('Counts')\nplt.show()",
"_____no_output_____"
],
[
"f , ax = plt.subplots()\nax.hist(df.Age, bins=20, color='c')\nax.set_title('Histogram : Age')\nax.set_xlabel('Bins')\nax.set_ylabel('Counts')\nplt.show()",
"_____no_output_____"
],
[
"# Add subplots\nf , (ax1, ax2) =plt.subplots(1,2, figsize=(14,3))\n\nax1.hist(df.Fare, bins=20, color='c')\nax1.set_title('Histogram : Fare')\nax1.set_xlabel('Bins')\nax1.set_ylabel('Counts')\n\nax2.hist(df.Age, bins=20, color='tomato')\nax2.set_title('Histogram : Age')\nax2.set_xlabel('Bins')\nax2.set_ylabel('Counts')\n\nplt.show()",
"_____no_output_____"
],
[
"# Adding subplots\nf, ax_arr = plt.subplots(3,2, figsize=(14, 7))\n\n# plot 1\nax_arr[0,0].hist(df.Fare, bins=20, color='c')\nax_arr[0,0].set_title('Histogram : Fare')\nax_arr[0,0].set_xlabel('Bins')\nax_arr[0,0].set_ylabel('Counts')\n\n# plot 2\nax_arr[0,1].hist(df.Age, bins=20, color='c')\nax_arr[0,1].set_title('Histogram : Age')\nax_arr[0,1].set_xlabel('Bins')\nax_arr[0,1].set_ylabel('Counts')\n\n# plot 3\nax_arr[1,0].boxplot(df.Fare.values)\nax_arr[1,0].set_title('Boxplot : Fare')\nax_arr[1,0].set_xlabel('Fare')\nax_arr[1,0].set_ylabel('Fare')\n\n# plot 4\nax_arr[1,1].hist(df.Age.values)\nax_arr[1,1].set_title('Boxplot : Age')\nax_arr[1,1].set_xlabel('Age')\nax_arr[1,1].set_ylabel('Age')\n\n# plot 5\nax_arr[2,0].scatter(df.Age, df.Fare, color='c', alpha=0.15)\nax_arr[2,0].set_title('Scatter Plot : Age vs Fare')\nax_arr[2,0].set_xlabel('Age')\nax_arr[2,0].set_ylabel('Age')\n\n# remove empty subplot\nax_arr[2,1].axis('off')\n\nplt.tight_layout()\n\nplt.show()",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acef36f6a136f06bc1f490d5625acc6322e778a
| 7,070 |
ipynb
|
Jupyter Notebook
|
_build/jupyter_execute/KDP-2_operands_spectrometer.ipynb
|
Jonas231/OpticalDesignDocu_o
|
c99cad0621666aa708e0f4d8b1c3e9d8460d46f6
|
[
"MIT"
] | null | null | null |
_build/jupyter_execute/KDP-2_operands_spectrometer.ipynb
|
Jonas231/OpticalDesignDocu_o
|
c99cad0621666aa708e0f4d8b1c3e9d8460d46f6
|
[
"MIT"
] | null | null | null |
_build/jupyter_execute/KDP-2_operands_spectrometer.ipynb
|
Jonas231/OpticalDesignDocu_o
|
c99cad0621666aa708e0f4d8b1c3e9d8460d46f6
|
[
"MIT"
] | null | null | null | 33.507109 | 344 | 0.471429 |
[
[
[
"# Spectrometer\n\nThese are the operands of spectrometer.",
"_____no_output_____"
]
],
[
[
"import pandas as pd\nimport itables\nfrom itables import init_notebook_mode, show\nimport itables.options as opt\n\ninit_notebook_mode(all_interactive=True)\n\nopt.lengthMenu = [50, 100, 200, 500]\n#opt.classes = [\"display\", \"cell-border\"]\n#opt.classes = [\"display\", \"nowrap\"]\n\nopt.columnDefs = [{\"className\": \"dt-left\", \"targets\": \"_all\"}, {\"width\": \"500px\", \"targets\": 4}]\n#opt.maxBytes = 0\n#pd.get_option('display.max_columns')\n#pd.get_option('display.max_rows')\n\n\n",
"_____no_output_____"
],
[
"import os\ncwd = os.getcwd()\nfilename = os.path.join(cwd, os.path.join('Excel', 'KDP-2_optimization_operands.xlsx'))\n\ndf_spec = pd.read_excel(filename, sheet_name = \"spectrometer\", header = 1, index_col = 0)\ndf_spec = df_spec.dropna() # drop nan values\n\n",
"_____no_output_____"
],
[
"\ndf_spec\n",
"_____no_output_____"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code"
]
] |
4acef6999c8d6338a05de317b3361d076bb48382
| 21,397 |
ipynb
|
Jupyter Notebook
|
analysis/paper_main_eso.ipynb
|
RafalSkolasinski/simulation-codes-arxiv-1806.01815
|
fc5c27c6c625cc7f45cf5d7627eaafa3d3df4b9e
|
[
"BSD-2-Clause"
] | null | null | null |
analysis/paper_main_eso.ipynb
|
RafalSkolasinski/simulation-codes-arxiv-1806.01815
|
fc5c27c6c625cc7f45cf5d7627eaafa3d3df4b9e
|
[
"BSD-2-Clause"
] | null | null | null |
analysis/paper_main_eso.ipynb
|
RafalSkolasinski/simulation-codes-arxiv-1806.01815
|
fc5c27c6c625cc7f45cf5d7627eaafa3d3df4b9e
|
[
"BSD-2-Clause"
] | null | null | null | 209.77451 | 19,052 | 0.9184 |
[
[
[
"import xarray as xr\nimport numpy as np\nimport deepdish as dd\n\nfrom codes import misc\nimport matplotlib.pyplot as plt\n%matplotlib inline",
"_____no_output_____"
]
],
[
[
"# main figure 5",
"_____no_output_____"
]
],
[
[
"data = xr.open_dataset(b'data/paper_eso.nc')\ndata = data.sel(delta_epsilon=-0.05, shape='square', grid=0.5)\ndata = misc.reduce_dimensions(data)",
"_____no_output_____"
],
[
"labels = {\n 'both': 'Total',\n 'direct': 'Direct Rashba',\n 'rashba': 'Rashba'\n}\n \n\nplt.figure(figsize=(3, 3))\nfor soi in data.soi:\n x = data.sel(soi=soi).Ex.data\n y = data.sel(soi=soi).energy_so \n plt.plot(1000*x, 1000 * y, label=labels[str(soi.data)])\n\n \nplt.xlabel(r'$E_x$ (V/$\\mu$m)', fontsize=16)\nplt.ylabel(r'$E_\\mathrm{so} (meV)$', fontsize=16)\nplt.xlim([0,10])\nplt.tick_params(labelsize=15)\nplt.tick_params(labelsize=15)\nplt.legend(fontsize=12,frameon=False)\n\n\nplt.savefig('figures/fig_5inset.pdf',transparent=True, bbox_inches='tight', pad_inches=0.1)\nplt.savefig('figures/fig_5inset.png',transparent=True, bbox_inches='tight', pad_inches=0.1)",
"_____no_output_____"
]
]
] |
[
"code",
"markdown",
"code"
] |
[
[
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4acefd2767bf21bdc8dccb1332bb9325dccefedd
| 9,087 |
ipynb
|
Jupyter Notebook
|
02_CommandLine/20220220_Electiva_ComandLine_LinuxROS.ipynb
|
drozooecci/02_ElectivaI
|
22a85f92e8625f2f9201a47ac7de5c26ceefaf9a
|
[
"MIT"
] | null | null | null |
02_CommandLine/20220220_Electiva_ComandLine_LinuxROS.ipynb
|
drozooecci/02_ElectivaI
|
22a85f92e8625f2f9201a47ac7de5c26ceefaf9a
|
[
"MIT"
] | null | null | null |
02_CommandLine/20220220_Electiva_ComandLine_LinuxROS.ipynb
|
drozooecci/02_ElectivaI
|
22a85f92e8625f2f9201a47ac7de5c26ceefaf9a
|
[
"MIT"
] | null | null | null | 28.756329 | 308 | 0.545725 |
[
[
[
"# Electiva Técnica I - Introducción *Robot Operating System* (ROS1)\n\n### David Rozo Osorio, I.M, M.Sc.",
"_____no_output_____"
],
[
"## Introducción a Linux System\n\n- Objetivo: comprender el funcionamiento de un sistema operativo tipo Linux.\n\n\n- Procedimiento:\n 1. Características de la máquina virtual.\n 2. Introducción.\n 3. Características del S.O. - Gráfico.\n 4. Operaciones básicas con la Terminal.",
"_____no_output_____"
],
[
"### Características para una máquina virtual\n\nEl proceso de instalación puede ser implementado de 2 formas:\n\n- **Instalación con arranque paralelo**. En el siguiente [link](https://www.youtube.com/watch?v=x1ykDpSzpKU) puede visualizar el proceso. Nota: este proceso debe ser desarrollado con cuidado porque en caso de que se presente algún incidentes puede perder toda la información que tenga en computador.\n- **Máquina virtual** [link descarga](https://www.virtualbox.org/). En el siguiente [link](https://www.youtube.com/watch?v=x5MhydijWmc) puede visualizar el proceso (recomendado). Configuración máquina virtual (óptimo) [mínimo]:\n - Disco virtual 25GB o más. [15 GB].\n - Memoria RAM 4 GB o más. [2 GB].\n - CPU 2 o más. [1].\n - Se requiere de una instalación completa (no de prueba).\n - Se recomienda instalar la versión en Inglés de Ubuntu.\n - Ubuntu 20.04 https://ubuntu.com/download/desktop",
"_____no_output_____"
],
[
"<p style=\"page-break-after:always;\"></p>",
"_____no_output_____"
],
[
"### Introducción Ubuntu\n\nSistema Operativo basado en Unix, desarrollados como parte del Proyecto GNU.\n\n- Multitasking.\n- Multiusuario.\n- Software de uso libre y código abierto.\n- Distribuciones populares:\n - Ubuntu\n - Debian\n - Fedora\n\nEj. Android, utiliza un núcleo tipo Linux.",
"_____no_output_____"
],
[
"<img src=\"imag/ubuntuGUI.png\" width=400 height=400 />",
"_____no_output_____"
],
[
"<img src=\"imag/ubuntuFileSystem.png\" width=400 height=400 />",
"_____no_output_____"
],
[
"<img src=\"imag/ubuntuGUISoftware.png\" width=400 height=400 />",
"_____no_output_____"
],
[
"### CommandLine \n\n- `ls`: List Directory\n - `ls -a`: Extra parameter that shows all hidden elements in a folder.\n - `ls --help`: Extra parameters that present all extensions. \n \n- `cd`: Change Directory.\n - `cd ..`: Change directory to the previous.\n - `cd --`: Change to HOME directory.\n\n- `clear`: erase all content of the terminal window.\n- `pwd`: Current terminal path.\n- `mkdir`: Create a folder.\n- `rm`: Delete file.\n- `rmdir`: Delete folder.\n- `mv`: Move file.\n- `cp`: Copy file.\n- `wget`: Command to download files from the internet. \n- `sudo`: Run a command Admin Mode.\n- `apt-get`: Install Package Ubuntu.\n- `nano`: Text editor in terminal.\n- `sudo apt update`: update the Operating System.\n- `sudo apt upgrade`: erase older Operating System Files.\n\nNota: `man`: Open a manual of a command.\n",
"_____no_output_____"
],
[
"## Introducción a ROS1\n\n- Objetivo: comprender el funcionamiento y las características preliminares de ROS1\n\n\n- Procedimiento:\n 1. ROS - Master\n 2. ROS - Node operations\n 3. ROS - Topic operations\n 4. ROS - Launch\n 5. ROS - Demo",
"_____no_output_____"
],
[
"<img src=\"imag/ROSArq.png\" width=400 height=400 />",
"_____no_output_____"
],
[
"### CommandLine\n\n- `roscore`: inicia el ROS-Master, el servidor de parámetros y el nodo de logging.\n```bash\n$ roscore\n```\n- `rosnode`: comando que explora características de los nodos. Cada comando tiene una serie de modificadores o atributos que permiten realizar operaciones. Para identificar todos los atributos disponibles, se utiliza el modificador `-h` signficado \"*help*\"\n \n - **Ej**. Listar los nodos activos.\n```bash\n$ rosnode list\n```\n\n- `rostopic`: comando que provee información de los *topics* activos.\n\n - **Ej.** Listar *topics* activos.\n```bash\n$ rostopic list\n```\n - **Ej**. Imprimir información del *topic*.\n```bash\n$ rostopic echo /<topic_name>\n```\n - **Ej.** Publicar información en un *topic*.\n```bash\n$ rostopic pub <topic_name> <msg_type> data\n$ rostopic pub /hello std_msgs/String \"Hello\"\n```\n```bash\n$ rostopic pub -1 /hello std_msgs/String \"Hello\"\n$ rostopic pub /hello std_msgs/String -r1 -- \"Hello\"\n```\n\n- `rosmsg`: comando que provee información sobre los tipos de mensajes.\n - **Ej.** mensaje tipo *string*.\n```bash\n$ rosmsg show std_msgs/String\n```\n - **Ej.** mensaje tipo *Twist*.\n```bash\n$ rosmsg show geometry_msgs/Twist\n```\n\n- `rosrun`: comando que permite ejecutar nodos\n```bash\n$ rosrun <ros_pkg_name> <ros_node_name>\n$ rosrun roscpp_tutorials talker\n``` \n\n- `rqt_graph`: nodo que construye un gráfico de los *topic* y *node* activos.\n```bash\n$ rqt_graph\n```",
"_____no_output_____"
],
[
"### ROS Demo (Talker and Listener)\n\n- Iniciar ROS Master.\n```bash\n$ roscore\n```\n\n- Ejecutar nodos.\n\n - Iniciar el nodo \"talker\"\n```bash\n$ rosrun roscpp_tutorials talker\n```\n - Ejecutar el nodo \"listener\"\n```bash\n$ rosrun roscpp_tutorials listener\n```\n\n- Ejecutar comandos para *topics*\n```bash\n$ rostopic list\n```",
"_____no_output_____"
],
[
"### ROS Demo (Turtlesim)\n\n- Iniciar ROS Master.\n```bash\n$ roscore\n```\n\n- Ejecutar nodos.\n\n - Iniciar el nodo \"turtlesim_node\"\n```bash\n$ rosrun turtlesim turtlesim_node\n```\n - Ejecutar el nodo \"teleop\"\n```bash\n$ rosrun turtlesim turtle_teleop_key\n```\n - Ejecutar el nodo \"movimiento autónomo\"\n```bash\n$ rosrun turtlesim draw_square\n```\n\n- Ejecutar comandos para *topics*\n```bash\n$ rostopic list\n```\n```bash\n$ rostopic pub -1 /turtle1/cmd_vel geometry_msgs/Twist '[2.0,0.0,0.0]' '[0.0,0.0,1.0]'\n$ rostopic pub /turtle1/cmd_vel geometry_msgs/Twist -r1 -- '[2.0,0.0,0.0]' '[0.0,0.0,1.0]'\n```\n\n- Ejecutar el nodo \"*Graph*\"\n```bash\n$ rosrun rqt_graph rqt_graph\n```",
"_____no_output_____"
],
[
"## Referencias\n\n- Understanding ROS Nodes: http://wiki.ros.org/ROS/Tutorials/UnderstandingNodes\n\n- Understanding ROS Topics: http://wiki.ros.org/ROS/Tutorials/UnderstandingTopics\n\n- Lentin Joseph, Robot Operating System (ROS) for Absolute Beginners, Apress. 2018.\n\n- Ramkumar Gandhinathan, Lentin Joseph, ROS Robotics Projects, 2ed, Packt, 2019.\n\n- Morgan Quigley, Brian Gerkey, and William D. Smart., Programming Robots with ROS, O’ Reilly Media, Inc., 2016.",
"_____no_output_____"
]
]
] |
[
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
4aceff9341748d942d8684ab4150b12ad481962c
| 330,132 |
ipynb
|
Jupyter Notebook
|
code/Chapter 06 - Causal Emergence and the Emergence of Scale.ipynb
|
ajayjeswani12/NEW
|
44c9fdf03cbb6e37c8a2c43ff8bf3ac2a686ba82
|
[
"MIT"
] | 63 |
2019-07-19T09:30:18.000Z
|
2022-03-27T18:35:51.000Z
|
code/Chapter 06 - Causal Emergence and the Emergence of Scale.ipynb
|
ajayjeswani12/NEW
|
44c9fdf03cbb6e37c8a2c43ff8bf3ac2a686ba82
|
[
"MIT"
] | 1 |
2020-05-30T11:19:36.000Z
|
2020-05-30T11:19:36.000Z
|
code/Chapter 06 - Causal Emergence and the Emergence of Scale.ipynb
|
ajayjeswani12/NEW
|
44c9fdf03cbb6e37c8a2c43ff8bf3ac2a686ba82
|
[
"MIT"
] | 17 |
2019-07-22T18:52:03.000Z
|
2021-10-02T08:09:31.000Z
| 343.172557 | 53,472 | 0.911345 |
[
[
[
"from ei_net import *\nfrom ce_net import *\nimport matplotlib.pyplot as plt\n\nimport datetime as dt\n\n%matplotlib inline",
"_____no_output_____"
],
[
"##########################################\n############ PLOTTING SETUP ##############\nEI_cmap = \"Greys\"\nwhere_to_save_pngs = \"../figs/pngs/\"\nwhere_to_save_pdfs = \"../figs/pdfs/\"\nsave = True\nplt.rc('axes', axisbelow=True)\n##########################################\n##########################################",
"_____no_output_____"
]
],
[
[
"# The emergence of informative higher scales in complex networks",
"_____no_output_____"
],
[
"# Chapter 06 - Causal Emergence and the Emergence of Scale\n\n## Network Macroscales\nFirst we must introduce how to recast a network, $G$, at a higher scale. This is represented by a new network, $G_M$. Within $G_M$, a micro-node is a node that was present in the original $G$, whereas a macro-node is defined as a node, $v_M$, that represents a subgraph, $S_i$, from the original $G$ (replacing the subgraph within the network). Since the original network has been dimensionally reduced by grouping nodes together, $G_M$ will always have fewer nodes than $G$.\n\nA macro-node $\\mu$ is defined by some $W^{out}_{\\mu}$, derived from the edge weights of the various nodes within the subgraph it represents. One can think of a macro-node as being a summary statistic of the underlying subgraph's behavior, a statistic that takes the form of a single node. Ultimately there are many ways of representing a subgraph, that is, building a macro-node, and some ways are more accurate than others in capturing the subgraph's behavior, depending on the connectivity. To decide whether or not a macro-node is an accurate summary of its underlying subgraph, we check whether random walkers behave identically on $G$ and $G_M$. We do this because many important analyses and algorithms---such as using PageRank for determining a node's centrality or InfoMap for community discovery---are based on random walking.\n\nSpecifically, we define the *inaccuracy* of a macroscale as the Kullback-Leibler divergence between the expected distribution of random walkers on $G$ vs. $G_M$, given some identical initial distribution on each. The expected distribution over $G$ at some future time $t$ is $P_m(t)$, while the distribution over $G_M$ at some future time $t$ is $P_M(t)$. To compare the two, the distribution $P_m(t)$ is summed over the same nodes in the macroscale $G_M$, resulting in the distribution $P_{M|m}(t)$ (the microscale given the macroscale). We can then define the macroscale inaccuracy over some series of time steps $T$ as:\n\n$$ \\text{inaccuracy} = \\sum_{t=0}^T \\text{D}_{_{KL}}[P_{M}(t) || P_{M|m}(t)] $$\n\nThis measure addresses the extent to which a random dynamical process on the microscale topology will be recapitulated on a dimensionally-reduced topology.\n\nWhat constitutes an accurate macroscale depends on the connectivity of the subgraph that gets grouped into a macro-node. The $W^{out}_{\\mu}$ can be constructed based on the collective $W^{out}$ of the subgraph. For instance, in some cases one could just coarse-grain a subgraph by using its average $W^{out}$ as the $W^{out}_{\\mu}$ of some new macro-node $\\mu$. However, it may be that the subgraph has dependencies not captured by such a coarse-grain. Indeed, this is similar to the recent discovery that when constructing networks from data it is often necessary to explicitly model higher-order dependencies by using higher-order nodes so that the dynamics of random walks to stay true to the original data. We therefore introduce *higher-order macro-nodes* (HOMs), which draw on similar techniques to accurately represent subgraphs as single nodes.\n____________",
"_____no_output_____"
],
[
"\n<img src=\"../figs/pngs/CoarseGraining.png\" width=800>\n\n- Top: The original network, $G$ along with its adjacency matrix (left). The shaded oval indicates that subgraph $S$ member nodes $v_B$ and $v_C$ will be grouped together, forming a macro-node, ${\\mu}$. All macro-nodes are some transformation of the original adjacency matrix via recasting it as a new adjacency matrix (right). The manner of this recasting depends on the type of macro-node. \n- Bottom left: The simplest form of a macro-node is when $W^{out}_{\\mu}$ is an average of the $W^{out}_{i}$ of each node in the subgraph. \n- Bottom center left: A macro-node that represents some path-dependency, such as input from $A$. Here, in averaging to create the $W^{out}_{\\mu}$ the out-weights of nodes $v_B$ and $v_C$ are weighted by their input from $v_A$. \n- Bottom center right: A macro-node that represents the subgraph's output over the network's stationary dynamics. Each node has some associated ${\\pi}_{i}$, which is the probability of ${v}_{i}$ in the stationary distribution of the network. The $W^{out}_{\\mu}$ of a $\\mu | \\pi$ macro-node is created by weighting each $W^{out}_{i}$ of the micro-nodes in the subgraph $S$ by $\\frac{{\\pi}_{i}}{\\sum_{k \\in S} {\\pi}_{k}}$. \n- Bottom right: A macro-node with a single timestep delay between input $\\mu | j$ and its output $\\mu | \\pi$, each constructed using the same techniques as its components. However, $\\mu | j$ always deterministically outputs to $\\mu | \\pi$. ",
"_____no_output_____"
],
[
"Different subgraph connectivities require different types of HOMs to accurately represent. For instance, HOMs can be based on the input weights to the macro-node, which take the form $\\mu | j$. In these cases the $W^{out}_{\\mu|j}$ is a weighted average of each node's $W^{out}$ in the subgraph, where the weight is based on the input weight to each node in the subgraph. Another type of HOM that generally leads to accurate macro-nodes over time is when the $W^{out}_{\\mu}$ is based on the stationary output from the subgraph to the rest of the network, which we represent as $\\mu | \\pi$. These types of HOMs may sometimes have minor inaccuracies given some initial state, but will almost always trend toward perfect accuracy as the network approaches its stationary dynamics. \n\nSubgraphs with complex internal dynamics can require a more complex type of HOM in order to preserve the network's accuracy. For instance, in cases where subgraphs have a delay between their inputs and outputs, this can be represented by a combination of $\\mu | j$ and $\\mu | \\pi$, which when combined captures that delay. In these cases the macro-node $\\mu$ has two components, one of which acts as a buffer over a timestep. This means that macro-nodes can possess memory even when constructed from networks that are at the microscale memoryless, and in fact this type of HOM is sometimes necessary to accurately capture the microscale dynamics.\n\nWe present these types of macro-nodes not as an exhaustive list of all possible HOMs, but rather as examples of how to construct higher scales in a network by representing subgraphs as nodes, and also sometimes using higher-order dependencies to ensure those nodes are accurate. This approach offers a complete generalization of previous work on coarse-grains and also black boxes, while simultaneously solving the previously unresolved issue of macroscale accuracy by using higher-order dependencies. The types of macro-nodes formed by subgraphs also provides substantive information about the network, such as whether the macroscale of a network possesses memory or path-dependency.\n\n## Causal emergence reveals the scale of networks\n\nCausal emergence occurs when a recast network, $G_M$ (a macroscale), has more $EI$ than the original network, $G$ (the microscale). In general, networks with lower effectiveness (low $EI$ given their size) have a higher potential for causal emergence, since they can be recast to reduce their uncertainty. Searching across groupings allows the identification or approximation of a macroscale that maximizes the $EI$. \n\nChecking all possible groupings is computationally intractable for all but the smallest networks. Therefore, in order to find macro-nodes which increase the $EI$, we use a greedy algorithm that groups nodes together and checks if the $EI$ increases. By choosing a node and then pairing it iteratively with its surrounding nodes we can grow macro-nodes until pairings no longer increase the $EI$, and then move on to a new node.\n\nBy generating undirected preferential attachment networks and varying the degree of preferential attachment, $\\alpha$, we observe a crucial relationship between preferential attachment and causal emergence. One of the central results in network science has been the identification of \"scale-free\" networks. Our results show that networks that are not \"scale-free\" can be further separated into micro-, meso-, and macroscales depending on their connectivity. This scale can be identified based on their degree of causal emergence. In cases of sublinear preferential attachment ($\\alpha < 1.0$) networks lack higher scales. Linear preferential attachment ($\\alpha=1.0$) produces networks that are scale-free, which is the zone of preferential attachment right before the network develops higher scales. Such higher scales only exist in cases of superlinear preferential attachment ($\\alpha > 1.0$). And past $\\alpha > 3.0$ the network begins to converge to a macroscale where almost all the nodes are grouped into a single macro-node. The greatest degree of causal emergence is found in mesoscale networks, which is when $\\alpha$ is between 1.5 and 3.0, when networks possess a rich array of macro-nodes.\n\nCorrespondingly the size of $G_M$ decreases as $\\alpha$ increases and the network develops an informative higher scale, which can be seen in the ratio of macroscale network size, $N_M$, to the original network size, $N$. As discussed in previous sections, on the upper end of the spectrum of $\\alpha$ the resulting network will approximate a hub-and-spoke, star-like network. Star-like networks have higher degeneracy and thus less $EI$, and because of this, we expect that there are more opportunities to increase the network's $EI$ through grouping nodes into macro-nodes. Indeed, the ideal grouping of a star network is when $N_M=2$ and $EI$ is 1 bit. This result is similar to recent advances in spectral coarse-graining that also observe that the ideal coarse-graining of a star network is to collapse it into a two-node network, grouping all the spokes into a single macro-node, which is what happens to star networks that are recast as macroscales.\n\nOur results offer a principled and general approach to such community detection by asking when there is an informational gain from replacing a subgraph with a single node. Therefore we can define *causal communities* as being when a cluster of nodes, or some subgraph, forms a viable macro-node. Fundamentally causal communities represent noise at the microscale. The closer a subgraph is to complete noise, the greater the gain in $EI$ by replacing it with a macro-node. Minimizing the noise in a given network also identifies the optimal scale to represent that network. However, there must be some structure that can be revealed by noise minimization in the first place. In cases of random networks that form a single large component which lacks any such structure, causal emergence does not occur.\n____________",
"_____no_output_____"
],
[
"## 6.1 Causal Emergence in Preferential Attachment Networks",
"_____no_output_____"
]
],
[
[
"def preferential_attachment_network(N, alpha=1.0, m=1):\n \"\"\"\n Generates a network based off of a preferential attachment \n growth rule. Under this growth rule, new nodes place their \n $m$ edges to nodes already present in the graph, G, with \n a probability proportional to $k^\\alpha$.\n \n Params\n ------\n N (int): the desired number of nodes in the final network\n alpha (float): the exponent of preferential attachment. \n When alpha is less than 1.0, we describe it\n as sublinear preferential attachment. At\n alpha > 1.0, it is superlinear preferential\n attachment. And at alpha=1.0, the network \n was grown under linear preferential attachment,\n as in the case of Barabasi-Albert networks.\n m (int): the number of new links that each new node joins\n the network with.\n \n Returns\n -------\n G (nx.Graph): a graph grown under preferential attachment.\n \n \"\"\"\n G = nx.Graph()\n G = nx.complete_graph(m+1)\n\n for node_i in range(m+1,N):\n degrees = np.array(list(dict(G.degree()).values()))\n probs = (degrees**alpha) / sum(degrees**alpha)\n eijs = np.random.choice(G.number_of_nodes(), \n size=(m,), replace=False, p=probs)\n for node_j in eijs:\n G.add_edge(node_i, node_j)\n\n return G",
"_____no_output_____"
],
[
"Nvals = sorted([30,60,90,120,150])\nalphas= np.linspace(-1,5,25)\nNiter = 2\n\nm = 1\npa_ce = {'alpha' :[], \n 'N_micro':[],\n 'N_macro':[],\n 'EI_micro':[],\n 'EI_macro':[],\n 'CE' :[],\n 'N_frac' :[],\n 'runtime' :[]}",
"_____no_output_____"
]
],
[
[
"### Note: the following cell was run on a super-computing cluster. It is included as an example computation.",
"_____no_output_____"
]
],
[
[
"for N in Nvals:\n for alpha in alphas:\n for _ in range(Niter):\n G = preferential_attachment_network(N,alpha,m)\n\n startT = dt.datetime.now()\n CE = causal_emergence(G, printt=False)\n finisH = dt.datetime.now()\n\n diff = finisH-startT\n diff = diff.total_seconds()\n\n pa_ce['alpha'].append(alpha)\n pa_ce['N_micro'].append(N)\n pa_ce['N_macro'].append(CE['G_macro'].number_of_nodes())\n pa_ce['EI_micro'].append(CE['EI_micro'])\n pa_ce['EI_macro'].append(CE['EI_macro'])\n pa_ce['CE'].append(CE['EI_macro']-CE['EI_micro'])\n pa_ce['N_frac'].append(CE['G_macro'].number_of_nodes()/N)\n pa_ce['runtime'].append(diff)\n \nNCE = pa_ce.copy()",
"_____no_output_____"
],
[
"# import cmocean as cmo\n# colorz = cmo.cm.amp(np.linspace(0.2,0.9,len(Nvals)))\ncolorz = plt.cm.viridis(np.linspace(0,0.9,len(Nvals)))",
"_____no_output_____"
],
[
"mult=0.95\n\nfig,ax=plt.subplots(1,1,figsize=(5.0*mult,4.5*mult))\n\nplt.subplots_adjust(wspace=0.24, hspace=0.11)\nymax_so_far = 0\nxmin_so_far = 0\nxmax_so_far = 0\n\nfor i,Nn in enumerate(Nvals):\n col = colorz[i]\n means = [np.mean(NCE[Nn][i]['CE']) for i in NCE[Nn].keys()]\n stdvs = [np.std(NCE[Nn][i]['CE']) for i in NCE[Nn].keys()]\n alphs = list(NCE[Nn].keys())\n\n alphs = np.array([(alphs[i]+alphs[i+1])/2\n for i in range(0,len(alphs)-1,2)])\n means = np.array([(means[i]+means[i+1])/2\n for i in range(0,len(means)-1,2)])\n stdvs = np.array([(stdvs[i]+stdvs[i+1])/2\n for i in range(0,len(stdvs)-1,2)])\n \n xmin_so_far = min([xmin_so_far, min(alphs)])\n xmax_so_far = max([xmax_so_far, max(alphs)])\n ymax_so_far = max([ymax_so_far, max(means+stdvs)])\n\n ax.plot(alphs, means,\n markeredgecolor=col, color=col,\n markerfacecolor='w',\n markeredgewidth=1.5,markersize=5.0,\n linestyle='-',marker='o',linewidth=2.2,label='N = %i'%Nn)\n \n ax.fill_between(alphs, means-stdvs, means+stdvs,\n facecolor=col, alpha=0.2, \n edgecolors='w', linewidth=1)\n\ncols = [\"#a7d6ca\",\"#dbb9d1\",\"#d6cdae\",\"#a5c9e3\"] \nax.fill_between([-2,0.90],[-1,-1],[3,3], \n facecolor=cols[0],alpha=0.3,edgecolors='w',linewidth=0)\nax.fill_between([0.90,1.1],[-1,-1],[3,3],\n facecolor=cols[1],alpha=0.7,edgecolors='w',linewidth=0)\nax.fill_between([1.1,3.0],[-1,-1],[3,3],\n facecolor=cols[2],alpha=0.3,edgecolors='w',linewidth=0)\nax.fill_between([3.0,6],[-1,-1],[3,3],\n facecolor=cols[3],alpha=0.3,edgecolors='w',linewidth=0)\n \nax.text(-0.500, 2.65, '|', fontsize=14)\nax.text(0.9425, 2.65, '|', fontsize=14)\nax.text(0.9425, 2.72, '|', fontsize=14)\nax.text(0.9425, 2.79, '|', fontsize=14)\nax.text(2.4000, 2.65, '|', fontsize=14)\nax.text(4.2500, 2.65, '|', fontsize=14)\n \nax.text(-1.1, 2.81,'microscale',fontsize=12)\nax.text(0.35, 2.95,'scale-free',fontsize=12)\nax.text(1.70, 2.81,'mesoscale',fontsize=12)\nax.text(3.45, 2.81,'macroscale',fontsize=12)\n \nax.set_ylim(-0.025*ymax_so_far,ymax_so_far*1.05)\nax.set_xlim(-1.075,5*1.01)\nax.set_xlabel(r'$\\alpha$',fontsize=14)\nax.set_ylabel('Causal emergence',fontsize=14, labelpad=10)\nax.legend(loc=6,framealpha=0.99)\nax.set_xticks(np.linspace(-1,5,7))\nax.set_xticklabels([\"%i\"%i for i in np.linspace(-1,5,7)])\nax.grid(linestyle='-', linewidth=2.0, color='#999999', alpha=0.3)\n\nif save:\n plt.savefig(\n where_to_save_pngs+\\\n 'CE_pa_alpha_labs.png', \n dpi=425, bbox_inches='tight')\n plt.savefig(\n where_to_save_pdfs+\\\n 'CE_pa_alpha_labs.pdf', \n dpi=425, bbox_inches='tight')\n \nplt.show()",
"_____no_output_____"
],
[
"mult=0.95\n\nfig,ax=plt.subplots(1,1,figsize=(5.0*mult,4.5*mult))\n\nplt.subplots_adjust(wspace=0.24, hspace=0.11)\nymax_so_far = 0\nxmin_so_far = 0\nxmax_so_far = 0\n\nfor i,Nn in enumerate(Nvals):\n col = colorz[i]\n means = [np.mean(NCE[Nn][i]['N_frac']) for i in NCE[Nn].keys()]\n stdvs = [np.std(NCE[Nn][i]['N_frac']) for i in NCE[Nn].keys()]\n alphs = list(NCE[Nn].keys())\n\n alphs = np.array([(alphs[i]+alphs[i+1])/2 \n for i in range(0,len(alphs)-1,2)])\n means = np.array([(means[i]+means[i+1])/2 \n for i in range(0,len(means)-1,2)])\n stdvs = np.array([(stdvs[i]+stdvs[i+1])/2 \n for i in range(0,len(stdvs)-1,2)])\n\n xmin_so_far = min([xmin_so_far, min(alphs)])\n xmax_so_far = max([xmax_so_far, max(alphs)])\n ymax_so_far = max([ymax_so_far, max(means+stdvs)])\n \n ax.semilogy(alphs, means, markeredgecolor=col,\n color=col,markerfacecolor='w',\n markeredgewidth=1.5, markersize=5.0,\n linestyle='-',marker='o',linewidth=2.0,\n alpha=0.99,label='N = %i'%Nn)\n ax.fill_between(alphs, means-stdvs, means+stdvs, \n facecolor=col,alpha=0.2,\n edgecolors='w',linewidth=1)\n \ncols = [\"#a7d6ca\",\"#dbb9d1\",\"#d6cdae\",\"#a5c9e3\"] \nax.fill_between([-2,0.9],[-1,-1],[3,3],\n facecolor=cols[0],alpha=0.3,edgecolors='w',linewidth=0)\nax.fill_between([0.9,1.1],[-1,-1],[3,3],\n facecolor=cols[1],alpha=0.7,edgecolors='w',linewidth=0)\nax.fill_between([1.1,3.0],[-1,-1],[3,3],\n facecolor=cols[2],alpha=0.3,edgecolors='w',linewidth=0)\nax.fill_between([3.0,6],[-1,-1],[3,3],\n facecolor=cols[3],alpha=0.3,edgecolors='w',linewidth=0)\n \nax.text(-0.50, 1.036,'|', fontsize=14)\nax.text(0.935, 1.036,'|', fontsize=14)\nax.text(0.935, 1.170,'|', fontsize=14)\nax.text(0.935, 1.320,'|', fontsize=14)\nax.text(2.400, 1.036,'|', fontsize=14)\nax.text(4.250, 1.036,'|', fontsize=14)\n\nax.text(-1.1, 1.368, 'microscale', fontsize=12)\nax.text(0.35, 1.750, 'scale-free', fontsize=12)\nax.text(1.70, 1.368, 'mesoscale', fontsize=12)\nax.text(3.45, 1.368, 'macroscale', fontsize=12)\n \nax.set_ylim(0.009*ymax_so_far,ymax_so_far*1.075)\nax.set_xlim(-1.075,5*1.01)\nax.set_xlabel(r'$\\alpha$',fontsize=14)\nax.set_ylabel('Size ratio: macro to micro',\n fontsize=14, labelpad=2)\nax.legend(loc=6,framealpha=0.99)\nax.set_xticks(np.linspace(-1,5,7))\nax.set_xticklabels([\"%i\"%i for i in np.linspace(-1,5,7)])\nax.grid(linestyle='-', linewidth=2.0, color='#999999', alpha=0.3)\n\nif save:\n plt.savefig(\n where_to_save_pngs+\\\n 'Nfrac_pa_alpha_labs.png',\n dpi=425, bbox_inches='tight')\n plt.savefig(\n where_to_save_pdfs+\\\n 'Nfrac_pa_alpha_labs.pdf',\n dpi=425, bbox_inches='tight')\n\nplt.show()",
"_____no_output_____"
]
],
[
[
"_______________",
"_____no_output_____"
],
[
"## 6.2 Causal Emergence of Random Networks",
"_____no_output_____"
]
],
[
[
"Ns = [20,30,40,50]\nps = np.round(np.logspace(-3.25,-0.4,31),5)\nNiter = 40\n\ner_ce = {'p' :[], \n 'N_micro':[],\n 'N_macro':[],\n 'EI_micro':[],\n 'EI_macro':[],\n 'CE_mean' :[],\n 'CE_stdv' :[],\n 'N_frac' :[],\n 'runtime' :[]}\nER_CE = {N:er_ce for N in Ns}",
"_____no_output_____"
]
],
[
[
"### Note: the following cell was run on a super-computing cluster. It is included as an example computation.",
"_____no_output_____"
]
],
[
[
"for N in Ns: \n print(N, dt.datetime.now())\n er_ce = {'p' :[], \n 'N_micro':[],\n 'N_macro':[],\n 'EI_micro':[],\n 'EI_macro':[],\n 'CE_mean' :[],\n 'CE_stdv' :[],\n 'N_frac' :[],\n 'runtime' :[]}\n\n for p in ps:\n print('\\t',p)\n cee = []\n for rr in range(Niter):\n G = nx.erdos_renyi_graph(N,p)\n startT = dt.datetime.now()\n CE = causal_emergence(G,printt=False)\n finisH = dt.datetime.now()\n\n diff = finisH-startT\n diff = diff.total_seconds()\n ce = CE['EI_macro']-CE['EI_micro']\n cee.append(ce)\n \n er_ce['p'].append(p)\n er_ce['N_micro'].append(N)\n er_ce['N_macro'].append(CE['G_macro'].number_of_nodes())\n er_ce['EI_micro'].append(CE['EI_micro'])\n er_ce['EI_macro'].append(CE['EI_macro'])\n er_ce['CE_mean'].append(np.mean(cee))\n er_ce['CE_stdv'].append(np.std( cee))\n er_ce['runtime'].append(diff)\n \n ER_CE[N] = er_ce.copy()",
"_____no_output_____"
],
[
"# import cmocean as cmo\n# colors = cmo.cm.thermal(np.linspace(0.1,0.95,len(Ns)))\ncolors = plt.cm.viridis(np.linspace(0.0,1,len(Ns)))\n\ni = 0\nymax = 0\nplt.vlines(100, -1, 1, \n label=r'$\\langle k \\rangle=1$', linestyle='--',\n color=\"#333333\", linewidth=3.5, alpha=0.99)\nfor N in Ns:\n CE1 = np.array(ER_CE1[N]['CE_mean'].copy())\n CE2 = np.array(ER_CE2[N]['CE_mean'].copy())\n CE3 = np.array(ER_CE3[N]['CE_mean'].copy())\n CE4 = np.array(ER_CE4[N]['CE_mean'].copy())\n CE5 = np.array(ER_CE5[N]['CE_mean'].copy())\n CE6 = np.array(ER_CE6[N]['CE_mean'].copy())\n CEs = (CE1 + CE2 + CE3 + CE4 + CE5 + CE6)/6\n CEs = list(CEs)\n CEs = [(CEs[i] + CEs[i+1])/2 for i in range(0,len(CEs)-1)]\n CEs = [0] + CEs\n CEs.append(0)\n \n x1 = np.array(ER_CE1[N]['p'].copy())\n x2 = np.array(ER_CE2[N]['p'].copy())\n x3 = np.array(ER_CE3[N]['p'].copy())\n x4 = np.array(ER_CE4[N]['p'].copy())\n x5 = np.array(ER_CE5[N]['p'].copy())\n x6 = np.array(ER_CE6[N]['p'].copy())\n xx = (x1 + x2 + x3 + x4 + x5 + x6)/6\n xx = list(xx)\n xx = [(xx[i] + xx[i+1])/2 for i in range(0,len(xx)-1)]\n xx = [1e-4] + xx\n xx.append(1)\n \n std1 = np.array(ER_CE1[N]['CE_stdv'].copy())\n std2 = np.array(ER_CE2[N]['CE_stdv'].copy())\n std3 = np.array(ER_CE3[N]['CE_stdv'].copy())\n std4 = np.array(ER_CE4[N]['CE_stdv'].copy())\n std5 = np.array(ER_CE5[N]['CE_stdv'].copy())\n std6 = np.array(ER_CE6[N]['CE_stdv'].copy())\n stds = (std1 + std2 + std3 + std4 + std5 + std6)/6\n stds = list(stds)\n stds = [(stds[i] + stds[i+1])/2 for i in range(0,len(stds)-1)]\n stds = [0] + stds\n stds.append(0)\n\n ytop = np.array(CEs) + np.array(stds)\n ybot = np.array(CEs) - np.array(stds)\n ybot[ybot<0] = 0\n\n ymax = max([ymax, max(ytop)])\n \n plt.semilogx(xx, CEs, label='N=%i'%N, \n color=colors[i], linewidth=4.0, alpha=0.95)\n plt.vlines(1/(N-1), -1, 1, linestyle='--',\n color=colors[i], linewidth=3.5, alpha=0.95)\n i += 1\n\nplt.xlim(2.5e-4,max(xx))\nplt.ylim(-0.0015, ymax*0.6)\nplt.grid(linestyle='-', linewidth=2.5, alpha=0.3, color='#999999')\nplt.ylabel('Causal emergence', fontsize=14)\nplt.xlabel(r'$p$', fontsize=14)\nplt.legend(fontsize=12)\n\nif save:\n plt.savefig(\n where_to_save_pngs+\\\n 'CE_ER_p_N.png', dpi=425, bbox_inches='tight')\n plt.savefig(\n where_to_save_pdfs+\\\n 'CE_ER_p_N.pdf', dpi=425, bbox_inches='tight')\nplt.show()",
"/Users/brennan/anaconda3/lib/python3.7/site-packages/matplotlib/axes/_base.py:3099: UserWarning: Attempting to set identical left==right results\nin singular transformations; automatically expanding.\nleft=100.0, right=100.0\n self.set_xlim(upper, lower, auto=None)\n"
],
[
"# import cmocean as cmo\n# colors = cmo.cm.thermal(np.linspace(0.1,0.95,len(Ns)))\ncolors = plt.cm.viridis(np.linspace(0.0,1,len(Ns)))\n\ni = 0\nymax = 0\nplt.vlines(100, -1, 1, label=r'$\\langle k \\rangle=1$', linestyle='--',\n color=\"#333333\", linewidth=3.5, alpha=0.99)\nfor N in Ns:\n CE1 = np.array(ER_CE1[N]['CE_mean'].copy())\n CE2 = np.array(ER_CE2[N]['CE_mean'].copy())\n CE3 = np.array(ER_CE3[N]['CE_mean'].copy())\n CE4 = np.array(ER_CE4[N]['CE_mean'].copy())\n CE5 = np.array(ER_CE5[N]['CE_mean'].copy())\n CE6 = np.array(ER_CE6[N]['CE_mean'].copy())\n CEs = (CE1 + CE2 + CE3 + CE4 + CE5 + CE6)/6\n CEs = list(CEs)\n CEs = [(CEs[i] + CEs[i+1])/2 for i in range(0,len(CEs)-1)]\n CEs = [0] + CEs\n CEs.append(0)\n \n x1 = np.array(ER_CE1[N]['p'].copy())\n x2 = np.array(ER_CE2[N]['p'].copy())\n x3 = np.array(ER_CE3[N]['p'].copy())\n x4 = np.array(ER_CE4[N]['p'].copy())\n x5 = np.array(ER_CE5[N]['p'].copy())\n x6 = np.array(ER_CE6[N]['p'].copy())\n xx = (x1 + x2 + x3 + x4 + x5 + x6)/6\n xx = list(xx)\n xx = [(xx[i] + xx[i+1])/2 for i in range(0,len(xx)-1)]\n xx = [1e-4] + xx\n xx.append(1)\n \n std1 = np.array(ER_CE1[N]['CE_stdv'].copy())\n std2 = np.array(ER_CE2[N]['CE_stdv'].copy())\n std3 = np.array(ER_CE3[N]['CE_stdv'].copy())\n std4 = np.array(ER_CE4[N]['CE_stdv'].copy())\n std5 = np.array(ER_CE5[N]['CE_stdv'].copy())\n std6 = np.array(ER_CE6[N]['CE_stdv'].copy())\n stds = (std1 + std2 + std3 + std4 + std5 + std6)/6\n stds = list(stds)\n stds = [(stds[i] + stds[i+1])/2 for i in range(0,len(stds)-1)]\n stds = [0] + stds\n stds.append(0)\n\n ytop = np.array(CEs) + np.array(stds)\n ybot = np.array(CEs) - np.array(stds)\n ybot[ybot<0] = 0\n\n ymax = max([ymax, max(ytop)])\n \n plt.semilogx(xx, CEs, label='N=%i'%N, color=colors[i], \n linewidth=4.0, alpha=0.95)\n plt.fill_between(xx, ytop, ybot, facecolor=colors[i], \n linewidth=2.0, alpha=0.35, edgecolor='w')\n plt.vlines(1/(N-1), -1, 1, linestyle='--',\n color=colors[i], linewidth=3.5, alpha=0.95)\n i += 1\n\nplt.xlim(2.5e-4,max(xx))\nplt.ylim(-0.0015, ymax)\nplt.grid(linestyle='-', linewidth=2.5, \n alpha=0.3, color='#999999')\nplt.ylabel('Causal emergence', fontsize=14)\nplt.xlabel(r'$p$', fontsize=14)\nplt.legend(fontsize=12)\n\nif save:\n plt.savefig(\n where_to_save_pngs+\\\n 'CE_ER_p_N0.png', dpi=425, bbox_inches='tight')\n plt.savefig(\n where_to_save_pdfs+\\\n 'CE_ER_p_N0.pdf', dpi=425, bbox_inches='tight')\nplt.show()",
"/Users/brennan/anaconda3/lib/python3.7/site-packages/matplotlib/axes/_base.py:3099: UserWarning: Attempting to set identical left==right results\nin singular transformations; automatically expanding.\nleft=100.0, right=100.0\n self.set_xlim(upper, lower, auto=None)\n"
],
[
"for n in ER_CE1.keys():\n ER_CE1[n]['k'] = np.array(ER_CE1[n]['p'])*n\n \nfor n in ER_CE2.keys():\n ER_CE2[n]['k'] = np.array(ER_CE2[n]['p'])*n\n \nfor n in ER_CE3.keys():\n ER_CE3[n]['k'] = np.array(ER_CE3[n]['p'])*n\n \nfor n in ER_CE4.keys():\n ER_CE4[n]['k'] = np.array(ER_CE4[n]['p'])*n\n \nfor n in ER_CE5.keys():\n ER_CE5[n]['k'] = np.array(ER_CE5[n]['p'])*n\n \nfor n in ER_CE6.keys():\n ER_CE6[n]['k'] = np.array(ER_CE6[n]['p'])*n ",
"_____no_output_____"
],
[
"# import cmocean as cmo\n# colors = cmo.cm.thermal(np.linspace(0.1,0.95,len(Ns)))\ncolors = plt.cm.viridis(np.linspace(0.0,1,len(Ns)))\n\ni = 0\nymax = 0\n\nfor N in Ns:\n CE1 = np.array(ER_CE1[N]['CE_mean'].copy())\n CE2 = np.array(ER_CE2[N]['CE_mean'].copy())\n CE3 = np.array(ER_CE3[N]['CE_mean'].copy())\n CE4 = np.array(ER_CE4[N]['CE_mean'].copy())\n CE5 = np.array(ER_CE5[N]['CE_mean'].copy())\n CE6 = np.array(ER_CE6[N]['CE_mean'].copy())\n CEs = (CE1 + CE2 + CE3 + CE4 + CE5 + CE6)/6\n CEs = list(CEs)\n CEs = [(CEs[i] + CEs[i+1])/2 for i in range(0,len(CEs)-1)]\n CEs = [0] + CEs\n\n x1 = np.array(ER_CE1[N]['k'].copy())\n x2 = np.array(ER_CE2[N]['k'].copy())\n x3 = np.array(ER_CE3[N]['k'].copy())\n x4 = np.array(ER_CE4[N]['k'].copy())\n x5 = np.array(ER_CE5[N]['k'].copy())\n x6 = np.array(ER_CE6[N]['k'].copy())\n xx = (x1 + x2 + x3 + x4 + x5 + x6)/6\n xx = list(xx)\n xx = [(xx[i] + xx[i+1])/2 for i in range(0,len(xx)-1)]\n xx = [1e-4] + xx\n \n std1 = np.array(ER_CE1[N]['CE_stdv'].copy())\n std2 = np.array(ER_CE2[N]['CE_stdv'].copy())\n std3 = np.array(ER_CE3[N]['CE_stdv'].copy())\n std4 = np.array(ER_CE4[N]['CE_stdv'].copy())\n std5 = np.array(ER_CE5[N]['CE_stdv'].copy())\n std6 = np.array(ER_CE6[N]['CE_stdv'].copy())\n stds = (std1 + std2 + std3 + std4 + std5 + std6)/6\n stds = list(stds)\n stds = [(stds[i] + stds[i+1])/2 for i in range(0,len(stds)-1)]\n stds = [0] + stds\n\n ytop = np.array(CEs) + np.array(stds)\n ybot = np.array(CEs) - np.array(stds)\n ybot[ybot<0] = 0\n\n ymax = max([ymax, max(ytop)])\n \n plt.semilogx(xx, CEs, label='N=%i'%N, \n color=colors[i], linewidth=4.0, alpha=0.95)\n plt.fill_between(xx, ytop, ybot, \n facecolor=colors[i], \n linewidth=2.0, alpha=0.3, edgecolor='w')\n i += 1\n\nplt.vlines(1, -1, 1, linestyle='--',label=r'$\\langle k \\rangle=1$',\n color='k', linewidth=3.0, alpha=0.95)\n\nplt.xlim(1.0e-2,max(xx))\nplt.ylim(-0.0015, ymax*1.01)\nplt.grid(linestyle='-', linewidth=2.5, alpha=0.3, color='#999999')\nplt.ylabel('Causal emergence', fontsize=14)\nplt.xlabel(r'$\\langle k \\rangle$', fontsize=14)\nplt.legend(fontsize=12)\n\nif save:\n plt.savefig(\n where_to_save_pngs+\\\n 'CE_ER_k_N0.png', dpi=425, bbox_inches='tight')\n plt.savefig(\n where_to_save_pdfs+\\\n 'CE_ER_k_N0.pdf', dpi=425, bbox_inches='tight')\nplt.show()",
"_____no_output_____"
],
[
"# import cmocean as cmo\n# colors = cmo.cm.thermal(np.linspace(0.1,0.95,len(Ns)))\ncolors = plt.cm.viridis(np.linspace(0.0,1,len(Ns)))\n\ni = 0\nymax = 0\n\nfor N in Ns:\n CE1 = np.array(ER_CE1[N]['CE_mean'].copy())\n CE2 = np.array(ER_CE2[N]['CE_mean'].copy())\n CE3 = np.array(ER_CE3[N]['CE_mean'].copy())\n CE4 = np.array(ER_CE4[N]['CE_mean'].copy())\n CE5 = np.array(ER_CE5[N]['CE_mean'].copy())\n CE6 = np.array(ER_CE6[N]['CE_mean'].copy())\n CEs = (CE1 + CE2 + CE3 + CE4 + CE5 + CE6)/6\n CEs = list(CEs)\n CEs = [(CEs[i] + CEs[i+1])/2 for i in range(0,len(CEs)-1)]\n CEs = [0] + CEs\n \n x1 = np.array(ER_CE1[N]['k'].copy())\n x2 = np.array(ER_CE2[N]['k'].copy())\n x3 = np.array(ER_CE3[N]['k'].copy())\n x4 = np.array(ER_CE4[N]['k'].copy())\n x5 = np.array(ER_CE5[N]['k'].copy())\n x6 = np.array(ER_CE6[N]['k'].copy())\n xx = (x1 + x2 + x3 + x4 + x5 + x6)/6\n xx = list(xx)\n xx = [(xx[i] + xx[i+1])/2 for i in range(0,len(xx)-1)]\n xx = [1e-4] + xx\n \n std1 = np.array(ER_CE1[N]['CE_stdv'].copy())\n std2 = np.array(ER_CE2[N]['CE_stdv'].copy())\n std3 = np.array(ER_CE3[N]['CE_stdv'].copy())\n std4 = np.array(ER_CE4[N]['CE_stdv'].copy())\n std5 = np.array(ER_CE5[N]['CE_stdv'].copy())\n std6 = np.array(ER_CE6[N]['CE_stdv'].copy())\n stds = (std1 + std2 + std3 + std4 + std5 + std6)/6\n stds = list(stds)\n stds = [(stds[i] + stds[i+1])/2 for i in range(0,len(stds)-1)]\n stds = [0] + stds\n\n ytop = np.array(CEs) + np.array(stds)\n ybot = np.array(CEs) - np.array(stds)\n ybot[ybot<0] = 0\n\n ymax = max([ymax, max(ytop)])\n \n plt.semilogx(xx, CEs, label='N=%i'%N, \n color=colors[i], \n linewidth=4.0, alpha=0.95)\n i += 1\n\nplt.vlines(1, -1, 1, linestyle='--',\n label=r'$\\langle k \\rangle=1$',\n color='k', linewidth=3.0, alpha=0.95)\n\nplt.xlim(1.0e-2,max(xx))\nplt.ylim(-0.0015, ymax*0.6)\nplt.grid(linestyle='-', linewidth=2.5, \n alpha=0.3, color='#999999')\nplt.ylabel('Causal emergence', fontsize=14)\nplt.xlabel(r'$\\langle k \\rangle$', fontsize=14)\nplt.legend(fontsize=12)\n\nif save:\n plt.savefig(\n where_to_save_pngs+'CE_ER_k.png', \n dpi=425, bbox_inches='tight')\n plt.savefig(\n where_to_save_pdfs+'CE_ER_k.pdf', \n dpi=425, bbox_inches='tight')\nplt.show()",
"_____no_output_____"
]
],
[
[
"## End of Chapter 06. In [Chapter 07](https://nbviewer.jupyter.org/github/jkbren/einet/blob/master/code/Chapter%2007%20-%20Estimating%20Causal%20Emergence%20in%20Real%20Networks.ipynb) we'll estimate causal emergence in real networks.\n\n_______________",
"_____no_output_____"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"code",
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
]
] |
4acf048a28908cf04ff60129a661c73e76b0b670
| 66,141 |
ipynb
|
Jupyter Notebook
|
Python for ML/pandas.ipynb
|
ranjith-ms/ML_and_NN_course
|
ff22d43d1d5f94db45a169ded447d704ed100ea1
|
[
"Apache-2.0"
] | 1 |
2021-08-20T17:16:50.000Z
|
2021-08-20T17:16:50.000Z
|
Python for ML/pandas.ipynb
|
ranjith-ms/ML_and_NN_course
|
ff22d43d1d5f94db45a169ded447d704ed100ea1
|
[
"Apache-2.0"
] | null | null | null |
Python for ML/pandas.ipynb
|
ranjith-ms/ML_and_NN_course
|
ff22d43d1d5f94db45a169ded447d704ed100ea1
|
[
"Apache-2.0"
] | null | null | null | 66,141 | 66,141 | 0.494141 |
[
[
[
"import pandas as pd\n",
"_____no_output_____"
],
[
"from google.colab import drive\ndrive.mount('/content/drive')",
"Go to this URL in a browser: https://accounts.google.com/o/oauth2/auth?client_id=947318989803-6bn6qk8qdgf4n4g3pfee6491hc0brc4i.apps.googleusercontent.com&redirect_uri=urn%3aietf%3awg%3aoauth%3a2.0%3aoob&response_type=code&scope=email%20https%3a%2f%2fwww.googleapis.com%2fauth%2fdocs.test%20https%3a%2f%2fwww.googleapis.com%2fauth%2fdrive%20https%3a%2f%2fwww.googleapis.com%2fauth%2fdrive.photos.readonly%20https%3a%2f%2fwww.googleapis.com%2fauth%2fpeopleapi.readonly\n\nEnter your authorization code:\n··········\nMounted at /content/drive\n"
],
[
"%cd /content/drive/My Drive/Colab Notebooks/ML_and_NN_course/Python for ML",
"/content/drive/My Drive/Colab Notebooks/ML_and_NN_course/Python for ML\n"
]
],
[
[
"# Pandas \nPandas is a high-level data manipulation tool developed by Wes McKinney. It is built on the Numpy package and its key data structure is called the DataFrame. DataFrames allow you to store and manipulate tabular data in rows of observations and columns of variables.There are several ways to create a DataFrame.\n",
"_____no_output_____"
],
[
"## Dataframe\npd.read_csv() -> https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.read_csv.html",
"_____no_output_____"
]
],
[
[
"df=pd.read_csv('kc_house_data.csv') #specifying the directory\ndf.head()",
"_____no_output_____"
],
[
"#Specifying the URL\ndf1=pd.read_csv('https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv')",
"_____no_output_____"
],
[
"df1.head()",
"_____no_output_____"
],
[
"data=pd.DataFrame()\n\ndata['x']=[1,2,0,3,4] \ndata['y']=[1,2,0,3,4] \n\ndata.insert(0, 'Ones', 2)\ndata.head()",
"_____no_output_____"
],
[
"data = pd.DataFrame({'animal': ['alligator', 'bee', 'falcon', 'lion',\n 'monkey', 'parrot', 'shark', 'whale', 'zebra']})\ndata.head()",
"_____no_output_____"
]
],
[
[
"## dataframe.describe()\nhttps://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.describe.html",
"_____no_output_____"
]
],
[
[
"df.describe()",
"_____no_output_____"
]
],
[
[
"## shape:\nPandas .size, .shape and .ndim are used to return size, shape and dimensions of data frames and series.\n\n#### Syntax: dataframe.size\n\nReturn : Returns size of dataframe/series which is equivalent to total number of elements. That is rows x columns.\n\n#### Syntax: dataframe.shape\n\nReturn : Returns tuple of shape (Rows, columns) of dataframe/series\n\n#### Syntax: dataframe.ndim\n\nReturn : Returns dimension of dataframe/series\n",
"_____no_output_____"
]
],
[
[
"df.size",
"_____no_output_____"
],
[
"df.shape",
"_____no_output_____"
],
[
"df.ndim #bcz only row and columns",
"_____no_output_____"
]
],
[
[
"## df.columns",
"_____no_output_____"
]
],
[
[
"df.columns",
"_____no_output_____"
]
],
[
[
"## Selecting a column",
"_____no_output_____"
]
],
[
[
"a=df['lat'] \nb= df.lat\nb.head()",
"_____no_output_____"
]
],
[
[
"## Slicing",
"_____no_output_____"
]
],
[
[
"df.head()",
"_____no_output_____"
],
[
"a=df.iloc[:,0:2]\na.head()",
"_____no_output_____"
],
[
"a=df.iloc[2:,:5]\na.head()",
"_____no_output_____"
]
],
[
[
"## Mean and std",
"_____no_output_____"
]
],
[
[
"import numpy as np",
"_____no_output_____"
],
[
"mn=np.mean(df)",
"_____no_output_____"
],
[
"mn.shape",
"_____no_output_____"
],
[
"df.std()",
"_____no_output_____"
]
],
[
[
"## Droping a column\nhttps://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.drop.html",
"_____no_output_____"
]
],
[
[
"df.head()",
"_____no_output_____"
],
[
"df = df.drop(['lat', 'long'],axis=1)",
"_____no_output_____"
],
[
"df.head()",
"_____no_output_____"
],
[
"df['bedrooms'] +=2\ndf.head()",
"_____no_output_____"
]
],
[
[
"## dataframe to csv",
"_____no_output_____"
]
],
[
[
"df.to_csv(\"a.csv\")",
"_____no_output_____"
],
[
"",
"_____no_output_____"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
4acf09b963796f1aa51bb044bce964c8b1248db3
| 1,399 |
ipynb
|
Jupyter Notebook
|
New_Key_12.ipynb
|
microprediction/streamcreator
|
6e37c48252f75276189a9fd2a7230e7f4c54cd07
|
[
"MIT"
] | null | null | null |
New_Key_12.ipynb
|
microprediction/streamcreator
|
6e37c48252f75276189a9fd2a7230e7f4c54cd07
|
[
"MIT"
] | null | null | null |
New_Key_12.ipynb
|
microprediction/streamcreator
|
6e37c48252f75276189a9fd2a7230e7f4c54cd07
|
[
"MIT"
] | 1 |
2021-12-19T16:32:16.000Z
|
2021-12-19T16:32:16.000Z
| 26.396226 | 236 | 0.533953 |
[
[
[
"<a href=\"https://colab.research.google.com/github/microprediction/streamcreator/blob/main/New_Key_12.ipynb\" target=\"_parent\"><img src=\"https://colab.research.google.com/assets/colab-badge.svg\" alt=\"Open In Colab\"/></a>",
"_____no_output_____"
]
],
[
[
"!pip install microprediction",
"_____no_output_____"
],
[
"from microprediction import new_key\n# This will take a while. https://www.microprediction.com/contact-us if you are in a hurry\nwrite_key_12 = new_key(difficulty=12) # Strong enough to create you own streams\nprint(write_key_12)\n",
"_____no_output_____"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code"
]
] |
4acf0def4b5d9296da81f65bff05aaba81823e04
| 3,991 |
ipynb
|
Jupyter Notebook
|
notebooks/prepare_dataset.ipynb
|
alexander-pv/insects-recognition
|
40f7a29425ffaf04ac1fee319841484885dcbdd9
|
[
"MIT"
] | null | null | null |
notebooks/prepare_dataset.ipynb
|
alexander-pv/insects-recognition
|
40f7a29425ffaf04ac1fee319841484885dcbdd9
|
[
"MIT"
] | null | null | null |
notebooks/prepare_dataset.ipynb
|
alexander-pv/insects-recognition
|
40f7a29425ffaf04ac1fee319841484885dcbdd9
|
[
"MIT"
] | null | null | null | 22.421348 | 380 | 0.498121 |
[
[
[
"### Prepare dataset for training",
"_____no_output_____"
],
[
"To prepare your custom dataset for model training, create folder `datasets` in the project folder and place your images to the `datasets/your_dataset_name/raw` subfodler. To distinguish classes during data preparation, split your images to folders in `raw` subfolder. Each folder name in `raw` will be noted as a specific class name. Use only jpeg/jpg, png image formats.\n\nFor example, for `datasets/adelphocoris_net` the file structure can be organised as:\n\n adelphocoris_net/\n ├── raw\n │ ├── Adelphocoris_lineolatus\n │ ├── Adelphocoris_qudripunctatus\n │ ├── Adelphocoris_seticornis\n │ └── Adelphocoris_vandalicus\n\n",
"_____no_output_____"
]
],
[
[
"import os",
"_____no_output_____"
],
[
"os.chdir(os.path.join('..', 'src'))\nos.listdir()",
"_____no_output_____"
],
[
"import prepare_dataset",
"_____no_output_____"
],
[
"# Set folder name of a dataset to DATASET_FOLDER\nDATASET_FOLDER = 'test_data'\n# Prepare dataset\nprepare_dataset.register_image_dataset(data_folder=DATASET_FOLDER,\n work_dir=os.getcwd())",
"\r 0%| | 0/2 [00:00<?, ?it/s]"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
]
] |
4acf1194ebf52d5a1cd9d6a25ed88f2a5bec0f37
| 654,082 |
ipynb
|
Jupyter Notebook
|
module3-ridge-regression/LS_DS_213_assignment_ALEX_KAISER.ipynb
|
Lord-Kanzler/DS-Unit-2-Linear-Models
|
f76c27694cbc116ce84880fe7813edcacda5cc9e
|
[
"MIT"
] | null | null | null |
module3-ridge-regression/LS_DS_213_assignment_ALEX_KAISER.ipynb
|
Lord-Kanzler/DS-Unit-2-Linear-Models
|
f76c27694cbc116ce84880fe7813edcacda5cc9e
|
[
"MIT"
] | null | null | null |
module3-ridge-regression/LS_DS_213_assignment_ALEX_KAISER.ipynb
|
Lord-Kanzler/DS-Unit-2-Linear-Models
|
f76c27694cbc116ce84880fe7813edcacda5cc9e
|
[
"MIT"
] | null | null | null | 214.172233 | 78,382 | 0.849355 |
[
[
[
"<a href=\"https://colab.research.google.com/github/Lord-Kanzler/DS-Unit-2-Linear-Models/blob/master/module3-ridge-regression/LS_DS_213_assignment_ALEX_KAISER.ipynb\" target=\"_parent\"><img src=\"https://colab.research.google.com/assets/colab-badge.svg\" alt=\"Open In Colab\"/></a>",
"_____no_output_____"
],
[
"Lambda School Data Science\n\n*Unit 2, Sprint 1, Module 3*\n\n---",
"_____no_output_____"
],
[
"# Ridge Regression\n\n## Assignment\n\nWe're going back to our other **New York City** real estate dataset. Instead of predicting apartment rents, you'll predict property sales prices.\n\nBut not just for condos in Tribeca...\n\n- [ ] Use a subset of the data where `BUILDING_CLASS_CATEGORY` == `'01 ONE FAMILY DWELLINGS'` and the sale price was more than 100 thousand and less than 2 million.\n- [ ] Do train/test split. Use data from January — March 2019 to train. Use data from April 2019 to test.\n- [ ] Do one-hot encoding of categorical features.\n- [ ] Do feature selection with `SelectKBest`.\n- [ ] Fit a ridge regression model with multiple features. Use the `normalize=True` parameter (or do [feature scaling](https://scikit-learn.org/stable/modules/preprocessing.html) beforehand — use the scaler's `fit_transform` method with the train set, and the scaler's `transform` method with the test set)\n- [ ] Get mean absolute error for the test set.\n- [ ] As always, commit your notebook to your fork of the GitHub repo.\n\nThe [NYC Department of Finance](https://www1.nyc.gov/site/finance/taxes/property-rolling-sales-data.page) has a glossary of property sales terms and NYC Building Class Code Descriptions. The data comes from the [NYC OpenData](https://data.cityofnewyork.us/browse?q=NYC%20calendar%20sales) portal.\n\n\n## Stretch Goals\n\nDon't worry, you aren't expected to do all these stretch goals! These are just ideas to consider and choose from.\n\n- [ ] Add your own stretch goal(s) !\n- [ ] Instead of `Ridge`, try `LinearRegression`. Depending on how many features you select, your errors will probably blow up! 💥\n- [ ] Instead of `Ridge`, try [`RidgeCV`](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.RidgeCV.html).\n- [ ] Learn more about feature selection:\n - [\"Permutation importance\"](https://www.kaggle.com/dansbecker/permutation-importance)\n - [scikit-learn's User Guide for Feature Selection](https://scikit-learn.org/stable/modules/feature_selection.html)\n - [mlxtend](http://rasbt.github.io/mlxtend/) library\n - scikit-learn-contrib libraries: [boruta_py](https://github.com/scikit-learn-contrib/boruta_py) & [stability-selection](https://github.com/scikit-learn-contrib/stability-selection)\n - [_Feature Engineering and Selection_](http://www.feat.engineering/) by Kuhn & Johnson.\n- [ ] Try [statsmodels](https://www.statsmodels.org/stable/index.html) if you’re interested in more inferential statistical approach to linear regression and feature selection, looking at p values and 95% confidence intervals for the coefficients.\n- [ ] Read [_An Introduction to Statistical Learning_](http://faculty.marshall.usc.edu/gareth-james/ISL/ISLR%20Seventh%20Printing.pdf), Chapters 1-3, for more math & theory, but in an accessible, readable way.\n- [ ] Try [scikit-learn pipelines](https://scikit-learn.org/stable/modules/compose.html).",
"_____no_output_____"
]
],
[
[
"%%capture\nimport sys\n\n# If you're on Colab:\nif 'google.colab' in sys.modules:\n DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'\n !pip install category_encoders==2.*\n\n# If you're working locally:\nelse:\n DATA_PATH = '../data/'\n \n# Ignore this Numpy warning when using Plotly Express:\n# FutureWarning: Method .ptp is deprecated and will be removed in a future version. Use numpy.ptp instead.\nimport warnings\nwarnings.filterwarnings(action='ignore', category=FutureWarning, module='numpy')",
"_____no_output_____"
],
[
"import pandas as pd\nimport pandas_profiling\n\n# Read New York City property sales data\ndf = pd.read_csv(DATA_PATH+'condos/NYC_Citywide_Rolling_Calendar_Sales.csv')\n\n# Change column names: replace spaces with underscores\ndf.columns = [col.replace(' ', '_') for col in df]\n\n# SALE_PRICE was read as strings.\n# Remove symbols, convert to integer\ndf['SALE_PRICE'] = (\n df['SALE_PRICE']\n .str.replace('$','')\n .str.replace('-','')\n .str.replace(',','')\n .astype(int)\n)",
"_____no_output_____"
],
[
"# BOROUGH is a numeric column, but arguably should be a categorical feature,\n# so convert it from a number to a string\ndf['BOROUGH'] = df['BOROUGH'].astype(str)",
"_____no_output_____"
],
[
"# Reduce cardinality for NEIGHBORHOOD feature\n\n# Get a list of the top 10 neighborhoods\ntop10 = df['NEIGHBORHOOD'].value_counts()[:10].index\n\n# At locations where the neighborhood is NOT in the top 10, \n# replace the neighborhood with 'OTHER'\ndf.loc[~df['NEIGHBORHOOD'].isin(top10), 'NEIGHBORHOOD'] = 'OTHER'",
"_____no_output_____"
],
[
"import numpy as np\n\nprint(df.shape)\ndf.head()",
"(23040, 21)\n"
],
[
"df.describe(include='all')",
"_____no_output_____"
],
[
"df.dtypes",
"_____no_output_____"
],
[
"df['SALE_PRICE'].describe()",
"_____no_output_____"
],
[
"df['SALE_PRICE'][23035]",
"_____no_output_____"
],
[
"# subsetting\ndf = df.copy()\n\ndf_subset = df[(df['BUILDING_CLASS_CATEGORY'] == '01 ONE FAMILY DWELLINGS') & (df['SALE_PRICE'] >= 100000) & (df['SALE_PRICE'] < 20000000)]\n#everything_after_nov1st_2018 = df[(df['DATE']>=datetime.datetime(2018,11,1))]\n\ndf_subset.head()",
"_____no_output_____"
],
[
"df_subset['SALE_PRICE'].describe()",
"_____no_output_____"
],
[
"df_subset['SALE_PRICE'].shape",
"_____no_output_____"
],
[
"# hot encoding prep\ndf_subset.dtypes",
"_____no_output_____"
],
[
"df_subset.head()",
"_____no_output_____"
],
[
"# SALE_DATE to datetime\ndf_subset['SALE_DATE'] = pd.to_datetime(df_subset['SALE_DATE'])\ndf_subset.dtypes",
"/usr/local/lib/python3.6/dist-packages/ipykernel_launcher.py:1: SettingWithCopyWarning: \nA value is trying to be set on a copy of a slice from a DataFrame.\nTry using .loc[row_indexer,col_indexer] = value instead\n\nSee the caveats in the documentation: http://pandas.pydata.org/pandas-docs/stable/user_guide/indexing.html#returning-a-view-versus-a-copy\n \"\"\"Entry point for launching an IPython kernel.\n"
],
[
"# dropping columns\ndf_subset.drop(['EASE-MENT','ADDRESS', 'APARTMENT_NUMBER','BUILDING_CLASS_AT_TIME_OF_SALE'], axis= 1, inplace=True)",
"/usr/local/lib/python3.6/dist-packages/pandas/core/frame.py:4117: SettingWithCopyWarning: \nA value is trying to be set on a copy of a slice from a DataFrame\n\nSee the caveats in the documentation: http://pandas.pydata.org/pandas-docs/stable/user_guide/indexing.html#returning-a-view-versus-a-copy\n errors=errors,\n"
],
[
"# LAND_SQUARE_FEET str to float\ndf_subset['LAND_SQUARE_FEET'] = df_subset['LAND_SQUARE_FEET'].str.replace(',', '').astype(int)\ndf_subset['LAND_SQUARE_FEET'].head()",
"/usr/local/lib/python3.6/dist-packages/ipykernel_launcher.py:1: SettingWithCopyWarning: \nA value is trying to be set on a copy of a slice from a DataFrame.\nTry using .loc[row_indexer,col_indexer] = value instead\n\nSee the caveats in the documentation: http://pandas.pydata.org/pandas-docs/stable/user_guide/indexing.html#returning-a-view-versus-a-copy\n \"\"\"Entry point for launching an IPython kernel.\n"
],
[
"print(df_subset.shape)\ndf_subset.dtypes",
"(3240, 17)\n"
],
[
"# train/test split\nimport datetime as dt\ntrain = df_subset[(df_subset['SALE_DATE'] < dt.datetime(2019,4,1))]\ntest = df_subset[(df_subset['SALE_DATE'] >= dt.datetime(2019,4,1))]",
"_____no_output_____"
],
[
"# feature selection\ntarget = 'SALE_PRICE'\nfeatures = train.columns.drop([target]) \n\nX_train = train[features]\ny_train = train[target]\nX_test = test[features]\ny_test = test[target]",
"_____no_output_____"
],
[
"# hot encoding\nimport category_encoders as ce\nencoder = ce.OneHotEncoder(use_cat_names=True)\nX_train = encoder.fit_transform(X_train)\nX_train.head()",
"_____no_output_____"
],
[
"# transform\nX_test = encoder.transform(X_test)",
"_____no_output_____"
],
[
"X_test.head()",
"_____no_output_____"
],
[
"X_train.drop(['SALE_DATE'], axis= 1, inplace=True)\nX_test.drop(['SALE_DATE'], axis= 1, inplace=True)",
"_____no_output_____"
],
[
"df_subset.shape, X_train.shape, X_test.shape",
"_____no_output_____"
],
[
"# feature selection with SelectKBest\nimport warnings\nwarnings.filterwarnings(\"ignore\", category=RuntimeWarning) \n\nfrom sklearn.feature_selection import SelectKBest, f_regression\n\nselector = SelectKBest(score_func=f_regression, k=35)\nX_train_selected = selector.fit_transform(X_train, y_train)\nX_train_selected.shape",
"_____no_output_____"
],
[
"# Which features in selection\nselected_mask = selector.get_support()\nall_names = X_train.columns\nselected_names = all_names[selected_mask]\nunselected_names = all_names[~selected_mask]\n\nprint('Features selected:')\nfor name in selected_names:\n print(name)\n\nprint('\\n')\nprint('Features not selected:')\nfor name in unselected_names:\n print(name)",
"Features selected:\nBOROUGH_3\nBOROUGH_4\nBOROUGH_2\nBOROUGH_5\nBOROUGH_1\nNEIGHBORHOOD_OTHER\nNEIGHBORHOOD_FLUSHING-NORTH\nNEIGHBORHOOD_EAST NEW YORK\nNEIGHBORHOOD_FOREST HILLS\nNEIGHBORHOOD_BOROUGH PARK\nNEIGHBORHOOD_UPPER EAST SIDE (59-79)\nNEIGHBORHOOD_ASTORIA\nTAX_CLASS_AT_PRESENT_1\nTAX_CLASS_AT_PRESENT_1D\nBLOCK\nLOT\nBUILDING_CLASS_AT_PRESENT_A9\nBUILDING_CLASS_AT_PRESENT_A1\nBUILDING_CLASS_AT_PRESENT_A5\nBUILDING_CLASS_AT_PRESENT_A2\nBUILDING_CLASS_AT_PRESENT_A3\nBUILDING_CLASS_AT_PRESENT_A7\nBUILDING_CLASS_AT_PRESENT_S1\nBUILDING_CLASS_AT_PRESENT_A4\nBUILDING_CLASS_AT_PRESENT_A6\nBUILDING_CLASS_AT_PRESENT_A8\nBUILDING_CLASS_AT_PRESENT_S0\nBUILDING_CLASS_AT_PRESENT_B3\nZIP_CODE\nRESIDENTIAL_UNITS\nCOMMERCIAL_UNITS\nTOTAL_UNITS\nLAND_SQUARE_FEET\nGROSS_SQUARE_FEET\nYEAR_BUILT\n\n\nFeatures not selected:\nNEIGHBORHOOD_BEDFORD STUYVESANT\nBUILDING_CLASS_CATEGORY_01 ONE FAMILY DWELLINGS\nBUILDING_CLASS_AT_PRESENT_A0\nBUILDING_CLASS_AT_PRESENT_B2\nTAX_CLASS_AT_TIME_OF_SALE\n"
],
[
"# How many features should be selected\nfrom sklearn.linear_model import LinearRegression\nfrom sklearn.metrics import mean_absolute_error\n\nfor k in range(1, len(X_train.columns)+1):\n print(f'{k} features')\n \n selector = SelectKBest(score_func=f_regression, k=k)\n X_train_selected = selector.fit_transform(X_train, y_train)\n X_test_selected = selector.transform(X_test)\n\n model = LinearRegression()\n model.fit(X_train_selected, y_train)\n y_pred = model.predict(X_test_selected)\n mae = mean_absolute_error(y_test, y_pred)\n print(f'Test Mean Absolute Error: ${mae:,.0f} \\n')",
"1 features\nTest Mean Absolute Error: $276,211 \n\n2 features\nTest Mean Absolute Error: $282,820 \n\n3 features\nTest Mean Absolute Error: $246,634 \n\n4 features\nTest Mean Absolute Error: $244,564 \n\n5 features\nTest Mean Absolute Error: $240,443 \n\n6 features\nTest Mean Absolute Error: $237,674 \n\n7 features\nTest Mean Absolute Error: $234,000 \n\n8 features\nTest Mean Absolute Error: $234,604 \n\n9 features\nTest Mean Absolute Error: $235,697 \n\n10 features\nTest Mean Absolute Error: $235,594 \n\n11 features\nTest Mean Absolute Error: $232,381 \n\n12 features\nTest Mean Absolute Error: $230,926 \n\n13 features\nTest Mean Absolute Error: $233,176 \n\n14 features\nTest Mean Absolute Error: $233,595 \n\n15 features\nTest Mean Absolute Error: $234,772 \n\n16 features\nTest Mean Absolute Error: $230,639 \n\n17 features\nTest Mean Absolute Error: $230,609 \n\n18 features\nTest Mean Absolute Error: $231,231 \n\n19 features\nTest Mean Absolute Error: $223,650 \n\n20 features\nTest Mean Absolute Error: $224,226 \n\n21 features\nTest Mean Absolute Error: $224,226 \n\n22 features\nTest Mean Absolute Error: $224,340 \n\n23 features\nTest Mean Absolute Error: $224,423 \n\n24 features\nTest Mean Absolute Error: $224,137 \n\n25 features\nTest Mean Absolute Error: $224,012 \n\n26 features\nTest Mean Absolute Error: $223,943 \n\n27 features\nTest Mean Absolute Error: $223,897 \n\n28 features\nTest Mean Absolute Error: $220,383 \n\n29 features\nTest Mean Absolute Error: $221,292 \n\n30 features\nTest Mean Absolute Error: $221,996 \n\n31 features\nTest Mean Absolute Error: $222,136 \n\n32 features\nTest Mean Absolute Error: $219,495 \n\n33 features\nTest Mean Absolute Error: $219,535 \n\n34 features\nTest Mean Absolute Error: $219,732 \n\n35 features\nTest Mean Absolute Error: $218,793 \n\n36 features\nTest Mean Absolute Error: $219,606 \n\n37 features\nTest Mean Absolute Error: $218,749 \n\n38 features\nTest Mean Absolute Error: $218,863 \n\n39 features\nTest Mean Absolute Error: $218,863 \n\n40 features\nTest Mean Absolute Error: $218,783 \n\n"
]
],
[
[
"35 features\nTest Mean Absolute Error: $218,793 ",
"_____no_output_____"
]
],
[
[
"# imports\n%matplotlib inline\n\nimport matplotlib.pyplot as plt\nimport seaborn as sns\nfrom sklearn.linear_model import Ridge\nfrom IPython.display import display, HTML\nfrom sklearn.linear_model import RidgeCV\n",
"_____no_output_____"
],
[
"# ridge regression for multiple features\nfor alpha in [0.001, 0.01, 0.1, 1.0, 1, 100.0, 1000.0]:\n \n # Fit Ridge Regression model\n display(HTML(f'Ridge Regression, with alpha={alpha}'))\n model = Ridge(alpha=alpha, normalize=True)\n model.fit(X_train, y_train)\n y_pred = model.predict(X_test)\n\n # Get Test MAE\n mae = mean_absolute_error(y_test, y_pred)\n display(HTML(f'Test Mean Absolute Error: ${mae:,.0f}'))\n \n # Plot coefficients\n coefficients = pd.Series(model.coef_, X_train.columns)\n plt.figure(figsize=(16,8))\n coefficients.sort_values().plot.barh(color='grey')\n plt.xlim(-700000,7000000)\n plt.show()",
"_____no_output_____"
],
[
"coefficients = pd.Series(model.coef_, X_train.columns)\ncoefficients",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
]
] |
4acf2397dd6bf9d5074ff17fd181be275022acfe
| 9,530 |
ipynb
|
Jupyter Notebook
|
content/Chapter_19/02_Moment_Generating_Functions.ipynb
|
dcroce/jupyter-book
|
9ac4b502af8e8c5c3b96f5ec138602a0d3d8a624
|
[
"MIT"
] | null | null | null |
content/Chapter_19/02_Moment_Generating_Functions.ipynb
|
dcroce/jupyter-book
|
9ac4b502af8e8c5c3b96f5ec138602a0d3d8a624
|
[
"MIT"
] | null | null | null |
content/Chapter_19/02_Moment_Generating_Functions.ipynb
|
dcroce/jupyter-book
|
9ac4b502af8e8c5c3b96f5ec138602a0d3d8a624
|
[
"MIT"
] | null | null | null | 43.318182 | 400 | 0.57681 |
[
[
[
"# HIDDEN\nfrom datascience import *\nfrom prob140 import *\nimport numpy as np\nimport matplotlib.pyplot as plt\nplt.style.use('fivethirtyeight')\n%matplotlib inline\nimport math\nfrom scipy import stats",
"_____no_output_____"
]
],
[
[
"## Moment Generating Functions ##",
"_____no_output_____"
],
[
"The probability mass function and probability density, cdf, and survival functions are all ways of specifying the probability distribution of a random variable. They are all defined as probabilities or as probability per unit length, and thus have natural interpretations and visualizations.\n\nBut there are also more abstract ways of describing distributions. One that you have encountered is the probability generating function (pgf), which we defined for random variables with finitely many non-negative integer values.\n\nWe now define another such *transform* of a distribution. More general than the pgf, it is a powerful tool for studying distributions.\n\nLet $X$ be a random variable. The *moment generating function* (mgf) of $X$ is a function defined on the real numbers by the formula\n\n$$\nM_X(t) ~ = ~ E(e^{tX}) \n$$\n\nfor all $t$ for which the expectation is finite. It is a fact (which we will not prove) that the domain of the mgf has to be an interval, not necessarily finite but necessarily including 0 because $M_X(0) = 1$.\n\nFor $X$ with finitely many non-negative integer values, we had defined the pgf by $G_X(s) = E(s^X)$. Notice that this is a special case of the mgf with $s = e^t$ and hence positive. For a random variable $X$ that has both a pgf $G_X$ and an mgf $M_X$, the two functions are related by $M_X(\\log(s)) = G_X(s)$. Therefore the properties of $M_X$ near 0 reflect the properties of $G_X$ near 1.\n\nThis section presents three ways in which the mgf is useful. Other ways are demonstrated in the subsequent sections of this chapter. Much of what we say about mgf's will not be accompanied by complete proofs as the math required is beyond the scope of this class. But the results should seem reasonable, even without formal proofs.\n\nWe will list the three ways first, and then use them all in examples.",
"_____no_output_____"
],
[
"### Generating Moments ###\nFor non-negative integers $k$, the expectation $E(X^k)$ is called *$k$th moment* of $X$. You saw in Data 8 and again in this course that the mean $E(X)$ is the center of gravity of the probability histogram of $X$. In physics, the center of mass is called the *first moment*. The terminology of moments is used in probability theory as well.\n\nIn this course we are only going to work with mgf's that are finite in some interval around 0. The interval could be the entire real line. It is a fact that if the mgf is finite around 0 (not just to one side of 0), then all the moments exist.\n\nExpand $e^{tX}$ to see that\n\n$$\n\\begin{align*}\nM_X(t) ~ &= ~ E \\big{(} 1 + t \\frac{X}{1!} + t^2 \\frac{X^2}{2!} + t^3 \\frac{X^3}{3!} + \\cdots \\big{)} \\\\ \\\\\n&= ~ 1 + t \\frac{E(X)}{1!} + t^2 \\frac{E(X^2)}{2!} + t^3 \\frac{E(X^3)}{3!} + \\cdots\n\\end{align*}\n$$\n\nby blithely switching the expectation and the infinite sum. This requires justification, which we won't go into.\n\nContinue to set aside questions about whether we can switch infinite sums with other operations. Just go ahead and differentiate $M_X$ term by term. Let $M_X^{(n)}$ denote the $n$th derivative. Then\n\n$$\nM_X^{(1)} (t) ~ = ~ \\frac{d}{dt} M_X(t) ~ = \\frac{E(X)}{1!} + 2t \\frac{E(X^2)}{2!} + 3t^2 \\frac{E(X^3)}{3!} + \\cdots\n$$\n\nand hence\n$$\nM^{(1)} (0) ~ = ~ E(X)\n$$\n\nNow differentiate $M_X^{(1)}$ to see that $M_X^{(2)}(0) = E(X^2)$, and, by induction,\n\n$$\nM^{(n)} (0) ~ = ~ E(X^n), ~~~~ n = 1, 2, 3, \\ldots\n$$\n\nHence we can *generate the moments of $X$* by evaluating successive derivatives of $M_X$ at $t=0$. This is one way in which mgf's are helpful.",
"_____no_output_____"
],
[
"### Identifying the Distribution ###\nIn this class we have made heavy use of the first and second moments, and no use at all of the higher moments. That will continue to be the case. But mgf's do involve all the moments, and this results in a property that is very useful for proving facts about distributions. This property is valid if the mgf exists in an interval around 0, which we assumed earlier in this section.\n\n**If two distributions have the same mgf, then they must be the same distribution.** For example, if you recognize the mgf of a random variable as the mgf of a normal distribution, then the random variable must be normal.\n\nBy contrast, if you know the expectation of a random variable you can't identify the distribution of the random variable; even if you know both the mean and the SD (equivalently, the first and second moments), you can't identify the distribution. But if you know the moment generating function, and hence all the moments, then you can.",
"_____no_output_____"
],
[
"### Working Well with Sums ###\nThe third reason mgf's are useful is that like the pgf, the mgf of the sum of independent random variables is easily computed as a product.\n\nLet $X$ and $Y$ be independent. Then\n$$\nM_{X+Y} (t) ~ = ~ E(e^{t(X+Y)}) ~ = ~ E(e^{tX} \\cdot e^{tY})\n$$\n\nSo if $X$ and $Y$ are independent,\n$$\nM_{X+Y}(t) ~ = ~ M_X(t) M_Y(t)\n$$",
"_____no_output_____"
],
[
"It's time for some examples. Remember that the mgf of $X$ is the expectation of a function of $X$. In some cases we will calculate it using the non-linear function rule for expectations. In other cases we will use the multiplicative property of the mgf of the sum of independent random variables.",
"_____no_output_____"
],
[
"### MGFs of Some Discrete Random Variables ###\n\n#### Bernoulli $(p)$ ####\n$P(X = 1) = p$ and $P(X = 0) = 1 - p = q$. So\n\n$$\nM_X(t) ~ = ~ qe^{t \\cdot 0} + pe^{t \\cdot 1} ~ = ~ q + pe^t ~ = ~ 1 - p(e^t - 1) ~~~ \\text{for all } t \n$$\n\n#### Binomial $(n, p)$ ####\nA binomial random variable is the sum of $n$ i.i.d. indicators. So\n\n$$\nM_X(t) ~ = ~ (q + pe^t)^n ~~~ \\text{for all } t \n$$\n\n#### Poisson $(\\mu)$ ####\nThis one is an exercise.\n$$\nM_X(t) ~ = ~ e^{\\mu(e^t - 1)} ~~~ \\text{for all } t\n$$\n\nYou can also use this to show that the sum of independent Poisson variables is Poisson.",
"_____no_output_____"
],
[
"### MGF of a Gamma $(r, \\lambda )$ Random Variable ###\n\nLet $X$ have the gamma $(r, \\lambda)$ distribution. Then\n$$\n\\begin{align*}\nM_X(t) ~ &= ~ \\int_0^\\infty e^{tx} \\frac{\\lambda^r}{\\Gamma(r)} x^{r-1} e^{-\\lambda x} dx \\\\ \\\\\n&= ~ \\frac{\\lambda^r}{\\Gamma(r)} \\int_0^\\infty x^{r-1} e^{-(\\lambda - t)x} dx \\\\ \\\\\n&= ~ \\frac{\\lambda^r}{\\Gamma(r)} \\cdot \\frac{\\Gamma(r)}{(\\lambda - t)^r} ~~~~ t < \\lambda \\\\ \\\\\n&= \\big{(} \\frac{\\lambda}{\\lambda - t} \\big{)}^r ~~~~ t < \\lambda\n\\end{align*} \n$$\n\n#### Sums of Independent Gamma Variables with the Same Rate ####\nIf $X$ has gamma $(r, \\lambda)$ distribution and $Y$ independent of $X$ has gamma $(s, \\lambda)$ distribution, then\n$$\n\\begin{align*} \nM_{X+Y} (t) ~ &= ~ \\big{(} \\frac{\\lambda}{\\lambda - t} \\big{)}^r \\cdot \\big{(} \\frac{\\lambda}{\\lambda - t} \\big{)}^s ~~~~ t < \\lambda \\\\ \\\\\n&= ~ \\big{(} \\frac{\\lambda}{\\lambda - t} \\big{)}^{r+s} ~~~~ t < \\lambda\n\\end{align*}\n$$\n\nThat's the mgf of the gamma $(r+s, \\lambda)$ distribution. Because the mgf identifies the distribution, $X+Y$ must have the gamma $(r+s, \\lambda)$ distribution.",
"_____no_output_____"
]
]
] |
[
"code",
"markdown"
] |
[
[
"code"
],
[
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown",
"markdown"
]
] |
4acf36155dcc4291bf7db01e0e135f7374d1db51
| 265,603 |
ipynb
|
Jupyter Notebook
|
Differentiation.ipynb
|
DRWardrope/GeometryTests
|
84cd6ab8e5dbe4caf1854acc33c8638c349bfffc
|
[
"MIT"
] | null | null | null |
Differentiation.ipynb
|
DRWardrope/GeometryTests
|
84cd6ab8e5dbe4caf1854acc33c8638c349bfffc
|
[
"MIT"
] | null | null | null |
Differentiation.ipynb
|
DRWardrope/GeometryTests
|
84cd6ab8e5dbe4caf1854acc33c8638c349bfffc
|
[
"MIT"
] | null | null | null | 280.467793 | 52,972 | 0.892799 |
[
[
[
"# Test differentation\n\nTest differentiation of distance functions, by implementing gradient descent.",
"_____no_output_____"
]
],
[
[
"import numpy as np\nimport matplotlib.pyplot as plt\nfrom matplotlib import collections, lines, markers, path, patches\n%matplotlib inline",
"_____no_output_____"
],
[
"from geometry import *",
"_____no_output_____"
]
],
[
[
"## Set up 1-D hyperboloid manifold",
"_____no_output_____"
]
],
[
[
"theta = np.linspace(-0.323*np.pi, 0.323*np.pi, 1000)\nx0 = np.sinh(theta)\nx1 = np.cosh(theta)\nx = np.array([np.array([x0i, x1i]) for x0i, x1i in zip(x0, x1)])",
"_____no_output_____"
],
[
"target = np.array([0., 1.])\ninitial_point = np.array([np.sinh(0.9), np.cosh(0.9)])",
"_____no_output_____"
],
[
"def gradient_descent(pt_i, target, differential_fn, geometry=\"hyperbolic\", learning_rate=1.):\n '''\n Calculate local gradient of differential, given the current pt and the target.\n Inputs:\n Two (d+1)-dimensional vectors in ambient space co-ordinates, pt_i and target\n pt_i: (d+1)-dimensional vector in ambient space co-ordinates,\n the point to evaluate the gradient at.\n target: (d+1)-dimensional vectors in ambient space co-ordinates, the target point\n differential_fn: function that calculates the derivative\n learning_rate: dictates how far to step in gradient direction\n '''\n # Calculate gradient in ambient space co-ordinates\n step = differential_fn(pt_i, target, geometry)\n print(\"step =\",step)\n # Project this gradient onto tangent space\n projection = project_to_tangent(pt_i, step, geometry)\n print(\"projection on tangent space = \",projection)\n # Map to manifold and return this updated pt\n return exponential_map(-learning_rate*projection, pt_i, geometry)\n# return exponential_map(-projection, pt_i, geometry)",
"_____no_output_____"
],
[
"def error_differential_eucl(u, v, geometry=\"hyperbolic\"):\n '''\n Calculate differential of distance between points u and v, **with respect to u**,\n accounting for different geometries by implementing an appropriate metric.\n Inputs:\n u: (d+1)-dimensional vector, expressed in ambient space coordinates\n v: (d+1)-dimensional vector, expressed in ambient space coordinates\n geometry: specifies which metric to use (and hence how inner product calculated)\n Outputs:\n gradient of the distance in (d+1)-dimensional ambient space coordinates\n ''' \n if np.array_equal(u,v):\n return np.zeros(u.shape)\n # If u and v are different, calculate the gradient\n metric = get_metric(u.shape[0], geometry)\n print(\"u = {}, v = {}, u.v = {}\".format(u, v, dot(u, v, geometry)))\n if geometry == \"spherical\":\n coeff = -1./np.sqrt(1.-dot(u, v, geometry)**2)\n if geometry == \"hyperbolic\": \n coeff = -1./np.sqrt(dot(u, v, geometry)**2-1.) \n #return coeff*metric.dot(v)\n return coeff*v",
"_____no_output_____"
],
[
"updated_pts = [initial_point]\nfor i in range(1,10):\n updated_pts.append(\n gradient_descent( \n updated_pts[i-1], \n target,\n error_differential_eucl, \n geometry=\"hyperbolic\", \n learning_rate=0.1\n )\n )\n print(\"updated_pt = \", updated_pts[i])\n print(\"-\"*80)\nprint(updated_pts)\n",
"u = [1.02651673 1.43308639], v = [0. 1.], u.v = -1.4330863854487743\nstep = [-0. -0.97416825]\nproject_to_tangent: point_on_manifold = [1.02651673 1.43308639], displacement = [-0. -0.97416825], geometry = hyperbolic\nproject_to_tangent: xp_norm = 1.3960672530300118, xx_norm = -1.0\nprojection on tangent space = [1.43308639 1.02651673]\nexponential_map: norm_v_tan = 0.09999999999999999\nupdated_pt = [0.88810598 1.33743495]\n--------------------------------------------------------------------------------\nu = [0.88810598 1.33743495], v = [0. 1.], u.v = -1.3374349463048445\nstep = [-0. -1.12599174]\nproject_to_tangent: point_on_manifold = [0.88810598 1.33743495], displacement = [-0. -1.12599174], geometry = hyperbolic\nproject_to_tangent: xp_norm = 1.5059407020437068, xx_norm = -0.9999999999999996\nprojection on tangent space = [1.33743495 0.88810598]\nexponential_map: norm_v_tan = 0.10000000000000006\nupdated_pt = [0.7585837 1.25516901]\n--------------------------------------------------------------------------------\nu = [0.7585837 1.25516901], v = [0. 1.], u.v = -1.2551690056309428\nstep = [-0. -1.31824609]\nproject_to_tangent: point_on_manifold = [0.7585837 1.25516901], displacement = [-0. -1.31824609], geometry = hyperbolic\nproject_to_tangent: xp_norm = 1.6546216358026298, xx_norm = -0.9999999999999996\nprojection on tangent space = [1.25516901 0.7585837 ]\nexponential_map: norm_v_tan = 0.10000000000000003\nupdated_pt = [0.63665358 1.18546522]\n--------------------------------------------------------------------------------\nu = [0.63665358 1.18546522], v = [0. 1.], u.v = -1.1854652182422671\nstep = [-0. -1.57071291]\nproject_to_tangent: point_on_manifold = [0.63665358 1.18546522], displacement = [-0. -1.57071291], geometry = hyperbolic\nproject_to_tangent: xp_norm = 1.8620255213866685, xx_norm = -0.999999999999999\nprojection on tangent space = [1.18546522 0.63665358]\nexponential_map: norm_v_tan = 0.10000000000000019\nupdated_pt = [0.52109531 1.12762597]\n--------------------------------------------------------------------------------\nu = [0.52109531 1.12762597], v = [0. 1.], u.v = -1.1276259652063803\nstep = [-0. -1.91903475]\nproject_to_tangent: point_on_manifold = [0.52109531 1.12762597], displacement = [-0. -1.91903475], geometry = hyperbolic\nproject_to_tangent: xp_norm = 2.1639534137386565, xx_norm = -0.9999999999999993\nprojection on tangent space = [1.12762597 0.52109531]\nexponential_map: norm_v_tan = 0.10000000000000017\nupdated_pt = [0.41075233 1.08107237]\n--------------------------------------------------------------------------------\nu = [0.41075233 1.08107237], v = [0. 1.], u.v = -1.0810723718384543\nstep = [-0. -2.43455712]\nproject_to_tangent: point_on_manifold = [0.41075233 1.08107237], displacement = [-0. -2.43455712], geometry = hyperbolic\nproject_to_tangent: xp_norm = 2.6319324418321957, xx_norm = -0.9999999999999994\nprojection on tangent space = [1.08107237 0.41075233]\nexponential_map: norm_v_tan = 0.10000000000000021\nupdated_pt = [0.30452029 1.04533851]\n--------------------------------------------------------------------------------\nu = [0.30452029 1.04533851], v = [0. 1.], u.v = -1.04533851412886\nstep = [-0. -3.2838534]\nproject_to_tangent: point_on_manifold = [0.30452029 1.04533851], displacement = [-0. -3.2838534], geometry = hyperbolic\nproject_to_tangent: xp_norm = 3.432738430321758, xx_norm = -0.9999999999999994\nprojection on tangent space = [1.04533851 0.30452029]\nexponential_map: norm_v_tan = 0.10000000000000031\nupdated_pt = [0.201336 1.02006676]\n--------------------------------------------------------------------------------\nu = [0.201336 1.02006676], v = [0. 1.], u.v = -1.0200667556190754\nstep = [-0. -4.96682157]\nproject_to_tangent: point_on_manifold = [0.201336 1.02006676], displacement = [-0. -4.96682157], geometry = hyperbolic\nproject_to_tangent: xp_norm = 5.0664895634395215, xx_norm = -0.9999999999999996\nprojection on tangent space = [1.02006676 0.201336 ]\nexponential_map: norm_v_tan = 0.10000000000000053\nupdated_pt = [0.10016675 1.00500417]\n--------------------------------------------------------------------------------\nu = [0.10016675 1.00500417], v = [0. 1.], u.v = -1.0050041680558033\nstep = [-0. -9.98335276]\nproject_to_tangent: point_on_manifold = [0.10016675 1.00500417], displacement = [-0. -9.98335276], geometry = hyperbolic\nproject_to_tangent: xp_norm = 10.033311132254251, xx_norm = -0.9999999999999998\nprojection on tangent space = [1.00500417 0.10016675]\nexponential_map: norm_v_tan = 0.10000000000000116\nupdated_pt = [-2.65065747e-15 1.00000000e+00]\n--------------------------------------------------------------------------------\n[array([1.02651673, 1.43308639]), array([0.88810598, 1.33743495]), array([0.7585837 , 1.25516901]), array([0.63665358, 1.18546522]), array([0.52109531, 1.12762597]), array([0.41075233, 1.08107237]), array([0.30452029, 1.04533851]), array([0.201336 , 1.02006676]), array([0.10016675, 1.00500417]), array([-2.65065747e-15, 1.00000000e+00])]\n"
],
[
"fig = plt.figure(figsize=(12,12))\nax = plt.gca(xlim=[-1.2, 1.2], ylim=[0.9, 1.6])\nax.scatter(x0, x1, s=1)\nax.scatter(target[0], target[1], color='r', marker='*',s=200,label = \"Target\")\nfor i, pt in enumerate(updated_pts):\n ax.scatter(\n pt[0], pt[1], \n marker=markers.MarkerStyle('o',fillstyle=\"none\"), \n s=100, label=\"Update {}\".format(i)\n )\nax.legend()",
"_____no_output_____"
],
[
"updated_pts = [np.array([0., 1.])]\nfor i in range(1,10):\n updated_pts.append(\n gradient_descent( \n updated_pts[i-1], \n target,\n error_differential_eucl, \n geometry=\"hyperbolic\", \n learning_rate=0.1\n )\n )\n# print(\"updated_pt = \", updated_pts[i])\n# print(\"-\"*80)\nprint(updated_pts)\n",
"step = [0. 0.]\nproject_to_tangent: point_on_manifold = [0. 1.], displacement = [0. 0.], geometry = hyperbolic\nproject_to_tangent: xp_norm = 0.0, xx_norm = -1.0\nprojection on tangent space = [0. 0.]\nexponential_map: norm_v_tan = 0.0\nstep = [0. 0.]\nproject_to_tangent: point_on_manifold = [0. 1.], displacement = [0. 0.], geometry = hyperbolic\nproject_to_tangent: xp_norm = 0.0, xx_norm = -1.0\nprojection on tangent space = [0. 0.]\nexponential_map: norm_v_tan = 0.0\nstep = [0. 0.]\nproject_to_tangent: point_on_manifold = [0. 1.], displacement = [0. 0.], geometry = hyperbolic\nproject_to_tangent: xp_norm = 0.0, xx_norm = -1.0\nprojection on tangent space = [0. 0.]\nexponential_map: norm_v_tan = 0.0\nstep = [0. 0.]\nproject_to_tangent: point_on_manifold = [0. 1.], displacement = [0. 0.], geometry = hyperbolic\nproject_to_tangent: xp_norm = 0.0, xx_norm = -1.0\nprojection on tangent space = [0. 0.]\nexponential_map: norm_v_tan = 0.0\nstep = [0. 0.]\nproject_to_tangent: point_on_manifold = [0. 1.], displacement = [0. 0.], geometry = hyperbolic\nproject_to_tangent: xp_norm = 0.0, xx_norm = -1.0\nprojection on tangent space = [0. 0.]\nexponential_map: norm_v_tan = 0.0\nstep = [0. 0.]\nproject_to_tangent: point_on_manifold = [0. 1.], displacement = [0. 0.], geometry = hyperbolic\nproject_to_tangent: xp_norm = 0.0, xx_norm = -1.0\nprojection on tangent space = [0. 0.]\nexponential_map: norm_v_tan = 0.0\nstep = [0. 0.]\nproject_to_tangent: point_on_manifold = [0. 1.], displacement = [0. 0.], geometry = hyperbolic\nproject_to_tangent: xp_norm = 0.0, xx_norm = -1.0\nprojection on tangent space = [0. 0.]\nexponential_map: norm_v_tan = 0.0\nstep = [0. 0.]\nproject_to_tangent: point_on_manifold = [0. 1.], displacement = [0. 0.], geometry = hyperbolic\nproject_to_tangent: xp_norm = 0.0, xx_norm = -1.0\nprojection on tangent space = [0. 0.]\nexponential_map: norm_v_tan = 0.0\nstep = [0. 0.]\nproject_to_tangent: point_on_manifold = [0. 1.], displacement = [0. 0.], geometry = hyperbolic\nproject_to_tangent: xp_norm = 0.0, xx_norm = -1.0\nprojection on tangent space = [0. 0.]\nexponential_map: norm_v_tan = 0.0\n[array([0., 1.]), array([0., 1.]), array([0., 1.]), array([0., 1.]), array([0., 1.]), array([0., 1.]), array([0., 1.]), array([0., 1.]), array([0., 1.]), array([0., 1.])]\n"
],
[
"fig = plt.figure(figsize=(12,12))\nax = plt.gca(xlim=[-1.2, 1.2], ylim=[0.9, 1.6])\nax.scatter(x0, x1, s=1)\nax.scatter(target[0], target[1], color='r', marker='*',s=200,label = \"Target\")\nfor i, pt in enumerate(updated_pts):\n ax.scatter(\n pt[0], pt[1], \n marker=markers.MarkerStyle('o',fillstyle=\"none\"), \n s=100, label=\"Update {}\".format(i)\n )\nax.legend()",
"_____no_output_____"
],
[
"updated_pts = [initial_point]\nfor i in range(1,10):\n updated_pts.append(\n gradient_descent( \n updated_pts[i-1], \n target,\n error_differential_eucl, \n geometry=\"hyperbolic\", \n learning_rate=0.18\n )\n )\n print(\"****xhi =\", np.arcsinh(updated_pts[i][0]))\n\n\n print(\"updated_pt = \", updated_pts[i])\n print(\"-\"*80)\nprint(updated_pts)\n",
"u = [1.02651673 1.43308639], v = [0. 1.], u.v = -1.4330863854487743\nstep = [-0. -0.97416825]\nproject_to_tangent: point_on_manifold = [1.02651673 1.43308639], displacement = [-0. -0.97416825], geometry = hyperbolic\nproject_to_tangent: xp_norm = 1.3960672530300118, xx_norm = -1.0\nprojection on tangent space = [1.43308639 1.02651673]\nexponential_map: norm_v_tan = 0.17999999999999994\n****xhi = 0.7200000000000001\nupdated_pt = [0.78384048 1.27059273]\n--------------------------------------------------------------------------------\nu = [0.78384048 1.27059273], v = [0. 1.], u.v = -1.2705927333019296\nstep = [-0. -1.27576979]\nproject_to_tangent: point_on_manifold = [0.78384048 1.27059273], displacement = [-0. -1.27576979], geometry = hyperbolic\nproject_to_tangent: xp_norm = 1.6209838226402558, xx_norm = -0.9999999999999997\nprojection on tangent space = [1.27059273 0.78384048]\nexponential_map: norm_v_tan = 0.18000000000000008\n****xhi = 0.5399999999999999\nupdated_pt = [0.5666293 1.14937756]\n--------------------------------------------------------------------------------\nu = [0.5666293 1.14937756], v = [0. 1.], u.v = -1.149377557279424\nstep = [-0. -1.76482224]\nproject_to_tangent: point_on_manifold = [0.5666293 1.14937756], displacement = [-0. -1.76482224], geometry = hyperbolic\nproject_to_tangent: xp_norm = 2.0284470770025655, xx_norm = -1.0\nprojection on tangent space = [1.14937756 0.5666293 ]\nexponential_map: norm_v_tan = 0.17999999999999994\n****xhi = 0.35999999999999993\nupdated_pt = [0.36782654 1.06550287]\n--------------------------------------------------------------------------------\nu = [0.36782654 1.06550287], v = [0. 1.], u.v = -1.0655028703156857\nstep = [-0. -2.71867274]\nproject_to_tangent: point_on_manifold = [0.36782654 1.06550287], displacement = [-0. -2.71867274], geometry = hyperbolic\nproject_to_tangent: xp_norm = 2.8967536111449883, xx_norm = -1.0000000000000002\nprojection on tangent space = [1.06550287 0.36782654]\nexponential_map: norm_v_tan = 0.17999999999999988\n****xhi = 0.18000000000000008\nupdated_pt = [0.18097358 1.01624379]\n--------------------------------------------------------------------------------\nu = [0.18097358 1.01624379], v = [0. 1.], u.v = -1.0162437872665413\nstep = [-0. -5.52566857]\nproject_to_tangent: point_on_manifold = [0.18097358 1.01624379], displacement = [-0. -5.52566857], geometry = hyperbolic\nproject_to_tangent: xp_norm = 5.615426354172611, xx_norm = -1.0000000000000004\nprojection on tangent space = [1.01624379 0.18097358]\nexponential_map: norm_v_tan = 0.17999999999999874\n****xhi = 1.3600232051658168e-15\nupdated_pt = [1.36002321e-15 1.00000000e+00]\n--------------------------------------------------------------------------------\nu = [1.36002321e-15 1.00000000e+00], v = [0. 1.], u.v = -1.0000000000000004\nstep = [ -0. -33554432.]\nproject_to_tangent: point_on_manifold = [1.36002321e-15 1.00000000e+00], displacement = [ -0. -33554432.], geometry = hyperbolic\nproject_to_tangent: xp_norm = 33554432.000000015, xx_norm = -1.0000000000000009\nprojection on tangent space = [4.56348062e-08 0.00000000e+00]\nexponential_map: norm_v_tan = 8.214265108108516e-09\n****xhi = 1.3600232051658168e-15\nupdated_pt = [1.36002321e-15 1.00000000e+00]\n--------------------------------------------------------------------------------\nu = [1.36002321e-15 1.00000000e+00], v = [0. 1.], u.v = -1.0000000000000004\nstep = [ -0. -33554432.]\nproject_to_tangent: point_on_manifold = [1.36002321e-15 1.00000000e+00], displacement = [ -0. -33554432.], geometry = hyperbolic\nproject_to_tangent: xp_norm = 33554432.000000015, xx_norm = -1.0000000000000009\nprojection on tangent space = [4.56348062e-08 0.00000000e+00]\nexponential_map: norm_v_tan = 8.214265108108516e-09\n****xhi = 1.3600232051658168e-15\nupdated_pt = [1.36002321e-15 1.00000000e+00]\n--------------------------------------------------------------------------------\nu = [1.36002321e-15 1.00000000e+00], v = [0. 1.], u.v = -1.0000000000000004\nstep = [ -0. -33554432.]\nproject_to_tangent: point_on_manifold = [1.36002321e-15 1.00000000e+00], displacement = [ -0. -33554432.], geometry = hyperbolic\nproject_to_tangent: xp_norm = 33554432.000000015, xx_norm = -1.0000000000000009\nprojection on tangent space = [4.56348062e-08 0.00000000e+00]\nexponential_map: norm_v_tan = 8.214265108108516e-09\n****xhi = 1.3600232051658168e-15\nupdated_pt = [1.36002321e-15 1.00000000e+00]\n--------------------------------------------------------------------------------\nu = [1.36002321e-15 1.00000000e+00], v = [0. 1.], u.v = -1.0000000000000004\nstep = [ -0. -33554432.]\nproject_to_tangent: point_on_manifold = [1.36002321e-15 1.00000000e+00], displacement = [ -0. -33554432.], geometry = hyperbolic\nproject_to_tangent: xp_norm = 33554432.000000015, xx_norm = -1.0000000000000009\nprojection on tangent space = [4.56348062e-08 0.00000000e+00]\nexponential_map: norm_v_tan = 8.214265108108516e-09\n****xhi = 1.3600232051658168e-15\nupdated_pt = [1.36002321e-15 1.00000000e+00]\n--------------------------------------------------------------------------------\n[array([1.02651673, 1.43308639]), array([0.78384048, 1.27059273]), array([0.5666293 , 1.14937756]), array([0.36782654, 1.06550287]), array([0.18097358, 1.01624379]), array([1.36002321e-15, 1.00000000e+00]), array([1.36002321e-15, 1.00000000e+00]), array([1.36002321e-15, 1.00000000e+00]), array([1.36002321e-15, 1.00000000e+00]), array([1.36002321e-15, 1.00000000e+00])]\n"
],
[
"updated_pts = [initial_point]\nfor i in range(1,10):\n updated_pts.append(\n gradient_descent( \n updated_pts[i-1], \n target,\n error_differential_eucl, \n geometry=\"hyperbolic\", \n learning_rate=0.17\n )\n )\n print(\"****xhi =\", np.arcsinh(updated_pts[i][0]))\n\n\n print(\"updated_pt = \", updated_pts[i])\n print(\"-\"*80)\nprint(updated_pts)\n",
"u = [1.02651673 1.43308639], v = [0. 1.], u.v = -1.4330863854487743\nstep = [-0. -0.97416825]\nproject_to_tangent: point_on_manifold = [1.02651673 1.43308639], displacement = [-0. -0.97416825], geometry = hyperbolic\nproject_to_tangent: xp_norm = 1.3960672530300118, xx_norm = -1.0\nprojection on tangent space = [1.43308639 1.02651673]\nexponential_map: norm_v_tan = 0.16999999999999998\n****xhi = 0.73\nupdated_pt = [0.79658581 1.2784948 ]\n--------------------------------------------------------------------------------\nu = [0.79658581 1.2784948 ], v = [0. 1.], u.v = -1.2784947988821622\nstep = [-0. -1.25535754]\nproject_to_tangent: point_on_manifold = [0.79658581 1.2784948 ], displacement = [-0. -1.25535754], geometry = hyperbolic\nproject_to_tangent: xp_norm = 1.6049680835025524, xx_norm = -0.9999999999999993\nprojection on tangent space = [1.2784948 0.79658581]\nexponential_map: norm_v_tan = 0.17000000000000015\n****xhi = 0.5599999999999997\nupdated_pt = [0.58973172 1.16094078]\n--------------------------------------------------------------------------------\nu = [0.58973172 1.16094078], v = [0. 1.], u.v = -1.1609407820724575\nstep = [-0. -1.69568631]\nproject_to_tangent: point_on_manifold = [0.58973172 1.16094078], displacement = [-0. -1.69568631], geometry = hyperbolic\nproject_to_tangent: xp_norm = 1.9685913885883228, xx_norm = -0.9999999999999994\nprojection on tangent space = [1.16094078 0.58973172]\nexponential_map: norm_v_tan = 0.17000000000000018\n****xhi = 0.38999999999999946\nupdated_pt = [0.39996196 1.07701883]\n--------------------------------------------------------------------------------\nu = [0.39996196 1.07701883], v = [0. 1.], u.v = -1.077018834190403\nstep = [-0. -2.50023777]\nproject_to_tangent: point_on_manifold = [0.39996196 1.07701883], displacement = [-0. -2.50023777], geometry = hyperbolic\nproject_to_tangent: xp_norm = 2.6928031731296347, xx_norm = -0.999999999999999\nprojection on tangent space = [1.07701883 0.39996196]\nexponential_map: norm_v_tan = 0.17000000000000057\n****xhi = 0.21999999999999886\nupdated_pt = [0.22177897 1.02429776]\n--------------------------------------------------------------------------------\nu = [0.22177897 1.02429776], v = [0. 1.], u.v = -1.0242977642749287\nstep = [-0. -4.50899387]\nproject_to_tangent: point_on_manifold = [0.22177897 1.02429776], displacement = [-0. -4.50899387], geometry = hyperbolic\nproject_to_tangent: xp_norm = 4.6185523420281305, xx_norm = -0.9999999999999986\nprojection on tangent space = [1.02429776 0.22177897]\nexponential_map: norm_v_tan = 0.17000000000000265\n****xhi = 0.04999999999999618\nupdated_pt = [0.05002084 1.00125026]\n--------------------------------------------------------------------------------\nu = [0.05002084 1.00125026], v = [0. 1.], u.v = -1.0012502604383682\nstep = [ -0. -19.9916691]\nproject_to_tangent: point_on_manifold = [0.05002084 1.00125026], displacement = [ -0. -19.9916691], geometry = hyperbolic\nproject_to_tangent: xp_norm = 20.01666388955649, xx_norm = -0.9999999999999988\nprojection on tangent space = [1.00125026 0.05002084]\nexponential_map: norm_v_tan = 0.17000000000004153\n****xhi = -0.12000000000004536\nupdated_pt = [-0.12028821 1.00720864]\n--------------------------------------------------------------------------------\nu = [-0.12028821 1.00720864], v = [0. 1.], u.v = -1.0072086441482715\nstep = [-0. -8.31336688]\nproject_to_tangent: point_on_manifold = [-0.12028821 1.00720864], displacement = [-0. -8.31336688], geometry = hyperbolic\nproject_to_tangent: xp_norm = 8.373294985917648, xx_norm = -0.9999999999999989\nprojection on tangent space = [-1.00720864 0.12028821]\nexponential_map: norm_v_tan = 0.17000000000000662\n****xhi = 0.04999999999996126\nupdated_pt = [0.05002084 1.00125026]\n--------------------------------------------------------------------------------\nu = [0.05002084 1.00125026], v = [0. 1.], u.v = -1.0012502604383666\nstep = [ -0. -19.9916691]\nproject_to_tangent: point_on_manifold = [0.05002084 1.00125026], displacement = [ -0. -19.9916691], geometry = hyperbolic\nproject_to_tangent: xp_norm = 20.016663889568893, xx_norm = -0.9999999999999992\nprojection on tangent space = [1.00125026 0.05002084]\nexponential_map: norm_v_tan = 0.17000000000002835\n****xhi = -0.12000000000006707\nupdated_pt = [-0.12028821 1.00720864]\n--------------------------------------------------------------------------------\nu = [-0.12028821 1.00720864], v = [0. 1.], u.v = -1.007208644148275\nstep = [-0. -8.31336688]\nproject_to_tangent: point_on_manifold = [-0.12028821 1.00720864], displacement = [-0. -8.31336688], geometry = hyperbolic\nproject_to_tangent: xp_norm = 8.373294985915622, xx_norm = -1.0000000000000007\nprojection on tangent space = [-1.00720864 0.12028821]\nexponential_map: norm_v_tan = 0.16999999999999563\n****xhi = 0.049999999999928595\nupdated_pt = [0.05002084 1.00125026]\n--------------------------------------------------------------------------------\n[array([1.02651673, 1.43308639]), array([0.79658581, 1.2784948 ]), array([0.58973172, 1.16094078]), array([0.39996196, 1.07701883]), array([0.22177897, 1.02429776]), array([0.05002084, 1.00125026]), array([-0.12028821, 1.00720864]), array([0.05002084, 1.00125026]), array([-0.12028821, 1.00720864]), array([0.05002084, 1.00125026])]\n"
],
[
"fig = plt.figure(figsize=(12,12))\nax = plt.gca(xlim=[-1.2, 1.2], ylim=[0.9, 1.6])\nax.scatter(x0, x1, s=1)\nax.scatter(target[0], target[1], color='r', marker='*',s=200,label = \"Target\")\nfor i, pt in enumerate(updated_pts):\n ax.scatter(\n pt[0], pt[1], \n marker=markers.MarkerStyle('o',fillstyle=\"none\"), \n s=100, label=\"Update {}\".format(i)\n )\n ax.annotate(\"$\\chi$ = {:.3g}\".format(np.arccosh(pt[1])), xy=pt, \n xytext=pt-(0.05,0.05), fontsize=12)\n\nax.legend()",
"_____no_output_____"
]
],
[
[
"Oscillates. Why does $\\chi$ step down in units of learning rate?\n## Test with spherical coordinates",
"_____no_output_____"
]
],
[
[
"target2 = np.array([0., -1.])\ninitial_pt2 = np.array([1., 0.])\nupdated_pts = [initial_pt2]\nfor i in range(1,10):\n updated_pts.append(\n gradient_descent(updated_pts[i-1], \n target2, \n error_differential_eucl, \n geometry=\"spherical\", \n learning_rate=0.2\n )\n\n )\nprint(updated_pts)",
"u = [1. 0.], v = [ 0. -1.], u.v = 0.0\nstep = [-0. 1.]\nproject_to_tangent: point_on_manifold = [1. 0.], displacement = [-0. 1.], geometry = spherical\nproject_to_tangent: xp_norm = 0.0, xx_norm = 1.0\nprojection on tangent space = [-0. 1.]\nexponential_map: norm_v_tan = 0.2\nu = [ 0.98006658 -0.19866933], v = [ 0. -1.], u.v = 0.19866933079506122\nstep = [-0. 1.02033884]\nproject_to_tangent: point_on_manifold = [ 0.98006658 -0.19866933], displacement = [-0. 1.02033884], geometry = spherical\nproject_to_tangent: xp_norm = -0.2027100355086725, xx_norm = 1.0\nprojection on tangent space = [0.19866933 0.98006658]\nexponential_map: norm_v_tan = 0.20000000000000004\nu = [ 0.92106099 -0.38941834], v = [ 0. -1.], u.v = 0.3894183423086505\nstep = [-0. 1.08570443]\nproject_to_tangent: point_on_manifold = [ 0.92106099 -0.38941834], displacement = [-0. 1.08570443], geometry = spherical\nproject_to_tangent: xp_norm = -0.4227932187381619, xx_norm = 1.0\nprojection on tangent space = [0.38941834 0.92106099]\nexponential_map: norm_v_tan = 0.20000000000000004\nu = [ 0.82533561 -0.56464247], v = [ 0. -1.], u.v = 0.5646424733950354\nstep = [-0. 1.21162831]\nproject_to_tangent: point_on_manifold = [ 0.82533561 -0.56464247], displacement = [-0. 1.21162831], geometry = spherical\nproject_to_tangent: xp_norm = -0.6841368083416923, xx_norm = 1.0\nprojection on tangent space = [0.56464247 0.82533561]\nexponential_map: norm_v_tan = 0.20000000000000004\nu = [ 0.69670671 -0.71735609], v = [ 0. -1.], u.v = 0.7173560908995228\nstep = [-0. 1.4353242]\nproject_to_tangent: point_on_manifold = [ 0.69670671 -0.71735609], displacement = [-0. 1.4353242], geometry = spherical\nproject_to_tangent: xp_norm = -1.0296385570503641, xx_norm = 1.0000000000000002\nprojection on tangent space = [0.71735609 0.69670671]\nexponential_map: norm_v_tan = 0.20000000000000007\nu = [ 0.54030231 -0.84147098], v = [ 0. -1.], u.v = 0.8414709848078966\nstep = [-0. 1.85081572]\nproject_to_tangent: point_on_manifold = [ 0.54030231 -0.84147098], displacement = [-0. 1.85081572], geometry = spherical\nproject_to_tangent: xp_norm = -1.557407724654903, xx_norm = 1.0000000000000002\nprojection on tangent space = [0.84147098 0.54030231]\nexponential_map: norm_v_tan = 0.20000000000000007\nu = [ 0.36235775 -0.93203909], v = [ 0. -1.], u.v = 0.9320390859672265\nstep = [-0. 2.7597036]\nproject_to_tangent: point_on_manifold = [ 0.36235775 -0.93203909], displacement = [-0. 2.7597036], geometry = spherical\nproject_to_tangent: xp_norm = -2.572151622126322, xx_norm = 1.0000000000000002\nprojection on tangent space = [0.93203909 0.36235775]\nexponential_map: norm_v_tan = 0.20000000000000018\nu = [ 0.16996714 -0.98544973], v = [ 0. -1.], u.v = 0.9854497299884604\nstep = [-0. 5.88349008]\nproject_to_tangent: point_on_manifold = [ 0.16996714 -0.98544973], displacement = [-0. 5.88349008], geometry = spherical\nproject_to_tangent: xp_norm = -5.79788371548293, xx_norm = 1.0000000000000004\nprojection on tangent space = [0.98544973 0.16996714]\nexponential_map: norm_v_tan = 0.20000000000000112\nu = [-0.02919952 -0.9995736 ], v = [ 0. -1.], u.v = 0.9995736030415054\nstep = [-0. 34.24713561]\nproject_to_tangent: point_on_manifold = [-0.02919952 -0.9995736 ], displacement = [-0. 34.24713561], geometry = spherical\nproject_to_tangent: xp_norm = -34.232532735568924, xx_norm = 1.0000000000000007\nprojection on tangent space = [-0.9995736 0.02919952]\nexponential_map: norm_v_tan = 0.20000000000007603\n[array([1., 0.]), array([ 0.98006658, -0.19866933]), array([ 0.92106099, -0.38941834]), array([ 0.82533561, -0.56464247]), array([ 0.69670671, -0.71735609]), array([ 0.54030231, -0.84147098]), array([ 0.36235775, -0.93203909]), array([ 0.16996714, -0.98544973]), array([-0.02919952, -0.9995736 ]), array([ 0.16996714, -0.98544973])]\n"
],
[
"fig = plt.figure(figsize=(12,12))\nax = plt.gca(xlim=[-1.2, 1.2], ylim=[-1.2,1.2], xlabel=\"$x^0$\", ylabel=\"$x^1$\")\ncircle = patches.Circle((0,0), 1., edgecolor=\"k\", fill=False)\n#ax.add_collection(collections.PatchCollection(patch_list))\nax.add_artist(circle)\nax.scatter(target2[0], target2[1], color='r', marker='*',s=200,label = \"Target\")\nfor i, pt in enumerate(updated_pts):\n ax.scatter(\n pt[0], pt[1], \n marker=markers.MarkerStyle('o',fillstyle=\"none\"), \n s=100, label=\"Update {}\".format(i)\n )\nax.legend()",
"_____no_output_____"
]
],
[
[
"Solution seems to oscillate, but why? pt_N = pt_{N-2}, although pt_{N-1} was closer.",
"_____no_output_____"
]
],
[
[
"target2 = np.array([0., -1.])\ninitial_pt2 = np.array([np.cos(0.49*np.pi), np.sin(0.49*np.pi)])\nupdated_pts = [initial_pt2]\nfor i in range(1,8):\n updated_pts.append(\n gradient_descent(updated_pts[i-1], \n target2, \n error_differential_eucl, \n geometry=\"spherical\", \n learning_rate=.8\n )\n )\n print(\"****theta =\", np.arctan2(updated_pts[i][0], updated_pts[i][1]))\nprint(updated_pts)",
"u = [0.03141076 0.99950656], v = [ 0. -1.], u.v = -0.9995065603657316\nstep = [-0. 31.83622521]\nproject_to_tangent: point_on_manifold = [0.03141076 0.99950656], displacement = [-0. 31.83622521], geometry = spherical\nproject_to_tangent: xp_norm = 31.820515953775725, xx_norm = 1.0000000000000002\nprojection on tangent space = [-0.99950656 0.03141076]\nexponential_map: norm_v_tan = 0.800000000000047\n****theta = 0.8314159265359473\nu = [0.73888621 0.67383023], v = [ 0. -1.], u.v = -0.673830227298802\nstep = [-0. 1.35338837]\nproject_to_tangent: point_on_manifold = [0.73888621 0.67383023], displacement = [-0. 1.35338837], geometry = spherical\nproject_to_tangent: xp_norm = 0.9119539953800536, xx_norm = 0.9999999999999958\nprojection on tangent space = [-0.67383023 0.73888621]\nexponential_map: norm_v_tan = 0.7999999999999986\n****theta = 1.631415926535947\nu = [ 0.99816319 -0.06058248], v = [ 0. -1.], u.v = 0.060582479725725724\nstep = [-0. 1.00184019]\nproject_to_tangent: point_on_manifold = [ 0.99816319 -0.06058248], displacement = [-0. 1.00184019], geometry = spherical\nproject_to_tangent: xp_norm = -0.06069396272127466, xx_norm = 0.9999999999999979\nprojection on tangent space = [0.06058248 0.99816319]\nexponential_map: norm_v_tan = 0.7999999999999999\n****theta = 2.4314159265359474\nu = [ 0.65196778 -0.75824667], v = [ 0. -1.], u.v = 0.7582466674864073\nstep = [-0. 1.53381812]\nproject_to_tangent: point_on_manifold = [ 0.65196778 -0.75824667], displacement = [-0. 1.53381812], geometry = spherical\nproject_to_tangent: xp_norm = -1.1630124773169384, xx_norm = 0.9999999999999991\nprojection on tangent space = [0.75824667 0.65196778]\nexponential_map: norm_v_tan = 0.7999999999999997\n****theta = -3.0517693806436395\nu = [-0.08970254 -0.9959686 ], v = [ 0. -1.], u.v = 0.9959686014300924\nstep = [-0. 11.14795684]\nproject_to_tangent: point_on_manifold = [-0.08970254 -0.9959686 ], displacement = [-0. 11.14795684], geometry = spherical\nproject_to_tangent: xp_norm = -11.103014984947297, xx_norm = 0.9999999999999996\nprojection on tangent space = [-0.9959686 0.08970254]\nexponential_map: norm_v_tan = 0.7999999999999768\n****theta = 2.4314159265359696\nu = [ 0.65196778 -0.75824667], v = [ 0. -1.], u.v = 0.7582466674864216\nstep = [-0. 1.53381812]\nproject_to_tangent: point_on_manifold = [ 0.65196778 -0.75824667], displacement = [-0. 1.53381812], geometry = spherical\nproject_to_tangent: xp_norm = -1.1630124773169896, xx_norm = 0.9999999999999984\nprojection on tangent space = [0.75824667 0.65196778]\nexponential_map: norm_v_tan = 0.7999999999999993\n****theta = -3.0517693806436172\nu = [-0.08970254 -0.9959686 ], v = [ 0. -1.], u.v = 0.9959686014300901\nstep = [-0. 11.14795684]\nproject_to_tangent: point_on_manifold = [-0.08970254 -0.9959686 ], displacement = [-0. 11.14795684], geometry = spherical\nproject_to_tangent: xp_norm = -11.103014984944208, xx_norm = 0.999999999999999\nprojection on tangent space = [-0.9959686 0.08970254]\nexponential_map: norm_v_tan = 0.7999999999999527\n****theta = 2.431415926536016\n[array([0.03141076, 0.99950656]), array([0.73888621, 0.67383023]), array([ 0.99816319, -0.06058248]), array([ 0.65196778, -0.75824667]), array([-0.08970254, -0.9959686 ]), array([ 0.65196778, -0.75824667]), array([-0.08970254, -0.9959686 ]), array([ 0.65196778, -0.75824667])]\n"
],
[
"fig = plt.figure(figsize=(12,12))\nax = plt.gca(xlim=[-1.2, 1.2], ylim=[-1.2,1.2], xlabel=\"$x^0$\", ylabel=\"$x^1$\")\ncircle = patches.Circle((0,0), 1., edgecolor=\"k\", fill=False)\n#ax.add_collection(collections.PatchCollection(patch_list))\nax.add_artist(circle)\nax.scatter(target2[0], target2[1], color='r', marker='*',s=200,label = \"Target\")\nfor i, pt in enumerate(updated_pts):\n ax.scatter(\n pt[0], pt[1], \n marker=markers.MarkerStyle('o',fillstyle=\"none\"), \n s=100, label=\"Update {}\".format(i)\n )\nax.legend()",
"_____no_output_____"
]
],
[
[
"Same oscillatory behaviour.",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
]
] |
4acf3a19cc9a0951906f087df9d6d5fa5c06c939
| 2,698 |
ipynb
|
Jupyter Notebook
|
code/group-weeks-to-periods.ipynb
|
caporaso-lab/exmp-paper1
|
d6bb70b2f3b517ebe8671797abbcc5b5ae3713d5
|
[
"BSD-3-Clause"
] | 1 |
2021-03-10T00:17:58.000Z
|
2021-03-10T00:17:58.000Z
|
code/group-weeks-to-periods.ipynb
|
caporaso-lab/exmp-paper1
|
d6bb70b2f3b517ebe8671797abbcc5b5ae3713d5
|
[
"BSD-3-Clause"
] | null | null | null |
code/group-weeks-to-periods.ipynb
|
caporaso-lab/exmp-paper1
|
d6bb70b2f3b517ebe8671797abbcc5b5ae3713d5
|
[
"BSD-3-Clause"
] | null | null | null | 24.089286 | 91 | 0.547443 |
[
[
[
"import exmp\nimport numpy as np\nimport qiime2\nimport os.path\n\nfrom qiime2.plugins.feature_table.actions import group, filter_features\nfrom qiime2.plugins.diversity.actions import core_metrics_phylogenetic",
"_____no_output_____"
],
[
"grouping_metadata = exmp.load_sample_metadata_grouping()\ntable = exmp.load_table()",
"_____no_output_____"
],
[
"grouped_table = group(table, \n axis='sample', \n metadata=grouping_metadata.get_column('subject-id-period'), \n mode='median-ceiling').grouped_table",
"_____no_output_____"
],
[
"grouped_table.save(\n os.path.join(exmp.data_dir, \"table-grouped-by-period.qza\")) \n",
"_____no_output_____"
],
[
"sample_metadata_grouped_by_period = exmp.load_sample_metadata_grouped_by_period()\nsample_metadata_grouped_by_period.save(exmp.sample_md_grouped_by_period_fp)",
"_____no_output_____"
],
[
"results = core_metrics_phylogenetic(grouped_table, \n metadata=sample_metadata_grouped_by_period,\n phylogeny=exmp.load_phylogeny(),\n sampling_depth=5000)",
"_____no_output_____"
],
[
"for name, artifact in list(zip(results._fields, results)):\n artifact.save(os.path.join(exmp.cm_grouped_by_period_path, name))",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acf452d81bfc7e7a6b91e5711adbe1fd4b676cc
| 75,611 |
ipynb
|
Jupyter Notebook
|
_notebooks/2022-03-20-Python-Line.ipynb
|
leungadh/coding
|
2d31c1c6cc83c371fe3c6fe6d246bf9808d7e54b
|
[
"Apache-2.0"
] | null | null | null |
_notebooks/2022-03-20-Python-Line.ipynb
|
leungadh/coding
|
2d31c1c6cc83c371fe3c6fe6d246bf9808d7e54b
|
[
"Apache-2.0"
] | null | null | null |
_notebooks/2022-03-20-Python-Line.ipynb
|
leungadh/coding
|
2d31c1c6cc83c371fe3c6fe6d246bf9808d7e54b
|
[
"Apache-2.0"
] | null | null | null | 333.088106 | 36,424 | 0.931068 |
[
[
[
"\n# Python Line Plot\n> How to create line plot\n\n- toc: true\n- branch: master\n- badges: true\n- comments: true\n- author: Andy Leung\n- categories: [python, visual]",
"_____no_output_____"
],
[
"## Python Line Plot",
"_____no_output_____"
]
],
[
[
"%matplotlib inline\nimport matplotlib.pyplot as plt\nimport numpy as np",
"_____no_output_____"
]
],
[
[
"### Line plots\n\n**Showing the line plot of X and Y value**\n* x:The x-coordinate\n* y:The y-coordinate",
"_____no_output_____"
]
],
[
[
"k = np.linspace(0,10) #np.linspace (start, finish) equally divine into 50 numbers(default)\nkk = np.sin(x)\nk, kk",
"_____no_output_____"
],
[
"x = np.linspace(0,20)\ny1 = np.sin(x)\ny2 = np.sin(x - np.pi)\n\nplt.figure()\nplt.plot(x,y1)\nplt.plot(x,y2)\nplt.figure;",
"_____no_output_____"
]
],
[
[
"## Implementing plot with \n* Color \n* Linestyle\n* Linewidth\n* Marker, Marker size\n* Label\n* Legend",
"_____no_output_____"
]
],
[
[
"x = np.linspace(0, 20)\ny1 = np.sin(x)\ny2 = np.sin(x - np.pi)\n\nplt.figure()\n\nplt.plot(x,\n y1,\n color='black',\n linestyle='-',\n linewidth=2,\n marker='s',\n markersize=6,\n label='y1')\n\nplt.plot(x,\n y2,\n color='red',\n linestyle='--',\n linewidth=2,\n marker='^',\n markersize=6,\n label='y2')\n\nplt.legend()\n;",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
]
] |
4acf4998c48dd8646960e1b83385bb6397b722be
| 3,824 |
ipynb
|
Jupyter Notebook
|
0.17/_downloads/19f42385e184c24343e0e939f7de62b5/plot_forward_sensitivity_maps.ipynb
|
drammock/mne-tools.github.io
|
5d3a104d174255644d8d5335f58036e32695e85d
|
[
"BSD-3-Clause"
] | null | null | null |
0.17/_downloads/19f42385e184c24343e0e939f7de62b5/plot_forward_sensitivity_maps.ipynb
|
drammock/mne-tools.github.io
|
5d3a104d174255644d8d5335f58036e32695e85d
|
[
"BSD-3-Clause"
] | null | null | null |
0.17/_downloads/19f42385e184c24343e0e939f7de62b5/plot_forward_sensitivity_maps.ipynb
|
drammock/mne-tools.github.io
|
5d3a104d174255644d8d5335f58036e32695e85d
|
[
"BSD-3-Clause"
] | null | null | null | 42.488889 | 1,043 | 0.591789 |
[
[
[
"%matplotlib inline",
"_____no_output_____"
]
],
[
[
"\n# Display sensitivity maps for EEG and MEG sensors\n\n\nSensitivity maps can be produced from forward operators that\nindicate how well different sensor types will be able to detect\nneural currents from different regions of the brain.\n\nTo get started with forward modeling see `tut_forward`.\n\n\n",
"_____no_output_____"
]
],
[
[
"# Author: Eric Larson <[email protected]>\n#\n# License: BSD (3-clause)\n\nimport mne\nfrom mne.datasets import sample\nimport matplotlib.pyplot as plt\n\nprint(__doc__)\n\ndata_path = sample.data_path()\n\nraw_fname = data_path + '/MEG/sample/sample_audvis_raw.fif'\nfwd_fname = data_path + '/MEG/sample/sample_audvis-meg-eeg-oct-6-fwd.fif'\n\nsubjects_dir = data_path + '/subjects'\n\n# Read the forward solutions with surface orientation\nfwd = mne.read_forward_solution(fwd_fname)\nmne.convert_forward_solution(fwd, surf_ori=True, copy=False)\nleadfield = fwd['sol']['data']\nprint(\"Leadfield size : %d x %d\" % leadfield.shape)",
"_____no_output_____"
]
],
[
[
"Compute sensitivity maps\n\n",
"_____no_output_____"
]
],
[
[
"grad_map = mne.sensitivity_map(fwd, ch_type='grad', mode='fixed')\nmag_map = mne.sensitivity_map(fwd, ch_type='mag', mode='fixed')\neeg_map = mne.sensitivity_map(fwd, ch_type='eeg', mode='fixed')",
"_____no_output_____"
]
],
[
[
"Show gain matrix a.k.a. leadfield matrix with sensitivity map\n\n",
"_____no_output_____"
]
],
[
[
"picks_meg = mne.pick_types(fwd['info'], meg=True, eeg=False)\npicks_eeg = mne.pick_types(fwd['info'], meg=False, eeg=True)\n\nfig, axes = plt.subplots(2, 1, figsize=(10, 8), sharex=True)\nfig.suptitle('Lead field matrix (500 dipoles only)', fontsize=14)\nfor ax, picks, ch_type in zip(axes, [picks_meg, picks_eeg], ['meg', 'eeg']):\n im = ax.imshow(leadfield[picks, :500], origin='lower', aspect='auto',\n cmap='RdBu_r')\n ax.set_title(ch_type.upper())\n ax.set_xlabel('sources')\n ax.set_ylabel('sensors')\n plt.colorbar(im, ax=ax, cmap='RdBu_r')\nplt.show()\n\nplt.figure()\nplt.hist([grad_map.data.ravel(), mag_map.data.ravel(), eeg_map.data.ravel()],\n bins=20, label=['Gradiometers', 'Magnetometers', 'EEG'],\n color=['c', 'b', 'k'])\nplt.legend()\nplt.title('Normal orientation sensitivity')\nplt.xlabel('sensitivity')\nplt.ylabel('count')\nplt.show()\n\ngrad_map.plot(time_label='Gradiometer sensitivity', subjects_dir=subjects_dir,\n clim=dict(lims=[0, 50, 100]))",
"_____no_output_____"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4acf51f9f55685b0c213177f74cded706f839790
| 230,544 |
ipynb
|
Jupyter Notebook
|
plot_success_rate/.ipynb_checkpoints/calculate_successes-checkpoint.ipynb
|
adit98/good_robot
|
e7cae029c33bce1c9584c7328016579db4742221
|
[
"BSD-2-Clause"
] | 69 |
2019-09-30T13:42:02.000Z
|
2022-03-28T08:37:51.000Z
|
plot_success_rate/.ipynb_checkpoints/calculate_successes-checkpoint.ipynb
|
adit98/good_robot
|
e7cae029c33bce1c9584c7328016579db4742221
|
[
"BSD-2-Clause"
] | 5 |
2019-10-23T20:03:42.000Z
|
2021-07-10T09:43:50.000Z
|
plot_success_rate/.ipynb_checkpoints/calculate_successes-checkpoint.ipynb
|
adit98/good_robot
|
e7cae029c33bce1c9584c7328016579db4742221
|
[
"BSD-2-Clause"
] | 18 |
2019-11-17T20:57:46.000Z
|
2022-03-15T10:46:25.000Z
| 649.419718 | 82,336 | 0.9449 |
[
[
[
"import os\nimport numpy as np\nimport matplotlib.pyplot as plt\nfrom matplotlib.ticker import PercentFormatter\nfrom glob import glob\n%matplotlib inline",
"_____no_output_____"
]
],
[
[
"# Instructions for Use\n\nThe \"Main Functions\" section contains functions which return the success rate to be plotted as well as lower and upper bounds for uncertainty plotting.\n\nTo plot a given log, simply copy the code below two cells and replace `log_dir` with the path of the logs you wish to plot.\n\n```\nwindow = 1000\nmax_iter = None # Can set to an integer\nlog_dir = 'LOG_DIR'\ntitle = 'Stack 4 Blocks, Training'\nplot_it(log_dir, title, window=window, max_iter=max_iter)\n```",
"_____no_output_____"
],
[
"# Main Functions",
"_____no_output_____"
]
],
[
[
"def get_grasp_success_rate(actions, rewards=None, window=200, reward_threshold=0.5):\n \"\"\"Evaluate moving window of grasp success rate\n actions: Nx4 array of actions giving [id, rotation, i, j]\n \n \"\"\"\n grasps = actions[:, 0] == 1\n if rewards is None:\n places = actions[:, 0] == 2\n success_rate = np.zeros(actions.shape[0] - 1)\n lower = np.zeros_like(success_rate)\n upper = np.zeros_like(success_rate)\n for i in range(success_rate.shape[0]):\n start = max(i - window, 0)\n if rewards is None:\n successes = places[start+1: i+2][grasps[start:i+1]]\n else:\n successes = (rewards[start: i+1] > reward_threshold)[grasps[start:i+1]]\n success_rate[i] = successes.mean()\n var = np.sqrt(success_rate[i] * (1 - success_rate[i]) / successes.shape[0])\n lower[i] = success_rate[i] + 3*var\n upper[i] = success_rate[i] - 3*var\n lower = np.clip(lower, 0, 1)\n upper = np.clip(upper, 0, 1)\n return success_rate, lower, upper\n\ndef get_place_success_rate(stack_height, actions, include_push=False, window=200, hot_fix=False, max_height=4):\n \"\"\"\n stack_heights: length N array of integer stack heights\n actions: Nx4 array of actions giving [id, rotation, i, j]\n hot_fix: fix the stack_height bug, where the trial didn't end on successful pushes, which reached a stack of 4.\n \n where id=0 is a push, id=1 is grasp, and id=2 is place.\n \n \"\"\"\n if hot_fix:\n indices = np.logical_or(stack_height < 4, np.array([True] + list(stack_height[:-1] < 4)))\n actions = actions[:stack_height.shape[0]][indices]\n stack_height = stack_height[indices]\n \n if include_push:\n success_possible = actions[:, 0] == 2\n else:\n success_possible = np.logical_or(actions[:, 0] == 0, actions[:, 0] == 2)\n \n stack_height_increased = np.zeros_like(stack_height, np.bool)\n stack_height_increased[0] = False\n stack_height_increased[1:] = stack_height[1:] > stack_height[:-1]\n \n success_rate = np.zeros_like(stack_height)\n lower = np.zeros_like(success_rate)\n upper = np.zeros_like(success_rate)\n for i in range(stack_height.shape[0]):\n start = max(i - window, 0)\n successes = stack_height_increased[start:i+1][success_possible[start:i+1]]\n success_rate[i] = successes.mean()\n success_rate[np.isnan(success_rate)] = 0\n var = np.sqrt(success_rate[i] * (1 - success_rate[i]) / successes.shape[0])\n lower[i] = success_rate[i] + 3*var\n upper[i] = success_rate[i] - 3*var\n lower = np.clip(lower, 0, 1)\n upper = np.clip(upper, 0, 1)\n return success_rate, lower, upper\n\ndef get_action_efficiency(stack_height, window=200, ideal_actions_per_trial=6, max_height=4):\n \"\"\"Calculate the running action efficiency from successful trials.\n\n trials: array giving the number of trials up to iteration i (TODO: unused?)\n min_actions: ideal number of actions per trial\n \n Formula: successful_trial_count * ideal_actions_per_trial / window_size\n \"\"\"\n\n success = stack_height == max_height\n efficiency = np.zeros_like(stack_height, np.float64)\n lower = np.zeros_like(efficiency)\n upper = np.zeros_like(efficiency)\n for i in range(1, efficiency.shape[0]):\n start = max(i - window, 1)\n window_size = min(i, window)\n num_trials = success[start:i+1].sum()\n efficiency[i] = num_trials * ideal_actions_per_trial / window_size\n var = efficiency[i] / np.sqrt(window_size)\n lower[i] = efficiency[i] + 3*var\n upper[i] = efficiency[i] - 3*var\n lower = np.clip(lower, 0, 1)\n upper = np.clip(upper, 0, 1)\n return efficiency, lower, upper\n\ndef get_grasp_action_efficiency(actions, rewards, reward_threshold=0.5, window=200, ideal_actions_per_trial=3):\n \"\"\"Get grasp efficiency from when the trial count increases.\n \n \"\"\"\n grasps = actions[:, 0] == 1\n efficiency = np.zeros_like(rewards, np.float64)\n lower = np.zeros_like(efficiency)\n upper = np.zeros_like(efficiency)\n for i in range(efficiency.shape[0]):\n start = max(i - window, 0)\n window_size = min(i+1, window)\n successful = rewards[start: i+1] > reward_threshold\n successful_grasps = successful[grasps[start:start+successful.shape[0]]].sum()\n efficiency[i] = successful_grasps / window_size\n var = efficiency[i] / np.sqrt(window_size)\n lower[i] = efficiency[i] + 3*var\n upper[i] = efficiency[i] - 3*var\n lower = np.clip(lower, 0, 1)\n upper = np.clip(upper, 0, 1)\n return efficiency, lower, upper",
"_____no_output_____"
],
[
"def plot_it(log_dir, title, window=1000, colors=['tab:blue', 'tab:green', 'tab:orange'], alpha=0.35, mult=100, max_iter=None, place=False):\n if place:\n heights = np.loadtxt(os.path.join(log_dir, 'transitions', 'stack-height.log.txt'))\n rewards = None\n else:\n rewards = np.loadtxt(os.path.join(log_dir, 'transitions', 'reward-value.log.txt'))\n actions = np.loadtxt(os.path.join(log_dir, 'transitions', 'executed-action.log.txt'))\n trials = np.loadtxt(os.path.join(log_dir, 'transitions', 'trial.log.txt'))\n \n if max_iter is not None:\n if place:\n heights = heights[:max_iter]\n else:\n rewards = rewards[:max_iter]\n actions = actions[:max_iter]\n trials = trials[:max_iter]\n \n grasp_rate, grasp_lower, grasp_upper = get_grasp_success_rate(actions, rewards=rewards, window=window)\n if place:\n if 'row' in log_dir or 'row' in title.lower():\n place_rate, place_lower, place_upper = get_place_success_rate(heights, actions, include_push=True, hot_fix=True, window=window)\n else:\n place_rate, place_lower, place_upper = get_place_success_rate(heights, actions, window=window)\n eff, eff_lower, eff_upper = get_action_efficiency(heights, window=window)\n else:\n eff, eff_lower, eff_upper = get_grasp_action_efficiency(actions, rewards, window=window)\n\n plt.plot(mult*grasp_rate, color=colors[0], label='Grasp Success Rate')\n if place:\n plt.plot(mult*place_rate, color=colors[1], label='Place Success Rate')\n plt.plot(mult*eff, color=colors[2], label='Action Efficiency')\n\n plt.fill_between(np.arange(1, grasp_rate.shape[0]+1), \n mult*grasp_lower, mult*grasp_upper,\n color=colors[0], alpha=alpha)\n if place:\n plt.fill_between(np.arange(1, place_rate.shape[0]+1), \n mult*place_lower, mult*place_upper,\n color=colors[1], alpha=alpha)\n plt.fill_between(np.arange(1, eff.shape[0]+1), \n mult*eff_lower, mult*eff_upper,\n color=colors[2], alpha=alpha)\n\n ax = plt.gca()\n plt.xlabel('Iteration')\n plt.ylabel('Running Mean')\n plt.title(title)\n plt.legend()\n ax.yaxis.set_major_formatter(PercentFormatter())\n plt.savefig(log_dir + '_success_plot.pdf')",
"_____no_output_____"
]
],
[
[
"# Any-object Stacking",
"_____no_output_____"
]
],
[
[
"window = 1000\nmax_iter = None\nlog_dir = 'any-stack-v2-steps-37k'\ntitle = 'Stack 4 Blocks, Training'\nplot_it(log_dir, title, window=window, max_iter=max_iter)",
"/usr/local/lib/python3.7/site-packages/ipykernel_launcher.py:55: RuntimeWarning: Mean of empty slice.\n/usr/local/lib/python3.7/site-packages/ipykernel_launcher.py:57: RuntimeWarning: invalid value encountered in double_scalars\n"
]
],
[
[
"# Arranging Rows",
"_____no_output_____"
]
],
[
[
"window = 1000\nmax_iter = None\nlog_dir = '../logs/2019-09-13.19-55-21-train-rows-no-images-16.5k'\ntitle = 'Arrange 4 Blocks in Rows'\nplot_it(log_dir, title, window=window, max_iter=max_iter)",
"/usr/local/lib/python3.7/site-packages/ipykernel_launcher.py:55: RuntimeWarning: Mean of empty slice.\n/usr/local/lib/python3.7/site-packages/ipykernel_launcher.py:57: RuntimeWarning: invalid value encountered in double_scalars\n"
]
],
[
[
"# Push + Grasp",
"_____no_output_____"
]
],
[
[
"window = 200\nmax_iter = 5000\nlog_dir = 'train-grasp-place-split-efficientnet-21k-acc'\ntitle = 'Push + Grasp Training'\nplot_it(log_dir, title, window=window, max_iter=max_iter, place=False)",
"_____no_output_____"
]
]
] |
[
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4acf54c3bc16574acb8a1533f64308ccf5a2680b
| 172,047 |
ipynb
|
Jupyter Notebook
|
notebooks/archive/troubleshooting_in_man_filter.ipynb
|
allisoncstafford/walk_risk_engine
|
2afed49dd7d1f93c50a5f4c76e16a79a4e053ca6
|
[
"MIT"
] | 1 |
2019-11-21T00:38:52.000Z
|
2019-11-21T00:38:52.000Z
|
notebooks/archive/troubleshooting_in_man_filter.ipynb
|
allisoncstafford/walk_risk_engine
|
2afed49dd7d1f93c50a5f4c76e16a79a4e053ca6
|
[
"MIT"
] | 1 |
2021-06-02T00:42:49.000Z
|
2021-06-02T00:42:49.000Z
|
notebooks/archive/troubleshooting_in_man_filter.ipynb
|
allisoncstafford/walk_risk_engine
|
2afed49dd7d1f93c50a5f4c76e16a79a4e053ca6
|
[
"MIT"
] | null | null | null | 278.393204 | 46,676 | 0.914244 |
[
[
[
"# import necessary packages\nimport json\nimport requests\nimport pandas as pd\nimport polyline\nimport geopandas as gpd\nfrom shapely.geometry import LineString, Point\nimport numpy as np\nfrom itertools import product\nfrom haversine import haversine, Unit\nfrom shapely.ops import nearest_points\nimport os\nfrom matplotlib import pyplot as plt\n%matplotlib inline",
"_____no_output_____"
],
[
"def create_pt_grid(minx, miny, maxx, maxy):\n \"\"\"creates a grid of points (lat/longs) in the range specified. lat longs \n are rounded to hundredth place\n\n Args:\n minx: minimum longitude\n miny: minimum latitude\n maxx: maximum longitude\n maxy: maximum latitude\n\n Returns: DataFrame of all lat/long combinations in region\n \"\"\"\n lats = range(int(miny*1000), int(maxy*1000 +1))\n longs = range(int(minx*1000), int(maxx*1000 +1))\n ll_df = pd.DataFrame(product(lats, longs), \n columns=['lat1000', 'long1000'])\n ll_df['geometry'] = [Point(x, y) for x, y in zip(ll_df['long1000'], \n ll_df['lat1000'])]\n return ll_df",
"_____no_output_____"
],
[
"def get_pts_near_path(line, distance):\n \"\"\"returns all lat/longs within specified distance of line that are in \n manhattan\n \n Args:\n line: shapely linestring of route\n distance: maximum distance from path for returned points\n\n Returns:\n pandas dataframe of all points within distance from line\n \"\"\"\n # get line bounds\n (minx, miny, maxx, maxy) = line.bounds\n \n # extract max/min values with buffer area\n minx = round(minx, 3) -0.002\n miny = round(miny, 3) -0.002\n maxx = round(maxx, 3) + 0.002\n maxy = round(maxy, 3) + 0.002\n\n # load manhattan lat_longs\n manhattan_pts = pd.read_csv('models/man_lat_longs.csv')\n\n # brute force fix for floating point error\n manhattan_pts['latitude'] = manhattan_pts['lat1000']/1000\n manhattan_pts['longitdue'] = manhattan_pts['long1000']/1000\n manhattan_pts = manhattan_pts.loc[:, ['latitude', 'longitude']]\n \n # create a df of all lat, longs w/in bounds\n all_pts = create_pt_grid(minx, miny, maxx, maxy)\n \n # remove pts not in manhattan\n all_pts = pd.merge(all_pts, manhattan_pts, \n on=['latitude', 'longitude'],\n how='inner')\n\n # flag points in the grid in manhattan as on/within distance of path\n all_pts['on_path'] = get_on_path(all_pts['geometry'], distance, line)\n return pd.DataFrame(all_pts.loc[(all_pts['on_path']==True)])",
"_____no_output_____"
],
[
"practice_grid = create_pt_grid(-74.000, 40.750, -73.960183, 40.7800)",
"_____no_output_____"
],
[
"practice_grid_gdf = gpd.GeoDataFrame(practice_grid)",
"_____no_output_____"
],
[
"fig, ax = plt.subplots(figsize=(20,20))\npractice_grid_gdf.plot(ax=ax)",
"_____no_output_____"
],
[
"manhattan_pts = pd.read_csv('/Users/allisonhonold/ds0805/walk_proj/walk_risk_engine/data/csv/man_lat_longs.csv')",
"_____no_output_____"
],
[
"manhattan_pts.head()",
"_____no_output_____"
],
[
"fig5, ax5 = plt.subplots(figsize=(20,20))\nax5.scatter(manhattan_pts['long1000'], manhattan_pts['lat1000'], alpha=.3)",
"_____no_output_____"
],
[
"all_pts = pd.merge(practice_grid_gdf, manhattan_pts.loc[:,['lat1000', 'long1000']], \n on=['lat1000', 'long1000'],\n how='inner')",
"_____no_output_____"
],
[
"all_pts.head()",
"_____no_output_____"
],
[
"fig2, ax2 = plt.subplots(figsize=(20,20))\nall_pts.plot(ax=ax2)",
"_____no_output_____"
],
[
"manhattan_pts.shape",
"_____no_output_____"
],
[
"practice_grid_gdf.shape",
"_____no_output_____"
],
[
"practice_grid_gdf.head()",
"_____no_output_____"
],
[
"man_gdf = gpd.GeoDataFrame(manhattan_pts, geometry=[Point(x, y) for x, y in zip(manhattan_pts['long1000'], \n manhattan_pts['lat1000'])])\nfig3, ax3 = plt.subplots(figsize=(20,20))\nman_gdf.plot(ax=ax3, markersize=2, alpha=.5)",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acf56386c758367d7b00e75c68ffa085e6aae17
| 68,959 |
ipynb
|
Jupyter Notebook
|
Python Projects/Which Debts Are Worth the Bank's Effort_/notebook.ipynb
|
kingabzpro/Data-Scince-Project-on-DataCamp
|
f49890851cd9637fe267d18c074847c764ab7613
|
[
"MIT"
] | 1 |
2020-11-27T20:15:39.000Z
|
2020-11-27T20:15:39.000Z
|
Python Projects/Which Debts Are Worth the Bank's Effort_/notebook.ipynb
|
kingabzpro/Data-Scince-Project-on-DataCamp
|
f49890851cd9637fe267d18c074847c764ab7613
|
[
"MIT"
] | null | null | null |
Python Projects/Which Debts Are Worth the Bank's Effort_/notebook.ipynb
|
kingabzpro/Data-Scince-Project-on-DataCamp
|
f49890851cd9637fe267d18c074847c764ab7613
|
[
"MIT"
] | 1 |
2021-07-27T20:52:06.000Z
|
2021-07-27T20:52:06.000Z
| 68,959 | 68,959 | 0.775794 |
[
[
[
"## 1. Regression discontinuity: banking recovery\n<p>After a debt has been legally declared \"uncollectable\" by a bank, the account is considered \"charged-off.\" But that doesn't mean the bank <strong><em>walks away</em></strong> from the debt. They still want to collect some of the money they are owed. The bank will score the account to assess the expected recovery amount, that is, the expected amount that the bank may be able to receive from the customer in the future. This amount is a function of the probability of the customer paying, the total debt, and other factors that impact the ability and willingness to pay.</p>\n<p>The bank has implemented different recovery strategies at different thresholds (\\$1000, \\$2000, etc.) where the greater the expected recovery amount, the more effort the bank puts into contacting the customer. For low recovery amounts (Level 0), the bank just adds the customer's contact information to their automatic dialer and emailing system. For higher recovery strategies, the bank incurs more costs as they leverage human resources in more efforts to obtain payments. Each additional level of recovery strategy requires an additional \\$50 per customer so that customers in the Recovery Strategy Level 1 cost the company \\$50 more than those in Level 0. Customers in Level 2 cost \\$50 more than those in Level 1, etc. </p>\n<p><strong>The big question</strong>: does the extra amount that is recovered at the higher strategy level exceed the extra \\$50 in costs? In other words, was there a jump (also called a \"discontinuity\") of more than \\$50 in the amount recovered at the higher strategy level? We'll find out in this notebook.</p>\n<p>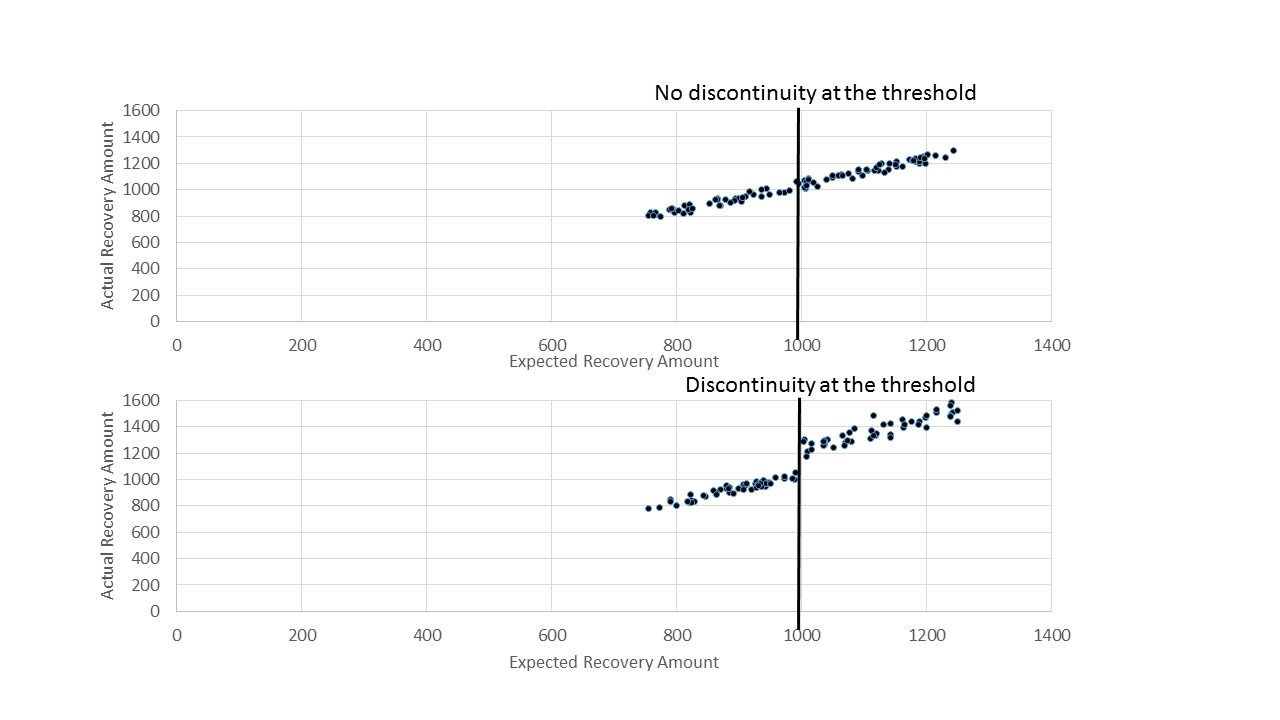</p>\n<p>First, we'll load the banking dataset and look at the first few rows of data. This lets us understand the dataset itself and begin thinking about how to analyze the data.</p>",
"_____no_output_____"
]
],
[
[
"# Import modules\nimport pandas as pd\nimport numpy as np\n\n# Read in dataset\ndf = pd.read_csv(\"datasets/bank_data.csv\")\n\n# Print the first few rows of the DataFrame\ndf.head()",
"_____no_output_____"
]
],
[
[
"## 2. Graphical exploratory data analysis\n<p>The bank has implemented different recovery strategies at different thresholds (\\$1000, \\$2000, \\$3000 and \\$5000) where the greater the Expected Recovery Amount, the more effort the bank puts into contacting the customer. Zeroing in on the first transition (between Level 0 and Level 1) means we are focused on the population with Expected Recovery Amounts between \\$0 and \\$2000 where the transition between Levels occurred at \\$1000. We know that the customers in Level 1 (expected recovery amounts between \\$1001 and \\$2000) received more attention from the bank and, by definition, they had higher Expected Recovery Amounts than the customers in Level 0 (between \\$1 and \\$1000).</p>\n<p>Here's a quick summary of the Levels and thresholds again:</p>\n<ul>\n<li>Level 0: Expected recovery amounts >\\$0 and <=\\$1000</li>\n<li>Level 1: Expected recovery amounts >\\$1000 and <=\\$2000</li>\n<li>The threshold of \\$1000 separates Level 0 from Level 1</li>\n</ul>\n<p>A key question is whether there are other factors besides Expected Recovery Amount that also varied systematically across the \\$1000 threshold. For example, does the customer age show a jump (discontinuity) at the \\$1000 threshold or does that age vary smoothly? We can examine this by first making a scatter plot of the age as a function of Expected Recovery Amount for a small window of Expected Recovery Amount, \\$0 to \\$2000. This range covers Levels 0 and 1.</p>",
"_____no_output_____"
]
],
[
[
"# Scatter plot of Age vs. Expected Recovery Amount\nfrom matplotlib import pyplot as plt\n%matplotlib inline\nplt.scatter(x=df['expected_recovery_amount'], y=df['age'], c=\"g\", s=2)\nplt.xlim(0, 2000)\nplt.ylim(0, 60)\nplt.xlabel(\"Expected Recovery Amount\")\nplt.ylabel(\"Age\")\nplt.legend(loc=2)\nplt.show()",
"/usr/local/lib/python3.6/dist-packages/matplotlib/figure.py:2267: UserWarning: This figure includes Axes that are not compatible with tight_layout, so results might be incorrect.\n warnings.warn(\"This figure includes Axes that are not compatible \"\n"
]
],
[
[
"## 3. Statistical test: age vs. expected recovery amount\n<p>We want to convince ourselves that variables such as age and sex are similar above and below the \\$1000 Expected Recovery Amount threshold. This is important because we want to be able to conclude that differences in the actual recovery amount are due to the higher Recovery Strategy and not due to some other difference like age or sex.</p>\n<p>The scatter plot of age versus Expected Recovery Amount did not show an obvious jump around \\$1000. We will now do statistical analysis examining the average age of the customers just above and just below the threshold. We can start by exploring the range from \\$900 to \\$1100.</p>\n<p>For determining if there is a difference in the ages just above and just below the threshold, we will use the Kruskal-Wallis test, a statistical test that makes no distributional assumptions.</p>",
"_____no_output_____"
]
],
[
[
"# Import stats module\nfrom scipy import stats\n\n# Compute average age just below and above the threshold\nera_900_1100 = df.loc[(df['expected_recovery_amount']<1100) & \n (df['expected_recovery_amount']>=900)]\nby_recovery_strategy = era_900_1100.groupby(['recovery_strategy'])\nby_recovery_strategy['age'].describe().unstack()\n\n# Perform Kruskal-Wallis test \nLevel_0_age = era_900_1100.loc[df['recovery_strategy']==\"Level 0 Recovery\"]['age']\nLevel_1_age = era_900_1100.loc[df['recovery_strategy']==\"Level 1 Recovery\"]['age']\nstats.kruskal(Level_0_age,Level_1_age) ",
"_____no_output_____"
]
],
[
[
"## 4. Statistical test: sex vs. expected recovery amount\n<p>We have seen that there is no major jump in the average customer age just above and just \nbelow the \\$1000 threshold by doing a statistical test as well as exploring it graphically with a scatter plot. </p>\n<p>We want to also test that the percentage of customers that are male does not jump across the \\$1000 threshold. We can start by exploring the range of \\$900 to \\$1100 and later adjust this range.</p>\n<p>We can examine this question statistically by developing cross-tabs as well as doing chi-square tests of the percentage of customers that are male vs. female.</p>",
"_____no_output_____"
]
],
[
[
"# Number of customers in each category\ncrosstab = pd.crosstab(df.loc[(df['expected_recovery_amount']<1100) & \n (df['expected_recovery_amount']>=900)]['recovery_strategy'], \n df['sex'])\nprint(crosstab)\n\n# Chi-square test\nchi2_stat, p_val, dof, ex = stats.chi2_contingency(crosstab)\nprint(p_val)",
"sex Female Male\nrecovery_strategy \nLevel 0 Recovery 32 57\nLevel 1 Recovery 39 55\n0.5377947810444592\n"
]
],
[
[
"## 5. Exploratory graphical analysis: recovery amount\n<p>We are now reasonably confident that customers just above and just below the \\$1000 threshold are, on average, similar in their average age and the percentage that are male. </p>\n<p>It is now time to focus on the key outcome of interest, the actual recovery amount.</p>\n<p>A first step in examining the relationship between the actual recovery amount and the expected recovery amount is to develop a scatter plot where we want to focus our attention at the range just below and just above the threshold. Specifically, we will develop a scatter plot of Expected Recovery Amount (X) versus Actual Recovery Amount (Y) for Expected Recovery Amounts between \\$900 to \\$1100. This range covers Levels 0 and 1. A key question is whether or not we see a discontinuity (jump) around the \\$1000 threshold.</p>",
"_____no_output_____"
]
],
[
[
"# Scatter plot of Actual Recovery Amount vs. Expected Recovery Amount \nplt.scatter(x=df['expected_recovery_amount'], y=df['actual_recovery_amount'], c=\"g\", s=2)\nplt.xlim(900, 1100)\nplt.ylim(0, 2000)\nplt.xlabel(\"Expected Recovery Amount\")\nplt.ylabel(\"Actual Recovery Amount\")\nplt.legend(loc=2)\n# ... YOUR CODE FOR TASK 5 ...",
"_____no_output_____"
]
],
[
[
"## 6. Statistical analysis: recovery amount\n<p>As we did with age, we can perform statistical tests to see if the actual recovery amount has a discontinuity above the \\$1000 threshold. We are going to do this for two different windows of the expected recovery amount \\$900 to \\$1100 and for a narrow range of \\$950 to \\$1050 to see if our results are consistent.</p>\n<p>Again, we will use the Kruskal-Wallis test.</p>\n<p>We will first compute the average actual recovery amount for those customers just below and just above the threshold using a range from \\$900 to \\$1100. Then we will perform a Kruskal-Wallis test to see if the actual recovery amounts are different just above and just below the threshold. Once we do that, we will repeat these steps for a smaller window of \\$950 to \\$1050.</p>",
"_____no_output_____"
]
],
[
[
"# Compute average actual recovery amount just below and above the threshold\nby_recovery_strategy['actual_recovery_amount'].describe().unstack()\n\n# Perform Kruskal-Wallis test\nLevel_0_actual = era_900_1100.loc[df['recovery_strategy']=='Level 0 Recovery']['actual_recovery_amount']\nLevel_1_actual = era_900_1100.loc[df['recovery_strategy']=='Level 1 Recovery']['actual_recovery_amount']\nstats.kruskal(Level_0_actual,Level_1_actual) \n\n# Repeat for a smaller range of $950 to $1050\nera_950_1050 = df.loc[(df['expected_recovery_amount']<1050) & \n (df['expected_recovery_amount']>=950)]\nLevel_0_actual = era_950_1050.loc[df['recovery_strategy']=='Level 0 Recovery']['actual_recovery_amount']\nLevel_1_actual = era_950_1050.loc[df['recovery_strategy']=='Level 1 Recovery']['actual_recovery_amount']\nstats.kruskal(Level_0_actual,Level_1_actual)",
"_____no_output_____"
]
],
[
[
"## 7. Regression modeling: no threshold\n<p>We now want to take a regression-based approach to estimate the program impact at the \\$1000 threshold using data that is just above and below the threshold. </p>\n<p>We will build two models. The first model does not have a threshold while the second will include a threshold.</p>\n<p>The first model predicts the actual recovery amount (dependent variable) as a function of the expected recovery amount (independent variable). We expect that there will be a strong positive relationship between these two variables. </p>\n<p>We will examine the adjusted R-squared to see the percent of variance explained by the model. In this model, we are not representing the threshold but simply seeing how the variable used for assigning the customers (expected recovery amount) relates to the outcome variable (actual recovery amount).</p>",
"_____no_output_____"
]
],
[
[
"# Import statsmodels\nimport statsmodels.api as sm\n\n# Define X and y\nX = era_900_1100['expected_recovery_amount']\ny = era_900_1100['actual_recovery_amount']\nX = sm.add_constant(X)\n\n# Build linear regression model\nmodel = sm.OLS(y, X).fit()\npredictions = model.predict(X)\n\n# Print out the model summary statistics\nmodel.summary()",
"_____no_output_____"
]
],
[
[
"## 8. Regression modeling: adding true threshold\n<p>From the first model, we see that the expected recovery amount's regression coefficient is statistically significant. </p>\n<p>The second model adds an indicator of the true threshold to the model (in this case at \\$1000). </p>\n<p>We will create an indicator variable (either a 0 or a 1) that represents whether or not the expected recovery amount was greater than \\$1000. When we add the true threshold to the model, the regression coefficient for the true threshold represents the additional amount recovered due to the higher recovery strategy. That is to say, the regression coefficient for the true threshold measures the size of the discontinuity for customers just above and just below the threshold.</p>\n<p>If the higher recovery strategy helped recovery more money, then the regression coefficient of the true threshold will be greater than zero. If the higher recovery strategy did not help recovery more money, then the regression coefficient will not be statistically significant.</p>",
"_____no_output_____"
]
],
[
[
"#Create indicator (0 or 1) for expected recovery amount >= $1000\ndf['indicator_1000'] = np.where(df['expected_recovery_amount']<1000, 0, 1)\nera_900_1100 = df.loc[(df['expected_recovery_amount']<1100) & \n (df['expected_recovery_amount']>=900)]\n\n# Define X and y\nX = era_900_1100[['expected_recovery_amount','indicator_1000']]\ny = era_900_1100['actual_recovery_amount']\nX = sm.add_constant(X)\n\n# Build linear regression model\nmodel = sm.OLS(y,X).fit()\n\n# Print the model summary\nmodel.summary()",
"_____no_output_____"
]
],
[
[
"## 9. Regression modeling: adjusting the window\n<p>The regression coefficient for the true threshold was statistically significant with an estimated impact of around \\$278. This is much larger than the \\$50 per customer needed to run this higher recovery strategy. </p>\n<p>Before showing this to our manager, we want to convince ourselves that this result wasn't due to choosing an expected recovery amount window of \\$900 to \\$1100. Let's repeat this analysis for the window from \\$950 to \\$1050 to see if we get similar results.</p>\n<p>The answer? Whether we use a wide (\\$900 to \\$1100) or narrower window (\\$950 to \\$1050), the incremental recovery amount at the higher recovery strategy is much greater than the \\$50 per customer it costs for the higher recovery strategy. So we conclude that the higher recovery strategy is worth the extra cost of \\$50 per customer.</p>",
"_____no_output_____"
]
],
[
[
"# Redefine era_950_1050 so the indicator variable is included\nera_950_1050 = df.loc[(df['expected_recovery_amount']<1050) & \n (df['expected_recovery_amount']>=950)]\n\n# Define X and y \nX = era_950_1050[['expected_recovery_amount','indicator_1000']]\ny = era_950_1050['actual_recovery_amount']\nX = sm.add_constant(X)\n\n# Build linear regression model\nmodel = sm.OLS(y,X).fit()\n\n# Print the model summary\nmodel.summary()",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
]
] |
4acf5b3ce14d68335ebcf83ac9c97d549449c45c
| 57,376 |
ipynb
|
Jupyter Notebook
|
.ipynb_checkpoints/bvp_solver-checkpoint.ipynb
|
rlabbe/numpy-cookbook
|
d62a1b33004ea6f93c0517221404578a0fe917c1
|
[
"MIT"
] | 8 |
2015-11-22T01:18:49.000Z
|
2020-01-11T14:58:42.000Z
|
.ipynb_checkpoints/bvp_solver-checkpoint.ipynb
|
rlabbe/numpy-cookbook
|
d62a1b33004ea6f93c0517221404578a0fe917c1
|
[
"MIT"
] | null | null | null |
.ipynb_checkpoints/bvp_solver-checkpoint.ipynb
|
rlabbe/numpy-cookbook
|
d62a1b33004ea6f93c0517221404578a0fe917c1
|
[
"MIT"
] | 9 |
2015-12-31T09:41:06.000Z
|
2022-01-26T16:30:23.000Z
| 182.726115 | 33,049 | 0.864734 |
[
[
[
"empty"
]
]
] |
[
"empty"
] |
[
[
"empty"
]
] |
4acf6024a83c4353bfb3d46ae2b442764122c353
| 1,447 |
ipynb
|
Jupyter Notebook
|
09.Common roadblocks when Web Scraping/RequestHeaders.ipynb
|
younus8imran/web-scraping
|
2f7df1b404c7838cd62a57e6dabd52de38c61f71
|
[
"MIT"
] | 1 |
2021-09-15T18:15:44.000Z
|
2021-09-15T18:15:44.000Z
|
09.Common roadblocks when Web Scraping/RequestHeaders.ipynb
|
younus8imran/web-scraping
|
2f7df1b404c7838cd62a57e6dabd52de38c61f71
|
[
"MIT"
] | null | null | null |
09.Common roadblocks when Web Scraping/RequestHeaders.ipynb
|
younus8imran/web-scraping
|
2f7df1b404c7838cd62a57e6dabd52de38c61f71
|
[
"MIT"
] | 3 |
2021-09-15T02:19:36.000Z
|
2022-02-04T17:23:15.000Z
| 18.0875 | 153 | 0.503801 |
[
[
[
"import requests",
"_____no_output_____"
],
[
"headers = {\"User-Agent\": \"Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/74.0.3729.169 Safari/537.36\"}",
"_____no_output_____"
],
[
"r = requests.get('https://www.youtube.com', headers = headers)",
"_____no_output_____"
],
[
"r.status_code",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code"
]
] |
4acf62deae1667de8e1b101f49a67edfdbcf1430
| 246,192 |
ipynb
|
Jupyter Notebook
|
src/scratchpad.ipynb
|
blu3r4y/ccc-linz-nov2018
|
67696a85417d01d8eb8a2fd22fc2f70c3e11a7f8
|
[
"MIT"
] | null | null | null |
src/scratchpad.ipynb
|
blu3r4y/ccc-linz-nov2018
|
67696a85417d01d8eb8a2fd22fc2f70c3e11a7f8
|
[
"MIT"
] | null | null | null |
src/scratchpad.ipynb
|
blu3r4y/ccc-linz-nov2018
|
67696a85417d01d8eb8a2fd22fc2f70c3e11a7f8
|
[
"MIT"
] | null | null | null | 23.572578 | 8,824 | 0.360251 |
[
[
[
"from scipy.ndimage.measurements import label\n\nimport numpy as np\nimport json",
"_____no_output_____"
],
[
"with open(r\"C:\\data\\Dropbox\\Projekte\\Code\\CCC_Linz18Fall\\data\\level5\\level5_2.json\", \"r\") as f:\n input = json.load(f)",
"_____no_output_____"
],
[
"grid = np.array(input[\"rows\"])\nplt.figure(figsize=(10, 10))\nplt.imshow(grid)",
"_____no_output_____"
],
[
"plt.figure(figsize=(10, 10))\n#plt.imshow(grid[grid == 24])\nplt.imshow(np.where(grid == 24, grid, 0))",
"_____no_output_____"
],
[
"mask, ncomponents = label(grid)\nprint(ncomponents)\nplt.figure(figsize=(10, 10))\nplt.imshow(mask)\n\nassert grid[0, 0] == 0",
"37\n"
],
[
"def get_building(grid, mask, building_index):\n r1, c1 = None, None\n r2, c2 = None, None\n for i, row in enumerate(mask):\n if any(row == building_index):\n fr = i\n fc_start = np.argmax(row == building_index)\n fc_end = len(row) - 1 - np.argmax(row[::-1] == building_index)\n \n # set upper left corner point (first match)\n if not r1 and not c1:\n r1, c1 = fr, fc_start\n \n # lower right corner point (last match)\n r2, c2 = fr, fc_end\n \n return r1, c1, r2, c2\n\ndef is_hotspot(size, r1, c1, r2, c2):\n return (r2 - r1) + 1 >= size \\\n and (c2 - c1) + 1 >= size\n\ndef get_center_point(r1, c1, r2, c2):\n rx = r1 + (r2 - r1) // 2\n cx = c1 + (c2 - c1) // 2\n return rx, cx",
"_____no_output_____"
],
[
"big_mask = np.zeros_like(mask)\niii = 1\n\ndef get_hotspots(grid, mask, building, ncomponent, size):\n r1, c1, r2, c2 = building\n hotspots_grid = np.zeros_like(mask)\n \n def _does_fit(row_, col_):\n # extract possible hotspot \n submatrix = mask[row_:row_ + size, col_:col_ + size]\n if submatrix.shape[0] != size or submatrix.shape[1] != size:\n return False\n # check if all cells are on the building\n return np.all(submatrix == ncomponent)\n \n for row in range(r1, r2 + 1):\n for col in range(c1, c2 + 1):\n if _does_fit(row, col):\n hotspots_grid[row:row + size, col:col + size] = 1\n big_mask[row:row + size, col:col + size] = iii\n \n hotspots_mask, nhotspots = label(hotspots_grid)\n \n #if np.any(hotspots_mask):\n # plt.figure(figsize=(10,10))\n # plt.imshow(hotspots_mask)\n # plt.show()\n \n # use the building algorithm again ...\n hotspots = []\n for nhotspots in range(1, nhotspots + 1):\n hotspot = get_building(hotspots_grid, hotspots_mask, nhotspots)\n hotspots.append(hotspot)\n \n # get center points of hotspots\n hotspots = [get_center_point(*a) for a in hotspots]\n \n # hotspot center must be in on the building\n hotspots = [e for e in hotspots if hotspots_grid[e[0], e[1]] == 1]\n \n return hotspots\n\nbuildings = []\n\n\nheights = sorted(np.unique(grid))\nfor height in heights[1:]:\n grid_on_height = np.where(grid == height, grid, 0)\n\n mask_on_height, ncomponents = label(grid_on_height)\n #plt.figure(figsize=(10,10))\n #plt.imshow(mask_on_height)\n #plt.show()\n\n # is the floor in the upper left corner?\n assert grid_on_height[0, 0] == 0\n\n for ncomponent in range(1, ncomponents + 1):\n building = get_building(grid_on_height, mask_on_height, ncomponent)\n iii += 1\n hotspots = get_hotspots(grid_on_height, mask_on_height, building, ncomponent, input[\"s\"])\n buildings.extend(hotspots)\n\n# sort by row and by col\nbuildings = sorted(buildings, key=lambda x: (x[0], x[1]))\n\n# prepend id and only output upper left corner\nbuildings = [(i, *a) for i, a in enumerate(buildings)]\n\nprint(buildings)\n# [' '.join([' '.join(f) for f in e]) for e in buildings]\nresult = ' '.join([' '.join(map(str, e)) for e in buildings])\nprint(result)\n\nplt.figure(figsize=(10,10))\nplt.imshow(big_mask)",
"[(0, 4, 12), (1, 4, 15), (2, 4, 38), (3, 5, 33), (4, 6, 15), (5, 8, 28), (6, 9, 25), (7, 10, 22), (8, 12, 24), (9, 15, 20), (10, 15, 40), (11, 15, 43), (12, 15, 47), (13, 17, 36), (14, 18, 2), (15, 18, 11), (16, 18, 42), (17, 19, 19), (18, 21, 33), (19, 22, 31), (20, 23, 36), (21, 24, 10), (22, 25, 6), (23, 25, 28), (24, 25, 32), (25, 26, 20), (26, 26, 31), (27, 26, 39), (28, 27, 5), (29, 27, 18), (30, 28, 29), (31, 28, 36), (32, 30, 17), (33, 30, 35), (34, 31, 24), (35, 31, 41), (36, 32, 14), (37, 32, 37), (38, 33, 18), (39, 35, 1), (40, 37, 5), (41, 37, 25), (42, 37, 28), (43, 39, 7), (44, 40, 4), (45, 40, 30), (46, 40, 38), (47, 41, 18), (48, 41, 22), (49, 43, 10), (50, 43, 32), (51, 43, 36), (52, 44, 6), (53, 44, 8), (54, 44, 37), (55, 45, 17), (56, 45, 23), (57, 45, 28), (58, 45, 34), (59, 46, 15), (60, 46, 42), (61, 47, 18), (62, 48, 15), (63, 49, 30), (64, 49, 41)]\n0 4 12 1 4 15 2 4 38 3 5 33 4 6 15 5 8 28 6 9 25 7 10 22 8 12 24 9 15 20 10 15 40 11 15 43 12 15 47 13 17 36 14 18 2 15 18 11 16 18 42 17 19 19 18 21 33 19 22 31 20 23 36 21 24 10 22 25 6 23 25 28 24 25 32 25 26 20 26 26 31 27 26 39 28 27 5 29 27 18 30 28 29 31 28 36 32 30 17 33 30 35 34 31 24 35 31 41 36 32 14 37 32 37 38 33 18 39 35 1 40 37 5 41 37 25 42 37 28 43 39 7 44 40 4 45 40 30 46 40 38 47 41 18 48 41 22 49 43 10 50 43 32 51 43 36 52 44 6 53 44 8 54 44 37 55 45 17 56 45 23 57 45 28 58 45 34 59 46 15 60 46 42 61 47 18 62 48 15 63 49 30 64 49 41\n"
],
[
"for row in mask:\n if any(row == 2):\n print(1)",
"1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n1\n"
],
[
"(6 - 2 + 1) // 2",
"_____no_output_____"
],
[
"plt.imshow(grid)\nnp.unique(labeled)",
"_____no_output_____"
],
[
"structure = np.ones((3, 3), dtype=np.int)\n\nfor x in range(grid.shape[0]):\n for y in range(grid.shape[1]):\n #print((x, y))",
"(0, 0)\n(0, 1)\n(0, 2)\n(0, 3)\n(0, 4)\n(0, 5)\n(0, 6)\n(0, 7)\n(0, 8)\n(0, 9)\n(0, 10)\n(0, 11)\n(0, 12)\n(0, 13)\n(0, 14)\n(0, 15)\n(0, 16)\n(0, 17)\n(0, 18)\n(0, 19)\n(0, 20)\n(0, 21)\n(0, 22)\n(0, 23)\n(0, 24)\n(0, 25)\n(0, 26)\n(0, 27)\n(0, 28)\n(0, 29)\n(0, 30)\n(0, 31)\n(0, 32)\n(0, 33)\n(0, 34)\n(0, 35)\n(0, 36)\n(0, 37)\n(0, 38)\n(0, 39)\n(0, 40)\n(0, 41)\n(0, 42)\n(0, 43)\n(0, 44)\n(0, 45)\n(0, 46)\n(0, 47)\n(0, 48)\n(0, 49)\n(0, 50)\n(0, 51)\n(0, 52)\n(0, 53)\n(0, 54)\n(0, 55)\n(0, 56)\n(0, 57)\n(0, 58)\n(0, 59)\n(0, 60)\n(0, 61)\n(0, 62)\n(0, 63)\n(0, 64)\n(0, 65)\n(0, 66)\n(0, 67)\n(0, 68)\n(0, 69)\n(0, 70)\n(0, 71)\n(0, 72)\n(0, 73)\n(0, 74)\n(0, 75)\n(0, 76)\n(0, 77)\n(0, 78)\n(0, 79)\n(0, 80)\n(0, 81)\n(0, 82)\n(0, 83)\n(0, 84)\n(0, 85)\n(0, 86)\n(0, 87)\n(0, 88)\n(0, 89)\n(0, 90)\n(0, 91)\n(0, 92)\n(0, 93)\n(0, 94)\n(0, 95)\n(0, 96)\n(0, 97)\n(0, 98)\n(0, 99)\n(1, 0)\n(1, 1)\n(1, 2)\n(1, 3)\n(1, 4)\n(1, 5)\n(1, 6)\n(1, 7)\n(1, 8)\n(1, 9)\n(1, 10)\n(1, 11)\n(1, 12)\n(1, 13)\n(1, 14)\n(1, 15)\n(1, 16)\n(1, 17)\n(1, 18)\n(1, 19)\n(1, 20)\n(1, 21)\n(1, 22)\n(1, 23)\n(1, 24)\n(1, 25)\n(1, 26)\n(1, 27)\n(1, 28)\n(1, 29)\n(1, 30)\n(1, 31)\n(1, 32)\n(1, 33)\n(1, 34)\n(1, 35)\n(1, 36)\n(1, 37)\n(1, 38)\n(1, 39)\n(1, 40)\n(1, 41)\n(1, 42)\n(1, 43)\n(1, 44)\n(1, 45)\n(1, 46)\n(1, 47)\n(1, 48)\n(1, 49)\n(1, 50)\n(1, 51)\n(1, 52)\n(1, 53)\n(1, 54)\n(1, 55)\n(1, 56)\n(1, 57)\n(1, 58)\n(1, 59)\n(1, 60)\n(1, 61)\n(1, 62)\n(1, 63)\n(1, 64)\n(1, 65)\n(1, 66)\n(1, 67)\n(1, 68)\n(1, 69)\n(1, 70)\n(1, 71)\n(1, 72)\n(1, 73)\n(1, 74)\n(1, 75)\n(1, 76)\n(1, 77)\n(1, 78)\n(1, 79)\n(1, 80)\n(1, 81)\n(1, 82)\n(1, 83)\n(1, 84)\n(1, 85)\n(1, 86)\n(1, 87)\n(1, 88)\n(1, 89)\n(1, 90)\n(1, 91)\n(1, 92)\n(1, 93)\n(1, 94)\n(1, 95)\n(1, 96)\n(1, 97)\n(1, 98)\n(1, 99)\n(2, 0)\n(2, 1)\n(2, 2)\n(2, 3)\n(2, 4)\n(2, 5)\n(2, 6)\n(2, 7)\n(2, 8)\n(2, 9)\n(2, 10)\n(2, 11)\n(2, 12)\n(2, 13)\n(2, 14)\n(2, 15)\n(2, 16)\n(2, 17)\n(2, 18)\n(2, 19)\n(2, 20)\n(2, 21)\n(2, 22)\n(2, 23)\n(2, 24)\n(2, 25)\n(2, 26)\n(2, 27)\n(2, 28)\n(2, 29)\n(2, 30)\n(2, 31)\n(2, 32)\n(2, 33)\n(2, 34)\n(2, 35)\n(2, 36)\n(2, 37)\n(2, 38)\n(2, 39)\n(2, 40)\n(2, 41)\n(2, 42)\n(2, 43)\n(2, 44)\n(2, 45)\n(2, 46)\n(2, 47)\n(2, 48)\n(2, 49)\n(2, 50)\n(2, 51)\n(2, 52)\n(2, 53)\n(2, 54)\n(2, 55)\n(2, 56)\n(2, 57)\n(2, 58)\n(2, 59)\n(2, 60)\n(2, 61)\n(2, 62)\n(2, 63)\n(2, 64)\n(2, 65)\n(2, 66)\n(2, 67)\n(2, 68)\n(2, 69)\n(2, 70)\n(2, 71)\n(2, 72)\n(2, 73)\n(2, 74)\n(2, 75)\n(2, 76)\n(2, 77)\n(2, 78)\n(2, 79)\n(2, 80)\n(2, 81)\n(2, 82)\n(2, 83)\n(2, 84)\n(2, 85)\n(2, 86)\n(2, 87)\n(2, 88)\n(2, 89)\n(2, 90)\n(2, 91)\n(2, 92)\n(2, 93)\n(2, 94)\n(2, 95)\n(2, 96)\n(2, 97)\n(2, 98)\n(2, 99)\n(3, 0)\n(3, 1)\n(3, 2)\n(3, 3)\n(3, 4)\n(3, 5)\n(3, 6)\n(3, 7)\n(3, 8)\n(3, 9)\n(3, 10)\n(3, 11)\n(3, 12)\n(3, 13)\n(3, 14)\n(3, 15)\n(3, 16)\n(3, 17)\n(3, 18)\n(3, 19)\n(3, 20)\n(3, 21)\n(3, 22)\n(3, 23)\n(3, 24)\n(3, 25)\n(3, 26)\n(3, 27)\n(3, 28)\n(3, 29)\n(3, 30)\n(3, 31)\n(3, 32)\n(3, 33)\n(3, 34)\n(3, 35)\n(3, 36)\n(3, 37)\n(3, 38)\n(3, 39)\n(3, 40)\n(3, 41)\n(3, 42)\n(3, 43)\n(3, 44)\n(3, 45)\n(3, 46)\n(3, 47)\n(3, 48)\n(3, 49)\n(3, 50)\n(3, 51)\n(3, 52)\n(3, 53)\n(3, 54)\n(3, 55)\n(3, 56)\n(3, 57)\n(3, 58)\n(3, 59)\n(3, 60)\n(3, 61)\n(3, 62)\n(3, 63)\n(3, 64)\n(3, 65)\n(3, 66)\n(3, 67)\n(3, 68)\n(3, 69)\n(3, 70)\n(3, 71)\n(3, 72)\n(3, 73)\n(3, 74)\n(3, 75)\n(3, 76)\n(3, 77)\n(3, 78)\n(3, 79)\n(3, 80)\n(3, 81)\n(3, 82)\n(3, 83)\n(3, 84)\n(3, 85)\n(3, 86)\n(3, 87)\n(3, 88)\n(3, 89)\n(3, 90)\n(3, 91)\n(3, 92)\n(3, 93)\n(3, 94)\n(3, 95)\n(3, 96)\n(3, 97)\n(3, 98)\n(3, 99)\n(4, 0)\n(4, 1)\n(4, 2)\n(4, 3)\n(4, 4)\n(4, 5)\n(4, 6)\n(4, 7)\n(4, 8)\n(4, 9)\n(4, 10)\n(4, 11)\n(4, 12)\n(4, 13)\n(4, 14)\n(4, 15)\n(4, 16)\n(4, 17)\n(4, 18)\n(4, 19)\n(4, 20)\n(4, 21)\n(4, 22)\n(4, 23)\n(4, 24)\n(4, 25)\n(4, 26)\n(4, 27)\n(4, 28)\n(4, 29)\n(4, 30)\n(4, 31)\n(4, 32)\n(4, 33)\n(4, 34)\n(4, 35)\n(4, 36)\n(4, 37)\n(4, 38)\n(4, 39)\n(4, 40)\n(4, 41)\n(4, 42)\n(4, 43)\n(4, 44)\n(4, 45)\n(4, 46)\n(4, 47)\n(4, 48)\n(4, 49)\n(4, 50)\n(4, 51)\n(4, 52)\n(4, 53)\n(4, 54)\n(4, 55)\n(4, 56)\n(4, 57)\n(4, 58)\n(4, 59)\n(4, 60)\n(4, 61)\n(4, 62)\n(4, 63)\n(4, 64)\n(4, 65)\n(4, 66)\n(4, 67)\n(4, 68)\n(4, 69)\n(4, 70)\n(4, 71)\n(4, 72)\n(4, 73)\n(4, 74)\n(4, 75)\n(4, 76)\n(4, 77)\n(4, 78)\n(4, 79)\n(4, 80)\n(4, 81)\n(4, 82)\n(4, 83)\n(4, 84)\n(4, 85)\n(4, 86)\n(4, 87)\n(4, 88)\n(4, 89)\n(4, 90)\n(4, 91)\n(4, 92)\n(4, 93)\n(4, 94)\n(4, 95)\n(4, 96)\n(4, 97)\n(4, 98)\n(4, 99)\n(5, 0)\n(5, 1)\n(5, 2)\n(5, 3)\n(5, 4)\n(5, 5)\n(5, 6)\n(5, 7)\n(5, 8)\n(5, 9)\n(5, 10)\n(5, 11)\n(5, 12)\n(5, 13)\n(5, 14)\n(5, 15)\n(5, 16)\n(5, 17)\n(5, 18)\n(5, 19)\n(5, 20)\n(5, 21)\n(5, 22)\n(5, 23)\n(5, 24)\n(5, 25)\n(5, 26)\n(5, 27)\n(5, 28)\n(5, 29)\n(5, 30)\n(5, 31)\n(5, 32)\n(5, 33)\n(5, 34)\n(5, 35)\n(5, 36)\n(5, 37)\n(5, 38)\n(5, 39)\n(5, 40)\n(5, 41)\n(5, 42)\n(5, 43)\n(5, 44)\n(5, 45)\n(5, 46)\n(5, 47)\n(5, 48)\n(5, 49)\n(5, 50)\n(5, 51)\n(5, 52)\n(5, 53)\n(5, 54)\n(5, 55)\n(5, 56)\n(5, 57)\n(5, 58)\n(5, 59)\n(5, 60)\n(5, 61)\n(5, 62)\n(5, 63)\n(5, 64)\n(5, 65)\n(5, 66)\n(5, 67)\n(5, 68)\n(5, 69)\n(5, 70)\n(5, 71)\n(5, 72)\n(5, 73)\n(5, 74)\n(5, 75)\n(5, 76)\n(5, 77)\n(5, 78)\n(5, 79)\n(5, 80)\n(5, 81)\n(5, 82)\n(5, 83)\n(5, 84)\n(5, 85)\n(5, 86)\n(5, 87)\n(5, 88)\n(5, 89)\n(5, 90)\n(5, 91)\n(5, 92)\n(5, 93)\n(5, 94)\n(5, 95)\n(5, 96)\n(5, 97)\n(5, 98)\n(5, 99)\n(6, 0)\n(6, 1)\n(6, 2)\n(6, 3)\n(6, 4)\n(6, 5)\n(6, 6)\n(6, 7)\n(6, 8)\n(6, 9)\n(6, 10)\n(6, 11)\n(6, 12)\n(6, 13)\n(6, 14)\n(6, 15)\n(6, 16)\n(6, 17)\n(6, 18)\n(6, 19)\n(6, 20)\n(6, 21)\n(6, 22)\n(6, 23)\n(6, 24)\n(6, 25)\n(6, 26)\n(6, 27)\n(6, 28)\n(6, 29)\n(6, 30)\n(6, 31)\n(6, 32)\n(6, 33)\n(6, 34)\n(6, 35)\n(6, 36)\n(6, 37)\n(6, 38)\n(6, 39)\n(6, 40)\n(6, 41)\n(6, 42)\n(6, 43)\n(6, 44)\n(6, 45)\n(6, 46)\n(6, 47)\n(6, 48)\n(6, 49)\n(6, 50)\n(6, 51)\n(6, 52)\n(6, 53)\n(6, 54)\n(6, 55)\n(6, 56)\n(6, 57)\n(6, 58)\n(6, 59)\n(6, 60)\n(6, 61)\n(6, 62)\n(6, 63)\n(6, 64)\n(6, 65)\n(6, 66)\n(6, 67)\n(6, 68)\n(6, 69)\n(6, 70)\n(6, 71)\n(6, 72)\n(6, 73)\n(6, 74)\n(6, 75)\n(6, 76)\n(6, 77)\n(6, 78)\n(6, 79)\n(6, 80)\n(6, 81)\n(6, 82)\n(6, 83)\n(6, 84)\n(6, 85)\n(6, 86)\n(6, 87)\n(6, 88)\n(6, 89)\n(6, 90)\n(6, 91)\n(6, 92)\n(6, 93)\n(6, 94)\n(6, 95)\n(6, 96)\n(6, 97)\n(6, 98)\n(6, 99)\n(7, 0)\n(7, 1)\n(7, 2)\n(7, 3)\n(7, 4)\n(7, 5)\n(7, 6)\n(7, 7)\n(7, 8)\n(7, 9)\n(7, 10)\n(7, 11)\n(7, 12)\n(7, 13)\n(7, 14)\n(7, 15)\n(7, 16)\n(7, 17)\n(7, 18)\n(7, 19)\n(7, 20)\n(7, 21)\n(7, 22)\n(7, 23)\n(7, 24)\n(7, 25)\n(7, 26)\n(7, 27)\n(7, 28)\n(7, 29)\n(7, 30)\n(7, 31)\n(7, 32)\n(7, 33)\n(7, 34)\n(7, 35)\n(7, 36)\n(7, 37)\n(7, 38)\n(7, 39)\n(7, 40)\n(7, 41)\n(7, 42)\n(7, 43)\n(7, 44)\n(7, 45)\n(7, 46)\n(7, 47)\n(7, 48)\n(7, 49)\n(7, 50)\n(7, 51)\n(7, 52)\n(7, 53)\n(7, 54)\n(7, 55)\n(7, 56)\n(7, 57)\n(7, 58)\n(7, 59)\n(7, 60)\n(7, 61)\n(7, 62)\n(7, 63)\n(7, 64)\n(7, 65)\n(7, 66)\n(7, 67)\n(7, 68)\n(7, 69)\n(7, 70)\n(7, 71)\n(7, 72)\n(7, 73)\n(7, 74)\n(7, 75)\n(7, 76)\n(7, 77)\n(7, 78)\n(7, 79)\n(7, 80)\n(7, 81)\n(7, 82)\n(7, 83)\n(7, 84)\n(7, 85)\n(7, 86)\n(7, 87)\n(7, 88)\n(7, 89)\n(7, 90)\n(7, 91)\n(7, 92)\n(7, 93)\n(7, 94)\n(7, 95)\n(7, 96)\n(7, 97)\n(7, 98)\n(7, 99)\n(8, 0)\n(8, 1)\n(8, 2)\n(8, 3)\n(8, 4)\n(8, 5)\n(8, 6)\n(8, 7)\n(8, 8)\n(8, 9)\n(8, 10)\n(8, 11)\n(8, 12)\n(8, 13)\n(8, 14)\n(8, 15)\n(8, 16)\n(8, 17)\n(8, 18)\n(8, 19)\n(8, 20)\n(8, 21)\n(8, 22)\n(8, 23)\n(8, 24)\n(8, 25)\n(8, 26)\n(8, 27)\n(8, 28)\n(8, 29)\n(8, 30)\n(8, 31)\n(8, 32)\n(8, 33)\n(8, 34)\n(8, 35)\n(8, 36)\n(8, 37)\n(8, 38)\n(8, 39)\n(8, 40)\n(8, 41)\n(8, 42)\n(8, 43)\n(8, 44)\n(8, 45)\n(8, 46)\n(8, 47)\n(8, 48)\n(8, 49)\n(8, 50)\n(8, 51)\n(8, 52)\n(8, 53)\n(8, 54)\n(8, 55)\n(8, 56)\n(8, 57)\n(8, 58)\n(8, 59)\n(8, 60)\n(8, 61)\n(8, 62)\n(8, 63)\n(8, 64)\n(8, 65)\n(8, 66)\n(8, 67)\n(8, 68)\n(8, 69)\n(8, 70)\n(8, 71)\n(8, 72)\n(8, 73)\n(8, 74)\n(8, 75)\n(8, 76)\n(8, 77)\n(8, 78)\n(8, 79)\n(8, 80)\n(8, 81)\n(8, 82)\n(8, 83)\n(8, 84)\n(8, 85)\n(8, 86)\n(8, 87)\n(8, 88)\n(8, 89)\n(8, 90)\n(8, 91)\n(8, 92)\n(8, 93)\n(8, 94)\n(8, 95)\n(8, 96)\n(8, 97)\n(8, 98)\n(8, 99)\n(9, 0)\n(9, 1)\n(9, 2)\n(9, 3)\n(9, 4)\n(9, 5)\n(9, 6)\n(9, 7)\n(9, 8)\n(9, 9)\n(9, 10)\n(9, 11)\n(9, 12)\n(9, 13)\n(9, 14)\n(9, 15)\n(9, 16)\n(9, 17)\n(9, 18)\n(9, 19)\n(9, 20)\n(9, 21)\n(9, 22)\n(9, 23)\n(9, 24)\n(9, 25)\n(9, 26)\n(9, 27)\n(9, 28)\n(9, 29)\n(9, 30)\n(9, 31)\n(9, 32)\n(9, 33)\n(9, 34)\n(9, 35)\n(9, 36)\n(9, 37)\n(9, 38)\n(9, 39)\n(9, 40)\n(9, 41)\n(9, 42)\n(9, 43)\n(9, 44)\n(9, 45)\n(9, 46)\n(9, 47)\n(9, 48)\n(9, 49)\n(9, 50)\n(9, 51)\n(9, 52)\n(9, 53)\n(9, 54)\n(9, 55)\n(9, 56)\n(9, 57)\n(9, 58)\n(9, 59)\n(9, 60)\n(9, 61)\n(9, 62)\n(9, 63)\n(9, 64)\n(9, 65)\n(9, 66)\n(9, 67)\n(9, 68)\n(9, 69)\n(9, 70)\n(9, 71)\n(9, 72)\n(9, 73)\n(9, 74)\n(9, 75)\n(9, 76)\n(9, 77)\n(9, 78)\n(9, 79)\n(9, 80)\n(9, 81)\n(9, 82)\n(9, 83)\n(9, 84)\n(9, 85)\n(9, 86)\n(9, 87)\n(9, 88)\n(9, 89)\n(9, 90)\n(9, 91)\n(9, 92)\n(9, 93)\n(9, 94)\n(9, 95)\n(9, 96)\n(9, 97)\n(9, 98)\n(9, 99)\n(10, 0)\n(10, 1)\n(10, 2)\n(10, 3)\n(10, 4)\n(10, 5)\n(10, 6)\n(10, 7)\n(10, 8)\n(10, 9)\n(10, 10)\n(10, 11)\n(10, 12)\n(10, 13)\n(10, 14)\n(10, 15)\n(10, 16)\n(10, 17)\n(10, 18)\n(10, 19)\n(10, 20)\n(10, 21)\n(10, 22)\n(10, 23)\n(10, 24)\n(10, 25)\n(10, 26)\n(10, 27)\n(10, 28)\n(10, 29)\n(10, 30)\n(10, 31)\n(10, 32)\n(10, 33)\n(10, 34)\n(10, 35)\n(10, 36)\n(10, 37)\n(10, 38)\n(10, 39)\n(10, 40)\n(10, 41)\n(10, 42)\n(10, 43)\n(10, 44)\n(10, 45)\n(10, 46)\n(10, 47)\n(10, 48)\n(10, 49)\n(10, 50)\n(10, 51)\n(10, 52)\n(10, 53)\n(10, 54)\n(10, 55)\n(10, 56)\n(10, 57)\n(10, 58)\n(10, 59)\n(10, 60)\n(10, 61)\n(10, 62)\n(10, 63)\n(10, 64)\n(10, 65)\n(10, 66)\n(10, 67)\n(10, 68)\n(10, 69)\n(10, 70)\n(10, 71)\n(10, 72)\n(10, 73)\n(10, 74)\n(10, 75)\n(10, 76)\n(10, 77)\n(10, 78)\n(10, 79)\n(10, 80)\n(10, 81)\n(10, 82)\n(10, 83)\n(10, 84)\n(10, 85)\n(10, 86)\n(10, 87)\n(10, 88)\n(10, 89)\n(10, 90)\n(10, 91)\n(10, 92)\n(10, 93)\n(10, 94)\n(10, 95)\n(10, 96)\n(10, 97)\n(10, 98)\n(10, 99)\n(11, 0)\n(11, 1)\n(11, 2)\n(11, 3)\n(11, 4)\n(11, 5)\n(11, 6)\n(11, 7)\n(11, 8)\n(11, 9)\n(11, 10)\n(11, 11)\n(11, 12)\n(11, 13)\n(11, 14)\n(11, 15)\n(11, 16)\n(11, 17)\n(11, 18)\n(11, 19)\n(11, 20)\n(11, 21)\n(11, 22)\n(11, 23)\n(11, 24)\n(11, 25)\n(11, 26)\n(11, 27)\n(11, 28)\n(11, 29)\n(11, 30)\n(11, 31)\n(11, 32)\n(11, 33)\n(11, 34)\n(11, 35)\n(11, 36)\n(11, 37)\n(11, 38)\n(11, 39)\n(11, 40)\n(11, 41)\n(11, 42)\n(11, 43)\n(11, 44)\n(11, 45)\n(11, 46)\n(11, 47)\n(11, 48)\n(11, 49)\n(11, 50)\n(11, 51)\n(11, 52)\n(11, 53)\n(11, 54)\n(11, 55)\n(11, 56)\n(11, 57)\n(11, 58)\n(11, 59)\n(11, 60)\n(11, 61)\n(11, 62)\n(11, 63)\n(11, 64)\n(11, 65)\n(11, 66)\n(11, 67)\n(11, 68)\n(11, 69)\n(11, 70)\n(11, 71)\n(11, 72)\n(11, 73)\n(11, 74)\n(11, 75)\n(11, 76)\n(11, 77)\n(11, 78)\n(11, 79)\n(11, 80)\n(11, 81)\n(11, 82)\n(11, 83)\n(11, 84)\n(11, 85)\n(11, 86)\n(11, 87)\n(11, 88)\n(11, 89)\n(11, 90)\n(11, 91)\n(11, 92)\n(11, 93)\n(11, 94)\n(11, 95)\n(11, 96)\n(11, 97)\n(11, 98)\n(11, 99)\n(12, 0)\n(12, 1)\n(12, 2)\n(12, 3)\n(12, 4)\n(12, 5)\n(12, 6)\n(12, 7)\n(12, 8)\n(12, 9)\n(12, 10)\n(12, 11)\n(12, 12)\n(12, 13)\n(12, 14)\n(12, 15)\n(12, 16)\n(12, 17)\n(12, 18)\n(12, 19)\n(12, 20)\n(12, 21)\n(12, 22)\n(12, 23)\n(12, 24)\n(12, 25)\n(12, 26)\n(12, 27)\n(12, 28)\n(12, 29)\n(12, 30)\n(12, 31)\n(12, 32)\n(12, 33)\n(12, 34)\n(12, 35)\n(12, 36)\n(12, 37)\n(12, 38)\n(12, 39)\n(12, 40)\n(12, 41)\n(12, 42)\n(12, 43)\n(12, 44)\n(12, 45)\n(12, 46)\n(12, 47)\n(12, 48)\n(12, 49)\n(12, 50)\n(12, 51)\n(12, 52)\n(12, 53)\n(12, 54)\n(12, 55)\n(12, 56)\n(12, 57)\n(12, 58)\n(12, 59)\n(12, 60)\n(12, 61)\n(12, 62)\n(12, 63)\n(12, 64)\n(12, 65)\n(12, 66)\n(12, 67)\n(12, 68)\n(12, 69)\n(12, 70)\n(12, 71)\n(12, 72)\n(12, 73)\n(12, 74)\n(12, 75)\n(12, 76)\n(12, 77)\n(12, 78)\n(12, 79)\n(12, 80)\n(12, 81)\n(12, 82)\n(12, 83)\n(12, 84)\n(12, 85)\n(12, 86)\n(12, 87)\n(12, 88)\n(12, 89)\n(12, 90)\n(12, 91)\n(12, 92)\n(12, 93)\n(12, 94)\n(12, 95)\n(12, 96)\n(12, 97)\n(12, 98)\n(12, 99)\n(13, 0)\n(13, 1)\n(13, 2)\n(13, 3)\n(13, 4)\n(13, 5)\n(13, 6)\n(13, 7)\n(13, 8)\n(13, 9)\n(13, 10)\n(13, 11)\n(13, 12)\n(13, 13)\n(13, 14)\n(13, 15)\n(13, 16)\n(13, 17)\n(13, 18)\n(13, 19)\n(13, 20)\n(13, 21)\n(13, 22)\n(13, 23)\n(13, 24)\n(13, 25)\n(13, 26)\n(13, 27)\n(13, 28)\n(13, 29)\n(13, 30)\n(13, 31)\n(13, 32)\n(13, 33)\n(13, 34)\n(13, 35)\n(13, 36)\n(13, 37)\n(13, 38)\n(13, 39)\n(13, 40)\n(13, 41)\n(13, 42)\n(13, 43)\n(13, 44)\n(13, 45)\n(13, 46)\n(13, 47)\n(13, 48)\n(13, 49)\n(13, 50)\n(13, 51)\n(13, 52)\n(13, 53)\n(13, 54)\n(13, 55)\n(13, 56)\n(13, 57)\n(13, 58)\n(13, 59)\n(13, 60)\n(13, 61)\n(13, 62)\n(13, 63)\n(13, 64)\n(13, 65)\n(13, 66)\n(13, 67)\n(13, 68)\n(13, 69)\n(13, 70)\n(13, 71)\n(13, 72)\n(13, 73)\n(13, 74)\n(13, 75)\n(13, 76)\n(13, 77)\n(13, 78)\n(13, 79)\n(13, 80)\n(13, 81)\n(13, 82)\n(13, 83)\n(13, 84)\n(13, 85)\n(13, 86)\n(13, 87)\n(13, 88)\n(13, 89)\n(13, 90)\n(13, 91)\n(13, 92)\n(13, 93)\n(13, 94)\n(13, 95)\n(13, 96)\n(13, 97)\n(13, 98)\n(13, 99)\n(14, 0)\n(14, 1)\n(14, 2)\n(14, 3)\n(14, 4)\n(14, 5)\n(14, 6)\n(14, 7)\n(14, 8)\n(14, 9)\n(14, 10)\n(14, 11)\n(14, 12)\n(14, 13)\n(14, 14)\n(14, 15)\n(14, 16)\n(14, 17)\n(14, 18)\n(14, 19)\n(14, 20)\n(14, 21)\n(14, 22)\n(14, 23)\n(14, 24)\n(14, 25)\n(14, 26)\n(14, 27)\n(14, 28)\n(14, 29)\n(14, 30)\n(14, 31)\n(14, 32)\n(14, 33)\n(14, 34)\n(14, 35)\n(14, 36)\n(14, 37)\n(14, 38)\n(14, 39)\n(14, 40)\n(14, 41)\n(14, 42)\n(14, 43)\n(14, 44)\n(14, 45)\n(14, 46)\n(14, 47)\n(14, 48)\n(14, 49)\n(14, 50)\n(14, 51)\n(14, 52)\n(14, 53)\n(14, 54)\n(14, 55)\n(14, 56)\n(14, 57)\n(14, 58)\n(14, 59)\n(14, 60)\n(14, 61)\n(14, 62)\n(14, 63)\n(14, 64)\n(14, 65)\n(14, 66)\n(14, 67)\n(14, 68)\n(14, 69)\n(14, 70)\n(14, 71)\n(14, 72)\n(14, 73)\n(14, 74)\n(14, 75)\n(14, 76)\n(14, 77)\n(14, 78)\n(14, 79)\n(14, 80)\n(14, 81)\n(14, 82)\n(14, 83)\n(14, 84)\n(14, 85)\n(14, 86)\n(14, 87)\n(14, 88)\n(14, 89)\n(14, 90)\n(14, 91)\n(14, 92)\n(14, 93)\n(14, 94)\n(14, 95)\n(14, 96)\n(14, 97)\n(14, 98)\n(14, 99)\n(15, 0)\n(15, 1)\n(15, 2)\n(15, 3)\n(15, 4)\n(15, 5)\n(15, 6)\n(15, 7)\n(15, 8)\n(15, 9)\n(15, 10)\n(15, 11)\n(15, 12)\n(15, 13)\n(15, 14)\n(15, 15)\n(15, 16)\n(15, 17)\n(15, 18)\n(15, 19)\n(15, 20)\n(15, 21)\n(15, 22)\n(15, 23)\n(15, 24)\n(15, 25)\n(15, 26)\n(15, 27)\n(15, 28)\n(15, 29)\n(15, 30)\n(15, 31)\n(15, 32)\n(15, 33)\n(15, 34)\n(15, 35)\n(15, 36)\n(15, 37)\n(15, 38)\n(15, 39)\n(15, 40)\n(15, 41)\n(15, 42)\n(15, 43)\n(15, 44)\n(15, 45)\n(15, 46)\n(15, 47)\n(15, 48)\n(15, 49)\n(15, 50)\n(15, 51)\n(15, 52)\n(15, 53)\n(15, 54)\n(15, 55)\n(15, 56)\n(15, 57)\n(15, 58)\n(15, 59)\n(15, 60)\n(15, 61)\n(15, 62)\n(15, 63)\n(15, 64)\n(15, 65)\n(15, 66)\n(15, 67)\n(15, 68)\n(15, 69)\n(15, 70)\n(15, 71)\n(15, 72)\n(15, 73)\n(15, 74)\n(15, 75)\n(15, 76)\n(15, 77)\n(15, 78)\n(15, 79)\n(15, 80)\n(15, 81)\n(15, 82)\n(15, 83)\n(15, 84)\n(15, 85)\n(15, 86)\n(15, 87)\n(15, 88)\n(15, 89)\n(15, 90)\n(15, 91)\n(15, 92)\n(15, 93)\n(15, 94)\n(15, 95)\n(15, 96)\n(15, 97)\n(15, 98)\n(15, 99)\n(16, 0)\n(16, 1)\n(16, 2)\n(16, 3)\n(16, 4)\n(16, 5)\n(16, 6)\n(16, 7)\n(16, 8)\n(16, 9)\n(16, 10)\n(16, 11)\n(16, 12)\n(16, 13)\n(16, 14)\n(16, 15)\n(16, 16)\n(16, 17)\n(16, 18)\n(16, 19)\n(16, 20)\n(16, 21)\n(16, 22)\n(16, 23)\n(16, 24)\n(16, 25)\n(16, 26)\n(16, 27)\n(16, 28)\n(16, 29)\n(16, 30)\n(16, 31)\n(16, 32)\n(16, 33)\n(16, 34)\n(16, 35)\n(16, 36)\n(16, 37)\n(16, 38)\n(16, 39)\n(16, 40)\n(16, 41)\n(16, 42)\n(16, 43)\n(16, 44)\n(16, 45)\n(16, 46)\n(16, 47)\n(16, 48)\n(16, 49)\n(16, 50)\n(16, 51)\n(16, 52)\n(16, 53)\n(16, 54)\n(16, 55)\n(16, 56)\n(16, 57)\n(16, 58)\n(16, 59)\n(16, 60)\n(16, 61)\n(16, 62)\n(16, 63)\n(16, 64)\n(16, 65)\n(16, 66)\n(16, 67)\n(16, 68)\n(16, 69)\n(16, 70)\n(16, 71)\n(16, 72)\n(16, 73)\n(16, 74)\n(16, 75)\n(16, 76)\n(16, 77)\n(16, 78)\n(16, 79)\n(16, 80)\n(16, 81)\n(16, 82)\n(16, 83)\n(16, 84)\n(16, 85)\n(16, 86)\n(16, 87)\n(16, 88)\n(16, 89)\n(16, 90)\n(16, 91)\n(16, 92)\n(16, 93)\n(16, 94)\n(16, 95)\n(16, 96)\n(16, 97)\n(16, 98)\n(16, 99)\n(17, 0)\n(17, 1)\n(17, 2)\n(17, 3)\n(17, 4)\n(17, 5)\n(17, 6)\n(17, 7)\n(17, 8)\n(17, 9)\n(17, 10)\n(17, 11)\n(17, 12)\n(17, 13)\n(17, 14)\n(17, 15)\n(17, 16)\n(17, 17)\n(17, 18)\n(17, 19)\n(17, 20)\n(17, 21)\n(17, 22)\n(17, 23)\n(17, 24)\n(17, 25)\n(17, 26)\n(17, 27)\n(17, 28)\n(17, 29)\n(17, 30)\n(17, 31)\n(17, 32)\n(17, 33)\n(17, 34)\n(17, 35)\n(17, 36)\n(17, 37)\n(17, 38)\n(17, 39)\n(17, 40)\n(17, 41)\n(17, 42)\n(17, 43)\n(17, 44)\n(17, 45)\n(17, 46)\n(17, 47)\n(17, 48)\n(17, 49)\n(17, 50)\n(17, 51)\n(17, 52)\n(17, 53)\n(17, 54)\n(17, 55)\n(17, 56)\n(17, 57)\n(17, 58)\n(17, 59)\n(17, 60)\n(17, 61)\n(17, 62)\n(17, 63)\n(17, 64)\n(17, 65)\n(17, 66)\n(17, 67)\n(17, 68)\n(17, 69)\n(17, 70)\n(17, 71)\n(17, 72)\n(17, 73)\n(17, 74)\n(17, 75)\n(17, 76)\n(17, 77)\n(17, 78)\n(17, 79)\n(17, 80)\n(17, 81)\n(17, 82)\n(17, 83)\n(17, 84)\n(17, 85)\n(17, 86)\n(17, 87)\n(17, 88)\n(17, 89)\n(17, 90)\n(17, 91)\n(17, 92)\n(17, 93)\n(17, 94)\n(17, 95)\n(17, 96)\n(17, 97)\n(17, 98)\n(17, 99)\n(18, 0)\n(18, 1)\n(18, 2)\n(18, 3)\n(18, 4)\n(18, 5)\n(18, 6)\n(18, 7)\n(18, 8)\n(18, 9)\n(18, 10)\n(18, 11)\n(18, 12)\n(18, 13)\n(18, 14)\n(18, 15)\n(18, 16)\n(18, 17)\n(18, 18)\n(18, 19)\n(18, 20)\n(18, 21)\n(18, 22)\n(18, 23)\n(18, 24)\n(18, 25)\n(18, 26)\n(18, 27)\n(18, 28)\n(18, 29)\n(18, 30)\n(18, 31)\n(18, 32)\n(18, 33)\n(18, 34)\n(18, 35)\n(18, 36)\n(18, 37)\n(18, 38)\n(18, 39)\n(18, 40)\n(18, 41)\n(18, 42)\n(18, 43)\n(18, 44)\n(18, 45)\n(18, 46)\n(18, 47)\n(18, 48)\n(18, 49)\n(18, 50)\n(18, 51)\n(18, 52)\n(18, 53)\n(18, 54)\n(18, 55)\n(18, 56)\n(18, 57)\n(18, 58)\n(18, 59)\n(18, 60)\n(18, 61)\n(18, 62)\n(18, 63)\n(18, 64)\n(18, 65)\n(18, 66)\n(18, 67)\n(18, 68)\n(18, 69)\n(18, 70)\n(18, 71)\n(18, 72)\n(18, 73)\n(18, 74)\n(18, 75)\n(18, 76)\n(18, 77)\n(18, 78)\n(18, 79)\n(18, 80)\n(18, 81)\n(18, 82)\n(18, 83)\n(18, 84)\n(18, 85)\n(18, 86)\n(18, 87)\n(18, 88)\n(18, 89)\n(18, 90)\n(18, 91)\n(18, 92)\n(18, 93)\n(18, 94)\n(18, 95)\n(18, 96)\n(18, 97)\n(18, 98)\n(18, 99)\n(19, 0)\n(19, 1)\n(19, 2)\n(19, 3)\n(19, 4)\n(19, 5)\n(19, 6)\n(19, 7)\n(19, 8)\n(19, 9)\n(19, 10)\n(19, 11)\n(19, 12)\n(19, 13)\n(19, 14)\n(19, 15)\n(19, 16)\n(19, 17)\n(19, 18)\n(19, 19)\n(19, 20)\n(19, 21)\n(19, 22)\n(19, 23)\n(19, 24)\n(19, 25)\n(19, 26)\n(19, 27)\n(19, 28)\n(19, 29)\n(19, 30)\n(19, 31)\n(19, 32)\n(19, 33)\n(19, 34)\n(19, 35)\n(19, 36)\n(19, 37)\n(19, 38)\n(19, 39)\n(19, 40)\n(19, 41)\n(19, 42)\n(19, 43)\n(19, 44)\n(19, 45)\n(19, 46)\n(19, 47)\n(19, 48)\n(19, 49)\n(19, 50)\n(19, 51)\n(19, 52)\n(19, 53)\n(19, 54)\n(19, 55)\n(19, 56)\n(19, 57)\n(19, 58)\n(19, 59)\n(19, 60)\n(19, 61)\n(19, 62)\n(19, 63)\n(19, 64)\n(19, 65)\n(19, 66)\n(19, 67)\n(19, 68)\n(19, 69)\n(19, 70)\n(19, 71)\n(19, 72)\n(19, 73)\n(19, 74)\n(19, 75)\n(19, 76)\n(19, 77)\n(19, 78)\n(19, 79)\n(19, 80)\n(19, 81)\n(19, 82)\n(19, 83)\n(19, 84)\n(19, 85)\n(19, 86)\n(19, 87)\n(19, 88)\n(19, 89)\n(19, 90)\n(19, 91)\n(19, 92)\n(19, 93)\n(19, 94)\n(19, 95)\n(19, 96)\n(19, 97)\n(19, 98)\n(19, 99)\n(20, 0)\n(20, 1)\n(20, 2)\n(20, 3)\n(20, 4)\n(20, 5)\n(20, 6)\n(20, 7)\n(20, 8)\n(20, 9)\n(20, 10)\n(20, 11)\n(20, 12)\n(20, 13)\n(20, 14)\n(20, 15)\n(20, 16)\n(20, 17)\n(20, 18)\n(20, 19)\n(20, 20)\n(20, 21)\n(20, 22)\n(20, 23)\n(20, 24)\n(20, 25)\n(20, 26)\n(20, 27)\n(20, 28)\n(20, 29)\n(20, 30)\n(20, 31)\n(20, 32)\n(20, 33)\n(20, 34)\n(20, 35)\n(20, 36)\n(20, 37)\n(20, 38)\n(20, 39)\n(20, 40)\n(20, 41)\n(20, 42)\n(20, 43)\n(20, 44)\n(20, 45)\n(20, 46)\n(20, 47)\n(20, 48)\n(20, 49)\n(20, 50)\n(20, 51)\n(20, 52)\n(20, 53)\n(20, 54)\n(20, 55)\n(20, 56)\n(20, 57)\n(20, 58)\n(20, 59)\n(20, 60)\n(20, 61)\n(20, 62)\n(20, 63)\n(20, 64)\n(20, 65)\n(20, 66)\n(20, 67)\n(20, 68)\n(20, 69)\n(20, 70)\n(20, 71)\n(20, 72)\n(20, 73)\n(20, 74)\n(20, 75)\n(20, 76)\n(20, 77)\n(20, 78)\n(20, 79)\n(20, 80)\n(20, 81)\n(20, 82)\n(20, 83)\n(20, 84)\n(20, 85)\n(20, 86)\n(20, 87)\n(20, 88)\n(20, 89)\n(20, 90)\n(20, 91)\n(20, 92)\n(20, 93)\n(20, 94)\n(20, 95)\n(20, 96)\n(20, 97)\n(20, 98)\n(20, 99)\n(21, 0)\n(21, 1)\n(21, 2)\n(21, 3)\n(21, 4)\n(21, 5)\n(21, 6)\n(21, 7)\n(21, 8)\n(21, 9)\n(21, 10)\n(21, 11)\n(21, 12)\n(21, 13)\n(21, 14)\n(21, 15)\n(21, 16)\n(21, 17)\n(21, 18)\n(21, 19)\n(21, 20)\n(21, 21)\n(21, 22)\n(21, 23)\n(21, 24)\n(21, 25)\n(21, 26)\n(21, 27)\n(21, 28)\n(21, 29)\n(21, 30)\n(21, 31)\n(21, 32)\n(21, 33)\n(21, 34)\n(21, 35)\n(21, 36)\n(21, 37)\n(21, 38)\n(21, 39)\n(21, 40)\n(21, 41)\n(21, 42)\n(21, 43)\n(21, 44)\n(21, 45)\n(21, 46)\n(21, 47)\n(21, 48)\n(21, 49)\n(21, 50)\n(21, 51)\n(21, 52)\n(21, 53)\n(21, 54)\n(21, 55)\n(21, 56)\n(21, 57)\n(21, 58)\n(21, 59)\n(21, 60)\n(21, 61)\n(21, 62)\n(21, 63)\n(21, 64)\n(21, 65)\n(21, 66)\n(21, 67)\n(21, 68)\n(21, 69)\n(21, 70)\n(21, 71)\n(21, 72)\n(21, 73)\n(21, 74)\n(21, 75)\n(21, 76)\n(21, 77)\n(21, 78)\n(21, 79)\n(21, 80)\n(21, 81)\n(21, 82)\n(21, 83)\n(21, 84)\n(21, 85)\n(21, 86)\n(21, 87)\n(21, 88)\n(21, 89)\n(21, 90)\n(21, 91)\n(21, 92)\n(21, 93)\n(21, 94)\n(21, 95)\n(21, 96)\n(21, 97)\n(21, 98)\n(21, 99)\n(22, 0)\n(22, 1)\n(22, 2)\n(22, 3)\n(22, 4)\n(22, 5)\n(22, 6)\n(22, 7)\n(22, 8)\n(22, 9)\n(22, 10)\n(22, 11)\n(22, 12)\n(22, 13)\n(22, 14)\n(22, 15)\n(22, 16)\n(22, 17)\n(22, 18)\n(22, 19)\n(22, 20)\n(22, 21)\n(22, 22)\n(22, 23)\n(22, 24)\n(22, 25)\n(22, 26)\n(22, 27)\n(22, 28)\n(22, 29)\n(22, 30)\n(22, 31)\n(22, 32)\n(22, 33)\n(22, 34)\n(22, 35)\n(22, 36)\n(22, 37)\n(22, 38)\n(22, 39)\n(22, 40)\n(22, 41)\n(22, 42)\n(22, 43)\n(22, 44)\n(22, 45)\n(22, 46)\n(22, 47)\n(22, 48)\n(22, 49)\n(22, 50)\n(22, 51)\n(22, 52)\n(22, 53)\n(22, 54)\n(22, 55)\n(22, 56)\n(22, 57)\n(22, 58)\n(22, 59)\n(22, 60)\n(22, 61)\n(22, 62)\n(22, 63)\n(22, 64)\n(22, 65)\n(22, 66)\n(22, 67)\n(22, 68)\n(22, 69)\n(22, 70)\n(22, 71)\n(22, 72)\n(22, 73)\n(22, 74)\n(22, 75)\n(22, 76)\n(22, 77)\n(22, 78)\n(22, 79)\n(22, 80)\n(22, 81)\n(22, 82)\n(22, 83)\n(22, 84)\n(22, 85)\n(22, 86)\n(22, 87)\n(22, 88)\n(22, 89)\n(22, 90)\n(22, 91)\n(22, 92)\n(22, 93)\n(22, 94)\n(22, 95)\n(22, 96)\n(22, 97)\n(22, 98)\n(22, 99)\n(23, 0)\n(23, 1)\n(23, 2)\n(23, 3)\n(23, 4)\n(23, 5)\n(23, 6)\n(23, 7)\n(23, 8)\n(23, 9)\n(23, 10)\n(23, 11)\n(23, 12)\n(23, 13)\n(23, 14)\n(23, 15)\n(23, 16)\n(23, 17)\n(23, 18)\n(23, 19)\n(23, 20)\n(23, 21)\n(23, 22)\n(23, 23)\n(23, 24)\n(23, 25)\n(23, 26)\n(23, 27)\n(23, 28)\n(23, 29)\n(23, 30)\n(23, 31)\n(23, 32)\n(23, 33)\n(23, 34)\n(23, 35)\n(23, 36)\n(23, 37)\n(23, 38)\n(23, 39)\n(23, 40)\n(23, 41)\n(23, 42)\n(23, 43)\n(23, 44)\n(23, 45)\n(23, 46)\n(23, 47)\n(23, 48)\n(23, 49)\n(23, 50)\n(23, 51)\n(23, 52)\n(23, 53)\n(23, 54)\n(23, 55)\n(23, 56)\n(23, 57)\n(23, 58)\n(23, 59)\n(23, 60)\n(23, 61)\n(23, 62)\n(23, 63)\n(23, 64)\n(23, 65)\n(23, 66)\n(23, 67)\n(23, 68)\n(23, 69)\n(23, 70)\n(23, 71)\n(23, 72)\n(23, 73)\n(23, 74)\n(23, 75)\n(23, 76)\n(23, 77)\n(23, 78)\n(23, 79)\n(23, 80)\n(23, 81)\n(23, 82)\n(23, 83)\n(23, 84)\n(23, 85)\n(23, 86)\n(23, 87)\n(23, 88)\n(23, 89)\n(23, 90)\n(23, 91)\n(23, 92)\n(23, 93)\n(23, 94)\n(23, 95)\n(23, 96)\n(23, 97)\n(23, 98)\n(23, 99)\n(24, 0)\n(24, 1)\n(24, 2)\n(24, 3)\n(24, 4)\n(24, 5)\n(24, 6)\n(24, 7)\n(24, 8)\n(24, 9)\n(24, 10)\n(24, 11)\n(24, 12)\n(24, 13)\n(24, 14)\n(24, 15)\n(24, 16)\n(24, 17)\n(24, 18)\n(24, 19)\n(24, 20)\n(24, 21)\n(24, 22)\n(24, 23)\n(24, 24)\n(24, 25)\n(24, 26)\n(24, 27)\n(24, 28)\n(24, 29)\n(24, 30)\n(24, 31)\n(24, 32)\n(24, 33)\n(24, 34)\n(24, 35)\n(24, 36)\n(24, 37)\n(24, 38)\n(24, 39)\n(24, 40)\n(24, 41)\n(24, 42)\n(24, 43)\n(24, 44)\n(24, 45)\n(24, 46)\n(24, 47)\n(24, 48)\n(24, 49)\n(24, 50)\n(24, 51)\n(24, 52)\n(24, 53)\n(24, 54)\n(24, 55)\n(24, 56)\n(24, 57)\n(24, 58)\n(24, 59)\n(24, 60)\n(24, 61)\n(24, 62)\n(24, 63)\n(24, 64)\n(24, 65)\n(24, 66)\n(24, 67)\n(24, 68)\n(24, 69)\n(24, 70)\n(24, 71)\n(24, 72)\n(24, 73)\n(24, 74)\n(24, 75)\n(24, 76)\n(24, 77)\n(24, 78)\n(24, 79)\n(24, 80)\n(24, 81)\n(24, 82)\n(24, 83)\n(24, 84)\n(24, 85)\n(24, 86)\n(24, 87)\n(24, 88)\n(24, 89)\n(24, 90)\n(24, 91)\n(24, 92)\n(24, 93)\n(24, 94)\n(24, 95)\n(24, 96)\n(24, 97)\n(24, 98)\n(24, 99)\n(25, 0)\n(25, 1)\n(25, 2)\n(25, 3)\n(25, 4)\n(25, 5)\n(25, 6)\n(25, 7)\n(25, 8)\n(25, 9)\n(25, 10)\n(25, 11)\n(25, 12)\n(25, 13)\n(25, 14)\n(25, 15)\n(25, 16)\n(25, 17)\n(25, 18)\n(25, 19)\n(25, 20)\n(25, 21)\n(25, 22)\n(25, 23)\n(25, 24)\n(25, 25)\n(25, 26)\n(25, 27)\n(25, 28)\n(25, 29)\n(25, 30)\n(25, 31)\n(25, 32)\n(25, 33)\n(25, 34)\n(25, 35)\n(25, 36)\n(25, 37)\n(25, 38)\n(25, 39)\n(25, 40)\n(25, 41)\n(25, 42)\n(25, 43)\n(25, 44)\n(25, 45)\n(25, 46)\n(25, 47)\n(25, 48)\n(25, 49)\n(25, 50)\n(25, 51)\n(25, 52)\n(25, 53)\n(25, 54)\n(25, 55)\n(25, 56)\n(25, 57)\n(25, 58)\n(25, 59)\n(25, 60)\n(25, 61)\n(25, 62)\n(25, 63)\n(25, 64)\n(25, 65)\n(25, 66)\n(25, 67)\n(25, 68)\n(25, 69)\n(25, 70)\n(25, 71)\n(25, 72)\n(25, 73)\n(25, 74)\n(25, 75)\n(25, 76)\n(25, 77)\n(25, 78)\n(25, 79)\n(25, 80)\n(25, 81)\n(25, 82)\n(25, 83)\n(25, 84)\n(25, 85)\n(25, 86)\n(25, 87)\n(25, 88)\n(25, 89)\n(25, 90)\n(25, 91)\n(25, 92)\n(25, 93)\n(25, 94)\n(25, 95)\n(25, 96)\n(25, 97)\n(25, 98)\n(25, 99)\n(26, 0)\n(26, 1)\n(26, 2)\n(26, 3)\n(26, 4)\n(26, 5)\n(26, 6)\n(26, 7)\n(26, 8)\n(26, 9)\n(26, 10)\n(26, 11)\n(26, 12)\n(26, 13)\n(26, 14)\n(26, 15)\n(26, 16)\n(26, 17)\n(26, 18)\n(26, 19)\n(26, 20)\n(26, 21)\n(26, 22)\n(26, 23)\n(26, 24)\n(26, 25)\n(26, 26)\n(26, 27)\n(26, 28)\n(26, 29)\n(26, 30)\n(26, 31)\n(26, 32)\n(26, 33)\n(26, 34)\n(26, 35)\n(26, 36)\n(26, 37)\n(26, 38)\n(26, 39)\n(26, 40)\n(26, 41)\n(26, 42)\n(26, 43)\n(26, 44)\n(26, 45)\n(26, 46)\n(26, 47)\n(26, 48)\n(26, 49)\n(26, 50)\n(26, 51)\n(26, 52)\n(26, 53)\n(26, 54)\n(26, 55)\n(26, 56)\n(26, 57)\n(26, 58)\n(26, 59)\n(26, 60)\n(26, 61)\n(26, 62)\n(26, 63)\n(26, 64)\n(26, 65)\n(26, 66)\n(26, 67)\n(26, 68)\n(26, 69)\n(26, 70)\n(26, 71)\n(26, 72)\n(26, 73)\n(26, 74)\n(26, 75)\n(26, 76)\n(26, 77)\n(26, 78)\n(26, 79)\n(26, 80)\n(26, 81)\n(26, 82)\n(26, 83)\n(26, 84)\n(26, 85)\n(26, 86)\n(26, 87)\n(26, 88)\n(26, 89)\n(26, 90)\n(26, 91)\n(26, 92)\n(26, 93)\n(26, 94)\n(26, 95)\n(26, 96)\n(26, 97)\n(26, 98)\n(26, 99)\n(27, 0)\n(27, 1)\n(27, 2)\n(27, 3)\n(27, 4)\n(27, 5)\n(27, 6)\n(27, 7)\n(27, 8)\n(27, 9)\n(27, 10)\n(27, 11)\n(27, 12)\n(27, 13)\n(27, 14)\n(27, 15)\n(27, 16)\n(27, 17)\n(27, 18)\n(27, 19)\n(27, 20)\n(27, 21)\n(27, 22)\n(27, 23)\n(27, 24)\n(27, 25)\n(27, 26)\n(27, 27)\n(27, 28)\n(27, 29)\n(27, 30)\n(27, 31)\n(27, 32)\n(27, 33)\n(27, 34)\n(27, 35)\n(27, 36)\n(27, 37)\n(27, 38)\n(27, 39)\n(27, 40)\n(27, 41)\n(27, 42)\n(27, 43)\n(27, 44)\n(27, 45)\n(27, 46)\n(27, 47)\n(27, 48)\n(27, 49)\n(27, 50)\n(27, 51)\n(27, 52)\n(27, 53)\n(27, 54)\n(27, 55)\n(27, 56)\n(27, 57)\n(27, 58)\n(27, 59)\n(27, 60)\n(27, 61)\n(27, 62)\n(27, 63)\n(27, 64)\n(27, 65)\n(27, 66)\n(27, 67)\n(27, 68)\n(27, 69)\n(27, 70)\n(27, 71)\n(27, 72)\n(27, 73)\n(27, 74)\n(27, 75)\n(27, 76)\n(27, 77)\n(27, 78)\n(27, 79)\n(27, 80)\n(27, 81)\n(27, 82)\n(27, 83)\n(27, 84)\n(27, 85)\n(27, 86)\n(27, 87)\n(27, 88)\n(27, 89)\n(27, 90)\n(27, 91)\n(27, 92)\n(27, 93)\n(27, 94)\n(27, 95)\n(27, 96)\n(27, 97)\n(27, 98)\n(27, 99)\n(28, 0)\n(28, 1)\n(28, 2)\n(28, 3)\n(28, 4)\n(28, 5)\n(28, 6)\n(28, 7)\n(28, 8)\n(28, 9)\n(28, 10)\n(28, 11)\n(28, 12)\n(28, 13)\n(28, 14)\n(28, 15)\n(28, 16)\n(28, 17)\n(28, 18)\n(28, 19)\n(28, 20)\n(28, 21)\n(28, 22)\n(28, 23)\n(28, 24)\n(28, 25)\n(28, 26)\n(28, 27)\n(28, 28)\n(28, 29)\n(28, 30)\n(28, 31)\n(28, 32)\n(28, 33)\n(28, 34)\n(28, 35)\n(28, 36)\n(28, 37)\n(28, 38)\n(28, 39)\n(28, 40)\n(28, 41)\n(28, 42)\n(28, 43)\n(28, 44)\n(28, 45)\n(28, 46)\n(28, 47)\n(28, 48)\n(28, 49)\n(28, 50)\n(28, 51)\n(28, 52)\n(28, 53)\n(28, 54)\n(28, 55)\n(28, 56)\n(28, 57)\n(28, 58)\n(28, 59)\n(28, 60)\n(28, 61)\n(28, 62)\n(28, 63)\n(28, 64)\n(28, 65)\n(28, 66)\n(28, 67)\n(28, 68)\n(28, 69)\n(28, 70)\n(28, 71)\n(28, 72)\n(28, 73)\n(28, 74)\n(28, 75)\n(28, 76)\n(28, 77)\n(28, 78)\n(28, 79)\n(28, 80)\n(28, 81)\n(28, 82)\n(28, 83)\n(28, 84)\n(28, 85)\n(28, 86)\n(28, 87)\n(28, 88)\n(28, 89)\n(28, 90)\n(28, 91)\n(28, 92)\n(28, 93)\n(28, 94)\n(28, 95)\n(28, 96)\n(28, 97)\n(28, 98)\n(28, 99)\n(29, 0)\n(29, 1)\n(29, 2)\n(29, 3)\n(29, 4)\n(29, 5)\n(29, 6)\n(29, 7)\n(29, 8)\n(29, 9)\n(29, 10)\n(29, 11)\n(29, 12)\n(29, 13)\n(29, 14)\n(29, 15)\n(29, 16)\n(29, 17)\n(29, 18)\n(29, 19)\n(29, 20)\n(29, 21)\n(29, 22)\n(29, 23)\n(29, 24)\n(29, 25)\n(29, 26)\n(29, 27)\n(29, 28)\n(29, 29)\n(29, 30)\n(29, 31)\n(29, 32)\n(29, 33)\n(29, 34)\n(29, 35)\n(29, 36)\n(29, 37)\n(29, 38)\n(29, 39)\n(29, 40)\n(29, 41)\n(29, 42)\n(29, 43)\n(29, 44)\n(29, 45)\n(29, 46)\n(29, 47)\n(29, 48)\n(29, 49)\n(29, 50)\n(29, 51)\n(29, 52)\n(29, 53)\n(29, 54)\n(29, 55)\n(29, 56)\n(29, 57)\n(29, 58)\n(29, 59)\n(29, 60)\n(29, 61)\n(29, 62)\n(29, 63)\n(29, 64)\n(29, 65)\n(29, 66)\n(29, 67)\n(29, 68)\n(29, 69)\n(29, 70)\n(29, 71)\n(29, 72)\n(29, 73)\n(29, 74)\n(29, 75)\n(29, 76)\n(29, 77)\n(29, 78)\n(29, 79)\n(29, 80)\n(29, 81)\n(29, 82)\n(29, 83)\n(29, 84)\n(29, 85)\n(29, 86)\n(29, 87)\n(29, 88)\n(29, 89)\n(29, 90)\n(29, 91)\n(29, 92)\n(29, 93)\n(29, 94)\n(29, 95)\n(29, 96)\n(29, 97)\n(29, 98)\n(29, 99)\n(30, 0)\n(30, 1)\n(30, 2)\n(30, 3)\n(30, 4)\n(30, 5)\n(30, 6)\n(30, 7)\n(30, 8)\n(30, 9)\n(30, 10)\n(30, 11)\n(30, 12)\n(30, 13)\n(30, 14)\n(30, 15)\n(30, 16)\n(30, 17)\n(30, 18)\n(30, 19)\n(30, 20)\n(30, 21)\n(30, 22)\n(30, 23)\n(30, 24)\n(30, 25)\n(30, 26)\n(30, 27)\n(30, 28)\n(30, 29)\n(30, 30)\n(30, 31)\n(30, 32)\n(30, 33)\n(30, 34)\n(30, 35)\n(30, 36)\n(30, 37)\n(30, 38)\n(30, 39)\n(30, 40)\n(30, 41)\n(30, 42)\n(30, 43)\n(30, 44)\n(30, 45)\n(30, 46)\n(30, 47)\n(30, 48)\n(30, 49)\n(30, 50)\n(30, 51)\n(30, 52)\n(30, 53)\n(30, 54)\n(30, 55)\n(30, 56)\n(30, 57)\n(30, 58)\n(30, 59)\n(30, 60)\n(30, 61)\n(30, 62)\n(30, 63)\n(30, 64)\n(30, 65)\n(30, 66)\n(30, 67)\n(30, 68)\n(30, 69)\n(30, 70)\n(30, 71)\n(30, 72)\n(30, 73)\n(30, 74)\n(30, 75)\n(30, 76)\n(30, 77)\n(30, 78)\n(30, 79)\n(30, 80)\n(30, 81)\n(30, 82)\n(30, 83)\n(30, 84)\n(30, 85)\n(30, 86)\n(30, 87)\n(30, 88)\n(30, 89)\n(30, 90)\n(30, 91)\n(30, 92)\n(30, 93)\n(30, 94)\n(30, 95)\n(30, 96)\n(30, 97)\n(30, 98)\n(30, 99)\n(31, 0)\n(31, 1)\n(31, 2)\n(31, 3)\n(31, 4)\n(31, 5)\n(31, 6)\n(31, 7)\n(31, 8)\n(31, 9)\n(31, 10)\n(31, 11)\n(31, 12)\n(31, 13)\n(31, 14)\n(31, 15)\n(31, 16)\n(31, 17)\n(31, 18)\n(31, 19)\n(31, 20)\n(31, 21)\n(31, 22)\n(31, 23)\n(31, 24)\n(31, 25)\n(31, 26)\n(31, 27)\n(31, 28)\n(31, 29)\n(31, 30)\n(31, 31)\n(31, 32)\n(31, 33)\n(31, 34)\n(31, 35)\n(31, 36)\n(31, 37)\n(31, 38)\n(31, 39)\n(31, 40)\n(31, 41)\n(31, 42)\n(31, 43)\n(31, 44)\n(31, 45)\n(31, 46)\n(31, 47)\n(31, 48)\n(31, 49)\n(31, 50)\n(31, 51)\n(31, 52)\n(31, 53)\n(31, 54)\n(31, 55)\n(31, 56)\n(31, 57)\n(31, 58)\n(31, 59)\n(31, 60)\n(31, 61)\n(31, 62)\n(31, 63)\n(31, 64)\n(31, 65)\n(31, 66)\n(31, 67)\n(31, 68)\n(31, 69)\n(31, 70)\n(31, 71)\n(31, 72)\n(31, 73)\n(31, 74)\n(31, 75)\n(31, 76)\n(31, 77)\n(31, 78)\n(31, 79)\n(31, 80)\n(31, 81)\n(31, 82)\n(31, 83)\n(31, 84)\n(31, 85)\n(31, 86)\n(31, 87)\n(31, 88)\n(31, 89)\n(31, 90)\n(31, 91)\n(31, 92)\n(31, 93)\n(31, 94)\n(31, 95)\n(31, 96)\n(31, 97)\n(31, 98)\n(31, 99)\n(32, 0)\n(32, 1)\n(32, 2)\n(32, 3)\n(32, 4)\n(32, 5)\n(32, 6)\n(32, 7)\n(32, 8)\n(32, 9)\n(32, 10)\n(32, 11)\n(32, 12)\n(32, 13)\n(32, 14)\n(32, 15)\n(32, 16)\n(32, 17)\n(32, 18)\n(32, 19)\n(32, 20)\n(32, 21)\n(32, 22)\n(32, 23)\n(32, 24)\n(32, 25)\n(32, 26)\n(32, 27)\n(32, 28)\n(32, 29)\n(32, 30)\n(32, 31)\n(32, 32)\n(32, 33)\n(32, 34)\n(32, 35)\n(32, 36)\n(32, 37)\n(32, 38)\n(32, 39)\n(32, 40)\n(32, 41)\n(32, 42)\n(32, 43)\n(32, 44)\n(32, 45)\n(32, 46)\n(32, 47)\n(32, 48)\n(32, 49)\n(32, 50)\n(32, 51)\n(32, 52)\n(32, 53)\n(32, 54)\n(32, 55)\n(32, 56)\n(32, 57)\n(32, 58)\n(32, 59)\n(32, 60)\n(32, 61)\n(32, 62)\n(32, 63)\n(32, 64)\n(32, 65)\n(32, 66)\n(32, 67)\n(32, 68)\n(32, 69)\n(32, 70)\n(32, 71)\n(32, 72)\n(32, 73)\n(32, 74)\n(32, 75)\n(32, 76)\n(32, 77)\n(32, 78)\n(32, 79)\n(32, 80)\n(32, 81)\n(32, 82)\n(32, 83)\n(32, 84)\n(32, 85)\n(32, 86)\n(32, 87)\n(32, 88)\n(32, 89)\n(32, 90)\n(32, 91)\n(32, 92)\n(32, 93)\n(32, 94)\n(32, 95)\n(32, 96)\n(32, 97)\n(32, 98)\n(32, 99)\n(33, 0)\n(33, 1)\n(33, 2)\n(33, 3)\n(33, 4)\n(33, 5)\n(33, 6)\n(33, 7)\n(33, 8)\n(33, 9)\n(33, 10)\n(33, 11)\n(33, 12)\n(33, 13)\n(33, 14)\n(33, 15)\n(33, 16)\n(33, 17)\n(33, 18)\n(33, 19)\n(33, 20)\n(33, 21)\n(33, 22)\n(33, 23)\n(33, 24)\n(33, 25)\n(33, 26)\n(33, 27)\n(33, 28)\n(33, 29)\n(33, 30)\n(33, 31)\n(33, 32)\n(33, 33)\n(33, 34)\n(33, 35)\n(33, 36)\n(33, 37)\n(33, 38)\n(33, 39)\n(33, 40)\n(33, 41)\n(33, 42)\n(33, 43)\n(33, 44)\n(33, 45)\n(33, 46)\n(33, 47)\n(33, 48)\n(33, 49)\n(33, 50)\n(33, 51)\n(33, 52)\n(33, 53)\n(33, 54)\n(33, 55)\n(33, 56)\n(33, 57)\n(33, 58)\n(33, 59)\n(33, 60)\n(33, 61)\n(33, 62)\n(33, 63)\n(33, 64)\n(33, 65)\n(33, 66)\n(33, 67)\n(33, 68)\n(33, 69)\n(33, 70)\n(33, 71)\n(33, 72)\n(33, 73)\n(33, 74)\n(33, 75)\n(33, 76)\n(33, 77)\n(33, 78)\n(33, 79)\n(33, 80)\n(33, 81)\n(33, 82)\n(33, 83)\n(33, 84)\n(33, 85)\n(33, 86)\n(33, 87)\n(33, 88)\n(33, 89)\n(33, 90)\n(33, 91)\n(33, 92)\n(33, 93)\n(33, 94)\n(33, 95)\n(33, 96)\n(33, 97)\n(33, 98)\n(33, 99)\n(34, 0)\n(34, 1)\n(34, 2)\n(34, 3)\n(34, 4)\n(34, 5)\n(34, 6)\n(34, 7)\n(34, 8)\n(34, 9)\n(34, 10)\n(34, 11)\n(34, 12)\n(34, 13)\n(34, 14)\n(34, 15)\n(34, 16)\n(34, 17)\n(34, 18)\n(34, 19)\n(34, 20)\n(34, 21)\n(34, 22)\n(34, 23)\n(34, 24)\n(34, 25)\n(34, 26)\n(34, 27)\n(34, 28)\n(34, 29)\n(34, 30)\n(34, 31)\n(34, 32)\n(34, 33)\n(34, 34)\n(34, 35)\n(34, 36)\n(34, 37)\n(34, 38)\n(34, 39)\n(34, 40)\n(34, 41)\n(34, 42)\n(34, 43)\n(34, 44)\n(34, 45)\n(34, 46)\n(34, 47)\n(34, 48)\n(34, 49)\n(34, 50)\n(34, 51)\n(34, 52)\n(34, 53)\n(34, 54)\n(34, 55)\n(34, 56)\n(34, 57)\n(34, 58)\n(34, 59)\n(34, 60)\n(34, 61)\n(34, 62)\n(34, 63)\n(34, 64)\n(34, 65)\n(34, 66)\n(34, 67)\n(34, 68)\n(34, 69)\n(34, 70)\n(34, 71)\n(34, 72)\n(34, 73)\n(34, 74)\n(34, 75)\n(34, 76)\n(34, 77)\n(34, 78)\n(34, 79)\n(34, 80)\n(34, 81)\n(34, 82)\n(34, 83)\n(34, 84)\n(34, 85)\n(34, 86)\n(34, 87)\n(34, 88)\n(34, 89)\n(34, 90)\n(34, 91)\n(34, 92)\n(34, 93)\n(34, 94)\n(34, 95)\n(34, 96)\n(34, 97)\n(34, 98)\n(34, 99)\n(35, 0)\n(35, 1)\n(35, 2)\n(35, 3)\n(35, 4)\n(35, 5)\n(35, 6)\n(35, 7)\n(35, 8)\n(35, 9)\n(35, 10)\n(35, 11)\n(35, 12)\n(35, 13)\n(35, 14)\n(35, 15)\n(35, 16)\n(35, 17)\n(35, 18)\n(35, 19)\n(35, 20)\n(35, 21)\n(35, 22)\n(35, 23)\n(35, 24)\n(35, 25)\n(35, 26)\n(35, 27)\n(35, 28)\n(35, 29)\n(35, 30)\n(35, 31)\n(35, 32)\n(35, 33)\n(35, 34)\n(35, 35)\n(35, 36)\n(35, 37)\n(35, 38)\n(35, 39)\n(35, 40)\n(35, 41)\n(35, 42)\n(35, 43)\n(35, 44)\n(35, 45)\n(35, 46)\n(35, 47)\n(35, 48)\n(35, 49)\n(35, 50)\n(35, 51)\n(35, 52)\n(35, 53)\n(35, 54)\n(35, 55)\n(35, 56)\n(35, 57)\n(35, 58)\n(35, 59)\n(35, 60)\n(35, 61)\n(35, 62)\n(35, 63)\n(35, 64)\n(35, 65)\n(35, 66)\n(35, 67)\n(35, 68)\n(35, 69)\n(35, 70)\n(35, 71)\n(35, 72)\n(35, 73)\n(35, 74)\n(35, 75)\n(35, 76)\n(35, 77)\n(35, 78)\n(35, 79)\n(35, 80)\n(35, 81)\n(35, 82)\n(35, 83)\n(35, 84)\n(35, 85)\n(35, 86)\n(35, 87)\n(35, 88)\n(35, 89)\n(35, 90)\n(35, 91)\n(35, 92)\n(35, 93)\n(35, 94)\n(35, 95)\n(35, 96)\n(35, 97)\n(35, 98)\n(35, 99)\n(36, 0)\n(36, 1)\n(36, 2)\n(36, 3)\n(36, 4)\n(36, 5)\n(36, 6)\n(36, 7)\n(36, 8)\n(36, 9)\n(36, 10)\n(36, 11)\n(36, 12)\n(36, 13)\n(36, 14)\n(36, 15)\n(36, 16)\n(36, 17)\n(36, 18)\n(36, 19)\n(36, 20)\n(36, 21)\n(36, 22)\n(36, 23)\n(36, 24)\n(36, 25)\n(36, 26)\n(36, 27)\n(36, 28)\n(36, 29)\n(36, 30)\n(36, 31)\n(36, 32)\n(36, 33)\n(36, 34)\n(36, 35)\n(36, 36)\n(36, 37)\n(36, 38)\n(36, 39)\n(36, 40)\n(36, 41)\n(36, 42)\n(36, 43)\n(36, 44)\n(36, 45)\n(36, 46)\n(36, 47)\n(36, 48)\n(36, 49)\n(36, 50)\n(36, 51)\n(36, 52)\n(36, 53)\n(36, 54)\n(36, 55)\n(36, 56)\n(36, 57)\n(36, 58)\n(36, 59)\n(36, 60)\n(36, 61)\n(36, 62)\n(36, 63)\n(36, 64)\n(36, 65)\n(36, 66)\n(36, 67)\n(36, 68)\n(36, 69)\n(36, 70)\n(36, 71)\n(36, 72)\n(36, 73)\n(36, 74)\n(36, 75)\n(36, 76)\n(36, 77)\n(36, 78)\n(36, 79)\n(36, 80)\n(36, 81)\n(36, 82)\n(36, 83)\n(36, 84)\n(36, 85)\n(36, 86)\n(36, 87)\n(36, 88)\n(36, 89)\n(36, 90)\n(36, 91)\n(36, 92)\n(36, 93)\n(36, 94)\n(36, 95)\n(36, 96)\n(36, 97)\n(36, 98)\n(36, 99)\n(37, 0)\n(37, 1)\n(37, 2)\n(37, 3)\n(37, 4)\n(37, 5)\n(37, 6)\n(37, 7)\n(37, 8)\n(37, 9)\n(37, 10)\n(37, 11)\n(37, 12)\n(37, 13)\n(37, 14)\n(37, 15)\n(37, 16)\n(37, 17)\n(37, 18)\n(37, 19)\n(37, 20)\n(37, 21)\n(37, 22)\n(37, 23)\n(37, 24)\n(37, 25)\n(37, 26)\n(37, 27)\n(37, 28)\n(37, 29)\n(37, 30)\n(37, 31)\n(37, 32)\n(37, 33)\n(37, 34)\n(37, 35)\n(37, 36)\n(37, 37)\n(37, 38)\n(37, 39)\n(37, 40)\n(37, 41)\n(37, 42)\n(37, 43)\n(37, 44)\n(37, 45)\n(37, 46)\n(37, 47)\n(37, 48)\n(37, 49)\n(37, 50)\n(37, 51)\n(37, 52)\n(37, 53)\n(37, 54)\n(37, 55)\n(37, 56)\n(37, 57)\n(37, 58)\n(37, 59)\n(37, 60)\n(37, 61)\n(37, 62)\n(37, 63)\n(37, 64)\n(37, 65)\n(37, 66)\n(37, 67)\n(37, 68)\n(37, 69)\n(37, 70)\n(37, 71)\n(37, 72)\n(37, 73)\n(37, 74)\n(37, 75)\n(37, 76)\n(37, 77)\n(37, 78)\n(37, 79)\n(37, 80)\n(37, 81)\n(37, 82)\n(37, 83)\n(37, 84)\n(37, 85)\n(37, 86)\n(37, 87)\n(37, 88)\n(37, 89)\n(37, 90)\n(37, 91)\n(37, 92)\n(37, 93)\n(37, 94)\n(37, 95)\n(37, 96)\n(37, 97)\n(37, 98)\n(37, 99)\n(38, 0)\n(38, 1)\n(38, 2)\n(38, 3)\n(38, 4)\n(38, 5)\n(38, 6)\n(38, 7)\n(38, 8)\n(38, 9)\n(38, 10)\n(38, 11)\n(38, 12)\n(38, 13)\n(38, 14)\n(38, 15)\n(38, 16)\n(38, 17)\n(38, 18)\n(38, 19)\n(38, 20)\n(38, 21)\n(38, 22)\n(38, 23)\n(38, 24)\n(38, 25)\n(38, 26)\n(38, 27)\n(38, 28)\n(38, 29)\n(38, 30)\n(38, 31)\n(38, 32)\n(38, 33)\n(38, 34)\n(38, 35)\n(38, 36)\n(38, 37)\n(38, 38)\n(38, 39)\n(38, 40)\n(38, 41)\n(38, 42)\n(38, 43)\n(38, 44)\n(38, 45)\n(38, 46)\n(38, 47)\n(38, 48)\n(38, 49)\n(38, 50)\n(38, 51)\n(38, 52)\n(38, 53)\n(38, 54)\n(38, 55)\n(38, 56)\n(38, 57)\n(38, 58)\n(38, 59)\n(38, 60)\n(38, 61)\n(38, 62)\n(38, 63)\n(38, 64)\n(38, 65)\n(38, 66)\n(38, 67)\n(38, 68)\n(38, 69)\n(38, 70)\n(38, 71)\n(38, 72)\n(38, 73)\n(38, 74)\n(38, 75)\n(38, 76)\n(38, 77)\n(38, 78)\n(38, 79)\n(38, 80)\n(38, 81)\n(38, 82)\n(38, 83)\n(38, 84)\n(38, 85)\n(38, 86)\n(38, 87)\n(38, 88)\n(38, 89)\n(38, 90)\n(38, 91)\n(38, 92)\n(38, 93)\n(38, 94)\n(38, 95)\n(38, 96)\n(38, 97)\n(38, 98)\n(38, 99)\n(39, 0)\n(39, 1)\n(39, 2)\n(39, 3)\n(39, 4)\n(39, 5)\n(39, 6)\n(39, 7)\n(39, 8)\n(39, 9)\n(39, 10)\n(39, 11)\n(39, 12)\n(39, 13)\n(39, 14)\n(39, 15)\n(39, 16)\n(39, 17)\n(39, 18)\n(39, 19)\n(39, 20)\n(39, 21)\n(39, 22)\n(39, 23)\n(39, 24)\n(39, 25)\n(39, 26)\n(39, 27)\n(39, 28)\n(39, 29)\n(39, 30)\n(39, 31)\n(39, 32)\n(39, 33)\n(39, 34)\n(39, 35)\n(39, 36)\n(39, 37)\n(39, 38)\n(39, 39)\n(39, 40)\n(39, 41)\n(39, 42)\n(39, 43)\n(39, 44)\n(39, 45)\n(39, 46)\n(39, 47)\n(39, 48)\n(39, 49)\n(39, 50)\n(39, 51)\n(39, 52)\n(39, 53)\n(39, 54)\n(39, 55)\n(39, 56)\n(39, 57)\n(39, 58)\n(39, 59)\n(39, 60)\n(39, 61)\n(39, 62)\n(39, 63)\n(39, 64)\n(39, 65)\n(39, 66)\n(39, 67)\n(39, 68)\n(39, 69)\n(39, 70)\n(39, 71)\n(39, 72)\n(39, 73)\n(39, 74)\n(39, 75)\n(39, 76)\n(39, 77)\n(39, 78)\n(39, 79)\n(39, 80)\n(39, 81)\n(39, 82)\n(39, 83)\n(39, 84)\n(39, 85)\n(39, 86)\n(39, 87)\n(39, 88)\n(39, 89)\n(39, 90)\n(39, 91)\n(39, 92)\n(39, 93)\n(39, 94)\n(39, 95)\n(39, 96)\n(39, 97)\n(39, 98)\n(39, 99)\n(40, 0)\n(40, 1)\n(40, 2)\n(40, 3)\n(40, 4)\n(40, 5)\n(40, 6)\n(40, 7)\n(40, 8)\n(40, 9)\n(40, 10)\n(40, 11)\n(40, 12)\n(40, 13)\n(40, 14)\n(40, 15)\n(40, 16)\n(40, 17)\n(40, 18)\n(40, 19)\n(40, 20)\n(40, 21)\n(40, 22)\n(40, 23)\n(40, 24)\n(40, 25)\n(40, 26)\n(40, 27)\n(40, 28)\n(40, 29)\n(40, 30)\n(40, 31)\n(40, 32)\n(40, 33)\n(40, 34)\n(40, 35)\n(40, 36)\n(40, 37)\n(40, 38)\n(40, 39)\n(40, 40)\n(40, 41)\n(40, 42)\n(40, 43)\n(40, 44)\n(40, 45)\n(40, 46)\n(40, 47)\n(40, 48)\n(40, 49)\n(40, 50)\n(40, 51)\n(40, 52)\n(40, 53)\n(40, 54)\n(40, 55)\n(40, 56)\n(40, 57)\n(40, 58)\n(40, 59)\n(40, 60)\n(40, 61)\n(40, 62)\n(40, 63)\n(40, 64)\n(40, 65)\n(40, 66)\n(40, 67)\n(40, 68)\n(40, 69)\n(40, 70)\n(40, 71)\n(40, 72)\n(40, 73)\n(40, 74)\n(40, 75)\n(40, 76)\n(40, 77)\n(40, 78)\n(40, 79)\n(40, 80)\n(40, 81)\n(40, 82)\n(40, 83)\n(40, 84)\n(40, 85)\n(40, 86)\n(40, 87)\n(40, 88)\n(40, 89)\n(40, 90)\n(40, 91)\n(40, 92)\n(40, 93)\n(40, 94)\n(40, 95)\n(40, 96)\n(40, 97)\n(40, 98)\n(40, 99)\n(41, 0)\n(41, 1)\n(41, 2)\n(41, 3)\n(41, 4)\n(41, 5)\n(41, 6)\n(41, 7)\n(41, 8)\n(41, 9)\n(41, 10)\n(41, 11)\n(41, 12)\n(41, 13)\n(41, 14)\n(41, 15)\n(41, 16)\n(41, 17)\n(41, 18)\n(41, 19)\n(41, 20)\n(41, 21)\n(41, 22)\n(41, 23)\n(41, 24)\n(41, 25)\n(41, 26)\n(41, 27)\n(41, 28)\n(41, 29)\n(41, 30)\n(41, 31)\n(41, 32)\n(41, 33)\n(41, 34)\n(41, 35)\n(41, 36)\n(41, 37)\n(41, 38)\n(41, 39)\n(41, 40)\n(41, 41)\n(41, 42)\n(41, 43)\n(41, 44)\n(41, 45)\n(41, 46)\n(41, 47)\n(41, 48)\n(41, 49)\n(41, 50)\n(41, 51)\n(41, 52)\n(41, 53)\n(41, 54)\n(41, 55)\n(41, 56)\n(41, 57)\n(41, 58)\n(41, 59)\n(41, 60)\n(41, 61)\n(41, 62)\n(41, 63)\n(41, 64)\n(41, 65)\n(41, 66)\n(41, 67)\n(41, 68)\n(41, 69)\n(41, 70)\n(41, 71)\n(41, 72)\n(41, 73)\n(41, 74)\n(41, 75)\n(41, 76)\n(41, 77)\n(41, 78)\n(41, 79)\n(41, 80)\n(41, 81)\n(41, 82)\n(41, 83)\n(41, 84)\n(41, 85)\n(41, 86)\n(41, 87)\n(41, 88)\n(41, 89)\n(41, 90)\n(41, 91)\n(41, 92)\n(41, 93)\n(41, 94)\n(41, 95)\n(41, 96)\n(41, 97)\n(41, 98)\n(41, 99)\n(42, 0)\n(42, 1)\n(42, 2)\n(42, 3)\n(42, 4)\n(42, 5)\n(42, 6)\n(42, 7)\n(42, 8)\n(42, 9)\n(42, 10)\n(42, 11)\n(42, 12)\n(42, 13)\n(42, 14)\n(42, 15)\n(42, 16)\n(42, 17)\n(42, 18)\n(42, 19)\n(42, 20)\n(42, 21)\n(42, 22)\n(42, 23)\n(42, 24)\n(42, 25)\n(42, 26)\n(42, 27)\n(42, 28)\n(42, 29)\n(42, 30)\n(42, 31)\n(42, 32)\n(42, 33)\n(42, 34)\n(42, 35)\n(42, 36)\n(42, 37)\n(42, 38)\n(42, 39)\n(42, 40)\n(42, 41)\n(42, 42)\n(42, 43)\n(42, 44)\n(42, 45)\n(42, 46)\n(42, 47)\n(42, 48)\n(42, 49)\n(42, 50)\n(42, 51)\n(42, 52)\n(42, 53)\n(42, 54)\n(42, 55)\n(42, 56)\n(42, 57)\n(42, 58)\n(42, 59)\n(42, 60)\n(42, 61)\n(42, 62)\n(42, 63)\n(42, 64)\n(42, 65)\n(42, 66)\n(42, 67)\n(42, 68)\n(42, 69)\n(42, 70)\n(42, 71)\n(42, 72)\n(42, 73)\n(42, 74)\n(42, 75)\n(42, 76)\n(42, 77)\n(42, 78)\n(42, 79)\n(42, 80)\n(42, 81)\n(42, 82)\n(42, 83)\n(42, 84)\n(42, 85)\n(42, 86)\n(42, 87)\n(42, 88)\n(42, 89)\n(42, 90)\n(42, 91)\n(42, 92)\n(42, 93)\n(42, 94)\n(42, 95)\n(42, 96)\n(42, 97)\n(42, 98)\n(42, 99)\n(43, 0)\n(43, 1)\n(43, 2)\n(43, 3)\n(43, 4)\n(43, 5)\n(43, 6)\n(43, 7)\n(43, 8)\n(43, 9)\n(43, 10)\n(43, 11)\n(43, 12)\n(43, 13)\n(43, 14)\n(43, 15)\n(43, 16)\n(43, 17)\n(43, 18)\n(43, 19)\n(43, 20)\n(43, 21)\n(43, 22)\n(43, 23)\n(43, 24)\n(43, 25)\n(43, 26)\n(43, 27)\n(43, 28)\n(43, 29)\n(43, 30)\n(43, 31)\n(43, 32)\n(43, 33)\n(43, 34)\n(43, 35)\n(43, 36)\n(43, 37)\n(43, 38)\n(43, 39)\n(43, 40)\n(43, 41)\n(43, 42)\n(43, 43)\n(43, 44)\n(43, 45)\n(43, 46)\n(43, 47)\n(43, 48)\n(43, 49)\n(43, 50)\n(43, 51)\n(43, 52)\n(43, 53)\n(43, 54)\n(43, 55)\n(43, 56)\n(43, 57)\n(43, 58)\n(43, 59)\n(43, 60)\n(43, 61)\n(43, 62)\n(43, 63)\n(43, 64)\n(43, 65)\n(43, 66)\n(43, 67)\n(43, 68)\n(43, 69)\n(43, 70)\n(43, 71)\n(43, 72)\n(43, 73)\n(43, 74)\n(43, 75)\n(43, 76)\n(43, 77)\n(43, 78)\n(43, 79)\n(43, 80)\n(43, 81)\n(43, 82)\n(43, 83)\n(43, 84)\n(43, 85)\n(43, 86)\n(43, 87)\n(43, 88)\n(43, 89)\n(43, 90)\n(43, 91)\n(43, 92)\n(43, 93)\n(43, 94)\n(43, 95)\n(43, 96)\n(43, 97)\n(43, 98)\n(43, 99)\n(44, 0)\n(44, 1)\n(44, 2)\n(44, 3)\n(44, 4)\n(44, 5)\n(44, 6)\n(44, 7)\n(44, 8)\n(44, 9)\n(44, 10)\n(44, 11)\n(44, 12)\n(44, 13)\n(44, 14)\n(44, 15)\n(44, 16)\n(44, 17)\n(44, 18)\n(44, 19)\n(44, 20)\n(44, 21)\n(44, 22)\n(44, 23)\n(44, 24)\n(44, 25)\n(44, 26)\n(44, 27)\n(44, 28)\n(44, 29)\n(44, 30)\n(44, 31)\n(44, 32)\n(44, 33)\n(44, 34)\n(44, 35)\n(44, 36)\n(44, 37)\n(44, 38)\n(44, 39)\n(44, 40)\n(44, 41)\n(44, 42)\n(44, 43)\n(44, 44)\n(44, 45)\n(44, 46)\n(44, 47)\n(44, 48)\n(44, 49)\n(44, 50)\n(44, 51)\n(44, 52)\n(44, 53)\n(44, 54)\n(44, 55)\n(44, 56)\n(44, 57)\n(44, 58)\n(44, 59)\n(44, 60)\n(44, 61)\n(44, 62)\n(44, 63)\n(44, 64)\n(44, 65)\n(44, 66)\n(44, 67)\n(44, 68)\n(44, 69)\n(44, 70)\n(44, 71)\n(44, 72)\n(44, 73)\n(44, 74)\n(44, 75)\n(44, 76)\n(44, 77)\n(44, 78)\n(44, 79)\n(44, 80)\n(44, 81)\n(44, 82)\n(44, 83)\n(44, 84)\n(44, 85)\n(44, 86)\n(44, 87)\n(44, 88)\n(44, 89)\n(44, 90)\n(44, 91)\n(44, 92)\n(44, 93)\n(44, 94)\n(44, 95)\n(44, 96)\n(44, 97)\n(44, 98)\n(44, 99)\n(45, 0)\n(45, 1)\n(45, 2)\n(45, 3)\n(45, 4)\n(45, 5)\n(45, 6)\n(45, 7)\n(45, 8)\n(45, 9)\n(45, 10)\n(45, 11)\n(45, 12)\n(45, 13)\n(45, 14)\n(45, 15)\n(45, 16)\n(45, 17)\n(45, 18)\n(45, 19)\n(45, 20)\n(45, 21)\n(45, 22)\n(45, 23)\n(45, 24)\n(45, 25)\n(45, 26)\n(45, 27)\n(45, 28)\n(45, 29)\n(45, 30)\n(45, 31)\n(45, 32)\n(45, 33)\n(45, 34)\n(45, 35)\n(45, 36)\n(45, 37)\n(45, 38)\n(45, 39)\n(45, 40)\n(45, 41)\n(45, 42)\n(45, 43)\n(45, 44)\n(45, 45)\n(45, 46)\n(45, 47)\n(45, 48)\n(45, 49)\n(45, 50)\n(45, 51)\n(45, 52)\n(45, 53)\n(45, 54)\n(45, 55)\n(45, 56)\n(45, 57)\n(45, 58)\n(45, 59)\n(45, 60)\n(45, 61)\n(45, 62)\n(45, 63)\n(45, 64)\n(45, 65)\n(45, 66)\n(45, 67)\n(45, 68)\n(45, 69)\n(45, 70)\n(45, 71)\n(45, 72)\n(45, 73)\n(45, 74)\n(45, 75)\n(45, 76)\n(45, 77)\n(45, 78)\n(45, 79)\n(45, 80)\n(45, 81)\n(45, 82)\n(45, 83)\n(45, 84)\n(45, 85)\n(45, 86)\n(45, 87)\n(45, 88)\n(45, 89)\n(45, 90)\n(45, 91)\n(45, 92)\n(45, 93)\n(45, 94)\n(45, 95)\n(45, 96)\n(45, 97)\n(45, 98)\n(45, 99)\n(46, 0)\n(46, 1)\n(46, 2)\n(46, 3)\n(46, 4)\n(46, 5)\n(46, 6)\n(46, 7)\n(46, 8)\n(46, 9)\n(46, 10)\n(46, 11)\n(46, 12)\n(46, 13)\n(46, 14)\n(46, 15)\n(46, 16)\n(46, 17)\n(46, 18)\n(46, 19)\n(46, 20)\n(46, 21)\n(46, 22)\n(46, 23)\n(46, 24)\n(46, 25)\n(46, 26)\n(46, 27)\n(46, 28)\n(46, 29)\n(46, 30)\n(46, 31)\n(46, 32)\n(46, 33)\n(46, 34)\n(46, 35)\n(46, 36)\n(46, 37)\n(46, 38)\n(46, 39)\n(46, 40)\n(46, 41)\n(46, 42)\n(46, 43)\n(46, 44)\n(46, 45)\n(46, 46)\n(46, 47)\n(46, 48)\n(46, 49)\n(46, 50)\n(46, 51)\n(46, 52)\n(46, 53)\n(46, 54)\n(46, 55)\n(46, 56)\n(46, 57)\n(46, 58)\n(46, 59)\n(46, 60)\n(46, 61)\n(46, 62)\n(46, 63)\n(46, 64)\n(46, 65)\n(46, 66)\n(46, 67)\n(46, 68)\n(46, 69)\n(46, 70)\n(46, 71)\n(46, 72)\n(46, 73)\n(46, 74)\n(46, 75)\n(46, 76)\n(46, 77)\n(46, 78)\n(46, 79)\n(46, 80)\n(46, 81)\n(46, 82)\n(46, 83)\n(46, 84)\n(46, 85)\n(46, 86)\n(46, 87)\n(46, 88)\n(46, 89)\n(46, 90)\n(46, 91)\n(46, 92)\n(46, 93)\n(46, 94)\n(46, 95)\n(46, 96)\n(46, 97)\n(46, 98)\n(46, 99)\n(47, 0)\n(47, 1)\n(47, 2)\n(47, 3)\n(47, 4)\n(47, 5)\n(47, 6)\n(47, 7)\n(47, 8)\n(47, 9)\n(47, 10)\n(47, 11)\n(47, 12)\n(47, 13)\n(47, 14)\n(47, 15)\n(47, 16)\n(47, 17)\n(47, 18)\n(47, 19)\n(47, 20)\n(47, 21)\n(47, 22)\n(47, 23)\n(47, 24)\n(47, 25)\n(47, 26)\n(47, 27)\n(47, 28)\n(47, 29)\n(47, 30)\n(47, 31)\n(47, 32)\n(47, 33)\n(47, 34)\n(47, 35)\n(47, 36)\n(47, 37)\n(47, 38)\n(47, 39)\n(47, 40)\n(47, 41)\n(47, 42)\n(47, 43)\n(47, 44)\n(47, 45)\n(47, 46)\n(47, 47)\n(47, 48)\n(47, 49)\n(47, 50)\n(47, 51)\n(47, 52)\n(47, 53)\n(47, 54)\n(47, 55)\n(47, 56)\n(47, 57)\n(47, 58)\n(47, 59)\n(47, 60)\n(47, 61)\n(47, 62)\n(47, 63)\n(47, 64)\n(47, 65)\n(47, 66)\n(47, 67)\n(47, 68)\n(47, 69)\n(47, 70)\n(47, 71)\n(47, 72)\n(47, 73)\n(47, 74)\n(47, 75)\n(47, 76)\n(47, 77)\n(47, 78)\n(47, 79)\n(47, 80)\n(47, 81)\n(47, 82)\n(47, 83)\n(47, 84)\n(47, 85)\n(47, 86)\n(47, 87)\n(47, 88)\n(47, 89)\n(47, 90)\n(47, 91)\n(47, 92)\n(47, 93)\n(47, 94)\n(47, 95)\n(47, 96)\n(47, 97)\n(47, 98)\n(47, 99)\n(48, 0)\n(48, 1)\n(48, 2)\n(48, 3)\n(48, 4)\n(48, 5)\n(48, 6)\n(48, 7)\n(48, 8)\n(48, 9)\n(48, 10)\n(48, 11)\n(48, 12)\n(48, 13)\n(48, 14)\n(48, 15)\n(48, 16)\n(48, 17)\n(48, 18)\n(48, 19)\n(48, 20)\n(48, 21)\n(48, 22)\n(48, 23)\n(48, 24)\n(48, 25)\n(48, 26)\n(48, 27)\n(48, 28)\n(48, 29)\n(48, 30)\n(48, 31)\n(48, 32)\n(48, 33)\n(48, 34)\n(48, 35)\n(48, 36)\n(48, 37)\n(48, 38)\n(48, 39)\n(48, 40)\n(48, 41)\n(48, 42)\n(48, 43)\n(48, 44)\n(48, 45)\n(48, 46)\n(48, 47)\n(48, 48)\n(48, 49)\n(48, 50)\n(48, 51)\n(48, 52)\n(48, 53)\n(48, 54)\n(48, 55)\n(48, 56)\n(48, 57)\n(48, 58)\n(48, 59)\n(48, 60)\n(48, 61)\n(48, 62)\n(48, 63)\n(48, 64)\n(48, 65)\n(48, 66)\n(48, 67)\n(48, 68)\n(48, 69)\n(48, 70)\n(48, 71)\n(48, 72)\n(48, 73)\n(48, 74)\n(48, 75)\n(48, 76)\n(48, 77)\n(48, 78)\n(48, 79)\n(48, 80)\n(48, 81)\n(48, 82)\n(48, 83)\n(48, 84)\n(48, 85)\n(48, 86)\n(48, 87)\n(48, 88)\n(48, 89)\n(48, 90)\n(48, 91)\n(48, 92)\n(48, 93)\n(48, 94)\n(48, 95)\n(48, 96)\n(48, 97)\n(48, 98)\n(48, 99)\n(49, 0)\n(49, 1)\n(49, 2)\n(49, 3)\n(49, 4)\n(49, 5)\n(49, 6)\n(49, 7)\n(49, 8)\n(49, 9)\n(49, 10)\n(49, 11)\n(49, 12)\n(49, 13)\n(49, 14)\n(49, 15)\n(49, 16)\n(49, 17)\n(49, 18)\n(49, 19)\n(49, 20)\n(49, 21)\n(49, 22)\n(49, 23)\n(49, 24)\n(49, 25)\n(49, 26)\n(49, 27)\n(49, 28)\n(49, 29)\n(49, 30)\n(49, 31)\n(49, 32)\n(49, 33)\n(49, 34)\n(49, 35)\n(49, 36)\n(49, 37)\n(49, 38)\n(49, 39)\n(49, 40)\n(49, 41)\n(49, 42)\n(49, 43)\n(49, 44)\n(49, 45)\n(49, 46)\n(49, 47)\n(49, 48)\n(49, 49)\n(49, 50)\n(49, 51)\n(49, 52)\n(49, 53)\n(49, 54)\n(49, 55)\n(49, 56)\n(49, 57)\n(49, 58)\n(49, 59)\n(49, 60)\n(49, 61)\n(49, 62)\n(49, 63)\n(49, 64)\n(49, 65)\n(49, 66)\n(49, 67)\n(49, 68)\n(49, 69)\n(49, 70)\n(49, 71)\n(49, 72)\n(49, 73)\n(49, 74)\n(49, 75)\n(49, 76)\n(49, 77)\n(49, 78)\n(49, 79)\n(49, 80)\n(49, 81)\n(49, 82)\n(49, 83)\n(49, 84)\n(49, 85)\n(49, 86)\n(49, 87)\n(49, 88)\n(49, 89)\n(49, 90)\n(49, 91)\n(49, 92)\n(49, 93)\n(49, 94)\n(49, 95)\n(49, 96)\n(49, 97)\n(49, 98)\n(49, 99)\n(50, 0)\n(50, 1)\n(50, 2)\n(50, 3)\n(50, 4)\n(50, 5)\n(50, 6)\n(50, 7)\n(50, 8)\n(50, 9)\n(50, 10)\n(50, 11)\n(50, 12)\n(50, 13)\n(50, 14)\n(50, 15)\n(50, 16)\n(50, 17)\n(50, 18)\n(50, 19)\n(50, 20)\n(50, 21)\n(50, 22)\n(50, 23)\n(50, 24)\n(50, 25)\n(50, 26)\n(50, 27)\n(50, 28)\n(50, 29)\n(50, 30)\n(50, 31)\n(50, 32)\n(50, 33)\n(50, 34)\n(50, 35)\n(50, 36)\n(50, 37)\n(50, 38)\n(50, 39)\n(50, 40)\n(50, 41)\n(50, 42)\n(50, 43)\n(50, 44)\n(50, 45)\n(50, 46)\n(50, 47)\n(50, 48)\n(50, 49)\n(50, 50)\n(50, 51)\n(50, 52)\n(50, 53)\n(50, 54)\n(50, 55)\n(50, 56)\n(50, 57)\n(50, 58)\n(50, 59)\n(50, 60)\n(50, 61)\n(50, 62)\n(50, 63)\n(50, 64)\n(50, 65)\n(50, 66)\n(50, 67)\n(50, 68)\n(50, 69)\n(50, 70)\n(50, 71)\n(50, 72)\n(50, 73)\n(50, 74)\n(50, 75)\n(50, 76)\n(50, 77)\n(50, 78)\n(50, 79)\n(50, 80)\n(50, 81)\n(50, 82)\n(50, 83)\n(50, 84)\n(50, 85)\n(50, 86)\n(50, 87)\n(50, 88)\n(50, 89)\n(50, 90)\n(50, 91)\n(50, 92)\n(50, 93)\n(50, 94)\n(50, 95)\n(50, 96)\n(50, 97)\n(50, 98)\n(50, 99)\n(51, 0)\n(51, 1)\n(51, 2)\n(51, 3)\n(51, 4)\n(51, 5)\n(51, 6)\n(51, 7)\n(51, 8)\n(51, 9)\n(51, 10)\n(51, 11)\n(51, 12)\n(51, 13)\n(51, 14)\n(51, 15)\n(51, 16)\n(51, 17)\n(51, 18)\n(51, 19)\n(51, 20)\n(51, 21)\n(51, 22)\n(51, 23)\n(51, 24)\n(51, 25)\n(51, 26)\n(51, 27)\n(51, 28)\n(51, 29)\n(51, 30)\n(51, 31)\n(51, 32)\n(51, 33)\n(51, 34)\n(51, 35)\n(51, 36)\n(51, 37)\n(51, 38)\n(51, 39)\n(51, 40)\n(51, 41)\n(51, 42)\n(51, 43)\n(51, 44)\n(51, 45)\n(51, 46)\n(51, 47)\n(51, 48)\n(51, 49)\n(51, 50)\n(51, 51)\n(51, 52)\n(51, 53)\n(51, 54)\n(51, 55)\n(51, 56)\n(51, 57)\n(51, 58)\n(51, 59)\n(51, 60)\n(51, 61)\n(51, 62)\n(51, 63)\n(51, 64)\n(51, 65)\n(51, 66)\n(51, 67)\n(51, 68)\n(51, 69)\n(51, 70)\n(51, 71)\n(51, 72)\n(51, 73)\n(51, 74)\n(51, 75)\n(51, 76)\n(51, 77)\n(51, 78)\n(51, 79)\n(51, 80)\n(51, 81)\n(51, 82)\n(51, 83)\n(51, 84)\n(51, 85)\n(51, 86)\n(51, 87)\n(51, 88)\n(51, 89)\n(51, 90)\n(51, 91)\n(51, 92)\n(51, 93)\n(51, 94)\n(51, 95)\n(51, 96)\n(51, 97)\n(51, 98)\n(51, 99)\n(52, 0)\n(52, 1)\n(52, 2)\n(52, 3)\n(52, 4)\n(52, 5)\n(52, 6)\n(52, 7)\n(52, 8)\n(52, 9)\n(52, 10)\n(52, 11)\n(52, 12)\n(52, 13)\n(52, 14)\n(52, 15)\n(52, 16)\n(52, 17)\n(52, 18)\n(52, 19)\n(52, 20)\n(52, 21)\n(52, 22)\n(52, 23)\n(52, 24)\n(52, 25)\n(52, 26)\n(52, 27)\n(52, 28)\n(52, 29)\n(52, 30)\n(52, 31)\n(52, 32)\n(52, 33)\n(52, 34)\n(52, 35)\n(52, 36)\n(52, 37)\n(52, 38)\n(52, 39)\n(52, 40)\n(52, 41)\n(52, 42)\n(52, 43)\n(52, 44)\n(52, 45)\n(52, 46)\n(52, 47)\n(52, 48)\n(52, 49)\n(52, 50)\n(52, 51)\n(52, 52)\n(52, 53)\n(52, 54)\n(52, 55)\n(52, 56)\n(52, 57)\n(52, 58)\n(52, 59)\n(52, 60)\n(52, 61)\n(52, 62)\n(52, 63)\n(52, 64)\n(52, 65)\n(52, 66)\n(52, 67)\n(52, 68)\n(52, 69)\n(52, 70)\n(52, 71)\n(52, 72)\n(52, 73)\n(52, 74)\n(52, 75)\n(52, 76)\n(52, 77)\n(52, 78)\n(52, 79)\n(52, 80)\n(52, 81)\n(52, 82)\n(52, 83)\n(52, 84)\n(52, 85)\n(52, 86)\n(52, 87)\n(52, 88)\n(52, 89)\n(52, 90)\n(52, 91)\n(52, 92)\n(52, 93)\n(52, 94)\n(52, 95)\n(52, 96)\n(52, 97)\n(52, 98)\n(52, 99)\n(53, 0)\n(53, 1)\n(53, 2)\n(53, 3)\n(53, 4)\n(53, 5)\n(53, 6)\n(53, 7)\n(53, 8)\n(53, 9)\n(53, 10)\n(53, 11)\n(53, 12)\n(53, 13)\n(53, 14)\n(53, 15)\n(53, 16)\n(53, 17)\n(53, 18)\n(53, 19)\n(53, 20)\n(53, 21)\n(53, 22)\n(53, 23)\n(53, 24)\n(53, 25)\n(53, 26)\n(53, 27)\n(53, 28)\n(53, 29)\n(53, 30)\n(53, 31)\n(53, 32)\n(53, 33)\n(53, 34)\n(53, 35)\n(53, 36)\n(53, 37)\n(53, 38)\n(53, 39)\n(53, 40)\n(53, 41)\n(53, 42)\n(53, 43)\n(53, 44)\n(53, 45)\n(53, 46)\n(53, 47)\n(53, 48)\n(53, 49)\n(53, 50)\n(53, 51)\n(53, 52)\n(53, 53)\n(53, 54)\n(53, 55)\n(53, 56)\n(53, 57)\n(53, 58)\n(53, 59)\n(53, 60)\n(53, 61)\n(53, 62)\n(53, 63)\n(53, 64)\n(53, 65)\n(53, 66)\n(53, 67)\n(53, 68)\n(53, 69)\n(53, 70)\n(53, 71)\n(53, 72)\n(53, 73)\n(53, 74)\n(53, 75)\n(53, 76)\n(53, 77)\n(53, 78)\n(53, 79)\n(53, 80)\n(53, 81)\n(53, 82)\n(53, 83)\n(53, 84)\n(53, 85)\n(53, 86)\n(53, 87)\n(53, 88)\n(53, 89)\n(53, 90)\n(53, 91)\n(53, 92)\n(53, 93)\n(53, 94)\n(53, 95)\n(53, 96)\n(53, 97)\n(53, 98)\n(53, 99)\n(54, 0)\n(54, 1)\n(54, 2)\n(54, 3)\n(54, 4)\n(54, 5)\n(54, 6)\n(54, 7)\n(54, 8)\n(54, 9)\n(54, 10)\n(54, 11)\n(54, 12)\n(54, 13)\n(54, 14)\n(54, 15)\n(54, 16)\n(54, 17)\n(54, 18)\n(54, 19)\n(54, 20)\n(54, 21)\n(54, 22)\n(54, 23)\n(54, 24)\n(54, 25)\n(54, 26)\n(54, 27)\n(54, 28)\n(54, 29)\n(54, 30)\n(54, 31)\n(54, 32)\n(54, 33)\n(54, 34)\n(54, 35)\n(54, 36)\n(54, 37)\n(54, 38)\n(54, 39)\n(54, 40)\n(54, 41)\n(54, 42)\n(54, 43)\n(54, 44)\n(54, 45)\n(54, 46)\n(54, 47)\n(54, 48)\n(54, 49)\n(54, 50)\n(54, 51)\n(54, 52)\n(54, 53)\n(54, 54)\n(54, 55)\n(54, 56)\n(54, 57)\n(54, 58)\n(54, 59)\n(54, 60)\n(54, 61)\n(54, 62)\n(54, 63)\n(54, 64)\n(54, 65)\n(54, 66)\n(54, 67)\n(54, 68)\n(54, 69)\n(54, 70)\n(54, 71)\n(54, 72)\n(54, 73)\n(54, 74)\n(54, 75)\n(54, 76)\n(54, 77)\n(54, 78)\n(54, 79)\n(54, 80)\n(54, 81)\n(54, 82)\n(54, 83)\n(54, 84)\n(54, 85)\n(54, 86)\n(54, 87)\n(54, 88)\n(54, 89)\n(54, 90)\n(54, 91)\n(54, 92)\n(54, 93)\n(54, 94)\n(54, 95)\n(54, 96)\n(54, 97)\n(54, 98)\n(54, 99)\n(55, 0)\n(55, 1)\n(55, 2)\n(55, 3)\n(55, 4)\n(55, 5)\n(55, 6)\n(55, 7)\n(55, 8)\n(55, 9)\n(55, 10)\n(55, 11)\n(55, 12)\n(55, 13)\n(55, 14)\n(55, 15)\n(55, 16)\n(55, 17)\n(55, 18)\n(55, 19)\n(55, 20)\n(55, 21)\n(55, 22)\n(55, 23)\n(55, 24)\n(55, 25)\n(55, 26)\n(55, 27)\n(55, 28)\n(55, 29)\n(55, 30)\n(55, 31)\n(55, 32)\n(55, 33)\n(55, 34)\n(55, 35)\n(55, 36)\n(55, 37)\n(55, 38)\n(55, 39)\n(55, 40)\n(55, 41)\n(55, 42)\n(55, 43)\n(55, 44)\n(55, 45)\n(55, 46)\n(55, 47)\n(55, 48)\n(55, 49)\n(55, 50)\n(55, 51)\n(55, 52)\n(55, 53)\n(55, 54)\n(55, 55)\n(55, 56)\n(55, 57)\n(55, 58)\n(55, 59)\n(55, 60)\n(55, 61)\n(55, 62)\n(55, 63)\n(55, 64)\n(55, 65)\n(55, 66)\n(55, 67)\n(55, 68)\n(55, 69)\n(55, 70)\n(55, 71)\n(55, 72)\n(55, 73)\n(55, 74)\n(55, 75)\n(55, 76)\n(55, 77)\n(55, 78)\n(55, 79)\n(55, 80)\n(55, 81)\n(55, 82)\n(55, 83)\n(55, 84)\n(55, 85)\n(55, 86)\n(55, 87)\n(55, 88)\n(55, 89)\n(55, 90)\n(55, 91)\n(55, 92)\n(55, 93)\n(55, 94)\n(55, 95)\n(55, 96)\n(55, 97)\n(55, 98)\n(55, 99)\n(56, 0)\n(56, 1)\n(56, 2)\n(56, 3)\n(56, 4)\n(56, 5)\n(56, 6)\n(56, 7)\n(56, 8)\n(56, 9)\n(56, 10)\n(56, 11)\n(56, 12)\n(56, 13)\n(56, 14)\n(56, 15)\n(56, 16)\n(56, 17)\n(56, 18)\n(56, 19)\n(56, 20)\n(56, 21)\n(56, 22)\n(56, 23)\n(56, 24)\n(56, 25)\n(56, 26)\n(56, 27)\n(56, 28)\n(56, 29)\n(56, 30)\n(56, 31)\n(56, 32)\n(56, 33)\n(56, 34)\n(56, 35)\n(56, 36)\n(56, 37)\n(56, 38)\n(56, 39)\n(56, 40)\n(56, 41)\n(56, 42)\n(56, 43)\n(56, 44)\n(56, 45)\n(56, 46)\n(56, 47)\n(56, 48)\n(56, 49)\n(56, 50)\n(56, 51)\n(56, 52)\n(56, 53)\n(56, 54)\n(56, 55)\n(56, 56)\n(56, 57)\n(56, 58)\n(56, 59)\n(56, 60)\n(56, 61)\n(56, 62)\n(56, 63)\n(56, 64)\n(56, 65)\n(56, 66)\n(56, 67)\n(56, 68)\n(56, 69)\n(56, 70)\n(56, 71)\n(56, 72)\n(56, 73)\n(56, 74)\n(56, 75)\n(56, 76)\n(56, 77)\n(56, 78)\n(56, 79)\n(56, 80)\n(56, 81)\n(56, 82)\n(56, 83)\n(56, 84)\n(56, 85)\n(56, 86)\n(56, 87)\n(56, 88)\n(56, 89)\n(56, 90)\n(56, 91)\n(56, 92)\n(56, 93)\n(56, 94)\n(56, 95)\n(56, 96)\n(56, 97)\n(56, 98)\n(56, 99)\n(57, 0)\n(57, 1)\n(57, 2)\n(57, 3)\n(57, 4)\n(57, 5)\n(57, 6)\n(57, 7)\n(57, 8)\n(57, 9)\n(57, 10)\n(57, 11)\n(57, 12)\n(57, 13)\n(57, 14)\n(57, 15)\n(57, 16)\n(57, 17)\n(57, 18)\n(57, 19)\n(57, 20)\n(57, 21)\n(57, 22)\n(57, 23)\n(57, 24)\n(57, 25)\n(57, 26)\n(57, 27)\n(57, 28)\n(57, 29)\n(57, 30)\n(57, 31)\n(57, 32)\n(57, 33)\n(57, 34)\n(57, 35)\n(57, 36)\n(57, 37)\n(57, 38)\n(57, 39)\n(57, 40)\n(57, 41)\n(57, 42)\n(57, 43)\n(57, 44)\n(57, 45)\n(57, 46)\n(57, 47)\n(57, 48)\n(57, 49)\n(57, 50)\n(57, 51)\n(57, 52)\n(57, 53)\n(57, 54)\n(57, 55)\n(57, 56)\n(57, 57)\n(57, 58)\n(57, 59)\n(57, 60)\n(57, 61)\n(57, 62)\n(57, 63)\n(57, 64)\n(57, 65)\n(57, 66)\n(57, 67)\n(57, 68)\n(57, 69)\n(57, 70)\n(57, 71)\n(57, 72)\n(57, 73)\n(57, 74)\n(57, 75)\n(57, 76)\n(57, 77)\n(57, 78)\n(57, 79)\n(57, 80)\n(57, 81)\n(57, 82)\n(57, 83)\n(57, 84)\n(57, 85)\n(57, 86)\n(57, 87)\n(57, 88)\n(57, 89)\n(57, 90)\n(57, 91)\n(57, 92)\n(57, 93)\n(57, 94)\n(57, 95)\n(57, 96)\n(57, 97)\n(57, 98)\n(57, 99)\n(58, 0)\n(58, 1)\n(58, 2)\n(58, 3)\n(58, 4)\n(58, 5)\n(58, 6)\n(58, 7)\n(58, 8)\n(58, 9)\n(58, 10)\n(58, 11)\n(58, 12)\n(58, 13)\n(58, 14)\n(58, 15)\n(58, 16)\n(58, 17)\n(58, 18)\n(58, 19)\n(58, 20)\n(58, 21)\n(58, 22)\n(58, 23)\n(58, 24)\n(58, 25)\n(58, 26)\n(58, 27)\n(58, 28)\n(58, 29)\n(58, 30)\n(58, 31)\n(58, 32)\n(58, 33)\n(58, 34)\n(58, 35)\n(58, 36)\n(58, 37)\n(58, 38)\n(58, 39)\n(58, 40)\n(58, 41)\n(58, 42)\n(58, 43)\n(58, 44)\n(58, 45)\n(58, 46)\n(58, 47)\n(58, 48)\n(58, 49)\n(58, 50)\n(58, 51)\n(58, 52)\n(58, 53)\n(58, 54)\n(58, 55)\n(58, 56)\n(58, 57)\n(58, 58)\n(58, 59)\n(58, 60)\n(58, 61)\n(58, 62)\n(58, 63)\n(58, 64)\n(58, 65)\n(58, 66)\n(58, 67)\n(58, 68)\n(58, 69)\n(58, 70)\n(58, 71)\n(58, 72)\n(58, 73)\n(58, 74)\n(58, 75)\n(58, 76)\n(58, 77)\n(58, 78)\n(58, 79)\n(58, 80)\n(58, 81)\n(58, 82)\n(58, 83)\n(58, 84)\n(58, 85)\n(58, 86)\n(58, 87)\n(58, 88)\n(58, 89)\n(58, 90)\n(58, 91)\n(58, 92)\n(58, 93)\n(58, 94)\n(58, 95)\n(58, 96)\n(58, 97)\n(58, 98)\n(58, 99)\n(59, 0)\n(59, 1)\n(59, 2)\n(59, 3)\n(59, 4)\n(59, 5)\n(59, 6)\n(59, 7)\n(59, 8)\n(59, 9)\n(59, 10)\n(59, 11)\n(59, 12)\n(59, 13)\n(59, 14)\n(59, 15)\n(59, 16)\n(59, 17)\n(59, 18)\n(59, 19)\n(59, 20)\n(59, 21)\n(59, 22)\n(59, 23)\n(59, 24)\n(59, 25)\n(59, 26)\n(59, 27)\n(59, 28)\n(59, 29)\n(59, 30)\n(59, 31)\n(59, 32)\n(59, 33)\n(59, 34)\n(59, 35)\n(59, 36)\n(59, 37)\n(59, 38)\n(59, 39)\n(59, 40)\n(59, 41)\n(59, 42)\n(59, 43)\n(59, 44)\n(59, 45)\n(59, 46)\n(59, 47)\n(59, 48)\n(59, 49)\n(59, 50)\n(59, 51)\n(59, 52)\n(59, 53)\n(59, 54)\n(59, 55)\n(59, 56)\n(59, 57)\n(59, 58)\n(59, 59)\n(59, 60)\n(59, 61)\n(59, 62)\n(59, 63)\n(59, 64)\n(59, 65)\n(59, 66)\n(59, 67)\n(59, 68)\n(59, 69)\n(59, 70)\n(59, 71)\n(59, 72)\n(59, 73)\n(59, 74)\n(59, 75)\n(59, 76)\n(59, 77)\n(59, 78)\n(59, 79)\n(59, 80)\n(59, 81)\n(59, 82)\n(59, 83)\n(59, 84)\n(59, 85)\n(59, 86)\n(59, 87)\n(59, 88)\n(59, 89)\n(59, 90)\n(59, 91)\n(59, 92)\n(59, 93)\n(59, 94)\n(59, 95)\n(59, 96)\n(59, 97)\n(59, 98)\n(59, 99)\n(60, 0)\n(60, 1)\n(60, 2)\n(60, 3)\n(60, 4)\n(60, 5)\n(60, 6)\n(60, 7)\n(60, 8)\n(60, 9)\n(60, 10)\n(60, 11)\n(60, 12)\n(60, 13)\n(60, 14)\n(60, 15)\n(60, 16)\n(60, 17)\n(60, 18)\n(60, 19)\n(60, 20)\n(60, 21)\n(60, 22)\n(60, 23)\n(60, 24)\n(60, 25)\n(60, 26)\n(60, 27)\n(60, 28)\n(60, 29)\n(60, 30)\n(60, 31)\n(60, 32)\n(60, 33)\n(60, 34)\n(60, 35)\n(60, 36)\n(60, 37)\n(60, 38)\n(60, 39)\n(60, 40)\n(60, 41)\n(60, 42)\n(60, 43)\n(60, 44)\n(60, 45)\n(60, 46)\n(60, 47)\n(60, 48)\n(60, 49)\n(60, 50)\n(60, 51)\n(60, 52)\n(60, 53)\n(60, 54)\n(60, 55)\n(60, 56)\n(60, 57)\n(60, 58)\n(60, 59)\n(60, 60)\n(60, 61)\n(60, 62)\n(60, 63)\n(60, 64)\n(60, 65)\n(60, 66)\n(60, 67)\n(60, 68)\n(60, 69)\n(60, 70)\n(60, 71)\n(60, 72)\n(60, 73)\n(60, 74)\n(60, 75)\n(60, 76)\n(60, 77)\n(60, 78)\n(60, 79)\n(60, 80)\n(60, 81)\n(60, 82)\n(60, 83)\n(60, 84)\n(60, 85)\n(60, 86)\n(60, 87)\n(60, 88)\n(60, 89)\n(60, 90)\n(60, 91)\n(60, 92)\n(60, 93)\n(60, 94)\n(60, 95)\n(60, 96)\n(60, 97)\n(60, 98)\n(60, 99)\n(61, 0)\n(61, 1)\n(61, 2)\n(61, 3)\n(61, 4)\n(61, 5)\n(61, 6)\n(61, 7)\n(61, 8)\n(61, 9)\n(61, 10)\n(61, 11)\n(61, 12)\n(61, 13)\n(61, 14)\n(61, 15)\n(61, 16)\n(61, 17)\n(61, 18)\n(61, 19)\n(61, 20)\n(61, 21)\n(61, 22)\n(61, 23)\n(61, 24)\n(61, 25)\n(61, 26)\n(61, 27)\n(61, 28)\n(61, 29)\n(61, 30)\n(61, 31)\n(61, 32)\n(61, 33)\n(61, 34)\n(61, 35)\n(61, 36)\n(61, 37)\n(61, 38)\n(61, 39)\n(61, 40)\n(61, 41)\n(61, 42)\n(61, 43)\n(61, 44)\n(61, 45)\n(61, 46)\n(61, 47)\n(61, 48)\n(61, 49)\n(61, 50)\n(61, 51)\n(61, 52)\n(61, 53)\n(61, 54)\n(61, 55)\n(61, 56)\n(61, 57)\n(61, 58)\n(61, 59)\n(61, 60)\n(61, 61)\n(61, 62)\n(61, 63)\n(61, 64)\n(61, 65)\n(61, 66)\n(61, 67)\n(61, 68)\n(61, 69)\n(61, 70)\n(61, 71)\n(61, 72)\n(61, 73)\n(61, 74)\n(61, 75)\n(61, 76)\n(61, 77)\n(61, 78)\n(61, 79)\n(61, 80)\n(61, 81)\n(61, 82)\n(61, 83)\n(61, 84)\n(61, 85)\n(61, 86)\n(61, 87)\n(61, 88)\n(61, 89)\n(61, 90)\n(61, 91)\n(61, 92)\n(61, 93)\n(61, 94)\n(61, 95)\n(61, 96)\n(61, 97)\n(61, 98)\n(61, 99)\n(62, 0)\n(62, 1)\n(62, 2)\n(62, 3)\n(62, 4)\n(62, 5)\n(62, 6)\n(62, 7)\n(62, 8)\n(62, 9)\n(62, 10)\n(62, 11)\n(62, 12)\n(62, 13)\n(62, 14)\n(62, 15)\n(62, 16)\n(62, 17)\n(62, 18)\n(62, 19)\n(62, 20)\n(62, 21)\n(62, 22)\n(62, 23)\n(62, 24)\n(62, 25)\n(62, 26)\n(62, 27)\n(62, 28)\n(62, 29)\n(62, 30)\n(62, 31)\n(62, 32)\n(62, 33)\n(62, 34)\n(62, 35)\n(62, 36)\n(62, 37)\n(62, 38)\n(62, 39)\n(62, 40)\n(62, 41)\n(62, 42)\n(62, 43)\n(62, 44)\n(62, 45)\n(62, 46)\n(62, 47)\n(62, 48)\n(62, 49)\n(62, 50)\n(62, 51)\n(62, 52)\n(62, 53)\n(62, 54)\n(62, 55)\n(62, 56)\n(62, 57)\n(62, 58)\n(62, 59)\n(62, 60)\n(62, 61)\n(62, 62)\n(62, 63)\n(62, 64)\n(62, 65)\n(62, 66)\n(62, 67)\n(62, 68)\n(62, 69)\n(62, 70)\n(62, 71)\n(62, 72)\n(62, 73)\n(62, 74)\n(62, 75)\n(62, 76)\n(62, 77)\n(62, 78)\n(62, 79)\n(62, 80)\n(62, 81)\n(62, 82)\n(62, 83)\n(62, 84)\n(62, 85)\n(62, 86)\n(62, 87)\n(62, 88)\n(62, 89)\n(62, 90)\n(62, 91)\n(62, 92)\n(62, 93)\n(62, 94)\n(62, 95)\n(62, 96)\n(62, 97)\n(62, 98)\n(62, 99)\n(63, 0)\n(63, 1)\n(63, 2)\n(63, 3)\n(63, 4)\n(63, 5)\n(63, 6)\n(63, 7)\n(63, 8)\n(63, 9)\n(63, 10)\n(63, 11)\n(63, 12)\n(63, 13)\n(63, 14)\n(63, 15)\n(63, 16)\n(63, 17)\n(63, 18)\n(63, 19)\n(63, 20)\n(63, 21)\n(63, 22)\n(63, 23)\n(63, 24)\n(63, 25)\n(63, 26)\n(63, 27)\n(63, 28)\n(63, 29)\n(63, 30)\n(63, 31)\n(63, 32)\n(63, 33)\n(63, 34)\n(63, 35)\n(63, 36)\n(63, 37)\n(63, 38)\n(63, 39)\n(63, 40)\n(63, 41)\n(63, 42)\n(63, 43)\n(63, 44)\n(63, 45)\n(63, 46)\n(63, 47)\n(63, 48)\n(63, 49)\n(63, 50)\n(63, 51)\n(63, 52)\n(63, 53)\n(63, 54)\n(63, 55)\n(63, 56)\n(63, 57)\n(63, 58)\n(63, 59)\n(63, 60)\n(63, 61)\n(63, 62)\n(63, 63)\n(63, 64)\n(63, 65)\n(63, 66)\n(63, 67)\n(63, 68)\n(63, 69)\n(63, 70)\n(63, 71)\n(63, 72)\n(63, 73)\n(63, 74)\n(63, 75)\n(63, 76)\n(63, 77)\n(63, 78)\n(63, 79)\n(63, 80)\n(63, 81)\n(63, 82)\n(63, 83)\n(63, 84)\n(63, 85)\n(63, 86)\n(63, 87)\n(63, 88)\n(63, 89)\n(63, 90)\n(63, 91)\n(63, 92)\n(63, 93)\n(63, 94)\n(63, 95)\n(63, 96)\n(63, 97)\n(63, 98)\n(63, 99)\n(64, 0)\n(64, 1)\n(64, 2)\n(64, 3)\n(64, 4)\n(64, 5)\n(64, 6)\n(64, 7)\n(64, 8)\n(64, 9)\n(64, 10)\n(64, 11)\n(64, 12)\n(64, 13)\n(64, 14)\n(64, 15)\n(64, 16)\n(64, 17)\n(64, 18)\n(64, 19)\n(64, 20)\n(64, 21)\n(64, 22)\n(64, 23)\n(64, 24)\n(64, 25)\n(64, 26)\n(64, 27)\n(64, 28)\n(64, 29)\n(64, 30)\n(64, 31)\n(64, 32)\n(64, 33)\n(64, 34)\n(64, 35)\n(64, 36)\n(64, 37)\n(64, 38)\n(64, 39)\n(64, 40)\n(64, 41)\n(64, 42)\n(64, 43)\n(64, 44)\n(64, 45)\n(64, 46)\n(64, 47)\n(64, 48)\n(64, 49)\n(64, 50)\n(64, 51)\n(64, 52)\n(64, 53)\n(64, 54)\n(64, 55)\n(64, 56)\n(64, 57)\n(64, 58)\n(64, 59)\n(64, 60)\n(64, 61)\n(64, 62)\n(64, 63)\n(64, 64)\n(64, 65)\n(64, 66)\n(64, 67)\n(64, 68)\n(64, 69)\n(64, 70)\n(64, 71)\n(64, 72)\n(64, 73)\n(64, 74)\n(64, 75)\n(64, 76)\n(64, 77)\n(64, 78)\n(64, 79)\n(64, 80)\n(64, 81)\n(64, 82)\n(64, 83)\n(64, 84)\n(64, 85)\n(64, 86)\n(64, 87)\n(64, 88)\n(64, 89)\n(64, 90)\n(64, 91)\n(64, 92)\n(64, 93)\n(64, 94)\n(64, 95)\n(64, 96)\n(64, 97)\n(64, 98)\n(64, 99)\n(65, 0)\n(65, 1)\n(65, 2)\n(65, 3)\n(65, 4)\n(65, 5)\n(65, 6)\n(65, 7)\n(65, 8)\n(65, 9)\n(65, 10)\n(65, 11)\n(65, 12)\n(65, 13)\n(65, 14)\n(65, 15)\n(65, 16)\n(65, 17)\n(65, 18)\n(65, 19)\n(65, 20)\n(65, 21)\n(65, 22)\n(65, 23)\n(65, 24)\n(65, 25)\n(65, 26)\n(65, 27)\n(65, 28)\n(65, 29)\n(65, 30)\n(65, 31)\n(65, 32)\n(65, 33)\n(65, 34)\n(65, 35)\n(65, 36)\n(65, 37)\n(65, 38)\n(65, 39)\n(65, 40)\n(65, 41)\n(65, 42)\n(65, 43)\n(65, 44)\n(65, 45)\n(65, 46)\n(65, 47)\n(65, 48)\n(65, 49)\n(65, 50)\n(65, 51)\n(65, 52)\n(65, 53)\n(65, 54)\n(65, 55)\n(65, 56)\n(65, 57)\n(65, 58)\n(65, 59)\n(65, 60)\n(65, 61)\n(65, 62)\n(65, 63)\n(65, 64)\n(65, 65)\n(65, 66)\n(65, 67)\n(65, 68)\n(65, 69)\n(65, 70)\n(65, 71)\n(65, 72)\n(65, 73)\n(65, 74)\n(65, 75)\n(65, 76)\n(65, 77)\n(65, 78)\n(65, 79)\n(65, 80)\n(65, 81)\n(65, 82)\n(65, 83)\n(65, 84)\n(65, 85)\n(65, 86)\n(65, 87)\n(65, 88)\n(65, 89)\n(65, 90)\n(65, 91)\n(65, 92)\n(65, 93)\n(65, 94)\n(65, 95)\n(65, 96)\n(65, 97)\n(65, 98)\n(65, 99)\n(66, 0)\n(66, 1)\n(66, 2)\n(66, 3)\n(66, 4)\n(66, 5)\n(66, 6)\n(66, 7)\n(66, 8)\n(66, 9)\n(66, 10)\n(66, 11)\n(66, 12)\n(66, 13)\n(66, 14)\n(66, 15)\n(66, 16)\n(66, 17)\n(66, 18)\n(66, 19)\n(66, 20)\n(66, 21)\n(66, 22)\n(66, 23)\n(66, 24)\n(66, 25)\n(66, 26)\n(66, 27)\n(66, 28)\n(66, 29)\n(66, 30)\n(66, 31)\n(66, 32)\n(66, 33)\n(66, 34)\n(66, 35)\n(66, 36)\n(66, 37)\n(66, 38)\n(66, 39)\n(66, 40)\n(66, 41)\n(66, 42)\n(66, 43)\n(66, 44)\n(66, 45)\n(66, 46)\n(66, 47)\n(66, 48)\n(66, 49)\n(66, 50)\n(66, 51)\n(66, 52)\n(66, 53)\n(66, 54)\n(66, 55)\n(66, 56)\n(66, 57)\n(66, 58)\n(66, 59)\n(66, 60)\n(66, 61)\n(66, 62)\n(66, 63)\n(66, 64)\n(66, 65)\n(66, 66)\n(66, 67)\n(66, 68)\n(66, 69)\n(66, 70)\n(66, 71)\n(66, 72)\n(66, 73)\n(66, 74)\n(66, 75)\n(66, 76)\n(66, 77)\n(66, 78)\n(66, 79)\n(66, 80)\n(66, 81)\n(66, 82)\n(66, 83)\n(66, 84)\n(66, 85)\n(66, 86)\n(66, 87)\n(66, 88)\n(66, 89)\n(66, 90)\n(66, 91)\n(66, 92)\n(66, 93)\n(66, 94)\n(66, 95)\n(66, 96)\n(66, 97)\n(66, 98)\n(66, 99)\n(67, 0)\n(67, 1)\n(67, 2)\n(67, 3)\n(67, 4)\n(67, 5)\n(67, 6)\n(67, 7)\n(67, 8)\n(67, 9)\n(67, 10)\n(67, 11)\n(67, 12)\n(67, 13)\n(67, 14)\n(67, 15)\n(67, 16)\n(67, 17)\n(67, 18)\n(67, 19)\n(67, 20)\n(67, 21)\n(67, 22)\n(67, 23)\n(67, 24)\n(67, 25)\n(67, 26)\n(67, 27)\n(67, 28)\n(67, 29)\n(67, 30)\n(67, 31)\n(67, 32)\n(67, 33)\n(67, 34)\n(67, 35)\n(67, 36)\n(67, 37)\n(67, 38)\n(67, 39)\n(67, 40)\n(67, 41)\n(67, 42)\n(67, 43)\n(67, 44)\n(67, 45)\n(67, 46)\n(67, 47)\n(67, 48)\n(67, 49)\n(67, 50)\n(67, 51)\n(67, 52)\n(67, 53)\n(67, 54)\n(67, 55)\n(67, 56)\n(67, 57)\n(67, 58)\n(67, 59)\n(67, 60)\n(67, 61)\n(67, 62)\n(67, 63)\n(67, 64)\n(67, 65)\n(67, 66)\n(67, 67)\n(67, 68)\n(67, 69)\n(67, 70)\n(67, 71)\n(67, 72)\n(67, 73)\n(67, 74)\n(67, 75)\n(67, 76)\n(67, 77)\n(67, 78)\n(67, 79)\n(67, 80)\n(67, 81)\n(67, 82)\n(67, 83)\n(67, 84)\n(67, 85)\n(67, 86)\n(67, 87)\n(67, 88)\n(67, 89)\n(67, 90)\n(67, 91)\n(67, 92)\n(67, 93)\n(67, 94)\n(67, 95)\n(67, 96)\n(67, 97)\n(67, 98)\n(67, 99)\n(68, 0)\n(68, 1)\n(68, 2)\n(68, 3)\n(68, 4)\n(68, 5)\n(68, 6)\n(68, 7)\n(68, 8)\n(68, 9)\n(68, 10)\n(68, 11)\n(68, 12)\n(68, 13)\n(68, 14)\n(68, 15)\n(68, 16)\n(68, 17)\n(68, 18)\n(68, 19)\n(68, 20)\n(68, 21)\n(68, 22)\n(68, 23)\n(68, 24)\n(68, 25)\n(68, 26)\n(68, 27)\n(68, 28)\n(68, 29)\n(68, 30)\n(68, 31)\n(68, 32)\n(68, 33)\n(68, 34)\n(68, 35)\n(68, 36)\n(68, 37)\n(68, 38)\n(68, 39)\n(68, 40)\n(68, 41)\n(68, 42)\n(68, 43)\n(68, 44)\n(68, 45)\n(68, 46)\n(68, 47)\n(68, 48)\n(68, 49)\n(68, 50)\n(68, 51)\n(68, 52)\n(68, 53)\n(68, 54)\n(68, 55)\n(68, 56)\n(68, 57)\n(68, 58)\n(68, 59)\n(68, 60)\n(68, 61)\n(68, 62)\n(68, 63)\n(68, 64)\n(68, 65)\n(68, 66)\n(68, 67)\n(68, 68)\n(68, 69)\n(68, 70)\n(68, 71)\n(68, 72)\n(68, 73)\n(68, 74)\n(68, 75)\n(68, 76)\n(68, 77)\n(68, 78)\n(68, 79)\n(68, 80)\n(68, 81)\n(68, 82)\n(68, 83)\n(68, 84)\n(68, 85)\n(68, 86)\n(68, 87)\n(68, 88)\n(68, 89)\n(68, 90)\n(68, 91)\n(68, 92)\n(68, 93)\n(68, 94)\n(68, 95)\n(68, 96)\n(68, 97)\n(68, 98)\n(68, 99)\n(69, 0)\n(69, 1)\n(69, 2)\n(69, 3)\n(69, 4)\n(69, 5)\n(69, 6)\n(69, 7)\n(69, 8)\n(69, 9)\n(69, 10)\n(69, 11)\n(69, 12)\n(69, 13)\n(69, 14)\n(69, 15)\n(69, 16)\n(69, 17)\n(69, 18)\n(69, 19)\n(69, 20)\n(69, 21)\n(69, 22)\n(69, 23)\n(69, 24)\n(69, 25)\n(69, 26)\n(69, 27)\n(69, 28)\n(69, 29)\n(69, 30)\n(69, 31)\n(69, 32)\n(69, 33)\n(69, 34)\n(69, 35)\n(69, 36)\n(69, 37)\n(69, 38)\n(69, 39)\n(69, 40)\n(69, 41)\n(69, 42)\n(69, 43)\n(69, 44)\n(69, 45)\n(69, 46)\n(69, 47)\n(69, 48)\n(69, 49)\n(69, 50)\n(69, 51)\n(69, 52)\n(69, 53)\n(69, 54)\n(69, 55)\n(69, 56)\n(69, 57)\n(69, 58)\n(69, 59)\n(69, 60)\n(69, 61)\n(69, 62)\n(69, 63)\n(69, 64)\n(69, 65)\n(69, 66)\n(69, 67)\n(69, 68)\n(69, 69)\n(69, 70)\n(69, 71)\n(69, 72)\n(69, 73)\n(69, 74)\n(69, 75)\n(69, 76)\n(69, 77)\n(69, 78)\n(69, 79)\n(69, 80)\n(69, 81)\n(69, 82)\n(69, 83)\n(69, 84)\n(69, 85)\n(69, 86)\n(69, 87)\n(69, 88)\n(69, 89)\n(69, 90)\n(69, 91)\n(69, 92)\n(69, 93)\n(69, 94)\n(69, 95)\n(69, 96)\n(69, 97)\n(69, 98)\n(69, 99)\n(70, 0)\n(70, 1)\n(70, 2)\n(70, 3)\n(70, 4)\n(70, 5)\n(70, 6)\n(70, 7)\n(70, 8)\n(70, 9)\n(70, 10)\n(70, 11)\n(70, 12)\n(70, 13)\n(70, 14)\n(70, 15)\n(70, 16)\n(70, 17)\n(70, 18)\n(70, 19)\n(70, 20)\n(70, 21)\n(70, 22)\n(70, 23)\n(70, 24)\n(70, 25)\n(70, 26)\n(70, 27)\n(70, 28)\n(70, 29)\n(70, 30)\n(70, 31)\n(70, 32)\n(70, 33)\n(70, 34)\n(70, 35)\n(70, 36)\n(70, 37)\n(70, 38)\n(70, 39)\n(70, 40)\n(70, 41)\n(70, 42)\n(70, 43)\n(70, 44)\n(70, 45)\n(70, 46)\n(70, 47)\n(70, 48)\n(70, 49)\n(70, 50)\n(70, 51)\n(70, 52)\n(70, 53)\n(70, 54)\n(70, 55)\n(70, 56)\n(70, 57)\n(70, 58)\n(70, 59)\n(70, 60)\n(70, 61)\n(70, 62)\n(70, 63)\n(70, 64)\n(70, 65)\n(70, 66)\n(70, 67)\n(70, 68)\n(70, 69)\n(70, 70)\n(70, 71)\n(70, 72)\n(70, 73)\n(70, 74)\n(70, 75)\n(70, 76)\n(70, 77)\n(70, 78)\n(70, 79)\n(70, 80)\n(70, 81)\n(70, 82)\n(70, 83)\n(70, 84)\n(70, 85)\n(70, 86)\n(70, 87)\n(70, 88)\n(70, 89)\n(70, 90)\n(70, 91)\n(70, 92)\n(70, 93)\n(70, 94)\n(70, 95)\n(70, 96)\n(70, 97)\n(70, 98)\n(70, 99)\n(71, 0)\n(71, 1)\n(71, 2)\n(71, 3)\n(71, 4)\n(71, 5)\n(71, 6)\n(71, 7)\n(71, 8)\n(71, 9)\n(71, 10)\n(71, 11)\n(71, 12)\n(71, 13)\n(71, 14)\n(71, 15)\n(71, 16)\n(71, 17)\n(71, 18)\n(71, 19)\n(71, 20)\n(71, 21)\n(71, 22)\n(71, 23)\n(71, 24)\n(71, 25)\n(71, 26)\n(71, 27)\n(71, 28)\n(71, 29)\n(71, 30)\n(71, 31)\n(71, 32)\n(71, 33)\n(71, 34)\n(71, 35)\n(71, 36)\n(71, 37)\n(71, 38)\n(71, 39)\n(71, 40)\n(71, 41)\n(71, 42)\n(71, 43)\n(71, 44)\n(71, 45)\n(71, 46)\n(71, 47)\n(71, 48)\n(71, 49)\n(71, 50)\n(71, 51)\n(71, 52)\n(71, 53)\n(71, 54)\n(71, 55)\n(71, 56)\n(71, 57)\n(71, 58)\n(71, 59)\n(71, 60)\n(71, 61)\n(71, 62)\n(71, 63)\n(71, 64)\n(71, 65)\n(71, 66)\n(71, 67)\n(71, 68)\n(71, 69)\n(71, 70)\n(71, 71)\n(71, 72)\n(71, 73)\n(71, 74)\n(71, 75)\n(71, 76)\n(71, 77)\n(71, 78)\n(71, 79)\n(71, 80)\n(71, 81)\n(71, 82)\n(71, 83)\n(71, 84)\n(71, 85)\n(71, 86)\n(71, 87)\n(71, 88)\n(71, 89)\n(71, 90)\n(71, 91)\n(71, 92)\n(71, 93)\n(71, 94)\n(71, 95)\n(71, 96)\n(71, 97)\n(71, 98)\n(71, 99)\n(72, 0)\n(72, 1)\n(72, 2)\n(72, 3)\n(72, 4)\n(72, 5)\n(72, 6)\n(72, 7)\n(72, 8)\n(72, 9)\n(72, 10)\n(72, 11)\n(72, 12)\n(72, 13)\n(72, 14)\n(72, 15)\n(72, 16)\n(72, 17)\n(72, 18)\n(72, 19)\n(72, 20)\n(72, 21)\n(72, 22)\n(72, 23)\n(72, 24)\n(72, 25)\n(72, 26)\n(72, 27)\n(72, 28)\n(72, 29)\n(72, 30)\n(72, 31)\n(72, 32)\n(72, 33)\n(72, 34)\n(72, 35)\n(72, 36)\n(72, 37)\n(72, 38)\n(72, 39)\n(72, 40)\n(72, 41)\n(72, 42)\n(72, 43)\n(72, 44)\n(72, 45)\n(72, 46)\n(72, 47)\n(72, 48)\n(72, 49)\n(72, 50)\n(72, 51)\n(72, 52)\n(72, 53)\n(72, 54)\n(72, 55)\n(72, 56)\n(72, 57)\n(72, 58)\n(72, 59)\n(72, 60)\n(72, 61)\n(72, 62)\n(72, 63)\n(72, 64)\n(72, 65)\n(72, 66)\n(72, 67)\n(72, 68)\n(72, 69)\n(72, 70)\n(72, 71)\n(72, 72)\n(72, 73)\n(72, 74)\n(72, 75)\n(72, 76)\n(72, 77)\n(72, 78)\n(72, 79)\n(72, 80)\n(72, 81)\n(72, 82)\n(72, 83)\n(72, 84)\n(72, 85)\n(72, 86)\n(72, 87)\n(72, 88)\n(72, 89)\n(72, 90)\n(72, 91)\n(72, 92)\n(72, 93)\n(72, 94)\n(72, 95)\n(72, 96)\n(72, 97)\n(72, 98)\n(72, 99)\n(73, 0)\n(73, 1)\n(73, 2)\n(73, 3)\n(73, 4)\n(73, 5)\n(73, 6)\n(73, 7)\n(73, 8)\n(73, 9)\n(73, 10)\n(73, 11)\n(73, 12)\n(73, 13)\n(73, 14)\n(73, 15)\n(73, 16)\n(73, 17)\n(73, 18)\n(73, 19)\n(73, 20)\n(73, 21)\n(73, 22)\n(73, 23)\n(73, 24)\n(73, 25)\n(73, 26)\n(73, 27)\n(73, 28)\n(73, 29)\n(73, 30)\n(73, 31)\n(73, 32)\n(73, 33)\n(73, 34)\n(73, 35)\n(73, 36)\n(73, 37)\n(73, 38)\n(73, 39)\n(73, 40)\n(73, 41)\n(73, 42)\n(73, 43)\n(73, 44)\n(73, 45)\n(73, 46)\n(73, 47)\n(73, 48)\n(73, 49)\n(73, 50)\n(73, 51)\n(73, 52)\n(73, 53)\n(73, 54)\n(73, 55)\n(73, 56)\n(73, 57)\n(73, 58)\n(73, 59)\n(73, 60)\n(73, 61)\n(73, 62)\n(73, 63)\n(73, 64)\n(73, 65)\n(73, 66)\n(73, 67)\n(73, 68)\n(73, 69)\n(73, 70)\n(73, 71)\n(73, 72)\n(73, 73)\n(73, 74)\n(73, 75)\n(73, 76)\n(73, 77)\n(73, 78)\n(73, 79)\n(73, 80)\n(73, 81)\n(73, 82)\n(73, 83)\n(73, 84)\n(73, 85)\n(73, 86)\n(73, 87)\n(73, 88)\n(73, 89)\n(73, 90)\n(73, 91)\n(73, 92)\n(73, 93)\n(73, 94)\n(73, 95)\n(73, 96)\n(73, 97)\n(73, 98)\n(73, 99)\n(74, 0)\n(74, 1)\n(74, 2)\n(74, 3)\n(74, 4)\n(74, 5)\n(74, 6)\n(74, 7)\n(74, 8)\n(74, 9)\n(74, 10)\n(74, 11)\n(74, 12)\n(74, 13)\n(74, 14)\n(74, 15)\n(74, 16)\n(74, 17)\n(74, 18)\n(74, 19)\n(74, 20)\n(74, 21)\n(74, 22)\n(74, 23)\n(74, 24)\n(74, 25)\n(74, 26)\n(74, 27)\n(74, 28)\n(74, 29)\n(74, 30)\n(74, 31)\n(74, 32)\n(74, 33)\n(74, 34)\n(74, 35)\n(74, 36)\n(74, 37)\n(74, 38)\n(74, 39)\n(74, 40)\n(74, 41)\n(74, 42)\n(74, 43)\n(74, 44)\n(74, 45)\n(74, 46)\n(74, 47)\n(74, 48)\n(74, 49)\n(74, 50)\n(74, 51)\n(74, 52)\n(74, 53)\n(74, 54)\n(74, 55)\n(74, 56)\n(74, 57)\n(74, 58)\n(74, 59)\n(74, 60)\n(74, 61)\n(74, 62)\n(74, 63)\n(74, 64)\n(74, 65)\n(74, 66)\n(74, 67)\n(74, 68)\n(74, 69)\n(74, 70)\n(74, 71)\n(74, 72)\n(74, 73)\n(74, 74)\n(74, 75)\n(74, 76)\n(74, 77)\n(74, 78)\n(74, 79)\n(74, 80)\n(74, 81)\n(74, 82)\n(74, 83)\n(74, 84)\n(74, 85)\n(74, 86)\n(74, 87)\n(74, 88)\n(74, 89)\n(74, 90)\n(74, 91)\n(74, 92)\n(74, 93)\n(74, 94)\n(74, 95)\n(74, 96)\n(74, 97)\n(74, 98)\n(74, 99)\n(75, 0)\n(75, 1)\n(75, 2)\n(75, 3)\n(75, 4)\n(75, 5)\n(75, 6)\n(75, 7)\n(75, 8)\n(75, 9)\n(75, 10)\n(75, 11)\n(75, 12)\n(75, 13)\n(75, 14)\n(75, 15)\n(75, 16)\n(75, 17)\n(75, 18)\n(75, 19)\n(75, 20)\n(75, 21)\n(75, 22)\n(75, 23)\n(75, 24)\n(75, 25)\n(75, 26)\n(75, 27)\n(75, 28)\n(75, 29)\n(75, 30)\n(75, 31)\n(75, 32)\n(75, 33)\n(75, 34)\n(75, 35)\n(75, 36)\n(75, 37)\n(75, 38)\n(75, 39)\n(75, 40)\n(75, 41)\n(75, 42)\n(75, 43)\n(75, 44)\n(75, 45)\n(75, 46)\n(75, 47)\n(75, 48)\n(75, 49)\n(75, 50)\n(75, 51)\n(75, 52)\n(75, 53)\n(75, 54)\n(75, 55)\n(75, 56)\n(75, 57)\n(75, 58)\n(75, 59)\n(75, 60)\n(75, 61)\n(75, 62)\n(75, 63)\n(75, 64)\n(75, 65)\n(75, 66)\n(75, 67)\n(75, 68)\n(75, 69)\n(75, 70)\n(75, 71)\n(75, 72)\n(75, 73)\n(75, 74)\n(75, 75)\n(75, 76)\n(75, 77)\n(75, 78)\n(75, 79)\n(75, 80)\n(75, 81)\n(75, 82)\n(75, 83)\n(75, 84)\n(75, 85)\n(75, 86)\n(75, 87)\n(75, 88)\n(75, 89)\n(75, 90)\n(75, 91)\n(75, 92)\n(75, 93)\n(75, 94)\n(75, 95)\n(75, 96)\n(75, 97)\n(75, 98)\n(75, 99)\n(76, 0)\n(76, 1)\n(76, 2)\n(76, 3)\n(76, 4)\n(76, 5)\n(76, 6)\n(76, 7)\n(76, 8)\n(76, 9)\n(76, 10)\n(76, 11)\n(76, 12)\n(76, 13)\n(76, 14)\n(76, 15)\n(76, 16)\n(76, 17)\n(76, 18)\n(76, 19)\n(76, 20)\n(76, 21)\n(76, 22)\n(76, 23)\n(76, 24)\n(76, 25)\n(76, 26)\n(76, 27)\n(76, 28)\n(76, 29)\n(76, 30)\n(76, 31)\n(76, 32)\n(76, 33)\n(76, 34)\n(76, 35)\n(76, 36)\n(76, 37)\n(76, 38)\n(76, 39)\n(76, 40)\n(76, 41)\n(76, 42)\n(76, 43)\n(76, 44)\n(76, 45)\n(76, 46)\n(76, 47)\n(76, 48)\n(76, 49)\n(76, 50)\n(76, 51)\n(76, 52)\n(76, 53)\n(76, 54)\n(76, 55)\n(76, 56)\n(76, 57)\n(76, 58)\n(76, 59)\n(76, 60)\n(76, 61)\n(76, 62)\n(76, 63)\n(76, 64)\n(76, 65)\n(76, 66)\n(76, 67)\n(76, 68)\n(76, 69)\n(76, 70)\n(76, 71)\n(76, 72)\n(76, 73)\n(76, 74)\n(76, 75)\n(76, 76)\n(76, 77)\n(76, 78)\n(76, 79)\n(76, 80)\n(76, 81)\n(76, 82)\n(76, 83)\n(76, 84)\n(76, 85)\n(76, 86)\n(76, 87)\n(76, 88)\n(76, 89)\n(76, 90)\n(76, 91)\n(76, 92)\n(76, 93)\n(76, 94)\n(76, 95)\n(76, 96)\n(76, 97)\n(76, 98)\n(76, 99)\n(77, 0)\n(77, 1)\n(77, 2)\n(77, 3)\n(77, 4)\n(77, 5)\n(77, 6)\n(77, 7)\n(77, 8)\n(77, 9)\n(77, 10)\n(77, 11)\n(77, 12)\n(77, 13)\n(77, 14)\n(77, 15)\n(77, 16)\n(77, 17)\n(77, 18)\n(77, 19)\n(77, 20)\n(77, 21)\n(77, 22)\n(77, 23)\n(77, 24)\n(77, 25)\n(77, 26)\n(77, 27)\n(77, 28)\n(77, 29)\n(77, 30)\n(77, 31)\n(77, 32)\n(77, 33)\n(77, 34)\n(77, 35)\n(77, 36)\n(77, 37)\n(77, 38)\n(77, 39)\n(77, 40)\n(77, 41)\n(77, 42)\n(77, 43)\n(77, 44)\n(77, 45)\n(77, 46)\n(77, 47)\n(77, 48)\n(77, 49)\n(77, 50)\n(77, 51)\n(77, 52)\n(77, 53)\n(77, 54)\n(77, 55)\n(77, 56)\n(77, 57)\n(77, 58)\n(77, 59)\n(77, 60)\n(77, 61)\n(77, 62)\n(77, 63)\n(77, 64)\n(77, 65)\n(77, 66)\n(77, 67)\n(77, 68)\n(77, 69)\n(77, 70)\n(77, 71)\n(77, 72)\n(77, 73)\n(77, 74)\n(77, 75)\n(77, 76)\n(77, 77)\n(77, 78)\n(77, 79)\n(77, 80)\n(77, 81)\n(77, 82)\n(77, 83)\n(77, 84)\n(77, 85)\n(77, 86)\n(77, 87)\n(77, 88)\n(77, 89)\n(77, 90)\n(77, 91)\n(77, 92)\n(77, 93)\n(77, 94)\n(77, 95)\n(77, 96)\n(77, 97)\n(77, 98)\n(77, 99)\n(78, 0)\n(78, 1)\n(78, 2)\n(78, 3)\n(78, 4)\n(78, 5)\n(78, 6)\n(78, 7)\n(78, 8)\n(78, 9)\n(78, 10)\n(78, 11)\n(78, 12)\n(78, 13)\n(78, 14)\n(78, 15)\n(78, 16)\n(78, 17)\n(78, 18)\n(78, 19)\n(78, 20)\n(78, 21)\n(78, 22)\n(78, 23)\n(78, 24)\n(78, 25)\n(78, 26)\n(78, 27)\n(78, 28)\n(78, 29)\n(78, 30)\n(78, 31)\n(78, 32)\n(78, 33)\n(78, 34)\n(78, 35)\n(78, 36)\n(78, 37)\n(78, 38)\n(78, 39)\n(78, 40)\n(78, 41)\n(78, 42)\n(78, 43)\n(78, 44)\n(78, 45)\n(78, 46)\n(78, 47)\n(78, 48)\n(78, 49)\n(78, 50)\n(78, 51)\n(78, 52)\n(78, 53)\n(78, 54)\n(78, 55)\n(78, 56)\n(78, 57)\n(78, 58)\n(78, 59)\n(78, 60)\n(78, 61)\n(78, 62)\n(78, 63)\n(78, 64)\n(78, 65)\n(78, 66)\n(78, 67)\n(78, 68)\n(78, 69)\n(78, 70)\n(78, 71)\n(78, 72)\n(78, 73)\n(78, 74)\n(78, 75)\n(78, 76)\n(78, 77)\n(78, 78)\n(78, 79)\n(78, 80)\n(78, 81)\n(78, 82)\n(78, 83)\n(78, 84)\n(78, 85)\n(78, 86)\n(78, 87)\n(78, 88)\n(78, 89)\n(78, 90)\n(78, 91)\n(78, 92)\n(78, 93)\n(78, 94)\n(78, 95)\n(78, 96)\n(78, 97)\n(78, 98)\n(78, 99)\n(79, 0)\n(79, 1)\n(79, 2)\n(79, 3)\n(79, 4)\n(79, 5)\n(79, 6)\n(79, 7)\n(79, 8)\n(79, 9)\n(79, 10)\n(79, 11)\n(79, 12)\n(79, 13)\n(79, 14)\n(79, 15)\n(79, 16)\n(79, 17)\n(79, 18)\n(79, 19)\n(79, 20)\n(79, 21)\n(79, 22)\n(79, 23)\n(79, 24)\n(79, 25)\n(79, 26)\n(79, 27)\n(79, 28)\n(79, 29)\n(79, 30)\n(79, 31)\n(79, 32)\n(79, 33)\n(79, 34)\n(79, 35)\n(79, 36)\n(79, 37)\n(79, 38)\n(79, 39)\n(79, 40)\n(79, 41)\n(79, 42)\n(79, 43)\n(79, 44)\n(79, 45)\n(79, 46)\n(79, 47)\n(79, 48)\n(79, 49)\n(79, 50)\n(79, 51)\n(79, 52)\n(79, 53)\n(79, 54)\n(79, 55)\n(79, 56)\n(79, 57)\n(79, 58)\n(79, 59)\n(79, 60)\n(79, 61)\n(79, 62)\n(79, 63)\n(79, 64)\n(79, 65)\n(79, 66)\n(79, 67)\n(79, 68)\n(79, 69)\n(79, 70)\n(79, 71)\n(79, 72)\n(79, 73)\n(79, 74)\n(79, 75)\n(79, 76)\n(79, 77)\n(79, 78)\n(79, 79)\n(79, 80)\n(79, 81)\n(79, 82)\n(79, 83)\n(79, 84)\n(79, 85)\n(79, 86)\n(79, 87)\n(79, 88)\n(79, 89)\n(79, 90)\n(79, 91)\n(79, 92)\n(79, 93)\n(79, 94)\n(79, 95)\n(79, 96)\n(79, 97)\n(79, 98)\n(79, 99)\n(80, 0)\n(80, 1)\n(80, 2)\n(80, 3)\n(80, 4)\n(80, 5)\n(80, 6)\n(80, 7)\n(80, 8)\n(80, 9)\n(80, 10)\n(80, 11)\n(80, 12)\n(80, 13)\n(80, 14)\n(80, 15)\n(80, 16)\n(80, 17)\n(80, 18)\n(80, 19)\n(80, 20)\n(80, 21)\n(80, 22)\n(80, 23)\n(80, 24)\n(80, 25)\n(80, 26)\n(80, 27)\n(80, 28)\n(80, 29)\n(80, 30)\n(80, 31)\n(80, 32)\n(80, 33)\n(80, 34)\n(80, 35)\n(80, 36)\n(80, 37)\n(80, 38)\n(80, 39)\n(80, 40)\n(80, 41)\n(80, 42)\n(80, 43)\n(80, 44)\n(80, 45)\n(80, 46)\n(80, 47)\n(80, 48)\n(80, 49)\n(80, 50)\n(80, 51)\n(80, 52)\n(80, 53)\n(80, 54)\n(80, 55)\n(80, 56)\n(80, 57)\n(80, 58)\n(80, 59)\n(80, 60)\n(80, 61)\n(80, 62)\n(80, 63)\n(80, 64)\n(80, 65)\n(80, 66)\n(80, 67)\n(80, 68)\n(80, 69)\n(80, 70)\n(80, 71)\n(80, 72)\n(80, 73)\n(80, 74)\n(80, 75)\n(80, 76)\n(80, 77)\n(80, 78)\n(80, 79)\n(80, 80)\n(80, 81)\n(80, 82)\n(80, 83)\n(80, 84)\n(80, 85)\n(80, 86)\n(80, 87)\n(80, 88)\n(80, 89)\n(80, 90)\n(80, 91)\n(80, 92)\n(80, 93)\n(80, 94)\n(80, 95)\n(80, 96)\n(80, 97)\n(80, 98)\n(80, 99)\n(81, 0)\n(81, 1)\n(81, 2)\n(81, 3)\n(81, 4)\n(81, 5)\n(81, 6)\n(81, 7)\n(81, 8)\n(81, 9)\n(81, 10)\n(81, 11)\n(81, 12)\n(81, 13)\n(81, 14)\n(81, 15)\n(81, 16)\n(81, 17)\n(81, 18)\n(81, 19)\n(81, 20)\n(81, 21)\n(81, 22)\n(81, 23)\n(81, 24)\n(81, 25)\n(81, 26)\n(81, 27)\n(81, 28)\n(81, 29)\n(81, 30)\n(81, 31)\n(81, 32)\n(81, 33)\n(81, 34)\n(81, 35)\n(81, 36)\n(81, 37)\n(81, 38)\n(81, 39)\n(81, 40)\n(81, 41)\n(81, 42)\n(81, 43)\n(81, 44)\n(81, 45)\n(81, 46)\n(81, 47)\n(81, 48)\n(81, 49)\n(81, 50)\n(81, 51)\n(81, 52)\n(81, 53)\n(81, 54)\n(81, 55)\n(81, 56)\n(81, 57)\n(81, 58)\n(81, 59)\n(81, 60)\n(81, 61)\n(81, 62)\n(81, 63)\n(81, 64)\n(81, 65)\n(81, 66)\n(81, 67)\n(81, 68)\n(81, 69)\n(81, 70)\n(81, 71)\n(81, 72)\n(81, 73)\n(81, 74)\n(81, 75)\n(81, 76)\n(81, 77)\n(81, 78)\n(81, 79)\n(81, 80)\n(81, 81)\n(81, 82)\n(81, 83)\n(81, 84)\n(81, 85)\n(81, 86)\n(81, 87)\n(81, 88)\n(81, 89)\n(81, 90)\n(81, 91)\n(81, 92)\n(81, 93)\n(81, 94)\n(81, 95)\n(81, 96)\n(81, 97)\n(81, 98)\n(81, 99)\n(82, 0)\n(82, 1)\n(82, 2)\n(82, 3)\n(82, 4)\n(82, 5)\n(82, 6)\n(82, 7)\n(82, 8)\n(82, 9)\n(82, 10)\n(82, 11)\n(82, 12)\n(82, 13)\n(82, 14)\n(82, 15)\n(82, 16)\n(82, 17)\n(82, 18)\n(82, 19)\n(82, 20)\n(82, 21)\n(82, 22)\n(82, 23)\n(82, 24)\n(82, 25)\n(82, 26)\n(82, 27)\n(82, 28)\n(82, 29)\n(82, 30)\n(82, 31)\n(82, 32)\n(82, 33)\n(82, 34)\n(82, 35)\n(82, 36)\n(82, 37)\n(82, 38)\n(82, 39)\n(82, 40)\n(82, 41)\n(82, 42)\n(82, 43)\n(82, 44)\n(82, 45)\n(82, 46)\n(82, 47)\n(82, 48)\n(82, 49)\n(82, 50)\n(82, 51)\n(82, 52)\n(82, 53)\n(82, 54)\n(82, 55)\n(82, 56)\n(82, 57)\n(82, 58)\n(82, 59)\n(82, 60)\n(82, 61)\n(82, 62)\n(82, 63)\n(82, 64)\n(82, 65)\n(82, 66)\n(82, 67)\n(82, 68)\n(82, 69)\n(82, 70)\n(82, 71)\n(82, 72)\n(82, 73)\n(82, 74)\n(82, 75)\n(82, 76)\n(82, 77)\n(82, 78)\n(82, 79)\n(82, 80)\n(82, 81)\n(82, 82)\n(82, 83)\n(82, 84)\n(82, 85)\n(82, 86)\n(82, 87)\n(82, 88)\n(82, 89)\n(82, 90)\n(82, 91)\n(82, 92)\n(82, 93)\n(82, 94)\n(82, 95)\n(82, 96)\n(82, 97)\n(82, 98)\n(82, 99)\n(83, 0)\n(83, 1)\n(83, 2)\n(83, 3)\n(83, 4)\n(83, 5)\n(83, 6)\n(83, 7)\n(83, 8)\n(83, 9)\n(83, 10)\n(83, 11)\n(83, 12)\n(83, 13)\n(83, 14)\n(83, 15)\n(83, 16)\n(83, 17)\n(83, 18)\n(83, 19)\n(83, 20)\n(83, 21)\n(83, 22)\n(83, 23)\n(83, 24)\n(83, 25)\n(83, 26)\n(83, 27)\n(83, 28)\n(83, 29)\n(83, 30)\n(83, 31)\n(83, 32)\n(83, 33)\n(83, 34)\n(83, 35)\n(83, 36)\n(83, 37)\n(83, 38)\n(83, 39)\n(83, 40)\n(83, 41)\n(83, 42)\n(83, 43)\n(83, 44)\n(83, 45)\n(83, 46)\n(83, 47)\n(83, 48)\n(83, 49)\n(83, 50)\n(83, 51)\n(83, 52)\n(83, 53)\n(83, 54)\n(83, 55)\n(83, 56)\n(83, 57)\n(83, 58)\n(83, 59)\n(83, 60)\n(83, 61)\n(83, 62)\n(83, 63)\n(83, 64)\n(83, 65)\n(83, 66)\n(83, 67)\n(83, 68)\n(83, 69)\n(83, 70)\n(83, 71)\n(83, 72)\n(83, 73)\n(83, 74)\n(83, 75)\n(83, 76)\n(83, 77)\n(83, 78)\n(83, 79)\n(83, 80)\n(83, 81)\n(83, 82)\n(83, 83)\n(83, 84)\n(83, 85)\n(83, 86)\n(83, 87)\n(83, 88)\n(83, 89)\n(83, 90)\n(83, 91)\n(83, 92)\n(83, 93)\n(83, 94)\n(83, 95)\n(83, 96)\n(83, 97)\n(83, 98)\n(83, 99)\n(84, 0)\n(84, 1)\n(84, 2)\n(84, 3)\n(84, 4)\n(84, 5)\n(84, 6)\n(84, 7)\n(84, 8)\n(84, 9)\n(84, 10)\n(84, 11)\n(84, 12)\n(84, 13)\n(84, 14)\n(84, 15)\n(84, 16)\n(84, 17)\n(84, 18)\n(84, 19)\n(84, 20)\n(84, 21)\n(84, 22)\n(84, 23)\n(84, 24)\n(84, 25)\n(84, 26)\n(84, 27)\n(84, 28)\n(84, 29)\n(84, 30)\n(84, 31)\n(84, 32)\n(84, 33)\n(84, 34)\n(84, 35)\n(84, 36)\n(84, 37)\n(84, 38)\n(84, 39)\n(84, 40)\n(84, 41)\n(84, 42)\n(84, 43)\n(84, 44)\n(84, 45)\n(84, 46)\n(84, 47)\n(84, 48)\n(84, 49)\n(84, 50)\n(84, 51)\n(84, 52)\n(84, 53)\n(84, 54)\n(84, 55)\n(84, 56)\n(84, 57)\n(84, 58)\n(84, 59)\n(84, 60)\n(84, 61)\n(84, 62)\n(84, 63)\n(84, 64)\n(84, 65)\n(84, 66)\n(84, 67)\n(84, 68)\n(84, 69)\n(84, 70)\n(84, 71)\n(84, 72)\n(84, 73)\n(84, 74)\n(84, 75)\n(84, 76)\n(84, 77)\n(84, 78)\n(84, 79)\n(84, 80)\n(84, 81)\n(84, 82)\n(84, 83)\n(84, 84)\n(84, 85)\n(84, 86)\n(84, 87)\n(84, 88)\n(84, 89)\n(84, 90)\n(84, 91)\n(84, 92)\n(84, 93)\n(84, 94)\n(84, 95)\n(84, 96)\n(84, 97)\n(84, 98)\n(84, 99)\n(85, 0)\n(85, 1)\n(85, 2)\n(85, 3)\n(85, 4)\n(85, 5)\n(85, 6)\n(85, 7)\n(85, 8)\n(85, 9)\n(85, 10)\n(85, 11)\n(85, 12)\n(85, 13)\n(85, 14)\n(85, 15)\n(85, 16)\n(85, 17)\n(85, 18)\n(85, 19)\n(85, 20)\n(85, 21)\n(85, 22)\n(85, 23)\n(85, 24)\n(85, 25)\n(85, 26)\n(85, 27)\n(85, 28)\n(85, 29)\n(85, 30)\n(85, 31)\n(85, 32)\n(85, 33)\n(85, 34)\n(85, 35)\n(85, 36)\n(85, 37)\n(85, 38)\n(85, 39)\n(85, 40)\n(85, 41)\n(85, 42)\n(85, 43)\n(85, 44)\n(85, 45)\n(85, 46)\n(85, 47)\n(85, 48)\n(85, 49)\n(85, 50)\n(85, 51)\n(85, 52)\n(85, 53)\n(85, 54)\n(85, 55)\n(85, 56)\n(85, 57)\n(85, 58)\n(85, 59)\n(85, 60)\n(85, 61)\n(85, 62)\n(85, 63)\n(85, 64)\n(85, 65)\n(85, 66)\n(85, 67)\n(85, 68)\n(85, 69)\n(85, 70)\n(85, 71)\n(85, 72)\n(85, 73)\n(85, 74)\n(85, 75)\n(85, 76)\n(85, 77)\n(85, 78)\n(85, 79)\n(85, 80)\n(85, 81)\n(85, 82)\n(85, 83)\n(85, 84)\n(85, 85)\n(85, 86)\n(85, 87)\n(85, 88)\n(85, 89)\n(85, 90)\n(85, 91)\n(85, 92)\n(85, 93)\n(85, 94)\n(85, 95)\n(85, 96)\n(85, 97)\n(85, 98)\n(85, 99)\n(86, 0)\n(86, 1)\n(86, 2)\n(86, 3)\n(86, 4)\n(86, 5)\n(86, 6)\n(86, 7)\n(86, 8)\n(86, 9)\n(86, 10)\n(86, 11)\n(86, 12)\n(86, 13)\n(86, 14)\n(86, 15)\n(86, 16)\n(86, 17)\n(86, 18)\n(86, 19)\n(86, 20)\n(86, 21)\n(86, 22)\n(86, 23)\n(86, 24)\n(86, 25)\n(86, 26)\n(86, 27)\n(86, 28)\n(86, 29)\n(86, 30)\n(86, 31)\n(86, 32)\n(86, 33)\n(86, 34)\n(86, 35)\n(86, 36)\n(86, 37)\n(86, 38)\n(86, 39)\n(86, 40)\n(86, 41)\n(86, 42)\n(86, 43)\n(86, 44)\n(86, 45)\n(86, 46)\n(86, 47)\n(86, 48)\n(86, 49)\n(86, 50)\n(86, 51)\n(86, 52)\n(86, 53)\n(86, 54)\n(86, 55)\n(86, 56)\n(86, 57)\n(86, 58)\n(86, 59)\n(86, 60)\n(86, 61)\n(86, 62)\n(86, 63)\n(86, 64)\n(86, 65)\n(86, 66)\n(86, 67)\n(86, 68)\n(86, 69)\n(86, 70)\n(86, 71)\n(86, 72)\n(86, 73)\n(86, 74)\n(86, 75)\n(86, 76)\n(86, 77)\n(86, 78)\n(86, 79)\n(86, 80)\n(86, 81)\n(86, 82)\n(86, 83)\n(86, 84)\n(86, 85)\n(86, 86)\n(86, 87)\n(86, 88)\n(86, 89)\n(86, 90)\n(86, 91)\n(86, 92)\n(86, 93)\n(86, 94)\n(86, 95)\n(86, 96)\n(86, 97)\n(86, 98)\n(86, 99)\n(87, 0)\n(87, 1)\n(87, 2)\n(87, 3)\n(87, 4)\n(87, 5)\n(87, 6)\n(87, 7)\n(87, 8)\n(87, 9)\n(87, 10)\n(87, 11)\n(87, 12)\n(87, 13)\n(87, 14)\n(87, 15)\n(87, 16)\n(87, 17)\n(87, 18)\n(87, 19)\n(87, 20)\n(87, 21)\n(87, 22)\n(87, 23)\n(87, 24)\n(87, 25)\n(87, 26)\n(87, 27)\n(87, 28)\n(87, 29)\n(87, 30)\n(87, 31)\n(87, 32)\n(87, 33)\n(87, 34)\n(87, 35)\n(87, 36)\n(87, 37)\n(87, 38)\n(87, 39)\n(87, 40)\n(87, 41)\n(87, 42)\n(87, 43)\n(87, 44)\n(87, 45)\n(87, 46)\n(87, 47)\n(87, 48)\n(87, 49)\n(87, 50)\n(87, 51)\n(87, 52)\n(87, 53)\n(87, 54)\n(87, 55)\n(87, 56)\n(87, 57)\n(87, 58)\n(87, 59)\n(87, 60)\n(87, 61)\n(87, 62)\n(87, 63)\n(87, 64)\n(87, 65)\n(87, 66)\n(87, 67)\n(87, 68)\n(87, 69)\n(87, 70)\n(87, 71)\n(87, 72)\n(87, 73)\n(87, 74)\n(87, 75)\n(87, 76)\n(87, 77)\n(87, 78)\n(87, 79)\n(87, 80)\n(87, 81)\n(87, 82)\n(87, 83)\n(87, 84)\n(87, 85)\n(87, 86)\n(87, 87)\n(87, 88)\n(87, 89)\n(87, 90)\n(87, 91)\n(87, 92)\n(87, 93)\n(87, 94)\n(87, 95)\n(87, 96)\n(87, 97)\n(87, 98)\n(87, 99)\n(88, 0)\n(88, 1)\n(88, 2)\n(88, 3)\n(88, 4)\n(88, 5)\n(88, 6)\n(88, 7)\n(88, 8)\n(88, 9)\n(88, 10)\n(88, 11)\n(88, 12)\n(88, 13)\n(88, 14)\n(88, 15)\n(88, 16)\n(88, 17)\n(88, 18)\n(88, 19)\n(88, 20)\n(88, 21)\n(88, 22)\n(88, 23)\n(88, 24)\n(88, 25)\n(88, 26)\n(88, 27)\n(88, 28)\n(88, 29)\n(88, 30)\n(88, 31)\n(88, 32)\n(88, 33)\n(88, 34)\n(88, 35)\n(88, 36)\n(88, 37)\n(88, 38)\n(88, 39)\n(88, 40)\n(88, 41)\n(88, 42)\n(88, 43)\n(88, 44)\n(88, 45)\n(88, 46)\n(88, 47)\n(88, 48)\n(88, 49)\n(88, 50)\n(88, 51)\n(88, 52)\n(88, 53)\n(88, 54)\n(88, 55)\n(88, 56)\n(88, 57)\n(88, 58)\n(88, 59)\n(88, 60)\n(88, 61)\n(88, 62)\n(88, 63)\n(88, 64)\n(88, 65)\n(88, 66)\n(88, 67)\n(88, 68)\n(88, 69)\n(88, 70)\n(88, 71)\n(88, 72)\n(88, 73)\n(88, 74)\n(88, 75)\n(88, 76)\n(88, 77)\n(88, 78)\n(88, 79)\n(88, 80)\n(88, 81)\n(88, 82)\n(88, 83)\n(88, 84)\n(88, 85)\n(88, 86)\n(88, 87)\n(88, 88)\n(88, 89)\n(88, 90)\n(88, 91)\n(88, 92)\n(88, 93)\n(88, 94)\n(88, 95)\n(88, 96)\n(88, 97)\n(88, 98)\n(88, 99)\n(89, 0)\n(89, 1)\n(89, 2)\n(89, 3)\n(89, 4)\n(89, 5)\n(89, 6)\n(89, 7)\n(89, 8)\n(89, 9)\n(89, 10)\n(89, 11)\n(89, 12)\n(89, 13)\n(89, 14)\n(89, 15)\n(89, 16)\n(89, 17)\n(89, 18)\n(89, 19)\n(89, 20)\n(89, 21)\n(89, 22)\n(89, 23)\n(89, 24)\n(89, 25)\n(89, 26)\n(89, 27)\n(89, 28)\n(89, 29)\n(89, 30)\n(89, 31)\n(89, 32)\n(89, 33)\n(89, 34)\n(89, 35)\n(89, 36)\n(89, 37)\n(89, 38)\n(89, 39)\n(89, 40)\n(89, 41)\n(89, 42)\n(89, 43)\n(89, 44)\n(89, 45)\n(89, 46)\n(89, 47)\n(89, 48)\n(89, 49)\n(89, 50)\n(89, 51)\n(89, 52)\n(89, 53)\n(89, 54)\n(89, 55)\n(89, 56)\n(89, 57)\n(89, 58)\n(89, 59)\n(89, 60)\n(89, 61)\n(89, 62)\n(89, 63)\n(89, 64)\n(89, 65)\n(89, 66)\n(89, 67)\n(89, 68)\n(89, 69)\n(89, 70)\n(89, 71)\n(89, 72)\n(89, 73)\n(89, 74)\n(89, 75)\n(89, 76)\n(89, 77)\n(89, 78)\n(89, 79)\n(89, 80)\n(89, 81)\n(89, 82)\n(89, 83)\n(89, 84)\n(89, 85)\n(89, 86)\n(89, 87)\n(89, 88)\n(89, 89)\n(89, 90)\n(89, 91)\n(89, 92)\n(89, 93)\n(89, 94)\n(89, 95)\n(89, 96)\n(89, 97)\n(89, 98)\n(89, 99)\n(90, 0)\n(90, 1)\n(90, 2)\n(90, 3)\n(90, 4)\n(90, 5)\n(90, 6)\n(90, 7)\n(90, 8)\n(90, 9)\n(90, 10)\n(90, 11)\n(90, 12)\n(90, 13)\n(90, 14)\n(90, 15)\n(90, 16)\n(90, 17)\n(90, 18)\n(90, 19)\n(90, 20)\n(90, 21)\n(90, 22)\n(90, 23)\n(90, 24)\n(90, 25)\n(90, 26)\n(90, 27)\n(90, 28)\n(90, 29)\n(90, 30)\n(90, 31)\n(90, 32)\n(90, 33)\n(90, 34)\n(90, 35)\n(90, 36)\n(90, 37)\n(90, 38)\n(90, 39)\n(90, 40)\n(90, 41)\n(90, 42)\n(90, 43)\n(90, 44)\n(90, 45)\n(90, 46)\n(90, 47)\n(90, 48)\n(90, 49)\n(90, 50)\n(90, 51)\n(90, 52)\n(90, 53)\n(90, 54)\n(90, 55)\n(90, 56)\n(90, 57)\n(90, 58)\n(90, 59)\n(90, 60)\n(90, 61)\n(90, 62)\n(90, 63)\n(90, 64)\n(90, 65)\n(90, 66)\n(90, 67)\n(90, 68)\n(90, 69)\n(90, 70)\n(90, 71)\n(90, 72)\n(90, 73)\n(90, 74)\n(90, 75)\n(90, 76)\n(90, 77)\n(90, 78)\n(90, 79)\n(90, 80)\n(90, 81)\n(90, 82)\n(90, 83)\n(90, 84)\n(90, 85)\n(90, 86)\n(90, 87)\n(90, 88)\n(90, 89)\n(90, 90)\n(90, 91)\n(90, 92)\n(90, 93)\n(90, 94)\n(90, 95)\n(90, 96)\n(90, 97)\n(90, 98)\n(90, 99)\n(91, 0)\n(91, 1)\n(91, 2)\n(91, 3)\n(91, 4)\n(91, 5)\n(91, 6)\n(91, 7)\n(91, 8)\n(91, 9)\n(91, 10)\n(91, 11)\n(91, 12)\n(91, 13)\n(91, 14)\n(91, 15)\n(91, 16)\n(91, 17)\n(91, 18)\n(91, 19)\n(91, 20)\n(91, 21)\n(91, 22)\n(91, 23)\n(91, 24)\n(91, 25)\n(91, 26)\n(91, 27)\n(91, 28)\n(91, 29)\n(91, 30)\n(91, 31)\n(91, 32)\n(91, 33)\n(91, 34)\n(91, 35)\n(91, 36)\n(91, 37)\n(91, 38)\n(91, 39)\n(91, 40)\n(91, 41)\n(91, 42)\n(91, 43)\n(91, 44)\n(91, 45)\n(91, 46)\n(91, 47)\n(91, 48)\n(91, 49)\n(91, 50)\n(91, 51)\n(91, 52)\n(91, 53)\n(91, 54)\n(91, 55)\n(91, 56)\n(91, 57)\n(91, 58)\n(91, 59)\n(91, 60)\n(91, 61)\n(91, 62)\n(91, 63)\n(91, 64)\n(91, 65)\n(91, 66)\n(91, 67)\n(91, 68)\n(91, 69)\n(91, 70)\n(91, 71)\n(91, 72)\n(91, 73)\n(91, 74)\n(91, 75)\n(91, 76)\n(91, 77)\n(91, 78)\n(91, 79)\n(91, 80)\n(91, 81)\n(91, 82)\n(91, 83)\n(91, 84)\n(91, 85)\n(91, 86)\n(91, 87)\n(91, 88)\n(91, 89)\n(91, 90)\n(91, 91)\n(91, 92)\n(91, 93)\n(91, 94)\n(91, 95)\n(91, 96)\n(91, 97)\n(91, 98)\n(91, 99)\n(92, 0)\n(92, 1)\n(92, 2)\n(92, 3)\n(92, 4)\n(92, 5)\n(92, 6)\n(92, 7)\n(92, 8)\n(92, 9)\n(92, 10)\n(92, 11)\n(92, 12)\n(92, 13)\n(92, 14)\n(92, 15)\n(92, 16)\n(92, 17)\n(92, 18)\n(92, 19)\n(92, 20)\n(92, 21)\n(92, 22)\n(92, 23)\n(92, 24)\n(92, 25)\n(92, 26)\n(92, 27)\n(92, 28)\n(92, 29)\n(92, 30)\n(92, 31)\n(92, 32)\n(92, 33)\n(92, 34)\n(92, 35)\n(92, 36)\n(92, 37)\n(92, 38)\n(92, 39)\n(92, 40)\n(92, 41)\n(92, 42)\n(92, 43)\n(92, 44)\n(92, 45)\n(92, 46)\n(92, 47)\n(92, 48)\n(92, 49)\n(92, 50)\n(92, 51)\n(92, 52)\n(92, 53)\n(92, 54)\n(92, 55)\n(92, 56)\n(92, 57)\n(92, 58)\n(92, 59)\n(92, 60)\n(92, 61)\n(92, 62)\n(92, 63)\n(92, 64)\n(92, 65)\n(92, 66)\n(92, 67)\n(92, 68)\n(92, 69)\n(92, 70)\n(92, 71)\n(92, 72)\n(92, 73)\n(92, 74)\n(92, 75)\n(92, 76)\n(92, 77)\n(92, 78)\n(92, 79)\n(92, 80)\n(92, 81)\n(92, 82)\n(92, 83)\n(92, 84)\n(92, 85)\n(92, 86)\n(92, 87)\n(92, 88)\n(92, 89)\n(92, 90)\n(92, 91)\n(92, 92)\n(92, 93)\n(92, 94)\n(92, 95)\n(92, 96)\n(92, 97)\n(92, 98)\n(92, 99)\n(93, 0)\n(93, 1)\n(93, 2)\n(93, 3)\n(93, 4)\n(93, 5)\n(93, 6)\n(93, 7)\n(93, 8)\n(93, 9)\n(93, 10)\n(93, 11)\n(93, 12)\n(93, 13)\n(93, 14)\n(93, 15)\n(93, 16)\n(93, 17)\n(93, 18)\n(93, 19)\n(93, 20)\n(93, 21)\n(93, 22)\n(93, 23)\n(93, 24)\n(93, 25)\n(93, 26)\n(93, 27)\n(93, 28)\n(93, 29)\n(93, 30)\n(93, 31)\n(93, 32)\n(93, 33)\n(93, 34)\n(93, 35)\n(93, 36)\n(93, 37)\n(93, 38)\n(93, 39)\n(93, 40)\n(93, 41)\n(93, 42)\n(93, 43)\n(93, 44)\n(93, 45)\n(93, 46)\n(93, 47)\n(93, 48)\n(93, 49)\n(93, 50)\n(93, 51)\n(93, 52)\n(93, 53)\n(93, 54)\n(93, 55)\n(93, 56)\n(93, 57)\n(93, 58)\n(93, 59)\n(93, 60)\n(93, 61)\n(93, 62)\n(93, 63)\n(93, 64)\n(93, 65)\n(93, 66)\n(93, 67)\n(93, 68)\n(93, 69)\n(93, 70)\n(93, 71)\n(93, 72)\n(93, 73)\n(93, 74)\n(93, 75)\n(93, 76)\n(93, 77)\n(93, 78)\n(93, 79)\n(93, 80)\n(93, 81)\n(93, 82)\n(93, 83)\n(93, 84)\n(93, 85)\n(93, 86)\n(93, 87)\n(93, 88)\n(93, 89)\n(93, 90)\n(93, 91)\n(93, 92)\n(93, 93)\n(93, 94)\n(93, 95)\n(93, 96)\n(93, 97)\n(93, 98)\n(93, 99)\n(94, 0)\n(94, 1)\n(94, 2)\n(94, 3)\n(94, 4)\n(94, 5)\n(94, 6)\n(94, 7)\n(94, 8)\n(94, 9)\n(94, 10)\n(94, 11)\n(94, 12)\n(94, 13)\n(94, 14)\n(94, 15)\n(94, 16)\n(94, 17)\n(94, 18)\n(94, 19)\n(94, 20)\n(94, 21)\n(94, 22)\n(94, 23)\n(94, 24)\n(94, 25)\n(94, 26)\n(94, 27)\n(94, 28)\n(94, 29)\n(94, 30)\n(94, 31)\n(94, 32)\n(94, 33)\n(94, 34)\n(94, 35)\n(94, 36)\n(94, 37)\n(94, 38)\n(94, 39)\n(94, 40)\n(94, 41)\n(94, 42)\n(94, 43)\n(94, 44)\n(94, 45)\n(94, 46)\n(94, 47)\n(94, 48)\n(94, 49)\n(94, 50)\n(94, 51)\n(94, 52)\n(94, 53)\n(94, 54)\n(94, 55)\n(94, 56)\n(94, 57)\n(94, 58)\n(94, 59)\n(94, 60)\n(94, 61)\n(94, 62)\n(94, 63)\n(94, 64)\n(94, 65)\n(94, 66)\n(94, 67)\n(94, 68)\n(94, 69)\n(94, 70)\n(94, 71)\n(94, 72)\n(94, 73)\n(94, 74)\n(94, 75)\n(94, 76)\n(94, 77)\n(94, 78)\n(94, 79)\n(94, 80)\n(94, 81)\n(94, 82)\n(94, 83)\n(94, 84)\n(94, 85)\n(94, 86)\n(94, 87)\n(94, 88)\n(94, 89)\n(94, 90)\n(94, 91)\n(94, 92)\n(94, 93)\n(94, 94)\n(94, 95)\n(94, 96)\n(94, 97)\n(94, 98)\n(94, 99)\n(95, 0)\n(95, 1)\n(95, 2)\n(95, 3)\n(95, 4)\n(95, 5)\n(95, 6)\n(95, 7)\n(95, 8)\n(95, 9)\n(95, 10)\n(95, 11)\n(95, 12)\n(95, 13)\n(95, 14)\n(95, 15)\n(95, 16)\n(95, 17)\n(95, 18)\n(95, 19)\n(95, 20)\n(95, 21)\n(95, 22)\n(95, 23)\n(95, 24)\n(95, 25)\n(95, 26)\n(95, 27)\n(95, 28)\n(95, 29)\n(95, 30)\n(95, 31)\n(95, 32)\n(95, 33)\n(95, 34)\n(95, 35)\n(95, 36)\n(95, 37)\n(95, 38)\n(95, 39)\n(95, 40)\n(95, 41)\n(95, 42)\n(95, 43)\n(95, 44)\n(95, 45)\n(95, 46)\n(95, 47)\n(95, 48)\n(95, 49)\n(95, 50)\n(95, 51)\n(95, 52)\n(95, 53)\n(95, 54)\n(95, 55)\n(95, 56)\n(95, 57)\n(95, 58)\n(95, 59)\n(95, 60)\n(95, 61)\n(95, 62)\n(95, 63)\n(95, 64)\n(95, 65)\n(95, 66)\n(95, 67)\n(95, 68)\n(95, 69)\n(95, 70)\n(95, 71)\n(95, 72)\n(95, 73)\n(95, 74)\n(95, 75)\n(95, 76)\n(95, 77)\n(95, 78)\n(95, 79)\n(95, 80)\n(95, 81)\n(95, 82)\n(95, 83)\n(95, 84)\n(95, 85)\n(95, 86)\n(95, 87)\n(95, 88)\n(95, 89)\n(95, 90)\n(95, 91)\n(95, 92)\n(95, 93)\n(95, 94)\n(95, 95)\n(95, 96)\n(95, 97)\n(95, 98)\n(95, 99)\n(96, 0)\n(96, 1)\n(96, 2)\n(96, 3)\n(96, 4)\n(96, 5)\n(96, 6)\n(96, 7)\n(96, 8)\n(96, 9)\n(96, 10)\n(96, 11)\n(96, 12)\n(96, 13)\n(96, 14)\n(96, 15)\n(96, 16)\n(96, 17)\n(96, 18)\n(96, 19)\n(96, 20)\n(96, 21)\n(96, 22)\n(96, 23)\n(96, 24)\n(96, 25)\n(96, 26)\n(96, 27)\n(96, 28)\n(96, 29)\n(96, 30)\n(96, 31)\n(96, 32)\n(96, 33)\n(96, 34)\n(96, 35)\n(96, 36)\n(96, 37)\n(96, 38)\n(96, 39)\n(96, 40)\n(96, 41)\n(96, 42)\n(96, 43)\n(96, 44)\n(96, 45)\n(96, 46)\n(96, 47)\n(96, 48)\n(96, 49)\n(96, 50)\n(96, 51)\n(96, 52)\n(96, 53)\n(96, 54)\n(96, 55)\n(96, 56)\n(96, 57)\n(96, 58)\n(96, 59)\n(96, 60)\n(96, 61)\n(96, 62)\n(96, 63)\n(96, 64)\n(96, 65)\n(96, 66)\n(96, 67)\n(96, 68)\n(96, 69)\n(96, 70)\n(96, 71)\n(96, 72)\n(96, 73)\n(96, 74)\n(96, 75)\n(96, 76)\n(96, 77)\n(96, 78)\n(96, 79)\n(96, 80)\n(96, 81)\n(96, 82)\n(96, 83)\n(96, 84)\n(96, 85)\n(96, 86)\n(96, 87)\n(96, 88)\n(96, 89)\n(96, 90)\n(96, 91)\n(96, 92)\n(96, 93)\n(96, 94)\n(96, 95)\n(96, 96)\n(96, 97)\n(96, 98)\n(96, 99)\n(97, 0)\n(97, 1)\n(97, 2)\n(97, 3)\n(97, 4)\n(97, 5)\n(97, 6)\n(97, 7)\n(97, 8)\n(97, 9)\n(97, 10)\n(97, 11)\n(97, 12)\n(97, 13)\n(97, 14)\n(97, 15)\n(97, 16)\n(97, 17)\n(97, 18)\n(97, 19)\n(97, 20)\n(97, 21)\n(97, 22)\n(97, 23)\n(97, 24)\n(97, 25)\n(97, 26)\n(97, 27)\n(97, 28)\n(97, 29)\n(97, 30)\n(97, 31)\n(97, 32)\n(97, 33)\n(97, 34)\n(97, 35)\n(97, 36)\n(97, 37)\n(97, 38)\n(97, 39)\n(97, 40)\n(97, 41)\n(97, 42)\n(97, 43)\n(97, 44)\n(97, 45)\n(97, 46)\n(97, 47)\n(97, 48)\n(97, 49)\n(97, 50)\n(97, 51)\n(97, 52)\n(97, 53)\n(97, 54)\n(97, 55)\n(97, 56)\n(97, 57)\n(97, 58)\n(97, 59)\n(97, 60)\n(97, 61)\n(97, 62)\n(97, 63)\n(97, 64)\n(97, 65)\n(97, 66)\n(97, 67)\n(97, 68)\n(97, 69)\n(97, 70)\n(97, 71)\n(97, 72)\n(97, 73)\n(97, 74)\n(97, 75)\n(97, 76)\n(97, 77)\n(97, 78)\n(97, 79)\n(97, 80)\n(97, 81)\n(97, 82)\n(97, 83)\n(97, 84)\n(97, 85)\n(97, 86)\n(97, 87)\n(97, 88)\n(97, 89)\n(97, 90)\n(97, 91)\n(97, 92)\n(97, 93)\n(97, 94)\n(97, 95)\n(97, 96)\n(97, 97)\n(97, 98)\n(97, 99)\n(98, 0)\n(98, 1)\n(98, 2)\n(98, 3)\n(98, 4)\n(98, 5)\n(98, 6)\n(98, 7)\n(98, 8)\n(98, 9)\n(98, 10)\n(98, 11)\n(98, 12)\n(98, 13)\n(98, 14)\n(98, 15)\n(98, 16)\n(98, 17)\n(98, 18)\n(98, 19)\n(98, 20)\n(98, 21)\n(98, 22)\n(98, 23)\n(98, 24)\n(98, 25)\n(98, 26)\n(98, 27)\n(98, 28)\n(98, 29)\n(98, 30)\n(98, 31)\n(98, 32)\n(98, 33)\n(98, 34)\n(98, 35)\n(98, 36)\n(98, 37)\n(98, 38)\n(98, 39)\n(98, 40)\n(98, 41)\n(98, 42)\n(98, 43)\n(98, 44)\n(98, 45)\n(98, 46)\n(98, 47)\n(98, 48)\n(98, 49)\n(98, 50)\n(98, 51)\n(98, 52)\n(98, 53)\n(98, 54)\n(98, 55)\n(98, 56)\n(98, 57)\n(98, 58)\n(98, 59)\n(98, 60)\n(98, 61)\n(98, 62)\n(98, 63)\n(98, 64)\n(98, 65)\n(98, 66)\n(98, 67)\n(98, 68)\n(98, 69)\n(98, 70)\n(98, 71)\n(98, 72)\n(98, 73)\n(98, 74)\n(98, 75)\n(98, 76)\n(98, 77)\n(98, 78)\n(98, 79)\n(98, 80)\n(98, 81)\n(98, 82)\n(98, 83)\n(98, 84)\n(98, 85)\n(98, 86)\n(98, 87)\n(98, 88)\n(98, 89)\n(98, 90)\n(98, 91)\n(98, 92)\n(98, 93)\n(98, 94)\n(98, 95)\n(98, 96)\n(98, 97)\n(98, 98)\n(98, 99)\n(99, 0)\n(99, 1)\n(99, 2)\n(99, 3)\n(99, 4)\n(99, 5)\n(99, 6)\n(99, 7)\n(99, 8)\n(99, 9)\n(99, 10)\n(99, 11)\n(99, 12)\n(99, 13)\n(99, 14)\n(99, 15)\n(99, 16)\n(99, 17)\n(99, 18)\n(99, 19)\n(99, 20)\n(99, 21)\n(99, 22)\n(99, 23)\n(99, 24)\n(99, 25)\n(99, 26)\n(99, 27)\n(99, 28)\n(99, 29)\n(99, 30)\n(99, 31)\n(99, 32)\n(99, 33)\n(99, 34)\n(99, 35)\n(99, 36)\n(99, 37)\n(99, 38)\n(99, 39)\n(99, 40)\n(99, 41)\n(99, 42)\n(99, 43)\n(99, 44)\n(99, 45)\n(99, 46)\n(99, 47)\n(99, 48)\n(99, 49)\n(99, 50)\n(99, 51)\n(99, 52)\n(99, 53)\n(99, 54)\n(99, 55)\n(99, 56)\n(99, 57)\n(99, 58)\n(99, 59)\n(99, 60)\n(99, 61)\n(99, 62)\n(99, 63)\n(99, 64)\n(99, 65)\n(99, 66)\n(99, 67)\n(99, 68)\n(99, 69)\n(99, 70)\n(99, 71)\n(99, 72)\n(99, 73)\n(99, 74)\n(99, 75)\n(99, 76)\n(99, 77)\n(99, 78)\n(99, 79)\n(99, 80)\n(99, 81)\n(99, 82)\n(99, 83)\n(99, 84)\n(99, 85)\n(99, 86)\n(99, 87)\n(99, 88)\n(99, 89)\n(99, 90)\n(99, 91)\n(99, 92)\n(99, 93)\n(99, 94)\n(99, 95)\n(99, 96)\n(99, 97)\n(99, 98)\n(99, 99)\n"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acf6f0ce92cd143a292244c7593d8ba32abb374
| 57,727 |
ipynb
|
Jupyter Notebook
|
notebooks/retrieve_poetry.ipynb
|
M-Puig/Erato
|
97f7cf913c54aacc55bbc76f4eadc55641a44c2d
|
[
"MIT"
] | null | null | null |
notebooks/retrieve_poetry.ipynb
|
M-Puig/Erato
|
97f7cf913c54aacc55bbc76f4eadc55641a44c2d
|
[
"MIT"
] | null | null | null |
notebooks/retrieve_poetry.ipynb
|
M-Puig/Erato
|
97f7cf913c54aacc55bbc76f4eadc55641a44c2d
|
[
"MIT"
] | null | null | null | 48.510084 | 1,642 | 0.567291 |
[
[
[
"# Retrieve Poetry\n## Poetry Retriever using the Poly-encoder Transformer architecture (Humeau et al., 2019) for retrieval",
"_____no_output_____"
]
],
[
[
"# This notebook is based on :\n# https://aritter.github.io/CS-7650/\n# This Project was developed at the Georgia Institute of Technology by Ashutosh Baheti ([email protected]), \n# borrowing from the Neural Machine Translation Project (Project 2) \n# of the UC Berkeley NLP course https://cal-cs288.github.io/sp20/",
"_____no_output_____"
],
[
"from __future__ import absolute_import\nfrom __future__ import division\nfrom __future__ import print_function\nfrom __future__ import unicode_literals\n\nimport torch\nfrom torch.jit import script, trace\nimport torch.nn as nn\nfrom torch import optim\nimport torch.nn.functional as F\nimport numpy as np\nimport csv\nimport random\nimport re\nimport os\nimport unicodedata\nimport codecs\nfrom io import open\nimport itertools\nimport math\nimport pickle\nimport statistics\nimport sys\nfrom functools import partial\n\nfrom torch.utils.data import Dataset, DataLoader\nfrom torch.nn.utils.rnn import pad_sequence\nimport tqdm\nimport nltk\n#from google.colab import files",
"_____no_output_____"
],
[
"# General util functions\ndef make_dir_if_not_exists(directory):\n\tif not os.path.exists(directory):\n\t\tlogging.info(\"Creating new directory: {}\".format(directory))\n\t\tos.makedirs(directory)\n\ndef print_list(l, K=None):\n\t# If K is given then only print first K\n\tfor i, e in enumerate(l):\n\t\tif i == K:\n\t\t\tbreak\n\t\tprint(e)\n\tprint()\n\ndef remove_multiple_spaces(string):\n\treturn re.sub(r'\\s+', ' ', string).strip()\n\ndef save_in_pickle(save_object, save_file):\n\twith open(save_file, \"wb\") as pickle_out:\n\t\tpickle.dump(save_object, pickle_out)\n\ndef load_from_pickle(pickle_file):\n\twith open(pickle_file, \"rb\") as pickle_in:\n\t\treturn pickle.load(pickle_in)\n\ndef save_in_txt(list_of_strings, save_file):\n\twith open(save_file, \"w\") as writer:\n\t\tfor line in list_of_strings:\n\t\t\tline = line.strip()\n\t\t\twriter.write(f\"{line}\\n\")\n\ndef load_from_txt(txt_file):\n\twith open(txt_file, \"r\") as reader:\n\t\tall_lines = list()\n\t\tfor line in reader:\n\t\t\tline = line.strip()\n\t\t\tall_lines.append(line)\n\t\treturn all_lines",
"_____no_output_____"
],
[
"import pandas as pd\n\nprint(torch.cuda.is_available())\nif torch.cuda.is_available():\n device = torch.device(\"cuda\")\nelse:\n device = torch.device(\"cpu\")\nprint(\"Using device:\", device)",
"True\nUsing device: cuda\n"
],
[
"bert_model_name = 'distilbert-base-uncased' \n# Bert Imports\nfrom transformers import DistilBertTokenizer, DistilBertModel\n#bert_model = DistilBertModel.from_pretrained(bert_model_name)\ntokenizer = DistilBertTokenizer.from_pretrained(bert_model_name)",
"_____no_output_____"
]
],
[
[
"## Load Data",
"_____no_output_____"
],
[
"### Poetry Database",
"_____no_output_____"
]
],
[
[
"data_file = '../data/with_epoque.csv'\ndata = pd.read_csv(data_file)\nprint(len(data))\nprint(data.head())",
"573\n author \\\n0 WILLIAM SHAKESPEARE \n1 DUCHESS OF NEWCASTLE MARGARET CAVENDISH \n2 THOMAS BASTARD \n3 EDMUND SPENSER \n4 RICHARD BARNFIELD \n\n content \\\n0 Let the bird of loudest lay\\nOn the sole Arabi... \n1 Sir Charles into my chamber coming in,\\nWhen I... \n2 Our vice runs beyond all that old men saw,\\nAn... \n3 Lo I the man, whose Muse whilome did maske,\\nA... \n4 Long have I longd to see my love againe,\\nStil... \n\n poem name age type \n0 The Phoenix and the Turtle Renaissance Mythology & Folklore \n1 An Epilogue to the Above Renaissance Mythology & Folklore \n2 Book 7, Epigram 42 Renaissance Mythology & Folklore \n3 from The Faerie Queene: Book I, Canto I Renaissance Mythology & Folklore \n4 Sonnet 16 Renaissance Mythology & Folklore \n"
]
],
[
[
"## Dataset Preparation",
"_____no_output_____"
]
],
[
[
"def make_data_training(df, char_max_line = 20):\n inputs = []\n context = []\n targets = []\n for i,rows in df.iterrows():\n splitted = rows['content'].split('\\r\\n')\n for line in splitted:\n if len(line.strip()) > 0 and len(line.split(' ')) <= char_max_line:\n inputs.append(line)\n targets.append(line)\n context.append(' '.join([str(rows['poem name'])]))\n \n return pd.DataFrame(list(zip(inputs, context, targets)),columns =['text', 'context','target'])\n\n\n#Defining torch dataset class for poems\nclass PoemDataset(Dataset):\n def __init__(self, df):\n self.df = df\n\n def __len__(self):\n return len(self.df)\n\n def __getitem__(self, idx):\n return self.df.iloc[idx]",
"_____no_output_____"
],
[
"df = make_data_training(data, char_max_line = 30)",
"_____no_output_____"
],
[
"pad_word = \"<pad>\"\nbos_word = \"<bos>\"\neos_word = \"<eos>\"\nunk_word = \"<unk>\"\nsep_word = \"sep\"\n\npad_id = 0\nbos_id = 1\neos_id = 2\nunk_id = 3\nsep_id = 4\n \ndef normalize_sentence(s):\n s = re.sub(r\"([.!?])\", r\" \\1\", s)\n s = re.sub(r\"[^a-zA-Z.!?]+\", r\" \", s)\n s = re.sub(r\"\\s+\", r\" \", s).strip()\n return s\n\nclass Vocabulary:\n def __init__(self):\n self.word_to_id = {pad_word: pad_id, bos_word: bos_id, eos_word:eos_id, unk_word: unk_id, sep_word: sep_id}\n self.word_count = {}\n self.id_to_word = {pad_id: pad_word, bos_id: bos_word, eos_id: eos_word, unk_id: unk_word, sep_id: sep_word}\n self.num_words = 5\n \n def get_ids_from_sentence(self, sentence):\n sentence = normalize_sentence(sentence)\n sent_ids = [bos_id] + [self.word_to_id[word.lower()] if word.lower() in self.word_to_id \\\n else unk_id for word in sentence.split()] + \\\n [eos_id]\n return sent_ids\n \n def tokenized_sentence(self, sentence):\n sent_ids = self.get_ids_from_sentence(sentence)\n return [self.id_to_word[word_id] for word_id in sent_ids]\n\n def decode_sentence_from_ids(self, sent_ids):\n words = list()\n for i, word_id in enumerate(sent_ids):\n if word_id in [bos_id, eos_id, pad_id]:\n # Skip these words\n continue\n else:\n words.append(self.id_to_word[word_id])\n return ' '.join(words)\n\n def add_words_from_sentence(self, sentence):\n sentence = normalize_sentence(sentence)\n for word in sentence.split():\n if word not in self.word_to_id:\n # add this word to the vocabulary\n self.word_to_id[word] = self.num_words\n self.id_to_word[self.num_words] = word\n self.word_count[word] = 1\n self.num_words += 1\n else:\n # update the word count\n self.word_count[word] += 1\n\nvocab = Vocabulary()\nfor src in df['text']:\n vocab.add_words_from_sentence(src.lower())\n\nprint(f\"Total words in the vocabulary = {vocab.num_words}\")",
"Total words in the vocabulary = 319\n"
],
[
"class Poem_dataset(Dataset):\n \"\"\"Single-Turn version of Cornell Movie Dialog Cropus dataset.\"\"\"\n\n def __init__(self, poems, context,vocab, device):\n \"\"\"\n Args:\n conversations: list of tuple (src_string, tgt_string) \n - src_string: String of the source sentence\n - tgt_string: String of the target sentence\n vocab: Vocabulary object that contains the mapping of \n words to indices\n device: cpu or cuda\n \"\"\"\n l = []\n \n for i in range(len(poems)):\n l.append( ( context[i] + ' sep ' + poems[i] , poems[i] ))\n \n self.conversations = l.copy()\n self.vocab = vocab\n self.device = device\n\n def encode(src, tgt):\n src_ids = self.vocab.get_ids_from_sentence(src)\n tgt_ids = self.vocab.get_ids_from_sentence(tgt)\n return (src_ids, tgt_ids)\n\n # We will pre-tokenize the conversations and save in id lists for later use\n self.tokenized_conversations = [encode(src, tgt) for src, tgt in self.conversations]\n \n def __len__(self):\n return len(self.conversations)\n\n def __getitem__(self, idx):\n if torch.is_tensor(idx):\n idx = idx.tolist()\n\n return {\"conv_ids\":self.tokenized_conversations[idx], \"conv\":self.conversations[idx]}\n\ndef collate_fn(data):\n \"\"\"Creates mini-batch tensors from the list of tuples (src_seq, tgt_seq).\n We should build a custom collate_fn rather than using default collate_fn,\n because merging sequences (including padding) is not supported in default.\n Seqeuences are padded to the maximum length of mini-batch sequences (dynamic padding).\n Args:\n data: list of dicts {\"conv_ids\":(src_ids, tgt_ids), \"conv\":(src_str, trg_str)}.\n - src_ids: list of src piece ids; variable length.\n - tgt_ids: list of tgt piece ids; variable length.\n - src_str: String of src\n - tgt_str: String of tgt\n Returns: dict { \"conv_ids\": (src_ids, tgt_ids), \n \"conv\": (src_str, tgt_str), \n \"conv_tensors\": (src_seqs, tgt_seqs)}\n src_seqs: torch tensor of shape (src_padded_length, batch_size).\n tgt_seqs: torch tensor of shape (tgt_padded_length, batch_size).\n src_padded_length = length of the longest src sequence from src_ids\n tgt_padded_length = length of the longest tgt sequence from tgt_ids\n \"\"\"\n # Sort conv_ids based on decreasing order of the src_lengths.\n # This is required for efficient GPU computations.\n src_ids = [torch.LongTensor(e[\"conv_ids\"][0]) for e in data]\n tgt_ids = [torch.LongTensor(e[\"conv_ids\"][1]) for e in data]\n src_str = [e[\"conv\"][0] for e in data]\n tgt_str = [e[\"conv\"][1] for e in data]\n data = list(zip(src_ids, tgt_ids, src_str, tgt_str))\n data.sort(key=lambda x: len(x[0]), reverse=True)\n src_ids, tgt_ids, src_str, tgt_str = zip(*data)\n\n\n # Pad the src_ids and tgt_ids using token pad_id to create src_seqs and tgt_seqs\n \n # Implementation tip: You can use the nn.utils.rnn.pad_sequence utility\n # function to combine a list of variable-length sequences with padding.\n \n # YOUR CODE HERE\n src_seqs = nn.utils.rnn.pad_sequence(src_ids, padding_value = pad_id,\n batch_first = False)\n tgt_seqs = nn.utils.rnn.pad_sequence(tgt_ids, padding_value = pad_id, \n batch_first = False)\n \n src_padded_length = len(src_seqs[0])\n tgt_padded_length = len(tgt_seqs[0])\n return {\"conv_ids\":(src_ids, tgt_ids), \"conv\":(src_str, tgt_str), \"conv_tensors\":(src_seqs.to(device), tgt_seqs.to(device))}",
"_____no_output_____"
],
[
"# Create the DataLoader for all_conversations\n\nall_poems = df['text'].tolist()\ncontext = df['context'].tolist()\n\ndataset = Poem_dataset(all_poems, context, vocab, device)\n\nbatch_size = 5\n\ndata_loader = DataLoader(dataset=dataset, batch_size=batch_size, \n shuffle=True, collate_fn=collate_fn)",
"_____no_output_____"
],
[
"\nfor src, tgt in dataset.conversations[:3]:\n sentence = src\n word_tokens = vocab.tokenized_sentence(sentence)\n # Automatically adds bos_id and eos_id before and after sentence ids respectively\n word_ids = vocab.get_ids_from_sentence(sentence)\n print(sentence)\n print(word_tokens)\n print(word_ids)\n print(vocab.decode_sentence_from_ids(word_ids))\n print()\n\nword = \"the\"\nword_id = vocab.word_to_id[word.lower()]\nprint(f\"Word = {word}\")\nprint(f\"Word ID = {word_id}\")\nprint(f\"Word decoded from ID = {vocab.decode_sentence_from_ids([word_id])}\")",
"Written in her French Psalter sep No crooked leg, no bleared eye,\nNo part deformed out of kind,\nNor yet so ugly half can be\nAs is the inward suspicious mind.\n['<bos>', '<unk>', 'in', 'her', '<unk>', '<unk>', 'sep', 'no', 'crooked', 'leg', 'no', 'bleared', 'eye', 'no', 'part', 'deformed', 'out', 'of', 'kind', 'nor', 'yet', 'so', 'ugly', 'half', 'can', 'be', 'as', 'is', 'the', 'inward', 'suspicious', 'mind', '.', '<eos>']\n[1, 3, 89, 232, 3, 3, 4, 5, 6, 7, 5, 8, 9, 5, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 2]\n<unk> in her <unk> <unk> sep no crooked leg no bleared eye no part deformed out of kind nor yet so ugly half can be as is the inward suspicious mind .\n\nSong of the Witches: Double, double toil and trouble sep Notes:\nMacbeth: IV.i 10-19; 35-38\n['<bos>', 'song', 'of', 'the', '<unk>', '<unk>', '<unk>', '<unk>', 'and', '<unk>', 'sep', 'notes', 'macbeth', 'iv', '.i', '<eos>']\n[1, 281, 13, 24, 3, 3, 3, 3, 45, 3, 4, 29, 30, 31, 32, 2]\nsong of the <unk> <unk> <unk> <unk> and <unk> sep notes macbeth iv .i\n\nAmoretti IV: \"New yeare forth looking out of Janus gate\" sep New yeare forth looking out of Janus gate,\nDoth seeme to promise hope of new delight:\nAnd bidding thold Adieu, his pass\n['<bos>', '<unk>', 'iv', 'new', 'yeare', 'forth', 'looking', 'out', 'of', 'janus', 'gate', 'sep', 'new', 'yeare', 'forth', 'looking', 'out', 'of', 'janus', 'gate', 'doth', 'seeme', 'to', 'promise', 'hope', 'of', 'new', 'delight', 'and', 'bidding', 'thold', 'adieu', 'his', 'pass', '<eos>']\n[1, 3, 31, 33, 34, 35, 36, 12, 13, 37, 38, 4, 33, 34, 35, 36, 12, 13, 37, 38, 39, 40, 41, 42, 43, 13, 33, 44, 45, 46, 47, 48, 49, 50, 2]\n<unk> iv new yeare forth looking out of janus gate sep new yeare forth looking out of janus gate doth seeme to promise hope of new delight and bidding thold adieu his pass\n\nWord = the\nWord ID = 24\nWord decoded from ID = the\n"
],
[
"# Test one batch of training data\nfirst_batch = next(iter(data_loader))\nprint(f\"Testing first training batch of size {len(first_batch['conv'][0])}\")\nprint(f\"List of source strings:\")\nprint_list(first_batch[\"conv\"][0])\nprint(f\"Tokenized source ids:\")\nprint_list(first_batch[\"conv_ids\"][0])\nprint(f\"Padded source ids as tensor (shape {first_batch['conv_tensors'][0].size()}):\")\nprint(first_batch[\"conv_tensors\"][0])",
"Testing first training batch of size 5\nList of source strings:\nThe Poem that Took the Place of a Mountain sep Wallace Stevens, \"The Poem that Took the Place of a Mountain\" from The Collected Poems. Copyright 1954 by Wallace Stevens. Reprinted by permission of Random House, Inc.\nArs Poetica sep Archibald MacLeish, Ars Poetica from Collected Poems 1917-1982. Copyright 1985 by The Estate of Archibald MacLeish. Reprinted with the permission of Houghton Mifflin Company. All rights reserved.\nWritten in her French Psalter sep No crooked leg, no bleared eye,\nNo part deformed out of kind,\nNor yet so ugly half can be\nAs is the inward suspicious mind.\nThe Eemis Stane sep Hugh MacDiarmid, The Eemis Stane from Selected Poetry. Copyright 1992 by Alan Riach and Michael Grieve. Reprinted with the permission of New Directions Publishing Corporation.\nTo a Dead Lover sep Originally published in Poetry, August 1922.\n\nTokenized source ids:\ntensor([ 1, 24, 109, 110, 111, 24, 112, 13, 90, 113, 4, 107, 108, 24,\n 109, 110, 111, 24, 112, 13, 90, 113, 55, 24, 114, 73, 28, 57,\n 58, 107, 108, 28, 59, 58, 60, 13, 115, 116, 117, 28, 2])\ntensor([ 1, 227, 228, 4, 144, 145, 227, 228, 55, 114, 73, 28, 57, 58,\n 24, 147, 13, 144, 145, 28, 59, 92, 24, 60, 13, 148, 149, 150,\n 28, 84, 85, 86, 28, 2])\ntensor([ 1, 3, 89, 232, 3, 3, 4, 5, 6, 7, 5, 8, 9, 5,\n 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23,\n 24, 25, 26, 27, 28, 2])\ntensor([ 1, 24, 250, 251, 4, 99, 100, 24, 250, 251, 55, 94, 102, 28,\n 57, 58, 103, 104, 45, 105, 106, 28, 59, 92, 24, 60, 13, 33,\n 61, 62, 63, 28, 2])\ntensor([ 1, 41, 90, 3, 242, 4, 198, 95, 89, 102, 311, 28, 2])\n\nPadded source ids as tensor (shape torch.Size([41, 5])):\ntensor([[ 1, 1, 1, 1, 1],\n [ 24, 227, 3, 24, 41],\n [109, 228, 89, 250, 90],\n [110, 4, 232, 251, 3],\n [111, 144, 3, 4, 242],\n [ 24, 145, 3, 99, 4],\n [112, 227, 4, 100, 198],\n [ 13, 228, 5, 24, 95],\n [ 90, 55, 6, 250, 89],\n [113, 114, 7, 251, 102],\n [ 4, 73, 5, 55, 311],\n [107, 28, 8, 94, 28],\n [108, 57, 9, 102, 2],\n [ 24, 58, 5, 28, 0],\n [109, 24, 10, 57, 0],\n [110, 147, 11, 58, 0],\n [111, 13, 12, 103, 0],\n [ 24, 144, 13, 104, 0],\n [112, 145, 14, 45, 0],\n [ 13, 28, 15, 105, 0],\n [ 90, 59, 16, 106, 0],\n [113, 92, 17, 28, 0],\n [ 55, 24, 18, 59, 0],\n [ 24, 60, 19, 92, 0],\n [114, 13, 20, 24, 0],\n [ 73, 148, 21, 60, 0],\n [ 28, 149, 22, 13, 0],\n [ 57, 150, 23, 33, 0],\n [ 58, 28, 24, 61, 0],\n [107, 84, 25, 62, 0],\n [108, 85, 26, 63, 0],\n [ 28, 86, 27, 28, 0],\n [ 59, 28, 28, 2, 0],\n [ 58, 2, 2, 0, 0],\n [ 60, 0, 0, 0, 0],\n [ 13, 0, 0, 0, 0],\n [115, 0, 0, 0, 0],\n [116, 0, 0, 0, 0],\n [117, 0, 0, 0, 0],\n [ 28, 0, 0, 0, 0],\n [ 2, 0, 0, 0, 0]], device='cuda:0')\n"
],
[
"def transformer_collate_fn(batch, tokenizer):\n bert_vocab = tokenizer.get_vocab()\n bert_pad_token = bert_vocab['[PAD]']\n bert_unk_token = bert_vocab['[UNK]']\n bert_cls_token = bert_vocab['[CLS]']\n inputs, masks_input, outputs, masks_output = [], [], [], []\n\n sentences, masks_sentences, targets, masks_targets = [], [], [], []\n for data in batch:\n\n tokenizer_output = tokenizer([data['text']])\n tokenized_sent = tokenizer_output['input_ids'][0]\n \n tokenizer_target = tokenizer([data['target']])\n tokenized_sent_target = tokenizer_target['input_ids'][0]\n \n mask_sentence = tokenizer_output['attention_mask'][0]\n mask_target = tokenizer_target['attention_mask'][0]\n sentences.append(torch.tensor(tokenized_sent))\n targets.append(torch.tensor(tokenized_sent_target))\n masks_targets.append(torch.tensor(mask_targets))\n masks_sentences.append(torch.tensor(mask_sentences))\n sentences = pad_sequence(sentences, batch_first=True, padding_value=bert_pad_token)\n targets = pad_sequence(targets, batch_first=True, padding_value=bert_pad_token)\n masks = pad_sequence(masks, batch_first=True, padding_value=0.0)\n return sentences, targets, masks",
"_____no_output_____"
],
[
"#create pytorch dataloaders from train_dataset, val_dataset, and test_datset\nbatch_size=5\ntrain_dataloader = DataLoader(dataset,batch_size=batch_size,collate_fn=partial(transformer_collate_fn, tokenizer=tokenizer), shuffle = True)",
"_____no_output_____"
],
[
"#tokenizer.batch_decode(transformer_collate_fn(train_dataset,tokenizer)[0], skip_special_tokens=True)",
"_____no_output_____"
]
],
[
[
"## Polyencoder Model",
"_____no_output_____"
]
],
[
[
"#torch.cuda.empty_cache()\n#bert1 = DistilBertModel.from_pretrained(bert_model_name)\n#bert2 = DistilBertModel.from_pretrained(bert_model_name)\n\nbert = DistilBertModel.from_pretrained(bert_model_name)",
"Some weights of the model checkpoint at distilbert-base-uncased were not used when initializing DistilBertModel: ['vocab_projector.weight', 'vocab_transform.bias', 'vocab_projector.bias', 'vocab_transform.weight', 'vocab_layer_norm.weight', 'vocab_layer_norm.bias']\n- This IS expected if you are initializing DistilBertModel from the checkpoint of a model trained on another task or with another architecture (e.g. initializing a BertForSequenceClassification model from a BertForPreTraining model).\n- This IS NOT expected if you are initializing DistilBertModel from the checkpoint of a model that you expect to be exactly identical (initializing a BertForSequenceClassification model from a BertForSequenceClassification model).\n"
],
[
"#Double Bert\nclass RetrieverPolyencoder(nn.Module):\n def __init__(self, contextBert, candidateBert, vocab, max_len = 300, hidden_dim = 768, out_dim = 64, num_layers = 2, dropout=0.1, device=device):\n super().__init__()\n\n self.device = device\n self.hidden_dim = hidden_dim\n self.max_len = max_len\n self.out_dim = out_dim\n \n # Context layers\n self.contextBert = contextBert\n self.contextDropout = nn.Dropout(dropout)\n self.contextFc = nn.Linear(self.hidden_dim, self.out_dim)\n \n # Candidates layers\n self.candidatesBert = candidateBert\n self.pos_emb = nn.Embedding(self.max_len, self.hidden_dim)\n self.candidatesDropout = nn.Dropout(dropout)\n self.candidatesFc = nn.Linear(self.hidden_dim, self.out_dim)\n \n self.att_dropout = nn.Dropout(dropout)\n\n\n def attention(self, q, k, v, vMask=None):\n w = torch.matmul(q, k.transpose(-1, -2))\n if vMask is not None:\n w *= vMask.unsqueeze(1)\n w = F.softmax(w, -1)\n w = self.att_dropout(w)\n score = torch.matmul(w, v)\n return score\n\n def score(self, context, context_mask, responses, responses_mask):\n \"\"\"Run the model on the source and compute the loss on the target.\n\n Args:\n source: An integer tensor with shape (max_source_sequence_length,\n batch_size) containing subword indices for the source sentences.\n target: An integer tensor with shape (max_target_sequence_length,\n batch_size) containing subword indices for the target sentences.\n\n Returns:\n A scalar float tensor representing cross-entropy loss on the current batch\n divided by the number of target tokens in the batch.\n Many of the target tokens will be pad tokens. You should mask the loss \n from these tokens using appropriate mask on the target tokens loss.\n \"\"\"\n batch_size, nb_cand, seq_len = responses.shape\n # Context\n context_encoded = self.contextBert(context,context_mask)[-1]\n pos_emb = self.pos_emb(torch.arange(self.max_len).to(self.device))\n context_att = self.attention(pos_emb, context_encoded, context_encoded, context_mask)\n\n # Response\n responses_encoded = self.candidatesBert(responses.view(-1,responses.shape[2]), responses_mask.view(-1,responses.shape[2]))[-1][:,0,:]\n responses_encoded = responses_encoded.view(batch_size,nb_cand,-1)\n \n context_emb = self.attention(responses_encoded, context_att, context_att).squeeze() \n dot_product = (context_emb*responses_encoded).sum(-1)\n \n return dot_product\n\n \n def compute_loss(self, context, context_mask, response, response_mask):\n \"\"\"Run the model on the source and compute the loss on the target.\n\n Args:\n source: An integer tensor with shape (max_source_sequence_length,\n batch_size) containing subword indices for the source sentences.\n target: An integer tensor with shape (max_target_sequence_length,\n batch_size) containing subword indices for the target sentences.\n\n Returns:\n A scalar float tensor representing cross-entropy loss on the current batch\n divided by the number of target tokens in the batch.\n Many of the target tokens will be pad tokens. You should mask the loss \n from these tokens using appropriate mask on the target tokens loss.\n \"\"\"\n batch_size = context.shape[0]\n \n # Context\n context_encoded = self.contextBert(context,context_mask)[-1]\n pos_emb = self.pos_emb(torch.arange(self.max_len).to(self.device))\n context_att = self.attention(pos_emb, context_encoded, context_encoded, context_mask)\n\n # Response\n response_encoded = self.candidatesBert(response, response_mask)[-1][:,0,:]\n \n response_encoded = response_encoded.unsqueeze(0).expand(batch_size, batch_size, response_encoded.shape[1]) \n context_emb = self.attention(response_encoded, context_att, context_att).squeeze() \n dot_product = (context_emb*response_encoded).sum(-1)\n mask = torch.eye(batch_size).to(self.device)\n loss = F.log_softmax(dot_product, dim=-1) * mask\n loss = (-loss.sum(dim=1)).mean()\n return loss",
"_____no_output_____"
],
[
"#Single Bert\nclass RetrieverPolyencoder_single(nn.Module):\n def __init__(self, bert, max_len = 300, hidden_dim = 768, out_dim = 64, num_layers = 2, dropout=0.1, device=device):\n super().__init__()\n\n self.device = device\n self.hidden_dim = hidden_dim\n self.max_len = max_len\n self.out_dim = out_dim\n self.bert = bert\n \n # Context layers\n self.contextDropout = nn.Dropout(dropout)\n \n # Candidates layers\n self.pos_emb = nn.Embedding(self.max_len, self.hidden_dim)\n self.candidatesDropout = nn.Dropout(dropout)\n \n self.att_dropout = nn.Dropout(dropout)\n\n\n def attention(self, q, k, v, vMask=None):\n w = torch.matmul(q, k.transpose(-1, -2))\n if vMask is not None:\n w *= vMask.unsqueeze(1)\n w = F.softmax(w, -1)\n w = self.att_dropout(w)\n score = torch.matmul(w, v)\n return score\n\n def score(self, context, context_mask, responses, responses_mask):\n \"\"\"Run the model on the source and compute the loss on the target.\n\n Args:\n source: An integer tensor with shape (max_source_sequence_length,\n batch_size) containing subword indices for the source sentences.\n target: An integer tensor with shape (max_target_sequence_length,\n batch_size) containing subword indices for the target sentences.\n\n Returns:\n A scalar float tensor representing cross-entropy loss on the current batch\n divided by the number of target tokens in the batch.\n Many of the target tokens will be pad tokens. You should mask the loss \n from these tokens using appropriate mask on the target tokens loss.\n \"\"\"\n batch_size, nb_cand, seq_len = responses.shape\n # Context\n context_encoded = self.bert(context,context_mask)[0][:,0,:]\n pos_emb = self.pos_emb(torch.arange(self.max_len).to(self.device))\n context_att = self.attention(pos_emb, context_encoded, context_encoded, context_mask)\n\n # Response\n responses_encoded = self.bert(responses.view(-1,responses.shape[2]), responses_mask.view(-1,responses.shape[2]))[0][:,0,:]\n responses_encoded = responses_encoded.view(batch_size,nb_cand,-1)\n response_encoded = self.candidatesFc(response_encoded)\n \n context_emb = self.attention(responses_encoded, context_att, context_att).squeeze() \n dot_product = (context_emb*responses_encoded).sum(-1)\n \n return dot_product\n\n \n def compute_loss(self, context, context_mask, response, response_mask):\n \"\"\"Run the model on the source and compute the loss on the target.\n\n Args:\n source: An integer tensor with shape (max_source_sequence_length,\n batch_size) containing subword indices for the source sentences.\n target: An integer tensor with shape (max_target_sequence_length,\n batch_size) containing subword indices for the target sentences.\n\n Returns:\n A scalar float tensor representing cross-entropy loss on the current batch\n divided by the number of target tokens in the batch.\n Many of the target tokens will be pad tokens. You should mask the loss \n from these tokens using appropriate mask on the target tokens loss.\n \"\"\"\n batch_size = context.shape[0]\n seq_len = response.shape[1]\n \n # Context\n context_encoded = self.bert(context,context_mask)[0][:,0,:]\n pos_emb = self.pos_emb(torch.arange(self.max_len).to(self.device))\n context_att = self.attention(pos_emb, context_encoded, context_encoded, context_mask)\n\n # Response\n print(response.shape)\n response_encoded = self.bert(response, response_mask)[0][:,0,:]\n print(response_encoded.shape)\n response_encoded = response_encoded.view(batch_size, -1)\n\n \n response_encoded = response_encoded.unsqueeze(0).expand(batch_size, batch_size, response_encoded.shape[1]) \n context_emb = self.attention(response_encoded, context_att, context_att).squeeze() \n dot_product = (context_emb*response_encoded).sum(-1)\n mask = torch.eye(batch_size).to(self.device)\n loss = F.log_softmax(dot_product, dim=-1) * mask\n loss = (-loss.sum(dim=1)).mean()\n return loss",
"_____no_output_____"
],
[
"#Bi-encoder\nclass RetrieverBiencoder(nn.Module):\n def __init__(self, bert):\n super().__init__()\n self.bert = bert\n \n def score(self, context, context_mask, responses, responses_mask):\n\n context_vec = self.bert(context, context_mask)[0][:,0,:] # [bs,dim]\n\n batch_size, res_length = response.shape\n\n responses_vec = self.bert(responses_input_ids, responses_input_masks)[0][:,0,:] # [bs,dim]\n responses_vec = responses_vec.view(batch_size, 1, -1)\n\n responses_vec = responses_vec.squeeze(1) \n context_vec = context_vec.unsqueeze(1)\n dot_product = torch.matmul(context_vec, responses_vec.permute(0, 2, 1)).squeeze()\n return dot_product\n \n def compute_loss(self, context, context_mask, response, response_mask):\n\n context_vec = self.bert(context, context_mask)[0] # [bs,dim]\n\n batch_size, res_length = response.shape\n\n responses_vec = self.bert(response, response_mask)[0][:,0,:] # [bs,dim]\n #responses_vec = responses_vec.view(batch_size, 1, -1)\n \n print(context_vec.shape)\n print(responses_vec.shape)\n\n responses_vec = responses_vec.squeeze(1)\n dot_product = torch.matmul(context_vec, responses_vec.t()) # [bs, bs]\n mask = torch.eye(context.size(0)).to(context_mask.device)\n loss = F.log_softmax(dot_product, dim=-1) * mask\n loss = (-loss.sum(dim=1)).mean()\n return loss\n",
"_____no_output_____"
],
[
"def train(model, data_loader, num_epochs, model_file, learning_rate=0.0001):\n \"\"\"Train the model for given µnumber of epochs and save the trained model in \n the final model_file.\n \"\"\"\n\n decoder_learning_ratio = 5.0\n #encoder_parameter_names = ['word_embedding', 'encoder']\n encoder_parameter_names = ['encode_emb', 'encode_gru', 'l1', 'l2']\n \n encoder_named_params = list(filter(lambda kv: any(key in kv[0] for key in encoder_parameter_names), model.named_parameters()))\n decoder_named_params = list(filter(lambda kv: not any(key in kv[0] for key in encoder_parameter_names), model.named_parameters()))\n encoder_params = [e[1] for e in encoder_named_params]\n decoder_params = [e[1] for e in decoder_named_params]\n optimizer = torch.optim.AdamW([{'params': encoder_params},\n {'params': decoder_params, 'lr': learning_rate * decoder_learning_ratio}], lr=learning_rate)\n \n clip = 50.0\n for epoch in tqdm.notebook.trange(num_epochs, desc=\"training\", unit=\"epoch\"):\n # print(f\"Total training instances = {len(train_dataset)}\")\n # print(f\"train_data_loader = {len(train_data_loader)} {1180 > len(train_data_loader)/20}\")\n with tqdm.notebook.tqdm(\n data_loader,\n desc=\"epoch {}\".format(epoch + 1),\n unit=\"batch\",\n total=len(data_loader)) as batch_iterator:\n model.train()\n total_loss = 0.0\n for i, batch_data in enumerate(batch_iterator, start=1):\n source, mask_source, target, mask_target = batch_data[\"conv_tensors\"]\n optimizer.zero_grad()\n loss = model.compute_loss(source, mask_source, target, mask_target)\n total_loss += loss.item()\n loss.backward()\n # Gradient clipping before taking the step\n _ = nn.utils.clip_grad_norm_(model.parameters(), clip)\n optimizer.step()\n\n batch_iterator.set_postfix(mean_loss=total_loss / i, current_loss=loss.item())\n # Save the model after training \n torch.save(model.state_dict(), model_file)",
"_____no_output_____"
],
[
"# You are welcome to adjust these parameters based on your model implementation.\nnum_epochs = 10\nbatch_size = 32\nlearning_rate = 0.001\n# Reloading the data_loader to increase batch_size\n\nbaseline_model = RetrieverBiencoder(bert).to(device)\ntrain(baseline_model, train_dataloader, num_epochs, \"baseline_model.pt\",learning_rate=learning_rate)\n# Download the trained model to local for future use\n#files.download('baseline_model.pt')",
"_____no_output_____"
],
[
"baseline_model = RetrieverPolyencoder(bert1,bert2,vocab).to(device)\nbaseline_model.load_state_dict(torch.load(\"baseline_model3.pt\", map_location=device))",
"_____no_output_____"
],
[
"vals = transformer_collate_fn(all_conversations[0:100],tokenizer)",
"_____no_output_____"
],
[
"i=3",
"_____no_output_____"
],
[
"scores = baseline_model.score(vals[0][i].unsqueeze(0).cuda(),vals[1][i].unsqueeze(0).cuda(),vals[2].unsqueeze(0).cuda(),vals[3].unsqueeze(0).cuda()).detach().cpu().numpy()",
"_____no_output_____"
],
[
"all_conversations[i][0]",
"_____no_output_____"
],
[
"all_conversations[np.argmax(scores)][1]",
"_____no_output_____"
],
[
"max_v = 100\nvals = transformer_collate_fn(all_conversations[0:max_v],tokenizer)\ncorrect = 0\nfor i in range(max_v):\n scores = baseline_model.score(vals[0][i].unsqueeze(0).cuda(),vals[1][i].unsqueeze(0).cuda(),vals[2].unsqueeze(0).cuda(),vals[3].unsqueeze(0).cuda()).detach().cpu().numpy()\n if np.argmax(scores)==i:\n correct+=1\n print(all_conversations[i][0])\n print(all_conversations[np.argmax(scores)][1]+\"\\n\")",
"_____no_output_____"
],
[
"print(correct/max_v)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acf8aa5c7eb54cba0a317d0230ad206bf1e765b
| 4,629 |
ipynb
|
Jupyter Notebook
|
notes/operator-overloading.ipynb
|
Victor-Palacios/msds689
|
c7ec2afe2ec159d6ed8bdd31eda26aeaac8cb2ce
|
[
"MIT"
] | 52 |
2019-01-07T23:49:55.000Z
|
2022-03-11T05:43:02.000Z
|
notes/operator-overloading.ipynb
|
Victor-Palacios/msds689
|
c7ec2afe2ec159d6ed8bdd31eda26aeaac8cb2ce
|
[
"MIT"
] | null | null | null |
notes/operator-overloading.ipynb
|
Victor-Palacios/msds689
|
c7ec2afe2ec159d6ed8bdd31eda26aeaac8cb2ce
|
[
"MIT"
] | 76 |
2019-01-26T02:59:18.000Z
|
2022-03-05T20:33:19.000Z
| 23.617347 | 354 | 0.477209 |
[
[
[
"# Operator overloading in Python\n\nReview [Object-oriented programming](https://github.com/parrt/msds501/blob/master/notes/OO.ipynb)\n\nNote: We *overload* operators but *override* methods in a subclass definition\n\nPython allows class definitions to implement functions that are called when standard operator symbols such as + and / are applied to objects of that type. This is extremely useful for mathematical libraries such as numpy, but is often abused. Note that you could redefine subtraction to be multiplication when someone used the `-` sign. (Yikes!)\n\nImagine we have a simple `Point` class:",
"_____no_output_____"
]
],
[
[
"import numpy as np\n\nclass Point:\n def __init__(self, x, y):\n self.x = x\n self.y = y\n \n def distance(self, other):\n return np.sqrt( (self.x - other.x)**2 + (self.y - other.y)**2 )\n \n def __str__(self): # convert to string, such as when printing\n return f\"({self.x},{self.y})\"\n\n def __repr__(self): # used to convert to string in notebooks\n return f\"({self.x},{self.y})\"\n\nPoint(3,4)",
"_____no_output_____"
]
],
[
[
"Here's an extension to `Point` that supports + for `Point` addition:",
"_____no_output_____"
]
],
[
[
"import numpy as np\n\nclass Point:\n def __init__(self, x, y):\n self.x = x\n self.y = y\n \n def distance(self, other):\n return np.sqrt( (self.x - other.x)**2 + (self.y - other.y)**2 )\n \n def __add__(self,other):\n x = self.x + other.x\n y = self.y + other.y\n return Point(x,y)\n \n def __str__(self):\n return f\"({self.x},{self.y})\"\n\n def __repr__(self): # used to convert to string in notebooks\n return f\"({self.x},{self.y})\"",
"_____no_output_____"
],
[
"p = Point(3,4)\nq = Point(5,6)\nprint(p, q)\nprint(p + q) # calls p.__add__(q) or Point.__add__(p,q)\nprint(Point.__add__(p,q))",
"(3,4) (5,6)\n(8,10)\n(8,10)\n"
]
],
[
[
"## Exercise",
"_____no_output_____"
],
[
"Add a method to implement the `-` subtraction operator for `Point` so that the following code works:\n\n```\np = Point(5,4)\nq = Point(1,5)\nprint(p, q)\nprint(p - q)\n```",
"_____no_output_____"
]
],
[
[
" def __sub__(self,other):\n x = self.x - other.x\n y = self.y - other.y\n return Point(x,y)",
"_____no_output_____"
],
[
"A = [9,1,3,1,9]\ndef unique(A):\n seen = set()\n for a in A:\n if a not in seen:\n seen.add(a)\n return list(seen)\n\nunique(A)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code"
]
] |
4acf91889bdfcfab8e8eef62dbc53e6e64441bc7
| 13,059 |
ipynb
|
Jupyter Notebook
|
notebooks/.ipynb_checkpoints/Intro_08_Xarray-Advanced_cruise_data_case_study-checkpoint.ipynb
|
cgentemann/ocean_python_tutorial
|
4a974ff1707450dd42709bc268c1e2f5cdf8ad5e
|
[
"Apache-2.0"
] | 1 |
2021-12-01T00:38:41.000Z
|
2021-12-01T00:38:41.000Z
|
notebooks/Intro_08_Xarray-Advanced_cruise_data_case_study.ipynb
|
cgentemann/ocean_python_tutorial
|
4a974ff1707450dd42709bc268c1e2f5cdf8ad5e
|
[
"Apache-2.0"
] | null | null | null |
notebooks/Intro_08_Xarray-Advanced_cruise_data_case_study.ipynb
|
cgentemann/ocean_python_tutorial
|
4a974ff1707450dd42709bc268c1e2f5cdf8ad5e
|
[
"Apache-2.0"
] | 4 |
2019-08-30T15:33:25.000Z
|
2021-03-10T08:05:09.000Z
| 29.214765 | 273 | 0.583659 |
[
[
[
"# Cruise collocation with gridded data \n \nAuthors\n* [Dr Chelle Gentemann](mailto:[email protected]) - Earth and Space Research, USA\n* [Dr Marisol Garcia-Reyes](mailto:[email protected]) - Farallon Institute, USA \n-------------\n",
"_____no_output_____"
],
[
"# Structure of this tutorial\n\n1. Opening data\n1. Collocating satellite data with a cruise dataset\n\n",
"_____no_output_____"
],
[
"-------------------\n\n## Import python packages\n\nYou are going to want numpy, pandas, matplotlib.pyplot and xarray",
"_____no_output_____"
]
],
[
[
"import warnings\nwarnings.simplefilter('ignore') # filter some warning messages\n\nimport numpy as np\nimport pandas as pd\nimport matplotlib.pyplot as plt\nimport xarray as xr\nimport cartopy.crs as ccrs\n",
"_____no_output_____"
]
],
[
[
"## A nice cartopy tutorial is [here](http://earthpy.org/tag/visualization.html)",
"_____no_output_____"
],
[
"# Collocating Saildrone cruise data with satellite SSTs \n\n* read in the Saildrone data ",
"_____no_output_____"
],
[
"## The NCEI trajectory format uses 'obs' as the coordinate. This is an example of an 'older' style of data formatting that doesn't really mesh well with modern software capabilities. \n\n* So, let's change that by using [.swap_dims](http://xarray.pydata.org/en/stable/generated/xarray.DataArray.swap_dims.html) to change the coordinate from `obs` to `time`\n* Another thing, `latitude` and `longitude` are just long and annoying, lets [.rename](http://xarray.pydata.org/en/stable/generated/xarray.Dataset.rename.html) them to `lat` and `lon`\n\n* Finally, the first and last part of the cruise the USV is being towed, so let's only include data from `2018-04-12T02` to `2018-06-10T18`\n",
"_____no_output_____"
]
],
[
[
"#use first url if not online\n#url = '../data/saildrone-gen_4-baja_2018-sd1002-20180411T180000-20180611T055959-1_minutes-v1.nc'\nurl = 'https://podaac-opendap.jpl.nasa.gov/opendap/hyrax/allData/insitu/L2/saildrone/Baja/saildrone-gen_4-baja_2018-sd1002-20180411T180000-20180611T055959-1_minutes-v1.nc'\n\nds_usv = ",
"_____no_output_____"
],
[
"ds_usv2 = ds_usv.isel(trajectory=0)\\\n .swap_dims({'obs':'time'})\\\n .rename({'longitude':'lon','latitude':'lat'})\n\nds_usv_subset = ds_usv2.sel(time=slice('2018-04-12T02','2018-06-10T18')) \n\nstart_time = pd.to_datetime(str(ds_usv_subset.time.min().data)).strftime('%Y-%m-%dT%H:%m:%SZ') \nend_time = pd.to_datetime(str(ds_usv_subset.time.max().data)).strftime('%Y-%m-%dT%H:%m:%SZ') \n\nprint('start: ',start_time,'end: ',end_time)",
"_____no_output_____"
]
],
[
[
"## Let's open 2 months of 0.2 km AVHRR OI SST data\n`xarray`can open multiple files at once using string pattern matching. `xr.open_mfdataset()` \n \n## Now open multiple files (lazy) using [.open_mfdataset](http://xarray.pydata.org/en/stable/generated/xarray.open_mfdataset.html#xarray.open_mfdataset)\n\n* use the option `coords = 'minimal'`",
"_____no_output_____"
]
],
[
[
"files = '../data/avhrr_oi/*.nc'\n\nds_sst = xr.open_mfdataset(files,coords = 'minimal')\n",
"_____no_output_____"
]
],
[
[
"# Let's see what one day looks like\n\n* add coastlines `ax.coastlines()`\n* add gridlines `ax.gridlines()`",
"_____no_output_____"
]
],
[
[
"sst = ds_sst.sst[0,:,:]",
"_____no_output_____"
],
[
"ax = plt.axes(projection=ccrs.Orthographic(-80, 35))\n\nsst.plot(ax=ax, transform=ccrs.PlateCarree())\n",
"_____no_output_____"
]
],
[
[
"# Change the figure\n* colormap `cmap='jet'\n* colorscale `vmin=-1,vmax=34`\n* add land `ax.stock_imag()",
"_____no_output_____"
]
],
[
[
"ax = plt.axes(projection=ccrs.Orthographic(-80, 35))\n\nsst.plot(ax=ax, transform=ccrs.PlateCarree())\n\n\n\n",
"_____no_output_____"
]
],
[
[
"# Look at the sst data and notice the longitude range",
"_____no_output_____"
],
[
"## Again with the 0-360 vs -180-180. Change it up below!\n* `ds_sst.coords['lon'] = np.mod(ds_sst.coords['lon'] + 180,360) - 180`\n* remember to sort by lon, `.sortby(ds_sst.lon)`\n* Also, look at the coordinates, there is an extra one `zlev`. Drop it using .isel\n\n",
"_____no_output_____"
]
],
[
[
"ds_sst.coords['lon'] = #change lon -180to180\n\nds_sst = ds_sst # sort lon\n\nds_sst = ds_sst #isel zlev\n\nds_sst",
"_____no_output_____"
]
],
[
[
"`open_mfdataset` even puts them in the right order for you.",
"_____no_output_____"
]
],
[
[
"ds_sst.time",
"_____no_output_____"
]
],
[
[
"How big is all this data uncompressed? Will it fit into memory?\nUse `.nbytes` / 1e9 to convert it into gigabytes",
"_____no_output_____"
]
],
[
[
"print('file size (GB):', ds_sst.nbytes / 1e9) ",
"_____no_output_____"
]
],
[
[
"# Xarray interpolation won't run on chunked dimensions. \n1. First let's subset the data to make it smaller to deal with by using the cruise lat/lons\n * Find the max/min of the lat/lon using `.lon.min().data`\n\n1. Now load the data into memory (de-Dask-ify) it using `.load()` \n",
"_____no_output_____"
]
],
[
[
"#Step 1 from above\nlon_min,lon_max = ds_usv_subset.lon.min().data,ds_usv_subset.lon.max().data\n\nlat_min,lat_max = ds_usv_subset.lat.min().data,ds_usv_subset.lat.max().data\n\nsubset = ds_sst.sel(lon=slice(lon_min,lon_max),\n lat=slice(lat_min,lat_max))\n\nprint('file size (GB):', subset.nbytes / 1e9) ",
"_____no_output_____"
],
[
"#Step 2 from above\n\nsubset.load()",
"_____no_output_____"
]
],
[
[
"# Collocate USV data with SST data\nThere are different options when you interpolate. First, let's just do a linear interpolation using [.interp()](http://xarray.pydata.org/en/stable/generated/xarray.Dataset.interp.html#xarray.Dataset.interp)\n\n`Dataset.interp(coords=None, method='linear', assume_sorted=False, kwargs={}, **coords_kwargs))`\n\n",
"_____no_output_____"
]
],
[
[
"ds_collocated = subset.interp(lat=ds_usv_subset.lat,lon=ds_usv_subset.lon,time=ds_usv_subset.time,method='linear')",
"_____no_output_____"
]
],
[
[
"# Collocate USV data with SST data\nThere are different options when you interpolate. First, let's just do a nearest point rather than interpolate the data\n`method = 'nearest'`",
"_____no_output_____"
]
],
[
[
"ds_collocated_nearest = subset.interp(lat=ds_usv_subset.lat,lon=ds_usv_subset.lon,time=ds_usv_subset.time,method='nearest')",
"_____no_output_____"
]
],
[
[
"## Now, calculate the different in SSTs and print the [.mean()](http://xarray.pydata.org/en/stable/generated/xarray.DataArray.mean.html#xarray.DataArray.mean) and [.std()](http://xarray.pydata.org/en/stable/generated/xarray.DataArray.std.html#xarray.DataArray.std)\nFor the satellite data we need to use `sst` and for the USV data we need to use `TEMP_CTD_MEAN`",
"_____no_output_____"
]
],
[
[
"dif = ds_collocated_nearest.sst-ds_usv_subset.TEMP_CTD_MEAN\nprint('mean difference = ',dif.mean().data)\nprint('STD = ',dif.std().data)",
"_____no_output_____"
]
],
[
[
"# xarray can do more!\n\n* concatentaion\n* open network located files with openDAP\n* import and export Pandas DataFrames\n* .nc dump to \n* groupby_bins\n* resampling and reduction\n\nFor more details, read this blog post: http://continuum.io/blog/xray-dask\n",
"_____no_output_____"
]
],
[
[
"#ds_collocated_nearest.to_netcdf('./data/new file.nc')",
"_____no_output_____"
]
],
[
[
"## Where can I find more info?\n\n### For more information about xarray\n\n- Read the [online documentation](http://xarray.pydata.org/)\n- Ask questions on [StackOverflow](http://stackoverflow.com/questions/tagged/python-xarray)\n- View the source code and file bug reports on [GitHub](http://github.com/pydata/xarray/)\n\n### For more doing data analysis with Python:\n\n- Thomas Wiecki, [A modern guide to getting started with Data Science and Python](http://twiecki.github.io/blog/2014/11/18/python-for-data-science/)\n- Wes McKinney, [Python for Data Analysis](http://shop.oreilly.com/product/0636920023784.do) (book)\n\n### Packages building on xarray for the geophysical sciences\n\nFor analyzing GCM output:\n\n- [xgcm](https://github.com/xgcm/xgcm) by Ryan Abernathey\n- [oogcm](https://github.com/lesommer/oocgcm) by Julien Le Sommer\n- [MPAS xarray](https://github.com/pwolfram/mpas_xarray) by Phil Wolfram\n- [marc_analysis](https://github.com/darothen/marc_analysis) by Daniel Rothenberg\n\nOther tools:\n\n- [windspharm](https://github.com/ajdawson/windspharm): wind spherical harmonics by Andrew Dawson\n- [eofs](https://github.com/ajdawson/eofs): empirical orthogonal functions by Andrew Dawson\n- [infinite-diff](https://github.com/spencerahill/infinite-diff) by Spencer Hill \n- [aospy](https://github.com/spencerahill/aospy) by Spencer Hill and Spencer Clark\n- [regionmask](https://github.com/mathause/regionmask) by Mathias Hauser\n- [salem](https://github.com/fmaussion/salem) by Fabien Maussion\n\nResources for teaching and learning xarray in geosciences:\n- [Fabien's teaching repo](https://github.com/fmaussion/teaching): courses that combine teaching climatology and xarray\n",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown"
] |
[
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
]
] |
4acf93a4a6f9b1101a3c3aca3b2eb522f03accaa
| 8,424 |
ipynb
|
Jupyter Notebook
|
notebook/Untitled.ipynb
|
abbiyanaila/torchwisdom
|
56dc95ebca3f6861c7009cb4fa0c034e260236b1
|
[
"MIT"
] | 1 |
2019-04-29T12:35:07.000Z
|
2019-04-29T12:35:07.000Z
|
notebook/Untitled.ipynb
|
abbiyanaila/torchwisdom
|
56dc95ebca3f6861c7009cb4fa0c034e260236b1
|
[
"MIT"
] | null | null | null |
notebook/Untitled.ipynb
|
abbiyanaila/torchwisdom
|
56dc95ebca3f6861c7009cb4fa0c034e260236b1
|
[
"MIT"
] | null | null | null | 47.863636 | 1,798 | 0.593898 |
[
[
[
"from nltk.classify import NaiveBayesClassifier\nfrom nltk.corpus import stopwords\nstopset = list(set(stopwords.words('english')))\n",
"_____no_output_____"
],
[
"import re\nimport csv\n\nimport nltk.classify\n\n\ndef replaceTwoOrMore(s):\n pattern = re.compile(r\"(.)\\1{1,}\", re.DOTALL)\n return pattern.sub(r\"\\1\\1\", s)\n\n\ndef processTweet(tweet):\n tweet = tweet.lower()\n tweet = re.sub('((www\\.[^\\s]+)|(https?://[^\\s]+))', 'URL', tweet)\n tweet = re.sub('@[^\\s]+', 'AT_USER', tweet)\n tweet = re.sub('[\\s]+', ' ', tweet)\n tweet = re.sub(r'#([^\\s]+)', r'\\1', tweet)\n tweet = tweet.strip('\\'\"')\n return tweet\n\n\ndef getStopWordList(stopWordListFileName):\n stopWords = []\n stopWords.append('AT_USER')\n stopWords.append('URL')\n\n fp = open(stopWordListFileName, 'r')\n line = fp.readline()\n while line:\n word = line.strip()\n stopWords.append(word)\n line = fp.readline()\n fp.close()\n return stopWords\n\n\ndef getFeatureVector(tweet, stopWords):\n featureVector = []\n words = tweet.split()\n for w in words:\n\n w = replaceTwoOrMore(w)\n\n w = w.strip('\\'\"?,.')\n\n val = re.search(r\"^[a-zA-Z][a-zA-Z0-9]*[a-zA-Z]+[a-zA-Z0-9]*$\", w)\n\n if (w in stopWords or val is None):\n continue\n else:\n featureVector.append(w.lower())\n return featureVector\n\n\ndef extract_features(tweet):\n tweet_words = set(tweet)\n features = {}\n for word in featureList:\n features['contains(%s)' % word] = (word in tweet_words)\n return features\n\n\ninpTweets = csv.reader(open('data/training.csv', 'r'), delimiter=',', quotechar='|')\nstopWords = getStopWordList('data/stopwordsID.txt')\ncount = 0;\nfeatureList = []\ntweets = []\nfor row in inpTweets:\n sentiment = row[0]\n tweet = row[1]\n processedTweet = processTweet(tweet) # preprocessing\n featureVector = getFeatureVector(processedTweet, stopWords) # get feature vector\n featureList.extend(featureVector)\n tweets.append((featureVector, sentiment));\n\nfeatureList = list(set(featureList))\n\ntraining_set = nltk.classify.util.apply_features(extract_features, tweets)\n\nNBClassifier = nltk.NaiveBayesClassifier.train(training_set)\n\ntestTweet = 'pantai di lombok bersih bersih. pasirnya juga indah'\nprocessedTestTweet = processTweet(testTweet)\nsentiment = NBClassifier.classify(extract_features(getFeatureVector(processedTestTweet, stopWords)))\nprint(\"Test Tweets = %s, Sentiment = %s\\n\" % (testTweet, sentiment))\n\n# print(\"Show Most Informative Features\", NBClassifier.show_most_informative_features(32))\n# print()\n# print(\"Extract Features\", extract_features(testTweet.split()))\n\nprint(\"Akurasi Hasil Klasifikasi :\", (nltk.classify.accuracy(processedTestTweet, NBClassifier)) * 100)\n# print(\"Akurasi Hasil Klasifikasi :\", accuracy_score(testTweet, sentiment))\n\n# kal = getFeatureVector(processTweet(testTweet), stopWords)\n# kal = \" \".join(str(x) for x in kal)\n# print(kal)\n# d = {}\n# for word in kal.split():\n# word = int(word) if word.isdigit() else word\n# if word in d:\n# d[word] += 1\n# else:\n# d[word] = 1\n\n# print(d)",
"Test Tweets = pantai di lombok bersih bersih. pasirnya juga indah, Sentiment = positive\n\n"
]
]
] |
[
"code"
] |
[
[
"code",
"code"
]
] |
4acf9a2e0a8b105f640a7589fd2edc2a18dcc8f9
| 854,923 |
ipynb
|
Jupyter Notebook
|
_notebooks/2021-02-16-Kapitel-2-hotdogs.ipynb
|
Imotep460/FastAIBlog
|
5b368c05cdfdca0a2da1cdf1559585962ca9b9ca
|
[
"Apache-2.0"
] | null | null | null |
_notebooks/2021-02-16-Kapitel-2-hotdogs.ipynb
|
Imotep460/FastAIBlog
|
5b368c05cdfdca0a2da1cdf1559585962ca9b9ca
|
[
"Apache-2.0"
] | null | null | null |
_notebooks/2021-02-16-Kapitel-2-hotdogs.ipynb
|
Imotep460/FastAIBlog
|
5b368c05cdfdca0a2da1cdf1559585962ca9b9ca
|
[
"Apache-2.0"
] | null | null | null | 747.964129 | 673,338 | 0.946381 |
[
[
[
"<a href=\"https://colab.research.google.com/github/Imotep460/FastAIBlog/blob/master/_notebooks/2021-02-16-Kapitel-2%3A-hotdogs.ipynb\" target=\"_parent\"><img src=\"https://colab.research.google.com/assets/colab-badge.svg\" alt=\"Open In Colab\"/></a>",
"_____no_output_____"
]
],
[
[
"!pip install -Uqq fastbook\r\nimport fastbook\r\nfastbook.setup_book()",
"\u001b[K |████████████████████████████████| 727kB 5.9MB/s \n\u001b[K |████████████████████████████████| 194kB 33.0MB/s \n\u001b[K |████████████████████████████████| 51kB 5.3MB/s \n\u001b[K |████████████████████████████████| 1.2MB 29.8MB/s \n\u001b[K |████████████████████████████████| 61kB 7.6MB/s \n\u001b[?25hMounted at /content/gdrive\n"
],
[
"from fastbook import *\r\nfrom fastai.vision.widgets import *",
"_____no_output_____"
],
[
"configId = \"\"",
"_____no_output_____"
],
[
"subscriptionKey = \"\"",
"_____no_output_____"
],
[
"def search_images_bing_new(key, term, customConfigId, min_sz=128):\r\n\turl = 'https://api.bing.microsoft.com/v7.0/custom/images/search?' + 'q=' + term + '&' + 'customconfig=' + customConfigId + '&' + 'count=150'\r\n\tr = requests.get(url, headers={'Ocp-Apim-Subscription-Key': key})\r\n\tsearch_results = r.json()\r\n\treturn L([img[\"thumbnailUrl\"] + \".jpg\" for img in search_results[\"value\"][:150]])",
"_____no_output_____"
],
[
"hotdogImages = search_images_bing_new(subscriptionKey, \"hotdog\", configId)",
"_____no_output_____"
],
[
"firstHotdogImage = hotdogImages[0]\r\nhotdogDest = \"hotdog.jpg\"\r\ndownload_url(firstHotdogImage, hotdogDest)",
"_____no_output_____"
],
[
"hotdogImg = Image.open(hotdogDest)\nhotdogImg.to_thumb(128,128)",
"_____no_output_____"
],
[
"imagesPath = Path('images')\r\nif not imagesPath.exists:\r\n imagesPath.mkdir()",
"_____no_output_____"
],
[
"hotdogPath = Path(str(imagesPath) + '/hotdogs')\r\nif not hotdogPath.exists():\r\n hotdogPath.mkdir()\r\ndownload_images(hotdogPath, urls = hotdogImages)",
"_____no_output_____"
],
[
"hotdogImageFiles = get_image_files(hotdogPath)",
"_____no_output_____"
],
[
"failedHotdogs = verify_images(hotdogImageFiles)\r\nfailedHotdogs",
"_____no_output_____"
],
[
"foodImages = search_images_bing_new(subscriptionKey, \"food -hotdog\", configId)\r\nfoodPath = Path(str(imagesPath) + '/food')\r\nif not foodPath.exists():\r\n foodPath.exists()\r\ndownload_images(foodPath, urls=foodImages)\r\nfoodImageFiles = get_image_files(foodPath)\r\nfailedFood = verify_images(foodImageFiles)\r\nfailedFood",
"_____no_output_____"
],
[
"foods = DataBlock(\r\n blocks=(ImageBlock, CategoryBlock), \r\n get_items=get_image_files, \r\n splitter=RandomSplitter(valid_pct=0.2, seed=42),\r\n get_y=parent_label,\r\n item_tfms=Resize(128))\r\nfoods = foods.new(\r\n item_tfms=RandomResizedCrop(224, min_scale=0.5),\r\n batch_tfms=aug_transforms())",
"_____no_output_____"
],
[
"dls = foods.dataloaders(imagesPath)",
"_____no_output_____"
],
[
"dls.valid.show_batch(max_n=12, nrows=1)",
"_____no_output_____"
],
[
"learn = cnn_learner(dls, resnet18, metrics=error_rate)\r\nlearn.fine_tune(8)",
"Downloading: \"https://download.pytorch.org/models/resnet18-5c106cde.pth\" to /root/.cache/torch/hub/checkpoints/resnet18-5c106cde.pth\n"
],
[
"interp = ClassificationInterpretation.from_learner(learn)\r\ninterp.plot_confusion_matrix()",
"_____no_output_____"
],
[
"learn.export()",
"_____no_output_____"
],
[
"Image.open(\"hotdog.jpg\").to_thumb(128,128)",
"_____no_output_____"
],
[
"pred,pred_idx,probs = learn.predict(\"hotdog.jpg\")\r\nf'Prediction: {pred}; Probability: {probs[pred_idx]:.04f}'",
"_____no_output_____"
],
[
"Image.open(\"burger.jpg\").to_thumb(128,128)",
"_____no_output_____"
],
[
"pred,pred_idx,probs = learn.predict(\"burger.jpg\")\r\nf'Prediction: {pred}; Probability: {probs[pred_idx]:.04f}'",
"_____no_output_____"
],
[
"Image.open(\"dog.jpg\").to_thumb(128,128)",
"_____no_output_____"
],
[
"pred,pred_idx,probs = learn.predict(\"dog.jpg\")\r\nf'Prediction: {pred}; Probability: {probs[pred_idx]:.04f}'",
"_____no_output_____"
],
[
"Image.open(\"hotdogdog.jpg\").to_thumb(128,128)",
"_____no_output_____"
],
[
"pred,pred_idx,probs = learn.predict(\"hotdogdog.jpg\")\r\nf'Prediction: {pred}; Probability: {probs[pred_idx]:.04f}'",
"_____no_output_____"
],
[
"",
"_____no_output_____"
]
]
] |
[
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acfc21345bcb3e1b85e9e0ed36ed7afbd88f395
| 85,922 |
ipynb
|
Jupyter Notebook
|
conditional/.ipynb_checkpoints/main_conditional_disentangle_cifar_bs900_sratio_0_5_drop_0_5_run1_temp-checkpoint.ipynb
|
minhtannguyen/ffjord
|
f3418249eaa4647f4339aea8d814cf2ce33be141
|
[
"MIT"
] | null | null | null |
conditional/.ipynb_checkpoints/main_conditional_disentangle_cifar_bs900_sratio_0_5_drop_0_5_run1_temp-checkpoint.ipynb
|
minhtannguyen/ffjord
|
f3418249eaa4647f4339aea8d814cf2ce33be141
|
[
"MIT"
] | null | null | null |
conditional/.ipynb_checkpoints/main_conditional_disentangle_cifar_bs900_sratio_0_5_drop_0_5_run1_temp-checkpoint.ipynb
|
minhtannguyen/ffjord
|
f3418249eaa4647f4339aea8d814cf2ce33be141
|
[
"MIT"
] | null | null | null | 60.253857 | 1,074 | 0.475885 |
[
[
[
"import os\nos.environ['CUDA_VISIBLE_DEVICES']='0,1,2,3'",
"_____no_output_____"
],
[
"%run -p ../train_cnf_disentangle_cifar.py --data cifar10 --dims 64,64,64 --strides 1,1,1,1 --num_blocks 2 --layer_type concat --multiscale True --rademacher True --batch_size 900 --test_batch_size 500 --save ../experiments_published/cnf_conditional_disentangle_cifar10_bs900_sratio_0_125_drop_0_5_run1 --seed 1 --lr 0.001 --conditional True --controlled_tol False --train_mode semisup --log_freq 10 --weight_y 0.5 --condition_ratio 0.125 --dropout_rate 0.5\n#",
"/tancode/repos/tan-ffjord/train_cnf_disentangle_cifar.py\nimport argparse\nimport os\nimport time\nimport numpy as np\n\nimport torch\nimport torch.optim as optim\nimport torchvision.datasets as dset\nimport torchvision.transforms as tforms\nfrom torchvision.utils import save_image\n\nimport lib.layers as layers\nimport lib.utils as utils\nimport lib.multiscale_parallel as multiscale_parallel\nimport lib.modules as modules\nimport lib.thops as thops\n\nfrom train_misc import standard_normal_logprob\nfrom train_misc import set_cnf_options, count_nfe, count_parameters, count_total_time\nfrom train_misc import add_spectral_norm, spectral_norm_power_iteration\nfrom train_misc import create_regularization_fns, get_regularization, append_regularization_to_log\n\nfrom tensorboardX import SummaryWriter\n\nimport glob\n\n# go fast boi!!\ntorch.backends.cudnn.benchmark = True\nSOLVERS = [\"dopri5\", \"bdf\", \"rk4\", \"midpoint\", 'adams', 'explicit_adams']\nparser = argparse.ArgumentParser(\"Continuous Normalizing Flow\")\nparser.add_argument(\"--data\", choices=[\"mnist\", \"svhn\", \"cifar10\", 'lsun_church'], type=str, default=\"mnist\")\nparser.add_argument(\"--dims\", type=str, default=\"8,32,32,8\")\nparser.add_argument(\"--strides\", type=str, default=\"2,2,1,-2,-2\")\nparser.add_argument(\"--num_blocks\", type=int, default=1, help='Number of stacked CNFs.')\n\nparser.add_argument(\"--conv\", type=eval, default=True, choices=[True, False])\nparser.add_argument(\n \"--layer_type\", type=str, default=\"ignore\",\n choices=[\"ignore\", \"concat\", \"concat_v2\", \"squash\", \"concatsquash\", \"concatcoord\", \"hyper\", \"blend\"]\n)\nparser.add_argument(\"--divergence_fn\", type=str, default=\"approximate\", choices=[\"brute_force\", \"approximate\"])\nparser.add_argument(\n \"--nonlinearity\", type=str, default=\"softplus\", choices=[\"tanh\", \"relu\", \"softplus\", \"elu\", \"swish\"]\n)\n\nparser.add_argument(\"--seed\", type=int, default=0)\n\nparser.add_argument('--solver', type=str, default='dopri5', choices=SOLVERS)\nparser.add_argument('--atol', type=float, default=1e-5)\nparser.add_argument('--rtol', type=float, default=1e-5)\nparser.add_argument(\"--step_size\", type=float, default=None, help=\"Optional fixed step size.\")\n\nparser.add_argument('--test_solver', type=str, default=None, choices=SOLVERS + [None])\nparser.add_argument('--test_atol', type=float, default=None)\nparser.add_argument('--test_rtol', type=float, default=None)\n\nparser.add_argument(\"--imagesize\", type=int, default=None)\nparser.add_argument(\"--alpha\", type=float, default=1e-6)\nparser.add_argument('--time_length', type=float, default=1.0)\nparser.add_argument('--train_T', type=eval, default=True)\n\nparser.add_argument(\"--num_epochs\", type=int, default=1000)\nparser.add_argument(\"--batch_size\", type=int, default=200)\nparser.add_argument(\n \"--batch_size_schedule\", type=str, default=\"\", help=\"Increases the batchsize at every given epoch, dash separated.\"\n)\nparser.add_argument(\"--test_batch_size\", type=int, default=200)\nparser.add_argument(\"--lr\", type=float, default=1e-3)\nparser.add_argument(\"--warmup_iters\", type=float, default=1000)\nparser.add_argument(\"--weight_decay\", type=float, default=0.0)\nparser.add_argument(\"--spectral_norm_niter\", type=int, default=10)\nparser.add_argument(\"--weight_y\", type=float, default=0.5)\n\nparser.add_argument(\"--add_noise\", type=eval, default=True, choices=[True, False])\nparser.add_argument(\"--batch_norm\", type=eval, default=False, choices=[True, False])\nparser.add_argument('--residual', type=eval, default=False, choices=[True, False])\nparser.add_argument('--autoencode', type=eval, default=False, choices=[True, False])\nparser.add_argument('--rademacher', type=eval, default=True, choices=[True, False])\nparser.add_argument('--spectral_norm', type=eval, default=False, choices=[True, False])\nparser.add_argument('--multiscale', type=eval, default=False, choices=[True, False])\nparser.add_argument('--parallel', type=eval, default=False, choices=[True, False])\nparser.add_argument('--conditional', type=eval, default=False, choices=[True, False])\nparser.add_argument('--controlled_tol', type=eval, default=False, choices=[True, False])\nparser.add_argument(\"--train_mode\", choices=[\"semisup\", \"sup\", \"unsup\"], type=str, default=\"semisup\")\nparser.add_argument(\"--condition_ratio\", type=float, default=0.5)\nparser.add_argument(\"--dropout_rate\", type=float, default=0.0)\n\n\n# Regularizations\nparser.add_argument('--l1int', type=float, default=None, help=\"int_t ||f||_1\")\nparser.add_argument('--l2int', type=float, default=None, help=\"int_t ||f||_2\")\nparser.add_argument('--dl2int', type=float, default=None, help=\"int_t ||f^T df/dt||_2\")\nparser.add_argument('--JFrobint', type=float, default=None, help=\"int_t ||df/dx||_F\")\nparser.add_argument('--JdiagFrobint', type=float, default=None, help=\"int_t ||df_i/dx_i||_F\")\nparser.add_argument('--JoffdiagFrobint', type=float, default=None, help=\"int_t ||df/dx - df_i/dx_i||_F\")\n\nparser.add_argument(\"--time_penalty\", type=float, default=0, help=\"Regularization on the end_time.\")\nparser.add_argument(\n \"--max_grad_norm\", type=float, default=1e10,\n help=\"Max norm of graidents (default is just stupidly high to avoid any clipping)\"\n)\n\nparser.add_argument(\"--begin_epoch\", type=int, default=1)\nparser.add_argument(\"--resume\", type=str, default=None)\nparser.add_argument(\"--save\", type=str, default=\"experiments/cnf\")\nparser.add_argument(\"--load_dir\", type=str, default=None)\nparser.add_argument(\"--resume_load\", type=int, default=1)\nparser.add_argument(\"--val_freq\", type=int, default=1)\nparser.add_argument(\"--log_freq\", type=int, default=1)\n\nargs = parser.parse_args()\n\nif args.controlled_tol:\n import lib.odenvp_conditional_tol as odenvp\nelse:\n import lib.odenvp_conditional as odenvp\n \n# set seed\ntorch.manual_seed(args.seed)\nnp.random.seed(args.seed)\n\n# logger\nutils.makedirs(args.save)\nlogger = utils.get_logger(logpath=os.path.join(args.save, 'logs'), filepath=os.path.abspath(__file__)) # write to log file\nwriter = SummaryWriter(os.path.join(args.save, 'tensorboard')) # write to tensorboard\n\nif args.layer_type == \"blend\":\n logger.info(\"!! Setting time_length from None to 1.0 due to use of Blend layers.\")\n args.time_length = 1.0\n\nlogger.info(args)\n\n\ndef add_noise(x):\n \"\"\"\n [0, 1] -> [0, 255] -> add noise -> [0, 1]\n \"\"\"\n if args.add_noise:\n noise = x.new().resize_as_(x).uniform_()\n x = x * 255 + noise\n x = x / 256\n return x\n\n\ndef update_lr(optimizer, itr):\n iter_frac = min(float(itr + 1) / max(args.warmup_iters, 1), 1.0)\n lr = args.lr * iter_frac\n for param_group in optimizer.param_groups:\n param_group[\"lr\"] = lr\n\n\ndef get_train_loader(train_set, epoch):\n if args.batch_size_schedule != \"\":\n epochs = [0] + list(map(int, args.batch_size_schedule.split(\"-\")))\n n_passed = sum(np.array(epochs) <= epoch)\n current_batch_size = int(args.batch_size * n_passed)\n else:\n current_batch_size = args.batch_size\n train_loader = torch.utils.data.DataLoader(\n dataset=train_set, batch_size=current_batch_size, shuffle=True, drop_last=True, pin_memory=True\n )\n logger.info(\"===> Using batch size {}. Total {} iterations/epoch.\".format(current_batch_size, len(train_loader)))\n return train_loader\n\n\ndef get_dataset(args):\n trans = lambda im_size: tforms.Compose([tforms.Resize(im_size), tforms.ToTensor(), add_noise])\n\n if args.data == \"mnist\":\n im_dim = 1\n im_size = 28 if args.imagesize is None else args.imagesize\n train_set = dset.MNIST(root=\"../data\", train=True, transform=trans(im_size), download=True)\n test_set = dset.MNIST(root=\"../data\", train=False, transform=trans(im_size), download=True)\n elif args.data == \"svhn\":\n im_dim = 3\n im_size = 32 if args.imagesize is None else args.imagesize\n train_set = dset.SVHN(root=\"../data\", split=\"train\", transform=trans(im_size), download=True)\n test_set = dset.SVHN(root=\"../data\", split=\"test\", transform=trans(im_size), download=True)\n elif args.data == \"cifar10\":\n im_dim = 3\n im_size = 32 if args.imagesize is None else args.imagesize\n train_set = dset.CIFAR10(\n root=\"../data\", train=True, transform=tforms.Compose([\n tforms.Resize(im_size),\n tforms.RandomHorizontalFlip(),\n tforms.ToTensor(),\n add_noise,\n ]), download=True\n )\n test_set = dset.CIFAR10(root=\"../data\", train=False, transform=trans(im_size), download=True)\n elif args.data == 'celeba':\n im_dim = 3\n im_size = 64 if args.imagesize is None else args.imagesize\n train_set = dset.CelebA(\n train=True, transform=tforms.Compose([\n tforms.ToPILImage(),\n tforms.Resize(im_size),\n tforms.RandomHorizontalFlip(),\n tforms.ToTensor(),\n add_noise,\n ])\n )\n test_set = dset.CelebA(\n train=False, transform=tforms.Compose([\n tforms.ToPILImage(),\n tforms.Resize(im_size),\n tforms.ToTensor(),\n add_noise,\n ])\n )\n elif args.data == 'lsun_church':\n im_dim = 3\n im_size = 64 if args.imagesize is None else args.imagesize\n train_set = dset.LSUN(\n '../data', ['church_outdoor_train'], transform=tforms.Compose([\n tforms.Resize(96),\n tforms.RandomCrop(64),\n tforms.Resize(im_size),\n tforms.ToTensor(),\n add_noise,\n ])\n )\n test_set = dset.LSUN(\n '../data', ['church_outdoor_val'], transform=tforms.Compose([\n tforms.Resize(96),\n tforms.RandomCrop(64),\n tforms.Resize(im_size),\n tforms.ToTensor(),\n add_noise,\n ])\n ) \n elif args.data == 'imagenet_64':\n im_dim = 3\n im_size = 64 if args.imagesize is None else args.imagesize\n train_set = dset.ImageFolder(\n train=True, transform=tforms.Compose([\n tforms.ToPILImage(),\n tforms.Resize(im_size),\n tforms.RandomHorizontalFlip(),\n tforms.ToTensor(),\n add_noise,\n ])\n )\n test_set = dset.ImageFolder(\n train=False, transform=tforms.Compose([\n tforms.ToPILImage(),\n tforms.Resize(im_size),\n tforms.ToTensor(),\n add_noise,\n ])\n )\n \n data_shape = (im_dim, im_size, im_size)\n if not args.conv:\n data_shape = (im_dim * im_size * im_size,)\n\n test_loader = torch.utils.data.DataLoader(\n dataset=test_set, batch_size=args.test_batch_size, shuffle=False, drop_last=True\n )\n return train_set, test_loader, data_shape\n\n\ndef compute_bits_per_dim(x, model):\n zero = torch.zeros(x.shape[0], 1).to(x)\n\n # Don't use data parallelize if batch size is small.\n # if x.shape[0] < 200:\n # model = model.module\n \n z, delta_logp = model(x, zero) # run model forward\n\n logpz = standard_normal_logprob(z).view(z.shape[0], -1).sum(1, keepdim=True) # logp(z)\n logpx = logpz - delta_logp\n\n logpx_per_dim = torch.sum(logpx) / x.nelement() # averaged over batches\n bits_per_dim = -(logpx_per_dim - np.log(256)) / np.log(2)\n\n return bits_per_dim\n\ndef compute_bits_per_dim_conditional(x, y, model):\n zero = torch.zeros(x.shape[0], 1).to(x)\n y_onehot = thops.onehot(y, num_classes=model.module.y_class).to(x)\n\n # Don't use data parallelize if batch size is small.\n # if x.shape[0] < 200:\n # model = model.module\n \n z, delta_logp = model(x, zero) # run model forward\n \n dim_sup = int(args.condition_ratio * np.prod(z.size()[1:]))\n \n # prior\n mean, logs = model.module._prior(y_onehot)\n \n # compute the conditional nll\n logpz_sup = modules.GaussianDiag.logp(mean, logs, z[:, 0:dim_sup]).view(-1,1) # logp(z)_sup\n logpz_unsup = standard_normal_logprob(z[:, dim_sup:]).view(z.shape[0], -1).sum(1, keepdim=True)\n logpz = logpz_sup + logpz_unsup\n logpx = logpz - delta_logp\n\n logpx_per_dim = torch.sum(logpx) / x.nelement() # averaged over batches\n bits_per_dim = -(logpx_per_dim - np.log(256)) / np.log(2)\n \n # compute the marginal nll\n logpz_sup_marginal = []\n \n for indx in range(model.module.y_class):\n y_i = torch.full_like(y, indx).to(y)\n y_onehot = thops.onehot(y_i, num_classes=model.module.y_class).to(x)\n # prior\n mean, logs = model.module._prior(y_onehot)\n logpz_sup_marginal.append(modules.GaussianDiag.logp(mean, logs, z[:, 0:dim_sup]).view(-1,1) - torch.log(torch.tensor(model.module.y_class).to(z))) # logp(z)_sup\n \n logpz_sup_marginal = torch.cat(logpz_sup_marginal,dim=1)\n logpz_sup_marginal = torch.logsumexp(logpz_sup_marginal, 1, keepdim=True)\n \n logpz_marginal = logpz_sup_marginal + logpz_unsup\n \n logpx_marginal = logpz_marginal - delta_logp\n\n logpx_per_dim_marginal = torch.sum(logpx_marginal) / x.nelement() # averaged over batches\n bits_per_dim_marginal = -(logpx_per_dim_marginal - np.log(256)) / np.log(2)\n \n # dropout\n if args.dropout_rate > 0:\n zsup = model.module.dropout(z[:, 0:dim_sup])\n else:\n zsup = z[:, 0:dim_sup]\n \n # compute xentropy loss\n y_logits = model.module.project_class(zsup)\n loss_xent = model.module.loss_class(y_logits, y.to(x.get_device()))\n y_predicted = np.argmax(y_logits.cpu().detach().numpy(), axis=1)\n\n return bits_per_dim, loss_xent, y_predicted, bits_per_dim_marginal\n\ndef compute_marginal_bits_per_dim(x, y, model):\n zero = torch.zeros(x.shape[0], 1).to(x)\n\n # Don't use data parallelize if batch size is small.\n # if x.shape[0] < 200:\n # model = model.module\n \n z, delta_logp = model(x, zero) # run model forward\n \n dim_sup = int(args.condition_ratio * np.prod(z.size()[1:]))\n \n logpz_sup = []\n \n for indx in range(model.module.y_class):\n y_i = torch.full_like(y, indx).to(y)\n y_onehot = thops.onehot(y_i, num_classes=model.module.y_class).to(x)\n # prior\n mean, logs = model.module._prior(y_onehot)\n logpz_sup.append(modules.GaussianDiag.logp(mean, logs, z[:, 0:dim_sup]).view(-1,1) - torch.log(torch.tensor(model.module.y_class).to(z))) # logp(z)_sup\n \n logpz_sup = torch.cat(logpz_sup,dim=1)\n logpz_sup = torch.logsumexp(logpz_sup, 1, keepdim=True)\n \n logpz_unsup = standard_normal_logprob(z[:, dim_sup:]).view(z.shape[0], -1).sum(1, keepdim=True)\n logpz = logpz_sup + logpz_unsup\n \n logpx = logpz - delta_logp\n\n logpx_per_dim = torch.sum(logpx) / x.nelement() # averaged over batches\n bits_per_dim = -(logpx_per_dim - np.log(256)) / np.log(2)\n\n return bits_per_dim\n\ndef create_model(args, data_shape, regularization_fns):\n hidden_dims = tuple(map(int, args.dims.split(\",\")))\n strides = tuple(map(int, args.strides.split(\",\")))\n\n if args.multiscale:\n model = odenvp.ODENVP(\n (args.batch_size, *data_shape),\n n_blocks=args.num_blocks,\n intermediate_dims=hidden_dims,\n nonlinearity=args.nonlinearity,\n alpha=args.alpha,\n cnf_kwargs={\"T\": args.time_length, \"train_T\": args.train_T, \"regularization_fns\": regularization_fns, \"solver\": args.solver, \"atol\": args.atol, \"rtol\": args.rtol},\n condition_ratio=args.condition_ratio,\n dropout_rate=args.dropout_rate,)\n elif args.parallel:\n model = multiscale_parallel.MultiscaleParallelCNF(\n (args.batch_size, *data_shape),\n n_blocks=args.num_blocks,\n intermediate_dims=hidden_dims,\n alpha=args.alpha,\n time_length=args.time_length,\n )\n else:\n if args.autoencode:\n\n def build_cnf():\n autoencoder_diffeq = layers.AutoencoderDiffEqNet(\n hidden_dims=hidden_dims,\n input_shape=data_shape,\n strides=strides,\n conv=args.conv,\n layer_type=args.layer_type,\n nonlinearity=args.nonlinearity,\n )\n odefunc = layers.AutoencoderODEfunc(\n autoencoder_diffeq=autoencoder_diffeq,\n divergence_fn=args.divergence_fn,\n residual=args.residual,\n rademacher=args.rademacher,\n )\n cnf = layers.CNF(\n odefunc=odefunc,\n T=args.time_length,\n regularization_fns=regularization_fns,\n solver=args.solver,\n )\n return cnf\n else:\n\n def build_cnf():\n diffeq = layers.ODEnet(\n hidden_dims=hidden_dims,\n input_shape=data_shape,\n strides=strides,\n conv=args.conv,\n layer_type=args.layer_type,\n nonlinearity=args.nonlinearity,\n )\n odefunc = layers.ODEfunc(\n diffeq=diffeq,\n divergence_fn=args.divergence_fn,\n residual=args.residual,\n rademacher=args.rademacher,\n )\n cnf = layers.CNF(\n odefunc=odefunc,\n T=args.time_length,\n train_T=args.train_T,\n regularization_fns=regularization_fns,\n solver=args.solver,\n )\n return cnf\n\n chain = [layers.LogitTransform(alpha=args.alpha)] if args.alpha > 0 else [layers.ZeroMeanTransform()]\n chain = chain + [build_cnf() for _ in range(args.num_blocks)]\n if args.batch_norm:\n chain.append(layers.MovingBatchNorm2d(data_shape[0]))\n model = layers.SequentialFlow(chain)\n return model\n\n\nif __name__ == \"__main__\":\n\n # get deivce\n device = torch.device(\"cuda:0\" if torch.cuda.is_available() else \"cpu\")\n cvt = lambda x: x.type(torch.float32).to(device, non_blocking=True)\n\n # load dataset\n train_set, test_loader, data_shape = get_dataset(args)\n\n # build model\n regularization_fns, regularization_coeffs = create_regularization_fns(args)\n model = create_model(args, data_shape, regularization_fns)\n\n if args.spectral_norm: add_spectral_norm(model, logger)\n set_cnf_options(args, model)\n\n logger.info(model)\n logger.info(\"Number of trainable parameters: {}\".format(count_parameters(model)))\n \n writer.add_text('info', \"Number of trainable parameters: {}\".format(count_parameters(model)))\n\n # optimizer\n optimizer = optim.Adam(model.parameters(), lr=args.lr, weight_decay=args.weight_decay)\n \n # set initial iter\n itr = 1\n \n # set the meters\n time_epoch_meter = utils.RunningAverageMeter(0.97)\n time_meter = utils.RunningAverageMeter(0.97)\n loss_meter = utils.RunningAverageMeter(0.97) # track total loss\n nll_meter = utils.RunningAverageMeter(0.97) # track negative log-likelihood\n xent_meter = utils.RunningAverageMeter(0.97) # track xentropy score\n error_meter = utils.RunningAverageMeter(0.97) # track error score\n steps_meter = utils.RunningAverageMeter(0.97)\n grad_meter = utils.RunningAverageMeter(0.97)\n tt_meter = utils.RunningAverageMeter(0.97)\n nll_marginal_meter = utils.RunningAverageMeter(0.97) # track negative marginal log-likelihood\n \n if args.load_dir is not None:\n filelist = glob.glob(os.path.join(args.load_dir,\"*epoch_*_checkpt.pth\"))\n all_indx = []\n for infile in sorted(filelist):\n all_indx.append(int(infile.split('_')[-2]))\n \n indxmax = max(all_indx)\n indxstart = max(1, args.resume_load)\n \n for i in range(indxstart,indxmax+1):\n model = create_model(args, data_shape, regularization_fns)\n infile = os.path.join(args.load_dir,\"epoch_%i_checkpt.pth\"%i)\n print(infile)\n checkpt = torch.load(infile, map_location=lambda storage, loc: storage)\n model.load_state_dict(checkpt[\"state_dict\"])\n \n if torch.cuda.is_available():\n model = torch.nn.DataParallel(model).cuda()\n \n model.eval()\n \n with torch.no_grad():\n logger.info(\"validating...\")\n losses = []\n for (x, y) in test_loader:\n if args.data == \"colormnist\":\n y = y[0]\n \n if not args.conv:\n x = x.view(x.shape[0], -1)\n x = cvt(x)\n nll = compute_marginal_bits_per_dim(x, y, model)\n losses.append(nll.cpu().numpy())\n \n loss = np.mean(losses)\n writer.add_scalars('nll_marginal', {'validation': loss}, i)\n \n raise SystemExit(0)\n \n # restore parameters\n if args.resume is not None:\n checkpt = torch.load(args.resume, map_location=lambda storage, loc: storage)\n model.load_state_dict(checkpt[\"state_dict\"])\n if \"optim_state_dict\" in checkpt.keys():\n optimizer.load_state_dict(checkpt[\"optim_state_dict\"])\n # Manually move optimizer state to device.\n for state in optimizer.state.values():\n for k, v in state.items():\n if torch.is_tensor(v):\n state[k] = cvt(v)\n args.begin_epoch = checkpt['epoch'] + 1\n itr = checkpt['iter'] + 1\n time_epoch_meter.set(checkpt['epoch_time_avg'])\n time_meter.set(checkpt['time_train'])\n loss_meter.set(checkpt['loss_train'])\n nll_meter.set(checkpt['bits_per_dim_train'])\n xent_meter.set(checkpt['xent_train'])\n error_meter.set(checkpt['error_train'])\n steps_meter.set(checkpt['nfe_train'])\n grad_meter.set(checkpt['grad_train'])\n tt_meter.set(checkpt['total_time_train'])\n nll_marginal_meter.set(checkpt['bits_per_dim_marginal_train'])\n\n if torch.cuda.is_available():\n model = torch.nn.DataParallel(model).cuda()\n\n # For visualization.\n if args.conditional:\n dim_unsup = int((1.0 - args.condition_ratio) * np.prod(data_shape))\n fixed_y = torch.from_numpy(np.arange(model.module.y_class)).repeat(model.module.y_class).type(torch.long).to(device, non_blocking=True)\n fixed_y_onehot = thops.onehot(fixed_y, num_classes=model.module.y_class)\n with torch.no_grad():\n mean, logs = model.module._prior(fixed_y_onehot)\n fixed_z_sup = modules.GaussianDiag.sample(mean, logs)\n fixed_z_unsup = cvt(torch.randn(model.module.y_class**2, dim_unsup))\n fixed_z = torch.cat((fixed_z_sup, fixed_z_unsup),1)\n else:\n fixed_z = cvt(torch.randn(100, *data_shape))\n \n\n if args.spectral_norm and not args.resume: spectral_norm_power_iteration(model, 500)\n\n best_loss_nll = float(\"inf\")\n best_error_score = float(\"inf\")\n \n for epoch in range(args.begin_epoch, args.num_epochs + 1):\n start_epoch = time.time()\n model.train()\n train_loader = get_train_loader(train_set, epoch)\n for _, (x, y) in enumerate(train_loader):\n start = time.time()\n update_lr(optimizer, itr)\n optimizer.zero_grad()\n\n if not args.conv:\n x = x.view(x.shape[0], -1)\n\n # cast data and move to device\n x = cvt(x)\n \n # compute loss\n if args.conditional:\n loss_nll, loss_xent, y_predicted, loss_nll_marginal = compute_bits_per_dim_conditional(x, y, model)\n if args.train_mode == \"semisup\":\n loss = loss_nll + args.weight_y * loss_xent\n elif args.train_mode == \"sup\":\n loss = loss_xent\n elif args.train_mode == \"unsup\":\n loss = loss_nll\n else:\n raise ValueError('Choose supported train_mode: semisup, sup, unsup')\n error_score = 1. - np.mean(y_predicted.astype(int) == y.numpy()) \n \n else:\n loss = compute_bits_per_dim(x, model)\n loss_nll, loss_xent, error_score = loss, 0., 0.\n \n if regularization_coeffs:\n reg_states = get_regularization(model, regularization_coeffs)\n reg_loss = sum(\n reg_state * coeff for reg_state, coeff in zip(reg_states, regularization_coeffs) if coeff != 0\n )\n loss = loss + reg_loss\n total_time = count_total_time(model)\n loss = loss + total_time * args.time_penalty\n\n loss.backward()\n grad_norm = torch.nn.utils.clip_grad_norm_(model.parameters(), args.max_grad_norm)\n\n optimizer.step()\n\n if args.spectral_norm: spectral_norm_power_iteration(model, args.spectral_norm_niter)\n \n time_meter.update(time.time() - start)\n loss_meter.update(loss.item())\n nll_meter.update(loss_nll.item())\n if args.conditional:\n xent_meter.update(loss_xent.item())\n else:\n xent_meter.update(loss_xent)\n error_meter.update(error_score)\n steps_meter.update(count_nfe(model))\n grad_meter.update(grad_norm)\n tt_meter.update(total_time)\n nll_marginal_meter.update(loss_nll_marginal.item())\n \n # write to tensorboard\n writer.add_scalars('time', {'train_iter': time_meter.val}, itr)\n writer.add_scalars('loss', {'train_iter': loss_meter.val}, itr)\n writer.add_scalars('bits_per_dim', {'train_iter': nll_meter.val}, itr)\n writer.add_scalars('xent', {'train_iter': xent_meter.val}, itr)\n writer.add_scalars('error', {'train_iter': error_meter.val}, itr)\n writer.add_scalars('nfe', {'train_iter': steps_meter.val}, itr)\n writer.add_scalars('grad', {'train_iter': grad_meter.val}, itr)\n writer.add_scalars('total_time', {'train_iter': tt_meter.val}, itr)\n writer.add_scalars('bits_per_dim_marginal', {'train_iter': nll_marginal_meter.val}, itr)\n\n if itr % args.log_freq == 0:\n log_message = (\n \"Iter {:04d} | Time {:.4f}({:.4f}) | Bit/dim {:.4f}({:.4f}) | Xent {:.4f}({:.4f}) | Loss {:.4f}({:.4f}) | Error {:.4f}({:.4f}) \"\n \"Steps {:.0f}({:.2f}) | Grad Norm {:.4f}({:.4f}) | Total Time {:.2f}({:.2f})\".format(\n itr, time_meter.val, time_meter.avg, nll_meter.val, nll_meter.avg, xent_meter.val, xent_meter.avg, loss_meter.val, loss_meter.avg, error_meter.val, error_meter.avg, steps_meter.val, steps_meter.avg, grad_meter.val, grad_meter.avg, tt_meter.val, tt_meter.avg\n )\n )\n if regularization_coeffs:\n log_message = append_regularization_to_log(log_message, regularization_fns, reg_states)\n logger.info(log_message)\n writer.add_text('info', log_message, itr)\n\n itr += 1\n \n # compute test loss\n model.eval()\n if epoch % args.val_freq == 0:\n with torch.no_grad():\n # write to tensorboard\n writer.add_scalars('time', {'train_epoch': time_meter.avg}, epoch)\n writer.add_scalars('loss', {'train_epoch': loss_meter.avg}, epoch)\n writer.add_scalars('bits_per_dim', {'train_epoch': nll_meter.avg}, epoch)\n writer.add_scalars('xent', {'train_epoch': xent_meter.avg}, epoch)\n writer.add_scalars('error', {'train_epoch': error_meter.avg}, epoch)\n writer.add_scalars('nfe', {'train_epoch': steps_meter.avg}, epoch)\n writer.add_scalars('grad', {'train_epoch': grad_meter.avg}, epoch)\n writer.add_scalars('total_time', {'train_epoch': tt_meter.avg}, epoch)\n writer.add_scalars('bits_per_dim_marginal', {'train_epoch': nll_marginal_meter.avg}, epoch)\n \n start = time.time()\n logger.info(\"validating...\")\n writer.add_text('info', \"validating...\", epoch)\n losses_nll = []; losses_xent = []; losses = []\n total_correct = 0\n \n for (x, y) in test_loader:\n if not args.conv:\n x = x.view(x.shape[0], -1)\n x = cvt(x)\n if args.conditional:\n loss_nll, loss_xent, y_predicted, loss_nll_marginal = compute_bits_per_dim_conditional(x, y, model)\n if args.train_mode == \"semisup\":\n loss = loss_nll + args.weight_y * loss_xent\n elif args.train_mode == \"sup\":\n loss = loss_xent\n elif args.train_mode == \"unsup\":\n loss = loss_nll\n else:\n raise ValueError('Choose supported train_mode: semisup, sup, unsup')\n total_correct += np.sum(y_predicted.astype(int) == y.numpy())\n else:\n loss = compute_bits_per_dim(x, model)\n loss_nll, loss_xent = loss, 0.\n losses_nll.append(loss_nll.cpu().numpy()); losses.append(loss.cpu().numpy())\n if args.conditional: \n losses_xent.append(loss_xent.cpu().numpy())\n else:\n losses_xent.append(loss_xent)\n \n loss_nll = np.mean(losses_nll); loss_xent = np.mean(losses_xent); loss = np.mean(losses)\n error_score = 1. - total_correct / len(test_loader.dataset)\n time_epoch_meter.update(time.time() - start_epoch)\n \n # write to tensorboard\n test_time_spent = time.time() - start\n writer.add_scalars('time', {'validation': test_time_spent}, epoch)\n writer.add_scalars('epoch_time', {'validation': time_epoch_meter.val}, epoch)\n writer.add_scalars('bits_per_dim', {'validation': loss_nll}, epoch)\n writer.add_scalars('xent', {'validation': loss_xent}, epoch)\n writer.add_scalars('loss', {'validation': loss}, epoch)\n writer.add_scalars('error', {'validation': error_score}, epoch)\n writer.add_scalars('bits_per_dim_marginal', {'validation': loss_nll_marginal}, epoch)\n \n log_message = \"Epoch {:04d} | Time {:.4f}, Epoch Time {:.4f}({:.4f}), Bit/dim {:.4f}(best: {:.4f}), Xent {:.4f}, Loss {:.4f}, Error {:.4f}(best: {:.4f})\".format(epoch, time.time() - start, time_epoch_meter.val, time_epoch_meter.avg, loss_nll, best_loss_nll, loss_xent, loss, error_score, best_error_score)\n logger.info(log_message)\n writer.add_text('info', log_message, epoch)\n \n for name, param in model.named_parameters():\n writer.add_histogram(name, param.clone().cpu().data.numpy(), epoch)\n \n \n utils.makedirs(args.save)\n torch.save({\n \"args\": args,\n \"state_dict\": model.module.state_dict() if torch.cuda.is_available() else model.state_dict(),\n \"optim_state_dict\": optimizer.state_dict(),\n \"epoch\": epoch,\n \"iter\": itr-1,\n \"error\": error_score,\n \"loss\": loss,\n \"xent\": loss_xent,\n \"bits_per_dim\": loss_nll,\n \"bits_per_dim_marginal\": loss_nll_marginal,\n \"best_bits_per_dim\": best_loss_nll,\n \"best_error_score\": best_error_score,\n \"epoch_time\": time_epoch_meter.val,\n \"epoch_time_avg\": time_epoch_meter.avg,\n \"time\": test_time_spent,\n \"error_train\": error_meter.avg,\n \"loss_train\": loss_meter.avg,\n \"xent_train\": xent_meter.avg,\n \"bits_per_dim_train\": nll_meter.avg,\n \"bits_per_dim_marginal_train\": nll_marginal_meter.avg,\n \"total_time_train\": tt_meter.avg,\n \"time_train\": time_meter.avg,\n \"nfe_train\": steps_meter.avg,\n \"grad_train\": grad_meter.avg,\n }, os.path.join(args.save, \"epoch_%i_checkpt.pth\"%epoch))\n \n torch.save({\n \"args\": args,\n \"state_dict\": model.module.state_dict() if torch.cuda.is_available() else model.state_dict(),\n \"optim_state_dict\": optimizer.state_dict(),\n \"epoch\": epoch,\n \"iter\": itr-1,\n \"error\": error_score,\n \"loss\": loss,\n \"xent\": loss_xent,\n \"bits_per_dim\": loss_nll,\n \"best_bits_per_dim\": best_loss_nll,\n \"bits_per_dim_marginal\": loss_nll_marginal,\n \"best_error_score\": best_error_score,\n \"epoch_time\": time_epoch_meter.val,\n \"epoch_time_avg\": time_epoch_meter.avg,\n \"time\": test_time_spent,\n \"error_train\": error_meter.avg,\n \"loss_train\": loss_meter.avg,\n \"xent_train\": xent_meter.avg,\n \"bits_per_dim_train\": nll_meter.avg,\n \"bits_per_dim_marginal_train\": nll_marginal_meter.avg,\n \"total_time_train\": tt_meter.avg,\n \"time_train\": time_meter.avg,\n \"nfe_train\": steps_meter.avg,\n \"grad_train\": grad_meter.avg,\n }, os.path.join(args.save, \"current_checkpt.pth\"))\n \n if loss_nll < best_loss_nll:\n best_loss_nll = loss_nll\n utils.makedirs(args.save)\n torch.save({\n \"args\": args,\n \"state_dict\": model.module.state_dict() if torch.cuda.is_available() else model.state_dict(),\n \"optim_state_dict\": optimizer.state_dict(),\n \"epoch\": epoch,\n \"iter\": itr-1,\n \"error\": error_score,\n \"loss\": loss,\n \"xent\": loss_xent,\n \"bits_per_dim\": loss_nll,\n \"bits_per_dim_marginal\": loss_nll_marginal,\n \"best_bits_per_dim\": best_loss_nll,\n \"best_error_score\": best_error_score,\n \"epoch_time\": time_epoch_meter.val,\n \"epoch_time_avg\": time_epoch_meter.avg,\n \"time\": test_time_spent,\n \"error_train\": error_meter.avg,\n \"loss_train\": loss_meter.avg,\n \"xent_train\": xent_meter.avg,\n \"bits_per_dim_train\": nll_meter.avg,\n \"bits_per_dim_marginal_train\": nll_marginal_meter.avg,\n \"total_time_train\": tt_meter.avg,\n \"time_train\": time_meter.avg,\n \"nfe_train\": steps_meter.avg,\n \"grad_train\": grad_meter.avg,\n }, os.path.join(args.save, \"best_nll_checkpt.pth\"))\n \n if args.conditional:\n if error_score < best_error_score:\n best_error_score = error_score\n utils.makedirs(args.save)\n torch.save({\n \"args\": args,\n \"state_dict\": model.module.state_dict() if torch.cuda.is_available() else model.state_dict(),\n \"optim_state_dict\": optimizer.state_dict(),\n \"epoch\": epoch,\n \"iter\": itr-1,\n \"error\": error_score,\n \"loss\": loss,\n \"xent\": loss_xent,\n \"bits_per_dim\": loss_nll,\n \"bits_per_dim_marginal\": loss_nll_marginal,\n \"best_bits_per_dim\": best_loss_nll,\n \"best_error_score\": best_error_score,\n \"epoch_time\": time_epoch_meter.val,\n \"epoch_time_avg\": time_epoch_meter.avg,\n \"time\": test_time_spent,\n \"error_train\": error_meter.avg,\n \"loss_train\": loss_meter.avg,\n \"xent_train\": xent_meter.avg,\n \"bits_per_dim_train\": nll_meter.avg,\n \"bits_per_dim_marginal_train\": nll_marginal_meter.avg,\n \"total_time_train\": tt_meter.avg,\n \"time_train\": time_meter.avg,\n \"nfe_train\": steps_meter.avg,\n \"grad_train\": grad_meter.avg,\n }, os.path.join(args.save, \"best_error_checkpt.pth\"))\n \n\n # visualize samples and density\n with torch.no_grad():\n fig_filename = os.path.join(args.save, \"figs\", \"{:04d}.jpg\".format(epoch))\n utils.makedirs(os.path.dirname(fig_filename))\n generated_samples = model(fixed_z, reverse=True).view(-1, *data_shape)\n save_image(generated_samples, fig_filename, nrow=10)\n if args.data == \"mnist\":\n writer.add_images('generated_images', generated_samples.repeat(1,3,1,1), epoch)\n else:\n writer.add_images('generated_images', generated_samples.repeat(1,1,1,1), epoch)\n\nNamespace(JFrobint=None, JdiagFrobint=None, JoffdiagFrobint=None, add_noise=True, alpha=1e-06, atol=1e-05, autoencode=False, batch_norm=False, batch_size=900, batch_size_schedule='', begin_epoch=1, condition_ratio=0.125, conditional=True, controlled_tol=False, conv=True, data='cifar10', dims='64,64,64', divergence_fn='approximate', dl2int=None, dropout_rate=0.5, imagesize=None, l1int=None, l2int=None, layer_type='concat', load_dir=None, log_freq=10, lr=0.001, max_grad_norm=10000000000.0, multiscale=True, nonlinearity='softplus', num_blocks=2, num_epochs=1000, parallel=False, rademacher=True, residual=False, resume=None, resume_load=1, rtol=1e-05, save='../experiments_published/cnf_conditional_disentangle_cifar10_bs900_sratio_0_125_drop_0_5_run1', seed=1, solver='dopri5', spectral_norm=False, spectral_norm_niter=10, step_size=None, strides='1,1,1,1', test_atol=None, test_batch_size=500, test_rtol=None, test_solver=None, time_length=1.0, time_penalty=0, train_T=True, train_mode='semisup', val_freq=1, warmup_iters=1000, weight_decay=0.0, weight_y=0.5)\n"
]
]
] |
[
"code"
] |
[
[
"code",
"code"
]
] |
4acfdbfdeef432490101b39995d5f5ada2e0da94
| 45,855 |
ipynb
|
Jupyter Notebook
|
DeepLearning/ipython(guide)/classification_1_with_binary_CNN.ipynb
|
ZeinabTaghavi/text_multi_label_classification
|
dbbeafab16e201bdaed56c581562be68ddaeba7c
|
[
"MIT"
] | null | null | null |
DeepLearning/ipython(guide)/classification_1_with_binary_CNN.ipynb
|
ZeinabTaghavi/text_multi_label_classification
|
dbbeafab16e201bdaed56c581562be68ddaeba7c
|
[
"MIT"
] | null | null | null |
DeepLearning/ipython(guide)/classification_1_with_binary_CNN.ipynb
|
ZeinabTaghavi/text_multi_label_classification
|
dbbeafab16e201bdaed56c581562be68ddaeba7c
|
[
"MIT"
] | null | null | null | 45,855 | 45,855 | 0.64137 |
[
[
[
"A neural network consist of cnn layer (Kim,2014) and 4 fully connected layers.\r\n\r\nSource: https://github.com/jojonki/cnn-for-sentence-classification\r\n\r\n\r\n\r\n",
"_____no_output_____"
]
],
[
[
"from google.colab import drive\ndrive.mount('/content/drive')",
"Mounted at /content/drive\n"
],
[
"import os\nos.chdir('/content/drive/MyDrive/sharif/DeepLearning/ipython(guide)')",
"_____no_output_____"
],
[
"import numpy as np\nimport codecs\nimport os\nimport random\nimport pandas\nfrom keras import backend as K\nfrom keras.models import Model\nfrom keras.layers.embeddings import Embedding\nfrom keras.layers import Input, Dense, Lambda, Permute, Dropout\nfrom keras.layers import Conv2D, MaxPooling1D\nfrom keras.optimizers import SGD\nimport ast\nimport re\nfrom sklearn.preprocessing import MultiLabelBinarizer\nfrom sklearn.model_selection import train_test_split\nimport gensim\nfrom keras.models import load_model\nfrom keras.callbacks import EarlyStopping, ModelCheckpoint",
"_____no_output_____"
],
[
"limit_number = 750\ndata = pandas.read_csv('../Data/limited_to_'+str(limit_number)+'.csv',index_col=0,converters={'body': eval})\ndata = data.dropna().reset_index(drop=True)\nX = data[\"body\"].values.tolist()\ny = pandas.read_csv('../Data/limited_to_'+str(limit_number)+'.csv')\nlabels = []\ntag=[]\nfor item in y['tag']:\n labels += [i for i in re.sub('\\\"|\\[|\\]|\\'| |=','',item.lower()).split(\",\") if i!='' and i!=' ']\n tag.append([i for i in re.sub('\\\"|\\[|\\]|\\'| |=','',item.lower()).split(\",\") if i!='' and i!=' '])\nlabels = list(set(labels))\nmlb = MultiLabelBinarizer()\nY=mlb.fit_transform(tag)",
"_____no_output_____"
],
[
"len(labels)",
"_____no_output_____"
],
[
"sentence_maxlen = max(map(len, (d for d in X)))\nprint('sentence maxlen', sentence_maxlen)",
"sentence maxlen 300\n"
],
[
"freq_dist = pandas.read_csv('../Data/FreqDist_sorted.csv',index_col=False)\nvocab=[]\nfor item in freq_dist[\"word\"]:\n try:\n word=re.sub(r\"[\\u200c-\\u200f]\",\"\",item.replace(\" \",\"\"))\n vocab.append(word)\n except:\n pass\n \nprint(vocab[10])",
"زبان\n"
],
[
"vocab = sorted(vocab)\nvocab_size = len(vocab)",
"_____no_output_____"
],
[
"print('vocab size', len(vocab))\nw2i = {w:i for i,w in enumerate(vocab)}\n# i2w = {i:w for i,w in enumerate(vocab)}\nprint(w2i[\"زبان\"])",
"vocab size 225345\n129280\n"
],
[
"def vectorize(data, sentence_maxlen, w2i):\n vec_data = []\n \n for d in data:\n \n vec = [w2i[w] for w in d if w in w2i]\n pad_len = max(0, sentence_maxlen - len(vec))\n vec += [0] * pad_len\n vec_data.append(vec)\n # print(d)\n \n vec_data = np.array(vec_data)\n \n return vec_data\n\nvecX = vectorize(X, sentence_maxlen, w2i)\nvecY=Y",
"_____no_output_____"
],
[
"X_train, X_test, y_train, y_test = train_test_split(vecX, vecY, test_size=0.2)\nX_train, X_val, y_train, y_val = train_test_split(X_train, y_train, test_size=0.25)\nprint('train: ', X_train.shape , '\\ntest: ', X_test.shape , '\\nval: ', X_val.shape ,\"\\ny_tain:\",y_train.shape )\n# print(vecX[0])",
"train: (12935, 300) \ntest: (4312, 300) \nval: (4312, 300) \ny_tain: (12935, 78)\n"
],
[
"embd_dim = 300\n",
"_____no_output_____"
]
],
[
[
"# ***If the word2vec model is not generated before, we should run the next block.***",
"_____no_output_____"
]
],
[
[
"# embed_model = gensim.models.Word2Vec(X, size=embd_dim, window=5, min_count=5)\r\n# embed_model.save('word2vec_model')",
"_____no_output_____"
]
],
[
[
"# ***Otherwise, we can run the next block.***",
"_____no_output_____"
]
],
[
[
"embed_model=gensim.models.Word2Vec.load('word2vec_model')",
"_____no_output_____"
],
[
"word2vec_embd_w = np.zeros((vocab_size, embd_dim))\nfor word, i in w2i.items():\n if word in embed_model.wv.vocab:\n embedding_vector =embed_model[word]\n \n # words not found in embedding index will be all-zeros.\n word2vec_embd_w[i] = embedding_vector",
"/usr/local/lib/python3.6/dist-packages/ipykernel_launcher.py:4: DeprecationWarning: Call to deprecated `__getitem__` (Method will be removed in 4.0.0, use self.wv.__getitem__() instead).\n after removing the cwd from sys.path.\n"
],
[
"from keras.layers import LSTM\ndef Net(vocab_size, embd_size, sentence_maxlen, glove_embd_w):\n sentence = Input((sentence_maxlen,), name='SentenceInput')\n \n # embedding\n embd_layer = Embedding(input_dim=vocab_size, \n output_dim=embd_size, \n weights=[word2vec_embd_w], \n trainable=False,\n name='shared_embd')\n embd_sentence = embd_layer(sentence)\n embd_sentence = Permute((2,1))(embd_sentence)\n embd_sentence = Lambda(lambda x: K.expand_dims(x, -1))(embd_sentence)\n \n # cnn\n cnn = Conv2D(1, \n kernel_size=(5, sentence_maxlen),\n activation='relu')(embd_sentence)\n cnn = Lambda(lambda x: K.sum(x, axis=3))(cnn)\n cnn = MaxPooling1D(3)(cnn)\n cnn = Lambda(lambda x: K.sum(x, axis=2))(cnn)\n \n hidden1=Dense(400,activation=\"relu\")(cnn)\n hidden2=Dense(300,activation=\"relu\")(hidden1)\n hidden3=Dense(200,activation=\"relu\")(hidden2)\n hidden4=Dense(150,activation=\"relu\")(hidden3)\n out = Dense(len(labels), activation='sigmoid')(hidden4)\n \n sgd = SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)\n model = Model(inputs=sentence, outputs=out, name='sentence_claccification')\n model.compile(optimizer=sgd, loss='binary_crossentropy',metrics=[\"accuracy\", \"binary_accuracy\",\n \"categorical_accuracy\",])\n return model\n\nmodel = Net(vocab_size, embd_dim, sentence_maxlen,word2vec_embd_w)\nprint(model.summary())\n",
"Model: \"sentence_claccification\"\n_________________________________________________________________\nLayer (type) Output Shape Param # \n=================================================================\nSentenceInput (InputLayer) [(None, 300)] 0 \n_________________________________________________________________\nshared_embd (Embedding) (None, 300, 300) 67603500 \n_________________________________________________________________\npermute (Permute) (None, 300, 300) 0 \n_________________________________________________________________\nlambda (Lambda) (None, 300, 300, 1) 0 \n_________________________________________________________________\nconv2d (Conv2D) (None, 296, 1, 1) 1501 \n_________________________________________________________________\nlambda_1 (Lambda) (None, 296, 1) 0 \n_________________________________________________________________\nmax_pooling1d (MaxPooling1D) (None, 98, 1) 0 \n_________________________________________________________________\nlambda_2 (Lambda) (None, 98) 0 \n_________________________________________________________________\ndense (Dense) (None, 400) 39600 \n_________________________________________________________________\ndense_1 (Dense) (None, 300) 120300 \n_________________________________________________________________\ndense_2 (Dense) (None, 200) 60200 \n_________________________________________________________________\ndense_3 (Dense) (None, 150) 30150 \n_________________________________________________________________\ndense_4 (Dense) (None, 78) 11778 \n=================================================================\nTotal params: 67,867,029\nTrainable params: 263,529\nNon-trainable params: 67,603,500\n_________________________________________________________________\nNone\n"
],
[
"es = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=5) # Model stop training after 5 epoch where validation loss didnt decrease\nmc = ModelCheckpoint('best_cnn_4fc.h5', monitor='val_loss', mode='min', verbose=1, save_best_only=True) #You save model weight at the epoch where validation loss is minimal\nmodel.fit(X_train, y_train, batch_size=32,epochs=250,verbose=1,validation_data=(X_val, y_val),callbacks=[es,mc])#you can run for 1000 epoch btw model will stop after 5 epoch without better validation loss",
"Epoch 1/250\n405/405 [==============================] - 49s 122ms/step - loss: 0.1426 - accuracy: 0.0297 - binary_accuracy: 0.9651 - categorical_accuracy: 0.0297 - val_loss: 0.1100 - val_accuracy: 0.0343 - val_binary_accuracy: 0.9764 - val_categorical_accuracy: 0.0343\n\nEpoch 00001: val_loss improved from inf to 0.11000, saving model to best_cnn_4fc.h5\nEpoch 2/250\n405/405 [==============================] - 50s 124ms/step - loss: 0.1080 - accuracy: 0.0478 - binary_accuracy: 0.9766 - categorical_accuracy: 0.0478 - val_loss: 0.1075 - val_accuracy: 0.0596 - val_binary_accuracy: 0.9764 - val_categorical_accuracy: 0.0596\n\nEpoch 00002: val_loss improved from 0.11000 to 0.10752, saving model to best_cnn_4fc.h5\nEpoch 3/250\n405/405 [==============================] - 54s 132ms/step - loss: 0.1051 - accuracy: 0.0791 - binary_accuracy: 0.9766 - categorical_accuracy: 0.0791 - val_loss: 0.1044 - val_accuracy: 0.0837 - val_binary_accuracy: 0.9764 - val_categorical_accuracy: 0.0837\n\nEpoch 00003: val_loss improved from 0.10752 to 0.10438, saving model to best_cnn_4fc.h5\nEpoch 4/250\n405/405 [==============================] - 50s 124ms/step - loss: 0.1010 - accuracy: 0.1194 - binary_accuracy: 0.9766 - categorical_accuracy: 0.1194 - val_loss: 0.0995 - val_accuracy: 0.1491 - val_binary_accuracy: 0.9764 - val_categorical_accuracy: 0.1491\n\nEpoch 00004: val_loss improved from 0.10438 to 0.09951, saving model to best_cnn_4fc.h5\nEpoch 5/250\n405/405 [==============================] - 50s 124ms/step - loss: 0.0956 - accuracy: 0.1822 - binary_accuracy: 0.9767 - categorical_accuracy: 0.1822 - val_loss: 0.0938 - val_accuracy: 0.2201 - val_binary_accuracy: 0.9766 - val_categorical_accuracy: 0.2201\n\nEpoch 00005: val_loss improved from 0.09951 to 0.09384, saving model to best_cnn_4fc.h5\nEpoch 6/250\n405/405 [==============================] - 50s 124ms/step - loss: 0.0903 - accuracy: 0.2376 - binary_accuracy: 0.9769 - categorical_accuracy: 0.2376 - val_loss: 0.0890 - val_accuracy: 0.2632 - val_binary_accuracy: 0.9769 - val_categorical_accuracy: 0.2632\n\nEpoch 00006: val_loss improved from 0.09384 to 0.08898, saving model to best_cnn_4fc.h5\nEpoch 7/250\n405/405 [==============================] - 50s 125ms/step - loss: 0.0858 - accuracy: 0.2756 - binary_accuracy: 0.9773 - categorical_accuracy: 0.2756 - val_loss: 0.0855 - val_accuracy: 0.2843 - val_binary_accuracy: 0.9772 - val_categorical_accuracy: 0.2843\n\nEpoch 00007: val_loss improved from 0.08898 to 0.08549, saving model to best_cnn_4fc.h5\nEpoch 8/250\n405/405 [==============================] - 50s 123ms/step - loss: 0.0824 - accuracy: 0.3072 - binary_accuracy: 0.9776 - categorical_accuracy: 0.3072 - val_loss: 0.0824 - val_accuracy: 0.3152 - val_binary_accuracy: 0.9778 - val_categorical_accuracy: 0.3152\n\nEpoch 00008: val_loss improved from 0.08549 to 0.08243, saving model to best_cnn_4fc.h5\nEpoch 9/250\n405/405 [==============================] - 50s 124ms/step - loss: 0.0797 - accuracy: 0.3241 - binary_accuracy: 0.9781 - categorical_accuracy: 0.3241 - val_loss: 0.0800 - val_accuracy: 0.3367 - val_binary_accuracy: 0.9781 - val_categorical_accuracy: 0.3367\n\nEpoch 00009: val_loss improved from 0.08243 to 0.07998, saving model to best_cnn_4fc.h5\nEpoch 10/250\n405/405 [==============================] - 50s 123ms/step - loss: 0.0776 - accuracy: 0.3402 - binary_accuracy: 0.9783 - categorical_accuracy: 0.3402 - val_loss: 0.0786 - val_accuracy: 0.3564 - val_binary_accuracy: 0.9783 - val_categorical_accuracy: 0.3564\n\nEpoch 00010: val_loss improved from 0.07998 to 0.07861, saving model to best_cnn_4fc.h5\nEpoch 11/250\n405/405 [==============================] - 50s 123ms/step - loss: 0.0758 - accuracy: 0.3556 - binary_accuracy: 0.9786 - categorical_accuracy: 0.3556 - val_loss: 0.0771 - val_accuracy: 0.3553 - val_binary_accuracy: 0.9785 - val_categorical_accuracy: 0.3553\n\nEpoch 00011: val_loss improved from 0.07861 to 0.07711, saving model to best_cnn_4fc.h5\nEpoch 12/250\n405/405 [==============================] - 50s 122ms/step - loss: 0.0745 - accuracy: 0.3668 - binary_accuracy: 0.9788 - categorical_accuracy: 0.3668 - val_loss: 0.0760 - val_accuracy: 0.3650 - val_binary_accuracy: 0.9787 - val_categorical_accuracy: 0.3650\n\nEpoch 00012: val_loss improved from 0.07711 to 0.07600, saving model to best_cnn_4fc.h5\nEpoch 13/250\n405/405 [==============================] - 50s 124ms/step - loss: 0.0732 - accuracy: 0.3794 - binary_accuracy: 0.9789 - categorical_accuracy: 0.3794 - val_loss: 0.0753 - val_accuracy: 0.3755 - val_binary_accuracy: 0.9789 - val_categorical_accuracy: 0.3755\n\nEpoch 00013: val_loss improved from 0.07600 to 0.07525, saving model to best_cnn_4fc.h5\nEpoch 14/250\n405/405 [==============================] - 50s 123ms/step - loss: 0.0722 - accuracy: 0.3850 - binary_accuracy: 0.9791 - categorical_accuracy: 0.3850 - val_loss: 0.0744 - val_accuracy: 0.3994 - val_binary_accuracy: 0.9790 - val_categorical_accuracy: 0.3994\n\nEpoch 00014: val_loss improved from 0.07525 to 0.07436, saving model to best_cnn_4fc.h5\nEpoch 15/250\n405/405 [==============================] - 51s 126ms/step - loss: 0.0714 - accuracy: 0.3934 - binary_accuracy: 0.9792 - categorical_accuracy: 0.3934 - val_loss: 0.0738 - val_accuracy: 0.3915 - val_binary_accuracy: 0.9790 - val_categorical_accuracy: 0.3915\n\nEpoch 00015: val_loss improved from 0.07436 to 0.07383, saving model to best_cnn_4fc.h5\nEpoch 16/250\n405/405 [==============================] - 50s 123ms/step - loss: 0.0705 - accuracy: 0.3966 - binary_accuracy: 0.9793 - categorical_accuracy: 0.3966 - val_loss: 0.0740 - val_accuracy: 0.3933 - val_binary_accuracy: 0.9792 - val_categorical_accuracy: 0.3933\n\nEpoch 00016: val_loss did not improve from 0.07383\nEpoch 17/250\n405/405 [==============================] - 49s 121ms/step - loss: 0.0698 - accuracy: 0.4070 - binary_accuracy: 0.9795 - categorical_accuracy: 0.4070 - val_loss: 0.0742 - val_accuracy: 0.3908 - val_binary_accuracy: 0.9791 - val_categorical_accuracy: 0.3908\n\nEpoch 00017: val_loss did not improve from 0.07383\nEpoch 18/250\n405/405 [==============================] - 49s 121ms/step - loss: 0.0690 - accuracy: 0.4084 - binary_accuracy: 0.9796 - categorical_accuracy: 0.4084 - val_loss: 0.0722 - val_accuracy: 0.3994 - val_binary_accuracy: 0.9795 - val_categorical_accuracy: 0.3994\n\nEpoch 00018: val_loss improved from 0.07383 to 0.07217, saving model to best_cnn_4fc.h5\nEpoch 19/250\n405/405 [==============================] - 50s 123ms/step - loss: 0.0684 - accuracy: 0.4117 - binary_accuracy: 0.9798 - categorical_accuracy: 0.4117 - val_loss: 0.0722 - val_accuracy: 0.4024 - val_binary_accuracy: 0.9794 - val_categorical_accuracy: 0.4024\n\nEpoch 00019: val_loss did not improve from 0.07217\nEpoch 20/250\n405/405 [==============================] - 49s 122ms/step - loss: 0.0678 - accuracy: 0.4162 - binary_accuracy: 0.9798 - categorical_accuracy: 0.4162 - val_loss: 0.0717 - val_accuracy: 0.4047 - val_binary_accuracy: 0.9796 - val_categorical_accuracy: 0.4047\n\nEpoch 00020: val_loss improved from 0.07217 to 0.07174, saving model to best_cnn_4fc.h5\nEpoch 21/250\n405/405 [==============================] - 50s 123ms/step - loss: 0.0673 - accuracy: 0.4206 - binary_accuracy: 0.9799 - categorical_accuracy: 0.4206 - val_loss: 0.0711 - val_accuracy: 0.4045 - val_binary_accuracy: 0.9795 - val_categorical_accuracy: 0.4045\n\nEpoch 00021: val_loss improved from 0.07174 to 0.07109, saving model to best_cnn_4fc.h5\nEpoch 22/250\n405/405 [==============================] - 49s 121ms/step - loss: 0.0667 - accuracy: 0.4199 - binary_accuracy: 0.9799 - categorical_accuracy: 0.4199 - val_loss: 0.0715 - val_accuracy: 0.4058 - val_binary_accuracy: 0.9796 - val_categorical_accuracy: 0.4058\n\nEpoch 00022: val_loss did not improve from 0.07109\nEpoch 23/250\n405/405 [==============================] - 49s 120ms/step - loss: 0.0661 - accuracy: 0.4248 - binary_accuracy: 0.9800 - categorical_accuracy: 0.4248 - val_loss: 0.0710 - val_accuracy: 0.4079 - val_binary_accuracy: 0.9796 - val_categorical_accuracy: 0.4079\n\nEpoch 00023: val_loss improved from 0.07109 to 0.07103, saving model to best_cnn_4fc.h5\nEpoch 24/250\n405/405 [==============================] - 50s 123ms/step - loss: 0.0656 - accuracy: 0.4300 - binary_accuracy: 0.9802 - categorical_accuracy: 0.4300 - val_loss: 0.0702 - val_accuracy: 0.4142 - val_binary_accuracy: 0.9797 - val_categorical_accuracy: 0.4142\n\nEpoch 00024: val_loss improved from 0.07103 to 0.07025, saving model to best_cnn_4fc.h5\nEpoch 25/250\n405/405 [==============================] - 50s 123ms/step - loss: 0.0652 - accuracy: 0.4330 - binary_accuracy: 0.9802 - categorical_accuracy: 0.4330 - val_loss: 0.0703 - val_accuracy: 0.4142 - val_binary_accuracy: 0.9798 - val_categorical_accuracy: 0.4142\n\nEpoch 00025: val_loss did not improve from 0.07025\nEpoch 26/250\n405/405 [==============================] - 49s 120ms/step - loss: 0.0648 - accuracy: 0.4341 - binary_accuracy: 0.9803 - categorical_accuracy: 0.4341 - val_loss: 0.0700 - val_accuracy: 0.4163 - val_binary_accuracy: 0.9799 - val_categorical_accuracy: 0.4163\n\nEpoch 00026: val_loss improved from 0.07025 to 0.06999, saving model to best_cnn_4fc.h5\nEpoch 27/250\n405/405 [==============================] - 50s 123ms/step - loss: 0.0644 - accuracy: 0.4351 - binary_accuracy: 0.9803 - categorical_accuracy: 0.4351 - val_loss: 0.0697 - val_accuracy: 0.4137 - val_binary_accuracy: 0.9798 - val_categorical_accuracy: 0.4137\n\nEpoch 00027: val_loss improved from 0.06999 to 0.06973, saving model to best_cnn_4fc.h5\nEpoch 28/250\n405/405 [==============================] - 50s 123ms/step - loss: 0.0639 - accuracy: 0.4363 - binary_accuracy: 0.9804 - categorical_accuracy: 0.4363 - val_loss: 0.0696 - val_accuracy: 0.4167 - val_binary_accuracy: 0.9797 - val_categorical_accuracy: 0.4167\n\nEpoch 00028: val_loss improved from 0.06973 to 0.06964, saving model to best_cnn_4fc.h5\nEpoch 29/250\n405/405 [==============================] - 49s 122ms/step - loss: 0.0636 - accuracy: 0.4410 - binary_accuracy: 0.9804 - categorical_accuracy: 0.4410 - val_loss: 0.0700 - val_accuracy: 0.4116 - val_binary_accuracy: 0.9798 - val_categorical_accuracy: 0.4116\n\nEpoch 00029: val_loss did not improve from 0.06964\nEpoch 30/250\n405/405 [==============================] - 49s 120ms/step - loss: 0.0631 - accuracy: 0.4436 - binary_accuracy: 0.9805 - categorical_accuracy: 0.4436 - val_loss: 0.0700 - val_accuracy: 0.4114 - val_binary_accuracy: 0.9795 - val_categorical_accuracy: 0.4114\n\nEpoch 00030: val_loss did not improve from 0.06964\nEpoch 31/250\n405/405 [==============================] - 49s 120ms/step - loss: 0.0627 - accuracy: 0.4482 - binary_accuracy: 0.9805 - categorical_accuracy: 0.4482 - val_loss: 0.0700 - val_accuracy: 0.4179 - val_binary_accuracy: 0.9796 - val_categorical_accuracy: 0.4179\n\nEpoch 00031: val_loss did not improve from 0.06964\nEpoch 32/250\n405/405 [==============================] - 49s 120ms/step - loss: 0.0624 - accuracy: 0.4481 - binary_accuracy: 0.9806 - categorical_accuracy: 0.4481 - val_loss: 0.0689 - val_accuracy: 0.4184 - val_binary_accuracy: 0.9798 - val_categorical_accuracy: 0.4184\n\nEpoch 00032: val_loss improved from 0.06964 to 0.06892, saving model to best_cnn_4fc.h5\nEpoch 33/250\n405/405 [==============================] - 49s 122ms/step - loss: 0.0620 - accuracy: 0.4536 - binary_accuracy: 0.9807 - categorical_accuracy: 0.4536 - val_loss: 0.0691 - val_accuracy: 0.4230 - val_binary_accuracy: 0.9799 - val_categorical_accuracy: 0.4230\n\nEpoch 00033: val_loss did not improve from 0.06892\nEpoch 34/250\n405/405 [==============================] - 49s 120ms/step - loss: 0.0616 - accuracy: 0.4528 - binary_accuracy: 0.9807 - categorical_accuracy: 0.4528 - val_loss: 0.0690 - val_accuracy: 0.4195 - val_binary_accuracy: 0.9797 - val_categorical_accuracy: 0.4195\n\nEpoch 00034: val_loss did not improve from 0.06892\nEpoch 35/250\n405/405 [==============================] - 49s 121ms/step - loss: 0.0614 - accuracy: 0.4549 - binary_accuracy: 0.9807 - categorical_accuracy: 0.4549 - val_loss: 0.0685 - val_accuracy: 0.4290 - val_binary_accuracy: 0.9799 - val_categorical_accuracy: 0.4290\n\nEpoch 00035: val_loss improved from 0.06892 to 0.06855, saving model to best_cnn_4fc.h5\nEpoch 36/250\n405/405 [==============================] - 49s 122ms/step - loss: 0.0611 - accuracy: 0.4545 - binary_accuracy: 0.9807 - categorical_accuracy: 0.4545 - val_loss: 0.0689 - val_accuracy: 0.4290 - val_binary_accuracy: 0.9799 - val_categorical_accuracy: 0.4290\n\nEpoch 00036: val_loss did not improve from 0.06855\nEpoch 37/250\n405/405 [==============================] - 49s 120ms/step - loss: 0.0607 - accuracy: 0.4598 - binary_accuracy: 0.9808 - categorical_accuracy: 0.4598 - val_loss: 0.0686 - val_accuracy: 0.4290 - val_binary_accuracy: 0.9799 - val_categorical_accuracy: 0.4290\n\nEpoch 00037: val_loss did not improve from 0.06855\nEpoch 38/250\n405/405 [==============================] - 49s 120ms/step - loss: 0.0604 - accuracy: 0.4625 - binary_accuracy: 0.9808 - categorical_accuracy: 0.4625 - val_loss: 0.0684 - val_accuracy: 0.4325 - val_binary_accuracy: 0.9798 - val_categorical_accuracy: 0.4325\n\nEpoch 00038: val_loss improved from 0.06855 to 0.06840, saving model to best_cnn_4fc.h5\nEpoch 39/250\n405/405 [==============================] - 49s 122ms/step - loss: 0.0600 - accuracy: 0.4603 - binary_accuracy: 0.9809 - categorical_accuracy: 0.4603 - val_loss: 0.0689 - val_accuracy: 0.4360 - val_binary_accuracy: 0.9800 - val_categorical_accuracy: 0.4360\n\nEpoch 00039: val_loss did not improve from 0.06840\nEpoch 40/250\n405/405 [==============================] - 49s 120ms/step - loss: 0.0597 - accuracy: 0.4655 - binary_accuracy: 0.9810 - categorical_accuracy: 0.4655 - val_loss: 0.0689 - val_accuracy: 0.4242 - val_binary_accuracy: 0.9798 - val_categorical_accuracy: 0.4242\n\nEpoch 00040: val_loss did not improve from 0.06840\nEpoch 41/250\n405/405 [==============================] - 49s 121ms/step - loss: 0.0594 - accuracy: 0.4683 - binary_accuracy: 0.9810 - categorical_accuracy: 0.4683 - val_loss: 0.0684 - val_accuracy: 0.4318 - val_binary_accuracy: 0.9798 - val_categorical_accuracy: 0.4318\n\nEpoch 00041: val_loss improved from 0.06840 to 0.06838, saving model to best_cnn_4fc.h5\nEpoch 42/250\n405/405 [==============================] - 49s 122ms/step - loss: 0.0592 - accuracy: 0.4686 - binary_accuracy: 0.9811 - categorical_accuracy: 0.4686 - val_loss: 0.0688 - val_accuracy: 0.4177 - val_binary_accuracy: 0.9798 - val_categorical_accuracy: 0.4177\n\nEpoch 00042: val_loss did not improve from 0.06838\nEpoch 43/250\n405/405 [==============================] - 49s 120ms/step - loss: 0.0588 - accuracy: 0.4712 - binary_accuracy: 0.9811 - categorical_accuracy: 0.4712 - val_loss: 0.0683 - val_accuracy: 0.4397 - val_binary_accuracy: 0.9800 - val_categorical_accuracy: 0.4397\n\nEpoch 00043: val_loss improved from 0.06838 to 0.06830, saving model to best_cnn_4fc.h5\nEpoch 44/250\n405/405 [==============================] - 49s 122ms/step - loss: 0.0585 - accuracy: 0.4736 - binary_accuracy: 0.9812 - categorical_accuracy: 0.4736 - val_loss: 0.0683 - val_accuracy: 0.4228 - val_binary_accuracy: 0.9799 - val_categorical_accuracy: 0.4228\n\nEpoch 00044: val_loss improved from 0.06830 to 0.06826, saving model to best_cnn_4fc.h5\nEpoch 45/250\n405/405 [==============================] - 49s 122ms/step - loss: 0.0582 - accuracy: 0.4738 - binary_accuracy: 0.9812 - categorical_accuracy: 0.4738 - val_loss: 0.0682 - val_accuracy: 0.4290 - val_binary_accuracy: 0.9798 - val_categorical_accuracy: 0.4290\n\nEpoch 00045: val_loss improved from 0.06826 to 0.06823, saving model to best_cnn_4fc.h5\nEpoch 46/250\n405/405 [==============================] - 49s 122ms/step - loss: 0.0579 - accuracy: 0.4724 - binary_accuracy: 0.9813 - categorical_accuracy: 0.4724 - val_loss: 0.0686 - val_accuracy: 0.4272 - val_binary_accuracy: 0.9799 - val_categorical_accuracy: 0.4272\n\nEpoch 00046: val_loss did not improve from 0.06823\nEpoch 47/250\n405/405 [==============================] - 49s 121ms/step - loss: 0.0576 - accuracy: 0.4772 - binary_accuracy: 0.9813 - categorical_accuracy: 0.4772 - val_loss: 0.0681 - val_accuracy: 0.4392 - val_binary_accuracy: 0.9800 - val_categorical_accuracy: 0.4392\n\nEpoch 00047: val_loss improved from 0.06823 to 0.06809, saving model to best_cnn_4fc.h5\nEpoch 48/250\n405/405 [==============================] - 49s 121ms/step - loss: 0.0574 - accuracy: 0.4791 - binary_accuracy: 0.9813 - categorical_accuracy: 0.4791 - val_loss: 0.0682 - val_accuracy: 0.4388 - val_binary_accuracy: 0.9798 - val_categorical_accuracy: 0.4388\n\nEpoch 00048: val_loss did not improve from 0.06809\nEpoch 49/250\n405/405 [==============================] - 48s 120ms/step - loss: 0.0571 - accuracy: 0.4789 - binary_accuracy: 0.9813 - categorical_accuracy: 0.4789 - val_loss: 0.0688 - val_accuracy: 0.4223 - val_binary_accuracy: 0.9797 - val_categorical_accuracy: 0.4223\n\nEpoch 00049: val_loss did not improve from 0.06809\nEpoch 50/250\n405/405 [==============================] - 49s 120ms/step - loss: 0.0568 - accuracy: 0.4823 - binary_accuracy: 0.9814 - categorical_accuracy: 0.4823 - val_loss: 0.0684 - val_accuracy: 0.4432 - val_binary_accuracy: 0.9797 - val_categorical_accuracy: 0.4432\n\nEpoch 00050: val_loss did not improve from 0.06809\nEpoch 51/250\n405/405 [==============================] - 49s 122ms/step - loss: 0.0565 - accuracy: 0.4812 - binary_accuracy: 0.9815 - categorical_accuracy: 0.4812 - val_loss: 0.0688 - val_accuracy: 0.4365 - val_binary_accuracy: 0.9801 - val_categorical_accuracy: 0.4365\n\nEpoch 00051: val_loss did not improve from 0.06809\nEpoch 52/250\n405/405 [==============================] - 49s 120ms/step - loss: 0.0562 - accuracy: 0.4837 - binary_accuracy: 0.9815 - categorical_accuracy: 0.4837 - val_loss: 0.0684 - val_accuracy: 0.4348 - val_binary_accuracy: 0.9800 - val_categorical_accuracy: 0.4348\n\nEpoch 00052: val_loss did not improve from 0.06809\nEpoch 00052: early stopping\n"
]
],
[
[
"# ***If the model is generated before:***",
"_____no_output_____"
]
],
[
[
"model = load_model('best_cnn_4fc_with_binary.h5')\n# model.save('best_cnn_4fc_with_binary.h5')",
"_____no_output_____"
],
[
"pred=model.predict(X_test)\r\n# For evaluation: If the probability > 0.5 you can say that it belong to the class.",
"_____no_output_____"
],
[
"print(pred[0])#example",
"[1.28691900e-05 1.35993958e-03 8.81868327e-05 3.08060407e-06\n 1.21343424e-08 2.15166278e-06 5.58876200e-05 6.13067168e-05\n 6.20292553e-07 2.01195478e-04 5.23518429e-06 7.95147004e-11\n 6.75920386e-09 1.72682011e-08 9.49173398e-08 7.38362019e-07\n 3.91908571e-07 1.81894898e-02 1.27643347e-04 3.57568264e-04\n 2.41607428e-04 6.11127052e-06 1.60775457e-07 1.02964044e-03\n 5.03659248e-04 1.49760067e-01 7.94296375e-06 1.69244927e-07\n 3.16571482e-06 2.19076872e-04 1.47283077e-04 1.31869912e-02\n 2.23560278e-06 1.95117846e-05 2.65032053e-04 2.10514244e-07\n 4.25242570e-06 7.16469913e-07 6.15581084e-05 4.05709716e-06\n 6.52370930e-01 3.81792379e-05 9.69902612e-05 2.00494384e-07\n 2.13137269e-03 4.44282705e-06 4.90285075e-08 3.38226557e-04\n 1.03130937e-03 1.24948610e-06 3.26928102e-08 1.00508532e-06\n 5.09917736e-04 1.78045312e-08 2.34452413e-09 1.22595538e-05\n 1.09079480e-03 2.48280941e-07 1.48249567e-02 1.37757834e-05\n 7.76499510e-04 3.41236591e-03 2.26378441e-04 6.20567153e-05\n 2.09715245e-06 3.80242597e-07 1.29918316e-08 7.16418072e-05\n 6.42688337e-06 1.53124332e-04 7.26446660e-06 1.25186890e-01\n 7.54332214e-05 3.90789091e-06 1.52777460e-07 1.59919262e-04\n 1.14491195e-05 2.82711267e-06]\n"
],
[
"y_pred=[]\r\nmeasure = 9 * (np.mean(pred[0]) + 0.5*np.sqrt(np.var(pred[0])))\r\nfor l in pred:\r\n temp=[]\r\n for value in l:\r\n if value>= measure:\r\n temp.append(1)\r\n else:\r\n temp.append(0)\r\n y_pred.append(temp)\r\n",
"_____no_output_____"
],
[
"measure",
"_____no_output_____"
],
[
"from sklearn.metrics import classification_report,accuracy_score\r\n\r\nprint(\"accuracy=\",accuracy_score(y_test, y_pred))\r\nprint(classification_report(y_test, y_pred))",
"accuracy= 0.3587662337662338\n precision recall f1-score support\n\n 0 0.53 0.50 0.51 103\n 1 0.45 0.32 0.37 151\n 2 0.50 0.45 0.47 141\n 3 0.42 0.19 0.26 26\n 4 0.96 0.91 0.93 97\n 5 1.00 0.07 0.13 14\n 6 0.49 0.41 0.44 137\n 7 0.83 0.28 0.42 36\n 8 0.30 0.14 0.19 22\n 9 0.70 0.13 0.22 122\n 10 0.63 0.38 0.47 152\n 11 0.96 0.96 0.96 77\n 12 0.99 0.98 0.98 129\n 13 0.50 0.14 0.22 14\n 14 0.83 0.28 0.42 18\n 15 0.71 0.30 0.42 148\n 16 0.72 0.60 0.65 121\n 17 0.83 0.42 0.56 153\n 18 0.69 0.62 0.66 101\n 19 0.68 0.37 0.48 62\n 20 0.53 0.63 0.57 132\n 21 0.60 0.41 0.49 135\n 22 0.74 0.37 0.49 38\n 23 0.69 0.56 0.62 140\n 24 0.41 0.14 0.21 79\n 25 0.74 0.71 0.73 147\n 26 0.71 0.25 0.37 155\n 27 0.75 0.43 0.55 146\n 28 0.57 0.21 0.30 125\n 29 0.66 0.69 0.68 131\n 30 0.65 0.26 0.37 58\n 31 0.74 0.64 0.68 83\n 32 0.67 0.47 0.56 116\n 33 0.54 0.38 0.45 152\n 34 0.29 0.05 0.08 85\n 35 0.56 0.45 0.50 130\n 36 0.65 0.50 0.56 157\n 37 0.95 0.81 0.87 64\n 38 0.69 0.47 0.56 47\n 39 0.84 0.60 0.70 35\n 40 0.75 0.68 0.71 146\n 41 0.56 0.54 0.55 116\n 42 0.70 0.19 0.30 73\n 43 0.72 0.30 0.43 152\n 44 0.80 0.62 0.70 166\n 45 0.00 0.00 0.00 24\n 46 0.65 0.34 0.45 32\n 47 0.46 0.47 0.47 140\n 48 0.54 0.34 0.41 151\n 49 0.82 0.42 0.56 66\n 50 0.40 0.08 0.13 26\n 51 0.65 0.52 0.58 21\n 52 0.44 0.24 0.31 34\n 53 0.77 0.58 0.66 105\n 54 0.73 0.70 0.72 130\n 55 0.67 0.35 0.46 34\n 56 0.77 0.31 0.45 54\n 57 0.94 0.52 0.67 60\n 58 0.75 0.54 0.63 153\n 59 0.66 0.26 0.37 149\n 60 0.69 0.59 0.64 143\n 61 0.73 0.14 0.24 56\n 62 0.43 0.35 0.39 154\n 63 0.77 0.81 0.79 141\n 64 0.74 0.37 0.49 95\n 65 0.71 0.54 0.61 158\n 66 1.00 0.45 0.62 31\n 67 0.51 0.24 0.33 90\n 68 0.83 0.68 0.74 157\n 69 0.48 0.28 0.35 156\n 70 0.81 0.31 0.45 140\n 71 0.65 0.15 0.25 131\n 72 0.69 0.45 0.54 134\n 73 0.79 0.38 0.52 39\n 74 0.91 0.45 0.61 22\n 75 0.64 0.34 0.45 149\n 76 0.81 0.45 0.58 76\n 77 0.63 0.46 0.53 113\n\n micro avg 0.67 0.45 0.54 7796\n macro avg 0.67 0.42 0.50 7796\nweighted avg 0.67 0.45 0.52 7796\n samples avg 0.51 0.49 0.48 7796\n\n"
],
[
"",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4acfe3b5268c28cb02fc845cedbf0a3a36675295
| 54,236 |
ipynb
|
Jupyter Notebook
|
site/en/r2/guide/ragged_tensors.ipynb
|
NexusXi/docs
|
7c8aea728a4eca4ee719fe6cca36ee95eb7ad40d
|
[
"Apache-2.0"
] | 4 |
2019-08-20T11:59:23.000Z
|
2020-01-12T13:42:50.000Z
|
site/en/r2/guide/ragged_tensors.ipynb
|
NexusXi/docs
|
7c8aea728a4eca4ee719fe6cca36ee95eb7ad40d
|
[
"Apache-2.0"
] | null | null | null |
site/en/r2/guide/ragged_tensors.ipynb
|
NexusXi/docs
|
7c8aea728a4eca4ee719fe6cca36ee95eb7ad40d
|
[
"Apache-2.0"
] | null | null | null | 32.496105 | 306 | 0.4915 |
[
[
[
"##### Copyright 2018 The TensorFlow Authors.",
"_____no_output_____"
]
],
[
[
"#@title Licensed under the Apache License, Version 2.0 (the \"License\");\n# you may not use this file except in compliance with the License.\n# You may obtain a copy of the License at\n#\n# https://www.apache.org/licenses/LICENSE-2.0\n#\n# Unless required by applicable law or agreed to in writing, software\n# distributed under the License is distributed on an \"AS IS\" BASIS,\n# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.\n# See the License for the specific language governing permissions and\n# limitations under the License.",
"_____no_output_____"
]
],
[
[
"# Ragged Tensors\n\n<table class=\"tfo-notebook-buttons\" align=\"left\">\n <td>\n <a target=\"_blank\" href=\"https://www.tensorflow.org/beta/guide/ragged_tensors\"><img src=\"https://www.tensorflow.org/images/tf_logo_32px.png\" />View on TensorFlow.org</a>\n </td>\n <td>\n <a target=\"_blank\" href=\"https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r2/guide/ragged_tensors.ipynb\"><img src=\"https://www.tensorflow.org/images/colab_logo_32px.png\" />Run in Google Colab</a>\n </td>\n <td>\n <a target=\"_blank\" href=\"https://github.com/tensorflow/docs/blob/master/site/en/r2/guide/ragged_tensors.ipynb\"><img src=\"https://www.tensorflow.org/images/GitHub-Mark-32px.png\" />View source on GitHub</a>\n </td>\n <td>\n <a href=\"https://storage.googleapis.com/tensorflow_docs/docs/site/en/r2/guide/ragged_tensors.ipynb\"><img src=\"https://www.tensorflow.org/images/download_logo_32px.png\" />Download notebook</a>\n </td>\n</table>",
"_____no_output_____"
],
[
"## Setup",
"_____no_output_____"
]
],
[
[
"from __future__ import absolute_import, division, print_function, unicode_literals\n\nimport math\ntry:\n %tensorflow_version 2.x # Colab only.\nexcept Exception:\n pass\nimport tensorflow as tf",
"_____no_output_____"
]
],
[
[
"## Overview\n\nYour data comes in many shapes; your tensors should too.\n*Ragged tensors* are the TensorFlow equivalent of nested variable-length\nlists. They make it easy to store and process data with non-uniform shapes,\nincluding:\n\n* Variable-length features, such as the set of actors in a movie.\n* Batches of variable-length sequential inputs, such as sentences or video\n clips.\n* Hierarchical inputs, such as text documents that are subdivided into\n sections, paragraphs, sentences, and words.\n* Individual fields in structured inputs, such as protocol buffers.\n\n### What you can do with a ragged tensor\n\nRagged tensors are supported by more than a hundred TensorFlow operations,\nincluding math operations (such as `tf.add` and `tf.reduce_mean`), array operations\n(such as `tf.concat` and `tf.tile`), string manipulation ops (such as\n`tf.substr`), and many others:\n",
"_____no_output_____"
]
],
[
[
"digits = tf.ragged.constant([[3, 1, 4, 1], [], [5, 9, 2], [6], []])\nwords = tf.ragged.constant([[\"So\", \"long\"], [\"thanks\", \"for\", \"all\", \"the\", \"fish\"]])\nprint(tf.add(digits, 3))\nprint(tf.reduce_mean(digits, axis=1))\nprint(tf.concat([digits, [[5, 3]]], axis=0))\nprint(tf.tile(digits, [1, 2]))\nprint(tf.strings.substr(words, 0, 2))",
"_____no_output_____"
]
],
[
[
"There are also a number of methods and operations that are\nspecific to ragged tensors, including factory methods, conversion methods,\nand value-mapping operations.\nFor a list of supported ops, see the `tf.ragged` package\ndocumentation.\n\nAs with normal tensors, you can use Python-style indexing to access specific\nslices of a ragged tensor. For more information, see the section on\n**Indexing** below.",
"_____no_output_____"
]
],
[
[
"print(digits[0]) # First row",
"_____no_output_____"
],
[
"print(digits[:, :2]) # First two values in each row.",
"_____no_output_____"
],
[
"print(digits[:, -2:]) # Last two values in each row.",
"_____no_output_____"
]
],
[
[
"And just like normal tensors, you can use Python arithmetic and comparison\noperators to perform elementwise operations. For more information, see the section on\n**Overloaded Operators** below.",
"_____no_output_____"
]
],
[
[
"print(digits + 3)",
"_____no_output_____"
],
[
"print(digits + tf.ragged.constant([[1, 2, 3, 4], [], [5, 6, 7], [8], []]))",
"_____no_output_____"
]
],
[
[
"If you need to perform an elementwise transformation to the values of a `RaggedTensor`, you can use `tf.ragged.map_flat_values`, which takes a function plus one or more arguments, and applies the function to transform the `RaggedTensor`'s values.",
"_____no_output_____"
]
],
[
[
"times_two_plus_one = lambda x: x * 2 + 1\nprint(tf.ragged.map_flat_values(times_two_plus_one, digits))",
"_____no_output_____"
]
],
[
[
"### Constructing a ragged tensor\n\nThe simplest way to construct a ragged tensor is using\n`tf.ragged.constant`, which builds the\n`RaggedTensor` corresponding to a given nested Python `list`:",
"_____no_output_____"
]
],
[
[
"sentences = tf.ragged.constant([\n [\"Let's\", \"build\", \"some\", \"ragged\", \"tensors\", \"!\"],\n [\"We\", \"can\", \"use\", \"tf.ragged.constant\", \".\"]])\nprint(sentences)",
"_____no_output_____"
],
[
"paragraphs = tf.ragged.constant([\n [['I', 'have', 'a', 'cat'], ['His', 'name', 'is', 'Mat']],\n [['Do', 'you', 'want', 'to', 'come', 'visit'], [\"I'm\", 'free', 'tomorrow']],\n])\nprint(paragraphs)",
"_____no_output_____"
]
],
[
[
"Ragged tensors can also be constructed by pairing flat *values* tensors with\n*row-partitioning* tensors indicating how those values should be divided into\nrows, using factory classmethods such as `tf.RaggedTensor.from_value_rowids`,\n`tf.RaggedTensor.from_row_lengths`, and\n`tf.RaggedTensor.from_row_splits`.\n\n#### `tf.RaggedTensor.from_value_rowids`\nIf you know which row each value belongs in, then you can build a `RaggedTensor` using a `value_rowids` row-partitioning tensor:\n\n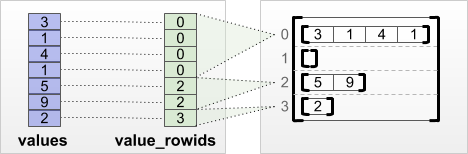",
"_____no_output_____"
]
],
[
[
"print(tf.RaggedTensor.from_value_rowids(\n values=[3, 1, 4, 1, 5, 9, 2, 6],\n value_rowids=[0, 0, 0, 0, 2, 2, 2, 3]))",
"_____no_output_____"
]
],
[
[
"#### `tf.RaggedTensor.from_row_lengths`\n\nIf you know how long each row is, then you can use a `row_lengths` row-partitioning tensor:\n\n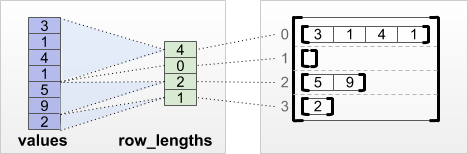",
"_____no_output_____"
]
],
[
[
"print(tf.RaggedTensor.from_row_lengths(\n values=[3, 1, 4, 1, 5, 9, 2, 6],\n row_lengths=[4, 0, 3, 1]))",
"_____no_output_____"
]
],
[
[
"#### `tf.RaggedTensor.from_row_splits`\n\nIf you know the index where each row starts and ends, then you can use a `row_splits` row-partitioning tensor:\n\n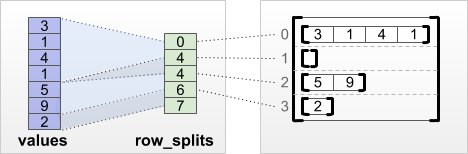",
"_____no_output_____"
]
],
[
[
"print(tf.RaggedTensor.from_row_splits(\n values=[3, 1, 4, 1, 5, 9, 2, 6],\n row_splits=[0, 4, 4, 7, 8]))",
"_____no_output_____"
]
],
[
[
"See the `tf.RaggedTensor` class documentation for a full list of factory methods.",
"_____no_output_____"
],
[
"### What you can store in a ragged tensor\n\nAs with normal `Tensor`s, the values in a `RaggedTensor` must all have the same\ntype; and the values must all be at the same nesting depth (the *rank* of the\ntensor):",
"_____no_output_____"
]
],
[
[
"print(tf.ragged.constant([[\"Hi\"], [\"How\", \"are\", \"you\"]])) # ok: type=string, rank=2",
"_____no_output_____"
],
[
"print(tf.ragged.constant([[[1, 2], [3]], [[4, 5]]])) # ok: type=int32, rank=3",
"_____no_output_____"
],
[
"try:\n tf.ragged.constant([[\"one\", \"two\"], [3, 4]]) # bad: multiple types\nexcept ValueError as exception:\n print(exception)",
"_____no_output_____"
],
[
"try:\n tf.ragged.constant([\"A\", [\"B\", \"C\"]]) # bad: multiple nesting depths\nexcept ValueError as exception:\n print(exception)",
"_____no_output_____"
]
],
[
[
"### Example use case\n\nThe following example demonstrates how `RaggedTensor`s can be used to construct\nand combine unigram and bigram embeddings for a batch of variable-length\nqueries, using special markers for the beginning and end of each sentence.\nFor more details on the ops used in this example, see the `tf.ragged` package documentation.",
"_____no_output_____"
]
],
[
[
"queries = tf.ragged.constant([['Who', 'is', 'Dan', 'Smith'],\n ['Pause'],\n ['Will', 'it', 'rain', 'later', 'today']])\n\n# Create an embedding table.\nnum_buckets = 1024\nembedding_size = 4\nembedding_table = tf.Variable(\n tf.random.truncated_normal([num_buckets, embedding_size],\n stddev=1.0 / math.sqrt(embedding_size)))\n\n# Look up the embedding for each word.\nword_buckets = tf.strings.to_hash_bucket_fast(queries, num_buckets)\nword_embeddings = tf.ragged.map_flat_values(\n tf.nn.embedding_lookup, embedding_table, word_buckets) # ①\n\n# Add markers to the beginning and end of each sentence.\nmarker = tf.fill([queries.nrows(), 1], '#')\npadded = tf.concat([marker, queries, marker], axis=1) # ②\n\n# Build word bigrams & look up embeddings.\nbigrams = tf.strings.join([padded[:, :-1],\n padded[:, 1:]],\n separator='+') # ③\n\nbigram_buckets = tf.strings.to_hash_bucket_fast(bigrams, num_buckets)\nbigram_embeddings = tf.ragged.map_flat_values(\n tf.nn.embedding_lookup, embedding_table, bigram_buckets) # ④\n\n# Find the average embedding for each sentence\nall_embeddings = tf.concat([word_embeddings, bigram_embeddings], axis=1) # ⑤\navg_embedding = tf.reduce_mean(all_embeddings, axis=1) # ⑥\nprint(avg_embedding)",
"_____no_output_____"
]
],
[
[
"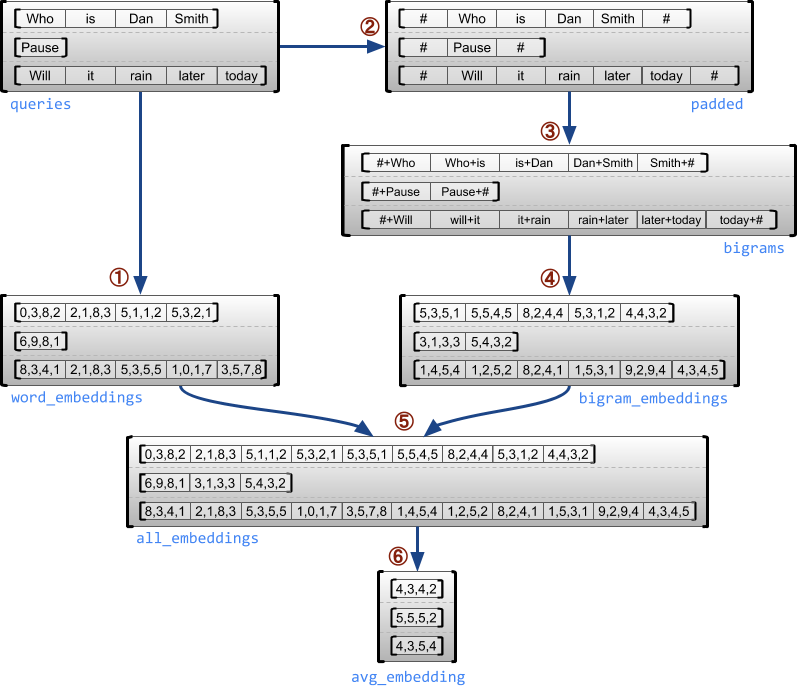",
"_____no_output_____"
],
[
"## Ragged tensors: definitions\n\n### Ragged and uniform dimensions\n\nA *ragged tensor* is a tensor with one or more *ragged dimensions*,\nwhich are dimensions whose slices may have different lengths. For example, the\ninner (column) dimension of `rt=[[3, 1, 4, 1], [], [5, 9, 2], [6], []]` is\nragged, since the column slices (`rt[0, :]`, ..., `rt[4, :]`) have different\nlengths. Dimensions whose slices all have the same length are called *uniform\ndimensions*.\n\nThe outermost dimension of a ragged tensor is always uniform, since it consists\nof a single slice (and so there is no possibility for differing slice lengths).\nIn addition to the uniform outermost dimension, ragged tensors may also have\nuniform inner dimensions. For example, we might store the word embeddings for\neach word in a batch of sentences using a ragged tensor with shape\n`[num_sentences, (num_words), embedding_size]`, where the parentheses around\n`(num_words)` indicate that the dimension is ragged.\n\n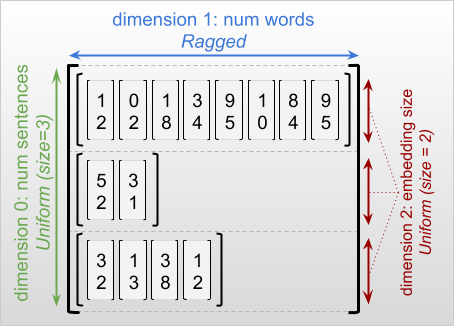\n\nRagged tensors may have multiple ragged dimensions. For example, we could store\na batch of structured text documents using a tensor with shape `[num_documents,\n(num_paragraphs), (num_sentences), (num_words)]` (where again parentheses are\nused to indicate ragged dimensions).\n\n#### Ragged tensor shape restrictions\n\nThe shape of a ragged tensor is currently restricted to have the following form:\n\n* A single uniform dimension\n* Followed by one or more ragged dimensions\n* Followed by zero or more uniform dimensions.\n\nNote: These restrictions are a consequence of the current implementation, and we\nmay relax them in the future.\n\n### Rank and ragged rank\n\nThe total number of dimensions in a ragged tensor is called its ***rank***, and\nthe number of ragged dimensions in a ragged tensor is called its ***ragged\nrank***. In graph execution mode (i.e., non-eager mode), a tensor's ragged rank\nis fixed at creation time: it can't depend\non runtime values, and can't vary dynamically for different session runs.\nA ***potentially ragged tensor*** is a value that might be\neither a `tf.Tensor` or a `tf.RaggedTensor`. The\nragged rank of a `tf.Tensor` is defined to be zero.\n\n### RaggedTensor shapes\n\nWhen describing the shape of a RaggedTensor, ragged dimensions are indicated by\nenclosing them in parentheses. For example, as we saw above, the shape of a 3-D\nRaggedTensor that stores word embeddings for each word in a batch of sentences\ncan be written as `[num_sentences, (num_words), embedding_size]`.\nThe `RaggedTensor.shape` attribute returns a `tf.TensorShape` for a\nragged tensor, where ragged dimensions have size `None`:\n",
"_____no_output_____"
]
],
[
[
"tf.ragged.constant([[\"Hi\"], [\"How\", \"are\", \"you\"]]).shape",
"_____no_output_____"
]
],
[
[
"The method `tf.RaggedTensor.bounding_shape` can be used to find a tight\nbounding shape for a given `RaggedTensor`:",
"_____no_output_____"
]
],
[
[
"print(tf.ragged.constant([[\"Hi\"], [\"How\", \"are\", \"you\"]]).bounding_shape())",
"_____no_output_____"
]
],
[
[
"## Ragged vs sparse tensors\n\nA ragged tensor should *not* be thought of as a type of sparse tensor, but\nrather as a dense tensor with an irregular shape.\n\nAs an illustrative example, consider how array operations such as `concat`,\n`stack`, and `tile` are defined for ragged vs. sparse tensors. Concatenating\nragged tensors joins each row to form a single row with the combined length:\n\n\n",
"_____no_output_____"
]
],
[
[
"ragged_x = tf.ragged.constant([[\"John\"], [\"a\", \"big\", \"dog\"], [\"my\", \"cat\"]])\nragged_y = tf.ragged.constant([[\"fell\", \"asleep\"], [\"barked\"], [\"is\", \"fuzzy\"]])\nprint(tf.concat([ragged_x, ragged_y], axis=1))",
"_____no_output_____"
]
],
[
[
"\nBut concatenating sparse tensors is equivalent to concatenating the corresponding dense tensors,\nas illustrated by the following example (where Ø indicates missing values):\n\n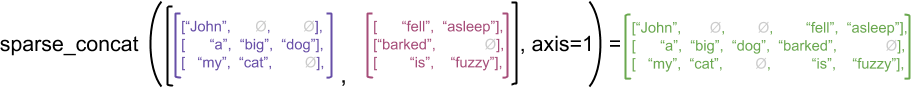\n",
"_____no_output_____"
]
],
[
[
"sparse_x = ragged_x.to_sparse()\nsparse_y = ragged_y.to_sparse()\nsparse_result = tf.sparse.concat(sp_inputs=[sparse_x, sparse_y], axis=1)\nprint(tf.sparse.to_dense(sparse_result, ''))",
"_____no_output_____"
]
],
[
[
"For another example of why this distinction is important, consider the\ndefinition of “the mean value of each row” for an op such as `tf.reduce_mean`.\nFor a ragged tensor, the mean value for a row is the sum of the\nrow’s values divided by the row’s width.\nBut for a sparse tensor, the mean value for a row is the sum of the\nrow’s values divided by the sparse tensor’s overall width (which is\ngreater than or equal to the width of the longest row).\n",
"_____no_output_____"
],
[
"## Overloaded operators\n\nThe `RaggedTensor` class overloads the standard Python arithmetic and comparison\noperators, making it easy to perform basic elementwise math:",
"_____no_output_____"
]
],
[
[
"x = tf.ragged.constant([[1, 2], [3], [4, 5, 6]])\ny = tf.ragged.constant([[1, 1], [2], [3, 3, 3]])\nprint(x + y)",
"_____no_output_____"
]
],
[
[
"Since the overloaded operators perform elementwise computations, the inputs to\nall binary operations must have the same shape, or be broadcastable to the same\nshape. In the simplest broadcasting case, a single scalar is combined\nelementwise with each value in a ragged tensor:",
"_____no_output_____"
]
],
[
[
"x = tf.ragged.constant([[1, 2], [3], [4, 5, 6]])\nprint(x + 3)",
"_____no_output_____"
]
],
[
[
"For a discussion of more advanced cases, see the section on\n**Broadcasting**.\n\nRagged tensors overload the same set of operators as normal `Tensor`s: the unary\noperators `-`, `~`, and `abs()`; and the binary operators `+`, `-`, `*`, `/`,\n`//`, `%`, `**`, `&`, `|`, `^`, `<`, `<=`, `>`, and `>=`. Note that, as with\nstandard `Tensor`s, binary `==` is not overloaded; you can use\n`tf.equal()` to check elementwise equality.",
"_____no_output_____"
],
[
"## Indexing\n\nRagged tensors support Python-style indexing, including multidimensional\nindexing and slicing. The following examples demonstrate ragged tensor indexing\nwith a 2-D and a 3-D ragged tensor.\n\n### Indexing a 2-D ragged tensor with 1 ragged dimension",
"_____no_output_____"
]
],
[
[
"queries = tf.ragged.constant(\n [['Who', 'is', 'George', 'Washington'],\n ['What', 'is', 'the', 'weather', 'tomorrow'],\n ['Goodnight']])\nprint(queries[1])",
"_____no_output_____"
],
[
"print(queries[1, 2]) # A single word",
"_____no_output_____"
],
[
"print(queries[1:]) # Everything but the first row",
"_____no_output_____"
],
[
"print(queries[:, :3]) # The first 3 words of each query",
"_____no_output_____"
],
[
"print(queries[:, -2:]) # The last 2 words of each query",
"_____no_output_____"
]
],
[
[
"### Indexing a 3-D ragged tensor with 2 ragged dimensions",
"_____no_output_____"
]
],
[
[
"rt = tf.ragged.constant([[[1, 2, 3], [4]],\n [[5], [], [6]],\n [[7]],\n [[8, 9], [10]]])",
"_____no_output_____"
],
[
"print(rt[1]) # Second row (2-D RaggedTensor)",
"_____no_output_____"
],
[
"print(rt[3, 0]) # First element of fourth row (1-D Tensor)",
"_____no_output_____"
],
[
"print(rt[:, 1:3]) # Items 1-3 of each row (3-D RaggedTensor)",
"_____no_output_____"
],
[
"print(rt[:, -1:]) # Last item of each row (3-D RaggedTensor)",
"_____no_output_____"
]
],
[
[
"`RaggedTensor`s supports multidimensional indexing and slicing, with one\nrestriction: indexing into a ragged dimension is not allowed. This case is\nproblematic because the indicated value may exist in some rows but not others.\nIn such cases, it's not obvious whether we should (1) raise an `IndexError`; (2)\nuse a default value; or (3) skip that value and return a tensor with fewer rows\nthan we started with. Following the\n[guiding principles of Python](https://www.python.org/dev/peps/pep-0020/)\n(\"In the face\nof ambiguity, refuse the temptation to guess\" ), we currently disallow this\noperation.",
"_____no_output_____"
],
[
"## Tensor Type Conversion\n\nThe `RaggedTensor` class defines methods that can be used to convert\nbetween `RaggedTensor`s and `tf.Tensor`s or `tf.SparseTensors`:",
"_____no_output_____"
]
],
[
[
"ragged_sentences = tf.ragged.constant([\n ['Hi'], ['Welcome', 'to', 'the', 'fair'], ['Have', 'fun']])\nprint(ragged_sentences.to_tensor(default_value=''))",
"_____no_output_____"
],
[
"print(ragged_sentences.to_sparse())",
"_____no_output_____"
],
[
"x = [[1, 3, -1, -1], [2, -1, -1, -1], [4, 5, 8, 9]]\nprint(tf.RaggedTensor.from_tensor(x, padding=-1))",
"_____no_output_____"
],
[
"st = tf.SparseTensor(indices=[[0, 0], [2, 0], [2, 1]],\n values=['a', 'b', 'c'],\n dense_shape=[3, 3])\nprint(tf.RaggedTensor.from_sparse(st))",
"_____no_output_____"
]
],
[
[
"## Evaluating ragged tensors\n\n### Eager execution\n\nIn eager execution mode, ragged tensors are evaluated immediately. To access the\nvalues they contain, you can:\n\n* Use the\n `tf.RaggedTensor.to_list()`\n method, which converts the ragged tensor to a Python `list`.",
"_____no_output_____"
]
],
[
[
"rt = tf.ragged.constant([[1, 2], [3, 4, 5], [6], [], [7]])\nprint(rt.to_list())",
"_____no_output_____"
]
],
[
[
"* Use Python indexing. If the tensor piece you select contains no ragged\n dimensions, then it will be returned as an `EagerTensor`. You can then use\n the `numpy()` method to access the value directly.",
"_____no_output_____"
]
],
[
[
"print(rt[1].numpy())",
"_____no_output_____"
]
],
[
[
"* Decompose the ragged tensor into its components, using the\n `tf.RaggedTensor.values`\n and\n `tf.RaggedTensor.row_splits`\n properties, or row-paritioning methods such as `tf.RaggedTensor.row_lengths()`\n and `tf.RaggedTensor.value_rowids()`.",
"_____no_output_____"
]
],
[
[
"print(rt.values)",
"_____no_output_____"
],
[
"print(rt.row_splits)",
"_____no_output_____"
]
],
[
[
"### Broadcasting\n\nBroadcasting is the process of making tensors with different shapes have\ncompatible shapes for elementwise operations. For more background on\nbroadcasting, see:\n\n* [Numpy: Broadcasting](https://docs.scipy.org/doc/numpy/user/basics.broadcasting.html)\n* `tf.broadcast_dynamic_shape`\n* `tf.broadcast_to`\n\nThe basic steps for broadcasting two inputs `x` and `y` to have compatible\nshapes are:\n\n1. If `x` and `y` do not have the same number of dimensions, then add outer\n dimensions (with size 1) until they do.\n\n2. For each dimension where `x` and `y` have different sizes:\n\n * If `x` or `y` have size `1` in dimension `d`, then repeat its values\n across dimension `d` to match the other input's size.\n\n * Otherwise, raise an exception (`x` and `y` are not broadcast\n compatible).",
"_____no_output_____"
],
[
"Where the size of a tensor in a uniform dimension is a single number (the size\nof slices across that dimension); and the size of a tensor in a ragged dimension\nis a list of slice lengths (for all slices across that dimension).\n\n#### Broadcasting examples",
"_____no_output_____"
]
],
[
[
"# x (2D ragged): 2 x (num_rows)\n# y (scalar)\n# result (2D ragged): 2 x (num_rows)\nx = tf.ragged.constant([[1, 2], [3]])\ny = 3\nprint(x + y)",
"_____no_output_____"
],
[
"# x (2d ragged): 3 x (num_rows)\n# y (2d tensor): 3 x 1\n# Result (2d ragged): 3 x (num_rows)\nx = tf.ragged.constant(\n [[10, 87, 12],\n [19, 53],\n [12, 32]])\ny = [[1000], [2000], [3000]]\nprint(x + y)",
"_____no_output_____"
],
[
"# x (3d ragged): 2 x (r1) x 2\n# y (2d ragged): 1 x 1\n# Result (3d ragged): 2 x (r1) x 2\nx = tf.ragged.constant(\n [[[1, 2], [3, 4], [5, 6]],\n [[7, 8]]],\n ragged_rank=1)\ny = tf.constant([[10]])\nprint(x + y)",
"_____no_output_____"
],
[
"# x (3d ragged): 2 x (r1) x (r2) x 1\n# y (1d tensor): 3\n# Result (3d ragged): 2 x (r1) x (r2) x 3\nx = tf.ragged.constant(\n [\n [\n [[1], [2]],\n [],\n [[3]],\n [[4]],\n ],\n [\n [[5], [6]],\n [[7]]\n ]\n ],\n ragged_rank=2)\ny = tf.constant([10, 20, 30])\nprint(x + y)",
"_____no_output_____"
]
],
[
[
"Here are some examples of shapes that do not broadcast:",
"_____no_output_____"
]
],
[
[
"# x (2d ragged): 3 x (r1)\n# y (2d tensor): 3 x 4 # trailing dimensions do not match\nx = tf.ragged.constant([[1, 2], [3, 4, 5, 6], [7]])\ny = tf.constant([[1, 2, 3, 4], [5, 6, 7, 8], [9, 10, 11, 12]])\ntry:\n x + y\nexcept tf.errors.InvalidArgumentError as exception:\n print(exception)",
"_____no_output_____"
],
[
"# x (2d ragged): 3 x (r1)\n# y (2d ragged): 3 x (r2) # ragged dimensions do not match.\nx = tf.ragged.constant([[1, 2, 3], [4], [5, 6]])\ny = tf.ragged.constant([[10, 20], [30, 40], [50]])\ntry:\n x + y\nexcept tf.errors.InvalidArgumentError as exception:\n print(exception)",
"_____no_output_____"
],
[
"# x (3d ragged): 3 x (r1) x 2\n# y (3d ragged): 3 x (r1) x 3 # trailing dimensions do not match\nx = tf.ragged.constant([[[1, 2], [3, 4], [5, 6]],\n [[7, 8], [9, 10]]])\ny = tf.ragged.constant([[[1, 2, 0], [3, 4, 0], [5, 6, 0]],\n [[7, 8, 0], [9, 10, 0]]])\ntry:\n x + y\nexcept tf.errors.InvalidArgumentError as exception:\n print(exception)",
"_____no_output_____"
]
],
[
[
"## RaggedTensor encoding\n\nRagged tensors are encoded using the `RaggedTensor` class. Internally, each\n`RaggedTensor` consists of:\n\n* A `values` tensor, which concatenates the variable-length rows into a\n flattened list.\n* A `row_splits` vector, which indicates how those flattened values are\n divided into rows. In particular, the values for row `rt[i]` are stored in\n the slice `rt.values[rt.row_splits[i]:rt.row_splits[i+1]]`.\n\n\n\n",
"_____no_output_____"
]
],
[
[
"rt = tf.RaggedTensor.from_row_splits(\n values=[3, 1, 4, 1, 5, 9, 2],\n row_splits=[0, 4, 4, 6, 7])\nprint(rt)",
"_____no_output_____"
]
],
[
[
"### Multiple ragged dimensions\n\nA ragged tensor with multiple ragged dimensions is encoded by using a nested\n`RaggedTensor` for the `values` tensor. Each nested `RaggedTensor` adds a single\nragged dimension.\n\n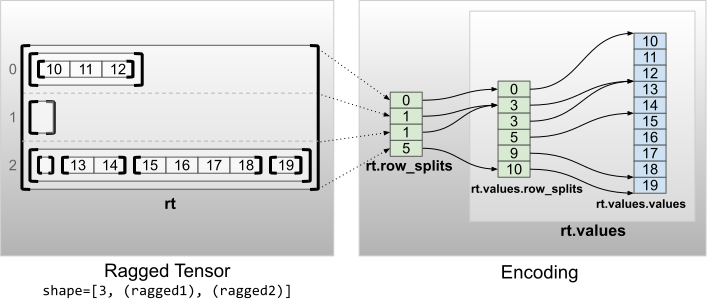",
"_____no_output_____"
]
],
[
[
"rt = tf.RaggedTensor.from_row_splits(\n values=tf.RaggedTensor.from_row_splits(\n values=[10, 11, 12, 13, 14, 15, 16, 17, 18, 19],\n row_splits=[0, 3, 3, 5, 9, 10]),\n row_splits=[0, 1, 1, 5])\nprint(rt)\nprint(\"Shape: {}\".format(rt.shape))\nprint(\"Number of ragged dimensions: {}\".format(rt.ragged_rank))",
"_____no_output_____"
]
],
[
[
"The factory function `tf.RaggedTensor.from_nested_row_splits` may be used to construct a\nRaggedTensor with multiple ragged dimensions directly, by providing a list of\n`row_splits` tensors:",
"_____no_output_____"
]
],
[
[
"rt = tf.RaggedTensor.from_nested_row_splits(\n flat_values=[10, 11, 12, 13, 14, 15, 16, 17, 18, 19],\n nested_row_splits=([0, 1, 1, 5], [0, 3, 3, 5, 9, 10]))\nprint(rt)",
"_____no_output_____"
]
],
[
[
"### Uniform Inner Dimensions\n\nRagged tensors with uniform inner dimensions are encoded by using a\nmultidimensional `tf.Tensor` for `values`.\n\n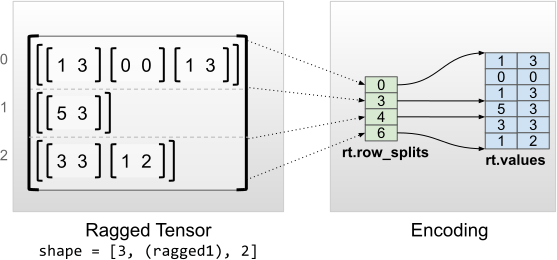",
"_____no_output_____"
]
],
[
[
"rt = tf.RaggedTensor.from_row_splits(\n values=[[1, 3], [0, 0], [1, 3], [5, 3], [3, 3], [1, 2]],\n row_splits=[0, 3, 4, 6])\nprint(rt)\nprint(\"Shape: {}\".format(rt.shape))\nprint(\"Number of ragged dimensions: {}\".format(rt.ragged_rank))",
"_____no_output_____"
]
],
[
[
"### Alternative row-partitioning schemes\n\nThe `RaggedTensor` class uses `row_splits` as the primary mechanism to store\ninformation about how the values are partitioned into rows. However,\n`RaggedTensor` also provides support for four alternative row-partitioning\nschemes, which can be more convenient to use depending on how your data is\nformatted. Internally, `RaggedTensor` uses these additional schemes to improve\nefficiency in some contexts.\n\n<dl>\n <dt>Row lengths</dt>\n <dd>`row_lengths` is a vector with shape `[nrows]`, which specifies the\n length of each row.</dd>\n\n <dt>Row starts</dt>\n <dd>`row_starts` is a vector with shape `[nrows]`, which specifies the start\n offset of each row. Equivalent to `row_splits[:-1]`.</dd>\n\n <dt>Row limits</dt>\n <dd>`row_limits` is a vector with shape `[nrows]`, which specifies the stop\n offset of each row. Equivalent to `row_splits[1:]`.</dd>\n\n <dt>Row indices and number of rows</dt>\n <dd>`value_rowids` is a vector with shape `[nvals]`, corresponding\n one-to-one with values, which specifies each value's row index. In\n particular, the row `rt[row]` consists of the values `rt.values[j]` where\n `value_rowids[j]==row`. \\\n `nrows` is an integer that specifies the number of rows in the\n `RaggedTensor`. In particular, `nrows` is used to indicate trailing empty\n rows.</dd>\n</dl>\n\nFor example, the following ragged tensors are equivalent:",
"_____no_output_____"
]
],
[
[
"values = [3, 1, 4, 1, 5, 9, 2, 6]\nprint(tf.RaggedTensor.from_row_splits(values, row_splits=[0, 4, 4, 7, 8, 8]))\nprint(tf.RaggedTensor.from_row_lengths(values, row_lengths=[4, 0, 3, 1, 0]))\nprint(tf.RaggedTensor.from_row_starts(values, row_starts=[0, 4, 4, 7, 8]))\nprint(tf.RaggedTensor.from_row_limits(values, row_limits=[4, 4, 7, 8, 8]))\nprint(tf.RaggedTensor.from_value_rowids(\n values, value_rowids=[0, 0, 0, 0, 2, 2, 2, 3], nrows=5))",
"_____no_output_____"
]
],
[
[
"The RaggedTensor class defines methods which can be used to construct\neach of these row-partitioning tensors.",
"_____no_output_____"
]
],
[
[
"rt = tf.ragged.constant([[3, 1, 4, 1], [], [5, 9, 2], [6], []])\nprint(\" values: {}\".format(rt.values))\nprint(\" row_splits: {}\".format(rt.row_splits))\nprint(\" row_lengths: {}\".format(rt.row_lengths()))\nprint(\" row_starts: {}\".format(rt.row_starts()))\nprint(\" row_limits: {}\".format(rt.row_limits()))\nprint(\"value_rowids: {}\".format(rt.value_rowids()))",
"_____no_output_____"
]
],
[
[
"(Note that `tf.RaggedTensor.values` and `tf.RaggedTensors.row_splits` are properties, while the remaining row-partitioning accessors are all methods. This reflects the fact that the `row_splits` are the primary underlying representation, and the other row-partitioning tensors must be computed.)",
"_____no_output_____"
],
[
"Some of the advantages and disadvantages of the different row-partitioning\nschemes are:\n\n+ **Efficient indexing**:\n The `row_splits`, `row_starts`, and `row_limits` schemes all enable\n constant-time indexing into ragged tensors. The `value_rowids` and\n `row_lengths` schemes do not.\n\n+ **Small encoding size**:\n The `value_rowids` scheme is more efficient when storing ragged tensors that\n have a large number of empty rows, since the size of the tensor depends only\n on the total number of values. On the other hand, the other four encodings\n are more efficient when storing ragged tensors with longer rows, since they\n require only one scalar value for each row.\n\n+ **Efficient concatenation**:\n The `row_lengths` scheme is more efficient when concatenating ragged\n tensors, since row lengths do not change when two tensors are concatenated\n together (but row splits and row indices do).\n\n+ **Compatibility**:\n The `value_rowids` scheme matches the\n [segmentation](../api_guides/python/math_ops.md#Segmentation)\n format used by operations such as `tf.segment_sum`. The `row_limits` scheme\n matches the format used by ops such as `tf.sequence_mask`.",
"_____no_output_____"
]
],
[
[
"",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code"
],
[
"markdown",
"markdown"
],
[
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown"
],
[
"code"
]
] |
4acff29b4dda609177429bde091bbe359ea9d9a0
| 658,836 |
ipynb
|
Jupyter Notebook
|
code/11_YOLO.ipynb
|
Akshatha-Jagadish/DL_topics
|
98aa979dde2021a20e7b561b83230ac0a475cf5e
|
[
"MIT"
] | null | null | null |
code/11_YOLO.ipynb
|
Akshatha-Jagadish/DL_topics
|
98aa979dde2021a20e7b561b83230ac0a475cf5e
|
[
"MIT"
] | null | null | null |
code/11_YOLO.ipynb
|
Akshatha-Jagadish/DL_topics
|
98aa979dde2021a20e7b561b83230ac0a475cf5e
|
[
"MIT"
] | null | null | null | 744.447458 | 537,630 | 0.9044 |
[
[
[
"#object detection using YOLO v4 codebasics video - also has paper links:\n#https://www.youtube.com/watch?v=IfRMV2MY9n0&list=PLeo1K3hjS3uu7CxAacxVndI4bE_o3BDtO&index=32\n\n# yolov4 repository on github: https://github.com/AlexeyAB/darknet#yolo-v4-v3-and-v2-for-windows-and-linux\n# for using yolov4 on colab: https://colab.research.google.com/drive/12QusaaRj_lUwCGDvQNfICpa7kA7_a2dE?usp=sharing",
"_____no_output_____"
],
[
"# clone darknet repo\n!git clone https://github.com/AlexeyAB/darknet",
"fatal: destination path 'darknet' already exists and is not an empty directory.\n"
],
[
"# change makefile to have GPU and OPENCV enabled\n%cd darknet\n!sed -i 's/OPENCV=0/OPENCV=1/' Makefile\n!sed -i 's/GPU=0/GPU=1/' Makefile\n!sed -i 's/CUDNN=0/CUDNN=1/' Makefile\n!sed -i 's/CUDNN_HALF=0/CUDNN_HALF=1/' Makefile",
"[Errno 20] Not a directory: 'darknet'\n/content/darknet\n"
],
[
"# verify CUDA\n!/usr/local/cuda/bin/nvcc --version",
"nvcc: NVIDIA (R) Cuda compiler driver\nCopyright (c) 2005-2020 NVIDIA Corporation\nBuilt on Mon_Oct_12_20:09:46_PDT_2020\nCuda compilation tools, release 11.1, V11.1.105\nBuild cuda_11.1.TC455_06.29190527_0\n"
],
[
"# make darknet (builds darknet so that you can then use the darknet executable file to run or train object detectors)\n!make",
"chmod +x *.sh\ng++ -std=c++11 -std=c++11 -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/image_opencv.cpp -o obj/image_opencv.o\n\u001b[01m\u001b[K./src/image_opencv.cpp:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kvoid draw_detections_cv_v3(void**, detection*, int, float, char**, image**, int, int)\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/image_opencv.cpp:946:23:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kvariable ‘\u001b[01m\u001b[Krgb\u001b[m\u001b[K’ set but not used [\u001b[01;35m\u001b[K-Wunused-but-set-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Krgb\u001b[m\u001b[K[3];\n \u001b[01;35m\u001b[K^~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/image_opencv.cpp:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kvoid draw_train_loss(char*, void**, int, float, float, int, int, float, int, char*, float, int, int, double)\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/image_opencv.cpp:1147:13:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kthis ‘\u001b[01m\u001b[Kif\u001b[m\u001b[K’ clause does not guard... [\u001b[01;35m\u001b[K-Wmisleading-indentation\u001b[m\u001b[K]\n \u001b[01;35m\u001b[Kif\u001b[m\u001b[K (iteration_old == 0)\n \u001b[01;35m\u001b[K^~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/image_opencv.cpp:1150:10:\u001b[m\u001b[K \u001b[01;36m\u001b[Knote: \u001b[m\u001b[K...this statement, but the latter is misleadingly indented as if it were guarded by the ‘\u001b[01m\u001b[Kif\u001b[m\u001b[K’\n \u001b[01;36m\u001b[Kif\u001b[m\u001b[K (iteration_old != 0){\n \u001b[01;36m\u001b[K^~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/image_opencv.cpp:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kvoid cv_draw_object(image, float*, int, int, int*, float*, int*, int, char**)\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/image_opencv.cpp:1444:14:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kbuff\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n char \u001b[01;35m\u001b[Kbuff\u001b[m\u001b[K[100];\n \u001b[01;35m\u001b[K^~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/image_opencv.cpp:1420:9:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kit_tb_res\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Kit_tb_res\u001b[m\u001b[K = cv::createTrackbar(it_trackbar_name, window_name, &it_trackbar_value, 1000);\n \u001b[01;35m\u001b[K^~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/image_opencv.cpp:1424:9:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Klr_tb_res\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Klr_tb_res\u001b[m\u001b[K = cv::createTrackbar(lr_trackbar_name, window_name, &lr_trackbar_value, 20);\n \u001b[01;35m\u001b[K^~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/image_opencv.cpp:1428:9:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kcl_tb_res\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Kcl_tb_res\u001b[m\u001b[K = cv::createTrackbar(cl_trackbar_name, window_name, &cl_trackbar_value, classes-1);\n \u001b[01;35m\u001b[K^~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/image_opencv.cpp:1431:9:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kbo_tb_res\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Kbo_tb_res\u001b[m\u001b[K = cv::createTrackbar(bo_trackbar_name, window_name, boxonly, 1);\n \u001b[01;35m\u001b[K^~~~~~~~~\u001b[m\u001b[K\ng++ -std=c++11 -std=c++11 -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/http_stream.cpp -o obj/http_stream.o\n\u001b[01m\u001b[K./src/http_stream.cpp:\u001b[m\u001b[K In member function ‘\u001b[01m\u001b[Kbool JSON_sender::write(const char*)\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/http_stream.cpp:253:21:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kn\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Kn\u001b[m\u001b[K = _write(client, outputbuf, outlen);\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/http_stream.cpp:\u001b[m\u001b[K In member function ‘\u001b[01m\u001b[Kbool MJPG_sender::write(const cv::Mat&)\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/http_stream.cpp:511:113:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%zu\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Ksize_t\u001b[m\u001b[K’, but argument 3 has type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n sprintf(head, \"--mjpegstream\\r\\nContent-Type: image/jpeg\\r\\nContent-Length: %zu\\r\\n\\r\\n\", outlen\u001b[01;35m\u001b[K)\u001b[m\u001b[K;\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/http_stream.cpp:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kvoid set_track_id(detection*, int, float, float, float, int, int, int)\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/http_stream.cpp:867:27:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kcomparison between signed and unsigned integer expressions [\u001b[01;35m\u001b[K-Wsign-compare\u001b[m\u001b[K]\n for (int i = 0; \u001b[01;35m\u001b[Ki < v.size()\u001b[m\u001b[K; ++i) {\n \u001b[01;35m\u001b[K~~^~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/http_stream.cpp:875:33:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kcomparison between signed and unsigned integer expressions [\u001b[01;35m\u001b[K-Wsign-compare\u001b[m\u001b[K]\n for (int old_id = 0; \u001b[01;35m\u001b[Kold_id < old_dets.size()\u001b[m\u001b[K; ++old_id) {\n \u001b[01;35m\u001b[K~~~~~~~^~~~~~~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/http_stream.cpp:894:31:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kcomparison between signed and unsigned integer expressions [\u001b[01;35m\u001b[K-Wsign-compare\u001b[m\u001b[K]\n for (int index = 0; \u001b[01;35m\u001b[Kindex < new_dets_num*old_dets.size()\u001b[m\u001b[K; ++index) {\n \u001b[01;35m\u001b[K~~~~~~^~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/http_stream.cpp:930:28:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kcomparison between signed and unsigned integer expressions [\u001b[01;35m\u001b[K-Wsign-compare\u001b[m\u001b[K]\n if (\u001b[01;35m\u001b[Kold_dets_dq.size() > deque_size\u001b[m\u001b[K) old_dets_dq.pop_front();\n \u001b[01;35m\u001b[K~~~~~~~~~~~~~~~~~~~^~~~~~~~~~~~\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/gemm.c -o obj/gemm.o\n\u001b[01m\u001b[K./src/gemm.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kconvolution_2d\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/gemm.c:2044:15:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kout_w\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n const int \u001b[01;35m\u001b[Kout_w\u001b[m\u001b[K = (w + 2 * pad - ksize) / stride + 1; // output_width=input_width for stride=1 and pad=1\n \u001b[01;35m\u001b[K^~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/gemm.c:2043:15:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kout_h\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n const int \u001b[01;35m\u001b[Kout_h\u001b[m\u001b[K = (h + 2 * pad - ksize) / stride + 1; // output_height=input_height for stride=1 and pad=1\n \u001b[01;35m\u001b[K^~~~~\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/utils.c -o obj/utils.o\n\u001b[01m\u001b[K./src/utils.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kcustom_hash\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/utils.c:1061:12:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Ksuggest parentheses around assignment used as truth value [\u001b[01;35m\u001b[K-Wparentheses\u001b[m\u001b[K]\n while (\u001b[01;35m\u001b[Kc\u001b[m\u001b[K = *str++)\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/dark_cuda.c -o obj/dark_cuda.o\n\u001b[01m\u001b[K./src/dark_cuda.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kcudnn_check_error_extended\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/dark_cuda.c:230:20:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kcomparison between ‘\u001b[01m\u001b[KcudaError_t {aka enum cudaError}\u001b[m\u001b[K’ and ‘\u001b[01m\u001b[Kenum <anonymous>\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wenum-compare\u001b[m\u001b[K]\n if (status \u001b[01;35m\u001b[K!=\u001b[m\u001b[K CUDNN_STATUS_SUCCESS)\n \u001b[01;35m\u001b[K^~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/dark_cuda.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kcublas_check_error_extended\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/dark_cuda.c:264:18:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kcomparison between ‘\u001b[01m\u001b[KcudaError_t {aka enum cudaError}\u001b[m\u001b[K’ and ‘\u001b[01m\u001b[Kenum cudaError_enum\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wenum-compare\u001b[m\u001b[K]\n if (status \u001b[01;35m\u001b[K!=\u001b[m\u001b[K CUDA_SUCCESS)\n \u001b[01;35m\u001b[K^~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/dark_cuda.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kpre_allocate_pinned_memory\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/dark_cuda.c:395:40:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%u\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kunsigned int\u001b[m\u001b[K’, but argument 2 has type ‘\u001b[01m\u001b[Klong unsigned int\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n printf(\"pre_allocate: size = \u001b[01;35m\u001b[K%Iu\u001b[m\u001b[K MB, num_of_blocks = %Iu, block_size = %Iu MB \\n\",\n \u001b[01;35m\u001b[K~~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%Ilu\u001b[m\u001b[K\n \u001b[32m\u001b[Ksize / (1024*1024)\u001b[m\u001b[K, num_of_blocks, pinned_block_size / (1024 * 1024));\n \u001b[32m\u001b[K~~~~~~~~~~~~~~~~~~\u001b[m\u001b[K \n\u001b[01m\u001b[K./src/dark_cuda.c:395:64:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%u\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kunsigned int\u001b[m\u001b[K’, but argument 3 has type ‘\u001b[01m\u001b[Ksize_t {aka const long unsigned int}\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n printf(\"pre_allocate: size = %Iu MB, num_of_blocks = \u001b[01;35m\u001b[K%Iu\u001b[m\u001b[K, block_size = %Iu MB \\n\",\n \u001b[01;35m\u001b[K~~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%Ilu\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/dark_cuda.c:395:82:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%u\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kunsigned int\u001b[m\u001b[K’, but argument 4 has type ‘\u001b[01m\u001b[Klong unsigned int\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n printf(\"pre_allocate: size = %Iu MB, num_of_blocks = %Iu, block_size = \u001b[01;35m\u001b[K%Iu\u001b[m\u001b[K MB \\n\",\n \u001b[01;35m\u001b[K~~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%Ilu\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/dark_cuda.c:405:37:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 2 has type ‘\u001b[01m\u001b[Ksize_t {aka const long unsigned int}\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n printf(\" Allocated \u001b[01;35m\u001b[K%d\u001b[m\u001b[K pinned block \\n\", pinned_block_size);\n \u001b[01;35m\u001b[K~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%ld\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/dark_cuda.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kcuda_make_array_pinned_preallocated\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/dark_cuda.c:426:43:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 2 has type ‘\u001b[01m\u001b[Ksize_t {aka long unsigned int}\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n printf(\"\\n Pinned block_id = \u001b[01;35m\u001b[K%d\u001b[m\u001b[K, filled = %f %% \\n\", pinned_block_id, filled);\n \u001b[01;35m\u001b[K~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%ld\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/dark_cuda.c:441:64:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 2 has type ‘\u001b[01m\u001b[Klong unsigned int\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n printf(\"Try to allocate new pinned memory, size = \u001b[01;35m\u001b[K%d\u001b[m\u001b[K MB \\n\", \u001b[32m\u001b[Ksize / (1024 * 1024)\u001b[m\u001b[K);\n \u001b[01;35m\u001b[K~^\u001b[m\u001b[K \u001b[32m\u001b[K~~~~~~~~~~~~~~~~~~~~\u001b[m\u001b[K\n \u001b[32m\u001b[K%ld\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/dark_cuda.c:447:63:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 2 has type ‘\u001b[01m\u001b[Klong unsigned int\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n printf(\"Try to allocate new pinned BLOCK, size = \u001b[01;35m\u001b[K%d\u001b[m\u001b[K MB \\n\", \u001b[32m\u001b[Ksize / (1024 * 1024)\u001b[m\u001b[K);\n \u001b[01;35m\u001b[K~^\u001b[m\u001b[K \u001b[32m\u001b[K~~~~~~~~~~~~~~~~~~~~\u001b[m\u001b[K\n \u001b[32m\u001b[K%ld\u001b[m\u001b[K\nAt top level:\n\u001b[01m\u001b[K./src/dark_cuda.c:287:23:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[K‘\u001b[01m\u001b[KswitchBlasHandle\u001b[m\u001b[K’ defined but not used [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n static cublasHandle_t \u001b[01;35m\u001b[KswitchBlasHandle\u001b[m\u001b[K[16];\n \u001b[01;35m\u001b[K^~~~~~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/dark_cuda.c:286:12:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[K‘\u001b[01m\u001b[KswitchBlasInit\u001b[m\u001b[K’ defined but not used [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n static int \u001b[01;35m\u001b[KswitchBlasInit\u001b[m\u001b[K[16] = { 0 };\n \u001b[01;35m\u001b[K^~~~~~~~~~~~~~\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/convolutional_layer.c -o obj/convolutional_layer.o\n\u001b[01m\u001b[K./src/convolutional_layer.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kforward_convolutional_layer\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/convolutional_layer.c:1342:32:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kt_intput_size\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n size_t \u001b[01;35m\u001b[Kt_intput_size\u001b[m\u001b[K = binary_transpose_align_input(k, n, state.workspace, &l.t_bit_input, ldb_align, l.bit_align);\n \u001b[01;35m\u001b[K^~~~~~~~~~~~~\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/list.c -o obj/list.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/image.c -o obj/image.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/activations.c -o obj/activations.o\n\u001b[01m\u001b[K./src/activations.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kactivate\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/activations.c:79:5:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kenumeration value ‘\u001b[01m\u001b[KRELU6\u001b[m\u001b[K’ not handled in switch [\u001b[01;35m\u001b[K-Wswitch\u001b[m\u001b[K]\n \u001b[01;35m\u001b[Kswitch\u001b[m\u001b[K(a){\n \u001b[01;35m\u001b[K^~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/activations.c:79:5:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kenumeration value ‘\u001b[01m\u001b[KSWISH\u001b[m\u001b[K’ not handled in switch [\u001b[01;35m\u001b[K-Wswitch\u001b[m\u001b[K]\n\u001b[01m\u001b[K./src/activations.c:79:5:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kenumeration value ‘\u001b[01m\u001b[KMISH\u001b[m\u001b[K’ not handled in switch [\u001b[01;35m\u001b[K-Wswitch\u001b[m\u001b[K]\n\u001b[01m\u001b[K./src/activations.c:79:5:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kenumeration value ‘\u001b[01m\u001b[KHARD_MISH\u001b[m\u001b[K’ not handled in switch [\u001b[01;35m\u001b[K-Wswitch\u001b[m\u001b[K]\n\u001b[01m\u001b[K./src/activations.c:79:5:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kenumeration value ‘\u001b[01m\u001b[KNORM_CHAN\u001b[m\u001b[K’ not handled in switch [\u001b[01;35m\u001b[K-Wswitch\u001b[m\u001b[K]\n\u001b[01m\u001b[K./src/activations.c:79:5:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kenumeration value ‘\u001b[01m\u001b[KNORM_CHAN_SOFTMAX\u001b[m\u001b[K’ not handled in switch [\u001b[01;35m\u001b[K-Wswitch\u001b[m\u001b[K]\n\u001b[01m\u001b[K./src/activations.c:79:5:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kenumeration value ‘\u001b[01m\u001b[KNORM_CHAN_SOFTMAX_MAXVAL\u001b[m\u001b[K’ not handled in switch [\u001b[01;35m\u001b[K-Wswitch\u001b[m\u001b[K]\n\u001b[01m\u001b[K./src/activations.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kgradient\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/activations.c:310:5:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kenumeration value ‘\u001b[01m\u001b[KSWISH\u001b[m\u001b[K’ not handled in switch [\u001b[01;35m\u001b[K-Wswitch\u001b[m\u001b[K]\n \u001b[01;35m\u001b[Kswitch\u001b[m\u001b[K(a){\n \u001b[01;35m\u001b[K^~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/activations.c:310:5:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kenumeration value ‘\u001b[01m\u001b[KMISH\u001b[m\u001b[K’ not handled in switch [\u001b[01;35m\u001b[K-Wswitch\u001b[m\u001b[K]\n\u001b[01m\u001b[K./src/activations.c:310:5:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kenumeration value ‘\u001b[01m\u001b[KHARD_MISH\u001b[m\u001b[K’ not handled in switch [\u001b[01;35m\u001b[K-Wswitch\u001b[m\u001b[K]\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/im2col.c -o obj/im2col.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/col2im.c -o obj/col2im.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/blas.c -o obj/blas.o\n\u001b[01m\u001b[K./src/blas.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kbackward_shortcut_multilayer_cpu\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/blas.c:207:21:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kout_index\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Kout_index\u001b[m\u001b[K = id;\n \u001b[01;35m\u001b[K^~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/blas.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kfind_sim\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/blas.c:597:59:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 2 has type ‘\u001b[01m\u001b[Ksize_t {aka long unsigned int}\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n printf(\" Error: find_sim(): sim isn't found: i = \u001b[01;35m\u001b[K%d\u001b[m\u001b[K, j = %d, z = %d \\n\", i, j, z);\n \u001b[01;35m\u001b[K~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%ld\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/blas.c:597:67:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 3 has type ‘\u001b[01m\u001b[Ksize_t {aka long unsigned int}\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n printf(\" Error: find_sim(): sim isn't found: i = %d, j = \u001b[01;35m\u001b[K%d\u001b[m\u001b[K, z = %d \\n\", i, j, z);\n \u001b[01;35m\u001b[K~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%ld\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/blas.c:597:75:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 4 has type ‘\u001b[01m\u001b[Ksize_t {aka long unsigned int}\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n printf(\" Error: find_sim(): sim isn't found: i = %d, j = %d, z = \u001b[01;35m\u001b[K%d\u001b[m\u001b[K \\n\", i, j, z);\n \u001b[01;35m\u001b[K~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%ld\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/blas.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kfind_P_constrastive\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/blas.c:611:68:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 2 has type ‘\u001b[01m\u001b[Ksize_t {aka long unsigned int}\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n printf(\" Error: find_P_constrastive(): P isn't found: i = \u001b[01;35m\u001b[K%d\u001b[m\u001b[K, j = %d, z = %d \\n\", i, j, z);\n \u001b[01;35m\u001b[K~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%ld\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/blas.c:611:76:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 3 has type ‘\u001b[01m\u001b[Ksize_t {aka long unsigned int}\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n printf(\" Error: find_P_constrastive(): P isn't found: i = %d, j = \u001b[01;35m\u001b[K%d\u001b[m\u001b[K, z = %d \\n\", i, j, z);\n \u001b[01;35m\u001b[K~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%ld\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/blas.c:611:84:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 4 has type ‘\u001b[01m\u001b[Ksize_t {aka long unsigned int}\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n printf(\" Error: find_P_constrastive(): P isn't found: i = %d, j = %d, z = \u001b[01;35m\u001b[K%d\u001b[m\u001b[K \\n\", i, j, z);\n \u001b[01;35m\u001b[K~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%ld\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/blas.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[KP_constrastive_f\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/blas.c:651:79:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 3 has type ‘\u001b[01m\u001b[Ksize_t {aka long unsigned int}\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n fprintf(stderr, \" Error: in P_constrastive must be i != l, while i = \u001b[01;35m\u001b[K%d\u001b[m\u001b[K, l = %d \\n\", i, l);\n \u001b[01;35m\u001b[K~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%ld\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/blas.c:651:87:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 4 has type ‘\u001b[01m\u001b[Ksize_t {aka long unsigned int}\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n fprintf(stderr, \" Error: in P_constrastive must be i != l, while i = %d, l = \u001b[01;35m\u001b[K%d\u001b[m\u001b[K \\n\", i, l);\n \u001b[01;35m\u001b[K~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%ld\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/blas.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[KP_constrastive\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/blas.c:785:79:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 3 has type ‘\u001b[01m\u001b[Ksize_t {aka long unsigned int}\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n fprintf(stderr, \" Error: in P_constrastive must be i != l, while i = \u001b[01;35m\u001b[K%d\u001b[m\u001b[K, l = %d \\n\", i, l);\n \u001b[01;35m\u001b[K~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%ld\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/blas.c:785:87:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 4 has type ‘\u001b[01m\u001b[Ksize_t {aka long unsigned int}\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n fprintf(stderr, \" Error: in P_constrastive must be i != l, while i = %d, l = \u001b[01;35m\u001b[K%d\u001b[m\u001b[K \\n\", i, l);\n \u001b[01;35m\u001b[K~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%ld\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/crop_layer.c -o obj/crop_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/dropout_layer.c -o obj/dropout_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/maxpool_layer.c -o obj/maxpool_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/softmax_layer.c -o obj/softmax_layer.o\n\u001b[01m\u001b[K./src/softmax_layer.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kmake_contrastive_layer\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/softmax_layer.c:203:101:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 9 has type ‘\u001b[01m\u001b[Ksize_t {aka const long unsigned int}\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n fprintf(stderr, \"contrastive %4d x%4d x%4d x emb_size %4d x batch: %4d classes = %4d, step = \u001b[01;35m\u001b[K%4d\u001b[m\u001b[K \\n\", w, h, l.n, l.embedding_size, batch, l.classes, step);\n \u001b[01;35m\u001b[K~~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%4ld\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/softmax_layer.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kforward_contrastive_layer\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/softmax_layer.c:244:27:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kvariable ‘\u001b[01m\u001b[Kmax_truth\u001b[m\u001b[K’ set but not used [\u001b[01;35m\u001b[K-Wunused-but-set-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Kmax_truth\u001b[m\u001b[K = 0;\n \u001b[01;35m\u001b[K^~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/softmax_layer.c:423:71:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat ‘\u001b[01m\u001b[K%d\u001b[m\u001b[K’ expects argument of type ‘\u001b[01m\u001b[Kint\u001b[m\u001b[K’, but argument 2 has type ‘\u001b[01m\u001b[Ksize_t {aka const long unsigned int}\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wformat=\u001b[m\u001b[K]\n printf(\" Error: too large number of bboxes: contr_size = \u001b[01;35m\u001b[K%d\u001b[m\u001b[K > max_contr_size = %d \\n\", contr_size, max_contr_size);\n \u001b[01;35m\u001b[K~^\u001b[m\u001b[K\n \u001b[32m\u001b[K%ld\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/data.c -o obj/data.o\n\u001b[01m\u001b[K./src/data.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kload_data_detection\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/data.c:1297:24:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kx\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int k, \u001b[01;35m\u001b[Kx\u001b[m\u001b[K, y;\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/data.c:1090:43:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kvariable ‘\u001b[01m\u001b[Kr_scale\u001b[m\u001b[K’ set but not used [\u001b[01;35m\u001b[K-Wunused-but-set-variable\u001b[m\u001b[K]\n float r1 = 0, r2 = 0, r3 = 0, r4 = 0, \u001b[01;35m\u001b[Kr_scale\u001b[m\u001b[K = 0;\n \u001b[01;35m\u001b[K^~~~~~~\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/matrix.c -o obj/matrix.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/network.c -o obj/network.o\n\u001b[01m\u001b[K./src/network.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Ktrain_network_waitkey\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/network.c:435:13:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kema_period\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Kema_period\u001b[m\u001b[K = (net.max_batches - ema_start_point - 1000) * (1.0 - net.ema_alpha);\n \u001b[01;35m\u001b[K^~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/network.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kresize_network\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/network.c:660:42:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kpassing argument 1 of ‘\u001b[01m\u001b[KcudaHostAlloc\u001b[m\u001b[K’ from incompatible pointer type [\u001b[01;35m\u001b[K-Wincompatible-pointer-types\u001b[m\u001b[K]\n if (cudaSuccess == cudaHostAlloc(\u001b[01;35m\u001b[K&\u001b[m\u001b[Knet->input_pinned_cpu, size * sizeof(float), cudaHostRegisterMapped))\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\nIn file included from \u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime.h:96:0\u001b[m\u001b[K,\n from \u001b[01m\u001b[Kinclude/darknet.h:41\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/network.c:1\u001b[m\u001b[K:\n\u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime_api.h:4811:39:\u001b[m\u001b[K \u001b[01;36m\u001b[Knote: \u001b[m\u001b[Kexpected ‘\u001b[01m\u001b[Kvoid **\u001b[m\u001b[K’ but argument is of type ‘\u001b[01m\u001b[Kfloat **\u001b[m\u001b[K’\n extern __host__ cudaError_t CUDARTAPI \u001b[01;36m\u001b[KcudaHostAlloc\u001b[m\u001b[K(void **pHost, size_t size, unsigned int flags);\n \u001b[01;36m\u001b[K^~~~~~~~~~~~~\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/connected_layer.c -o obj/connected_layer.o\n\u001b[01m\u001b[K./src/connected_layer.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kforward_connected_layer_gpu\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/connected_layer.c:346:11:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kone\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Kone\u001b[m\u001b[K = 1; // alpha[0], beta[0]\n \u001b[01;35m\u001b[K^~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/connected_layer.c:344:13:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kc\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float * \u001b[01;35m\u001b[Kc\u001b[m\u001b[K = l.output_gpu;\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/connected_layer.c:343:13:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kb\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float * \u001b[01;35m\u001b[Kb\u001b[m\u001b[K = l.weights_gpu;\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/connected_layer.c:342:13:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Ka\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float * \u001b[01;35m\u001b[Ka\u001b[m\u001b[K = state.input;\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/connected_layer.c:341:9:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kn\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Kn\u001b[m\u001b[K = l.outputs;\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/connected_layer.c:340:9:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kk\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Kk\u001b[m\u001b[K = l.inputs;\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/connected_layer.c:339:9:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Km\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Km\u001b[m\u001b[K = l.batch;\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/cost_layer.c -o obj/cost_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/parser.c -o obj/parser.o\n\u001b[01m\u001b[K./src/parser.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kparse_network_cfg_custom\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/parser.c:1777:42:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kpassing argument 1 of ‘\u001b[01m\u001b[KcudaHostAlloc\u001b[m\u001b[K’ from incompatible pointer type [\u001b[01;35m\u001b[K-Wincompatible-pointer-types\u001b[m\u001b[K]\n if (cudaSuccess == cudaHostAlloc(\u001b[01;35m\u001b[K&\u001b[m\u001b[Knet.input_pinned_cpu, size * sizeof(float), cudaHostRegisterMapped)) net.input_pinned_cpu_flag = 1;\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\nIn file included from \u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime.h:96:0\u001b[m\u001b[K,\n from \u001b[01m\u001b[Kinclude/darknet.h:41\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/activations.h:3\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/activation_layer.h:4\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/parser.c:6\u001b[m\u001b[K:\n\u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime_api.h:4811:39:\u001b[m\u001b[K \u001b[01;36m\u001b[Knote: \u001b[m\u001b[Kexpected ‘\u001b[01m\u001b[Kvoid **\u001b[m\u001b[K’ but argument is of type ‘\u001b[01m\u001b[Kfloat **\u001b[m\u001b[K’\n extern __host__ cudaError_t CUDARTAPI \u001b[01;36m\u001b[KcudaHostAlloc\u001b[m\u001b[K(void **pHost, size_t size, unsigned int flags);\n \u001b[01;36m\u001b[K^~~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/parser.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Ksave_implicit_weights\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/parser.c:1909:9:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Ki\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Ki\u001b[m\u001b[K;\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/parser.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kget_classes_multipliers\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/parser.c:438:29:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kargument 1 range [18446744071562067968, 18446744073709551615] exceeds maximum object size 9223372036854775807 [\u001b[01;35m\u001b[K-Walloc-size-larger-than=\u001b[m\u001b[K]\n \u001b[01;35m\u001b[Kclasses_multipliers = (float *)calloc(classes_counters, sizeof(float))\u001b[m\u001b[K;\n \u001b[01;35m\u001b[K~~~~~~~~~~~~~~~~~~~~^~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~\u001b[m\u001b[K\nIn file included from \u001b[01m\u001b[K./src/parser.c:3:0\u001b[m\u001b[K:\n\u001b[01m\u001b[K/usr/include/stdlib.h:541:14:\u001b[m\u001b[K \u001b[01;36m\u001b[Knote: \u001b[m\u001b[Kin a call to allocation function ‘\u001b[01m\u001b[Kcalloc\u001b[m\u001b[K’ declared here\n extern void *\u001b[01;36m\u001b[Kcalloc\u001b[m\u001b[K (size_t __nmemb, size_t __size)\n \u001b[01;36m\u001b[K^~~~~~\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/option_list.c -o obj/option_list.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/darknet.c -o obj/darknet.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/detection_layer.c -o obj/detection_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/captcha.c -o obj/captcha.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/route_layer.c -o obj/route_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/writing.c -o obj/writing.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/box.c -o obj/box.o\n\u001b[01m\u001b[K./src/box.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kbox_iou_kind\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/box.c:154:5:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kenumeration value ‘\u001b[01m\u001b[KMSE\u001b[m\u001b[K’ not handled in switch [\u001b[01;35m\u001b[K-Wswitch\u001b[m\u001b[K]\n \u001b[01;35m\u001b[Kswitch\u001b[m\u001b[K(iou_kind) {\n \u001b[01;35m\u001b[K^~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/box.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kdiounms_sort\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/box.c:898:27:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kbeta_prob\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Kbeta_prob\u001b[m\u001b[K = pow(dets[j].prob[k], 2) / sum_prob;\n \u001b[01;35m\u001b[K^~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/box.c:897:27:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kalpha_prob\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Kalpha_prob\u001b[m\u001b[K = pow(dets[i].prob[k], 2) / sum_prob;\n \u001b[01;35m\u001b[K^~~~~~~~~~\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/nightmare.c -o obj/nightmare.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/normalization_layer.c -o obj/normalization_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/avgpool_layer.c -o obj/avgpool_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/coco.c -o obj/coco.o\n\u001b[01m\u001b[K./src/coco.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kvalidate_coco_recall\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/coco.c:248:11:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kbase\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n char *\u001b[01;35m\u001b[Kbase\u001b[m\u001b[K = \"results/comp4_det_test_\";\n \u001b[01;35m\u001b[K^~~~\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/dice.c -o obj/dice.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/yolo.c -o obj/yolo.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/detector.c -o obj/detector.o\n\u001b[01m\u001b[K./src/detector.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Ktrain_detector\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/detector.c:395:72:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Ksuggest parentheses around ‘\u001b[01m\u001b[K&&\u001b[m\u001b[K’ within ‘\u001b[01m\u001b[K||\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wparentheses\u001b[m\u001b[K]\n \u001b[01;35m\u001b[K(iteration >= (iter_save + 1000) || iteration % 1000 == 0) && net.max_batches < 10000\u001b[m\u001b[K)\n \u001b[01;35m\u001b[K~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~^~~~~~~~~~~~~~~~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/detector.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kprint_cocos\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/detector.c:495:29:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kformat not a string literal and no format arguments [\u001b[01;35m\u001b[K-Wformat-security\u001b[m\u001b[K]\n fprintf(fp, \u001b[01;35m\u001b[Kbuff\u001b[m\u001b[K);\n \u001b[01;35m\u001b[K^~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/detector.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Keliminate_bdd\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/detector.c:588:21:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kstatement with no effect [\u001b[01;35m\u001b[K-Wunused-value\u001b[m\u001b[K]\n \u001b[01;35m\u001b[Kfor\u001b[m\u001b[K (k; buf[k + n] != '\\0'; k++)\n \u001b[01;35m\u001b[K^~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/detector.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kvalidate_detector\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/detector.c:709:13:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kmkd2\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Kmkd2\u001b[m\u001b[K = make_directory(buff2, 0777);\n \u001b[01;35m\u001b[K^~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/detector.c:707:13:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kmkd\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Kmkd\u001b[m\u001b[K = make_directory(buff, 0777);\n \u001b[01;35m\u001b[K^~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/detector.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kvalidate_detector_map\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/detector.c:1326:24:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kvariable ‘\u001b[01m\u001b[Kcur_prob\u001b[m\u001b[K’ set but not used [\u001b[01;35m\u001b[K-Wunused-but-set-variable\u001b[m\u001b[K]\n double \u001b[01;35m\u001b[Kcur_prob\u001b[m\u001b[K = 0;\n \u001b[01;35m\u001b[K^~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/detector.c:1347:15:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kclass_recall\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Kclass_recall\u001b[m\u001b[K = (float)tp_for_thresh_per_class[i] / ((float)tp_for_thresh_per_class[i] + (float)(truth_classes_count[i] - tp_for_thresh_per_class[i]));\n \u001b[01;35m\u001b[K^~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/detector.c:1346:15:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kclass_precision\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Kclass_precision\u001b[m\u001b[K = (float)tp_for_thresh_per_class[i] / ((float)tp_for_thresh_per_class[i] + (float)fp_for_thresh_per_class[i]);\n \u001b[01;35m\u001b[K^~~~~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/detector.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kdraw_object\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/detector.c:1890:19:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kinv_loss\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Kinv_loss\u001b[m\u001b[K = 1.0 / max_val_cmp(0.01, avg_loss);\n \u001b[01;35m\u001b[K^~~~~~~~\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/layer.c -o obj/layer.o\n\u001b[01m\u001b[K./src/layer.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kfree_layer_custom\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/layer.c:208:68:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Ksuggest parentheses around ‘\u001b[01m\u001b[K&&\u001b[m\u001b[K’ within ‘\u001b[01m\u001b[K||\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wparentheses\u001b[m\u001b[K]\n if (l.delta_gpu && (l.optimized_memory < 1 || \u001b[01;35m\u001b[Kl.keep_delta_gpu && l.optimized_memory < 3\u001b[m\u001b[K)) cuda_free(l.delta_gpu), l.delta_gpu = NULL;\n \u001b[01;35m\u001b[K~~~~~~~~~~~~~~~~~^~~~~~~~~~~~~~~~~~~~~~~~~\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/compare.c -o obj/compare.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/classifier.c -o obj/classifier.o\n\u001b[01m\u001b[K./src/classifier.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Ktrain_classifier\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/classifier.c:146:9:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kcount\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Kcount\u001b[m\u001b[K = 0;\n \u001b[01;35m\u001b[K^~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/classifier.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kpredict_classifier\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/classifier.c:855:13:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Ktime\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n clock_t \u001b[01;35m\u001b[Ktime\u001b[m\u001b[K;\n \u001b[01;35m\u001b[K^~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/classifier.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kdemo_classifier\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/classifier.c:1287:49:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Ktval_result\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n struct timeval tval_before, tval_after, \u001b[01;35m\u001b[Ktval_result\u001b[m\u001b[K;\n \u001b[01;35m\u001b[K^~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/classifier.c:1287:37:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Ktval_after\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n struct timeval tval_before, \u001b[01;35m\u001b[Ktval_after\u001b[m\u001b[K, tval_result;\n \u001b[01;35m\u001b[K^~~~~~~~~~\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/local_layer.c -o obj/local_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/swag.c -o obj/swag.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/shortcut_layer.c -o obj/shortcut_layer.o\n\u001b[01m\u001b[K./src/shortcut_layer.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kmake_shortcut_layer\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/shortcut_layer.c:55:15:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kscale\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Kscale\u001b[m\u001b[K = sqrt(2. / l.nweights);\n \u001b[01;35m\u001b[K^~~~~\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/representation_layer.c -o obj/representation_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/activation_layer.c -o obj/activation_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/rnn_layer.c -o obj/rnn_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/gru_layer.c -o obj/gru_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/rnn.c -o obj/rnn.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/rnn_vid.c -o obj/rnn_vid.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/crnn_layer.c -o obj/crnn_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/demo.c -o obj/demo.o\n\u001b[01m\u001b[K./src/demo.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kdetect_in_thread\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/demo.c:101:15:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kl\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n layer \u001b[01;35m\u001b[Kl\u001b[m\u001b[K = net.layers[net.n - 1];\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/tag.c -o obj/tag.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/cifar.c -o obj/cifar.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/go.c -o obj/go.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/batchnorm_layer.c -o obj/batchnorm_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/art.c -o obj/art.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/region_layer.c -o obj/region_layer.o\n\u001b[01m\u001b[K./src/region_layer.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kresize_region_layer\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/region_layer.c:63:9:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kold_h\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Kold_h\u001b[m\u001b[K = l->h;\n \u001b[01;35m\u001b[K^~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/region_layer.c:62:9:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kold_w\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Kold_w\u001b[m\u001b[K = l->w;\n \u001b[01;35m\u001b[K^~~~~\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/reorg_layer.c -o obj/reorg_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/reorg_old_layer.c -o obj/reorg_old_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/super.c -o obj/super.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/voxel.c -o obj/voxel.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/tree.c -o obj/tree.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/yolo_layer.c -o obj/yolo_layer.o\n\u001b[01m\u001b[K./src/yolo_layer.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kmake_yolo_layer\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/yolo_layer.c:68:38:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kpassing argument 1 of ‘\u001b[01m\u001b[KcudaHostAlloc\u001b[m\u001b[K’ from incompatible pointer type [\u001b[01;35m\u001b[K-Wincompatible-pointer-types\u001b[m\u001b[K]\n if (cudaSuccess == cudaHostAlloc(\u001b[01;35m\u001b[K&\u001b[m\u001b[Kl.output, batch*l.outputs*sizeof(float), cudaHostRegisterMapped)) l.output_pinned = 1;\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\nIn file included from \u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime.h:96:0\u001b[m\u001b[K,\n from \u001b[01m\u001b[Kinclude/darknet.h:41\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/activations.h:3\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/layer.h:4\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/yolo_layer.h:5\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/yolo_layer.c:1\u001b[m\u001b[K:\n\u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime_api.h:4811:39:\u001b[m\u001b[K \u001b[01;36m\u001b[Knote: \u001b[m\u001b[Kexpected ‘\u001b[01m\u001b[Kvoid **\u001b[m\u001b[K’ but argument is of type ‘\u001b[01m\u001b[Kfloat **\u001b[m\u001b[K’\n extern __host__ cudaError_t CUDARTAPI \u001b[01;36m\u001b[KcudaHostAlloc\u001b[m\u001b[K(void **pHost, size_t size, unsigned int flags);\n \u001b[01;36m\u001b[K^~~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/yolo_layer.c:75:38:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kpassing argument 1 of ‘\u001b[01m\u001b[KcudaHostAlloc\u001b[m\u001b[K’ from incompatible pointer type [\u001b[01;35m\u001b[K-Wincompatible-pointer-types\u001b[m\u001b[K]\n if (cudaSuccess == cudaHostAlloc(\u001b[01;35m\u001b[K&\u001b[m\u001b[Kl.delta, batch*l.outputs*sizeof(float), cudaHostRegisterMapped)) l.delta_pinned = 1;\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\nIn file included from \u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime.h:96:0\u001b[m\u001b[K,\n from \u001b[01m\u001b[Kinclude/darknet.h:41\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/activations.h:3\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/layer.h:4\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/yolo_layer.h:5\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/yolo_layer.c:1\u001b[m\u001b[K:\n\u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime_api.h:4811:39:\u001b[m\u001b[K \u001b[01;36m\u001b[Knote: \u001b[m\u001b[Kexpected ‘\u001b[01m\u001b[Kvoid **\u001b[m\u001b[K’ but argument is of type ‘\u001b[01m\u001b[Kfloat **\u001b[m\u001b[K’\n extern __host__ cudaError_t CUDARTAPI \u001b[01;36m\u001b[KcudaHostAlloc\u001b[m\u001b[K(void **pHost, size_t size, unsigned int flags);\n \u001b[01;36m\u001b[K^~~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/yolo_layer.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kresize_yolo_layer\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/yolo_layer.c:106:42:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kpassing argument 1 of ‘\u001b[01m\u001b[KcudaHostAlloc\u001b[m\u001b[K’ from incompatible pointer type [\u001b[01;35m\u001b[K-Wincompatible-pointer-types\u001b[m\u001b[K]\n if (cudaSuccess != cudaHostAlloc(\u001b[01;35m\u001b[K&\u001b[m\u001b[Kl->output, l->batch*l->outputs * sizeof(float), cudaHostRegisterMapped)) {\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\nIn file included from \u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime.h:96:0\u001b[m\u001b[K,\n from \u001b[01m\u001b[Kinclude/darknet.h:41\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/activations.h:3\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/layer.h:4\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/yolo_layer.h:5\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/yolo_layer.c:1\u001b[m\u001b[K:\n\u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime_api.h:4811:39:\u001b[m\u001b[K \u001b[01;36m\u001b[Knote: \u001b[m\u001b[Kexpected ‘\u001b[01m\u001b[Kvoid **\u001b[m\u001b[K’ but argument is of type ‘\u001b[01m\u001b[Kfloat **\u001b[m\u001b[K’\n extern __host__ cudaError_t CUDARTAPI \u001b[01;36m\u001b[KcudaHostAlloc\u001b[m\u001b[K(void **pHost, size_t size, unsigned int flags);\n \u001b[01;36m\u001b[K^~~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/yolo_layer.c:115:42:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kpassing argument 1 of ‘\u001b[01m\u001b[KcudaHostAlloc\u001b[m\u001b[K’ from incompatible pointer type [\u001b[01;35m\u001b[K-Wincompatible-pointer-types\u001b[m\u001b[K]\n if (cudaSuccess != cudaHostAlloc(\u001b[01;35m\u001b[K&\u001b[m\u001b[Kl->delta, l->batch*l->outputs * sizeof(float), cudaHostRegisterMapped)) {\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\nIn file included from \u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime.h:96:0\u001b[m\u001b[K,\n from \u001b[01m\u001b[Kinclude/darknet.h:41\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/activations.h:3\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/layer.h:4\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/yolo_layer.h:5\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/yolo_layer.c:1\u001b[m\u001b[K:\n\u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime_api.h:4811:39:\u001b[m\u001b[K \u001b[01;36m\u001b[Knote: \u001b[m\u001b[Kexpected ‘\u001b[01m\u001b[Kvoid **\u001b[m\u001b[K’ but argument is of type ‘\u001b[01m\u001b[Kfloat **\u001b[m\u001b[K’\n extern __host__ cudaError_t CUDARTAPI \u001b[01;36m\u001b[KcudaHostAlloc\u001b[m\u001b[K(void **pHost, size_t size, unsigned int flags);\n \u001b[01;36m\u001b[K^~~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/yolo_layer.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kprocess_batch\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/yolo_layer.c:426:25:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kvariable ‘\u001b[01m\u001b[Kbest_match_t\u001b[m\u001b[K’ set but not used [\u001b[01;35m\u001b[K-Wunused-but-set-variable\u001b[m\u001b[K]\n int \u001b[01;35m\u001b[Kbest_match_t\u001b[m\u001b[K = 0;\n \u001b[01;35m\u001b[K^~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/yolo_layer.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kforward_yolo_layer\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/yolo_layer.c:707:11:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kavg_anyobj\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Kavg_anyobj\u001b[m\u001b[K = 0;\n \u001b[01;35m\u001b[K^~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/yolo_layer.c:706:11:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kavg_obj\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Kavg_obj\u001b[m\u001b[K = 0;\n \u001b[01;35m\u001b[K^~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/yolo_layer.c:705:11:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kavg_cat\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Kavg_cat\u001b[m\u001b[K = 0;\n \u001b[01;35m\u001b[K^~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/yolo_layer.c:704:11:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Krecall75\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Krecall75\u001b[m\u001b[K = 0;\n \u001b[01;35m\u001b[K^~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/yolo_layer.c:703:11:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Krecall\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Krecall\u001b[m\u001b[K = 0;\n \u001b[01;35m\u001b[K^~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/yolo_layer.c:702:11:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Ktot_ciou_loss\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Ktot_ciou_loss\u001b[m\u001b[K = 0;\n \u001b[01;35m\u001b[K^~~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/yolo_layer.c:701:11:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Ktot_diou_loss\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Ktot_diou_loss\u001b[m\u001b[K = 0;\n \u001b[01;35m\u001b[K^~~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/yolo_layer.c:698:11:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Ktot_ciou\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Ktot_ciou\u001b[m\u001b[K = 0;\n \u001b[01;35m\u001b[K^~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/yolo_layer.c:697:11:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Ktot_diou\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Ktot_diou\u001b[m\u001b[K = 0;\n \u001b[01;35m\u001b[K^~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/yolo_layer.c:696:11:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Ktot_giou\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n float \u001b[01;35m\u001b[Ktot_giou\u001b[m\u001b[K = 0;\n \u001b[01;35m\u001b[K^~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/yolo_layer.c:668:12:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kunused variable ‘\u001b[01m\u001b[Kn\u001b[m\u001b[K’ [\u001b[01;35m\u001b[K-Wunused-variable\u001b[m\u001b[K]\n int b, \u001b[01;35m\u001b[Kn\u001b[m\u001b[K;\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/gaussian_yolo_layer.c -o obj/gaussian_yolo_layer.o\n\u001b[01m\u001b[K./src/gaussian_yolo_layer.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kmake_gaussian_yolo_layer\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/gaussian_yolo_layer.c:72:38:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kpassing argument 1 of ‘\u001b[01m\u001b[KcudaHostAlloc\u001b[m\u001b[K’ from incompatible pointer type [\u001b[01;35m\u001b[K-Wincompatible-pointer-types\u001b[m\u001b[K]\n if (cudaSuccess == cudaHostAlloc(\u001b[01;35m\u001b[K&\u001b[m\u001b[Kl.output, batch*l.outputs * sizeof(float), cudaHostRegisterMapped)) l.output_pinned = 1;\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\nIn file included from \u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime.h:96:0\u001b[m\u001b[K,\n from \u001b[01m\u001b[Kinclude/darknet.h:41\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/gaussian_yolo_layer.h:5\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/gaussian_yolo_layer.c:7\u001b[m\u001b[K:\n\u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime_api.h:4811:39:\u001b[m\u001b[K \u001b[01;36m\u001b[Knote: \u001b[m\u001b[Kexpected ‘\u001b[01m\u001b[Kvoid **\u001b[m\u001b[K’ but argument is of type ‘\u001b[01m\u001b[Kfloat **\u001b[m\u001b[K’\n extern __host__ cudaError_t CUDARTAPI \u001b[01;36m\u001b[KcudaHostAlloc\u001b[m\u001b[K(void **pHost, size_t size, unsigned int flags);\n \u001b[01;36m\u001b[K^~~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/gaussian_yolo_layer.c:79:38:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kpassing argument 1 of ‘\u001b[01m\u001b[KcudaHostAlloc\u001b[m\u001b[K’ from incompatible pointer type [\u001b[01;35m\u001b[K-Wincompatible-pointer-types\u001b[m\u001b[K]\n if (cudaSuccess == cudaHostAlloc(\u001b[01;35m\u001b[K&\u001b[m\u001b[Kl.delta, batch*l.outputs * sizeof(float), cudaHostRegisterMapped)) l.delta_pinned = 1;\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\nIn file included from \u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime.h:96:0\u001b[m\u001b[K,\n from \u001b[01m\u001b[Kinclude/darknet.h:41\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/gaussian_yolo_layer.h:5\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/gaussian_yolo_layer.c:7\u001b[m\u001b[K:\n\u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime_api.h:4811:39:\u001b[m\u001b[K \u001b[01;36m\u001b[Knote: \u001b[m\u001b[Kexpected ‘\u001b[01m\u001b[Kvoid **\u001b[m\u001b[K’ but argument is of type ‘\u001b[01m\u001b[Kfloat **\u001b[m\u001b[K’\n extern __host__ cudaError_t CUDARTAPI \u001b[01;36m\u001b[KcudaHostAlloc\u001b[m\u001b[K(void **pHost, size_t size, unsigned int flags);\n \u001b[01;36m\u001b[K^~~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/gaussian_yolo_layer.c:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kresize_gaussian_yolo_layer\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/gaussian_yolo_layer.c:111:42:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kpassing argument 1 of ‘\u001b[01m\u001b[KcudaHostAlloc\u001b[m\u001b[K’ from incompatible pointer type [\u001b[01;35m\u001b[K-Wincompatible-pointer-types\u001b[m\u001b[K]\n if (cudaSuccess != cudaHostAlloc(\u001b[01;35m\u001b[K&\u001b[m\u001b[Kl->output, l->batch*l->outputs * sizeof(float), cudaHostRegisterMapped)) {\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\nIn file included from \u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime.h:96:0\u001b[m\u001b[K,\n from \u001b[01m\u001b[Kinclude/darknet.h:41\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/gaussian_yolo_layer.h:5\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/gaussian_yolo_layer.c:7\u001b[m\u001b[K:\n\u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime_api.h:4811:39:\u001b[m\u001b[K \u001b[01;36m\u001b[Knote: \u001b[m\u001b[Kexpected ‘\u001b[01m\u001b[Kvoid **\u001b[m\u001b[K’ but argument is of type ‘\u001b[01m\u001b[Kfloat **\u001b[m\u001b[K’\n extern __host__ cudaError_t CUDARTAPI \u001b[01;36m\u001b[KcudaHostAlloc\u001b[m\u001b[K(void **pHost, size_t size, unsigned int flags);\n \u001b[01;36m\u001b[K^~~~~~~~~~~~~\u001b[m\u001b[K\n\u001b[01m\u001b[K./src/gaussian_yolo_layer.c:120:42:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kpassing argument 1 of ‘\u001b[01m\u001b[KcudaHostAlloc\u001b[m\u001b[K’ from incompatible pointer type [\u001b[01;35m\u001b[K-Wincompatible-pointer-types\u001b[m\u001b[K]\n if (cudaSuccess != cudaHostAlloc(\u001b[01;35m\u001b[K&\u001b[m\u001b[Kl->delta, l->batch*l->outputs * sizeof(float), cudaHostRegisterMapped)) {\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\nIn file included from \u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime.h:96:0\u001b[m\u001b[K,\n from \u001b[01m\u001b[Kinclude/darknet.h:41\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/gaussian_yolo_layer.h:5\u001b[m\u001b[K,\n from \u001b[01m\u001b[K./src/gaussian_yolo_layer.c:7\u001b[m\u001b[K:\n\u001b[01m\u001b[K/usr/local/cuda/include/cuda_runtime_api.h:4811:39:\u001b[m\u001b[K \u001b[01;36m\u001b[Knote: \u001b[m\u001b[Kexpected ‘\u001b[01m\u001b[Kvoid **\u001b[m\u001b[K’ but argument is of type ‘\u001b[01m\u001b[Kfloat **\u001b[m\u001b[K’\n extern __host__ cudaError_t CUDARTAPI \u001b[01;36m\u001b[KcudaHostAlloc\u001b[m\u001b[K(void **pHost, size_t size, unsigned int flags);\n \u001b[01;36m\u001b[K^~~~~~~~~~~~~\u001b[m\u001b[K\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/upsample_layer.c -o obj/upsample_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/lstm_layer.c -o obj/lstm_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/conv_lstm_layer.c -o obj/conv_lstm_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/scale_channels_layer.c -o obj/scale_channels_layer.o\ngcc -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF -c ./src/sam_layer.c -o obj/sam_layer.o\nnvcc -gencode arch=compute_35,code=sm_35 -gencode arch=compute_50,code=[sm_50,compute_50] -gencode arch=compute_52,code=[sm_52,compute_52] -gencode arch=compute_61,code=[sm_61,compute_61] -gencode arch=compute_70,code=[sm_70,compute_70] -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF --compiler-options \"-Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF\" -c ./src/convolutional_kernels.cu -o obj/convolutional_kernels.o\nnvcc warning : The 'compute_35', 'compute_37', 'compute_50', 'sm_35', 'sm_37' and 'sm_50' architectures are deprecated, and may be removed in a future release (Use -Wno-deprecated-gpu-targets to suppress warning).\nnvcc -gencode arch=compute_35,code=sm_35 -gencode arch=compute_50,code=[sm_50,compute_50] -gencode arch=compute_52,code=[sm_52,compute_52] -gencode arch=compute_61,code=[sm_61,compute_61] -gencode arch=compute_70,code=[sm_70,compute_70] -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF --compiler-options \"-Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF\" -c ./src/activation_kernels.cu -o obj/activation_kernels.o\nnvcc warning : The 'compute_35', 'compute_37', 'compute_50', 'sm_35', 'sm_37' and 'sm_50' architectures are deprecated, and may be removed in a future release (Use -Wno-deprecated-gpu-targets to suppress warning).\n./src/activation_kernels.cu(263): warning: variable \"MISH_THRESHOLD\" was declared but never referenced\n\n./src/activation_kernels.cu(263): warning: variable \"MISH_THRESHOLD\" was declared but never referenced\n\n./src/activation_kernels.cu(263): warning: variable \"MISH_THRESHOLD\" was declared but never referenced\n\n./src/activation_kernels.cu(263): warning: variable \"MISH_THRESHOLD\" was declared but never referenced\n\n./src/activation_kernels.cu(263): warning: variable \"MISH_THRESHOLD\" was declared but never referenced\n\nnvcc -gencode arch=compute_35,code=sm_35 -gencode arch=compute_50,code=[sm_50,compute_50] -gencode arch=compute_52,code=[sm_52,compute_52] -gencode arch=compute_61,code=[sm_61,compute_61] -gencode arch=compute_70,code=[sm_70,compute_70] -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF --compiler-options \"-Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF\" -c ./src/im2col_kernels.cu -o obj/im2col_kernels.o\nnvcc warning : The 'compute_35', 'compute_37', 'compute_50', 'sm_35', 'sm_37' and 'sm_50' architectures are deprecated, and may be removed in a future release (Use -Wno-deprecated-gpu-targets to suppress warning).\n./src/im2col_kernels.cu(1354): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1361): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1364): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1389): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1354): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1361): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1364): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1389): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1354): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1361): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1364): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1389): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1354): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1361): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1364): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1389): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1354): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1361): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1364): warning: unrecognized #pragma in device code\n\n./src/im2col_kernels.cu(1389): warning: unrecognized #pragma in device code\n\nnvcc -gencode arch=compute_35,code=sm_35 -gencode arch=compute_50,code=[sm_50,compute_50] -gencode arch=compute_52,code=[sm_52,compute_52] -gencode arch=compute_61,code=[sm_61,compute_61] -gencode arch=compute_70,code=[sm_70,compute_70] -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF --compiler-options \"-Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF\" -c ./src/col2im_kernels.cu -o obj/col2im_kernels.o\nnvcc warning : The 'compute_35', 'compute_37', 'compute_50', 'sm_35', 'sm_37' and 'sm_50' architectures are deprecated, and may be removed in a future release (Use -Wno-deprecated-gpu-targets to suppress warning).\nnvcc -gencode arch=compute_35,code=sm_35 -gencode arch=compute_50,code=[sm_50,compute_50] -gencode arch=compute_52,code=[sm_52,compute_52] -gencode arch=compute_61,code=[sm_61,compute_61] -gencode arch=compute_70,code=[sm_70,compute_70] -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF --compiler-options \"-Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF\" -c ./src/blas_kernels.cu -o obj/blas_kernels.o\nnvcc warning : The 'compute_35', 'compute_37', 'compute_50', 'sm_35', 'sm_37' and 'sm_50' architectures are deprecated, and may be removed in a future release (Use -Wno-deprecated-gpu-targets to suppress warning).\n./src/blas_kernels.cu(1086): warning: variable \"out_index\" was declared but never referenced\n\n./src/blas_kernels.cu(1130): warning: variable \"step\" was set but never used\n\n./src/blas_kernels.cu(1736): warning: variable \"stage_id\" was declared but never referenced\n\n./src/blas_kernels.cu(1086): warning: variable \"out_index\" was declared but never referenced\n\n./src/blas_kernels.cu(1130): warning: variable \"step\" was set but never used\n\n./src/blas_kernels.cu(1736): warning: variable \"stage_id\" was declared but never referenced\n\n./src/blas_kernels.cu(1086): warning: variable \"out_index\" was declared but never referenced\n\n./src/blas_kernels.cu(1130): warning: variable \"step\" was set but never used\n\n./src/blas_kernels.cu(1736): warning: variable \"stage_id\" was declared but never referenced\n\n./src/blas_kernels.cu(1086): warning: variable \"out_index\" was declared but never referenced\n\n./src/blas_kernels.cu(1130): warning: variable \"step\" was set but never used\n\n./src/blas_kernels.cu(1736): warning: variable \"stage_id\" was declared but never referenced\n\n./src/blas_kernels.cu(1086): warning: variable \"out_index\" was declared but never referenced\n\n./src/blas_kernels.cu(1130): warning: variable \"step\" was set but never used\n\n./src/blas_kernels.cu(1736): warning: variable \"stage_id\" was declared but never referenced\n\n\u001b[01m\u001b[K./src/blas_kernels.cu:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kvoid backward_shortcut_multilayer_gpu(int, int, int, int*, float**, float*, float*, float*, float*, int, float*, float**, WEIGHTS_NORMALIZATION_T)\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/blas_kernels.cu:1130:5:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kvariable ‘\u001b[01m\u001b[Kstep\u001b[m\u001b[K’ set but not used [\u001b[01;35m\u001b[K-Wunused-but-set-variable\u001b[m\u001b[K]\n \u001b[01;35m\u001b[Kint \u001b[m\u001b[Kstep = 0;\n \u001b[01;35m\u001b[K^~~~\u001b[m\u001b[K\nnvcc -gencode arch=compute_35,code=sm_35 -gencode arch=compute_50,code=[sm_50,compute_50] -gencode arch=compute_52,code=[sm_52,compute_52] -gencode arch=compute_61,code=[sm_61,compute_61] -gencode arch=compute_70,code=[sm_70,compute_70] -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF --compiler-options \"-Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF\" -c ./src/crop_layer_kernels.cu -o obj/crop_layer_kernels.o\nnvcc warning : The 'compute_35', 'compute_37', 'compute_50', 'sm_35', 'sm_37' and 'sm_50' architectures are deprecated, and may be removed in a future release (Use -Wno-deprecated-gpu-targets to suppress warning).\nnvcc -gencode arch=compute_35,code=sm_35 -gencode arch=compute_50,code=[sm_50,compute_50] -gencode arch=compute_52,code=[sm_52,compute_52] -gencode arch=compute_61,code=[sm_61,compute_61] -gencode arch=compute_70,code=[sm_70,compute_70] -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF --compiler-options \"-Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF\" -c ./src/dropout_layer_kernels.cu -o obj/dropout_layer_kernels.o\nnvcc warning : The 'compute_35', 'compute_37', 'compute_50', 'sm_35', 'sm_37' and 'sm_50' architectures are deprecated, and may be removed in a future release (Use -Wno-deprecated-gpu-targets to suppress warning).\nnvcc -gencode arch=compute_35,code=sm_35 -gencode arch=compute_50,code=[sm_50,compute_50] -gencode arch=compute_52,code=[sm_52,compute_52] -gencode arch=compute_61,code=[sm_61,compute_61] -gencode arch=compute_70,code=[sm_70,compute_70] -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF --compiler-options \"-Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF\" -c ./src/maxpool_layer_kernels.cu -o obj/maxpool_layer_kernels.o\nnvcc warning : The 'compute_35', 'compute_37', 'compute_50', 'sm_35', 'sm_37' and 'sm_50' architectures are deprecated, and may be removed in a future release (Use -Wno-deprecated-gpu-targets to suppress warning).\nnvcc -gencode arch=compute_35,code=sm_35 -gencode arch=compute_50,code=[sm_50,compute_50] -gencode arch=compute_52,code=[sm_52,compute_52] -gencode arch=compute_61,code=[sm_61,compute_61] -gencode arch=compute_70,code=[sm_70,compute_70] -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF --compiler-options \"-Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF\" -c ./src/network_kernels.cu -o obj/network_kernels.o\nnvcc warning : The 'compute_35', 'compute_37', 'compute_50', 'sm_35', 'sm_37' and 'sm_50' architectures are deprecated, and may be removed in a future release (Use -Wno-deprecated-gpu-targets to suppress warning).\n./src/network_kernels.cu(379): warning: variable \"l\" was declared but never referenced\n\n./src/network_kernels.cu(379): warning: variable \"l\" was declared but never referenced\n\n./src/network_kernels.cu(379): warning: variable \"l\" was declared but never referenced\n\n./src/network_kernels.cu(379): warning: variable \"l\" was declared but never referenced\n\n./src/network_kernels.cu(379): warning: variable \"l\" was declared but never referenced\n\n\u001b[01m\u001b[K./src/network_kernels.cu:\u001b[m\u001b[K In function ‘\u001b[01m\u001b[Kfloat train_network_datum_gpu(network, float*, float*)\u001b[m\u001b[K’:\n\u001b[01m\u001b[K./src/network_kernels.cu:379:7:\u001b[m\u001b[K \u001b[01;35m\u001b[Kwarning: \u001b[m\u001b[Kvariable ‘\u001b[01m\u001b[Kl\u001b[m\u001b[K’ set but not used [\u001b[01;35m\u001b[K-Wunused-but-set-variable\u001b[m\u001b[K]\n \u001b[01;35m\u001b[K \u001b[m\u001b[K layer l = net.layers[net.n - 1];\n \u001b[01;35m\u001b[K^\u001b[m\u001b[K\nnvcc -gencode arch=compute_35,code=sm_35 -gencode arch=compute_50,code=[sm_50,compute_50] -gencode arch=compute_52,code=[sm_52,compute_52] -gencode arch=compute_61,code=[sm_61,compute_61] -gencode arch=compute_70,code=[sm_70,compute_70] -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF --compiler-options \"-Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF\" -c ./src/avgpool_layer_kernels.cu -o obj/avgpool_layer_kernels.o\nnvcc warning : The 'compute_35', 'compute_37', 'compute_50', 'sm_35', 'sm_37' and 'sm_50' architectures are deprecated, and may be removed in a future release (Use -Wno-deprecated-gpu-targets to suppress warning).\ng++ -std=c++11 -std=c++11 -Iinclude/ -I3rdparty/stb/include -DOPENCV `pkg-config --cflags opencv4 2> /dev/null || pkg-config --cflags opencv` -DGPU -I/usr/local/cuda/include/ -DCUDNN -DCUDNN_HALF -Wall -Wfatal-errors -Wno-unused-result -Wno-unknown-pragmas -fPIC -Ofast -DOPENCV -DGPU -DCUDNN -I/usr/local/cudnn/include -DCUDNN_HALF obj/image_opencv.o obj/http_stream.o obj/gemm.o obj/utils.o obj/dark_cuda.o obj/convolutional_layer.o obj/list.o obj/image.o obj/activations.o obj/im2col.o obj/col2im.o obj/blas.o obj/crop_layer.o obj/dropout_layer.o obj/maxpool_layer.o obj/softmax_layer.o obj/data.o obj/matrix.o obj/network.o obj/connected_layer.o obj/cost_layer.o obj/parser.o obj/option_list.o obj/darknet.o obj/detection_layer.o obj/captcha.o obj/route_layer.o obj/writing.o obj/box.o obj/nightmare.o obj/normalization_layer.o obj/avgpool_layer.o obj/coco.o obj/dice.o obj/yolo.o obj/detector.o obj/layer.o obj/compare.o obj/classifier.o obj/local_layer.o obj/swag.o obj/shortcut_layer.o obj/representation_layer.o obj/activation_layer.o obj/rnn_layer.o obj/gru_layer.o obj/rnn.o obj/rnn_vid.o obj/crnn_layer.o obj/demo.o obj/tag.o obj/cifar.o obj/go.o obj/batchnorm_layer.o obj/art.o obj/region_layer.o obj/reorg_layer.o obj/reorg_old_layer.o obj/super.o obj/voxel.o obj/tree.o obj/yolo_layer.o obj/gaussian_yolo_layer.o obj/upsample_layer.o obj/lstm_layer.o obj/conv_lstm_layer.o obj/scale_channels_layer.o obj/sam_layer.o obj/convolutional_kernels.o obj/activation_kernels.o obj/im2col_kernels.o obj/col2im_kernels.o obj/blas_kernels.o obj/crop_layer_kernels.o obj/dropout_layer_kernels.o obj/maxpool_layer_kernels.o obj/network_kernels.o obj/avgpool_layer_kernels.o -o darknet -lm -pthread `pkg-config --libs opencv4 2> /dev/null || pkg-config --libs opencv` -L/usr/local/cuda/lib64 -lcuda -lcudart -lcublas -lcurand -L/usr/local/cudnn/lib64 -lcudnn -lstdc++\n"
],
[
"# define helper functions\ndef imShow(path):\n import cv2\n import matplotlib.pyplot as plt\n %matplotlib inline\n\n image = cv2.imread(path)\n height, width = image.shape[:2]\n resized_image = cv2.resize(image,(3*width, 3*height), interpolation = cv2.INTER_CUBIC)\n\n fig = plt.gcf()\n fig.set_size_inches(18, 10)\n plt.axis(\"off\")\n plt.imshow(cv2.cvtColor(resized_image, cv2.COLOR_BGR2RGB))\n plt.show()\n\n# use this to upload files\ndef upload():\n from google.colab import files\n uploaded = files.upload() \n for name, data in uploaded.items():\n with open(name, 'wb') as f:\n f.write(data)\n print ('saved file', name)\n\n# use this to download a file \ndef download(path):\n from google.colab import files\n files.download(path)",
"_____no_output_____"
],
[
"# run darknet detection on test images\n!./darknet detector test cfg/coco.data cfg/yolov4.cfg yolov4.weights data/person.jpg",
"_____no_output_____"
],
[
"# show image using our helper function\nimShow('predictions.jpg')",
"_____no_output_____"
],
[
"",
"_____no_output_____"
]
]
] |
[
"code"
] |
[
[
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code",
"code"
]
] |
4ad004859f05776033071b99a4f8fc5252f350f5
| 104,068 |
ipynb
|
Jupyter Notebook
|
code/chap14.ipynb
|
kmccracken8/ModSimPy
|
ca82f07b9fcebe2237856a1bb0f083d43b34023e
|
[
"MIT"
] | null | null | null |
code/chap14.ipynb
|
kmccracken8/ModSimPy
|
ca82f07b9fcebe2237856a1bb0f083d43b34023e
|
[
"MIT"
] | null | null | null |
code/chap14.ipynb
|
kmccracken8/ModSimPy
|
ca82f07b9fcebe2237856a1bb0f083d43b34023e
|
[
"MIT"
] | null | null | null | 125.081731 | 32,860 | 0.871315 |
[
[
[
"# Modeling and Simulation in Python\n\nChapter 14\n\nCopyright 2017 Allen Downey\n\nLicense: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)",
"_____no_output_____"
]
],
[
[
"# Configure Jupyter so figures appear in the notebook\n%matplotlib inline\n\n# Configure Jupyter to display the assigned value after an assignment\n%config InteractiveShell.ast_node_interactivity='last_expr_or_assign'\n\n# import functions from the modsim.py module\nfrom modsim import *",
"_____no_output_____"
]
],
[
[
"### Code from previous chapters",
"_____no_output_____"
]
],
[
[
"def make_system(beta, gamma):\n \"\"\"Make a system object for the SIR model.\n \n beta: contact rate in days\n gamma: recovery rate in days\n \n returns: System object\n \"\"\"\n init = State(S=89, I=1, R=0)\n init /= np.sum(init)\n\n t0 = 0\n t_end = 7 * 14\n\n return System(init=init, t0=t0, t_end=t_end,\n beta=beta, gamma=gamma)",
"_____no_output_____"
],
[
"def update_func(state, t, system):\n \"\"\"Update the SIR model.\n \n state: State (s, i, r)\n t: time\n system: System object\n \n returns: State (sir)\n \"\"\"\n s, i, r = state\n\n infected = system.beta * i * s \n recovered = system.gamma * i\n \n s -= infected\n i += infected - recovered\n r += recovered\n \n return State(S=s, I=i, R=r)",
"_____no_output_____"
],
[
"def run_simulation(system, update_func):\n \"\"\"Runs a simulation of the system.\n \n system: System object\n update_func: function that updates state\n \n returns: TimeFrame\n \"\"\"\n unpack(system)\n \n frame = TimeFrame(columns=init.index)\n frame.row[t0] = init\n \n for t in linrange(t0, t_end):\n frame.row[t+1] = update_func(frame.row[t], t, system)\n \n return frame",
"_____no_output_____"
],
[
"def calc_total_infected(results):\n \"\"\"Fraction of population infected during the simulation.\n \n results: DataFrame with columns S, I, R\n \n returns: fraction of population\n \"\"\"\n return get_first_value(results.S) - get_last_value(results.S)",
"_____no_output_____"
],
[
"def sweep_beta(beta_array, gamma):\n \"\"\"Sweep a range of values for beta.\n \n beta_array: array of beta values\n gamma: recovery rate\n \n returns: SweepSeries that maps from beta to total infected\n \"\"\"\n sweep = SweepSeries()\n for beta in beta_array:\n system = make_system(beta, gamma)\n results = run_simulation(system, update_func)\n sweep[system.beta] = calc_total_infected(results)\n return sweep",
"_____no_output_____"
]
],
[
[
"## SweepFrame\n\nThe following sweeps two parameters and stores the results in a `SweepFrame`",
"_____no_output_____"
]
],
[
[
"def sweep_parameters(beta_array, gamma_array):\n \"\"\"Sweep a range of values for beta and gamma.\n \n beta_array: array of infection rates\n gamma_array: array of recovery rates\n \n returns: SweepFrame with one row for each beta\n and one column for each gamma\n \"\"\"\n frame = SweepFrame(columns=gamma_array)\n for gamma in gamma_array:\n frame[gamma] = sweep_beta(beta_array, gamma)\n return frame",
"_____no_output_____"
]
],
[
[
"Here's what the results look like.",
"_____no_output_____"
]
],
[
[
"beta_array = linspace(0.1, 0.9, 11)\ngamma_array = linspace(0.1, 0.7, 4)\nframe = sweep_parameters(beta_array, gamma_array)\nframe.head()",
"_____no_output_____"
]
],
[
[
"And here's how we can plot the results.",
"_____no_output_____"
]
],
[
[
"for gamma in gamma_array:\n label = 'gamma = ' + str(gamma)\n plot(frame[gamma], label=label)\n \ndecorate(xlabel='Contacts per day (beta)',\n ylabel='Fraction infected',\n loc='upper left')",
"_____no_output_____"
]
],
[
[
"It's often useful to separate the code that generates results from the code that plots the results, so we can run the simulations once, save the results, and then use them for different analysis, visualization, etc.",
"_____no_output_____"
],
[
"### Contact number",
"_____no_output_____"
],
[
"After running `sweep_parameters`, we have a `SweepFrame` with one row for each value of `beta` and one column for each value of `gamma`.",
"_____no_output_____"
]
],
[
[
"frame.shape",
"_____no_output_____"
]
],
[
[
"The following loop shows how we can loop through the columns and rows of the `SweepFrame`. With 11 rows and 4 columns, there are 44 elements.",
"_____no_output_____"
]
],
[
[
"for gamma in frame.columns:\n series = frame[gamma]\n for beta in series.index:\n frac_infected = series[beta]\n print(beta, gamma, frac_infected)",
"0.1 0.1 0.0846929424381071\n0.18 0.1 0.7086227853695759\n0.26 0.1 0.9007802517781114\n0.33999999999999997 0.1 0.9568878995442757\n0.42000000000000004 0.1 0.9770452570735504\n0.5 0.1 0.9845958628261559\n0.58 0.1 0.9874003453175401\n0.66 0.1 0.9884042490643622\n0.74 0.1 0.9887434214062726\n0.82 0.1 0.9888495150524135\n0.9 0.1 0.9888795705171926\n0.1 0.3 0.0054435591223862545\n0.18 0.3 0.015914069144794984\n0.26 0.3 0.055379762106819386\n0.33999999999999997 0.3 0.2678641677332422\n0.42000000000000004 0.3 0.5245629358439001\n0.5 0.3 0.6860504839161878\n0.58 0.3 0.7883785563390235\n0.66 0.3 0.8550657464101674\n0.74 0.3 0.8994791356903035\n0.82 0.3 0.9294693026191699\n0.9 0.3 0.9498533103273188\n0.1 0.5 0.0027357655411521797\n0.18 0.5 0.006118341358324897\n0.26 0.5 0.011639469321666152\n0.33999999999999997 0.5 0.022114766524234164\n0.42000000000000004 0.5 0.04781622666891572\n0.5 0.5 0.13243803845818214\n0.58 0.5 0.30326419264834004\n0.66 0.5 0.4641102273186152\n0.74 0.5 0.5884769725281787\n0.82 0.5 0.6827496109784223\n0.9 0.5 0.7545952983288148\n0.1 0.7 0.001826769346999102\n0.18 0.7 0.003782561608418833\n0.26 0.7 0.0064266722107564345\n0.33999999999999997 0.7 0.010190551933453973\n0.42000000000000004 0.7 0.015945826561526877\n0.5 0.7 0.025707925046422053\n0.58 0.7 0.04500775311679983\n0.66 0.7 0.09069406882939202\n0.74 0.7 0.18979521165624091\n0.82 0.7 0.3183431867354656\n0.9 0.7 0.4369993744563133\n"
]
],
[
[
"Now we can wrap that loop in a function and plot the results. For each element of the `SweepFrame`, we have `beta`, `gamma`, and `frac_infected`, and we plot `beta/gamma` on the x-axis and `frac_infected` on the y-axis.",
"_____no_output_____"
]
],
[
[
"def plot_sweep_frame(frame):\n \"\"\"Plot the values from a SweepFrame.\n \n For each (beta, gamma), compute the contact number,\n beta/gamma\n \n frame: SweepFrame with one row per beta, one column per gamma\n \"\"\"\n for gamma in frame.columns:\n series = frame[gamma]\n for beta in series.index:\n frac_infected = series[beta]\n plot(beta/gamma, frac_infected, 'ro')",
"_____no_output_____"
]
],
[
[
"Here's what it looks like:",
"_____no_output_____"
]
],
[
[
"plot_sweep_frame(frame)\n\ndecorate(xlabel='Contact number (beta/gamma)',\n ylabel='Fraction infected',\n legend=False)\n\nsavefig('figs/chap06-fig03.pdf')",
"Saving figure to file figs/chap06-fig03.pdf\n"
]
],
[
[
"It turns out that the ratio `beta/gamma`, called the \"contact number\" is sufficient to predict the total number of infections; we don't have to know `beta` and `gamma` separately.\n\nWe can see that in the previous plot: when we plot the fraction infected versus the contact number, the results fall close to a curve.",
"_____no_output_____"
],
[
"### Analysis",
"_____no_output_____"
],
[
"In the book we figured out the relationship between $c$ and $s_{\\infty}$ analytically. Now we can compute it for a range of values:",
"_____no_output_____"
]
],
[
[
"s_inf_array = linspace(0.0001, 0.9999, 101);",
"_____no_output_____"
],
[
"c_array = log(s_inf_array) / (s_inf_array - 1);",
"_____no_output_____"
]
],
[
[
"`total_infected` is the change in $s$ from the beginning to the end.",
"_____no_output_____"
]
],
[
[
"frac_infected = 1 - s_inf_array\nfrac_infected_series = Series(frac_infected, index=c_array);",
"_____no_output_____"
]
],
[
[
"Now we can plot the analytic results and compare them to the simulations.",
"_____no_output_____"
]
],
[
[
"plot_sweep_frame(frame)\nplot(frac_infected_series, label='Analysis')\n\ndecorate(xlabel='Contact number (c)',\n ylabel='Fraction infected')\n\nsavefig('figs/chap06-fig04.pdf')",
"Saving figure to file figs/chap06-fig04.pdf\n"
]
],
[
[
"The agreement is generally good, except for values of `c` less than 1.",
"_____no_output_____"
],
[
"## Exercises",
"_____no_output_____"
],
[
"**Exercise:** If we didn't know about contact numbers, we might have explored other possibilities, like the difference between `beta` and `gamma`, rather than their ratio.\n\nWrite a version of `plot_sweep_frame`, called `plot_sweep_frame_difference`, that plots the fraction infected versus the difference `beta-gamma`.\n\nWhat do the results look like, and what does that imply? ",
"_____no_output_____"
]
],
[
[
"def plot_sweep_frame_difference(frame):\n \"\"\"Plot the values from a SweepFrame.\n \n For each (beta, gamma), compute the contact number,\n beta/gamma\n \n frame: SweepFrame with one row per beta, one column per gamma\n \"\"\"\n for gamma in frame.columns:\n series = frame[gamma]\n for beta in series.index:\n frac_infected = series[beta]\n plot(beta - gamma, frac_infected, 'ro')",
"_____no_output_____"
],
[
"plot_sweep_frame_difference(frame)\n\ndecorate(xlabel='Contact difference (beta - gamma)',\n ylabel='Fraction infected',\n legend=False)\n\nsavefig('figs/chap06-fig03.pdf')",
"Saving figure to file figs/chap06-fig03.pdf\n"
],
[
"# Solution goes here",
"_____no_output_____"
]
],
[
[
"**Exercise:** Suppose you run a survey at the end of the semester and find that 26% of students had the Freshman Plague at some point.\n\nWhat is your best estimate of `c`?\n\nHint: if you print `frac_infected_series`, you can read off the answer. ",
"_____no_output_____"
]
],
[
[
"print(frac_infected_series)",
"9.211261 0.999900\n4.642296 0.989902\n3.987365 0.979904\n3.612133 0.969906\n3.350924 0.959908\n3.151808 0.949910\n2.991711 0.939912\n2.858363 0.929914\n2.744467 0.919916\n2.645332 0.909918\n2.557767 0.899920\n2.479505 0.889922\n2.408879 0.879924\n2.344627 0.869926\n2.285771 0.859928\n2.231541 0.849930\n2.181315 0.839932\n2.134590 0.829934\n2.090947 0.819936\n2.050040 0.809938\n2.011573 0.799940\n1.975299 0.789942\n1.941002 0.779944\n1.908499 0.769946\n1.877628 0.759948\n1.848249 0.749950\n1.820238 0.739952\n1.793487 0.729954\n1.767898 0.719956\n1.743384 0.709958\n ... \n1.181034 0.290042\n1.173263 0.280044\n1.165630 0.270046\n1.158132 0.260048\n1.150765 0.250050\n1.143524 0.240052\n1.136407 0.230054\n1.129409 0.220056\n1.122527 0.210058\n1.115758 0.200060\n1.109099 0.190062\n1.102547 0.180064\n1.096099 0.170066\n1.089751 0.160068\n1.083503 0.150070\n1.077350 0.140072\n1.071291 0.130074\n1.065323 0.120076\n1.059444 0.110078\n1.053651 0.100080\n1.047943 0.090082\n1.042317 0.080084\n1.036772 0.070086\n1.031305 0.060088\n1.025914 0.050090\n1.020598 0.040092\n1.015356 0.030094\n1.010185 0.020096\n1.005083 0.010098\n1.000050 0.000100\nLength: 101, dtype: float64\n"
],
[
"# Alternative solution\n\n\"\"\"We can use `np.interp` to look up `s_inf` and\nestimate the corresponding value of `c`, but it only\nworks if the index of the series is sorted in ascending\norder. So we have to use `sort_index` first.\n\"\"\"\n\nfrac_infected_series.sort_index(inplace=True)\nnp.interp(0.26, frac_infected_series, frac_infected_series.index)",
"_____no_output_____"
]
]
] |
[
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code",
"markdown",
"code"
] |
[
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code",
"code",
"code",
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown"
],
[
"code"
],
[
"markdown",
"markdown",
"markdown"
],
[
"code",
"code",
"code"
],
[
"markdown"
],
[
"code",
"code"
]
] |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.